Cbx2, a PcG Family Gene, Plays a Regulatory Role in Medaka Gonadal Development
Abstract
1. Introduction
2. Results
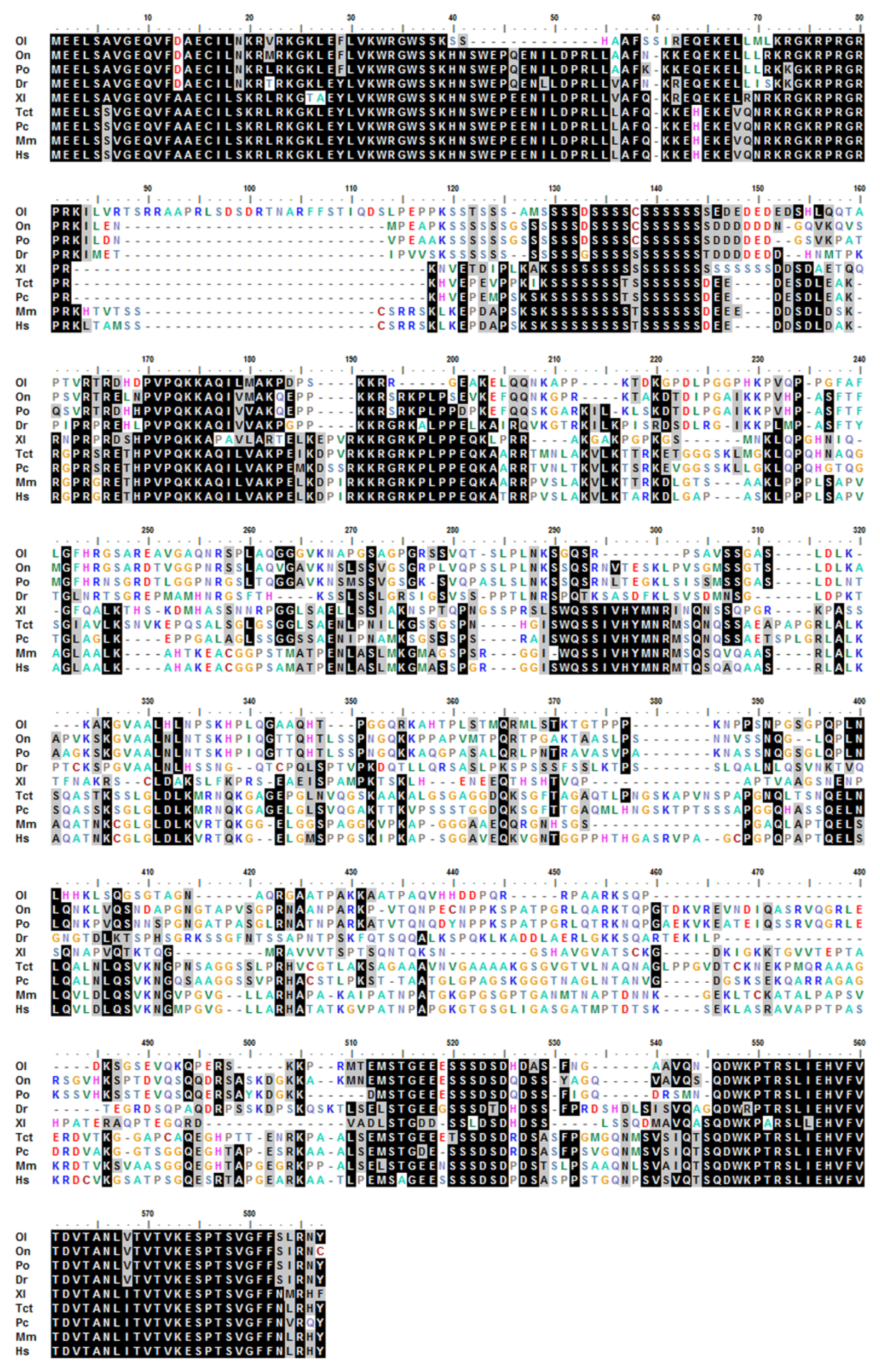
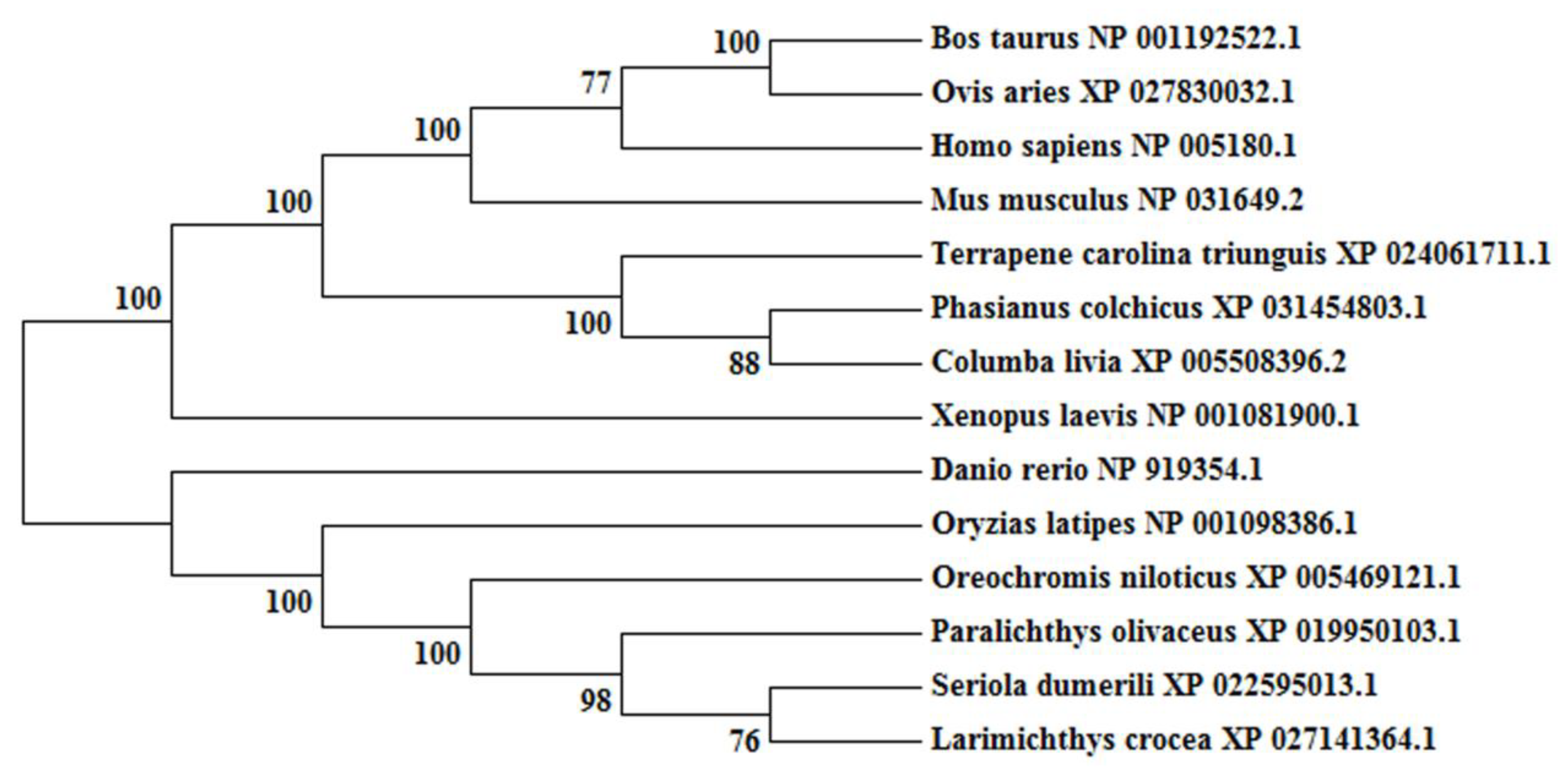
2.1. Molecular Characterization and Phylogenetic Analyses of cbx2
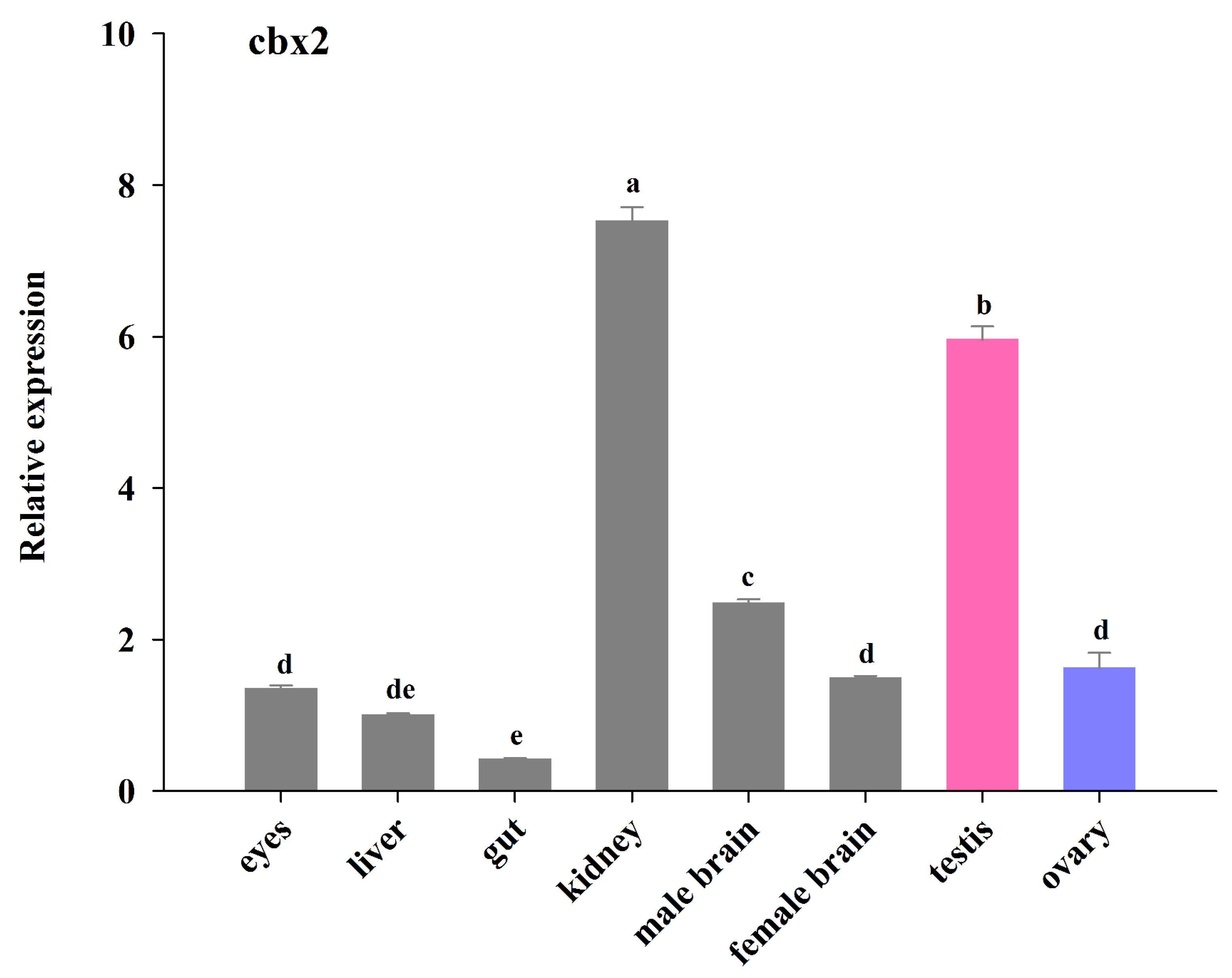
2.2. Tissue Expression of cbx2 mRNA
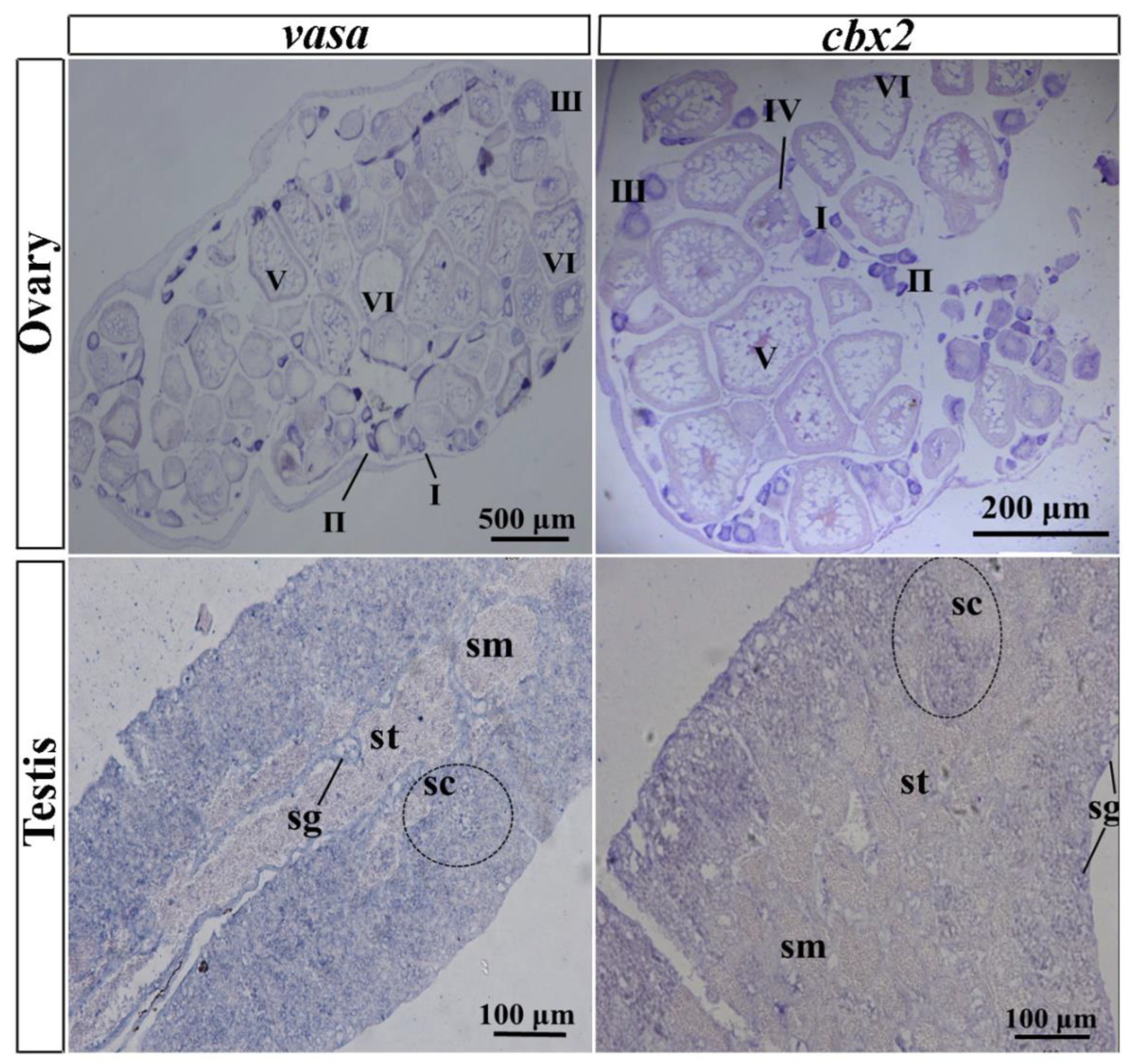
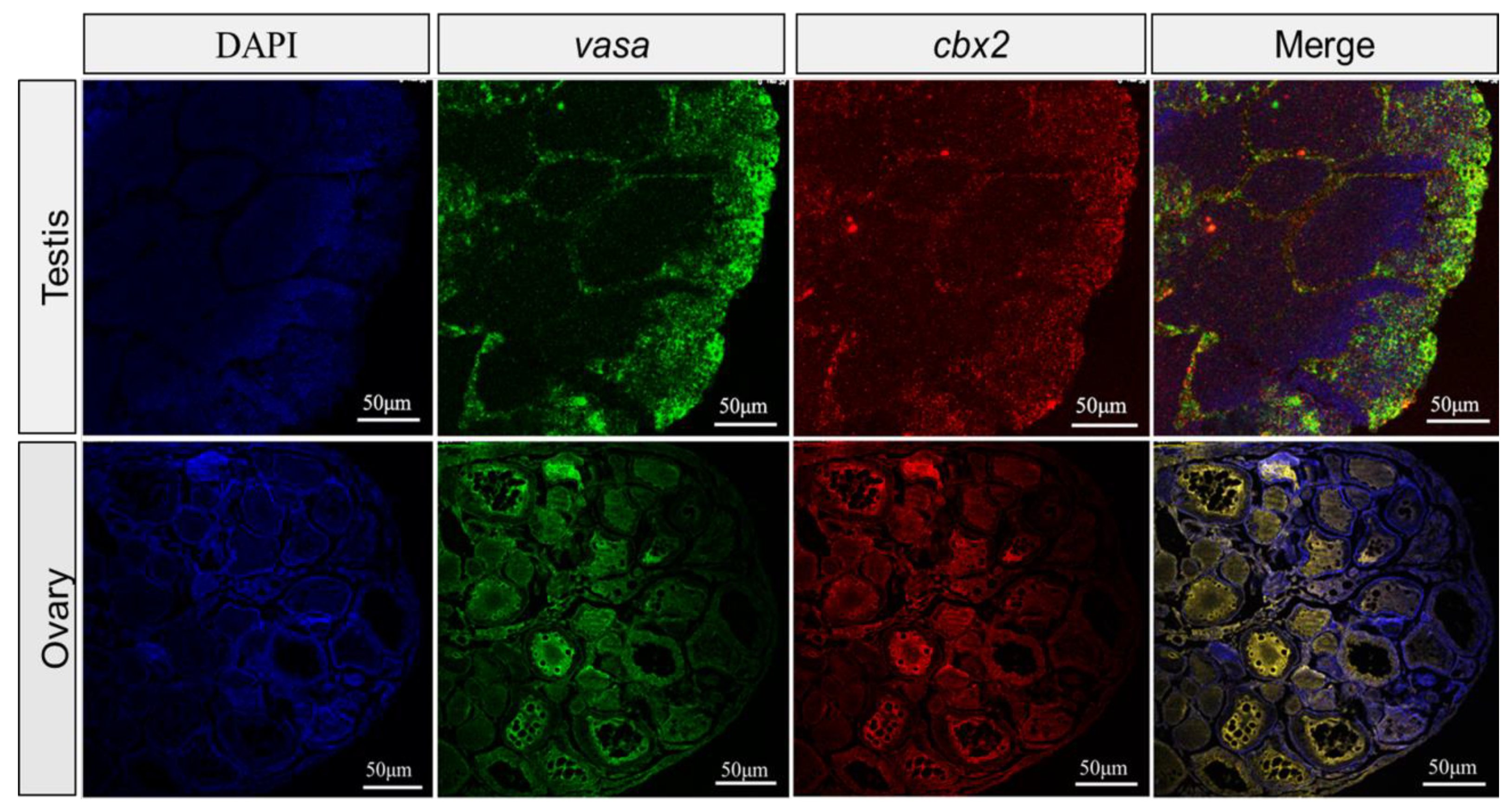
2.3. Gonadal Localization of cbx2 mRNA by ISH and FISH
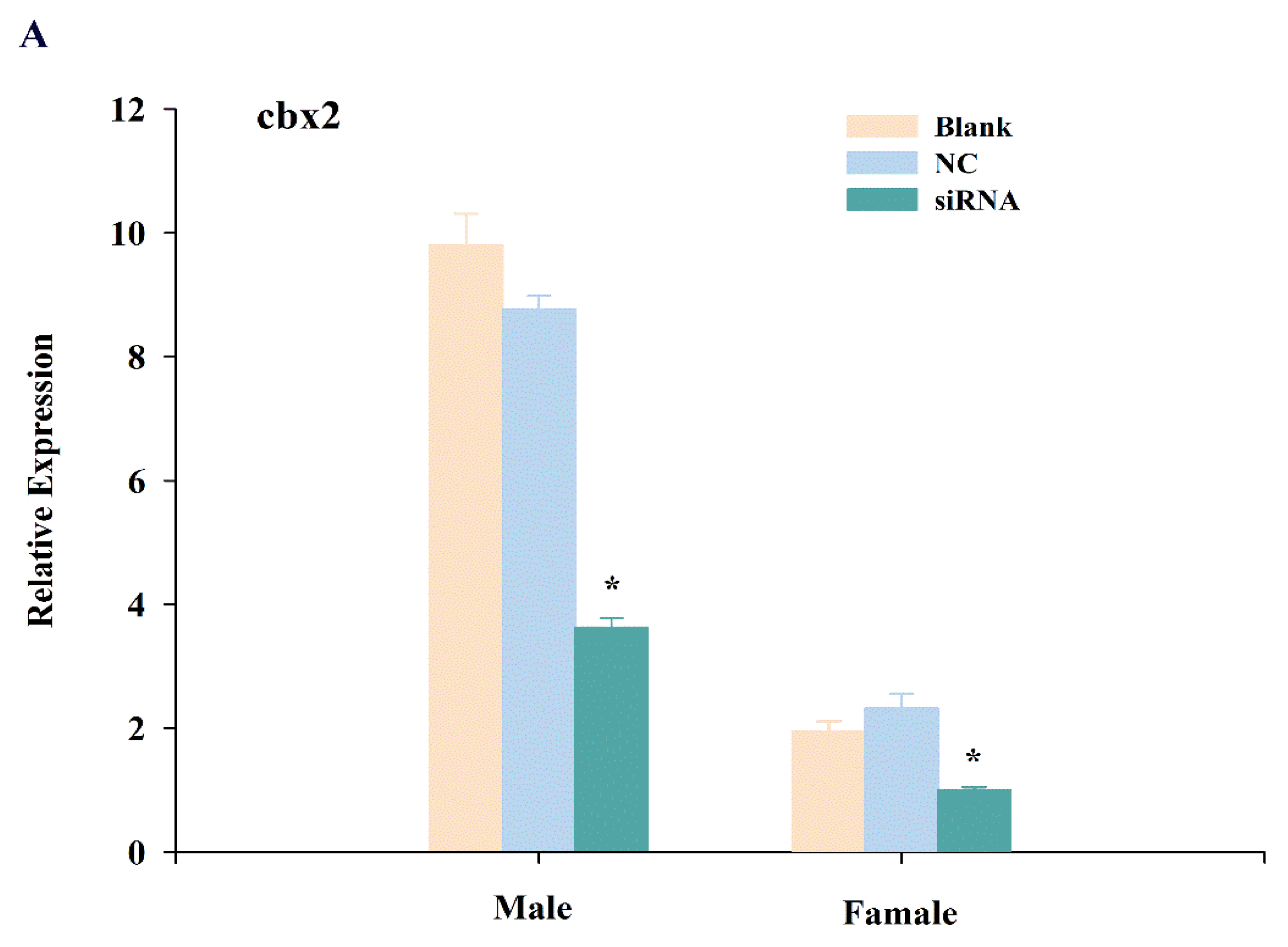
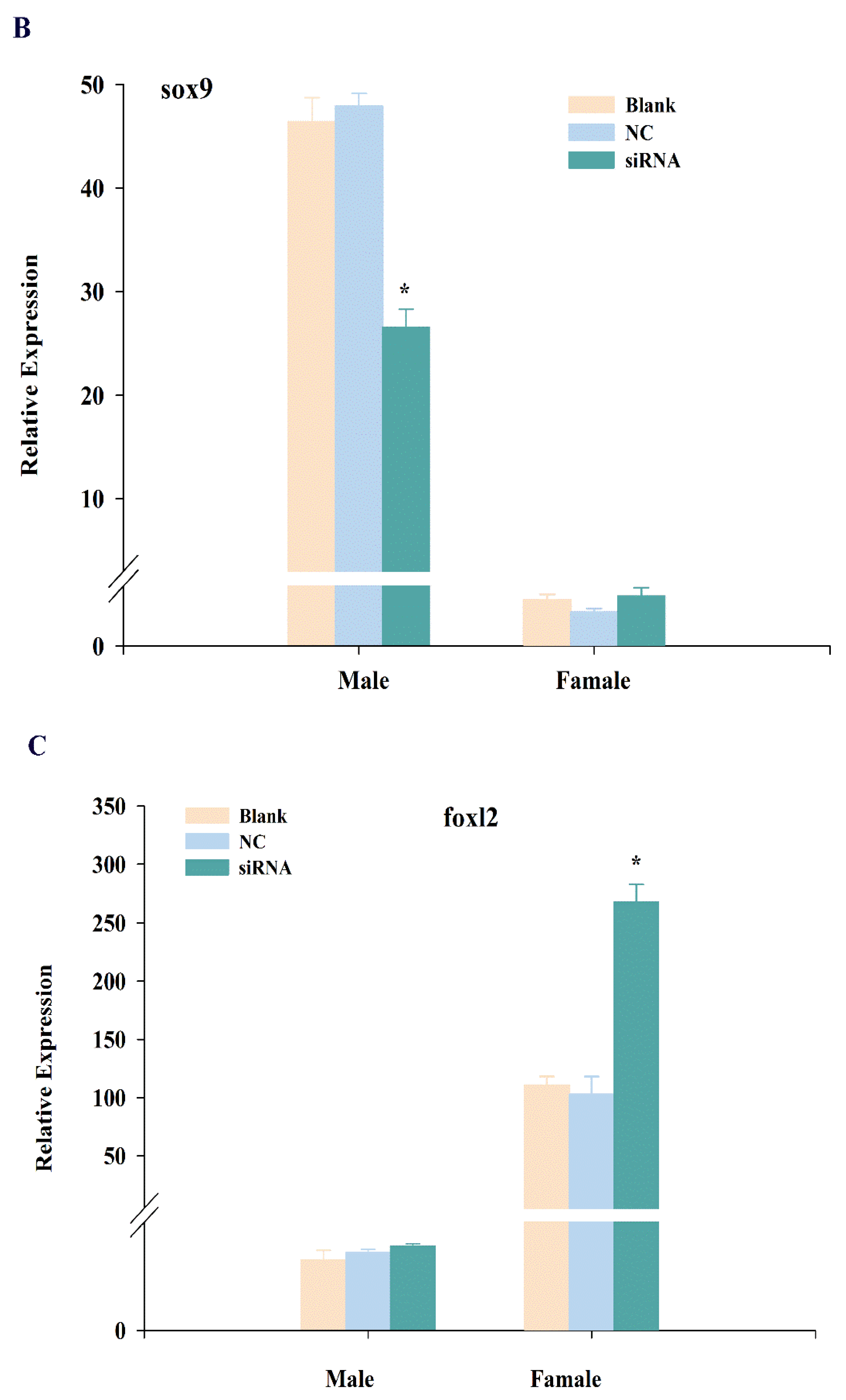
2.4. Expression Changes of cbx2, sox9, and foxl2 mRNA after RNA Interference
3. Discussion
4. Materials and Methods
4.1. Experimental Materials
4.2. Molecular Cloning and Sequence Analysis
4.3. Real-Time PCR of cbx2 in Different Tissues
4.4. Gonadal Expression of cbx2 by ISH and FISH
4.5. RNA Interference
Author Contributions
Funding
Conflicts of Interest
References
- Lewis, E.B. A gene complex controlling segmentation in Drosophila. Nature 1978, 276, 565–570. [Google Scholar] [CrossRef]
- Simon, J.A.; Kingston, R.E. Mechanisms of polycomb gene silencing: Knowns and unknowns. Nat. Rev. Mol. Cell Biol. 2009, 10, 697–708. [Google Scholar] [CrossRef]
- King, H.W.; Fursova, N.A.; Blackledge, N.P.; Klose, R.J. Polycomb repressive complex 1 shapes the nucleosome landscape but not accessibility at target genes. Genome Res. 2018, 28, 1494–1507. [Google Scholar] [CrossRef]
- Boyer, L.A.; Plath, K.; Zeitlinger, J.; Brambrink, T.; Medeiros, L.A.; Lee, T.I.; Levine, S.S.; Wernig, M.; Tajonar, A.; Ray, M.K.; et al. Polycomb complexes repress developmental regulators in murine embryonic stem cells. Nature 2006, 441, 349–353. [Google Scholar] [CrossRef] [PubMed]
- Kerppola, T.K. Polycomb group complexes-many combinations, many functions. Trends Cell Biol. 2009, 19, 692–704. [Google Scholar] [CrossRef] [PubMed]
- Di Croce, L.; Helin, K. Transcriptional regulation by Polycomb group proteins. Nat. Struct. Mol. Biol. 2013, 20, 1147–1155. [Google Scholar] [CrossRef]
- Katoh-Fukui, Y.; Miyabayashi, K.; Komatsu, T.; Owaki, A.; Baba, T.; Shima, Y.; Kidokoro, T.; Kanai, Y.; Schedl, A.; Wilhelm, D.; et al. Cbx2, a polycomb group gene, is required for Sry gene expression in mice. Endocrinology 2012, 153, 913–924. [Google Scholar] [CrossRef] [PubMed]
- Plys, A.J.; Davis, C.P.; Kim, J.; Rizki, G.; Keenen, M.M.; Marr, S.K.; Kingston, R.E. Phase separation of polycomb-repressive complex 1 is governed by a charged disordered region of CBX2. Genes Dev. 2019, 33, 799–813. [Google Scholar] [CrossRef]
- Ma, R.G.; Zhang, Y.; Sun, T.T.; Cheng, B. Epigenetic regulation by polycomb group complexes: Focus on roles of CBX proteins. J. Zhejiang Univ. Sci. B 2014, 15, 412–428. [Google Scholar] [CrossRef]
- Biason-Lauber, A.; Konrad, D.; Meyer, M.; DeBeaufort, C.; Schoenle, E.J. Ovaries and female phenotype in a girl with 46, XY karyotype and mutations in the CBX2 gene. Am. J. Hum. Genet. 2009, 84, 658–663. [Google Scholar] [CrossRef]
- Zhen, C.Y.; Duc, H.N.; Kokotovic, M.; Phiel, C.J.; Ren, X. Cbx2 stably associates with mitotic chromosomes via a PRC2-or PRC1-independent mechanism and is needed for recruiting PRC1 complex to mitotic chromosomes. Mol. Biol. Cell 2014, 25, 3726–3739. [Google Scholar] [CrossRef] [PubMed]
- Katoh-Fukui, Y.; Owaki, A.; Toyama, Y.; Kusaka, M.; Shinohara, Y.; Maekawa, M.; Toshimori, K.; Morohashi, K.I. Mouse polycomb M33 is required for splenic vascular and adrenal gland formation through regulating Ad4BP/SF1 expression. Blood 2005, 106, 1612–1620. [Google Scholar] [CrossRef] [PubMed]
- Sproll, P.; Eid, W.; Biason-Lauber, A. CBX2-dependent transcriptional landscape: Implications for human sex development and its defects. Sci. Rep. 2019, 9, 16552. [Google Scholar] [CrossRef] [PubMed]
- Kurokawa, H.; Saito, D.; Nakamura, S.; Katoh-Fukui, Y.; Ohta, K.; Baba, T.; Morohashi, K.; Tanaka, M. Germ cells are essential for sexual dimorphism in the medaka gonad. Proc. Natl. Acad. Sci. USA 2007, 104, 16958–16963. [Google Scholar] [CrossRef] [PubMed]
- Nanda, I.; Kondo, M.; Hornung, U.; Asakawa, S.; Winkler, C.; Shimizu, A.; Shan, Z.; Haaf, T.; Shimizu, N.; Shima, A.; et al. A duplicated copy of Dmrt1 in the sex-determining region of the Y chromosome of the medaka, Oryzias latipes. Proc. Natl. Acad. Sci. USA 2002, 99, 11778–11783. [Google Scholar] [CrossRef] [PubMed]
- Matsuda, M.; Nagahama, Y.; Shinomiya, A.; Sato, T.; Matsuda, C.; Kobayashi, T.; Morrey, C.E.; Shibata, N.; Asakawa, S.; Shimizu, N.; et al. DMY is a Y-specific DM-domain gene required for male development in the medaka fish. Nature 2002, 417, 559–563. [Google Scholar] [CrossRef]
- Kawaguchi, T.; Machida, S.; Kurumizaka, H.; Tagami, H.; Nakayama, J.I. Phosphorylation of CBX2 controls its nucleosome-binding specificity. J. Biochem. 2017, 162, 343–355. [Google Scholar] [CrossRef]
- Senthilkumar, R.; Mishra, R.K. Novel motifs distinguish multiple homologues of polycombin vertebrates: Expansion and diversification of the epigenetic toolkit. BMC Genom. 2009, 10, 549–566. [Google Scholar] [CrossRef]
- Wang, R.; Taylor, A.B.; Leal, B.Z.; Chadwell, L.V.; Ilangovan, U.; Robinson, A.K.; Schirf, V.; Hart, P.J.; Lafer, E.M.; Demeler, B.; et al. Polycomb group targeting through different binding partners of RING1B C-terminal domain. Structure 2010, 18, 966–975. [Google Scholar] [CrossRef]
- Fagerberg, L.; Hallström, B.; Oksvold, P.; Kampf, C.; Djureinovic, D.; Odeberg, J.; Habuka, M.; Tahmasebpoor, S.; Danielsson, A.; Edlund, K.; et al. Analysis of the human tissue-specific expression by genome-wide integration of transcriptomics and antibody-based proteomics. Mol. Cell. Proteom. 2014, 13, 397–406. [Google Scholar] [CrossRef]
- Klauke, K.; Radulović, V.; Broekhuis, M.; Weersing, E.; Zwart, E.; Olthof, S.; Bystrykh, L. Polycomb Cbx family members mediate the balance between haematopoietic stem cell self-renewal and differentiation. Nat. Cell Biol. 2013, 15, 353–374. [Google Scholar] [CrossRef] [PubMed]
- Kato, Y.; Koseki, H.; Vidal, M.; Nakauchi, H.; Iwama, A. Unique composition of polycomb repressive complex 1 in hematopoietic stem cells. Int. J. Hematol. 2007, 85, 179–181. [Google Scholar] [CrossRef]
- Sun, D.H.; Cao, X.T.; Wang, C.M. Polycomb chromobox Cbx2 enhances antiviral innate immunity by promoting Jmjd3-mediated demethylation of H3K27 at the Ifnb promoter. Protein Cell 2019, 10, 285–294. [Google Scholar] [CrossRef] [PubMed]
- Li, M.Y.; Shen, Q.; Xu, H.Y.; Wong, F.M.; Cui, J.Z.; Li, Z.D.; Hong, N.; Wang, L.; Zhao, H.B.; Ma, B.; et al. Differential conservation and divergence of fertility genes Boule and Dazl in the rainbow trout. PLoS ONE 2011, 6, e15910. [Google Scholar] [CrossRef]
- Noguchi, K.; Shiurba, R.; Higashinakagawa, T. Nuclear translocation of mouse polycomb M33 protein in regenerating liver. Biochem. Biophys. Res. Commun. 2002, 291, 508–515. [Google Scholar] [CrossRef]
- Hirose, S.; Komoike, Y.; Higashinakagawa, T. Identification of a nuclear localization signal in mouse polycomb protein, M33. Zool. Sci. 2006, 23, 785–791. [Google Scholar] [CrossRef]
- Baumann, C.; Rabindranath, D.L.F. Role of polycomb group protein Cbx2/M33 in meiosis onset and maintenance of chromosome stability in the mammalian germline. Genes 2011, 2, 59–80. [Google Scholar] [CrossRef]
- Johnsen, H.; Tveiten, H.; Torgersen, J.S.; Andersen, O. Divergent and sex-dimorphic expression of the paralogs of the Sox9-Amh-Cyp19a1 regulatory cascade in developing and adult-atlantic cod (Gadus morhua L.). Mol. Reprod. Dev. 2013, 80, 358–370. [Google Scholar] [CrossRef]
- Wang, X.Y.; Simpson, E.R.; Brown, K.A. Aromatase overexpression in dysfunctional adipose tissue links obesity to postmenopausal breast cancer. J. Steroid Biochem. Mol. Biol. 2015, 153, 35–44. [Google Scholar] [CrossRef]
- Chiang, E.F.; Pai, C.I.; Wyatt, M.; Yan, Y.L.; Postlethwait, J.; Chung, B. Two Sox9 genes on duplicated zebrafish chromosomes:expression of similar transcription activators in distinct sites. Dev. Biol. 2001, 231, 149–163. [Google Scholar] [CrossRef]
- Wang, H.P.; Wu, T.T.; Qin, F.; Wang, L.H.; Wang, Z.Z. Molecular cloning of Foxl2 gene and the effects of endocrine-disrupting chemicals on its mRNA level in rare minnow, Gobiocypris rarus. Fish Physiol. Biochem. 2012, 38, 653–664. [Google Scholar] [CrossRef] [PubMed]
- Sridevi, P.; Senthilkumaran, B. Cloning and differential expression of Foxl2 during ovarian development and recrudescence of the catfish, Clarias gariepinus. Gen. Comp. Endocrinol. 2011, 174, 259–268. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.S.; Kobayashi, T.; Zhou, L.Y.; Paul-Prasanth, B.; Ijiri, S.; Sakai, F.; Okubo, K.; Morohashi, K.; Nagahama, Y. Foxl2 up-regulates aromatase gene transcription in a female-specific manner by binding to the promoter as well as interacting with Ad4 binding protein/steroidogenic factor 1. Mol. Endocrinol. 2007, 21, 712–725. [Google Scholar] [CrossRef] [PubMed]
- Katoh-Fukui, Y.; Tsuchiya, R.; Shiroishi, T.; Nakahara, Y.; Hashimoto, N.; Noguchi, K.; Higashinakagawa, T. Male-to-female sex reversal in M33 mutant mice. Nature 1998, 393, 688–692. [Google Scholar] [CrossRef]
- Eid, W.; Opitz, L.; Biason-Lauber, A. Genome-wide identification of CBX2 targets: Insights in the human sex development network. Mol. Endocrinol. 2015, 29, 247–257. [Google Scholar] [CrossRef]


| Primer | Primer Sequence (5′→3′) |
|---|---|
| cbx2-F | GCCCAAGTCCAGCACCTCA |
| cbx2-R | GCTCCTTCGCCTCACCTCT |
| cbx2-spel-F | GGACTAGTTGGTGGGCGACTCCTTGA |
| cbx2-sphl-R | CATGCATGCACCTGAACCCTTCCAAACAC |
| vasa-spel-F | GGACTAGTCCCGCTTGTTGAATTTGG |
| vasa-sphl-R | CATGCATGCTGTGCGAGTCGTTGGAGAA |
| sox9-F | AAACTGGCCGACCAATAC |
| sox9-R | CTCAGCCTCCTCCACAAA |
| foxl2-F | TCCTACACGTCCTGCCAGAT |
| foxl2-R | CCCATGCCGTTGTAAGAGTT |
| 18s-F | CTGAGAAACGGCTACCACAG |
| 18s-R | CAGCAACTTTAAGATACGC |
| Gene | Direction | Sequence |
|---|---|---|
| cbx2-siRNA-1(251) | Sense (5′-3′) | CCGACUCUGACCGCACUAATT |
| Anti-sense (3′-5′) | UUAGUGCGGUCAGAGUCGGTT | |
| cbx2-siRNA-2(853) | Sense (5′-3′) | GCGCUGCACCUGAACCCUUTT |
| Anti-sense (3′-5′) | AAGGGUUCAGGUGCAGCGCTT | |
| cbx2-siRNA-3(1337) | Sense (5′-3′) | GCCUCAUCGAGCACGUGUUTT |
| Anti-sense (3′-5′) | AACACGUGCUCGAUGAGGCTT |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chao, Q.; Shen, F.; Xue, Y.; Wu, J.; Zhang, J. Cbx2, a PcG Family Gene, Plays a Regulatory Role in Medaka Gonadal Development. Int. J. Mol. Sci. 2020, 21, 1288. https://doi.org/10.3390/ijms21041288
Chao Q, Shen F, Xue Y, Wu J, Zhang J. Cbx2, a PcG Family Gene, Plays a Regulatory Role in Medaka Gonadal Development. International Journal of Molecular Sciences. 2020; 21(4):1288. https://doi.org/10.3390/ijms21041288
Chicago/Turabian StyleChao, Qinghe, Fengfeng Shen, Yidong Xue, Jikui Wu, and Junling Zhang. 2020. "Cbx2, a PcG Family Gene, Plays a Regulatory Role in Medaka Gonadal Development" International Journal of Molecular Sciences 21, no. 4: 1288. https://doi.org/10.3390/ijms21041288
APA StyleChao, Q., Shen, F., Xue, Y., Wu, J., & Zhang, J. (2020). Cbx2, a PcG Family Gene, Plays a Regulatory Role in Medaka Gonadal Development. International Journal of Molecular Sciences, 21(4), 1288. https://doi.org/10.3390/ijms21041288




