Construction and Analysis of GmFAD2-1A and GmFAD2-2A Soybean Fatty Acid Desaturase Mutants Based on CRISPR/Cas9 Technology
Abstract
1. Introduction
2. Results
2.1. Target Design
2.2. Verification of Expression Vector
2.3. Molecular Detection of Transgenic Offspring
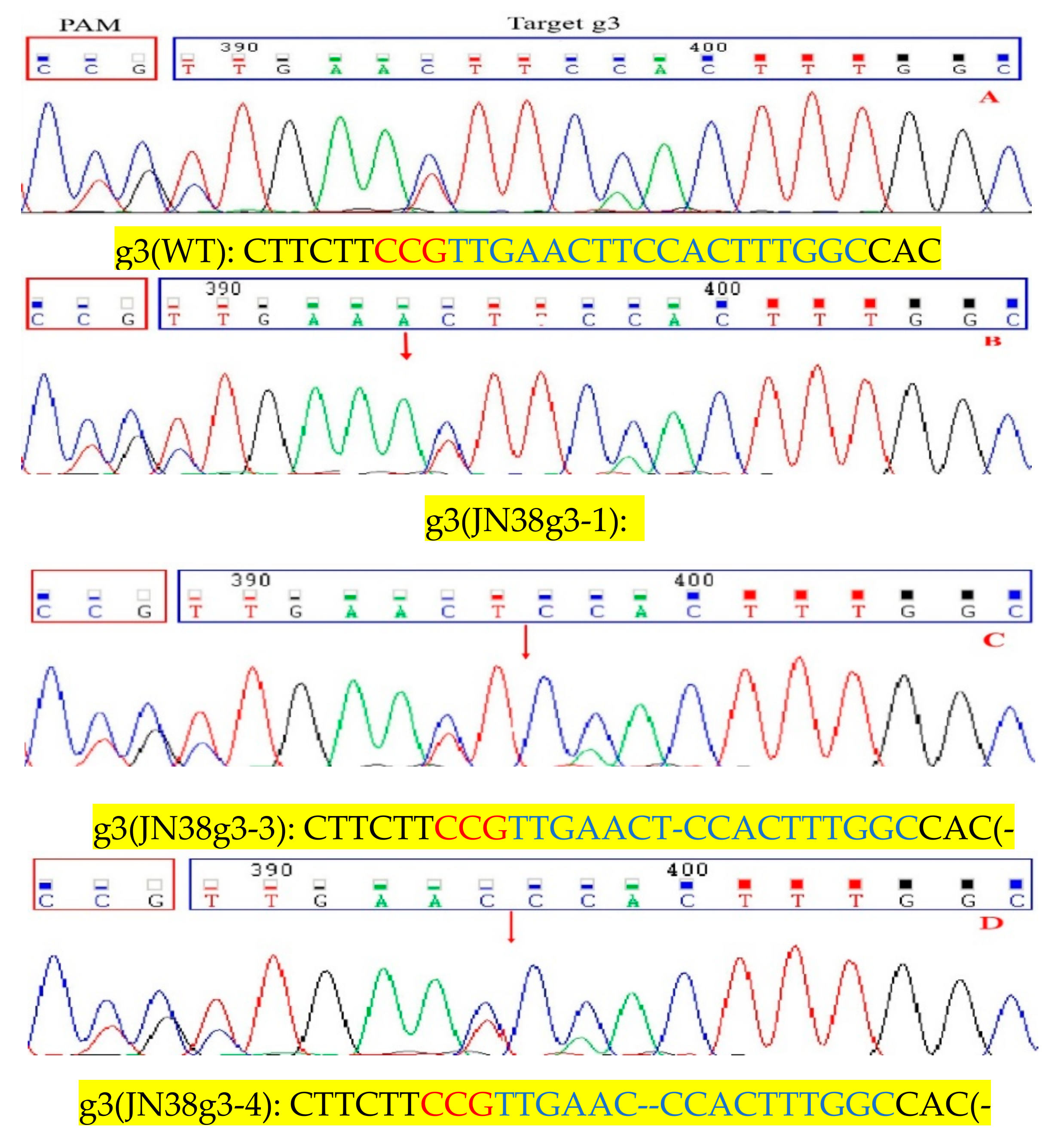
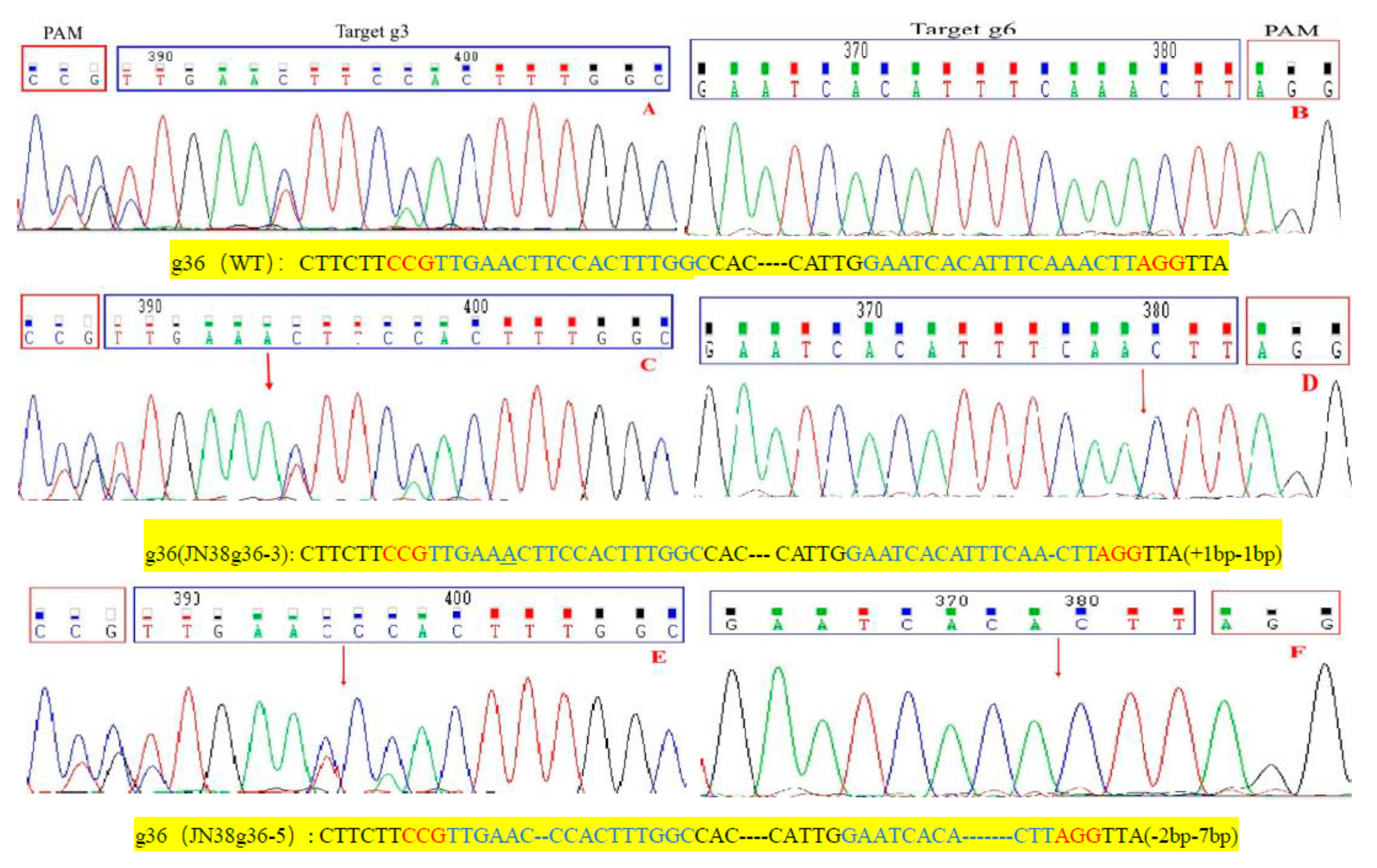
2.4. Sequence Analysis of the Target Mutation of Positive Plants
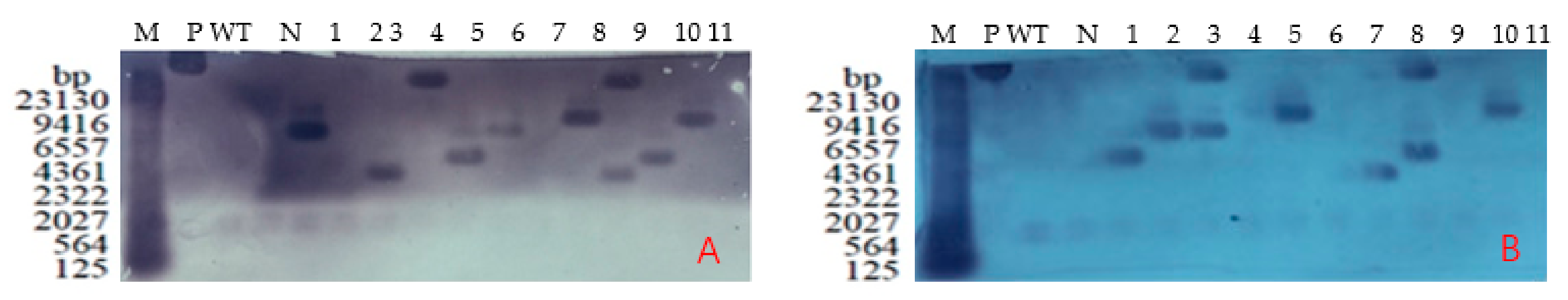
2.5. Southern Blotting Detection of Positive T3 Plants
2.6. Detection of T3 Generation Transgenic Plants by Real-Time PCR
2.7. Determination of Oleic Acid Content in Transgenic T2, T3 Positive Seeds
2.8. Significance Analysis of Difference in Oleic Acid Content in Offspring Grains
2.9. Analysis of Agronomic Characters of Converted Materials
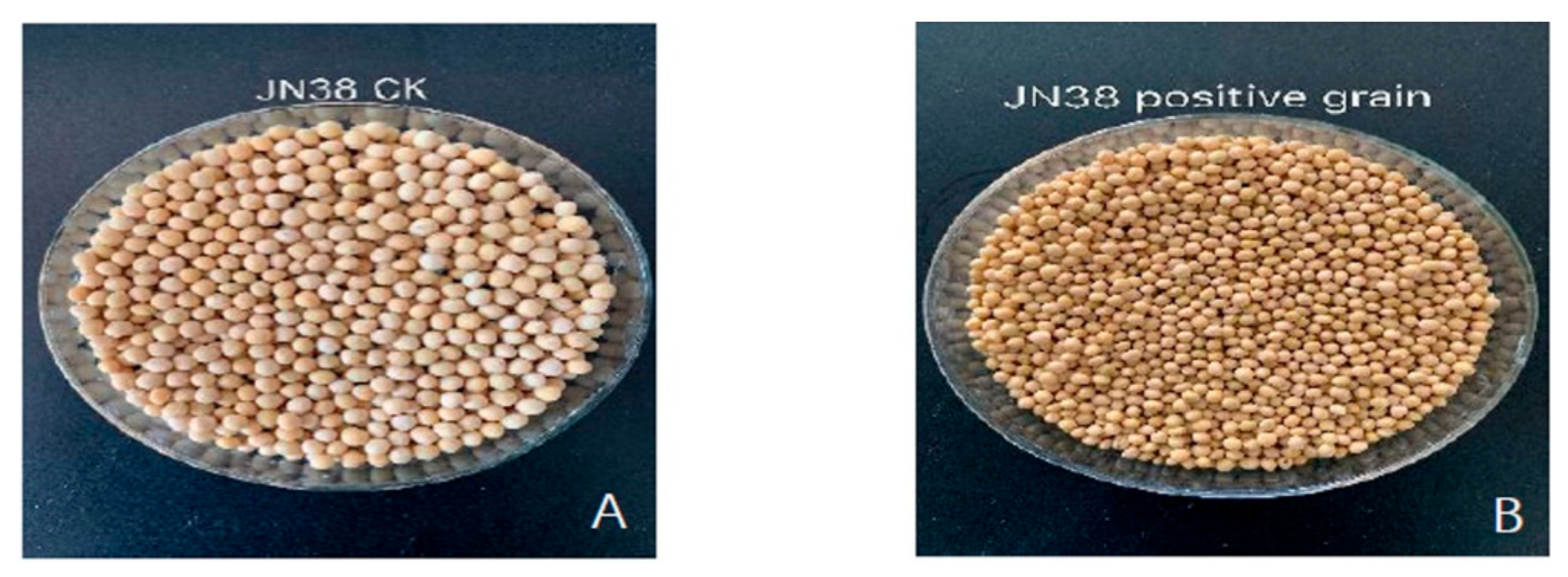
2.10. Phenotypic Analysis of Genetically Edited Progeny
3. Discussion
4. Materials and Methods
4.1. Materials
4.2. Target Design and Detection of gRNA Target Efficiency by Enzyme Activity in Vitro
4.3. Construction of the CRISPR/Cas9 Expression Vector
4.4. Detection of Transgenic Plant Progeny
4.5. Editing Efficiency Test
4.6. Southern Blotting Detection of T3 Transgenic Plants
4.7. Quantitative Real-Time PCR detection of T3 Transgenic Plants
4.8. Determination of Oleic Acid Content in T2, T3 Soybean Seeds Using a Near-Infrared Grain Analyser
4.9. Significant Analysis of Differences in Oleic Acid Content
4.10. Investigation and Phenotypic Analysis of Agronomic Traits in Offspring
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Chapman, K.D.; Ohlrogge, J.B. Compartmentation of Triacylglycerol Accumulation in Plants. J. Biol. Chem. 2012, 287, 2288–2294. [Google Scholar] [CrossRef] [PubMed]
- Haslam, R.P.; Ruiz-Lopez, N.; Eastmond, P.; Moloney, M.; Sayanova, O.; Napier, J.A. The modification of plant oil composition via metabolic engineering—better nutrition by design. Plant Biotechnol. J. 2013, 11, 157–168. [Google Scholar] [CrossRef] [PubMed]
- Zainal, N.A.; Zulkifli, N.W.M.; Gulzar, M.; Masjuki, H.H. A review on the chemistry, production, and technological potential of bio-based lubricants. Renew. Sustain. Energy Rev. 2018, 82, 80–102. [Google Scholar] [CrossRef]
- Haun, W.; Coffman, A.; Clasen, B.M.; Demorest, Z.L.; Lowy, A.; Ray, E.; Retterath, A.; Stoddars, T.; Juillerat, A.; Cedrone, F.; et al. Improved soybean oil quality by targeted mutagenesis of the fatty acid desaturase 2 gene family. Plant Biotechnol. J. 2014, 12, 934–940. [Google Scholar] [CrossRef]
- Pham, A.T.; Lee, J.D.; Shannon, J.G.; Bilyeu, K.D. A novel FAD2-1 A allele in a soybean plant introduction offers an alternate means to produce soybean seed oil with 85% oleic acid content. Theor. Appl. Genet. 2011, 5, 793–802. [Google Scholar] [CrossRef]
- Wagner, N.; Mroczka, A.; Roberts, P.D.; Schreckengost, W.; Voelker, T. RNAi trigger fragment truncation attenuates soybean FAD2-1 transcript suppression and yields intermediate oil phenotypes. Plant Biotechnol. J. 2011, 9, 723–728. [Google Scholar] [CrossRef]
- Buhr, T.; Sato, S.; Ebrahim, F.; Xing, A.; Zhou, Y.; Mathiesen, M.; Schweiger, B.; Kinney, A.; Staswick, P.; Tom, C. Ribozyme termination of RNA transcripts down-regulate seed fatty acid genes in transgenic soybean. Plant J. 2012, 30, 155–163. [Google Scholar] [CrossRef]
- Monsanto Company. Vistive Gold [EB/OL]. Monsanto Company. 2013. Available online: https://www.vistivegold.com/ (accessed on 2 February 2020).
- Pioneer Hi-Bred International, Inc. Plenish [EB/OL]. Pioneer Hi-Bred International. 2013. Available online: http://www.plenish.com/ (accessed on 2 February 2020).
- Cai, Y.; Chen, L.; Liu, X.; Guo, C.; Sun, S.; Wu, C.; Jiang, B.; Han, T.; Hou, W. CRISPR/Cas9-mediated targeted mutagenesis of GmFT2a delays flowering time in soya bean. Plant Biotechnol. J. 2018, 16, 176–185. [Google Scholar] [CrossRef]
- Char, S.N.; Neelakandan, A.K.; Nahampun, H.; Frame, B.; Main, M.; Spalding, M.H.; Becraft, P.W.; Meyers, B.C.; Walbot, V.; Wang, K.; et al. An Agrobacterium-delivered CRISPR/Cas9 system for high-frequency targeted mutagenesis in maize. Plant Biotechnol. J. 2017, 15, 257–268. [Google Scholar] [CrossRef]
- Shan, Q.; Zhang, Y.; Chen, K.; Zhang, K.; Gao, C. Creation of fragrant rice by targeted knockout of the OsBADH2 gene using TALEN technology. Plant Biotechnol. J. 2015, 13, 791–800. [Google Scholar] [CrossRef]
- Bortesi, L.; Fische, R. The CRISPR/Cas9 system for plant genome editing and beyond. Biotechnol. Adv. 2015, 33, 41–52. [Google Scholar] [CrossRef] [PubMed]
- Curtin, S.J.; Xiong, Y.; Michno, J.; Campbell, B.W.; Stec, A.O.; Čermák, T.; Starker, C.; Voytas, D.F.; Eamens, A.L.; Stupar, R.M. CRISPR/Cas9 and TALENs generate heritable mutations for genes involved in small RNA processing of Glycine max and Medicago truncatula. Plant Biotechnol. J. 2018, 16, 1125–1137. [Google Scholar] [CrossRef] [PubMed]
- Morineau, C.; Bellec, Y.; Tellier, F.; Gissot, L.; Kelemen, Z.; Nogué, F.; Faure, J. Selective gene dosage by CRISPR-Cas9 genome editing in hexaploid Camelina sativa. Plant Biotechnol. J. 2017, 15, 729–739. [Google Scholar] [CrossRef] [PubMed]
- Jiang, W.Z. Significant enhancement of fatty acid composition in seeds of the allohexaploid, Camelina sativa, using CRISPR/Cas9 gene editing. Plant Biotechnol. J. 2017, 15, 648–657. [Google Scholar] [CrossRef]
- Yan, L. Using CRISPR/Cas9 Technology to Create Soybean GmWRI1a Gene Mutant. Ph.D. Thesis, Northeast Agricultural University, Harbin, China, 2017. [Google Scholar]
- Hoshino, T.; Takagi, Y.; Anai, T. Novel GmFAD2-1b mutant alleles created by reverse genetics induce marked elevation of oleic acid content in soybean seeds in combination with GmFAD2-1a mutant alleles. Breed. Sci. 2010, 60, 419–425. [Google Scholar] [CrossRef]
- Liu, H.; Ding, Y.; Zhou, Y.; Jin, W.; Xie, K.; Chen, L.L. CRISPR-P 2.0: An Improved CRISPR-Cas9 Tool for Genome Editing in Plants. Mol. Plant 2017, 10, 530–532. [Google Scholar] [CrossRef]
- Zhao, C.; Zheng, X.; Qu, W. CRISPR-offinder: A CRISPR guide RNA design and off-target searching tool for user-defined protospacer adjacent motif. Int. J. Biol. Sci. 2017, 13, 470–478. [Google Scholar] [CrossRef]
- Ferreira, R.; Gatto, F.; Nielsen, J. Exploiting off-targeting in guide-RNAs for CRISPR systems for simultaneous editing of multiple genes. FEBS Lett. 2017, 591, 3288–3295. [Google Scholar] [CrossRef]
- Miao, X.F.; Xu, W.P.; Li, C.D. Analysis of fatty acid composition and content of soybean varieties in Northeast China. Soybean Sci. 2011, 30, 29–531. [Google Scholar]
- Song, X.K.; Zhang, Y.J.; Yan, L. Analysis of the characteristics of related variation of fatty acid components in Soybean. Acta Agric. Boreali Sin. 2011, 25, 68–73. [Google Scholar]
- Liu, X.Y.; Hu, C.X.; Yu, L.J. Analysis of fatty acid composition of soybean germplasm resources in China. Crop Var. Resour. 2009, 27, 40–42. [Google Scholar]
- Zou, X.; Han, F.X.; Chen, M.Y. QTL Mapping of Soybean Fatty Acid Contents. Chin. J. Crop. 2014, 40, 1595–1603. [Google Scholar]
- Jiang, W.Y. CRISPR-assisted editing of bacterial genomes. Nat. Biotechnol. 2013, 31, 233–239. [Google Scholar] [CrossRef] [PubMed]
- Jacobs, T.B.; LaFayette, P.R.; Schmitz, R.J.; Parrott, W.A. Targeted genome modifications in soybean with CRISPR/Cas9. BMC Biotechnol. 2015, 15, 16. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Xing, G.; Niu, L.; He, H.L.; Du, Q.; Guo, D.Q.; Yuan, Y.; Yang, X.D. Antisense RNA-mediated GmFAD2-1B gene silencing enhances the efficient accumulation of oleic acid in soybean seeds. Acta Agron. Sin. 2017, 43, 1588–1595. [Google Scholar] [CrossRef]
- Hou, Z.H.; Wu, Y.; Cheng, Q.; Dong, L.D.; Lu, S.J.; Nan, H.Y.; Gan, Z.R.; Liu, B.H. Using CRISPR/Cas9 Technology to Create Soybean High Oleic Acid Mutant Lines. Acta Agron. Sin. 2019, 45, 839–847. [Google Scholar]
- Wang, D.; Zhang, L.; Fan, S.; Han, X.; Wu, Y.; Wang, Y.; Yao, D.; Zhang, J. Near-infrared model of soybean fatty acid content. Soy Sci. 2017, 36, 295–299. [Google Scholar]
- Zhu, Z.; Chen, S.; Wu, X.; Xing, C.; Yuan, J. Determination of soybean routine quality parameters using near-infrared spectroscopy. Food Sci. Nutr. 2018, 6, 109–118. [Google Scholar] [CrossRef]
- Karn, A.; Heim, C.; Flint-Garci, S.; Bilyeu, K.; Gillman, J. Development of Rigorous Fatty Acid Near-Infrared Spectroscopy Quantitation Methods in Support of Soybean Oil Improvement. J. Am. Oil Chem. Soc. 2017, 94, 69–76. [Google Scholar] [CrossRef]
- Wang, X.R.; Liao, H.; Yan, X.L. Determination of protein and fat content in soybean seeds by near infrared spectroscopy. Soybean Sci. 2006, 24, 199–201. [Google Scholar]
- Yu, F.K.; Sun, J.M.; Han, F.X.; Ge, Y.N.; Zhang, J.Y.; Ma, L.; Zhang, J.W.; Yan, S.R.; Yang, H. Comparative analysis of fast detection methods for fatty acids in soybean seeds. Soybean Sci. 2011, 4, 26–631. [Google Scholar]


| Gene Target | GC% | Edited Efficiency % | Homozygous Mutation Efficiency % |
|---|---|---|---|
| g3 | 52% | 95% (19 plants) | 31.58% (6 plants) |
| g6 | 33.3% | 55.56% (10 plants) | 40% (4 plants) |
| g36 | - | 66.67% (28 plants) | 28.75% (8 plants) |
| T2 Generation Materials | Oleic | Linoleic Acid | Protein | Crude Fat | Oleic Acid Increase Percentage |
|---|---|---|---|---|---|
| JN38CK | 17.10 ± 0.05 | 62.91 ± 0.03 | 37.69 ± 0.05 | 21.96 ± 0.02 | - |
| g3CRA1 | 32.11 ± 0.02 | 48.45 ± 0.05 | 40.20 ± 0.05 | 22.66 ± 0.05 | 87.55 |
| g6CRA1 | 41.35 ± 0.04 | 38.81 ± 0.03 | 40.51 ± 0.03 | 22.21 ± 0.04 | 141.5 |
| g36CRA8 | 73.50 ± 0.02 | 12.23 ± 0.03 | 41.16 ± 0.02 | 20.63 ± 0.03 | 329.3 |
| T3 Generation Materials | Oleic | Linoleic Acid | Protein | Crude Fat | Oleic Acid Increase Percentage |
|---|---|---|---|---|---|
| JN38CK | 19.15 ± 0.03 | 56.58 ± 0.02 | 37.52 ± 0.02 | 21.02 ± 0.04 | - |
| g3CRA1 | 34.47 ± 0.02 | 47.49 ± 0.03 | 40.58 ± 0.03 | 22.79 ± 0.03 | 80.00 |
| g6CRA1 | 40.45 ± 0.04 | 41.69 ± 0.04 | 38.92 ± 0.05 | 23.92 ± 0.05 | 111.2 |
| g36CRA8 | 72.02 ± 0.02 | 17.27 ± 0.02 | 39.51 ± 0.03 | 21.61 ± 0.04 | 276.1 |
| Variety | Plant Height | Section Number | Number of Pods | 100-Grain Weight | Yield | Increase Production Ratio |
|---|---|---|---|---|---|---|
| JN38CK | 86.6 ± 2.89Aa | 15.6 ± 1.08Aa | 27.0 ± 3.59Aa | 18.32 ± 0.31Aa | 0.48 ± 0.04bc | - |
| g3CRA1 | 87.2 ± 3.05Aa | 16.8 ± 1.75Aa | 23.2 ± 4.52Aa | 18.85 ± 0.39Aa | 0.55 ± 0.02c | 14.58 |
| g3CRT5 | 91.5 ± 3.07Aa | 16.5 ± 1.13Aa | 22.8 ± 3.81Aa | 19.07 ± 0.28Aa | 0.47 ± 0.03c | −2.08 |
| g6CRA1 | 93.4 ± 3.25Aa | 14.0 ± 1.19Aa | 24.8 ± 4.05Aa | 19.33 ± 0.19Aa | 0.51 ± 0.07c | 6.25 |
| g6CRT5 | 85.5 ± 2.91Aa | 16.3 ± 1.25Aa | 25.7 ± 3.69Aa | 18.72 ± 0.21Aa | 0.58 ± 0.14a | 20.8 |
| g36CRA8 | 92.3 ± 2.65Aa | 15.8 ± 1.23Aa | 28.2 ± 3.59Aa | 19.21 ± 0.12Aa | 0.49 ± 0.09c | 2.08 |
| g36CRT11 | 91.8 ± 2.92Aa | 16.7 ± 1.05Aa | 29.1 ± 3.72Aa | 18.85 ± 0.24Aa | 0.53 ± 0.05c | 10.4 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wu, N.; Lu, Q.; Wang, P.; Zhang, Q.; Zhang, J.; Qu, J.; Wang, N. Construction and Analysis of GmFAD2-1A and GmFAD2-2A Soybean Fatty Acid Desaturase Mutants Based on CRISPR/Cas9 Technology. Int. J. Mol. Sci. 2020, 21, 1104. https://doi.org/10.3390/ijms21031104
Wu N, Lu Q, Wang P, Zhang Q, Zhang J, Qu J, Wang N. Construction and Analysis of GmFAD2-1A and GmFAD2-2A Soybean Fatty Acid Desaturase Mutants Based on CRISPR/Cas9 Technology. International Journal of Molecular Sciences. 2020; 21(3):1104. https://doi.org/10.3390/ijms21031104
Chicago/Turabian StyleWu, Nan, Qiang Lu, Piwu Wang, Qi Zhang, Jun Zhang, Jing Qu, and Nan Wang. 2020. "Construction and Analysis of GmFAD2-1A and GmFAD2-2A Soybean Fatty Acid Desaturase Mutants Based on CRISPR/Cas9 Technology" International Journal of Molecular Sciences 21, no. 3: 1104. https://doi.org/10.3390/ijms21031104
APA StyleWu, N., Lu, Q., Wang, P., Zhang, Q., Zhang, J., Qu, J., & Wang, N. (2020). Construction and Analysis of GmFAD2-1A and GmFAD2-2A Soybean Fatty Acid Desaturase Mutants Based on CRISPR/Cas9 Technology. International Journal of Molecular Sciences, 21(3), 1104. https://doi.org/10.3390/ijms21031104





