Focus on DNA Glycosylases—A Set of Tightly Regulated Enzymes with a High Potential as Anticancer Drug Targets
Abstract
1. Introduction
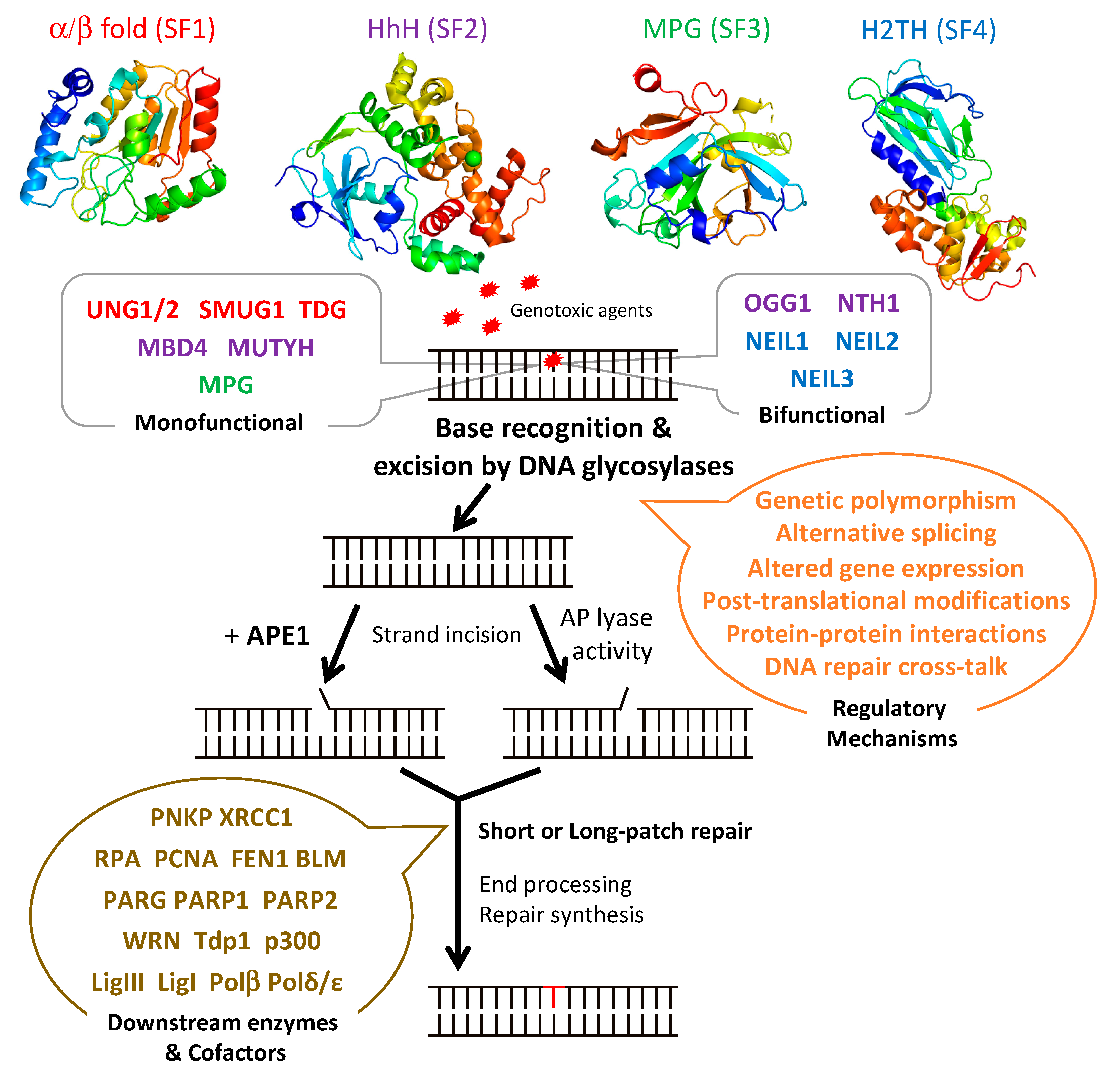
2. DNA Glycosylases and BER
3. Regulation of DNA Glycosylases
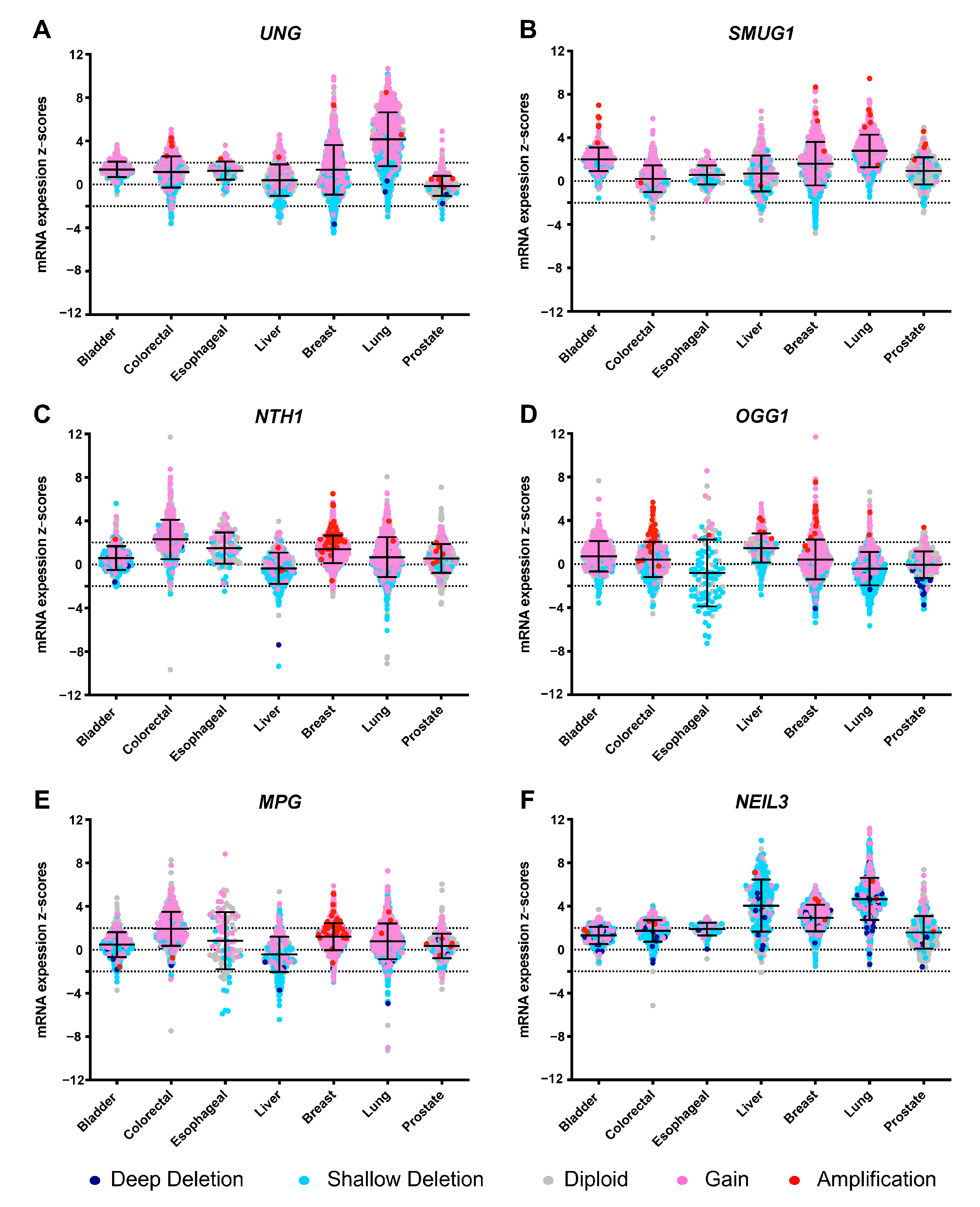
4. Modulating DNA Glycosylase Levels as a Means of Studying Their Implication in Cancer Development
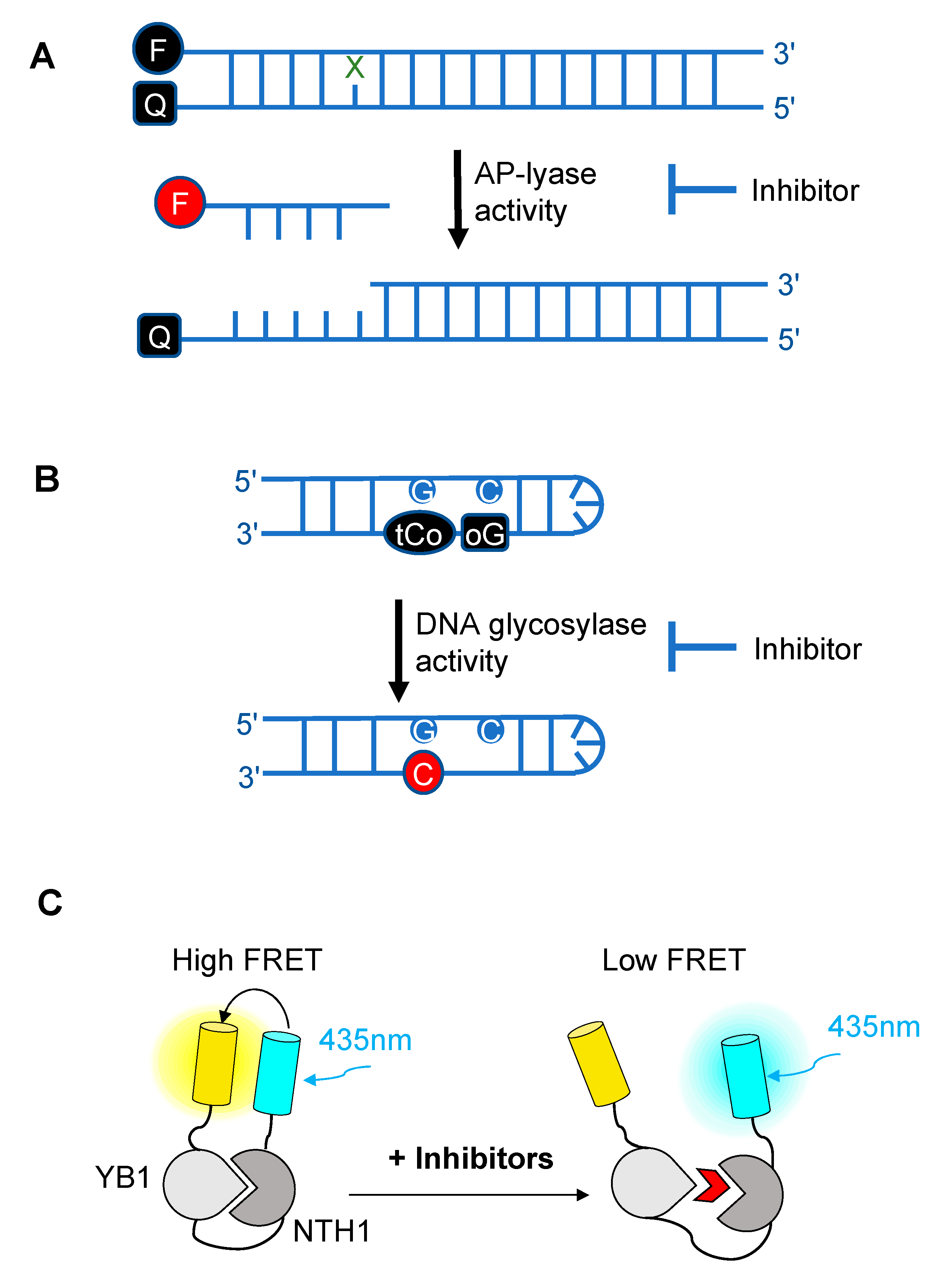
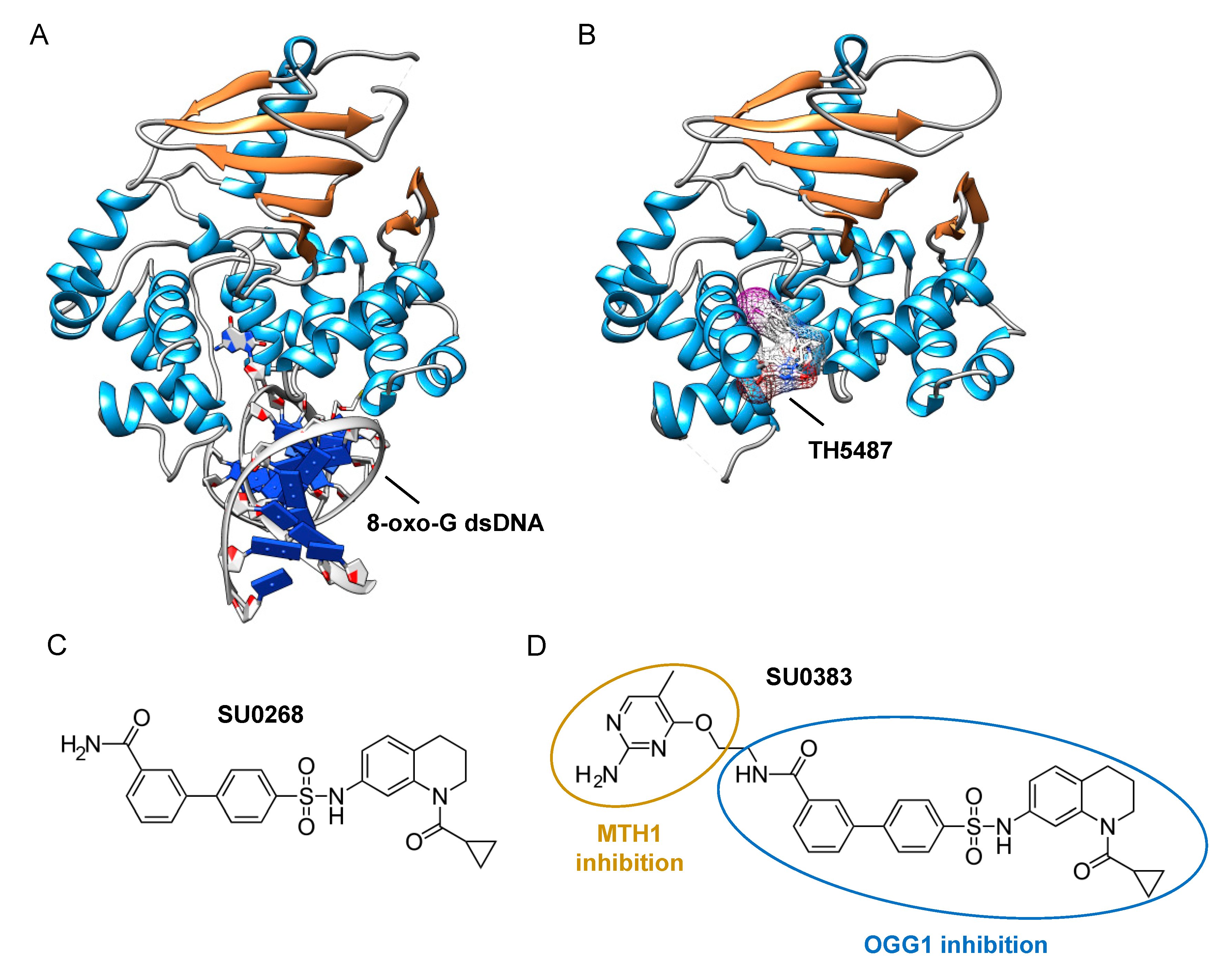
5. Finding Inhibitors of DNA Glycosylases
6. Conclusions
Funding
Conflicts of Interest
References
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [PubMed]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef] [PubMed]
- Hayes, J.D.; Dinkova-Kostova, A.T.; Tew, K.D. Oxidative Stress in Cancer. Cancer Cell 2020, 38, 167–197. [Google Scholar] [CrossRef] [PubMed]
- Van Loon, B.; Markkanen, E.; Hübscher, U. Oxygen as a friend and enemy: How to combat the mutational potential of 8-oxo-guanine. DNA Repair 2010, 9, 604–616. [Google Scholar] [CrossRef]
- Tubbs, A.; Nussenzweig, A. Endogenous DNA Damage as a Source of Genomic Instability in Cancer. Cell 2017, 168, 644–656. [Google Scholar] [CrossRef]
- Turgeon, M.-O.; Perry, N.J.S.; Poulogiannis, G. DNA Damage, Repair, and Cancer Metabolism. Front. Oncol. 2018, 8, 15. [Google Scholar] [CrossRef]
- DeBerardinis, R.J.; Chandel, N.S. Fundamentals of cancer metabolism. Sci. Adv. 2016, 2, e1600200. [Google Scholar] [CrossRef]
- Chae, Y.K.; Anker, J.F.; Carneiro, B.A.; Chandra, S.; Kaplan, J.; Kalyan, A.; Santa-Maria, C.A.; Platanias, L.C.; Giles, F.J. Genomic landscape of DNA repair genes in cancer. Oncotarget 2016, 7, 23312–23321. [Google Scholar] [CrossRef]
- Moretton, A.; Loizou, J.I. Interplay between Cellular Metabolism and the DNA Damage Response in Cancer. Cancers 2020, 12, 2051. [Google Scholar] [CrossRef]
- Liu, J. The dualistic origin of human tumors. Semin. Cancer Biol. 2018, 53, 1–16. [Google Scholar] [CrossRef]
- Herbein, G.; Nehme, Z. Polyploid Giant Cancer Cells, a Hallmark of Oncoviruses and a New Therapeutic Challenge. Front. Oncol. 2020, 10, 567116. [Google Scholar] [CrossRef] [PubMed]
- Ye, C.J.; Sharpe, Z.; Alemara, S.; Mackenzie, S.; Liu, G.; Abdallah, B.; Horne, S.; Regan, S.; Heng, H.H. Micronuclei and Genome Chaos: Changing the System Inheritance. Genes 2019, 10, 366. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Niu, N.; Zhang, J.; Qi, L.; Shen, W.; Donkena, K.V.; Feng, Z.; Liu, J. Polyploid Giant Cancer Cells (PGCCs): The Evil Roots of Cancer. Curr. Cancer Drug Targets 2019, 19, 360–367. [Google Scholar] [CrossRef] [PubMed]
- Alhmoud, J.F.; Woolley, J.F.; Al Moustafa, A.-E.; Malki, M.I. DNA Damage/Repair Management in Cancers. Cancers 2020, 12, 1050. [Google Scholar] [CrossRef]
- Falzone, L.; Salomone, S.; Libra, M. Evolution of Cancer Pharmacological Treatments at the Turn of the Third Millennium. Front. Pharm. 2018, 9, 1300. [Google Scholar] [CrossRef]
- Urruticoechea, A.; Alemany, R.; Balart, J.; Villanueva, A.; Viñals, F.; Capellá, G. Recent advances in cancer therapy: An overview. Curr. Pharm. Des. 2010, 16, 3–10. [Google Scholar] [CrossRef]
- Chabner, B.A.; Roberts, T.G.J. Timeline: Chemotherapy and the war on cancer. Nat. Rev. Cancer 2005, 5, 65–72. [Google Scholar] [CrossRef]
- DeVita, V.T.J.; Chu, E. A history of cancer chemotherapy. Cancer Res. 2008, 68, 8643–8653. [Google Scholar] [CrossRef]
- Galmarini, D.; Galmarini, C.M.; Galmarini, F.C. Cancer chemotherapy: A critical analysis of its 60 years of history. Crit. Rev. Oncol. Hematol. 2012, 84, 181–199. [Google Scholar] [CrossRef]
- Roos, W.P.; Thomas, A.D.; Kaina, B. DNA damage and the balance between survival and death in cancer biology. Nat. Rev. Cancer 2016, 16, 20–33. [Google Scholar] [CrossRef]
- Tang, H.M.; Tang, H.L. Anastasis: Recovery from the brink of cell death. Royal Soc. Open Sci. 2018, 5, 180442. [Google Scholar] [CrossRef] [PubMed]
- Wu, Z.; Li, S.; Tang, X.; Wang, Y.; Guo, W.; Cao, G.; Chen, K.; Zhang, M.; Guan, M.; Yang, D. Copy Number Amplification of DNA Damage Repair Pathways Potentiates Therapeutic Resistance in Cancer. Theranostics 2020, 10, 3939–3951. [Google Scholar] [CrossRef] [PubMed]
- Sarasin, A.; Kauffmann, A. Overexpression of DNA repair genes is associated with metastasis: A new hypothesis. Mutat. Res. 2008, 659, 49–55. [Google Scholar] [CrossRef] [PubMed]
- Bukowski, K.; Kciuk, M.; Kontek, R. Mechanisms of Multidrug Resistance in Cancer Chemotherapy. Int. J. Mol. Sci. 2020, 21, 3233. [Google Scholar] [CrossRef]
- Curtin, N.J. DNA repair dysregulation from cancer driver to therapeutic target. Nat. Rev. Cancer 2012, 12, 801–817. [Google Scholar] [CrossRef]
- Zhou, J.; Kang, Y.; Chen, L.; Wang, H.; Liu, J.; Zeng, S.; Yu, L. The Drug-Resistance Mechanisms of Five Platinum-Based Antitumor Agents. Front. Pharmacol. 2020, 11, 343. [Google Scholar] [CrossRef]
- Carusillo, A.; Mussolino, C. DNA Damage: From Threat to Treatment. Cells 2020, 9, 1665. [Google Scholar] [CrossRef]
- Helleday, T.; Petermann, E.; Lundin, C.; Hodgson, B.; Sharma, R.A. DNA repair pathways as targets for cancer therapy. Nat. Rev. Cancer 2008, 193–204. [Google Scholar] [CrossRef]
- Jackson, S.P.; Bartek, J. The DNA-damage response in human biology and disease. Nature 2009, 461, 1071–1078. [Google Scholar] [CrossRef]
- Lee, K.J.; Mann, E.; Wright, G.; Piett, C.G.; Nagel, Z.D.; Gassman, N.R. Exploiting DNA repair defects in triple negative breast cancer to improve cell killing. Adv. Med. Oncol. 2020, 12. [Google Scholar] [CrossRef]
- D’Andrea, A.D. Mechanisms of PARP inhibitor sensitivity and resistance. DNA Repair 2018, 71, 172–176. [Google Scholar] [CrossRef] [PubMed]
- Javle, M.; Curtin, N.J. The potential for poly (ADP-ribose) polymerase inhibitors in cancer therapy. Adv. Med. Oncol. 2011, 3, 257–267. [Google Scholar] [CrossRef] [PubMed]
- Kamel, D.; Gray, C.; Walia, J.S.; Kumar, V. PARP Inhibitor Drugs in the Treatment of Breast, Ovarian, Prostate and Pancreatic Cancers: An Update of Clinical Trials. Curr. Drug Targets 2018, 19, 21–37. [Google Scholar] [CrossRef] [PubMed]
- Bhattacharjee, S.; Nandi, S. Synthetic lethality in DNA repair network: A novel avenue in targeted cancer therapy and combination therapeutics. IUBMB Life 2017, 69, 929–937. [Google Scholar] [CrossRef] [PubMed]
- Huang, A.; Garraway, L.A.; Ashworth, A.; Weber, B. Synthetic lethality as an engine for cancer drug target discovery. Nat. Rev. Drug Discov. 2020, 19, 23–38. [Google Scholar] [CrossRef] [PubMed]
- Moeller, B.J.; Arap, W.; Pasqualini, R. Targeting synthetic lethality in DNA damage repair pathways as an anti-cancer strategy. Curr. Drug Targets 2010, 11, 1336–1340. [Google Scholar] [CrossRef] [PubMed]
- Yap, T.A.; Plummer, R.; Azad, N.S.; Helleday, T. The DNA Damaging Revolution: PARP Inhibitors and Beyond. Am. Soc. Clin. Oncol. Educ. Book 2019, 39, 185–195. [Google Scholar] [CrossRef]
- Lord, C.J.; Tutt, A.N.J.; Ashworth, A. Synthetic lethality and cancer therapy: Lessons learned from the development of PARP inhibitors. Annu. Rev. Med. 2015, 66, 455–470. [Google Scholar] [CrossRef]
- Hoeijmakers, J.H. Genome maintenance mechanisms for preventing cancer. Nature 2001, 411, 366–374. [Google Scholar] [CrossRef]
- Chatterjee, N.; Walker, G.C. Mechanisms of DNA damage, repair, and mutagenesis. Env. Mol. Mutagen. 2017, 58, 235–263. [Google Scholar] [CrossRef]
- Dizdaroglu, M.; Coskun, E.; Jaruga, P. Repair of oxidatively induced DNA damage by DNA glycosylases: Mechanisms of action, substrate specificities and excision kinetics. Mutat. Res. 2017, 771, 99–127. [Google Scholar] [CrossRef] [PubMed]
- Krokan, H.E.; Bjoras, M. Base excision repair. Cold Spring Harb. Perspect. Biol. 2013, 5, a012583. [Google Scholar] [CrossRef] [PubMed]
- Dizdaroglu, M. Oxidatively Induced DNA Damage and Its Repair in Cancer. Mutat. Res. Rev. Mutat. Res. 2015, 763, 212–245. [Google Scholar] [CrossRef] [PubMed]
- Mechetin, G.V.; Endutkin, A.V.; Diatlova, E.A.; Zharkov, D.O. Inhibitors of DNA Glycosylases as Prospective Drugs. Int. J. Mol. Sci. 2020, 21, 3118. [Google Scholar] [CrossRef] [PubMed]
- Visnes, T.; Grube, M.; Hanna, B.M.F.; Benitez-Buelga, C.; Cázares-Körner, A.; Helleday, T. Targeting BER enzymes in cancer therapy. DNA Repair 2018, 71, 118–126. [Google Scholar] [CrossRef] [PubMed]
- Bauer, N.C.; Corbett, A.H.; Doetsch, P.W. The current state of eukaryotic DNA base damage and repair. Nucleic Acids Res. 2015, 43, 10083–10101. [Google Scholar] [CrossRef]
- Wallace, S.S. Base excision repair: A critical player in many games. DNA Repair 2014, 19, 14–26. [Google Scholar] [CrossRef]
- Hegde, M.L.; Hazra, T.K.; Mitra, S. Early steps in the DNA base excision/single-strand interruption repair pathway in mammalian cells. Cell Res. 2008, 18, 27–47. [Google Scholar] [CrossRef]
- Jacobs, A.L.; Schar, P. DNA glycosylases: In DNA repair and beyond. Chromosoma 2012, 121, 1–20. [Google Scholar] [CrossRef]
- McCullough, A.K.; Dodson, M.L.; Lloyd, R.S. Initiation of base excision repair: Glycosylase mechanisms and structures. Annu. Rev. Biochem. 1999, 68, 255–285. [Google Scholar] [CrossRef]
- Kim, Y.-J.; Wilson, D.M., 3rd. Overview of base excision repair biochemistry. Curr. Mol. Pharm. 2012, 5, 3–13. [Google Scholar] [CrossRef] [PubMed]
- Dizdaroglu, M. Base-excision repair of oxidative DNA damage by DNA glycosylases. Mutat. Res. Fundam. Mol. Mech. Mutagenesis 2005, 591, 45–59. [Google Scholar] [CrossRef] [PubMed]
- Sampath, H.; McCullough, A.K.; Lloyd, R.S. Regulation of DNA glycosylases and their role in limiting disease. Free Radic. Res. 2012, 46, 460–478. [Google Scholar] [CrossRef] [PubMed]
- Hollis, T.; Ichikawa, Y.; Ellenberger, T. DNA bending and a flip-out mechanism for base excision by the helix-hairpin-helix DNA glycosylase, Escherichia coli AlkA. EMBO J. 2000, 19, 758–766. [Google Scholar] [CrossRef] [PubMed]
- Thayer, M.M.; Ahern, H.; Xing, D.; Cunningham, R.P.; Tainer, J.A. Novel DNA binding motifs in the DNA repair enzyme endonuclease III crystal structure. EMBO J. 1995, 14, 4108–4120. [Google Scholar] [CrossRef]
- Bandaru, V.; Sunkara, S.; Wallace, S.S.; Bond, J.P. A novel human DNA glycosylase that removes oxidative DNA damage and is homologous to Escherichia coli endonuclease VIII. DNA Repair 2002, 1, 517–529. [Google Scholar] [CrossRef]
- Doublié, S.; Bandaru, V.; Bond, J.P.; Wallace, S.S. The crystal structure of human endonuclease VIII-like 1 (NEIL1) reveals a zincless finger motif required for glycosylase activity. Proc. Natl. Acad. Sci. USA 2004, 101, 10284–10289. [Google Scholar] [CrossRef]
- Dalhus, B.; Laerdahl, J.K.; Backe, P.H.; Bjørås, M. DNA base repair--recognition and initiation of catalysis. FEMS Microbiol. Rev. 2009, 33, 1044–1078. [Google Scholar] [CrossRef]
- Whitaker, A.M.; Schaich, M.A.; Smith, M.R.; Flynn, T.S.; Freudenthal, B.D. Base excision repair of oxidative DNA damage: From mechanism to disease. Front. Biosci. 2017, 22, 1493–1522. [Google Scholar]
- Nilsen, H.; Otterlei, M.; Haug, T.; Solum, K.; Nagelhus, T.A.; Skorpen, F.; Krokan, H.E. Nuclear and mitochondrial uracil-DNA glycosylases are generated by alternative splicing and transcription from different positions in the UNG gene. Nucleic Acids Res. 1997, 25, 750–755. [Google Scholar] [CrossRef]
- Sarno, A.; Lundbæk, M.; Liabakk, N.B.; Aas, P.A.; Mjelle, R.; Hagen, L.; Sousa, M.M.L.; Krokan, H.E.; Kavli, B. Uracil-DNA glycosylase UNG1 isoform variant supports class switch recombination and repairs nuclear genomic uracil. Nucleic Acids Res. 2019, 47, 4569–4585. [Google Scholar] [CrossRef] [PubMed]
- Hwang, J.K.; Alt, F.W.; Yeap, L.-S. Related Mechanisms of Antibody Somatic Hypermutation and Class Switch Recombination. Microbiol. Spectr. 2015, 3, MDNA3-0037-2014. [Google Scholar] [PubMed]
- Pilzecker, B.; Jacobs, H. Mutating for Good: DNA Damage Responses during Somatic Hypermutation. Front. Immunol. 2019, 10, 438. [Google Scholar] [CrossRef] [PubMed]
- Bellacosa, A.; Drohat, A.C. Role of base excision repair in maintaining the genetic and epigenetic integrity of CpG sites. DNA Repair 2015, 32, 33–42. [Google Scholar] [CrossRef] [PubMed]
- Ito, S.; Kuraoka, I. Epigenetic modifications in DNA could mimic oxidative DNA damage: A double-edged sword. DNA Repair 2015, 32, 52–57. [Google Scholar] [CrossRef]
- Roth, R.B.; Samson, L.D. 3-Methyladenine DNA glycosylase-deficient Aag null mice display unexpected bone marrow alkylation resistance. Cancer Res. 2002, 62, 656–660. [Google Scholar]
- Prakash, A.; Doublié, S.; Wallace, S.S. The Fpg/Nei family of DNA glycosylases: Substrates, structures, and search for damage. Prog. Mol. Biol. Transl. Sci. 2012, 110, 71–91. [Google Scholar]
- Brenerman, B.M.; Illuzzi, J.L.; Wilson, D.M., 3rd. Base excision repair capacity in informing healthspan. Carcinogenesis 2014, 35, 2643–2652. [Google Scholar] [CrossRef]
- Mol, C.D.; Arvai, A.S.; Slupphaug, G.; Kavli, B.; Alseth, I.; Krokan, H.E.; Tainer, J.A. Crystal structure and mutational analysis of human uracil-DNA glycosylase: Structural basis for specificity and catalysis. Cell 1995, 80, 869–878. [Google Scholar] [CrossRef]
- Mol, C.D.; Arvai, A.S.; Sanderson, R.J.; Slupphaug, G.; Kavli, B.; Krokan, H.E.; Mosbaugh, D.W.; Tainer, J.A. Crystal structure of human uracil-DNA glycosylase in complex with a protein inhibitor: Protein mimicry of DNA. Cell 1995, 82, 701–708. [Google Scholar] [CrossRef]
- Schormann, N.; Ricciardi, R.; Chattopadhyay, D. Uracil-DNA glycosylases-structural and functional perspectives on an essential family of DNA repair enzymes. Protein Sci. 2014, 23, 1667–1685. [Google Scholar] [CrossRef] [PubMed]
- Kavli, B.; Sundheim, O.; Akbari, M.; Otterlei, M.; Nilsen, H.; Skorpen, F.; Aas, P.A.; Hagen, L.; Krokan, H.E.; Slupphaug, G. hUNG2 is the major repair enzyme for removal of uracil from U:A matches, U:G mismatches, and U in single-stranded DNA, with hSMUG1 as a broad specificity backup. J. Biol. Chem. 2002, 277, 39926–39936. [Google Scholar] [CrossRef] [PubMed]
- Lu, X.; Bocangel, D.; Nannenga, B.; Yamaguchi, H.; Appella, E.; Donehower, L.A. The p53-induced oncogenic phosphatase PPM1D interacts with uracil DNA glycosylase and suppresses base excision repair. Mol. Cell 2004, 15, 621–634. [Google Scholar] [CrossRef] [PubMed]
- Slupphaug, G.; Eftedal, I.; Kavli, B.; Bharati, S.; Helle, N.M.; Haug, T.; Levine, D.W.; Krokan, H.E. Properties of a recombinant human uracil-DNA glycosylase from the UNG gene and evidence that UNG encodes the major uracil-DNA glycosylase. Biochemistry 1995, 34, 128–138. [Google Scholar] [CrossRef] [PubMed]
- Hagen, L.; Kavli, B.; Sousa, M.M.L.; Torseth, K.; Liabakk, N.B.; Sundheim, O.; Pena-Diaz, J.; Otterlei, M.; Hørning, O.; Jensen, O.N.; et al. Cell cycle-specific UNG2 phosphorylations regulate protein turnover, activity and association with RPA. EMBO J. 2008, 27, 51–61. [Google Scholar] [CrossRef] [PubMed]
- Fischer, J.A.; Muller-Weeks, S.; Caradonna, S. Proteolytic degradation of the nuclear isoform of uracil-DNA glycosylase occurs during the S phase of the cell cycle. DNA Repair 2004, 3, 505–513. [Google Scholar] [CrossRef]
- Hardeland, U.; Kunz, C.; Focke, F.; Szadkowski, M.; Schär, P. Cell cycle regulation as a mechanism for functional separation of the apparently redundant uracil DNA glycosylases TDG and UNG2. Nucleic Acids Res. 2007, 35, 3859–3867. [Google Scholar] [CrossRef]
- Bao, Y.; Tong, L.; Song, B.; Liu, G.; Zhu, Q.; Lu, X.; Zhang, J.; Lu, Y.-F.; Wen, H.; Tian, Y.; et al. UNG2 deacetylation confers cancer cell resistance to hydrogen peroxide-induced cytotoxicity. Free Radic. Biol. Med. 2020, 160, 403–417. [Google Scholar] [CrossRef]
- Iveland, T.S.; Hagen, L.; Sharma, A.; Sousa, M.M.L.; Sarno, A.; Wollen, K.L.; Liabakk, N.B.; Slupphaug, G. HDACi mediate UNG2 depletion, dysregulated genomic uracil and altered expression of oncoproteins and tumor suppressors in B- and T-cell lines. J. Transl. Med. 2020, 18, 159. [Google Scholar] [CrossRef]
- Masaoka, A.; Matsubara, M.; Hasegawa, R.; Tanaka, T.; Kurisu, S.; Terato, H.; Ohyama, Y.; Karino, N.; Matsuda, A.; Ide, H. Mammalian 5-formyluracil-DNA glycosylase. 2. Role of SMUG1 uracil-DNA glycosylase in repair of 5-formyluracil and other oxidized and deaminated base lesions. Biochemistry 2003, 42, 5003–5012. [Google Scholar] [CrossRef]
- Goto, M.; Shinmura, K.; Matsushima, Y.; Ishino, K.; Yamada, H.; Totsuka, Y.; Matsuda, T.; Nakagama, H.; Sugimura, H. Human DNA glycosylase enzyme TDG repairs thymine mispaired with exocyclic etheno-DNA adducts. Free Radic. Biol. Med. 2014, 76, 136–146. [Google Scholar] [CrossRef] [PubMed]
- Hang, B.; Guliaev, A.B. Substrate specificity of human thymine-DNA glycosylase on exocyclic cytosine adducts. Chem. Biol. Interact. 2007, 165, 230–238. [Google Scholar] [CrossRef] [PubMed]
- Hardeland, U.; Steinacher, R.; Jiricny, J.; Schär, P. Modification of the human thymine-DNA glycosylase by ubiquitin-like proteins facilitates enzymatic turnover. EMBO J. 2002, 21, 1456–1464. [Google Scholar] [CrossRef] [PubMed]
- Maiti, A.; Morgan, M.T.; Pozharski, E.; Drohat, A.C. Crystal structure of human thymine DNA glycosylase bound to DNA elucidates sequence-specific mismatch recognition. Proc. Natl. Acad. Sci. USA 2008, 105, 8890–8895. [Google Scholar] [CrossRef]
- Tini, M.; Benecke, A.; Um, S.-J.; Torchia, J.; Evans, R.M.; Chambon, P. Association of CBP/p300 acetylase and thymine DNA glycosylase links DNA repair and transcription. Mol. Cell 2002, 9, 265–277. [Google Scholar] [CrossRef]
- Um, S.; Harbers, M.; Benecke, A.; Pierrat, B.; Losson, R.; Chambon, P. Retinoic acid receptors interact physically and functionally with the T:G mismatch-specific thymine-DNA glycosylase. J. Biol. Chem. 1998, 273, 20728–20736. [Google Scholar] [CrossRef]
- Mohan, R.D.; Litchfield, D.W.; Torchia, J.; Tini, M. Opposing regulatory roles of phosphorylation and acetylation in DNA mispair processing by thymine DNA glycosylase. Nucleic Acids Res. 2010, 38, 1135–1148. [Google Scholar] [CrossRef]
- Shibata, E.; Dar, A.; Dutta, A. CRL4Cdt2 E3 ubiquitin ligase and proliferating cell nuclear antigen (PCNA) cooperate to degrade thymine DNA glycosylase in S phase. J. Biol. Chem. 2014, 289, 23056–23064. [Google Scholar] [CrossRef]
- Slenn, T.J.; Morris, B.; Havens, C.G.; Freeman, R.M.J.; Takahashi, T.S.; Walter, J.C. Thymine DNA glycosylase is a CRL4Cdt2 substrate. J. Biol. Chem. 2014, 289, 23043–23055. [Google Scholar] [CrossRef]
- Manvilla, B.A.; Maiti, A.; Begley, M.C.; Toth, E.A.; Drohat, A.C. Crystal structure of human methyl-binding domain IV glycosylase bound to abasic DNA. J. Mol. Biol. 2012, 420, 164–175. [Google Scholar] [CrossRef]
- Otani, J.; Arita, K.; Kato, T.; Kinoshita, M.; Kimura, H.; Suetake, I.; Tajima, S.; Ariyoshi, M.; Shirakawa, M. Structural basis of the versatile DNA recognition ability of the methyl-CpG binding domain of methyl-CpG binding domain protein 4. J. Biol. Chem. 2013, 288, 6351–6362. [Google Scholar] [CrossRef] [PubMed]
- Yakovlev, D.A.; Kuznetsova, A.A.; Fedorova, O.S.; Kuznetsov, N.A. Search for Modified DNA Sites with the Human Methyl-CpG-Binding Enzyme MBD4. Acta Nat. 2017, 9, 88–98. [Google Scholar] [CrossRef]
- Bhakat, K.K.; Mokkapati, S.K.; Boldogh, I.; Hazra, T.K.; Mitra, S. Acetylation of human 8-oxoguanine-DNA glycosylase by p300 and its role in 8-oxoguanine repair in vivo. Mol. Cell. Biol. 2006, 26, 1654–1665. [Google Scholar] [CrossRef] [PubMed]
- Dantzer, F.; Luna, L.; Bjørås, M.; Seeberg, E. Human OGG1 undergoes serine phosphorylation and associates with the nuclear matrix and mitotic chromatin in vivo. Nucleic Acids Res. 2002, 30, 2349–2357. [Google Scholar] [CrossRef]
- Jaiswal, M.; LaRusso, N.F.; Nishioka, N.; Nakabeppu, Y.; Gores, G.J. Human Ogg1, a protein involved in the repair of 8-oxoguanine, is inhibited by nitric oxide. Cancer Res. 2001, 61, 6388–6393. [Google Scholar]
- Shinmura, K.; Yokota, J. The OGG1 gene encodes a repair enzyme for oxidatively damaged DNA and is involved in human carcinogenesis. Antioxid. Redox Signal. 2001, 3, 597–609. [Google Scholar] [CrossRef]
- Sidorenko, V.S.; Grollman, A.P.; Jaruga, P.; Dizdaroglu, M.; Zharkov, D.O. Substrate specificity and excision kinetics of natural polymorphic variants and phosphomimetic mutants of human 8-oxoguanine-DNA glycosylase. FEBS J. 2009, 276, 5149–5162. [Google Scholar] [CrossRef]
- Hu, J.; Imam, S.Z.; Hashiguchi, K.; de Souza-Pinto, N.C.; Bohr, V.A. Phosphorylation of human oxoguanine DNA glycosylase (alpha-OGG1) modulates its function. Nucleic Acids Res. 2005, 33, 3271–3282. [Google Scholar] [CrossRef]
- Luna, L.; Rolseth, V.; Hildrestrand, G.A.; Otterlei, M.; Dantzer, F.; Bjørås, M.; Seeberg, E. Dynamic relocalization of hOGG1 during the cell cycle is disrupted in cells harbouring the hOGG1-Cys326 polymorphic variant. Nucleic Acids Res. 2005, 33, 1813–1824. [Google Scholar] [CrossRef]
- Gu, Y.; Parker, A.; Wilson, T.M.; Bai, H.; Chang, D.-Y.; Lu, A.-L. Human MutY homolog, a DNA glycosylase involved in base excision repair, physically and functionally interacts with mismatch repair proteins human MutS homolog 2/human MutS homolog 6. J. Biol. Chem. 2002, 277, 11135–11142. [Google Scholar] [CrossRef]
- Kundu, S.; Brinkmeyer, M.K.; Eigenheer, R.A.; David, S.S. Ser 524 is a phosphorylation site in MUTYH and Ser 524 mutations alter 8-oxoguanine (OG): A mismatch recognition. DNA Repair 2010, 9, 1026–1037. [Google Scholar] [CrossRef] [PubMed]
- Markkanen, E.; Dorn, J.; Hübscher, U. MUTYH DNA glycosylase: The rationale for removing undamaged bases from the DNA. Front. Genet. 2013, 4, 18. [Google Scholar] [CrossRef] [PubMed]
- Parker, A.R.; O’Meally, R.N.; Sahin, F.; Su, G.H.; Racke, F.K.; Nelson, W.G.; DeWeese, T.L.; Eshleman, J.R. Defective human MutY phosphorylation exists in colorectal cancer cell lines with wild-type MutY alleles. J. Biol. Chem. 2003, 278, 47937–47945. [Google Scholar] [CrossRef] [PubMed]
- Dorn, J.; Ferrari, E.; Imhof, R.; Ziegler, N.; Hübscher, U. Regulation of human MutYH DNA glycosylase by the E3 ubiquitin ligase mule. J. Biol. Chem. 2014, 289, 7049–7058. [Google Scholar] [CrossRef]
- Eide, L.; Luna, L.; Gustad, E.C.; Henderson, P.T.; Essigmann, J.M.; Demple, B.; Seeberg, E. Human endonuclease III acts preferentially on DNA damage opposite guanine residues in DNA. Biochemistry 2001, 40, 6653–6659. [Google Scholar] [CrossRef]
- Marenstein, D.R.; Chan, M.K.; Altamirano, A.; Basu, A.K.; Boorstein, R.J.; Cunningham, R.P.; Teebor, G.W. Substrate Specificity of Human Endonuclease III (hNTH1): EFFECT OF HUMAN APE1 ON hNTH1 ACTIVITY. J. Biol. Chem. 2003, 278, 9005–9012. [Google Scholar] [CrossRef]
- Williams, S.C.; Parsons, J.L. NTH1 Is a New Target for Ubiquitylation-Dependent Regulation by TRIM26 Required for the Cellular Response to Oxidative Stress. Mol. Cell. Biol. 2018, 38. [Google Scholar] [CrossRef]
- Agnihotri, S.; Burrell, K.; Buczkowicz, P.; Remke, M.; Golbourn, B.; Chornenkyy, Y.; Gajadhar, A.; Fernandez, N.A.; Clarke, I.D.; Barszczyk, M.S.; et al. ATM regulates 3-methylpurine-DNA glycosylase and promotes therapeutic resistance to alkylating agents. Cancer Discov. 2014, 4, 1198–1213. [Google Scholar] [CrossRef]
- Lee, C.-Y.I.; Delaney, J.C.; Kartalou, M.; Lingaraju, G.M.; Maor-Shoshani, A.; Essigmann, J.M.; Samson, L.D. Recognition and processing of a new repertoire of DNA substrates by human 3-methyladenine DNA glycosylase (AAG). Biochemistry 2009, 48, 1850–1861. [Google Scholar] [CrossRef]
- Likhite, V.S.; Cass, E.I.; Anderson, S.D.; Yates, J.R.; Nardulli, A.M. Interaction of estrogen receptor alpha with 3-methyladenine DNA glycosylase modulates transcription and DNA repair. J. Biol. Chem. 2004, 279, 16875–16882. [Google Scholar] [CrossRef]
- O’Brien, P.J.; Ellenberger, T. Dissecting the broad substrate specificity of human 3-methyladenine-DNA glycosylase. J. Biol. Chem. 2004, 279, 9750–9757. [Google Scholar] [CrossRef] [PubMed]
- Hazra, T.K.; Izumi, T.; Boldogh, I.; Imhoff, B.; Kow, Y.W.; Jaruga, P.; Dizdaroglu, M.; Mitra, S. Identification and characterization of a human DNA glycosylase for repair of modified bases in oxidatively damaged DNA. Proc. Natl. Acad. Sci. USA 2002, 99, 3523. [Google Scholar] [CrossRef] [PubMed]
- Slyvka, A.; Mierzejewska, K.; Bochtler, M. Nei-like 1 (NEIL1) excises 5-carboxylcytosine directly and stimulates TDG-mediated 5-formyl and 5-carboxylcytosine excision. Sci. Rep. 2017, 7, 9001. [Google Scholar] [CrossRef] [PubMed]
- Bhakat, K.K.; Hazra, T.K.; Mitra, S. Acetylation of the human DNA glycosylase NEIL2 and inhibition of its activity. Nucleic Acids Res. 2004, 32, 3033–3039. [Google Scholar] [CrossRef]
- Hazra, T.K.; Kow, Y.W.; Hatahet, Z.; Imhoff, B.; Boldogh, I.; Mokkapati, S.K.; Mitra, S.; Izumi, T. Identification and characterization of a novel human DNA glycosylase for repair of cytosine-derived lesions. J. Biol. Chem. 2002, 277, 30417–30420. [Google Scholar] [CrossRef]
- Krokeide, S.Z.; Laerdahl, J.K.; Salah, M.; Luna, L.; Cederkvist, F.H.; Fleming, A.M.; Burrows, C.J.; Dalhus, B.; Bjørås, M. Human NEIL3 is mainly a monofunctional DNA glycosylase removing spiroimindiohydantoin and guanidinohydantoin. DNA Repair 2013, 12, 1159–1164. [Google Scholar] [CrossRef]
- Liu, M.; Doublié, S.; Wallace, S.S. Neil3, the final frontier for the DNA glycosylases that recognize oxidative damage. Mutat. Res. 2013, 743–744, 4–11. [Google Scholar] [CrossRef]
- Liu, M.; Bandaru, V.; Bond, J.P.; Jaruga, P.; Zhao, X.; Christov, P.P.; Burrows, C.J.; Rizzo, C.J.; Dizdaroglu, M.; Wallace, S.S. The mouse ortholog of NEIL3 is a functional DNA glycosylase in vitro and in vivo. Proc. Natl. Acad. Sci. USA 2010, 107, 4925–4930. [Google Scholar] [CrossRef]
- Sweasy, J.B.; Lang, T.; DiMaio, D. Is base excision repair a tumor suppressor mechanism? Cell Cycle 2006, 5, 250–259. [Google Scholar] [CrossRef][Green Version]
- Marsden, C.G.; Dragon, J.A.; Wallace, S.S.; Sweasy, J.B. Base Excision Repair Variants in Cancer. Methods Enzym. 2017, 591, 119–157. [Google Scholar]
- Bravard, A.; Vacher, M.; Moritz, E.; Vaslin, L.; Hall, J.; Epe, B.; Radicella, J.P. Oxidation status of human OGG1-S326C polymorphic variant determines cellular DNA repair capacity. Cancer Res. 2009, 69, 3642–3649. [Google Scholar] [CrossRef] [PubMed]
- Montero-Montoya, R.D.; López-Vargas, R.; Méndez-Serrano, A.; Galicia-Alonso, I.; García-Vargas, G.; Serrano-García, L.; Beltrán-Portugal, R.; Rosado-Zaidi, S.; Albores-Medina, A.; Oropeza-Hernández, L.; et al. Increased micronucleus frequencies in reticulocytes of children exposed to industrial pollution: Oxidative stress and the OGG1 S326C polymorphism. Mutat. Res. 2020, 853, 503170. [Google Scholar] [CrossRef] [PubMed]
- Simonelli, V.; Camerini, S.; Mazzei, F.; Van Loon, B.; Allione, A.; D’Errico, M.; Barone, F.; Minoprio, A.; Ricceri, F.; Guarrera, S.; et al. Genotype-phenotype analysis of S326C OGG1 polymorphism: A risk factor for oxidative pathologies. Free Radic. Biol. Med. 2013, 63, 401–409. [Google Scholar] [CrossRef] [PubMed]
- Sugimura, H.; Kohno, T.; Wakai, K.; Nagura, K.; Genka, K.; Igarashi, H.; Morris, B.J.; Baba, S.; Ohno, Y.; Gao, C.; et al. hOGG1 Ser326Cys polymorphism and lung cancer susceptibility. Cancer Epidemiol. Biomark. Prev. 1999, 8, 669–674. [Google Scholar]
- Morreall, J.; Limpose, K.; Sheppard, C.; Kow, Y.W.; Werner, E.; Doetsch, P.W. Inactivation of a common OGG1 variant by TNF-alpha in mammalian cells. DNA Repair 2015, 26, 15–22. [Google Scholar] [CrossRef]
- Dherin, C.; Radicella, J.P.; Dizdaroglu, M.; Boiteux, S. Excision of oxidatively damaged DNA bases by the human alpha-hOgg1 protein and the polymorphic alpha-hOgg1(Ser326Cys) protein which is frequently found in human populations. Nucleic Acids Res. 1999, 27, 4001–4007. [Google Scholar] [CrossRef]
- Weren, R.D.; Ligtenberg, M.J.; Geurts van Kessel, A.; De Voer, R.M.; Hoogerbrugge, N.; Kuiper, R.P. NTHL1 and MUTYH polyposis syndromes: Two sides of the same coin? J. Pathol. 2018, 244, 135–142. [Google Scholar] [CrossRef]
- Grolleman, J.E.; de Voer, R.M.; Elsayed, F.A.; Nielsen, M.; Weren, R.D.A.; Palles, C.; Ligtenberg, M.J.L.; Vos, J.R.; Ten Broeke, S.W.; de Miranda, N.F.C.C.; et al. Mutational Signature Analysis Reveals NTHL1 Deficiency to Cause a Multi-tumor Phenotype. Cancer Cell 2019, 35, 256–266.e5. [Google Scholar] [CrossRef]
- Galick, H.A.; Kathe, S.; Liu, M.; Robey-Bond, S.; Kidane, D.; Wallace, S.S.; Sweasy, J.B. Germ-line variant of human NTH1 DNA glycosylase induces genomic instability and cellular transformation. Proc. Natl. Acad. Sci. USA 2013, 110, 14314–14319. [Google Scholar] [CrossRef]
- Galick, H.A.; Marsden, C.G.; Kathe, S.; Dragon, J.A.; Volk, L.; Nemec, A.A.; Wallace, S.S.; Prakash, A.; Doublié, S.; Sweasy, J.B. The NEIL1 G83D germline DNA glycosylase variant induces genomic instability and cellular transformation. Oncotarget 2017, 8, 85883–85895. [Google Scholar] [CrossRef]
- Boldinova, E.O.; Khairullin, R.F.; Makarova, A.V.; Zharkov, D.O. Isoforms of Base Excision Repair Enzymes Produced by Alternative Splicing. Int. J. Mol. Sci. 2019, 20, 3279. [Google Scholar] [CrossRef] [PubMed]
- Otterlei, M.; Haug, T.; Nagelhus, T.A.; Slupphaug, G.; Lindmo, T.; Krokan, H.E. Nuclear and mitochondrial splice forms of human uracil-DNA glycosylase contain a complex nuclear localisation signal and a strong classical mitochondrial localisation signal, respectively. Nucleic Acids Res. 1998, 26, 4611–4617. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, R.; Yamamoto, M.; Kusaka, H.; Masatsugu, H.; Matsuyama, S.; Tajima, T.; Ide, H.; Kubo, K. NEIL1 mRNA splicing variants are expressed in normal mouse organs. J. Radiat. Res. 2012, 53, 234–241. [Google Scholar] [CrossRef] [PubMed]
- Kohno, T.; Shinmura, K.; Tosaka, M.; Tani, M.; Kim, S.R.; Sugimura, H.; Nohmi, T.; Kasai, H.; Yokota, J. Genetic polymorphisms and alternative splicing of the hOGG1 gene, that is involved in the repair of 8-hydroxyguanine in damaged DNA. Oncogene 1998, 16, 3219–3225. [Google Scholar] [CrossRef]
- Shinmura, K.; Yamaguchi, S.; Saitoh, T.; Takeuchi-Sasaki, M.; Kim, S.R.; Nohmi, T.; Yokota, J. Adenine excisional repair function of MYH protein on the adenine:8-hydroxyguanine base pair in double-stranded DNA. Nucleic Acids Res. 2000, 28, 4912–4918. [Google Scholar] [CrossRef]
- Radicella, J.P.; Dherin, C.; Desmaze, C.; Fox, M.S.; Boiteux, S. Cloning and characterization of hOGG1, a human homolog of the OGG1 gene of Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. USA 1997, 94, 8010–8015. [Google Scholar] [CrossRef]
- Kuo, F.C.; Sklar, J. Augmented expression of a human gene for 8-oxoguanine DNA glycosylase (MutM) in B lymphocytes of the dark zone in lymph node germinal centers. J. Exp. Med. 1997, 186, 1547–1556. [Google Scholar] [CrossRef]
- Imai, K.; Sarker, A.H.; Akiyama, K.; Ikeda, S.; Yao, M.; Tsutsui, K.; Shohmori, T.; Seki, S. Genomic structure and sequence of a human homologue (NTHL1/NTH1) of Escherichia coli endonuclease III with those of the adjacent parts of TSC2 and SLC9A3R2 genes. Gene 1998, 222, 287–295. [Google Scholar] [CrossRef]
- Boldogh, I.; Milligan, D.; Lee, M.S.; Bassett, H.; Lloyd, R.S.; McCullough, A.K. hMYH cell cycle-dependent expression, subcellular localization and association with replication foci: Evidence suggesting replication-coupled repair of adenine:8-oxoguanine mispairs. Nucleic Acids Res. 2001, 29, 2802–2809. [Google Scholar] [CrossRef]
- Cerami, E.; Gao, J.; Dogrusoz, U.; Gross, B.E.; Sumer, S.O.; Aksoy, B.A.; Jacobsen, A.; Byrne, C.J.; Heuer, M.L.; Larsson, E.; et al. The cBio cancer genomics portal: An open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012, 2, 401–404. [Google Scholar] [CrossRef]
- Shao, X.; Lv, N.; Liao, J.; Long, J.; Xue, R.; Ai, N.; Xu, D.; Fan, X. Copy number variation is highly correlated with differential gene expression: A pan-cancer study. BMC Med. Genet. 2019, 20, 175. [Google Scholar] [CrossRef] [PubMed]
- Abdel-Fatah, T.; Arora, A.; Gorguc, I.; Abbotts, R.; Beebeejaun, S.; Storr, S.; Mohan, V.; Hawkes, C.; Soomro, I.; Lobo, D.N.; et al. Are DNA repair factors promising biomarkers for personalized therapy in gastric cancer? Antioxid. Redox Signal. 2013, 18, 2392–2398. [Google Scholar] [CrossRef] [PubMed]
- Hagiwara, A.; Kitajima, Y.; Sato, S.; Miyazaki, K. Allelic loss of the DNA repair gene OGG1 against oxidative damage in esophageal squamous cell carcinoma. Oncol. Rep. 2005, 13, 1009–1016. [Google Scholar] [CrossRef] [PubMed]
- Tran, O.T.; Tadesse, S.; Chu, C.; Kidane, D. Overexpression of NEIL3 associated with altered genome and poor survival in selected types of human cancer. Tumour Biol. 2020, 42. [Google Scholar] [CrossRef]
- Carter, R.J.; Parsons, J.L. Base Excision Repair, a Pathway Regulated by Posttranslational Modifications. Mol. Cell. Biol. 2016, 36, 1426–1437. [Google Scholar] [CrossRef]
- Limpose, K.L.; Corbett, A.H.; Doetsch, P.W. BERing the burden of damage: Pathway crosstalk and posttranslational modification of base excision repair proteins regulate DNA damage management. DNA Repair 2017, 56, 51–64. [Google Scholar] [CrossRef]
- Weiser, B.P.; Stivers, J.T.; Cole, P.A. Investigation of N-Terminal Phospho-Regulation of Uracil DNA Glycosylase Using Protein Semisynthesis. Biophys. J. 2017, 113, 393–401. [Google Scholar] [CrossRef]
- Su, S.; Zhang, Y.; Liu, P. Roles of Ubiquitination and SUMOylation in DNA Damage Response. Curr. Issues Mol. Biol. 2020, 35, 59–84. [Google Scholar] [CrossRef]
- Coey, C.T.; Drohat, A.C. Defining the impact of sumoylation on substrate binding and catalysis by thymine DNA glycosylase. Nucleic Acids Res. 2018, 46, 5159–5170. [Google Scholar] [CrossRef]
- Ma, K.W.; Au, S.W.N.; Waye, M.M.Y. Over-expression of SUMO-1 induces the up-regulation of heterogeneous nuclear ribonucleoprotein A2/B1 isoform B1 (hnRNP A2/B1 isoform B1) and uracil DNA glycosylase (UDG) in hepG2 cells. Cell Biochem. Funct. 2009, 27, 228–237. [Google Scholar] [CrossRef]
- Noren Hooten, N.; Kompaniez, K.; Barnes, J.; Lohani, A.; Evans, M.K. Poly(ADP-ribose) polymerase 1 (PARP-1) binds to 8-oxoguanine-DNA glycosylase (OGG1). J. Biol. Chem. 2011, 286, 44679–44690. [Google Scholar] [CrossRef] [PubMed]
- Akbari, M.; Solvang-Garten, K.; Hanssen-Bauer, A.; Lieske, N.V.; Pettersen, H.S.; Pettersen, G.K.; Wilson, D.M., 3rd; Krokan, H.E.; Otterlei, M. Direct interaction between XRCC1 and UNG2 facilitates rapid repair of uracil in DNA by XRCC1 complexes. DNA Repair 2010, 9, 785–795. [Google Scholar] [CrossRef] [PubMed]
- Campalans, A.; Marsin, S.; Nakabeppu, Y.; O’connor, T.R.; Boiteux, S.; Radicella, J.P. XRCC1 interactions with multiple DNA glycosylases: A model for its recruitment to base excision repair. DNA Repair 2005, 4, 826–835. [Google Scholar] [CrossRef] [PubMed]
- Bessho, T. Nucleotide excision repair 3′ endonuclease XPG stimulates the activity of base excision repairenzyme thymine glycol DNA glycosylase. Nucleic Acids Res. 1999, 27, 979–983. [Google Scholar] [CrossRef] [PubMed]
- Klungland, A.; Höss, M.; Gunz, D.; Constantinou, A.; Clarkson, S.G.; Doetsch, P.W.; Bolton, P.H.; Wood, R.D.; Lindahl, T. Base Excision Repair of Oxidative DNA Damage Activated by XPG Protein. Mol. Cell 1999, 3, 33–42. [Google Scholar] [CrossRef]
- Aamann, M.D.; Muftuoglu, M.; Bohr, V.A.; Stevnsner, T. Multiple interaction partners for Cockayne syndrome proteins: Implications for genome and transcriptome maintenance. Mech. Ageing Dev. 2013, 134, 212–224. [Google Scholar] [CrossRef]
- Aamann, M.D.; Hvitby, C.; Popuri, V.; Muftuoglu, M.; Lemminger, L.; Skeby, C.K.; Keijzers, G.; Ahn, B.; Bjørås, M.; Bohr, V.A.; et al. Cockayne Syndrome group B protein stimulates NEIL2 DNA glycosylase activity. Mech. Ageing Dev. 2014, 135, 1–14. [Google Scholar] [CrossRef]
- Kiraly, O.; Gong, G.; Roytman, M.D.; Yamada, Y.; Samson, L.D.; Engelward, B.P. DNA glycosylase activity and cell proliferation are key factors in modulating homologous recombination in vivo. Carcinogenesis 2014, 35, 2495–2502. [Google Scholar] [CrossRef]
- De Souza-Pinto, N.C.; Maynard, S.; Hashiguchi, K.; Hu, J.; Muftuoglu, M.; Bohr, V.A. The recombination protein RAD52 cooperates with the excision repair protein OGG1 for the repair of oxidative lesions in mammalian cells. Mol. Cell. Biol. 2009, 29, 4441–4454. [Google Scholar] [CrossRef]
- Kohno, K.; Izumi, H.; Uchiumi, T.; Ashizuka, M.; Kuwano, M. The pleiotropic functions of the Y-box-binding protein, YB-1. BioEssays 2003, 25, 691–698. [Google Scholar] [CrossRef]
- Lasham, A.; Print, C.G.; Woolley, A.G.; Dunn, S.E.; Braithwaite, A.W. YB-1: Oncoprotein, prognostic marker and therapeutic target? Biochem. J. 2013, 449, 11–23. [Google Scholar] [CrossRef] [PubMed]
- Senarisoy, M.; Barette, C.; Lacroix, F.; De Bonis, S.; Stelter, M.; Hans, F.; Kleman, J.-P.; Fauvarque, M.-O.; Timmins, J. Förster Resonance Energy Transfer Based Biosensor for Targeting the hNTH1–YB1 Interface as a Potential Anticancer Drug Target. ACS Chem. Biol. 2020, 15, 990–1003. [Google Scholar] [CrossRef] [PubMed]
- Bargou, R.C.; Jurchott, K.; Wagener, C.; Bergmann, S.; Metzner, S.; Bommert, K.; Mapara, M.Y.; Winzer, K.J.; Dietel, M.; Dorken, B.; et al. Nuclear localization and increased levels of transcription factor YB-1 in primary human breast cancers are associated with intrinsic MDR1 gene expression. Nat. Med. 1997, 3, 447–450. [Google Scholar] [CrossRef] [PubMed]
- Dey, A.; Robitaille, M.; Remke, M.; Maier, C.; Malhotra, A.; Gregorieff, A.; Wrana, J.L.; Taylor, M.D.; Angers, S.; Kenney, A.M. YB-1 is elevated in medulloblastoma and drives proliferation in Sonic hedgehog-dependent cerebellar granule neuron progenitor cells and medulloblastoma cells. Oncogene 2016, 35, 4256–4268. [Google Scholar] [CrossRef] [PubMed]
- Janz, M.; Harbeck, N.; Dettmar, P.; Berger, U.; Schmidt, A.; Jurchott, K.; Schmitt, M.; Royer, H.D. Y-box factor YB-1 predicts drug resistance and patient outcome in breast cancer independent of clinically relevant tumor biologic factors HER2, uPA and PAI-1. Int. J. Cancer 2002, 97, 278–282. [Google Scholar] [CrossRef]
- Oda, Y.; Ohishi, Y.; Saito, T.; Hinoshita, E.; Uchiumi, T.; Kinukawa, N.; Iwamoto, Y.; Kohno, K.; Kuwano, M.; Tsuneyoshi, M. Nuclear expression of Y-box-binding protein-1 correlates with P-glycoprotein and topoisomerase II alpha expression, and with poor prognosis in synovial sarcoma. J. Pathol. 2003, 199, 251–258. [Google Scholar] [CrossRef]
- Rubinstein, D.B.; Stortchevoi, A.; Boosalis, M.; Ashfaq, R.; Guillaume, T. Overexpression of DNA-binding Protein B Gene Product in Breast Cancer as Detected by in Vitro-generated Combinatorial Human Immunoglobulin Libraries. Cancer Res. 2002, 62, 4985. [Google Scholar]
- Yahata, H.; Kobayashi, H.; Kamura, T.; Amada, S.; Hirakawa, T.; Kohno, K.; Kuwano, M.; Nakano, H. Increased nuclear localization of transcription factor YB-1 in acquired cisplatin-resistant ovarian cancer. J. Cancer Res. Clin. Oncol. 2002, 128, 621–626. [Google Scholar] [CrossRef]
- Guay, D.; Garand, C.; Reddy, S.; Schmutte, C.; Lebel, M. The human endonuclease III enzyme is a relevant target to potentiate cisplatin cytotoxicity in Y-box-binding protein-1 overexpressing tumor cells. Cancer Sci. 2008, 99, 762–769. [Google Scholar] [CrossRef]
- Guay, D.; Evoy, A.A.; Paquet, E.; Garand, C.; Bachvarova, M.; Bachvarov, D.; Lebel, M. The strand separation and nuclease activities associated with YB-1 are dispensable for cisplatin resistance but overexpression of YB-1 in MCF7 and MDA-MB-231 breast tumor cells generates several chemoresistance signatures. Int. J. Biochem. Cell Biol. 2008, 40, 2492–2507. [Google Scholar] [CrossRef]
- Hollenbach, S.; Dhénaut, A.; Eckert, I.; Radicella, J.P.; Epe, B. Overexpression of Ogg1 in mammalian cells: Effects on induced and spontaneous oxidative DNA damage and mutagenesis. Carcinogenesis 1999, 20, 1863–1868. [Google Scholar] [CrossRef] [PubMed]
- Limpose, K.L.; Trego, K.S.; Li, Z.; Leung, S.W.; Sarker, A.H.; Shah, J.A.; Ramalingam, S.S.; Werner, E.M.; Dynan, W.S.; Cooper, P.K.; et al. Overexpression of the base excision repair NTHL1 glycosylase causes genomic instability and early cellular hallmarks of cancer. Nucleic Acids Res. 2018, 46, 4515–4532. [Google Scholar] [CrossRef] [PubMed]
- Kiwerska, K.; Szyfter, K. DNA repair in cancer initiation, progression, and therapy-a double-edged sword. J. Appl. Genet. 2019, 60, 329–334. [Google Scholar] [CrossRef] [PubMed]
- Allan, J.M.; Engelward, B.P.; Dreslin, A.J.; Wyatt, M.D.; Tomasz, M.; Samson, L.D. Mammalian 3-methyladenine DNA glycosylase protects against the toxicity and clastogenicity of certain chemotherapeutic DNA cross-linking agents. Cancer Res. 1998, 58, 3965–3973. [Google Scholar]
- Engelward, B.P.; Dreslin, A.; Christensen, J.; Huszar, D.; Kurahara, C.; Samson, L. Repair-deficient 3-methyladenine DNA glycosylase homozygous mutant mouse cells have increased sensitivity to alkylation-induced chromosome damage and cell killing. EMBO J. 1996, 15, 945–952. [Google Scholar] [CrossRef]
- Rinne, M.; Caldwell, D.; Kelley, M.R. Transient adenoviral N-methylpurine DNA glycosylase overexpression imparts chemotherapeutic sensitivity to human breast cancer cells. Mol. Cancer 2004, 3, 955–967. [Google Scholar]
- Trivedi, R.N.; Almeida, K.H.; Fornsaglio, J.L.; Schamus, S.; Sobol, R.W. The role of base excision repair in the sensitivity and resistance to temozolomide-mediated cell death. Cancer Res. 2005, 65, 6394–6400. [Google Scholar] [CrossRef]
- Klungland, A.; Rosewell, I.; Hollenbach, S.; Larsen, E.; Daly, G.; Epe, B.; Seeberg, E.; Lindahl, T.; Barnes, D.E. Accumulation of premutagenic DNA lesions in mice defective in removal of oxidative base damage. Proc. Natl. Acad. Sci. USA 1999, 96, 13300–13305. [Google Scholar] [CrossRef]
- Yang, N.; Chaudhry, M.A.; Wallace, S.S. Base excision repair by hNTH1 and hOGG1: A two edged sword in the processing of DNA damage in gamma-irradiated human cells. DNA Repair 2006, 5, 43–51. [Google Scholar] [CrossRef]
- Preston, T.J.; Henderson, J.T.; McCallum, G.P.; Wells, P.G. Base excision repair of reactive oxygen species-initiated 7,8-dihydro-8-oxo-2′-deoxyguanosine inhibits the cytotoxicity of platinum anticancer drugs. Mol. Cancer 2009, 8, 2015–2026. [Google Scholar] [CrossRef]
- Takao, M.; Kanno, S.-I.; Kobayashi, K.; Zhang, Q.-M.; Yonei, S.; van der Horst, G.T.J.; Yasui, A. A back-up glycosylase in Nth1 knock-out mice is a functional Nei (endonuclease VIII) homologue. J. Biol. Chem. 2002, 277, 42205–42213. [Google Scholar] [CrossRef] [PubMed]
- Ocampo, M.T.A.; Chaung, W.; Marenstein, D.R.; Chan, M.K.; Altamirano, A.; Basu, A.K.; Boorstein, R.J.; Cunningham, R.P.; Teebor, G.W. Targeted deletion of mNth1 reveals a novel DNA repair enzyme activity. Mol. Cell. Biol. 2002, 22, 6111–6121. [Google Scholar] [CrossRef] [PubMed]
- Chan, M.K.; Ocampo-Hafalla, M.T.; Vartanian, V.; Jaruga, P.; Kirkali, G.; Koenig, K.L.; Brown, S.; Lloyd, R.S.; Dizdaroglu, M.; Teebor, G.W. Targeted deletion of the genes encoding NTH1 and NEIL1 DNA N-glycosylases reveals the existence of novel carcinogenic oxidative damage to DNA. DNA Repair 2009, 8, 786–794. [Google Scholar] [CrossRef] [PubMed]
- Michel, M.; Visnes, T.; Homan, E.J.; Seashore-Ludlow, B.; Hedenström, M.; Wiita, E.; Vallin, K.; Paulin, C.B.J.; Zhang, J.; Wallner, O.; et al. Computational and Experimental Druggability Assessment of Human DNA Glycosylases. ACS Omega 2019, 4, 11642–11656. [Google Scholar] [CrossRef] [PubMed]
- Grin, I.R.; Rieger, R.A.; Zharkov, D.O. Inactivation of NEIL2 DNA glycosylase by pyridoxal phosphate reveals a loop important for substrate binding. Biochem. Biophys. Res. Commun. 2010, 394, 100–105. [Google Scholar] [CrossRef] [PubMed]
- Bliksrud, Y.T.; Ellingsen, A.; Bjørås, M. Fumarylacetoacetate inhibits the initial step of the base excision repair pathway: Implication for the pathogenesis of tyrosinemia type I. J. Inherit. Metab. Dis. 2013, 36, 773–778. [Google Scholar] [CrossRef]
- Biela, A.; Coste, F.; Culard, F.; Guerin, M.; Goffinont, S.; Gasteiger, K.; Cieśla, J.; Winczura, A.; Kazimierczuk, Z.; Gasparutto, D.; et al. Zinc finger oxidation of Fpg/Nei DNA glycosylases by 2-thioxanthine: Biochemical and X-ray structural characterization. Nucleic Acids Res. 2014, 42, 10748–10761. [Google Scholar] [CrossRef]
- Rieux, C.; Goffinont, S.; Coste, F.; Tber, Z.; Cros, J.; Roy, V.; Guérin, M.; Gaudon, V.; Bourg, S.; Biela, A.; et al. Thiopurine Derivative-Induced Fpg/Nei DNA Glycosylase Inhibition: Structural, Dynamic and Functional Insights. Int. J. Mol. Sci. 2020, 21, 2058. [Google Scholar] [CrossRef]
- Wang, H.-C.; Ho, C.-H.; Chou, C.-C.; Ko, T.-P.; Huang, M.-F.; Hsu, K.-C.; Wang, A.H.-J. Using structural-based protein engineering to modulate the differential inhibition effects of SAUGI on human and HSV uracil DNA glycosylase. Nucleic Acids Res. 2016, 44, 4440–4449. [Google Scholar] [CrossRef]
- Chung, S.; Parker, J.B.; Bianchet, M.; Amzel, L.M.; Stivers, J.T. Impact of linker strain and flexibility in the design of a fragment-based inhibitor. Nat. Chem. Biol. 2009, 5, 407–413. [Google Scholar] [CrossRef]
- Jiang, Y.L.; Krosky, D.J.; Seiple, L.; Stivers, J.T. Uracil-Directed Ligand Tethering: An Efficient Strategy for Uracil DNA Glycosylase (UNG) Inhibitor Development. J. Am. Chem. Soc. 2005, 127, 17412–17420. [Google Scholar] [CrossRef] [PubMed]
- Krosky, D.J.; Bianchet, M.A.; Seiple, L.; Chung, S.; Amzel, L.M.; Stivers, J.T. Mimicking damaged DNA with a small molecule inhibitor of human UNG2. Nucleic Acids Res. 2006, 34, 5872–5879. [Google Scholar] [CrossRef] [PubMed]
- Jacobs, A.C.; Calkins, M.J.; Jadhav, A.; Dorjsuren, D.; Maloney, D.; Simeonov, A.; Jaruga, P.; Dizdaroglu, M.; McCullough, A.K.; Stephen Lloyd, R. Inhibition of DNA glycosylases via small molecule purine analogs. PLoS ONE 2013, 8. [Google Scholar] [CrossRef] [PubMed]
- Kennedy, R.D.; Chen, C.C.; Stuckert, P.; Archila, E.M.; De la Vega, M.A.; Moreau, L.A.; Shimamura, A.; D’Andrea, A.D. Fanconi anemia pathway-deficient tumor cells are hypersensitive to inhibition of ataxia telangiectasia mutated. J. Clin. Investig. 2007, 117, 1440–1449. [Google Scholar] [CrossRef] [PubMed]
- Dorjsuren, D.; Kim, D.; Vyjayanti, V.N.; Maloney, D.J.; Jadhav, A.; Wilson, D.M., 3rd; Simeonov, A. Diverse small molecule inhibitors of human apurinic/apyrimidinic endonuclease APE1 identified from a screen of a large public collection. PLoS ONE 2012, 7, e47974. [Google Scholar] [CrossRef] [PubMed]
- Donley, N.; Jaruga, P.; Coskun, E.; Dizdaroglu, M.; McCullough, A.K.; Lloyd, R.S. Small Molecule Inhibitors of 8-Oxoguanine DNA Glycosylase-1 (OGG1). ACS Chem. Biol. 2015, 10, 2334–2343. [Google Scholar] [CrossRef]
- Tahara, Y.K.; Auld, D.; Ji, D.; Beharry, A.A.; Kietrys, A.M.; Wilson, D.L.; Jimenez, M.; King, D.; Nguyen, Z.; Kool, E.T. Potent and Selective Inhibitors of 8-Oxoguanine DNA Glycosylase. J. Am. Chem. Soc. 2018, 140, 2105–2114. [Google Scholar] [CrossRef]
- Dalhus, B.; Forsbring, M.; Helle, I.H.; Vik, E.S.; Forstrøm, R.J.; Backe, P.H.; Alseth, I.; Bjørås, M. Separation-of-function mutants unravel the dual-reaction mode of human 8-oxoguanine DNA glycosylase. Structure 2011, 19, 117–127. [Google Scholar] [CrossRef]
- Mancuso, P.; Tricarico, R.; Bhattacharjee, V.; Cosentino, L.; Kadariya, Y.; Jelinek, J.; Nicolas, E.; Einarson, M.; Beeharry, N.; Devarajan, K.; et al. Thymine DNA glycosylase as a novel target for melanoma. Oncogene 2019, 38, 3710–3728. [Google Scholar] [CrossRef]
- Visnes, T.; Cázares-Körner, A.; Hao, W.; Wallner, O.; Masuyer, G.; Loseva, O.; Mortusewicz, O.; Wiita, E.; Sarno, A.; Manoilov, A.; et al. Small-molecule inhibitor of OGG1 suppresses proinflammatory gene expression and inflammation. Science 2018, 362, 834–839. [Google Scholar] [CrossRef]
- Shigdel, U.K.; Ovchinnikov, V.; Lee, S.-J.; Shih, J.A.; Karplus, M.; Nam, K.; Verdine, G.L. The trajectory of intrahelical lesion recognition and extrusion by the human 8-oxoguanine DNA glycosylase. Nat. Commun. 2020, 11, 4437. [Google Scholar] [CrossRef] [PubMed]
- Tahara, Y.-K.; Kietrys, A.M.; Hebenbrock, M.; Lee, Y.; Wilson, D.L.; Kool, E.T. Dual Inhibitors of 8-Oxoguanine Surveillance by OGG1 and NUDT1. ACS Chem. Biol. 2019, 14, 2606–2615. [Google Scholar] [CrossRef] [PubMed]
- Edwards, S.K.; Ono, T.; Wang, S.; Jiang, W.; Franzini, R.M.; Jung, J.W.; Chan, K.M.; Kool, E.T. In Vitro Fluorogenic Real-Time Assay of the Repair of Oxidative DNA Damage. ChemBioChem 2015, 16, 1637–1646. [Google Scholar] [CrossRef] [PubMed]
- Sandin, P.; Börjesson, K.; Li, H.; Mårtensson, J.; Brown, T.; Wilhelmsson, L.M.; Albinsson, B. Characterization and use of an unprecedentedly bright and structurally non-perturbing fluorescent DNA base analogue. Nucleic Acids Res. 2008, 36, 157–167. [Google Scholar] [CrossRef] [PubMed]
- Nakabeppu, Y.; Ohta, E.; Abolhassani, N. MTH1 as a nucleotide pool sanitizing enzyme: Friend or foe? Free Radic. Biol. Med. 2017, 107, 151–158. [Google Scholar] [CrossRef] [PubMed]
- Kumar, A.; Kawamura, T.; Kawatani, M.; Osada, H.; Zhang, K.Y.J. Identification and structure-activity relationship of purine derivatives as novel MTH1 inhibitors. Chem. Biol. Drug Des. 2017, 89, 862–869. [Google Scholar] [CrossRef] [PubMed]
- Ji, D.; Beharry, A.A.; Ford, J.M.; Kool, E.T. A Chimeric ATP-Linked Nucleotide Enables Luminescence Signaling of Damage Surveillance by MTH1, a Cancer Target. J. Am. Chem Soc. 2016, 138, 9005–9008. [Google Scholar] [CrossRef] [PubMed]
- Eastman, A. Improving anticancer drug development begins with cell culture: Misinformation perpetrated by the misuse of cytotoxicity assays. Oncotarget 2017, 8, 8854–8866. [Google Scholar] [CrossRef]
- Gao, H.; Korn, J.M.; Ferretti, S.; Monahan, J.E.; Wang, Y.; Singh, M.; Zhang, C.; Schnell, C.; Yang, G.; Zhang, Y.; et al. High-throughput screening using patient-derived tumor xenografts to predict clinical trial drug response. Nat. Med. 2015, 21, 1318–1325. [Google Scholar] [CrossRef]
- Lagies, S.; Schlimpert, M.; Neumann, S.; Wäldin, A.; Kammerer, B.; Borner, C.; Peintner, L. Cells grown in three-dimensional spheroids mirror in vivo metabolic response of epithelial cells. Commun. Biol. 2020, 3, 246. [Google Scholar] [CrossRef]
- Rossi, G.; Manfrin, A.; Lutolf, M.P. Progress and potential in organoid research. Nat. Rev. Genet. 2018, 19, 671–687. [Google Scholar] [CrossRef] [PubMed]
- Drost, J.; van Boxtel, R.; Blokzijl, F.; Mizutani, T.; Sasaki, N.; Sasselli, V.; de Ligt, J.; Behjati, S.; Grolleman, J.E.; van Wezel, T.; et al. Use of CRISPR-modified human stem cell organoids to study the origin of mutational signatures in cancer. Science 2017, 358, 234–238. [Google Scholar] [CrossRef] [PubMed]
- Sontheimer-Phelps, A.; Hassell, B.A.; Ingber, D.E. Modelling cancer in microfluidic human organs-on-chips. Nat. Rev. Cancer 2019, 19, 65–81. [Google Scholar] [CrossRef] [PubMed]
| Structural Motif Superfamily | Name of the Glycosylase | Mono/Bifunc | Major Substrate Specificity | Reported PTMs | Ref. | |
|---|---|---|---|---|---|---|
| Alpha-beta fold (UDG) superfamily | Uracil-N glycosylase1 | UNG1 | Mono | U in any context, in ss and dsDNA | [69,70,71] | |
| Uracil-N glycosylase2 | UNG2 | Mono | Similar to UNG1 | Phospho Ubiquit Acetyl | [71,72,73,74,75,76,77,78,79] | |
| Single-strand-specific monofunctional uracil DNA glycosylase 1 | SMUG1 | Mono | ssU, U:G, U:A, 5-hydroxymethylU, in ss and dsDNA | Ubiquit | [72,80] | |
| Thymine DNA glycosylase | TDG | Mono | U:G, T:G, oxidized/deaminated 5-methylC:G, in dsDNA | Phospho Ubiquit Sumoyl Acetyl | [81,82,83,84,85,86,87,88,89] | |
| Helix-hairpin-helix (HhH) superfamily | Methyl-binding domain glycosylase 4 | MBD4 | Mono | U:G and T:G, 5-hydroxymethylU in CpG islands, in dsDNA | Phospho | [90,91,92] |
| 8-OxoG DNA glycosylase 1 | OGG1 | Bifunc | Oxidized purines (8-oxoG:C, FapyG:C), in dsDNA | Phospho Nitrosyl Acetyl Ubiquit | [93,94,95,96,97,98,99] | |
| MutY homolog DNA glycosylase | MUTYH | Mono | A opposite 8-oxo-G/C/G, in dsDNA | Phospho Ubiquit | [100,101,102,103,104] | |
| Endonuclease III-like 1 | NTH1 | Bifunc | Oxidized pyrimidines (Tg, 5-hydroxyC, 5-hydroxyU), FapyG, FapyA, in dsDNA | Ubiquit | [105,106,107] | |
| 3-methyl-purine glycosylase (MPG) superfamily | 3-methyl-purine glycosylase | MPG | Mono | 3-methylA, 1-methyl, 7-methylG, εA, ethenoA hypoxanthine, in ss and dsDNA | Phospho Acetyl | [108,109,110,111] |
| Hairpin-2- turn-hairpin (NEIL) superfamily | Endonuclease VIII-like glycosylase 1 | NEIL1 | Bifunc | Oxidized pyrimidines (Tg, 5-hydroxyU, 5,6-dihydroU, hydantoins Gh and Sp), FapyG, FapyA, in ss and dsDNA | [57,67,112,113] | |
| Endonuclease VIII-like glycosylase 2 | NEIL2 | Bifunc | Similar to NEIL1 in bubbles and loops | Acetyl | [114,115] | |
| Endonuclease VIII-like glycosylase 3 | NEIL3 | Bifunc | Similar to NEIL1 (FapyG, FapyA, Sp and Gh) in ssDNA | [41,116,117,118] | ||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hans, F.; Senarisoy, M.; Bhaskar Naidu, C.; Timmins, J. Focus on DNA Glycosylases—A Set of Tightly Regulated Enzymes with a High Potential as Anticancer Drug Targets. Int. J. Mol. Sci. 2020, 21, 9226. https://doi.org/10.3390/ijms21239226
Hans F, Senarisoy M, Bhaskar Naidu C, Timmins J. Focus on DNA Glycosylases—A Set of Tightly Regulated Enzymes with a High Potential as Anticancer Drug Targets. International Journal of Molecular Sciences. 2020; 21(23):9226. https://doi.org/10.3390/ijms21239226
Chicago/Turabian StyleHans, Fabienne, Muge Senarisoy, Chandini Bhaskar Naidu, and Joanna Timmins. 2020. "Focus on DNA Glycosylases—A Set of Tightly Regulated Enzymes with a High Potential as Anticancer Drug Targets" International Journal of Molecular Sciences 21, no. 23: 9226. https://doi.org/10.3390/ijms21239226
APA StyleHans, F., Senarisoy, M., Bhaskar Naidu, C., & Timmins, J. (2020). Focus on DNA Glycosylases—A Set of Tightly Regulated Enzymes with a High Potential as Anticancer Drug Targets. International Journal of Molecular Sciences, 21(23), 9226. https://doi.org/10.3390/ijms21239226





