Lipid A from Oligotropha carboxidovorans Lipopolysaccharide That Contains Two Galacturonic Acid Residues in the Backbone and Malic Acid A Tertiary Acyl Substituent
Abstract
1. Introduction
2. Results
2.1. Isolation and Chemical Analysis of O. carboxidovorans Lipid A
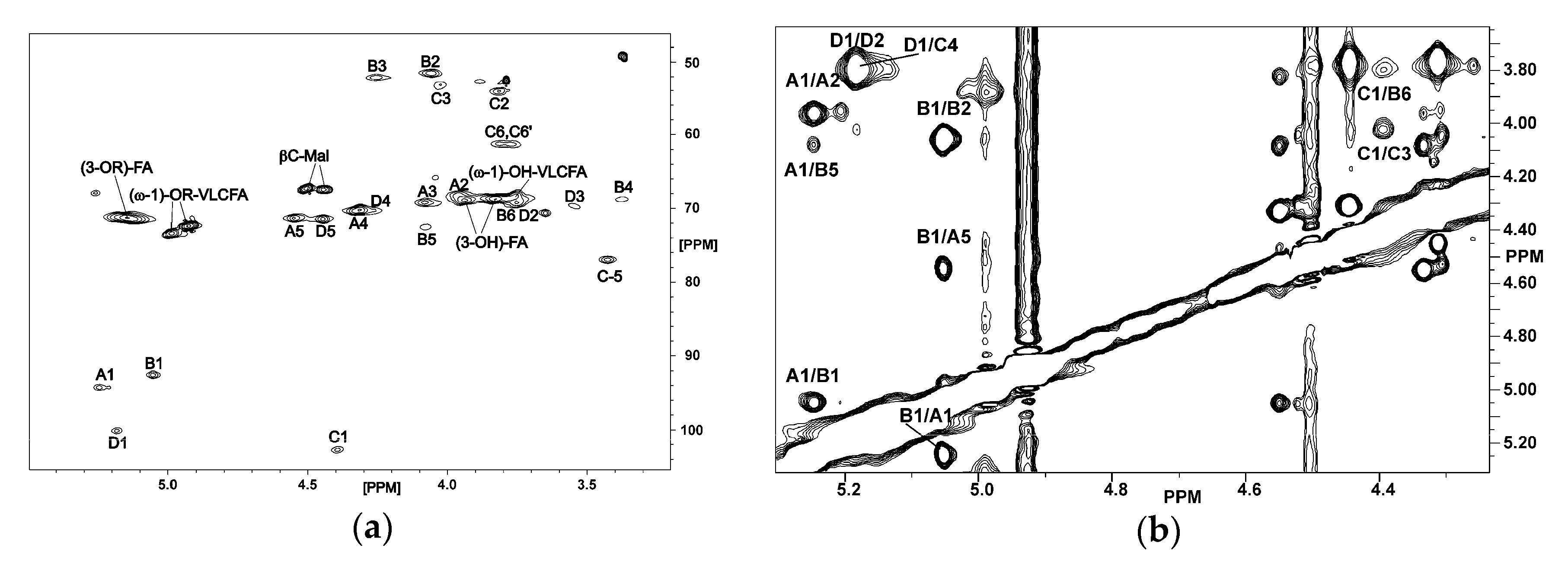
2.2. NMR Spectroscopy of O. carboxidovorans Lipid A
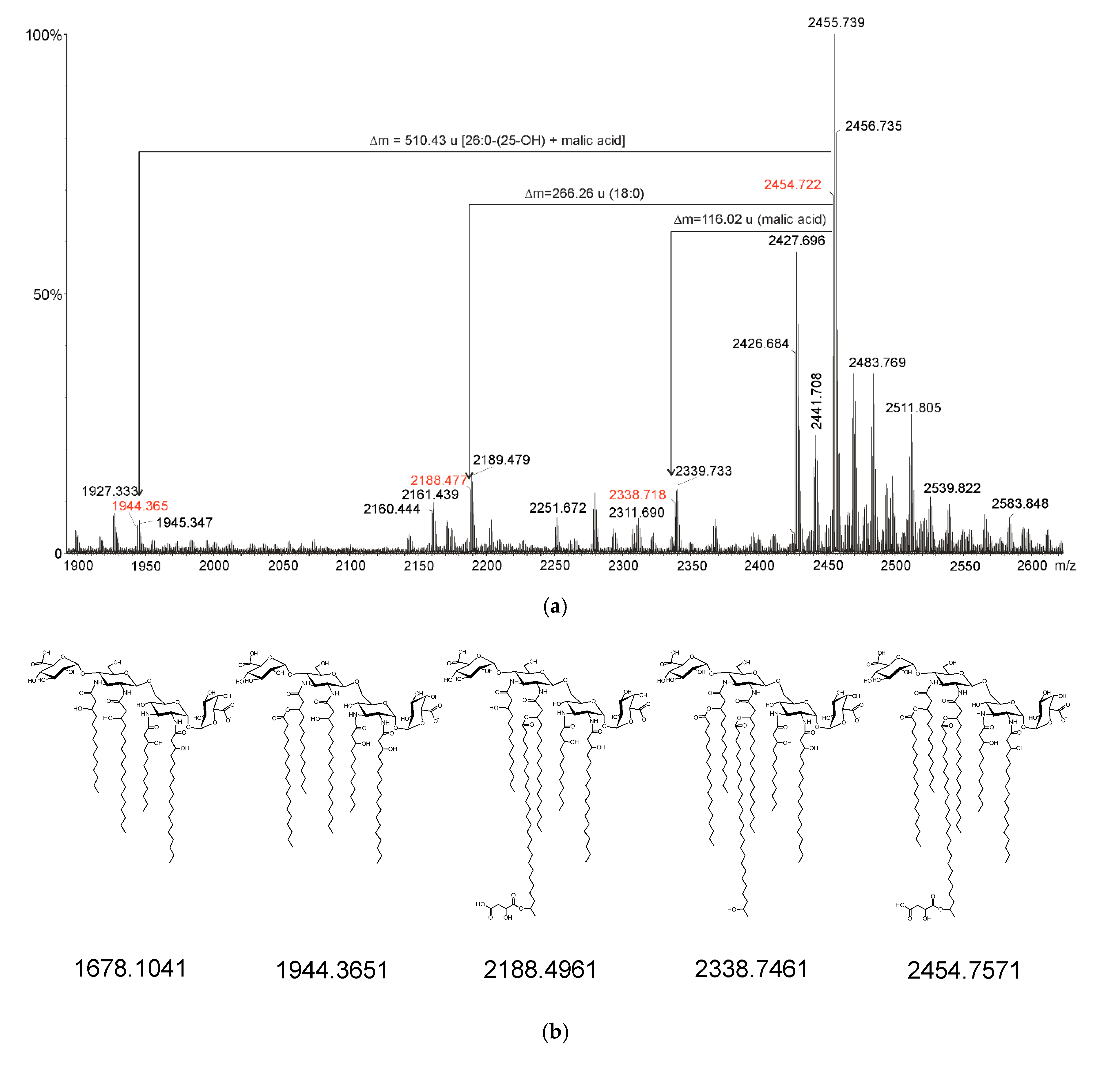
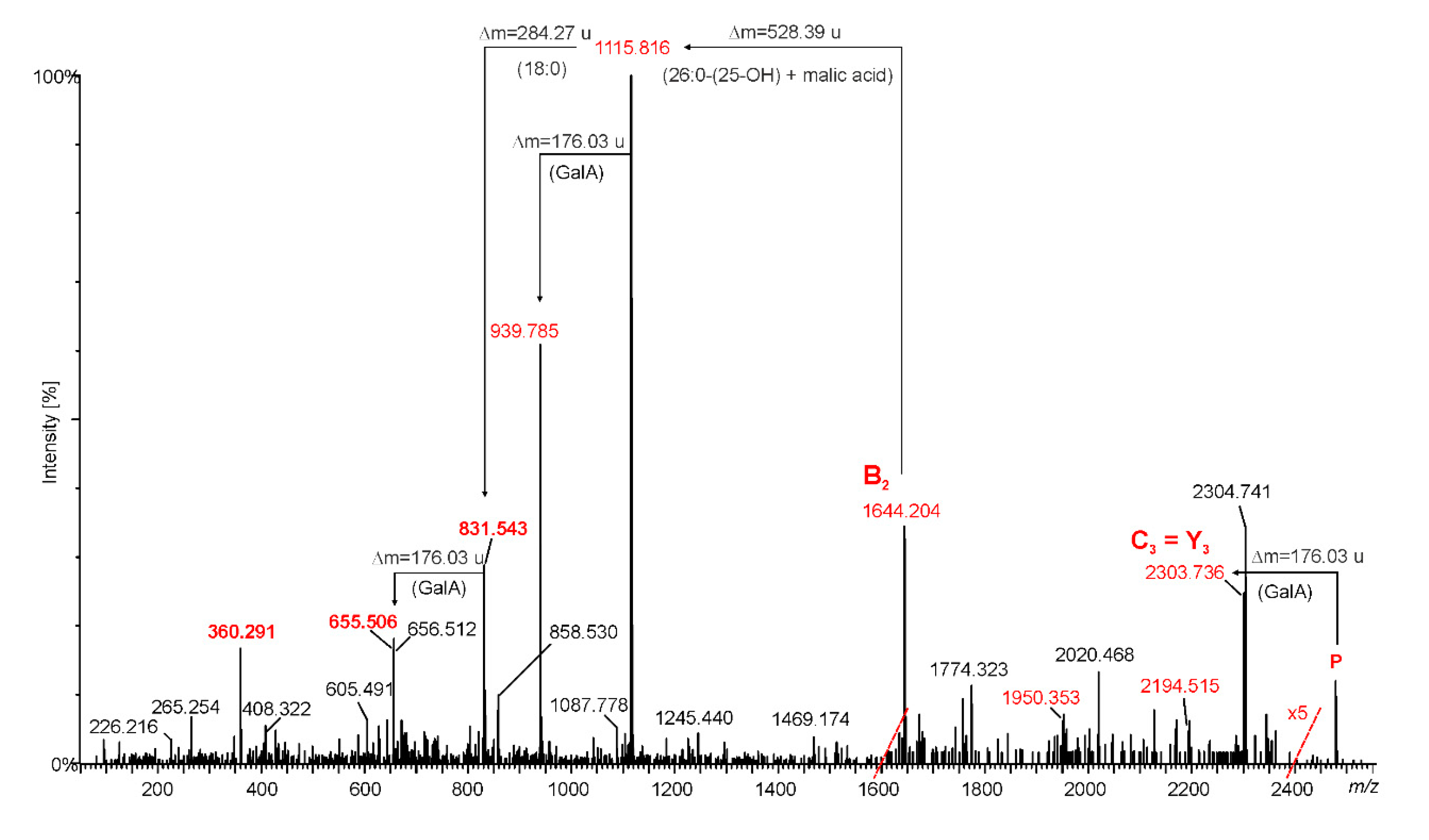
2.3. Mass Spectrometry of O. carboxidovorans Lipid A
2.4. Genomic Studies
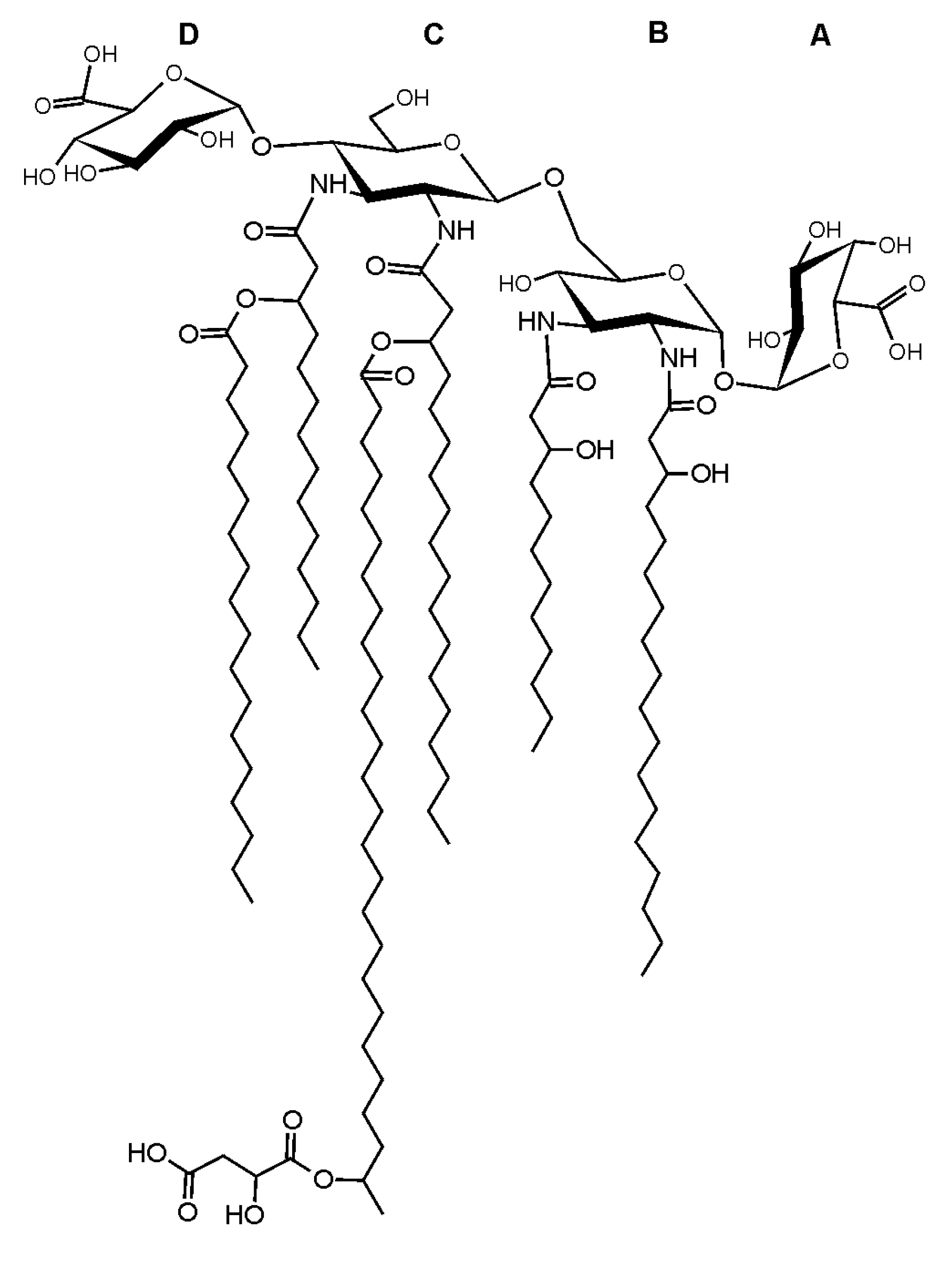
3. Discussion
4. Materials and Methods
4.1. Bacterial Strain and Culture Condition
4.2. LPS and Lipid A Isolation and Purification
4.3. Sugar and Fatty Acid Analysis
4.4. NMR Spectroscopy
4.5. Mass Spectrometry
4.6. Bioinformatics Analysis
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| EDTA | ethylenediaminetetraacetic acid |
| GlcpN3N | 2,3-diamino-2,3-dideoxyglucose |
| GalpA | galacturonic acid |
| MALDI-TOF | matrix assisted laser desorption/ionization—time of flight |
| NC | nitrocellulose |
| std | internal standard |
| Sug | sugar residue |
| THAP | 2′,4′,6′-trihydroxyacetophenone |
| TMSi | trimethylsilylating reagent |
| VLCFA | very long chain fatty acid |
References
- Meyer, O.; Schlegel, H.G. Reisolation of the carbon monoxide utilizing hydrogen bacterium Pseudomonas carboxydovorans (Kistner) comb. nov. Arch. Microbiol. 1978, 118, 35–43. [Google Scholar] [CrossRef] [PubMed]
- Meyer, O.; Stackebrandt, E.; Auling, G. Reclassification of ubiquinone Q-10 containing carbooxidotrophic bacteria: Transfer of “[Pseudomonas] carboxydovorans” OM5T to Oligotropha, gen. nov., as Oligotropha carboxidovorans, comb. nov., transfer of “[Alcaligenes] carboxydus” DSM 1086T to Carbophilus, gen. nov., as Carbophilus carboxidus, comb. nov., transfer of “[Pseudomonas] compransoris” DSM 1231T to Zavarzinia, gen. nov., as Zavarzinia compransoris, comb. nov., amended description of the new genera. Syst. Appl. Microbiol. 1993, 16, 390–395. [Google Scholar]
- Meyer, O.; Schlegel, H.G. Biology of aerobic carbon monoxide-oxidizing bacteria. Annu. Rev. Microbiol. 1983, 37, 277–310. [Google Scholar] [CrossRef] [PubMed]
- Fuhrmann, S.; Ferner, M.; Jefke, T.; Henne, A.; Gottschalk, G.; Meyer, O. Complete nucleotide sequence of the circular megaplasmid pHCG3 of Oligotropha carboxidovorans: Function in the chemolithoautotrophic utilization of CO, H(2) and CO(2). Gene 2003, 322, 67–75. [Google Scholar] [CrossRef] [PubMed]
- Hille, R.; Dingwall, S.; Wilcoxen, J. The aerobic CO dehydrogenase from Oligotropha carboxidovorans. J. Biol. Inorg. Chem. 2015, 20, 243–251. [Google Scholar] [CrossRef]
- Paul, D.; Bridges, S.S.; Burgess, S.C.; Dandass, Y.S.; Lawrence, M.L. Complete genome and comparative analysis of the chemolithoautotrophic bacterium Oligotropha carboxidovorans OM5. BMC Genom. 2010, 11, 511. [Google Scholar] [CrossRef][Green Version]
- Paul, D.; Kumar, R.; Nanduri, B.; French, T.; Pendarvis, K.; Brown, A.; Lawrence, M.L.; Burgess, S.C. Proteome and membrane fatty acid analyses on Oligotropha carboxidovorans OM5 grown under chemolithoautotrophic and heterotrophic conditions. PLoS ONE 2011, 6, e17111. [Google Scholar] [CrossRef][Green Version]
- Malik, A.; Kakii, K. Intergeneric coaggregations among Oligotropha carboxidovorans and Acinetobacter species present in activated sludge. FEMS Microbiol. Lett. 2003, 224, 23–28. [Google Scholar] [CrossRef]
- Choma, A.; Komaniecka, I.; Żebracki, K. Structure, biosynthesis and function of unusual lipids A from nodule-inducing and N2-fixing bacteria. Biochim. Biophys. Acta 2017, 1862, 196–209. [Google Scholar] [CrossRef]
- Choma, A.; Sowinski, P. Characterization of Mesorhizobium huakuii lipid A containing both d-galacturonic acid and phosphate residues. Eur J Biochem. 2004, 271, 1310–1322. [Google Scholar] [CrossRef]
- Brown, D.B.; Muszynski, A.; Carlson, R.W. Elucidation of a novel lipid A α-(1,1)-GalA transferase gene (rgtF) from Mesorhizobium loti: Heterologous expression of rgtF causes Rhizobium etli to synthesize lipid A with α-(1,1)-GalA. Glycobiology 2013, 23, 546–558. [Google Scholar] [CrossRef] [PubMed]
- Komaniecka, I.; Choma, A.; Lindner, B.; Holst, O. The structure of a novel neutral lipid A from the lipopolysaccharide of Bradyrhizobium elkanii containing three mannoses units in the backbone. Chem. Eur. J. 2010, 16, 2922–2929. [Google Scholar] [CrossRef]
- Komaniecka, I.; Choma, A.; Mazur, A.; Duda, K.A.; Lindner, B.; Schwudke, D.; Holst, O. Occurrence of an unusual hopanoid-containing lipid A among lipopolysaccharides from Bradyrhizobium species. J. Biol. Chem. 2014, 289, 35644–35655. [Google Scholar] [CrossRef] [PubMed]
- Silipo, A.; Vitiello, G.; Gully, D.; Sturiale, L.; Chaintreuil, C.; Fardoux, J.; Gargani, D.; Lee, H.I.; Kulkarni, G.; Busset, N.; et al. Covalently linked hopanoid-lipid A improves outer membrane resistance of a Bradyrhizobium symbiont of legumes. Nat. Commun. 2014, 5, 5106. [Google Scholar] [CrossRef] [PubMed]
- Mayer, H.; Krauss, J.H.; Urbanik-Sypniewska, T.; Puvanesarajah, V.; Stacey, G.; Auling, G. Lipid A with 2,3-diamino-2,3-dideoxy-glucose in lipopolysaccharides from slow-growing members of Rhizobiaceae and from “Pseudomonas carboxydovorans”. Arch. Microbiol. 1989, 151, 111–116. [Google Scholar] [CrossRef]
- Volland, S.; Rachinger, M.; Strittmatter, A.; Daniel, R.; Gottschalk, G.; Meyer, O. Complete genome sequences of the chemolithoautotrophic Oligotropha carboxidovorans strains OM4 and OM5. O. J. Bacteriol. 2011, 193, 5043. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Westphal, O.; Jann, K. Bacterial lipopolysaccharide. Extraction with phenol-water and further application of the procedure. In Methods in Carbohydrate; Whistler, R.L., Ed.; Academic Press: Cambridge, MA, USA, 1965; Volume 5, pp. 83–91. [Google Scholar]
- Johnson, K.G.; Perry, M.B. Improved techniques for the preparation of bacterial lipopolysaccharides. Can. J. Microbiol. 1976, 22, 29–34. [Google Scholar] [CrossRef]
- Domon, B.; Costello, C. A systematic nomenclature for carbohydrate fragmentations in FAB-MS/MS spectra of glycoconjugates. Glycoconjugate J. 1988, 5, 397–409. [Google Scholar] [CrossRef]
- Plötz, B.M.; Lindner, B.; Stetter, K.O.; Holst, O. Characterization of a novel lipid A containing D-galacturonic acid that replaces phosphate residues. The structure of the lipid a of the lipopolysaccharide from the hyperthermophilic bacterium Aquifex pyrophilus. J. Biol. Chem. 2000, 275, 11222–11228. [Google Scholar]
- Smit, J.; Kaltashov, J.A.; Cotter, R.J.; Vinogradov, E.; Perry, M.B.; Haider, H.; Qureshi, N. Structure of a novel lipid A obtained from the lipopolysaccharide of Caulobacter crescentus. Innate Immun. 2008, 14, 25–37. [Google Scholar] [CrossRef]
- Karbarz, M.J.; Six, D.A.; Raetz, C.R. Purification and characterization of the lipid A 1-phosphatase LpxE of Rhizobium leguminosarum. J. Biol. Chem. 2009, 284, 414–425. [Google Scholar] [CrossRef] [PubMed]
- Choma, A.; Komaniecka, I. Characterization of a novel lipid A structure isolated from Azospirillum lipoferum lipopolysaccharide. Carbohydr. Res. 2008, 343, 799–804. [Google Scholar] [CrossRef] [PubMed]
- Zamlynska, K.; Komaniecka, I.; Zebracki, K.; Mazur, A.; Sroka-Bartnicka, A.; Choma, A. Studies on lipid A isolated from Phyllobacterium trifolii PETP02T lipopolysaccharide. Ant. Leeuwen. 2017, 110, 1413–1433. [Google Scholar] [CrossRef]
- Jahnke, L.L.; Eder, W.; Huber, R.; Hope, J.M.; Hinrichs, K.-U.; Hayes, J.M.; Des Marais, D.J.; Cady, S.L.; Summons, R.E. Signature lipids and stable carbon isotope analyses of Octopus Spring hyperthermophilic communities compared with those of Aquificales representatives. Appl. Environ. Microbiol. 2001, 67, 5179–5189. [Google Scholar] [CrossRef] [PubMed]
- Letts, V.; Shaw, P.; Shapiro, L.; Henry, S. Synthesis and utilization of fatty acids by wild-type and fatty acid auxotrophs of Caulobacter crescentus. J. Bacteriol. 1982, 151, 1269–1278. [Google Scholar] [CrossRef]
- Bhat, U.R.; Mayer, H.; Yokota, A.; Hollingsworth, R.I.; Carlson, R.W. Occurrence of lipid A variants with 27-hydroxyoctacosanoic acid in lipopolysaccharides from members of the family Rhizobiaceae. J. Syst. Bacteriol. 1991, 173, 2155–2159. [Google Scholar] [CrossRef]
- Raetz, C.R.H.; Whitfield, C. Lipopolisaccharide endotoxins. Annu. Rev. Biochem. 2002, 71, 635–700. [Google Scholar] [CrossRef]
- Komaniecka, I.; Choma, A.; Zamlynska, K.; Sroka-Bartnicka, A.; Sowinski, P. Structure of O-specific polysaccharide of Oligotropha carboxidovorans OM5—A wastewater bacterium. Carbohydr. Res. 2017, 439, 30–34. [Google Scholar] [CrossRef]
- Que, N.L.S.; Lin, S.H.; Cotter, R.J.; Raetz, C.R.H. Purification and mass spectrometry of six lipid A species from the bacterial endosymbiont Rhizobium etli—Demonstration of a conserved distal unit and a variable proximal portion. J. Biol. Chem. 2000, 275, 28006–28016. [Google Scholar] [CrossRef]
- Choma, A.; Komaniecka, I.; Turska-Szewczuk, A.; Danikiewicz, W.; Spolnik, G. Structure of lipid A from a stem-nodulating bacterium Azorhizobium caulinodans. Carbohydr. Res. 2012, 352, 126–136. [Google Scholar] [CrossRef]
- Sawardeker, J.S.; Sloneker, J.H.; Jeanes, A.R. Quantitative determination of monosaccharides as their alditol acetates by gas liquid chromatography. Anal. Chem. 1965, 37, 1602–1604. [Google Scholar] [CrossRef]
- Silipo, A.; Molinaro, A.; Sturiale, L.; Dow, J.M.; Erbs, G.; Lanzetta, R.; Newman, M.A.; Parrilli, M. The elicitation of plant innate immunity by lipooligosaccharide of Xanthomonas campestris. J. Biol. Chem. 2005, 280, 33660–33668. [Google Scholar] [CrossRef] [PubMed]

| Component | Amount [µg/mg Lipid A] |
|---|---|
| 12:0(3-OH) | 39.7 |
| 14:0(3-OH) | 24.5 |
| 18:0(3-OH) | 95.7 |
| 16:0 | 5.7 |
| 18:0 | 93.7 |
| 18:1 | 4.6 |
| 22:0 | 33.2 |
| 24:0 | 6.5 |
| 26:0(25-OH) | 112.5 |
| 28:0(27-OH) | 18.2 |
| Spin System | 1J(C-1,H-1) [Hz] | H-1 C-1 | H-2 C-2 | H-3 C-3 | H-4 C-4 | H-5 C-5 | H-6; H-6’ C-6 |
|---|---|---|---|---|---|---|---|
| A, α-d-GalpA-(1→ | 173.8 | 5.248 | 3.96 | 4.08 | 4.333 | 4.55 | - |
| 94.24 | 68.62 | 69.13 | 70.34 | 71.3 | 171.5 | ||
| B, →6)-α-d-GlcpN3N-(1→ | 171.8 | 5.053 | 4.059 | 4.255 | 3.375 | 4.078 | 3.710; 3.710 |
| 92.57 | 51.65 | 52.23 | 68.73 | 72.48 | 68.7 | ||
| C, →4)-β-d-GlcpN3N-(1→ | 161.1 | 4.395 | 3.816 | 4.027 | 3.825 | 3.427 | 3.773; 3.812 |
| 102.69 | 54.1 | 53.41 | 75.22 | 76.94 | 61.24 | ||
| D, α-d-GalpA-(1→ | 170.6 | 5.189 | 3.796 | 3.773 | 4.316 | 4.446 | - |
| 100.02 | 68.68 | 68.74 | 70.15 | 71.46 | 170.5 |
| Fatty Acid | Group | 1H | 13C |
|---|---|---|---|
| Olefinic protons/carbons (double bond) | -CH2-HC=CH- | 5.350 | 130.08 |
| -CH2-HC=CH- | 2.016 | 27.49 | |
| Ist –(3-OR’)-FA | α1/α2 | 2.620/2.504 | 41 |
| β | 5.145 | 71.29 | |
| γ | 1.588 | 34.4 | |
| R-CONH-Sug | 171.51 | ||
| R-COO- | 174.53 | ||
| ω | 1.59 | 19.8 | |
| IInd –(3-OR”)-FA | α1/α2 | 2.614/2.401 | 40.9 |
| β | 5.262 | 68 | |
| γ | ND | ND | |
| R-CONH-Sug | 169.72 | ||
| R-COO- | ND | ||
| Ist –[(ω-1)-OR]-VLCFA | ω | 1.222 | 19.8 |
| ω-1 | 4.996 | 73.39 | |
| ω-2 | 1.600/1.486 | 36 | |
| ω-3 | 1.41 | 25.5 | |
| ω-4 and next CH2 groups | 1.59 | 29.8 | |
| R-(COO-) from malic acid | 173.3 | ||
| IInd –[(ω-1)-OR]-VLCFA | ω | 1.222 | 19.8 |
| ω-1 | 4.929 | 73.36 | |
| ω-2 | 1.600/1.488 | 36 | |
| ω-3 | 1.474 | 25.5 | |
| ω-4 and next CH2 groups | 1.59 | 29.8 | |
| R-(COO-) from malic acid | 170.84 | ||
| Ist –(3-OH)-FA with unsubstituted OH group | α1/α2 | 2.339/2.252 | 43.9 |
| β | 3.942 | 68.8 | |
| γ | 1.435 | 37.5 | |
| δ | 1.399 | 25.8 | |
| CH2 groups from acyl chain | 1.266 | 29.9 | |
| IInd – (3-OH)-FA with unsubstituted OH group | α1/α2 | 2.307/2.201 | 43.05 |
| β | 3.833 | 68.8 | |
| γ | 1.435 | 37.6 | |
| δ | 1.399 | 25.7 | |
| CH2 groups from acyl chain | 1.266 | 29.9 | |
| [(ω-1)-OH] VLCFA with unsubstituted OH group | ω-1 | 3.756 | 68.4 |
| ω-2 | 1.387/1.478 | 39.34 | |
| ω-3 | 1.28 | 26.05 | |
| ω-4 and next CH2 groups | 1.59 | 29.94 | |
| Ist Malic acid | α1/α2 | 2.860/2.717 | 39.41 |
| β | 4.505 | 67.16 | |
| COO- | 170.84 | ||
| R-COO- | 175.36 | ||
| IInd Malic acid | α1/α2 | 2.860/2.717 | 39.24 |
| β | 4.446 | 67.41 | |
| COO- | 170.58 | ||
| R-COO- | 173.3 |
| Gene Name (Putative Protein Function) | lpxE (Lipid A 1-Phosphatase) | lpxF (Lipid A 4’-Phosphatase) | rgtD (4′-GalA Transferase) | rgtF (α-(1,1)-GalA Transferase) | rgtE (Bactoprenyl/Phosphate GalA Transferase) |
|---|---|---|---|---|---|
| Reference sequence of: R. leguminosarum 3841 and/or M. loti MAFF303099 | RL_RS24225 (RL4708) | RL_RS08140 (RL1570) | RL_RS03600 (RL0684) | MAFF_RS01135 2 (mlr0011) | RL_RS07630 (RL1470) |
| O. carboxidovorans OM5 % Identity 1 | OCA5_c19740 31.97 | OCA5_cRS09935 30.96 | OCA5_RS09935 28.81 | OCA5_c19750 39.28 | OCA5_c19730 34.69 |
| Aquifex aeolicus VF5 % Identity 1 | ND | ND | aq_765 29.31 | aq_1695a 22.73 | aq_1899 39.82 |
| Caulobacter crescentus (vibroides) CB15 % Identity 1 | CC_3019 29.84 | ND | CC_0209 31.66 | CC_0468 33.27 | CC_0469 33.33 |
| Putative function ascribed on BLAST sequence similarity | phosphatase PAP2 family protein | phosphoesterase PA-phosphatase related | dolichyl-phosphate-mannose-protein mannosyltransferase | glycosyl transferase | glycosyl transferase, family 2 |
| Gene Name (Putative Protein Function) | gnnA (NAD-Dependent Oxidation of GlcN 3-OH of UDP-GlcNAc) | gnnB (Transamination to Form UDP-GlcNAc3N) | acpXL (Acyl Carrier Protein) | fabZXL (3R)-Hydroxyacyl-ACP Dehydratase | fabF2XL (Beta-Ketoacyl-ACP Synthase) | fabF1XL (Beta-Ketoacyl-ACP Synthase) | adh2XL (Zinc-Binding Dehydrogenase) | lpxXL (msbB) (VLCFA Acyltransferase to C2’ Position of Lipid A) |
|---|---|---|---|---|---|---|---|---|
| Reference sequence of M. loti MAFF303099 and/or S. meliloti 1021 (old description) | MAFF_RS12135 (mll2825I) | MAFF_RS30880 (mll7579) | MAFF_RS05885 (mlr1174) | SM_RS09960 (SMc04277) 2 | MAFF_RS05890 (mlr1176) | MAFF_RS05895 (mlr1177) | MAFF_RS05900 (mlr1178) | MAFF_RS05905 (mlr1179) |
| Oligotropha carboxydovorans OM5 (DSM1227) % Identity 1 | OCA5_c26370 32.37 | OCA5_c26360 55.41 | acpXL 78.49 | fabA 50.34 | OCA5_c15570 43.59 | fabF1 58.12 | OCA5_c15550 62.24 | OCA5_c15540 38.36 |
| Caulobacter crescentus CB15 % Identity 1 | CC_1225 25.29 | CC_3611 53.12 | CC_1677 39.29 | CCNA_01989 36.59 | CC_1678 23.76 | CC_1678 31.54 | CC_1569 30.41 | CC_2250 32.26 |
| CC-3719 30 | ||||||||
| Putative function ascribed on BLAST sequence similarity | oxidoreductase domain protein | DegT/DnrJ/EryC1/StrS aminotransferase | acyl carrier protein AcpXL | beta-hydroxyacylACP dehydratase | beta-ketoacyl synthase | 3-oxoacyl-[ACP] synthase 2 | alcohol dehydrogenase | VLCFA acyltransferase to C2’ position of lipid A |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Choma, A.; Zamłyńska, K.; Mazur, A.; Pastuszka, A.; Kaczyński, Z.; Komaniecka, I. Lipid A from Oligotropha carboxidovorans Lipopolysaccharide That Contains Two Galacturonic Acid Residues in the Backbone and Malic Acid A Tertiary Acyl Substituent. Int. J. Mol. Sci. 2020, 21, 7991. https://doi.org/10.3390/ijms21217991
Choma A, Zamłyńska K, Mazur A, Pastuszka A, Kaczyński Z, Komaniecka I. Lipid A from Oligotropha carboxidovorans Lipopolysaccharide That Contains Two Galacturonic Acid Residues in the Backbone and Malic Acid A Tertiary Acyl Substituent. International Journal of Molecular Sciences. 2020; 21(21):7991. https://doi.org/10.3390/ijms21217991
Chicago/Turabian StyleChoma, Adam, Katarzyna Zamłyńska, Andrzej Mazur, Anna Pastuszka, Zbigniew Kaczyński, and Iwona Komaniecka. 2020. "Lipid A from Oligotropha carboxidovorans Lipopolysaccharide That Contains Two Galacturonic Acid Residues in the Backbone and Malic Acid A Tertiary Acyl Substituent" International Journal of Molecular Sciences 21, no. 21: 7991. https://doi.org/10.3390/ijms21217991
APA StyleChoma, A., Zamłyńska, K., Mazur, A., Pastuszka, A., Kaczyński, Z., & Komaniecka, I. (2020). Lipid A from Oligotropha carboxidovorans Lipopolysaccharide That Contains Two Galacturonic Acid Residues in the Backbone and Malic Acid A Tertiary Acyl Substituent. International Journal of Molecular Sciences, 21(21), 7991. https://doi.org/10.3390/ijms21217991






