Doxorubicin-Induced Translocation of mtDNA into the Nuclear Genome of Human Lymphocytes Detected Using a Molecular-Cytogenetic Approach
Abstract
1. Introduction
2. Results
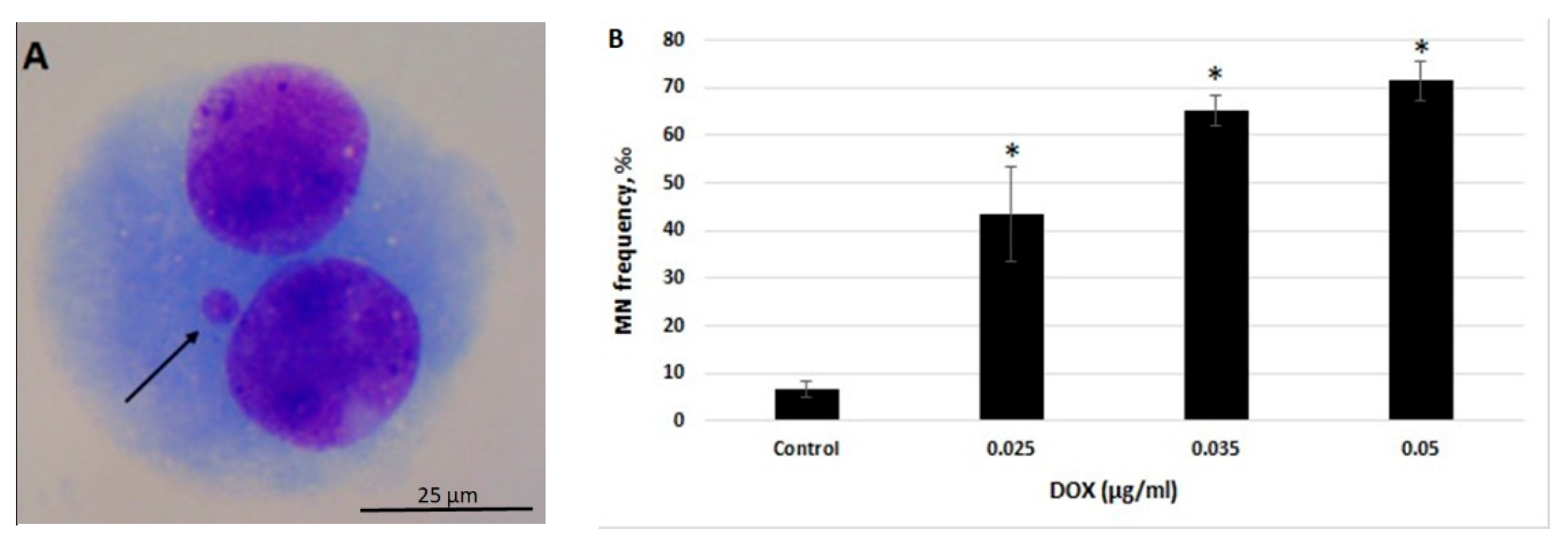
2.1. CBMN Assay in Human Whole Blood Lymphocytes
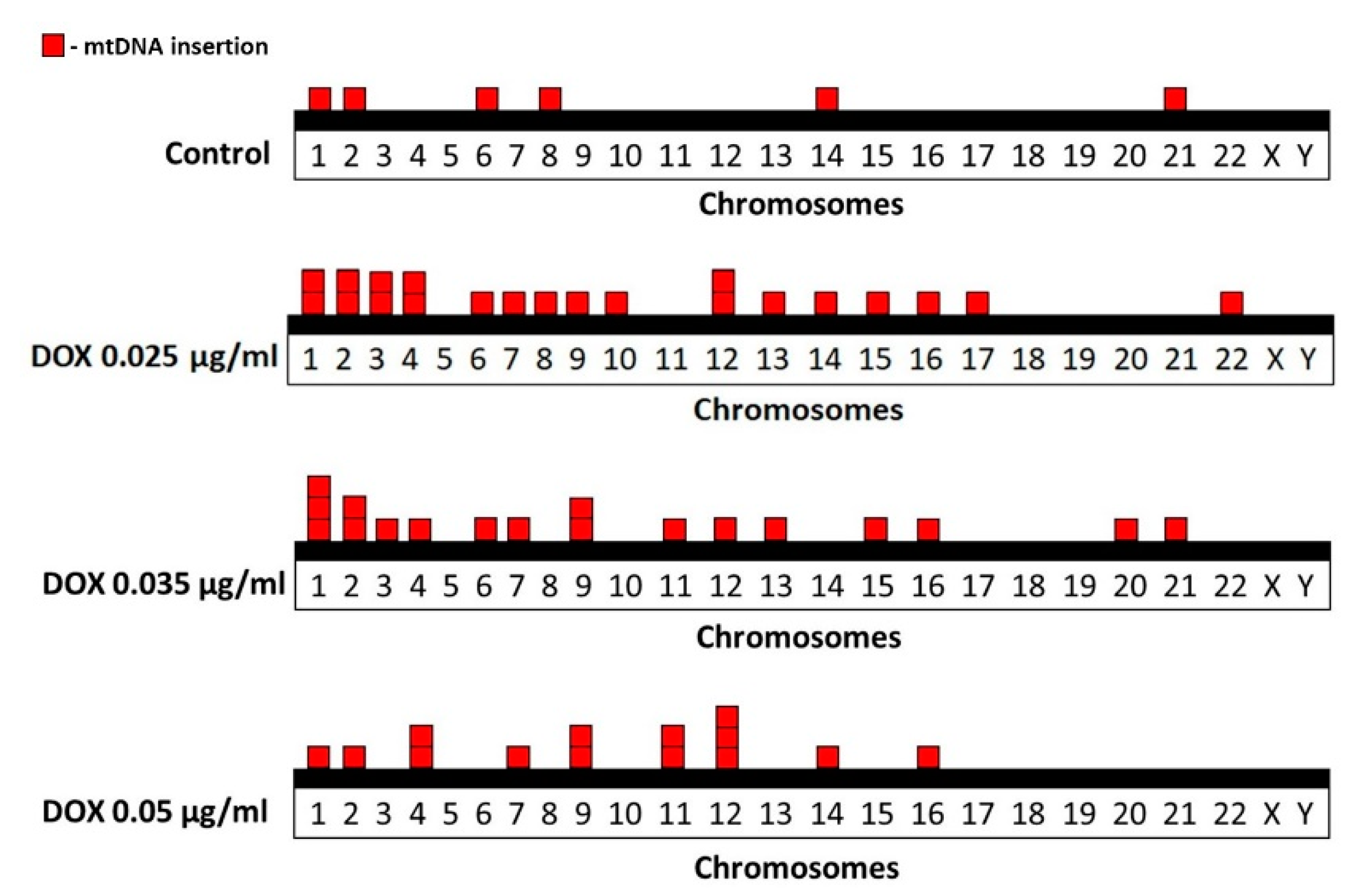
2.2. FISH Analysis of mtDNA Translocation in Metaphase Chromosomes
2.3. Correlation Studies of mtDNA Insertions
3. Discussion
4. Materials and Methods
4.1. Human Whole Blood Cultures
4.2. CBMN Assay
4.3. Metaphase Chromosome Preparation
4.4. FISH Probe Synthesis
4.5. FISH Analysis
4.6. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| CBMN CI DOX DSB FISH | Cytokinesis-blocked micronucleus Confidence interval Doxorubicin DNA double-stranded breaks Fluorescence in situ hybridization |
| mtDNA | Mitochondrial DNA |
| NHEJ NUMT PCR RPMI | Non-homologous end joining Nuclear DNA sequences of mitochondrial origin Polymerase chain reaction Roswell Park Memorial Institute |
References
- Leister, D. Origin, evolution and genetic effects of nuclear insertions of organelle DNA. Trends Genet. 2005, 21, 655–663. [Google Scholar] [CrossRef] [PubMed]
- Hazkani-Covo, E.; Zeller, M.; Martin, W. Molecular poltergeists: Mitochondrial DNA copies (numts) in sequenced nuclear genomes. PLoS Genet 2010, 6, e1000834. [Google Scholar] [CrossRef] [PubMed]
- Calabrese, M.; Simone, D.; Attimfonelli, M. Primates and mouse NumtS in the UCSC Genome Browser. BMC Bioinform. 2012, 4, S15. [Google Scholar] [CrossRef] [PubMed]
- Willett-Brozick, J.E.; Savul, S.A.; Richey, L.E.; Baysal, B.E. Germ line insertion of mtDNA at the breakpoint junction of a reciprocal constitutional translocation. Hum. Genet. 2001, 109, 216–223. [Google Scholar] [CrossRef] [PubMed]
- Turner, C.; Killoran, C.; Thomas, N.S.; Rosenberg, M.; Chuzhanova, N.A.; Johnston, J.; Kemel, Y.; Cooper, D.N.; Biesecker, L.G. Human genetic disease caused by de novo mitochondrial-nuclear DNA transfer. Hum. Genet. 2003, 112, 303–309. [Google Scholar] [CrossRef] [PubMed]
- Borensztajn, K.; Chafa, O.; Alhenc-Gelas, M.; Salha, S.; Reghis, A.; Fischer, A.M.; Tapon-Bretaudière, J. Characterization of two novel splice site mutations in human factor VII gene causing severe plasma factor VII deficiency and bleeding diathesis. Br. J. Haematol. 2002, 117, 168–171. [Google Scholar] [CrossRef] [PubMed]
- Goldin, E.; Stahl, S.; Cooney, A.M.; Kaneski, C.R.; Gupta, S.; Brady, R.O.; Ellis, J.R.; Schiffmann, R. Transfer of a mitochondrial DNA fragment to MCOLN1 causes an inherited case of mucolipidosis IV. Hum. Mutat. 2004, 24, 460–465. [Google Scholar] [CrossRef]
- Ju, Y.S.; Tubio, J.M.; Mifsud, W.; Fu, B.; Davies, H.R.; Ramakrishna, M.; Li, Y.; Yates, L.; Gundem, G.; Tarpey, P.S.; et al. Frequent somatic transfer of mitochondrial DNA into the nuclear genome of human cancer cells. Genome Res. 2015, 25, 814–824. [Google Scholar] [CrossRef]
- Srinivasainagendra, V.; Sandel, M.W.; Singh, B.; Sundaresan, A.; Mooga, V.P.; Bajpai, P.; Tiwari, H.K.; Singh, K.K. Migration of mitochondrial DNA in the nuclear genome of colorectal adenocarcinoma. Genome Med. 2017, 9, 31. [Google Scholar] [CrossRef]
- Singh, K.K.; Choudhury, A.R.; Tiwari, H.K. Numtogenesis as a mechanism for development of cancer. Semin. Cancer Biol. 2017, 47, 101–109. [Google Scholar] [CrossRef]
- Hazkani-Covo, E.; Covo, S. Numt-mediated double-strand break repair mitigates deletions during primate genome evolution. PLoS Genet. 2008, 4, e1000237. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Timmis, J.N. Cytoplasmic organelle DNA preferentially inserts into open chromatin. Genome Biol. Evol. 2013, 5, 1060–1064. [Google Scholar] [CrossRef] [PubMed]
- Ricchetti, M.; Fairhead, C.; Dujon, B. Mitochondrial DNA repairs double-strand breaks in yeast chromosomes. Nature 1999, 402, 96–100. [Google Scholar] [CrossRef] [PubMed]
- Decottignies, A. Capture of extranuclear DNA at fission yeast double-strand breaks. Genetics 2005, 171, 1535–1548. [Google Scholar] [CrossRef] [PubMed]
- Cortes-Funes, H.; Coronado, C. Role of anthracyclines in the era of targeted therapy. Cardiovasc. Toxicol. 2007, 7, 56–60. [Google Scholar] [CrossRef]
- Nathenson, M.J.; Conley, A.P.; Lin, H.; Fleming, N.; Ravi, V. Treatment of Recurrent or Metastatic Uterine Adenosarcoma. Sarcoma 2017, 2017, 4680273. [Google Scholar] [CrossRef]
- Yang, S.; Wang, X.Q. XLF-mediated NHEJ activity in hepatocellular carcinoma therapy resistance. BMC Cancer 2017, 17, 344. [Google Scholar] [CrossRef]
- Lough, A.N.; Faries, K.M.; Koo, D.H.; Hussain, A.; Roark, L.M.; Langewisch, T.L.; Backes, T.; Kremling, K.A.; Jiang, J.; Birchler, J.A.; et al. Cytogenetic and Sequence Analyses of Mitochondrial DNA Insertions in Nuclear Chromosomes of Maize. G3 Genes Genomes Genet. 2015, 5, 2229–2239. [Google Scholar] [CrossRef]
- Caro, P.; Gómez, J.; Arduini, A.; González-Sánchez, M.; González-García, M.; Borrás, C.; Viña, J.; Puertas, M.J.; Sastre, J.; Barja, G. Mitochondrial DNA sequences are present inside nuclear DNA in rat tissues and increase with age. Mitochondrion 2010, 10, 479–486. [Google Scholar] [CrossRef]
- Chondrou, V.; Trochoutsou, K.; Panayides, A.; Efthimiou, M.; Stephanou, G.; Demopoulos, N.A. Combined study on clastogenic, aneugenic and apoptotic properties of doxorubicin in human cells in vitro. J. Biol. Res. 2018, 25, 17. [Google Scholar] [CrossRef]
- Simone, D.; Calabrese, F.M.; Lang, M.; Gasparre, G.; Attimonelli, M. The reference human nuclear mitochondrial sequences compilation validated and implemented on the UCSC genome browser. BMC Genom. 2011, 12, 517. [Google Scholar] [CrossRef] [PubMed]
- Scherer, S. Guide to Human Genome. Available online: http://www.cshlp.org/ghg5_all/section/dna.shtml (accessed on 28 March 2020).
- Abdullaev, S.A.; Fomenko, L.A.; Kuznetsova, E.A.; Gaziev, A.I. Experimental detection of integration of mTDNA in the nuclear genome induced by ionizing radiation. Radiats. Biol. Radioecol. 2013, 53, 380–388. [Google Scholar] [PubMed]
- Yang, F.; Teves, S.S.; Kemp, C.J.; Henikoff, S. Doxorubicin, DNA torsion, and chromatin dynamics. Biochim. Biophys. Acta 2014, 1845, 84–89. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.; Stevens, J.B.; Horne, S.D.; Abdallah, B.Y.; Ye, K.J.; Bremer, S.W.; Ye, C.J.; Chen, D.J.; Heng, H.H. Genome chaos: Survival strategy during crisis. Cell Cycle. 2014, 13, 528–537. [Google Scholar] [CrossRef]
- Khiati, S.; Dalla Rosa, I.; Sourbier, C.; Ma, X.; Rao, V.A.; Neckers, L.M.; Zhang, H.; Pommier, Y. Mitochondrial topoisomerase I (top1mt) is a novel limiting factor of doxorubicin cardiotoxicity. Clin. Cancer Res. 2014, 20, 4873–4881. [Google Scholar] [CrossRef]
- Pillai, V.B.; Bindu, S.; Sharp, W.; Fang, Y.H.; Kim, G.; Gupta, M.; Samant, S.; Gupta, M.P. Sirt3 protects mitochondrial DNA damage and blocks the development of doxorubicin-induced cardiomyopathy in mice. Am. J. Physiol. Heart Circ. Physiol. 2016, 310, 962–972. [Google Scholar] [CrossRef]
- Dhawan, A.; Kayani, M.A.; Parry, J.M.; Parry, E.; Anderson, D. Aneugenic and clastogenic effects of doxorubicin in human lymphocytes. Mutagenesis 2003, 18, 487–490. [Google Scholar] [CrossRef]
- Hsu, C.C.; Tseng, L.M.; Lee, H.C. Role of mitochondrial dysfunction in cancer progression. Exp. Biol. Med. 2016, 241, 1281–1295. [Google Scholar] [CrossRef]
- De Araujo, L.F.; Fonseca, A.S.; Muys, B.R.; Plaça, J.R.; Bueno, R.B.; Lorenzi, J.C.; Santos, A.R.; Molfetta, G.A.; Zanette, D.L.; Souza, J.E.; et al. Mitochondrial genome instability in colorectal adenoma and adenocarcinoma. Tumour Biol. 2015, 36, 8869–8879. [Google Scholar] [CrossRef]
- Yuan, Y.; Ju, Y.S.; Kim, Y.; Li, J.; Wang, Y.; Yoon, C.J.; Yang, Y.; Martincorena, I.; Creighton, C.J.; Weinstein, J.N.; et al. Comprehensive molecular characterization of mitochondrial genomes in human cancers. Nat. Genet. 2020, 52, 342–352. [Google Scholar] [CrossRef]
- Baumgartner, A.; Schmid, T.E.; Cemeli, E.; Anderson, D. Parallel evaluation of doxorubicin-induced genetic damage in human lymphocytes and sperm using the comet assay and spectral karyotyping. Mutagenesis 2004, 19, 313–318. [Google Scholar] [CrossRef] [PubMed]
- Ricchetti, M.; Tekaia, F.; Dujon, B. Continued colonization of the human genome by mitochondrial DNA. PLoS Biol. 2004, 2, E273. [Google Scholar] [CrossRef] [PubMed]
- Yin, J.; Guo, J.; Zhang, Q.; Cui, L.; Zhang, L.; Zhang, T.; Zhao, J.; Li, J.; Middleton, A.; Carmichael, P.L.; et al. Doxorubicin-induced mitophagy and mitochondrial damage is associated with dysregulation of the PINK1/parkin pathway. Toxicol. Vitr. 2018, 51, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Campbell, C.L.; Thorsness, P.E. Escape of mitochondrial DNA to the nucleus in yme1 yeast is mediated by vacuolar-dependent turnover of abnormal mitochondrial compartments. J. Cell Sci. 1998, 111, 2455–2464. [Google Scholar]
- Wai, T.; García-Prieto, J.; Baker, M.J.; Merkwirth, C.; Benit, P.; Rustin, P.; Rupérez, F.J.; Barbas, C.; Ibañez, B.; Langer, T. Imbalanced OPA1 processing and mitochondrial fragmentation cause heart failure in mice. Science 2015, 350, aad0116. [Google Scholar] [CrossRef]
- Fenech, M. The in Vitro Micronucleus Technique. Mutat. Res. 2000, 455, 81–95. [Google Scholar] [CrossRef]
- Harutyunyan, T.; Hovhannisyan, G.; Sargsyan, A.; Grigoryan, B.; Al-Rikabi, A.H.; Weise, A.; Liehr, T.; Aroutiounian, R. Analysis of copy number variations induced by ultrashort electron beam radiation in human leukocytes in vitro. Mol. CytoGenet. 2019, 12, 18. [Google Scholar] [CrossRef]
- Ramos, A.; Santos, C.; Alvarez, L.; Nogués, R.; Aluja, M.P. Human mitochondrial DNA complete amplification and sequencing: A new validated primer set that prevents nuclear DNA sequences of mitochondrial origin co-amplification. Electrophoresis 2009, 30, 1587–1593. [Google Scholar] [CrossRef]
- MedCalc. Available online: https://www.medcalc.org/calc/odds_ratio.php (accessed on 5 May 2020).

| DOX Doses (µg/mL) | Frequency of mtDNA Insertion per Metaphase(Mean ± SD) | Odds Ratio (95% CI) |
|---|---|---|
| Control | 0.0033 ± 0.0015 | - |
| 0.025 | 0.0116 ± 0.0023 a | 3.53 (1.42–8.76) b |
| 0.035 | 0.0100 ± 0.0015 a | 3.02 (1.19–7.62) b |
| 0.05 | 0.0078 ± 0.0031 | 2.34 (0.89–6.11) |
| mtDNA Insertions | ||||
|---|---|---|---|---|
| Control | DOX 0.025 µg/mL | DOX 0.035 µg/mL | DOX 0.05 µg/mL | |
| Chromosome length | 0.321 | 0.631 ** | 0.502 * | 0.319 |
| Gene density (gene/Mb) | −0.153 | 0.134 | −0.06 | −0.01 |
| NUMTs per chromosome | 0.361 | 0.667 ** | 0.509 * | 0.376 |
| Number of MN | 0.886 | 0.985 * | 0.592 | 0.956 * |
| Product Length (bp) | Sequence (5’–3’) | Primers pair Length (bp) |
|---|---|---|
| 1822 | for tagccatgcactactcaccaga rev ggatgaggcaggaatcaaagac | 22 |
| 1758 | for ctgtatccgacatctggttcct rev gtttagctcagagcggtcaagt | 22 |
| 2543 | for acttaagggtcgaaggtggatt rev tcgatgttgaagcctgagacta | 22 |
| 3005 | for aagtcaccctagccatcattcta rev gatatcatagctcagaccatacc | 23 |
| 2709 | for ctgctggcatcactatactacta rev gattggtgggtcattatgtgttg | 23 |
| 1738 | for cttaccacaaggcacacctaca rev ggcacaatattggctaagaggg | 22 |
| 1866 | for gtctggcctatgagtgactaca rev cagttcttgtgagctttctcgg | 22 |
| 1853 | for ctccctctacatatttaccacaac rev aagtcctaggaaagtgacagcga | 24 |
| 1872 | for gcaggaatacctttcctcacag rev gtgcaagaataggaggtggagt | 22 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Harutyunyan, T.; Al-Rikabi, A.; Sargsyan, A.; Hovhannisyan, G.; Aroutiounian, R.; Liehr, T. Doxorubicin-Induced Translocation of mtDNA into the Nuclear Genome of Human Lymphocytes Detected Using a Molecular-Cytogenetic Approach. Int. J. Mol. Sci. 2020, 21, 7690. https://doi.org/10.3390/ijms21207690
Harutyunyan T, Al-Rikabi A, Sargsyan A, Hovhannisyan G, Aroutiounian R, Liehr T. Doxorubicin-Induced Translocation of mtDNA into the Nuclear Genome of Human Lymphocytes Detected Using a Molecular-Cytogenetic Approach. International Journal of Molecular Sciences. 2020; 21(20):7690. https://doi.org/10.3390/ijms21207690
Chicago/Turabian StyleHarutyunyan, Tigran, Ahmed Al-Rikabi, Anzhela Sargsyan, Galina Hovhannisyan, Rouben Aroutiounian, and Thomas Liehr. 2020. "Doxorubicin-Induced Translocation of mtDNA into the Nuclear Genome of Human Lymphocytes Detected Using a Molecular-Cytogenetic Approach" International Journal of Molecular Sciences 21, no. 20: 7690. https://doi.org/10.3390/ijms21207690
APA StyleHarutyunyan, T., Al-Rikabi, A., Sargsyan, A., Hovhannisyan, G., Aroutiounian, R., & Liehr, T. (2020). Doxorubicin-Induced Translocation of mtDNA into the Nuclear Genome of Human Lymphocytes Detected Using a Molecular-Cytogenetic Approach. International Journal of Molecular Sciences, 21(20), 7690. https://doi.org/10.3390/ijms21207690






