Exceptionally Long-Lived Individuals (ELLI) Demonstrate Slower Aging Rate Calculated by DNA Methylation Clocks as Possible Modulators for Healthy Longevity
Abstract
1. Introduction
2. Results
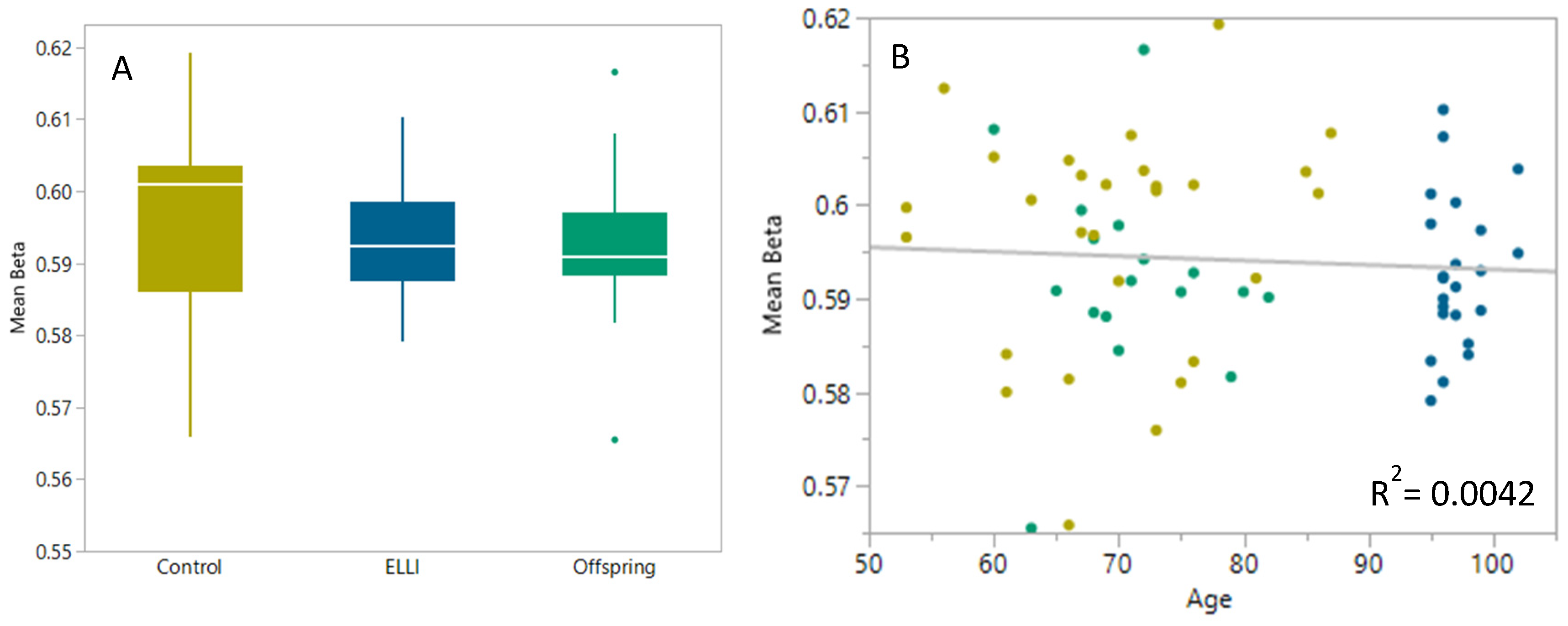
2.1. DNA Methylation
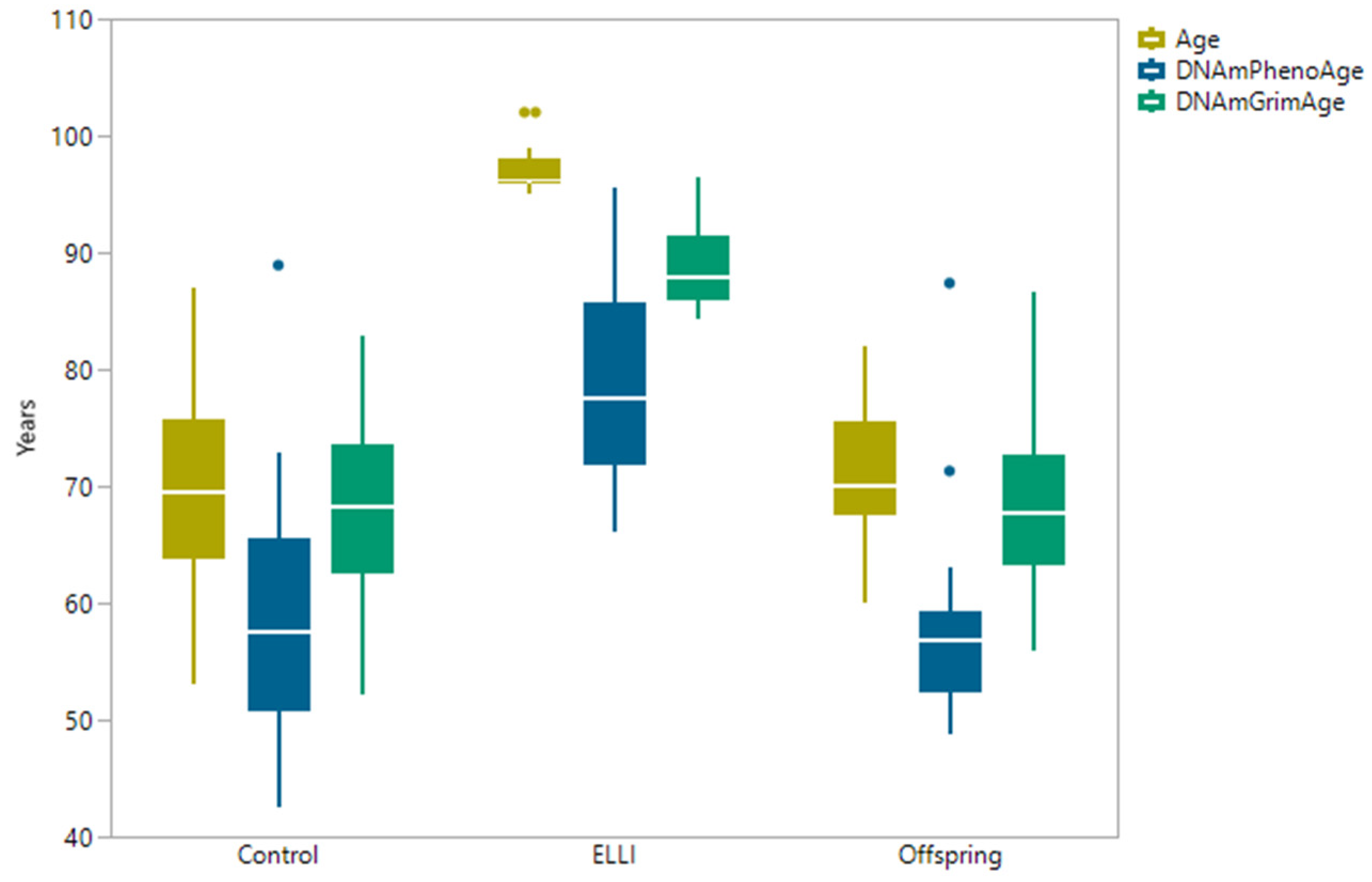
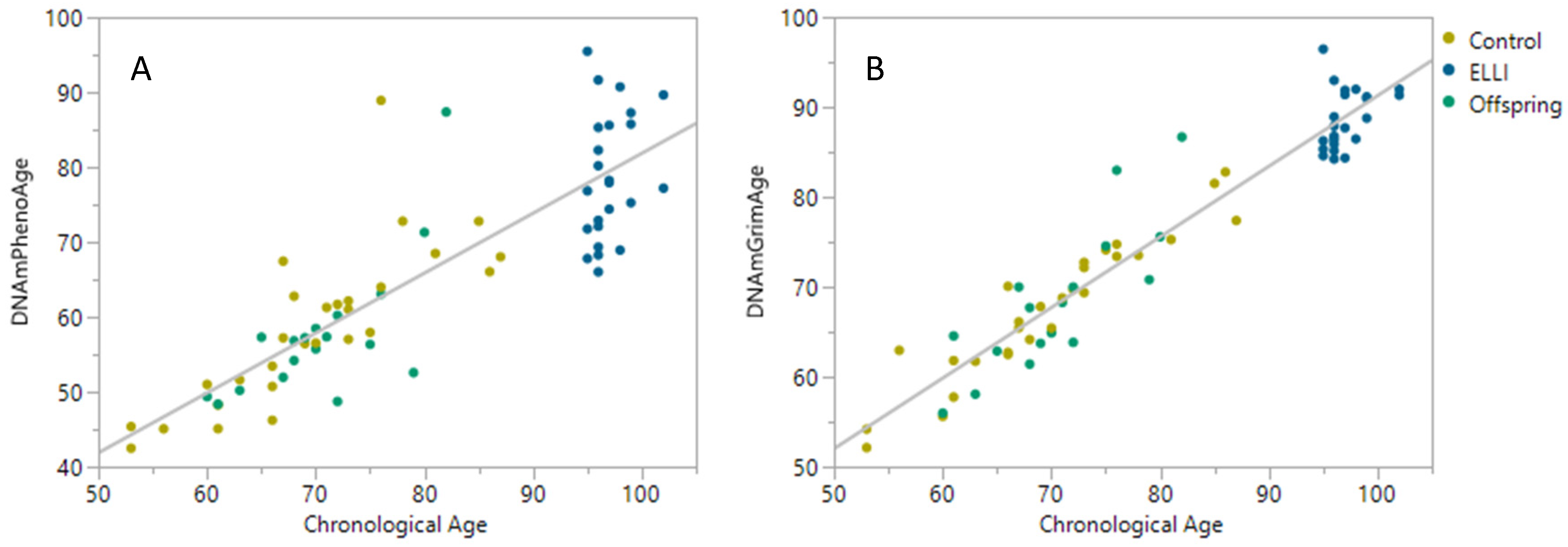
2.2. DNAm Age Clocks
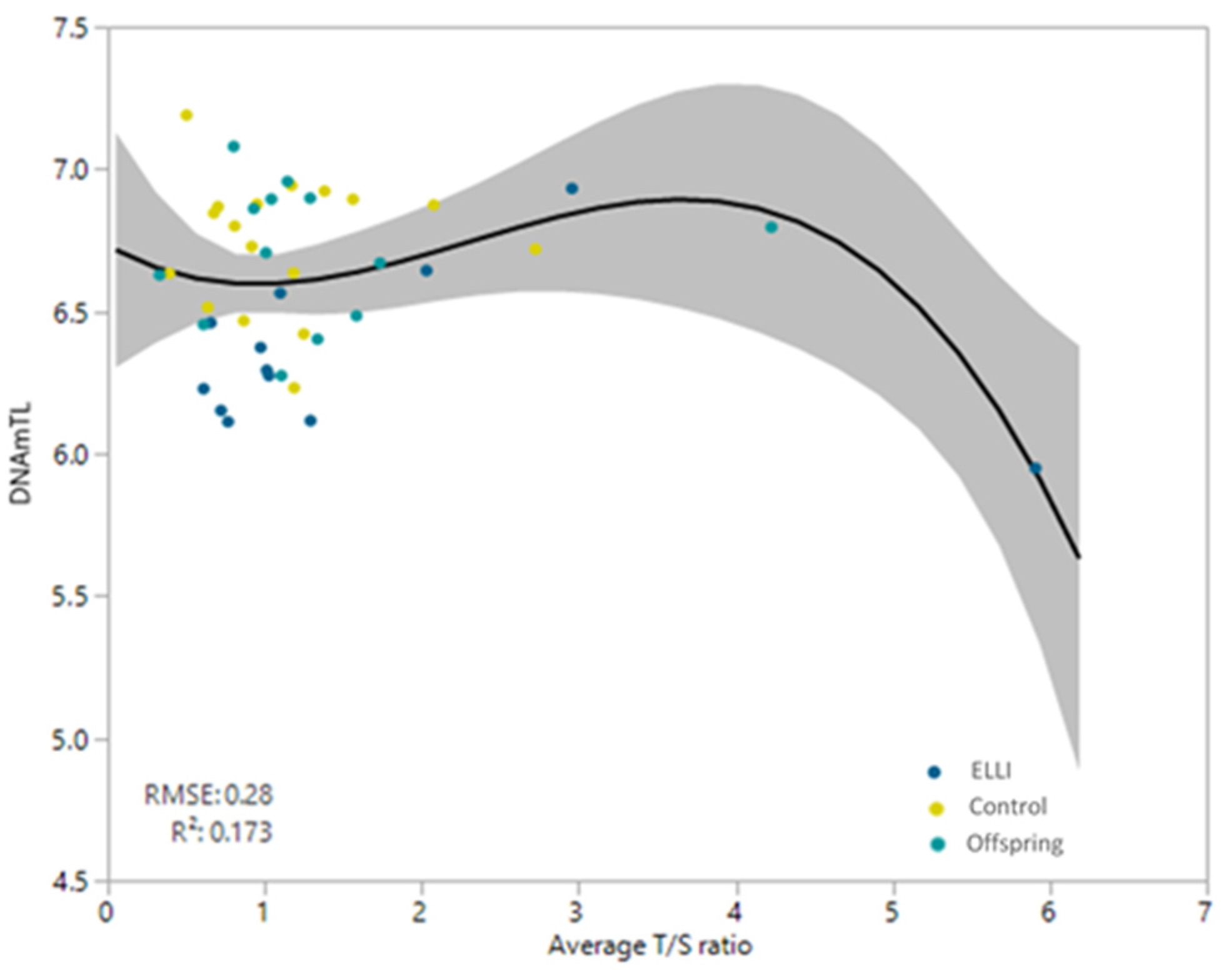
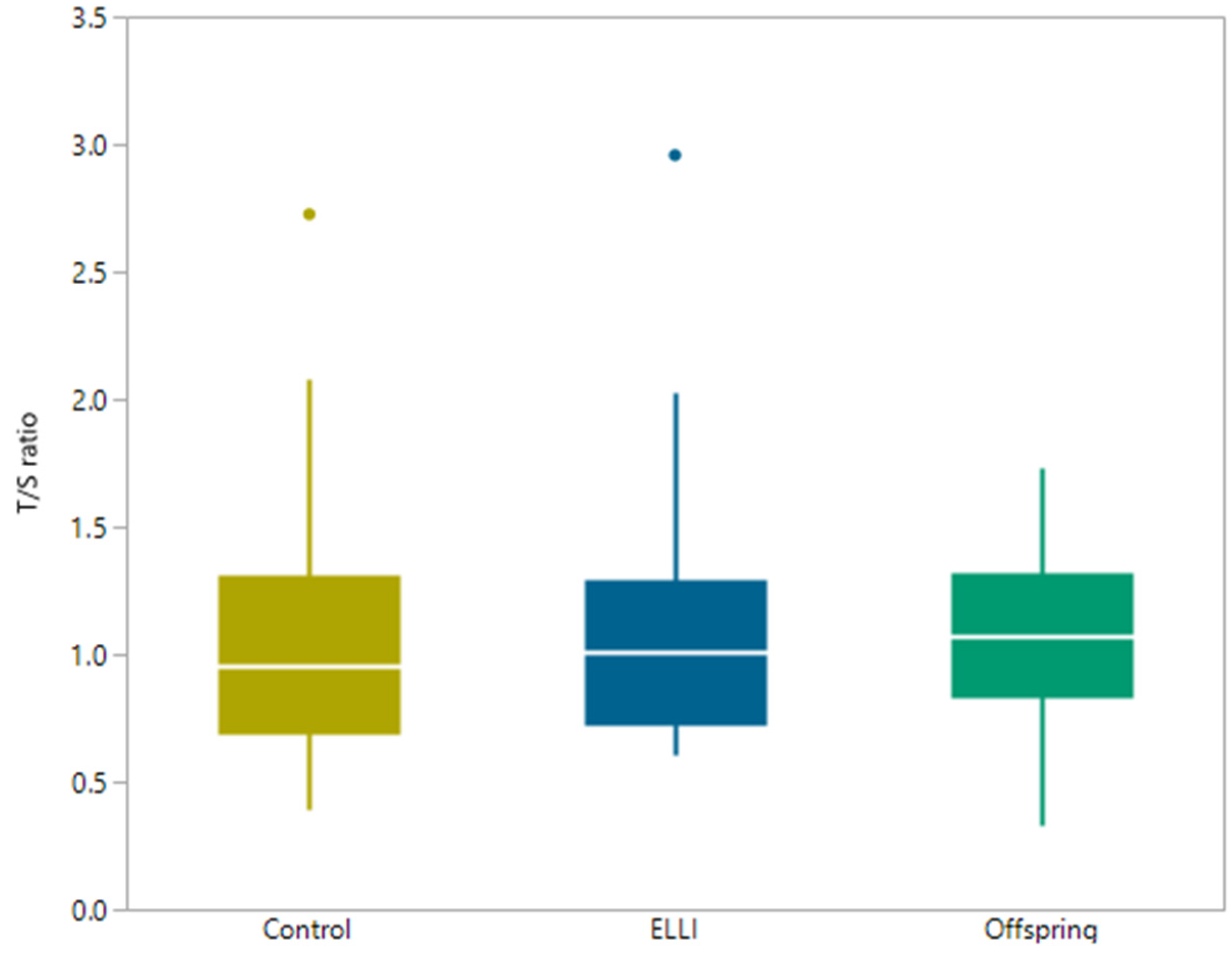
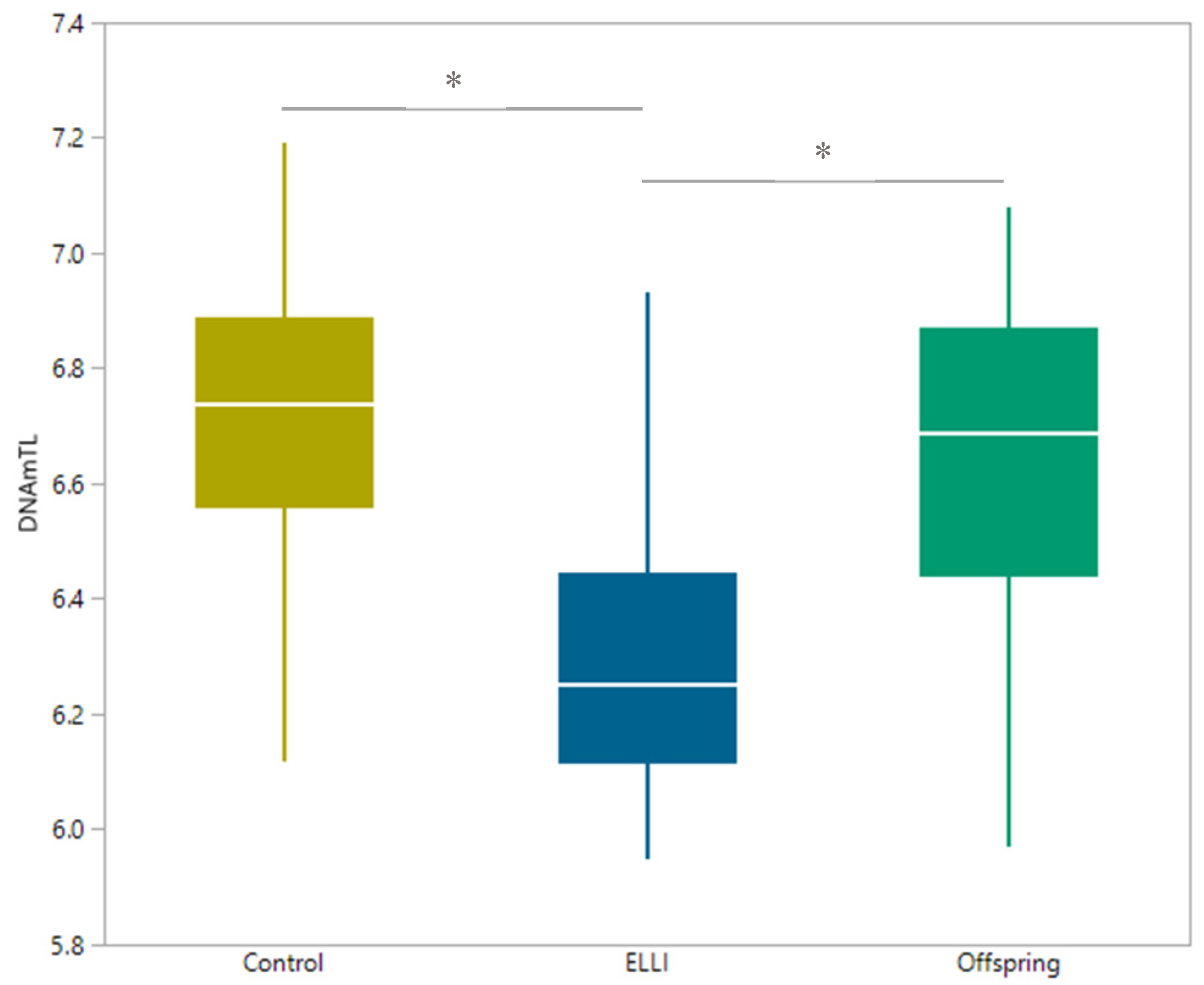
2.3. Telomere length
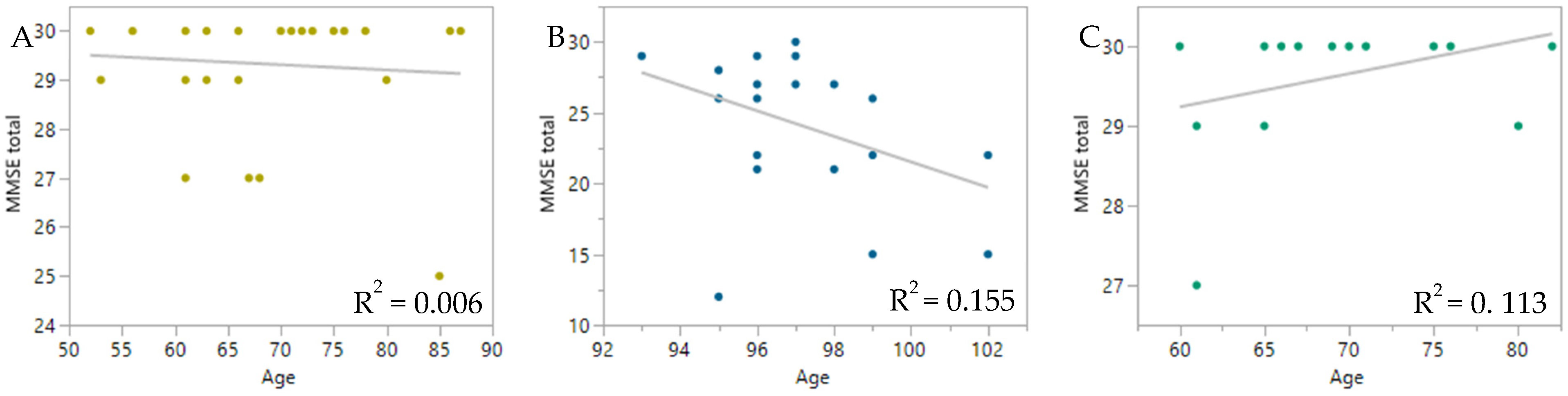
3. Discussion
4. Materials and Methods
4.1. Ethics Statement
4.2. Sample Collection and Preparation
4.3. DNA Methylation Analysis
4.4. Quantitative PCR for Relative Telomere Length Assessment
4.5. Statistical Analyses
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| IMECS | Israeli Multi-Ethnic Centenarian Study |
| ELLI | Exceptionally Long-Lived Individuals |
| DNAm | DNA methylation |
| qPCR | Quantitative Polymerase Chain Reaction |
| MMSE | Mini-Mental State Exam |
| CpG | Cytosine Guanine dinucleotide |
| TL | Telomere Length |
Appendix A
| Group | Number of Participants | Mean Age | STDEV | Min Age | Max Age |
|---|---|---|---|---|---|
| ELLI | 24 | 97.1 | 2.04 | 95 | 102 |
| Offspring | 18 | 70.4 | 6.25 | 60 | 82 |
| Control | 28 | 69.7 | 9.1 | 53 | 87 |
| Group | Gender | % Smokers | Education level | ||||
|---|---|---|---|---|---|---|---|
| F | M | Elementary | High School | College Degree | NA | ||
| ELLI | 12 | 12 | 57% | 4 | 10 | 7 | 3 |
| Offspring | 12 | 6 | 37% | 1 | 3 | 12 | 2 |
| Control | 20 | 8 | 39% | 0 | 7 | 21 | 0 |
| Age Measure | Group Pairs | Kruskal-Wallis P-Value | Wilcoxon Post-Hoc |
|---|---|---|---|
| DNAm PhenoAge | ELLI—Offspring | <0.0001 | <0.0001 |
| ELLI—Control | <0.0001 | ||
| Control—Offspring | 0.5969 | ||
| DNAm GrimAge | ELLI—Offspring | <0.0001 | <0.0001 |
| ELLI—Control | <0.0001 | ||
| Control—Offspring | 0.9731 |
| Measurement Pairs | Within Pairs P-Value | Among Pairs P-Value |
|---|---|---|
| DNAmPhenoAge—actual age | <0.0001 | <0.0001 |
| DNAmGrimAge—actual age | <0.0001 | <0.0001 |
| DNAmGrimAge—DNAmPhenoAge | 0.7812 | <0.0001 |
| Group | Median Average T/S | Minimum Average T/S | Maximum Average T/S |
|---|---|---|---|
| ELLI | 1.01 | 0.61 | 5.91 |
| Offspring | 1.11 | 0.33 | 4.23 |
| Control | 0.95 | 0.39 | 2.73 |
| Group Pairs | ANOVA P-Value | Tukey Post-Hoc |
|---|---|---|
| ELLI—Offspring | <0.0001 | 0.0003 |
| ELLI—Control | <0.0001 | |
| Control—Offspring | 0.3388 |
| Primer Name | Primer Sequence | Working Concentration |
|---|---|---|
| Telo-F | CGGTTTGTTTGGGTTTGGGTTTGGGTTTGGGTTTGGGTT | 10 µM |
| Telo-R | GGCTTGCCTTACCCTTACCCTTACCCTTACCCTTACCCT | 10 µM |
| IFNB1-F | GGTTACCTCCGAAACTGAAGA | 7.5 µM |
| IFNB1-R | CCTTTCATATGCAGTACATTAGCC | 7.5 µM |
| Reagent | Telo Reaction | IFNB1 Reaction |
|---|---|---|
| Ultra-Pure Water (Bio-Lab ltd, Jerusalem, Israel) | 6 µL | 4 µL |
| SYBR green (Thermo Fisher Scientific Baltics, Vilnius, Lithuania) | 10 µL | 10 µL |
| F primer (Sigma-Aldrich Israel ltd, Rehovot, Israel) | 0.2 µL | 0.6 µL |
| R primer (Sigma-Aldrich Israel ltd, Rehovot, Israel) | 1.8 µL | 1.4 µL |
| Genomic DNA (5 ng/µL) | 2 µL | 4 µL |
References
- Yashin, A.; De Benedictis, G.; Vaupel, J.; Tan, Q.; Andreev, K.; Iachine, I.; Bonafe, M.; Valensin, S.; De Luca, M.; Carotenuto, L.; et al. Genes and longevity: Lessons from studies of centenarians. J. Gerontol. Ser. A Boil. Sci. Med. Sci. 2000, 55, B319–B328. [Google Scholar] [CrossRef] [PubMed]
- Atzmon, G.; Cho, M.; Cawthon, R.M.; Budagov, T.; Katz, M.; Yang, X.; Siegel, G.; Bergman, A.; Huffman, D.M.; Schechter, C.B.; et al. Evolution in health and medicine Sackler colloquium: Genetic variation in human telomerase is associated with telomere length in Ashkenazi centenarians. Proc. Natl. Acad. Sci. USA 2010, 107, 1710–1717. [Google Scholar] [CrossRef] [PubMed]
- Freudenberg-Hua, Y.; Freudenberg, J.; Vacic, V.; Abhyankar, A.; Emde, A.-K.; Ben-Avraham, D.; Barzilai, N.; Oschwald, D.; Christen, E.; Koppel, J.; et al. Disease variants in genomes of 44 centenarians. Mol. Genet. Genom. Med. 2014, 2, 438–450. [Google Scholar] [CrossRef] [PubMed]
- Rubino, G.; Bulati, M.; Aiello, A.; Aprile, S.; Gambino, C.M.; Gervasi, F.; Caruso, C.; Accardi, G. Sicilian centenarian offspring are more resistant to immune ageing. Aging Clin. Exp. Res. 2018, 31, 125–133. [Google Scholar] [CrossRef] [PubMed]
- Stevenson, M.; Bae, H.; Schupf, N.; Andersen, S.; Zhang, Q.; Perls, T.; Sebastiani, P. Burden of disease variants in participants of the long life family Study. Aging 2015, 7, 123–132. [Google Scholar] [CrossRef] [PubMed]
- Giuliani, C.; Sazzini, M.; Pirazzini, C.; Bacalini, M.G.; Marasco, E.; Ruscone, G.A.G.; Fang, F.; Sarno, S.; Gentilini, D.; Di Blasio, A.M.; et al. Impact of demography and population dynamics on the genetic architecture of human longevity. Aging 2018, 10, 1947–1963. [Google Scholar] [CrossRef]
- Teixeira, L.; Araújo, L.; Jopp, D.; Ribeiro, O. Centenarians in Europe. Maturitas 2017, 104, 90–95. [Google Scholar] [CrossRef]
- Puca, A.A.; Spinelli, C.; Accardi, G.; Villa, F.; Caruso, C. Centenarians as a model to discover genetic and epigenetic signatures of healthy ageing. Mech. Ageing Dev. 2018, 174, 95–102. [Google Scholar] [CrossRef]
- Milman, S.; Barzilai, N. Dissecting the Mechanisms Underlying Unusually Successful Human Health Span and Life Span. Cold Spring Harb. Perspect. Med. 2015, 6, a025098. [Google Scholar] [CrossRef]
- Atzmon, G.; Rincon, M.; Schechter, C.B.; Shuldiner, A.R.; Lipton, R.B.; Bergman, A.; Barzilai, N. Lipoprotein genotype and conserved pathway for exceptional longevity in humans. PLoS Boil. 2006, 4, e113. [Google Scholar] [CrossRef]
- Barzilai, N.; Atzmon, G.; Schechter, C.; Schaefer, E.J.; Cupples, A.L.; Lipton, R.; Cheng, S.; Shuldiner, A.R. Unique Lipoprotein Phenotype and Genotype Associated With Exceptional Longevity. JAMA 2003, 290, 2030–2040. [Google Scholar] [CrossRef] [PubMed]
- Ben-Avraham, D.; Govindaraju, D.R.; Budagov, T.; Fradin, D.; Durda, P.; Liu, B.; Ott, S.; Gutman, D.; Sharvit, L.; Kaplan, R.; et al. The GH receptor exon 3 deletion is a marker of male-specific exceptional longevity associated with increased GH sensitivity and taller stature. Sci. Adv. 2017, 3, e1602025. [Google Scholar] [CrossRef] [PubMed]
- López-Otín, C.; Blasco, M.A.; Partridge, L.; Serrano, M.; Kroemer, G. The hallmarks of aging. Cell 2013, 153, 1194–1217. [Google Scholar] [CrossRef]
- Austriaco, N.R., Jr.; Guarente, L.P. Changes of telomere length cause reciprocal changes in the lifespan of mother cells in Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. USA 1997, 94, 9768–9772. [Google Scholar] [CrossRef]
- Teixeira, M.T. Saccharomyces cerevisiae as a Model to Study Replicative Senescence Triggered by Telomere Shortening. Front. Oncol. 2013, 3, 101. [Google Scholar] [CrossRef]
- Espejel, S.; Klatt, P.; Murcia, J.M.-D.; Martiín-Caballero, J.; Flores, J.M.; Taccioli, G.; De Murcia, G.; Blasco, M.A. Impact of telomerase ablation on organismal viability, aging, and tumorigenesis in mice lacking the DNA repair proteins PARP-1, Ku86, or DNA-PKcs. J. Cell Boil. 2004, 167, 627–638. [Google Scholar] [CrossRef]
- Honig, L.S.; Schupf, N.; Lee, J.H.; Tang, M.X.; Mayeux, R. Shorter telomeres are associated with mortality in those with APOE ϵ4 and dementia. Ann. Neurol. 2006, 60, 181–187. [Google Scholar] [CrossRef]
- Vera, E.; De Jesus, B.B.; Foronda, M.; Flores, J.M.; Blasco, M.A. The Rate of Increase of Short Telomeres Predicts Longevity in Mammals. Cell Rep. 2012, 2, 732–737. [Google Scholar] [CrossRef]
- Simons, M.J. Questioning causal involvement of telomeres in aging. Ageing Res. Rev. 2015, 24, 191–196. [Google Scholar] [CrossRef]
- Harley, C.B.; Futcher, A.B.; Greider, C.W. Telomeres shorten during ageing of human fibroblasts. Nature 1990, 345, 458–460. [Google Scholar] [CrossRef]
- Aviv, A.; Shay, J.W. Reflections on telomere dynamics and ageing-related diseases in humans. Philos. Trans. R. Soc. B Boil. Sci. 2018, 373, 20160436. [Google Scholar] [CrossRef]
- Sanders, J.L.; Newman, A.B. Telomere Length in Epidemiology: A Biomarker of Aging, Age-Related Disease, Both, or Neither? Epidemiologic Rev. 2013, 35, 112–131. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.; Duc, K.D.; Holcman, D.; Teixeira, M.T. The length of the shortest telomere as the major determinant of the onset of replicative senescence. Genetics 2013, 194, 847–857. [Google Scholar] [CrossRef] [PubMed]
- Savage, S.A. Beginning at the ends: Telomeres and human disease. F1000Research 2018, 7. [Google Scholar] [CrossRef]
- Levy, D.; Neuhausen, S.L.; Hunt, S.C.; Kimura, M.; Hwang, S.-J.; Chen, W.; Bis, J.C.; Fitzpatrick, A.L.; Smith, E.; Johnson, A.D.; et al. Genome-wide association identifies OBFC1 as a locus involved in human leukocyte telomere biology. Proc. Natl. Acad. Sci. USA 2010, 107, 9293–9298. [Google Scholar] [CrossRef] [PubMed]
- Terry, D.F.; Nolan, V.G.; Andersen, S.L.; Perls, T.T.; Cawthon, R. Association of longer telomeres with better health in centenarians. Journals Gerontol. Ser. A: Boil. Sci. Med Sci. 2008, 63, 809–812. [Google Scholar] [CrossRef] [PubMed]
- Gutman, D.; Sharvit, L.; Atzmon, G. Possible Mechanisms for Telomere Length Maintenance in Extremely Old People. Hered. Genet. 2014, 3, 1–2. [Google Scholar]
- Cawthon, R.M. Telomere measurement by quantitative PCR. Nucleic Acids Res. 2002, 30, e47. [Google Scholar] [CrossRef]
- Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001, 29, 45. [Google Scholar] [CrossRef]
- Cawthon, R.M.; Smith, K.R.; O’Brien, E.; Sivatchenko, A.; Kerber, R.A. Association between telomere length in blood and mortality in people aged 60 years or older. Lancet 2003, 361, 393–395. [Google Scholar] [CrossRef]
- Shekhidem, H.A.; Sharvit, L.; Leman, E.; Manov, I.; Roichman, A.; Holtze, S.; Huffman, D.M.; Cohen, H.Y.; Hildebrandt, T.B.; Shams, I.; et al. Telomeres and Longevity: A Cause or an Effect? Int. J. Mol. Sci. 2019, 20, 3233. [Google Scholar] [CrossRef] [PubMed]
- Vasilishina, A.; Kropotov, A.; Spivak, I.; Bernadotte, A. Relative human telomere length quantification by real-time PCR. Methods Mol. Biol. 2019, 1896, 39–44. [Google Scholar] [PubMed]
- Axelrad, M.D.; Budagov, T.; Atzmon, G. Telomere Length and Telomerase Activity; A Yin and Yang of Cell Senescence. J. Vis. Exp. 2013, 75, e50246. [Google Scholar] [CrossRef] [PubMed]
- Montpetit, A.J.; Alhareeri, A.A.; Montpetit, M.; Starkweather, A.R.; Elmore, L.W.; Filler, K.; Mohanraj, L.; Burton, C.W.; Menzies, V.S.; Lyon, D.E.; et al. Telomere length: A review of methods for measurement. Nurs. Res. 2014, 63, 289–299. [Google Scholar] [CrossRef]
- Hannum, G.; Guinney, J.; Zhao, L.; Zhang, L.; Hughes, G.; Sadda, S.; Klotzle, B.; Bibikova, M.; Fan, J.-B.; Gao, Y. Genome-wide methylation profiles reveal quantitative views of human aging rates. Mol. Cell 2013, 49, 359–367. [Google Scholar] [CrossRef]
- Horvath, S.; Pirazzini, C.; Bacalini, M.G.; Gentilini, D.; Di Blasio, A.M.; Delledonne, M.; Mari, D.; Arosio, B.; Monti, D.; Passarino, G.; et al. Decreased epigenetic age of PBMCs from Italian semi-supercentenarians and their offspring. Aging 2015, 7, 1159–1170. [Google Scholar] [CrossRef]
- Horvath, S. DNA methylation age of human tissues and cell types. Genome Boil. 2013, 14, R115. [Google Scholar] [CrossRef]
- Chen, B.H.; Marioni, R.E.; Colicino, E.; Peters, M.J.; Ward-Caviness, C.K.; Tsai, P.-C.; Roetker, N.S.; Just, A.C.; Demerath, E.W.; Guan, W.; et al. DNA methylation-based measures of biological age: Meta-analysis predicting time to death. Aging 2016, 8, 1844–1859. [Google Scholar] [CrossRef]
- Horvath, S.; Oshima, J.; Martin, G.M.; Lu, A.T.; Quach, A.; Cohen, H.; Felton, S.; Matsuyama, M.; Lowe, D.; Kabacik, S.; et al. Epigenetic clock for skin and blood cells applied to Hutchinson Gilford Progeria Syndrome and ex vivo studies. Aging 2018, 10, 1758. [Google Scholar] [CrossRef]
- Armstrong, N.J.; Mather, K.A.; Thalamuthu, A.; Wright, M.J.; Trollor, J.N.; Ames, D.; Brodaty, H.; Schofield, P.R.; Sachdev, P.S.; Kwok, J.B. Aging, exceptional longevity and comparisons of the Hannum and Horvath epigenetic clocks. Epigenomics 2017, 9, 689–700. [Google Scholar] [CrossRef]
- Ryan, J.; Wrigglesworth, J.; Loong, J.; Fransquet, P.D.; Woods, R.L.; Anderson, R. A systematic review and meta-analysis of environmental, lifestyle and health factors associated with DNA methylation age. J. Gerontol. Ser. A Boil. Sci. Med. Sci. 2019. [Google Scholar] [CrossRef] [PubMed]
- Lu, A.T.; Quach, A.; Wilson, J.G.; Reiner, A.P.; Aviv, A.; Raj, K.; Hou, L.; Baccarelli, A.A.; Li, Y.; Stewart, J.D.; et al. DNA methylation GrimAge strongly predicts lifespan and healthspan. Aging 2019, 11, 303–327. [Google Scholar] [CrossRef] [PubMed]
- Lu, A.T.; Seeboth, A.; Tsai, P.-C.; Sun, D.; Quach, A.; Reiner, A.P.; Kooperberg, C.; Ferrucci, L.; Hou, L.; Baccarelli, A.A.; et al. DNA methylation-based estimator of telomere length. Aging 2019, 11, 5895–5923. [Google Scholar] [CrossRef]
- Levine, M.E.; Lu, A.T.; Quach, A.; Chen, B.H.; Assimes, T.L.; Bandinelli, S.; Hou, L.; Baccarelli, A.A.; Stewart, J.D.; Li, Y.; et al. An epigenetic biomarker of aging for lifespan and healthspan. Aging 2018, 10, 573–591. [Google Scholar] [CrossRef]
- Fahy, G.M.; Brooke, R.T.; Watson, J.P.; Good, Z.; Vasanawala, S.S.; Maecker, H.; Leipold, M.D.; Lin, D.T.S.; Kobor, M.S.; Horvath, S. Reversal of epigenetic aging and immunosenescent trends in humans. Aging Cell 2019, 18, e13028. [Google Scholar] [CrossRef]
- Xiao, F.-H.; Wang, H.-T.; Kong, Q.-P. Dynamic DNA Methylation During Aging: A “Prophet” of Age-Related Outcomes. Front. Genet. 2019, 10, 107. [Google Scholar] [CrossRef]
- Bernadotte, A.; Mikhelson, V.M.; Spivak, I.M. Markers of cellular senescence. Telomere shortening as a marker of cellular senescence. Aging 2016, 8, 3–11. [Google Scholar] [CrossRef]
- Lin, Q.; Weidner, C.I.; Costa, I.G.; Marioni, R.E.; Ferreira, M.R.P.; Deary, I.J.; Wagner, W. DNA methylation levels at individual age-associated CpG sites can be indicative for life expectancy. Aging 2016, 8, 394–401. [Google Scholar] [CrossRef]
- Shiels, P.G.; Stenvinkel, P.; Kooman, J.P.; McGuinness, D. Circulating markers of ageing and allostatic load: A slow train coming. Pract. Lab. Med. 2017, 7, 49–54. [Google Scholar] [CrossRef]
- Olivieri, F.; Capri, M.; Bonafè, M.; Morsiani, C.; Jung, H.J.; Spazzafumo, L.; Viña, J.; Suh, Y. Circulating miRNAs and miRNA shuttles as biomarkers: Perspective trajectories of healthy and unhealthy aging. Mech. Ageing Dev. 2017, 165, 162–170. [Google Scholar] [CrossRef]
- Deelen, J.; Akker, E.B.V.D.; Trompet, S.; Van Heemst, D.; Mooijaart, S.P.; Slagboom, P.E.; Beekman, M. Employing biomarkers of healthy ageing for leveraging genetic studies into human longevity. Exp. Gerontol. 2016, 82, 166–174. [Google Scholar] [CrossRef] [PubMed]
- Barral, S.; Singh, J.; Fagan, E.; Cosentino, S.; Andersen-Toomey, S.L.; Wojczynski, M.K.; Feitosa, M.; Kammerer, C.M.; Schupf, N.; Long Life Family Study. Age-Related Biomarkers in LLFS Families With Exceptional Cognitive Abilities. J. Gerontol. Ser. A Boil. Sci. Med Sci. 2017, 72, 1683–1688. [Google Scholar]
- Aryee, M.J.; Jaffe, A.E.; Corrada-Bravo, H.; Ladd-Acosta, C.; Feinberg, A.P.; Hansen, K.D.; Irizarry, R.A. Minfi: A flexible and comprehensive Bioconductor package for the analysis of Infinium DNA methylation microarrays. Bioinformatics 2014, 30, 1363–1369. [Google Scholar] [CrossRef] [PubMed]
- Jaffe, A.E.; Murakami, P.; Lee, H.; Leek, J.T.; Fallin, M.D.; Feinberg, A.P.; Irizarry, R.A. Bump hunting to identify differentially methylated regions in epigenetic epidemiology studies. Int. J. Epidemiol. 2012, 41, 200–209. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gutman, D.; Rivkin, E.; Fadida, A.; Sharvit, L.; Hermush, V.; Rubin, E.; Kirshner, D.; Sabin, I.; Dwolatzky, T.; Atzmon, G. Exceptionally Long-Lived Individuals (ELLI) Demonstrate Slower Aging Rate Calculated by DNA Methylation Clocks as Possible Modulators for Healthy Longevity. Int. J. Mol. Sci. 2020, 21, 615. https://doi.org/10.3390/ijms21020615
Gutman D, Rivkin E, Fadida A, Sharvit L, Hermush V, Rubin E, Kirshner D, Sabin I, Dwolatzky T, Atzmon G. Exceptionally Long-Lived Individuals (ELLI) Demonstrate Slower Aging Rate Calculated by DNA Methylation Clocks as Possible Modulators for Healthy Longevity. International Journal of Molecular Sciences. 2020; 21(2):615. https://doi.org/10.3390/ijms21020615
Chicago/Turabian StyleGutman, Danielle, Elina Rivkin, Almog Fadida, Lital Sharvit, Vered Hermush, Elad Rubin, Dani Kirshner, Irina Sabin, Tzvi Dwolatzky, and Gil Atzmon. 2020. "Exceptionally Long-Lived Individuals (ELLI) Demonstrate Slower Aging Rate Calculated by DNA Methylation Clocks as Possible Modulators for Healthy Longevity" International Journal of Molecular Sciences 21, no. 2: 615. https://doi.org/10.3390/ijms21020615
APA StyleGutman, D., Rivkin, E., Fadida, A., Sharvit, L., Hermush, V., Rubin, E., Kirshner, D., Sabin, I., Dwolatzky, T., & Atzmon, G. (2020). Exceptionally Long-Lived Individuals (ELLI) Demonstrate Slower Aging Rate Calculated by DNA Methylation Clocks as Possible Modulators for Healthy Longevity. International Journal of Molecular Sciences, 21(2), 615. https://doi.org/10.3390/ijms21020615






