A Review of Starch Biosynthesis in Relation to the Building Block-Backbone Model
Abstract
1. Introduction
2. Components of Starch Granules
2.1. Amylopectin
2.2. Amylose
2.3. Other Components
3. Architecture of Starch Granules
4. The Amylopectin Backbone Model
5. The Backbone Model in Relation to the Granular Structure
6. Biosynthesis of the Starch Granule
7. Commitment of Carbon for Starch Biosynthesis
8. Starch Granule Initiation
8.1. Proteins Associated with Granule Initiation
8.2. Contribution of the Initiation Machinery to Granule Structure
9. Amylose Synthesis
9.1. Granule Bound Starch Synthase
9.2. Contribution of GBSS/PTST to the Building Block-Backbone Model
10. Amylopectin Synthesis
10.1. Starch Synthases
10.2. The Contribution of SS Isoforms to the Building Block-Backbone Model
10.3. Starch Branching Enzymes
10.4. The Contribution of SBE Isoforms to the Building Block-Backbone Model
10.5. Starch Debranching Enzymes
10.6. The Contribution of DBEs to the Building Block-Backbone Model
10.7. Other Enzyme Activities
10.7.1. α-Glucan Phosphorylation
10.7.2. α-Glucan Modification by DPE and SP
10.8. Enzyme Complexes
11. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Meyer, K.H.; Bernfeld, P. Recherches sur l’amidon V. L’amylopectine. Helv. Chim. Acta 1940, 23, 875–885. [Google Scholar] [CrossRef]
- Gunja-smith, Z.; Marshall, J.J.; Mercier, C.; Smith, E.E.; Whelan, W.J. A revision of the Meyer-Bernfeld model of glycogen and amylopectin. FEBS Lett. 1970, 12, 101–104. [Google Scholar] [CrossRef]
- Northcote, D.H. The Molecular Structure and Shape of Yeast Glycogen. Biochem. J. 1953, 53, 348–352. [Google Scholar] [CrossRef] [PubMed]
- Nikuni, Z. Starch and cooking (in Japanese). Jpn. Soc. Cook. Sci. 1969, 2, 6–14. [Google Scholar]
- French, D. Fine structure of starch and its relationship to the organisation of starch granules. J. Jpn. Soc. Starch Sci. 1972, 19, 8–25. [Google Scholar] [CrossRef]
- Robin, J.P.; Mercier, C.; Charbonnière, R.; Guilbot, A. Lintnerized starches. Gel filtration and enzymatic studies of insoluble residues from prolonged acid treatment of potato starch. Cereal Chem. 1974, 51, 389–406. [Google Scholar]
- Hizukuri, S. Polymodal distribution of the chain lengths of amylopectins, and its significance. Carbohydr. Res. 1986, 147, 342–347. [Google Scholar] [CrossRef]
- Matheson, N.K.; Caldwell, R.A. Modeling of α(1–4) chain arrangements in α(1–4)(1–6) glucans: The action and outcome of β-amylase and Pseudomonas stutzeri amylase on an α(1–4)(1–6) glucan model. Carbohydr. Polym. 2008, 72, 625–637. [Google Scholar] [CrossRef]
- Bertoft, E.; Laohaphatanalert, K.; Piyachomkwan, K.; Sriroth, K. The fine structure of cassava starch amylopectin. Part 2: Building block structure of clusters. Int. J. Biol. Macromol. 2010, 47, 325–335. [Google Scholar] [CrossRef]
- Bertoft, E. On the Building Block and Backbone Concepts of Amylopectin Structure. Cereal Chem. 2013, 90, 294–311. [Google Scholar] [CrossRef]
- Brust, H.; Orzechowski, S.; Fettke, J. Starch and glycogen analyses: Methods and techniques. Biomolecules 2020, 10, 1020. [Google Scholar] [CrossRef] [PubMed]
- Ball, S.; Guan, H.; James, M.; Myers, A.; Keeling, P.; Mouille, G.; Colonna, P.; Preiss, J. From Glycogen to Amylopectin: A Model for the Biogenesis of the Plant Starch Granule. Cell 1996, 86, 349–352. [Google Scholar] [CrossRef]
- Seung, D.; Echevarría-Poza, A.; Steuernagel, B.; Smith, A.M. Natural Polymorphisms in Arabidopsis Result in Wide Variation or Loss of the Amylose Component. Plant Physiol. 2020, 182, 870–881. [Google Scholar] [CrossRef] [PubMed]
- Bertoft, E. Understanding Starch Structure: Recent Progress. Agronomy 2017, 7, 56. [Google Scholar] [CrossRef]
- Donald, A.M. Plasticization and self assembly in the starch granule. Cereal Chem. 2001, 78, 307–314. [Google Scholar] [CrossRef]
- Doutch, J.; Gilbert, E.P. Characterisation of large scale structures in starch granules via small-angle neutron and X-ray scattering. Carbohydr. Polym. 2013, 91, 444–451. [Google Scholar] [CrossRef]
- Hizukuri, S.; Takeda, Y.; Yasuda, M.; Suzuki, A. Multi-branched nature of amylose and the action of debranching enzymes. Carbohydr. Res. 1981, 94, 205–213. [Google Scholar] [CrossRef]
- Hanashiro, I.; Sakaguchi, I.; Yamashita, H. Branched Structures of Rice Amylose Examined by Differential Fluorescence Detection of Side-chain Distribution. J. Appl. Glycosci. 2013, 60, 79–85. [Google Scholar] [CrossRef]
- Wang, K.; Hasjim, J.; Wu, A.C.; Henry, R.J.; Gilbert, R.G. Variation in Amylose Fine Structure of Starches from Different Botanical Sources. J. Agric. Food Chem. 2014. [Google Scholar] [CrossRef]
- Wang, K.; Hasjim, J.; Chi, A.; Li, E.; Henry, R.J.; Gilbert, R.G. Roles of GBSSI and SSIIa in determining amylose fine structure. Carbohydr. Polym. 2015, 127, 264–274. [Google Scholar] [CrossRef]
- Pérez, S.; Bertoft, E. The molecular structures of starch components and their contribution to the architecture of starch granules: A comprehensive review. Starch Staerke 2010, 62, 389–420. [Google Scholar] [CrossRef]
- Jenkins, P.J.; Donald, A.M. The influence of amylose on starch granule structure. Int. J. Biomacromolecules 1995, 17, 315–321. [Google Scholar] [CrossRef]
- Jane, J.; Shen, J.J. Internal structure of the potato starch granule revealed by chemical gelatinization. Carbohydr. Res. 1993, 247, 279–290. [Google Scholar] [CrossRef]
- Pan, D.D.; Jane, J. Internal Structure of Normal Maize Starch Granules Revealed by Chemical Surface Gelatinization. Biomacromolecules 2000, 1, 126–132. [Google Scholar] [CrossRef] [PubMed]
- Tester, R.F.; Morrison, W.R. Swelling and gelatinization of cereal starches. III. Some properties of waxy and normal nonwaxy barley starches. Cereal Chem. 1992, 69, 654–658. [Google Scholar]
- Swinkels, J.J.M. Composition and properties of commercial native starches. Starch Starke 1985, 37, 1–5. [Google Scholar] [CrossRef]
- Bettge, A.D.; Giroux, M.J.; Morris, C.F. Susceptibility of waxy starch granules to mechanical damage. Cereal Chem. 2000, 77, 750–753. [Google Scholar] [CrossRef]
- Hanashiro, I.; Itoh, K.; Kuratomi, Y.; Yamazaki, M.; Igarashi, T.; Matsugasako, J.; Takeda, Y. Granule-Bound Starch Synthase I is Responsible for Biosynthesis of Extra-Long Unit Chains of Amylopectin in Rice. Plant Cell Physiol. 2008, 49, 925–933. [Google Scholar] [CrossRef]
- Maddelein, M.; Libessarts, N.; Bellanger, F.; Delrue, B.; Hulst, C.D.; Koornhuyse, N.V.D.; Fontaine, T.; Wieruszeski, J.; Decq, A.; Ball, S. Toward an Understanding of the Biogenesis of the Starch Granule. J. Biol. Chem. 1994, 269, 25150–25157. [Google Scholar]
- Wang, Y.J.; White, P.; Pollak, L.; Jane, J. Characterization of starch structures of 17 maize endosperm mutant genotypes with Oh43 inbred line background. Cereal Chem. 1993, 70, 171–179. [Google Scholar]
- Klucinec, J.D.; Thompson, D.B. Structure of amylopectins from ae- containing maize starches. Cereal Chem. 2002, 79, 19–23. [Google Scholar] [CrossRef]
- Han, W.; Zhang, B.; Li, J.; Zhao, S.; Niu, M.; Jia, C. Understanding the fine structure of intermediate materials of maize starches. Food Chem. 2017, 233, 450–456. [Google Scholar] [CrossRef]
- Putseys, J.A.; Derde, L.J.; Lamberts, L.; Östman, E.; Bjorck, I.M.; Delcour, J.A. Functionality of short chain amylose-Lipid complexes in starch-Water systems and their impact on in vitro starch degradation. J. Agric. Food Chem. 2010, 58, 1939–1945. [Google Scholar] [CrossRef] [PubMed]
- Morrison, W.R. Lipids in Cereal Starches: A Review. J. Cereal Sci. 1988, 8, 1–15. [Google Scholar] [CrossRef]
- Hizukuri, S.; Tabata, S.; Kagoshima; Nikuni, Z. Studies on Starch Phosphate Part 1. Estimation of glucose-6-phosphate residues in starch and the presence of other bound phosphate(s). Starch/Stärke 1970, 22, 338–343. [Google Scholar] [CrossRef]
- Nitschke, F.; Wang, P.; Schmieder, P.; Girard, J.M.; Awrey, D.E.; Wang, T.; Israelian, J.; Zhao, X.; Turnbull, J.; Heydenreich, M.; et al. Hyperphosphorylation of glucosyl C6 carbons and altered structure of glycogen in the neurodegenerative epilepsy lafora disease. Cell Metab. 2013, 17, 756–767. [Google Scholar] [CrossRef] [PubMed]
- Blennow, A.; Engelsen, S.B.; Munck, L.; Møller, B.L. Starch molecular structure and phosphorylation investigated by a combined chromatographic and chemometric approach. Carbohydr. Polym. 2000, 41, 163–174. [Google Scholar] [CrossRef]
- Dhital, S.; Shrestha, A.K.; Hasjim, J.; Gidley, M.J. Physicochemical and Structural Properties of Maize and Potato Starches as a Function of Granule Size. J. Agric. Food Chem. 2011, 59, 10151–10161. [Google Scholar] [CrossRef]
- Blennow, A.; Sjöland, A.K.; Andersson, R.; Kristiansson, P. The distribution of elements in the native starch granule as studied by particle-induced X-ray emission and complementary methods. Anal. Biochem. 2005, 347, 327–329. [Google Scholar] [CrossRef]
- Denyer, K.; Sidebottom, C.; Hylton, C.M.; Smith, A.M. Soluble isoforms of starch synthase and starch-branching enzyme also occur within starch granules in developing pea embryos. Plant J. 1993, 4, 191–198. [Google Scholar] [CrossRef]
- Mu-Forster, A.C.; Huang, R.; Powers, J.R.; Harriman, R.W.; Knight, M.; Singletary, G.W.; Keeling, P.L.; Wasserman, B.P. Physical Association of Starch Biosynthetic Enzymes with Starch Granules of Maize Endosperm: Granule-Associated Forms of Starch Synthase I and Starch Branching Enzyme II. Plant Physiol. 1996, 111, 821–829. [Google Scholar] [CrossRef] [PubMed]
- Glaring, M.A.; Koch, C.B.; Blennow, A. Genotype-specific spatial distribution of starch molecules in the starch granule: A combined CLSM and SEM approach. Biomacromolecules 2006, 7, 2310–2320. [Google Scholar] [CrossRef] [PubMed]
- Fannon, J.E.; Hauber, R.J.; Bemiller, J.N. Surface pores of starch granules. Cereal Chem. 1992, 69, 284–288. [Google Scholar]
- Benmoussa, M.; Hamaker, B.R.; Huang, C.P.; Sherman, D.M.; Weil, C.F.; BeMiller, J.N. Elucidation of maize endosperm starch granule channel proteins and evidence for plastoskeletal structures in maize endosperm amyloplasts. J. Cereal Sci. 2010, 52, 22–29. [Google Scholar] [CrossRef]
- Jane, J.-L.; Kasemsuwan, T.; Leas, S.; Zobel, H.; Robyt, J.F. Anthology of Starch Granule Morphology by Scanning Electron Microscopy. Starch Stärke 1994, 46, 121–129. [Google Scholar] [CrossRef]
- Ellis, R.P.; Cochrane, M.P.; Dale, M.F.B.; Duþus, C.M.; Lynn, A.; Morrison, I.M.; Prentice, R.D.M.; Swanston, J.S.; Tiller, S. Starch Production and Industrial Use. J. Sci. Food Agric. 1998, 77, 289–311. [Google Scholar] [CrossRef]
- Lindeboom, N.; Chang, P.R.; Tyler, R.T. Analytical, Biochemical and Physicochemical Aspects of Starch Granule Size, with Emphasis on Small Granule Starches: A Review. Starch Stärke 2004, 56, 89–99. [Google Scholar] [CrossRef]
- Tetlow, I.; Emes, M. Starch Biosynthesis in the Developing Endosperms of Grasses and Cereals. Agronomy 2017, 7, 81. [Google Scholar] [CrossRef]
- Srichuwong, S.; Candra, T.; Mishima, T.; Isono, N.; Hisamatsu, M. Starches from different botanical sources I: Contribution of amylopectin fine structure to thermal properties and enzyme digestibility. Carbohydr. Polym. 2005, 60, 529–538. [Google Scholar] [CrossRef]
- Chanzy, H.; Putaux, J.; Dupeyre, D.; Davies, R.; Burghammer, M.; Montanari, S.; Riekel, C. Morphological and structural aspects of the giant starch granules from Phajus grandifolius. J. Struct. Biol. 2006, 154, 100–110. [Google Scholar] [CrossRef]
- Waterschoot, J.; Gomand, S.V.; Fierens, E.; Delcour, J.A. Production, structure, physicochemical and functional properties of maize, cassava, wheat, potato and rice starches. Starch Stärke 2015, 67, 14–29. [Google Scholar] [CrossRef]
- Fuentes, C.; Perez-Rea, D.; Bergenståhl, B.; Carballo, S.; Sjöö, M.; Nilsson, L. Physicochemical and structural properties of starch from five Andean crops grown in Bolivia. Int. J. Biol. Macromol. 2019, 125, 829–838. [Google Scholar] [CrossRef]
- Chen, P.; Yu, L.; Chen, L.; Li, X. Morphology and microstructure of Maize starches with different amylose/amylopectin Content. Starch Stärke 2006, 58, 611–615. [Google Scholar] [CrossRef]
- Fredriksson, H.; Silverio, J.; Andersson, R.; Eliasson, A.-C.; Åman, P. The influence of amylose and amylopectin characteristics on gelatinization and retrogradation properties of different starches. Carbohydr. Polym. 1998, 35, 119–134. [Google Scholar]
- Bathgate, G.N.; Palmer, G.H. The in vivo and in vitro degradation of barley and malt starch granules. J. Inst. Brew. 1973, 73, 402–406. [Google Scholar]
- Saccomanno, B.; Chambers, A.H.; Hayes, A.; Mackay, I.; Mcwilliam, S.C.; Trafford, K. Starch granule morphology in oat endosperm. J. Cereal Sci. 2017, 73, 46–54. [Google Scholar] [CrossRef]
- Chia, T.; Chirico, M.; King, R.; Ramirez-gonzalez, R.; Saccomanno, B.; Seung, D.; Simmonds, J.; Trick, M.; Uauy, C.; Verhoeven, T.; et al. A carbohydrate-binding protein, B-GRANULE CONTENT 1, influences starch granule size distribution in a dose-dependent manner in polyploid wheat. J. Exp. Bot. 2020, 71, 105–115. [Google Scholar] [CrossRef]
- Shapter, F.M.; Henry, R.J.; Lee, L.S. Endosperm and starch granule morphology in wild cereal relatives. Plant. Genet. Resour. 2008, 6, 85–97. [Google Scholar] [CrossRef]
- Paterson, A.H.; Bowers, J.E.; Bruggmann, R.; Dubchak, I.; Grimwood, J.; Gundlach, H.; Haberer, G.; Hellsten, U.; Mitros, T.; Poliakov, A.; et al. The Sorghum bicolor genome and the diversification of grasses. Nature 2009, 457, 551–556. [Google Scholar] [CrossRef]
- Goldstein, A.; Annor, G.; Putaux, J.; Hebelstrup, K.H.; Blennow, A.; Bertoft, E. International Journal of Biological Macromolecules Impact of full range of amylose contents on the architecture of starch granules. Int. J. Biol. Macromol. 2016, 89, 305–318. [Google Scholar] [CrossRef]
- Gallant, D.J.; Bouchet, B.; Baldwin, P.M. Microscopy of starch: Evidence of a new level of granule organization. Carbohydr. Polym. 1997, 32, 177–191. [Google Scholar] [CrossRef]
- De Pater, S.; Caspers, M.; Kottenhagen, M.; Meima, H.; Ter Stege, R.; De Vetten, N. Manipulation of starch granule size distribution in potato tubers by modulation of plastid division. Plant. Biotechnol. J. 2006, 4, 123–134. [Google Scholar] [CrossRef] [PubMed]
- Briarty, L.G.; Hughes, C.E.; Evers, A.D. The Developing Endosperm of Wheat—A Stereological Analysis. Ann. Bot. 1979, 44, 641–658. [Google Scholar] [CrossRef]
- Parker, M.L. The relationship between A-type and B-type starch granules in the developing endosperm of wheat. J. Cereal Sci. 1985, 3, 271–278. [Google Scholar] [CrossRef]
- Jenkins, P.J.; Cameron, R.E.; Donald, A.M. A Universal Feature in the Structure of Starch Granules from Different Botanical Sources. Starch Stärke 1993, 45, 417–420. [Google Scholar] [CrossRef]
- Ambigaipalan, P.; Hoover, R.; Donner, E.; Liu, Q.; Jaiswal, S.; Chibbar, R.; Nantanga, K.K.M.; Seetharaman, K. Structure of faba bean, black bean and pinto bean starches at different levels of granule organization and their physicochemical properties. Food Res. Int. 2011, 44, 2962–2974. [Google Scholar] [CrossRef]
- Pilling, E.; Smith, A.M. Growth ring formation in the starch granules of potato tubers. Plant. Physiol 2003, 132, 365–371. [Google Scholar] [CrossRef]
- Wang, S.; Blazek, J.; Gilbert, E.; Copeland, L. New insights on the mechanism of acid degradation of pea starch. Carbohydr. Polym. 2012, 87, 1941–1949. [Google Scholar] [CrossRef]
- Goldstein, A.; Annor, G.; Vamadevan, V.; Tetlow, I.; Kirkensgaard, J.J.K.; Mortensen, K.; Blennow, A.; Hebelstrup, K.H.; Bertoft, E. Influence of diurnal photosynthetic activity on the morphology, structure, and thermal properties of normal and waxy barley starch. Int. J. Biol. Macromol. 2017, 98, 188–200. [Google Scholar] [CrossRef]
- Baker, A.A.; Miles, M.J.; Helbert, W. Internal structure of the starch granule revealed by AFM. Carbohydr. Res. 2001, 330, 249–256. [Google Scholar] [CrossRef]
- Waigh, T.A.; Perry, P.; Riekel, C.; Gidley, M.J.; Donald, A.M. Chiral side-chain liquid-crystalline polymeric properties of starch. Macromolecules 1998, 31, 7980–7984. [Google Scholar] [CrossRef]
- Oostergetel, G.T.; van Bruggen, E.F.J. The crystalline domains in potato starch granules are arranged in a helical fashion. Carbohydr. Polym. 1993, 21, 7–12. [Google Scholar] [CrossRef]
- Van De Sand-Bakhuyzen, H. The Structure of Starch Grains from Wheat Grown Under Constant Conditions. Sci. Proc. 1926, 302, 2950–2951. [Google Scholar] [CrossRef]
- Buttrose, M. Ultrastructure of the developing aleurone cells of wheat grain. Aust. J. Biol. Sci. 1963, 16, 768–774. [Google Scholar] [CrossRef]
- Perez-Herrera, M.; Vasanthan, T.; Hoover, R. Characterization of Maize Starch Nanoparticles Prepared by Acid Hydrolysis. Cereal Chem. 2016, 93, 323–330. [Google Scholar] [CrossRef]
- Gernat, C.; Radosta, S.; Damaschun, G.; Schierbaum, F. Supramolecular Structure of Legume Starches Revealed by X-Ray Scattering. Starch Stärke 1990, 42, 175–178. [Google Scholar] [CrossRef]
- Tang, H.; Mitsunaga, T.; Kawamura, Y. Molecular arrangement in blocklets and starch granule architecture. Carbohydr. Polym. 2006, 63, 555–560. [Google Scholar] [CrossRef]
- Kozlov, S.S.; Krivandin, A.V.; Shatalova, O.V.; Noda, T.; Bertoft, E.; Fornal, J.; Yuryev, V.P. Structure of starches extracted from near-isogenic wheat lines. Part II. Molecular organization of amylopectin clusters. J. Therm. Anal. Calorim. 2007, 87, 575–584. [Google Scholar] [CrossRef]
- Imberty, A.; Buléon, A.; Tran, V.; Pérez, S. Recent advances in knowledge of starch structure. Starch Stärke 1991, 43, 375–384. [Google Scholar] [CrossRef]
- Buléon, A.; Pontoire, B.; Riekel, C.; Chanzy, H.; Helbert, W.; Vuong, R. Crystalline ultrastructure of starch granules revealed by synchrotron radiation microdiffraction mapping. Macromolecules 1997, 30, 3952–3954. [Google Scholar] [CrossRef]
- Buléon, A.; Colonna, P.; Planchot, V.; Ball, S. Starch granules: Structure and biosynthesis. Int. J. Biol. Macromol. 1998, 23, 85–112. [Google Scholar] [CrossRef] [PubMed]
- Gallant, D.J.; Bouchet, B.; Buléon, A.; Pérez, S. Physical characteristics of starch granules and susceptibility to enzymatic degradation. Eur. J. Clin. Nutr. 1992, 46, S3–S16. [Google Scholar] [PubMed]
- Pfannemüller, B. Influence of chain length of short monodisperse amyloses on the formation of A- and B-type X-ray diffraction patterns. Int. J. Biol. Macromol. 1987, 9, 105–108. [Google Scholar] [CrossRef]
- Zhang, P.; Hamaker, B.R. Banana starch structure and digestibility. Carbohydr. Polym. 2012, 87, 1552–1558. [Google Scholar]
- Biliaderis, C.G.; Grant, D.R.; Vose, J. Structural characterization of legume starches. II. Studies on acid-treated starches. Cereal Chem. 1981, 58, 502–507. [Google Scholar]
- Bogracheva, T.Y.; Morris, V.J.; Ring, S.G.; Hedley, C.L. The granular structure of C-type pea starch and its role in gelatinization. Biopolymers 1998, 45, 323–332. [Google Scholar] [CrossRef]
- Gérard, C.; Planchot, V.; Colonna, P.; Bertoft, E. Relationship between branching density and crystalline structure of A- and B-type maize mutant starches. Carbohydr. Res. 2000, 326, 130–144. [Google Scholar] [CrossRef]
- Lopez-Rubio, A.; Flanagan, B.M.; Gilbert, E.P.; Gidley, M.J. A novel approach for calculating starch crystallinity and its correlation with double helix content: A combined XRD and NMR study. Biopolymers 2008, 89, 761–768. [Google Scholar]
- Bertoft, E.; Piyachomkwan, K.; Chatakanonda, P.; Sriroth, K. Internal unit chain composition in amylopectins. Carbohydr. Polym. 2008, 74, 527–543. [Google Scholar] [CrossRef]
- Cooke, D.; Gidley, M.J. Loss of crystalline and molecular order during starch gelatinisation: Origin of the enthalpic transition. Carbohydr. Res. 1992, 227, 103–112. [Google Scholar] [CrossRef]
- Jane, J.-L.; Wong, K.S.; McPherson, A.E. Branch-structure difference in starches of A and B-type x-ray patterns revealed by their Naegeli dextrins. Carbohydr. Res. 1997, 300, 219–227. [Google Scholar] [CrossRef]
- Hanashiro, I.; Abe, J.; Hizukuri, S. A periodic distribution of the chain length of amylopectin as revealed by high-performance anion-exchange chromatography. Carbohydr. Res. 1996, 283, 151–159. [Google Scholar] [CrossRef]
- Wong, K.S.; Jane, J. Quantitative analysis of debranched amylopectin with a postcolumn enzyme reactor. J. Liq. Chromatogr. 1997, 20, 297–310. [Google Scholar] [CrossRef]
- O’Shea, M.G.; Morel, M.K. High resolution slab gel electrophoresis of 8-amino-1,3,6-pyrenetrisulfonic acid (APTS) tagged oligosaccharides using a DNA sequencer. Electrophoresis 1996, 17, 681–686. [Google Scholar]
- Hobson, P.N.; Whelan, W.J.; Peat, S. The enzymic synthesis and degradation of starch. Part XIV. R-enzyme. J. Chem. Soc. 1951, 333, 1451–1459. [Google Scholar] [CrossRef]
- Manners, D.J. Recent developments in our understanding of glycogen structure. Carbohydr. Polym. 1991, 16, 37–82. [Google Scholar] [CrossRef]
- Bertoft, E. Investigation of the fine structure of alpha-dextrins derived from amylopectin and their relation to the structure of waxy-maize starch. Carbohydr. Res. 1991, 212, 229–244. [Google Scholar]
- Bertoft, E. Hydrolysis of Amylopectin by the Alpha-Amylase of B. subtilis. Carbohydr. Res. 1986, 149, 379–387. [Google Scholar]
- Bertoft, E. Partial Characterization of amylopectin alpha-dextrins. Carbohydr. Res. 1989, 189, 181–183. [Google Scholar]
- Robyt, J.; French, D. Action pattern and specificity of an amylase from Bacillus subtilis. Arch. Biochem. Biophys. 1963, 100, 451–467. [Google Scholar]
- Umeki, K.; Yamamoto, T. Enzymatic determination of structure of singly branched hexaose dextrins formed by liquefying α-amylase of Bacillus subtilis. J. Biochem. 1972, 109, 101–109. [Google Scholar] [CrossRef] [PubMed]
- Bertoft, E.; Zhu, Q.; Andtfolk, H.; Jungner, M. Structural heterogeneity in waxy-rice starch. Carbohydr. Polym. 1999, 38, 349–359. [Google Scholar] [CrossRef]
- Bertoft, E.; Koch, K.; Åman, P. Structure of building blocks in amylopectins. Carbohydr. Res. 2012, 361, 105–113. [Google Scholar] [CrossRef] [PubMed]
- Haworth, W.; Hirst, E.; Isherwood, F. Polysaccharides. Part XXIII. Determination of the chain length of glycogen. J. Chem. Soc. Chem. Commun. 1937, 577–581. [Google Scholar] [CrossRef]
- Staudinger, H.; Husemann, E. Über hochpolymere Verbindungen. 150. Mitteilung. Über die Konstitution der Stärke. Justus Liebigs Ann. Chem. 1937, 527, 195–236. [Google Scholar] [CrossRef]
- Bertoft, E. Composition of building blocks in clusters from potato amylopectin. Carbohydr. Polym. 2007, 70, 123–136. [Google Scholar] [CrossRef]
- Bertoft, E.; Koch, K.; Åman, P. Building block organisation of clusters in amylopectin from different structural types. Int. J. Biol. Macromol. 2012, 50, 1212–1223. [Google Scholar] [CrossRef]
- Kong, X.; Corke, H.; Bertoft, E. Fine structure characterization of amylopectins from grain amaranth starch. Carbohydr. Res. 2009, 344, 1701–1708. [Google Scholar] [CrossRef]
- Zhu, F.; Corke, H.; Åman, P.; Bertoft, E. Structures of clusters in sweetpotato amylopectin. Carbohydr. Res. 2011, 346, 1112–1121. [Google Scholar] [CrossRef]
- Bertoft, E.; Källman, A.; Koch, K.; Andersson, R.; Åman, P. The building block structure of barley amylopectin. Int. J. Biol. Macromol. 2011, 49, 900–909. [Google Scholar] [CrossRef]
- Källman, A.; Bertoft, E.; Koch, K.; Sun, C.; Åman, P.; Andersson, R. Starch structure in developing barley endosperm. Int. J. Biol. Macromol. 2015, 81, 730–735. [Google Scholar] [CrossRef] [PubMed]
- Zobel, H.F.; French, A.D.; Hinkle, M.E. X-ray diffraction of oriented amylose fibers. II. Structure of V amyloses. Biopolymers 1967, 5, 837–845. [Google Scholar] [CrossRef]
- Vamadevan, V.; Bertoft, E.; Seetharaman, K. On the importance of organization of glucan chains on thermal properties of starch. Carbohydr. Polym. 2013, 92, 1653–1659. [Google Scholar] [CrossRef] [PubMed]
- Vamadevan, V.; Blennow, A.; Buleon, A.; Goldstein, A.; Bertoft, E. Distinct Properties and Structures among B-Crystalline Starch Granules. Starch 2017, 70, 1700240. [Google Scholar] [CrossRef]
- Chauhan, F.; Seetharaman, K. On the organization of chains in amylopectin. Starch Stärke 2013, 191–199. [Google Scholar] [CrossRef]
- Shen, X.; Bertoft, E.; Zhang, G.; Hamaker, B.R. Iodine binding to explore the conformational state of internal chains of amylopectin. Carbohydr. Polym. 2013, 98, 778–783. [Google Scholar] [CrossRef]
- Hanashiro, I.; Matsugasako, J.-I.; Egashira, T.; Takeda, Y. Structural characterization of long unit-chains of amylopectin. J. Appl. Glycosci. 2005, 52, 233–237. [Google Scholar]
- Yoo, S.; Jane, J.-L. Structural and physical characteristics of waxy and other wheat starches. Carbohydr. Polym. 2002, 49, 297–305. [Google Scholar]
- Noda, T.; Takigawa, S.; Matsuura-Endo, C.; Kim, S. Physicochemical properties and amylopectin structures of large, small, and extremely small potato starch granules. Carbohydr. Polym. 2005, 60, 245–251. [Google Scholar]
- Laohaphatanaleart, K.; Piyachomkwan, K.; Sriroth, K.; Santisopasri, V.; Bertoft, E. A study of the internal structure in cassava and rice amylopectin. Starch/Stärke 2009, 61, 557–569. [Google Scholar] [CrossRef]
- Takeda, Y.; Hizukuri, S.; Takeda, C.; Suzuki, A. Structures of branched molecules of amyloses of various origins, and molar fractions of branched and unbranched molecules. Carbohydr. Res. 1987, 165, 139–145. [Google Scholar] [CrossRef]
- Wikman, J.; Blennow, A.; Bul, A.; Putaux, J.; Serge, P.; Seetharaman, K.; Bertoft, E. Influence of Amylopectin Structure and Degree of Phosphorylation on the Molecular Composition of Potato Starch Lintners. Biopolymers 2014, 101, 257–271. [Google Scholar] [CrossRef] [PubMed]
- Badenhuizen, N.P.; Dutton, R.W. Growth of 14C-labelled starch granules in potato tubers as revealed by autoradiographs. Protoplasma 1956, 47, 156–163. [Google Scholar]
- Ao, Z.; Jane, J. Characterization and modeling of the A- and B-granule starches of wheat, triticale, and barley. Carbohydr. Polym. 2007, 67, 46–55. [Google Scholar] [CrossRef]
- O’Sullivan, A.; Perez, S. The relationship between internal chain length of amylopectin and crystallinity in starch. Biopolymers 1999, 50, 381–390. [Google Scholar]
- Palmer, T.N.; Macaskie, L.F.; Grewh, K. The Unit-Chain Distribution Profiles of Branched (1-4)-alpha-D-Glucans. Carbohydr. Res. 1983, 114, 338–342. [Google Scholar]
- Vamadevan, V.; Bertoft, E.; Soldatov, D.V.; Seetharaman, K. Impact on molecular organization of amylopectin in starch granules upon annealing. Carbohydr. Polym. 2013, 98, 1045–1055. [Google Scholar] [CrossRef]
- Zhu, F.; Mojel, R.; Li, G. Structure of black pepper (Piper nigrum) starch. Food Hydrocoll. 2017, 71, 102–107. [Google Scholar] [CrossRef]
- Putaux, J.; Molina-Boisseau, S.; Momaur, T.; Dufresne, A. Platelet Nanocrystals Resulting from the Disruption of Waxy Maize Starch Granules by Acid Hydrolysis. Biomacromolecules 2003, 4, 1198–1202. [Google Scholar] [CrossRef]
- Spinozzi, F.; Ferrero, C.; Pérez, S. The architecture of starch blocklets follows phyllotaxy. Sci. Rep. 2020, in press. [Google Scholar] [CrossRef]
- Vamadevan, V.; Bertoft, E. Food Hydrocolloids Observations on the impact of amylopectin and amylose structure on the swelling of starch granules. Food Hydrocoll. 2020, 103, 105663. [Google Scholar] [CrossRef]
- Goren, A.; Ashlock, D.; Tetlow, I.J. Starch formation inside plastids of higher plants. Protoplasma 2018, 255, 1–22. [Google Scholar] [CrossRef] [PubMed]
- Pfister, B.; Zeeman, S.C. Formation of starch in plant cells. Cell. Mol. Life Sci. 2016, 73, 2781–2807. [Google Scholar] [CrossRef] [PubMed]
- Pfister, B.; Zeeman, S.C.; Rugen, M.D.; Field, R.A.; Ebenhöh, O.; Raguin, A. Theoretical and experimental approaches to understand the biosynthesis of starch granules in a physiological context. Photosynth. Res. 2020, 145, 55–70. [Google Scholar] [CrossRef] [PubMed]
- Kartal, Ö.; Ebenhöh, O. A generic rate law for surface-active enzymes. FEBS Lett. 2013, 587, 2882–2890. [Google Scholar] [CrossRef]
- Ghosh, H.; Preiss, J. Adenosine diphosphate glucose pyrophosphorylase: A regulatory enzyme in the biosynthesis of starch in spinach leaf chloroplasts. J. Biol. Chem. 1966, 241, 4491–4504. [Google Scholar]
- Clarke, B.R.; Denyer, K.; Jenner, C.F.; Smith, A.M. The relationship between the rate of starch synthesis, the adenosine 5’-diphosphoglucose concentration and the amylose content of starch in developing pea embryos. Planta 1999, 209, 324–329. [Google Scholar] [CrossRef]
- Whelan, W.J. The Initiation of Glycogen Synthesis. Bioessays 1968, 5, 136–140. [Google Scholar] [CrossRef]
- Smythe, C.; Cohen, P. The discovery of glycogenin and the priming mechanism for glycogen biogenesis. Eur. J. Biochem. 1991, 200, 625–631. [Google Scholar] [CrossRef]
- Zeqiraj, E.; Tang, X.; Hunter, R.W.; Garcia-Rocha, M.; Judd, A.; Deak, M.; von Wilamowitz-Moellendorff, A.; Kurinov, I.; Guinovart, J.J.; Tyers, M.; et al. Structural basis for the recruitment of glycogen synthase by glycogenin. Proc. Natl. Acad. Sci. USA 2014, 111, E2831–E2840. [Google Scholar] [CrossRef]
- Ugalde, J.E.; Parodi, A.J.; Ugalde, R.A. De novo synthesis of bacterial glycogen: Agrobacterium tumefaciens glycogen synthase is involved in glucan initiation and elongation. Proc. Natl. Acad. Sci. USA 2003, 100, 10659–10663. [Google Scholar] [PubMed]
- Seung, D.; Smith, A.M. Starch granule initiation and morphogenesis—Progress in Arabidopsis and cereals. J. Exp. Bot. 2019, 70, 771–784. [Google Scholar] [CrossRef] [PubMed]
- Malinova, I.; Qasim, H.M.; Brust, H.; Fettke, J. Parameters of Starch Granule Genesis in Chloroplasts of Arabidopsis Thaliana. Front. Plant Sci. 2018, 9, 1–7. [Google Scholar] [CrossRef]
- Zhu, F.; Bertoft, E.; Wang, Y.; Emes, M.; Tetlow, I.; Seetharaman, K. Structure of Arabidopsis leaf starch is markedly altered following nocturnal degradation. Carbohydr. Polym. 2015, 117, 1002–1013. [Google Scholar] [CrossRef] [PubMed]
- MacNeill, G.J.; Mehrpouyan, S.; Minow, M.A.A.; Patterson, J.A.; Tetlow, I.J.; Emes, M.J. Starch as a source, starch as a sink: The bifunctional role of starch in carbon allocation. J. Exp. Bot. 2017, 68, 4433–4453. [Google Scholar] [CrossRef]
- Feike, D.; Seung, D.; Graf, A.; Bischof, S.; Ellick, T.; Coiro, M.; Soyk, S.; Eicke, S.; Mettler-Altmann, T.; Lu, K.J.; et al. The Starch Granule-Associated Protein EARLY STARVATION1 Is Required for the Control of Starch Degradation in Arabidopsis thaliana Leaves. Plant Cell 2016, 28, 1472–1489. [Google Scholar] [CrossRef]
- Ziegler, G.R.; Creek, J.A.; Runt, J. Spherulitic crystallization in starch as a model for starch granule initiation. Biomacromolecules 2005, 6, 1547–1554. [Google Scholar] [CrossRef]
- Vamadevan, V.; Hoover, R.; Bertoft, E.; Seetharaman, K. Hydrothermal Treatment and Iodine Binding Provide Insights into the Organization of Glucan Chains Within the Semi-Crystalline Lamellae of Corn Starch Granules. Biopolymers 2014, 101, 871–885. [Google Scholar] [CrossRef]
- Lundquist, P.K.; Mantegazza, O.; Stefanski, A.; Stühler, K.; Weber, A.P.M. Surveying the Oligomeric State of Arabidopsis thaliana Chloroplasts. Mol. Plant. 2017, 10, 197–211. [Google Scholar] [CrossRef]
- Szydlowski, N.; Ragel, P.; Raynaud, S.; Lucas, M.M.; Roldan, I.; Montero, M.; Munoz, F.J.; Ovecka, M.; Bahaji, A.; Planchot, V.; et al. Starch Granule Initiation in Arabidopsis Requires the Presence of Either Class IV or Class III Starch Synthases. Plant Cell 2009, 21, 2443–2457. [Google Scholar] [CrossRef]
- Leterrier, M.; Holappa, L.D.; Broglie, K.E.; Beckles, D.M. Cloning, characterisation and comparative analysis of a starch synthase IV gene in wheat: Functional and evolutionary implications. BMC Plant. Biol. 2008, 8, 98. [Google Scholar] [CrossRef]
- Lohmeier-Vogel, E.M.; Kerk, D.; Nimick, M.; Wrobel, S.; Vickerman, L.; Muench, D.G.; Moorhead, G.B. Arabidopsis At5g39790 encodes a chloroplast-localized, carbohydrate- binding, coiled-coil domain-containing putative scaffold protein. BMC Plant. Biol. 2008, 8, 1–14. [Google Scholar] [CrossRef]
- Seung, D.; Boudet, J.; Monroe, J.; Schreier, T.B.; David, L.C.; Abt, M.; Lu, K.-J.; Zanella, M.; Zeeman, S.C. Homologs of PROTEIN TARGETING TO STARCH Control Starch Granule Initiation in Arabidopsis Leaves. Plant Cell 2017, 29, 1657–1677. [Google Scholar] [CrossRef] [PubMed]
- Janeček, Š.; Svensson, B.; MacGregor, E.A. Structural and evolutionary aspects of two families of non-catalytic domains present in starch and glycogen binding proteins from microbes, plants and animals. Enzym. Microb. Technol. 2011, 49, 429–440. [Google Scholar] [CrossRef]
- Raynaud, S.; Ragel, P.; Rojas, T.; Mérida, Á. The N-terminal part of Arabidopsis thaliana starch synthase 4 determines the localization and activity of the enzyme. J. Biol. Chem. 2016, 291, 10759–10771. [Google Scholar] [CrossRef] [PubMed]
- Cuesta-Seijo, J.A.; Ruzanski, C.; Krucewicz, K.; Meier, S.; Hägglund, P.; Svensson, B.; Palcic, M.M. Functional and structural characterization of plastidic starch phosphorylase during barley endosperm development. PLoS ONE 2017, 12, e0175488. [Google Scholar] [CrossRef]
- Malinova, I.; Mahlow, S.; Alseekh, S.; Orawetz, T.; Fernie, A.R.; Baumann, O.; Steup, M.; Fettke, J. Double Knockout Mutants of Arabidopsis Grown under Normal Conditions Reveal that the Plastidial Phosphorylase Isozyme Participates in Transitory Starch Metabolism. Plant Physiol. 2014, 289, 9223–9246. [Google Scholar]
- Malinova, I.; Alseekh, S.; Feil, R.; Fernie, A.R.; Baumann, O.; Schöttler, M.A.; Lunn, J.E.; Fettke, J. Starch Synthase 4 and Plastidal Phosphorylase Differentially Affect Starch Granule Number and Morphology. Plant Physiol. 2017, 174, 73–85. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, Y.; Ono, M.; Sawada, T.; Crofts, N.; Fujita, N.; Steup, M. Characterization of the functional interactions of plastidial starch phosphorylase and starch branching enzymes from rice endosperm during reserve starch biosynthesis. Plant Sci. 2017, 264, 83–95. [Google Scholar] [CrossRef] [PubMed]
- Hwang, S.K.; Koper, K.; Satoh, H.; Okita, T.W. Rice endosperm starch phosphorylase (Pho1) assembles with disproportionating enzyme (Dpe1) to form a protein complex that enhances synthesis of malto-oligosaccharides. J. Biol. Chem. 2016, 291, 19994–20007. [Google Scholar] [CrossRef] [PubMed]
- Bresolin, N.S.; Li, Z.; Kosar-Hashemi, B.; Tetlow, I.J.; Chatterjee, M.; Rahman, S.; Morell, M.K.; Howitt, C.A. Characterisation of disproportionating enzyme from wheat endosperm. Planta 2006, 224, 20–31. [Google Scholar] [CrossRef] [PubMed]
- Vandromme, C.; Spriet, C.; Dauvillée, D.; Courseaux, A.; Putaux, J.L.; Wychowski, A.; Krzewinski, F.; Facon, M.; D’Hulst, C.; Wattebled, F. PII1: A protein involved in starch initiation that determines granule number and size in Arabidopsis chloroplast. New Phytol. 2019, 221, 356–370. [Google Scholar] [CrossRef] [PubMed]
- Seung, D.; Schreier, T.B.; Bürgy, L.; Eicke, S.; Zeeman, S.C. Two plastidial coiled-coil proteins are essential for normal starch granule initiation in Arabidopsis. Plant Cell 2018, 30, 1523–1542. [Google Scholar] [CrossRef] [PubMed]
- Abt, M.R.; Pfister, B.; Sharma, M.; Eicke, S.; Bürgy, L.; Neale, I.; Seung, D.; Zeeman, S. STARCH SYNTHASE 5, a Noncanonical Starch Synthase-Like Protein, Promotes Starch Granule Initiation in Arabidopsis. Plant Cell 2020. [Google Scholar] [CrossRef]
- Gidley, M.; Bulpin, P. Crystallisation of malto-oligosaccharides as models of the crystalline forms of starch: Minimum chain-length requirement for the formation of double helices. Carbohydr. Res. 1987, 161, 291–300. [Google Scholar] [CrossRef]
- Puteaux, J.; Potocki-Veronese, G.; Remaud-Simeon, M.; Buléon, A. Alpha-D-glucan-based dendritic nanoparticles prepared by in vitro enzymatic chain extension of glycogen. Biomacromolecules 2006, 7, 1720–1728. [Google Scholar] [CrossRef]
- Nakamura, Y.; Ono, M.; Utsumi, C.; Steup, M. Functional interaction between plastidial starch phosphorylase and starch branching enzymes from rice during the synthesis of branched maltodextrins. Plant Cell Physiol. 2012, 53, 869–878. [Google Scholar] [CrossRef]
- Shure, M.; Wessler, S.; Fedoroff, N. Molecular identification and isolation of the Waxy locus in maize. Cell 1983, 35, 225–233. [Google Scholar] [CrossRef]
- Ball, S.G.; Van De Wal, M.H.B.J.; Visser, R.G.F. Progress in understanding the biosynthesis of amylose. Trends Plant Sci. 1998, 3, 462–467. [Google Scholar] [CrossRef]
- Denyer, K.; Waite, D.; Motawia, S.; Møller, B.L.; Smith, A.M. Granule-bound starch synthase I in isolated starch granules elongates malto-oligosaccharides processively. Biochem. J. 1999, 340, 183–191. [Google Scholar] [CrossRef]
- Tsai, C.Y.; Salamini, F.; Nelson, O.E. Enzymes of Carbohydrate Metabolism in the Developing Endosperm of Maize. Plant Physiol. 1970, 46, 299–306. [Google Scholar] [CrossRef] [PubMed]
- Geddes, R.; Greenwood, C.T.; Mackenzie, S. Studies on the Biosynthesis of Starch Granules. Carbohydr. Res. 1965, 1, 71–82. [Google Scholar]
- Seung, D. Amylose in starch: Towards an understanding of biosynthesis, structure and function. New Phytol. 2020, in press. [Google Scholar]
- Denyer, K.; Hylton, C.M.; Jenner, C.F.; Smith, A.M. Identification of multiple isoforms of soluble and granule-bound starch synthase in developing wheat endosperm. Planta 1995, 196, 256–265. [Google Scholar] [CrossRef]
- Vrinten, P.L.; Nakamura, T. Wheat granule-bound starch synthase I and II are encoded by separate genes that are expressed in different tissues. Plant Physiol. 2000, 122, 255–264. [Google Scholar] [CrossRef]
- Edwards, A.; Vincken, J.; Suurs, L.C.J.M.; Visser, R.G.F.; Zeeman, S.; Smith, A.; Martin, C. Discrete Forms of Amylose Are Synthesized by Isoforms of GBSSI in Pea. Plant Cell 2002, 14, 1767–1785. [Google Scholar] [CrossRef]
- Denyer, K.; Clarke, B.; Hylton, C.; Tatge, H.; Smith, A.M. The elongation of amylose and amylopectin chains in isolated starch granules. Plant J. 1996, 10, 1135–1143. [Google Scholar]
- Zeeman, S.C.; Smith, S.M.; Smith, A.M. The Priming of Amylose Synthesis in Arabidopsis Leaves 1. Plant Physiol. 2002, 128, 1069–1076. [Google Scholar] [CrossRef]
- Grimaud, F.; Rogniaux, H.; James, M.G.; Myers, A.M.; Planchot, V. Proteome and phosphoproteome analysis of starch granule-associated proteins from normal maize and mutants affected in starch biosynthesis. J. Exp. Bot. 2008, 59, 3395–3406. [Google Scholar] [CrossRef]
- Liu, F.; Makhmoudova, A.; Lee, E.A.; Wait, R.; Emes, M.J.; Tetlow, I.J. The amylose extender mutant of maize conditions novel protein-protein interactions between starch biosynthetic enzymes in amyloplasts. J. Exp. Bot. 2009, 60, 4423–4440. [Google Scholar] [CrossRef]
- Ahmed, Z.; Tetlow, I.J.; Ahmed, R.; Morell, M.K.; Emes, M.J. Protein-protein interactions among enzymes of starch biosynthesis in high-amylose barley genotypes reveal differential roles of heteromeric enzyme complexes in the synthesis of A and B granules. Plant Sci. 2015, 233, 95–106. [Google Scholar] [CrossRef] [PubMed]
- Liu, D.R.; Huang, W.X.; Cai, X.L. Oligomerization of rice granule-bound starch synthase 1 modulates its activity regulation. Plant Sci. 2013, 210, 141–150. [Google Scholar] [CrossRef] [PubMed]
- Seung, D.; Soyk, S.; Coiro, M.; Maier, B.A.; Eicke, S.; Zeeman, S.C. Protein Targeting to Starch Is Required for Localising Granule-Bound Starch Synthase to Starch Granules and for Normal Amylose Synthesis in Arabidopsis. PLoS Biol. 2015, 13, e1002080. [Google Scholar] [CrossRef] [PubMed]
- Zhong, Y.; Blennow, A.; Kofoed-Enevoldsen, O.; Jiang, D.; Hebelstrup, K. Protein Targeting to Starch 1 is essential for starchy endosperm development in barley. J. Exp. Bot. 2019, 70, 485–496. [Google Scholar] [CrossRef]
- Wang, W.; Wei, X.; Jiao, G.; Chen, W.; Wu, Y.; Sheng, Z.; Hu, S.; Xie, L.; Wang, J.; Tang, S.; et al. GBSS-BINDING PROTEIN, encoding a CBM48 domain-containing protein, affects rice quality and yield. J. Integr. Plant Biol. 2020, 62, 948–966. [Google Scholar] [CrossRef]
- Guan, H.P.; Preiss, J. Differentiation of the Properties of the Branching Isozymes from Maize (Zea mays). Plant Physiol. 1993, 102, 1269–1273. [Google Scholar] [CrossRef]
- Kuriki, T.; Stewart, D.C.; Chem, J.B.; Kuriki, T.; Stewart, D.C.; Preiss, J. Construction of chimeric enzymes out of maize endosperm branching enzymes I and II: Activity and properties. J. Biol. Chem. 1997, 272, 28999–29004. [Google Scholar] [CrossRef]
- Tester, R.; Morrison, W. Swelling and gelatinization of cereal starches. I. Effects of amylopectin, amylose, and lipids. Cereal Chem. 1990, 67, 551–557. [Google Scholar]
- Recondo, E.; Leloir, L.F. Adenosine diphosphate glucose and starch synthesis. Biochem. Biophys. Res. Commun. 1961, 6, 85–88. [Google Scholar] [CrossRef]
- Cuesta-Seijo, J.A.; Nielsen, M.M.; Ruzanski, C.; Krucewicz, K.; Beeren, S.R.; Rydhal, M.G.; Yoshimura, Y.; Striebeck, A.; Motawia, M.S.; Willats, W.G.T.; et al. In vitro biochemical characterization of all barley endosperm starch synthases. Front. Plant Sci. 2016, 6, 1–17. [Google Scholar] [CrossRef]
- Larson, M.E.; Falconer, D.J.; Myers, A.M.; Barb, A.W. Direct characterization of the maize starch synthase IIa product shows maltodextrin elongation occurs at the non-reducing end. J. Biol. Chem. 2016, 291, 24951–24960. [Google Scholar] [CrossRef] [PubMed]
- Wilkens, C.; Cuesta-Seijo, J.; Palcic, M.; Svensson, B. Selectivity of the surface binding site (SBS) on barley starch synthase I. Biologia (Bratisl.) 2014, 69, 1118–1121. [Google Scholar] [CrossRef]
- Commuri, P.D.; Keeling, P.L. Chain-length specificities of maize starch synthase I enzyme: Studies of glucan affinity and catalytic properties. Plant J. 2001, 25, 475–486. [Google Scholar] [CrossRef] [PubMed]
- Liu, F.; Romanova, N.; Lee, E.A.; Ahmed, R.; Evans, M.; Gilbert, E.P.; Morell, M.K.; Emes, M.J.; Tetlow, I.J. Glucan affinity of starch synthase IIa determines binding of starch synthase I and starch-branching enzyme IIb to starch granules. Biochem. J. 2012, 448, 373–387. [Google Scholar] [CrossRef] [PubMed]
- Brust, H.; Lehmann, T.; D’Hulst, C.; Fettke, J. Analysis of the functional interaction of Arabidopsis starch synthase and branching enzyme isoforms reveals that the cooperative action of SSI and BEs results in glucans with polymodal chain length distribution similar to amylopectin. PLoS ONE 2014, 9, e102364. [Google Scholar] [CrossRef]
- Tyynela, J.; Schulman, A.H. An analysis of soluble starch synthase isozymes from the developing grains of normal and shx cv. Bomi barley (Hordeum vulgare). Physiol. Plant 1993, 89, 835–841. [Google Scholar] [CrossRef]
- Tetlow, I.J.; Beisel, K.G.; Cameron, S.; Makhmoudova, A.; Liu, F.; Bresolin, N.S.; Wait, R.; Morell, M.K.; Emes, M.J. Analysis of Protein Complexes in Wheat Amyloplasts Reveals Functional Interactions among Starch Biosynthetic Enzymes. Plant Physiol. 2008, 146, 1878–1891. [Google Scholar] [CrossRef]
- James, M.G.; Denyer, K.; Myers, A.M. Starch synthesis in the cereal endosperm. Curr. Opin. Plant. Biol. 2003, 6, 215–222. [Google Scholar] [CrossRef]
- Cuesta-Seijo, J.A.; Porcellinis, A.J.D.; Valente, A.H.; Striebeck, A.; Voss, C.; Marri, L.; Hansson, A.; Jansson, A.M.; Dinesen, M.H.; Fangel, J.U.; et al. Amylopectin Chain Length Dynamics and Activity Signatures of Key Carbon Metabolic Enzymes Highlight Early Maturation as Culprit for Yield Reduction of Barley Endosperm Starch after Heat Stress. Plant Cell Physiol. 2019, 60, 2692–2706. [Google Scholar] [CrossRef]
- Drummond, G.S.; Smith, E.E.; Whelan, W.J. Purification and Properties of Potato: α-1,4-Glucan 6-Glycosyltransferase (Q-Enzyme). Eur. J. Biochem. 1972, 26, 168–176. [Google Scholar] [CrossRef]
- Borovsky, D.; Smith, E.E.; Whelan, W.J.; French, D.; Kikumoto, S. The mechanism of Q-enzyme action and its influence on the structure of amylopectin. Arch. Biochem. Biophys. 1979, 198, 627–631. [Google Scholar] [CrossRef]
- Borovsky, D.; Smith, E.E.; Whelan, W.J. On The Mechanism of Amylose Branching by Potato Q-Enzyme. Eur. J. Biochem. 1976, 62, 307–312. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Hu, P.; Lin, L.; Chen, Z.; Liu, Q.; Wei, C. Gradually decreasing starch branching enzyme expression is responsible for the formation of heterogenous starch granules. Plant Physiol. 2018, 176, 582–595. [Google Scholar] [CrossRef]
- Larsson, C.T.; Hofvander, P.; Khoshnoodi, J.; Ek, B.; Rask, L.; Larsson, H. Three isoforms of starch synthase and two isoforms of branching enzyme are present in potato tuber starch. Plant Sci. 1996, 117, 9–16. [Google Scholar] [CrossRef]
- Rydberg, U.; Andersson, L.; Andersson, R. Comparison of starch branching enzyme I and II from potato. FEBS J. 2001, 268, 6140–6145. [Google Scholar] [CrossRef]
- Jobling, S.A.; Schwall, G.P.; Westcott, R.J.; Sidebottom, C.M.; Debet, M.; Gidley, M.J.; Jeffcoat, R.; Safford, R. A minor form of starch branching enzyme in potato (Solanum tuberosum L.) tubers has a major effect on starch structure: Cloning and characterisation of multiple forms of SBE A. Plant J. 1999, 18, 163–171. [Google Scholar] [CrossRef]
- Wolfe, K.H.; Gouy, M.; Yang, Y.W.; Sharp, P.M.; Li, W.H. Date of the monocot-dicot divergence estimated from chloroplast DNA sequence data. Proc. Natl. Acad. Sci. USA 1989, 86, 6201–6205. [Google Scholar] [CrossRef]
- Fisher, D.K.; Gao, M.; Kim, K.N.; Boyer, C.D.; Guiltinan, M.J. Allelic Analysis of the Maize amylose-extender Locus Suggests That Independent Genes Encode Starch-Branching Enzymes IIa and IIb. Plant Physiol. 1996, 110, 611–619. [Google Scholar] [CrossRef]
- Tetlow, I.J.; Emes, M.J. A review of starch-branching enzymes and their role in amylopectin biosynthesis. Iubmb Life 2014, 66, 546–558. [Google Scholar] [CrossRef]
- Takeda, Y.; Guan, H.P.; Preiss, J. Branching of amylose by the branching isoenzymes of maize endosperm. Carbohydr. Res. 1993, 240, 253–263. [Google Scholar] [CrossRef]
- Regina, A.; Kosar-Hashemi, B.; Li, Z.; Pedler, A.; Mukai, Y.; Yamamoto, M.; Gale, K.; Sharp, P.J.; Morell, M.K.; Rahman, S. Starch branching enzyme IIb in wheat is expressed at low levels in the endosperm compared to other cereals and encoded at a non-syntenic locus. Planta 2005, 222, 899–909. [Google Scholar] [CrossRef] [PubMed]
- Gao, M.; Fisher, D.K.; Kim, K.N.; Shannon, J.C.; Guiltinan, M.J. Independent genetic control of maize starch-branching enzymes IIa and IIb. Isolation and characterization of a Sbe2a cDNA. Plant Physiol. 1997, 114, 69–78. [Google Scholar] [CrossRef] [PubMed]
- Sawada, T.; Nakamura, Y.; Ohdan, T.; Saitoh, A.; Francisco, P.B.; Suzuki, E.; Fujita, N.; Shimonaga, T.; Fujiwara, S.; Tsuzuki, M.; et al. Diversity of reaction characteristics of glucan branching enzymes and the fine structure of α-glucan from various sources. Arch. Biochem. Biophys. 2014, 562, 9–21. [Google Scholar] [CrossRef]
- Blauth, S.L.; Kim, K.N.; Klucinec, J.; Shannon, J.C.; Thompson, D.; Guiltinan, M. Identification of mutator insertional mutants of starch-branching enzyme 1 (sbe1) in Zea mays L. Plant. Mol. Biol. 2002, 48, 287–297. [Google Scholar] [CrossRef]
- Regina, A.; Kosar-Hashemi, B.; Li, Z.; Rampling, L.; Cmiel, M.; Gianibelli, M.C.; Konik-Rose, C.; Larroque, O.; Rahman, S.; Morell, M.K. Multiple isoforms of starch branching enzyme-I in wheat: Lack of the major SBE-I isoform does not alter starch phenotype. Funct. Plant. Biol. 2004, 31, 591–601. [Google Scholar] [CrossRef] [PubMed]
- Satoh, H.; Nishi, A.; Yamashita, K.; Takemoto, Y.; Tanaka, Y.; Hosaka, Y.; Sakurai, A.; Fujita, N.; Nakamura, Y. Starch-branching enzyme I-deficient mutation specifically affects the structure and properties of starch in rice endosperm. Plant Physiol. 2003, 133, 1111–1121. [Google Scholar] [CrossRef]
- Xia, H.; Yandeau-Nelson, M.; Thompson, D.B.; Guiltinan, M.J. Deficiency of maize starch-branching enzyme I results in altered starch fine structure, decreased digestibility and reduced coleoptile growth during germination. BMC Plant. Biol. 2011, 11, 95. [Google Scholar] [CrossRef]
- Tunçay, H.; Findinier, J.; Duchêne, T.; Cogez, V.; Cousin, C.; Peltier, G.; Ball, S.G.; Dauvillée, D. A Forward Genetic Approach in Chlamydomonas reinhardtii as a Strategy for Exploring Starch Catabolism. PLoS ONE 2013, 8, e74763. [Google Scholar] [CrossRef]
- Dumez, S.; Wattebled, F.; Dauvillee, D.; Delvalle, D.; Planchot, V.; Ball, S.G.; D’Hulst, C. Mutants of Arabidopsis Lacking Starch Branching Enzyme II Substitute Plastidial Starch Synthesis by Cytoplasmic Maltose Accumulation. Plant Cell Online 2006, 18, 2694–2709. [Google Scholar] [CrossRef]
- Regina, A.; Bird, A.; Topping, D.; Bowden, S.; Freeman, J.; Barsby, T.; Kosar-Hashemi, B.; Li, Z.; Rahman, S.; Morell, M. High-amylose wheat generated by RNA interference improves indices of large-bowel health in rats. Proc. Natl. Acad. Sci. USA 2006, 103, 3546–3551. [Google Scholar] [CrossRef]
- Nishi, A.; Nakamura, Y.; Tanaka, N.; Satoh, H. Biochemical and genetic analysis of the effects of amylose-extender mutation in rice endosperm. Plant Physiol. 2001, 127, 459–472. [Google Scholar] [CrossRef]
- Boyer, C.D.; Preiss, J. Evidence for independent genetic control of the multiple forms of maize endosperm branching enzymes and starch synthases. Plant Physiol. 1981, 67, 1141–1145. [Google Scholar] [CrossRef] [PubMed]
- Banks, W.; Greenwood, C.T.; Muir, D. Studies on starches of high amylose-content. Die Stärke 1974, 9, 289–328. [Google Scholar] [CrossRef]
- Butardo, V.M.; Fitzgerald, M.A.; Bird, A.R.; Gidley, M.J.; Flanagan, B.M.; Larroque, O.; Resurreccion, A.P.; Laidlaw, H.K.C.; Jobling, S.A.; Morell, M.K.; et al. Impact of down-regulation of starch branching enzyme IIb in rice by artificial microRNA- and hairpin RNA-mediated RNA silencing. J. Exp. Bot. 2011, 62, 4927–4941. [Google Scholar] [CrossRef]
- Schwall, G.P.; Safford, R.; Westcott, R.J.; Jeffcoat, R.; Tayal, A.; Shi, Y.C.; Gidley, M.J.; Jobling, S.A. Production of very-high-amylose potato starch by inhibition of SBE A and B. Nat. Biotechnol. 2000, 18, 551–554. [Google Scholar] [CrossRef] [PubMed]
- Brummell, D.A.; Watson, L.M.; Zhou, J.; McKenzie, M.J.; Hallett, I.C.; Simmons, L.; Carpenter, M.; Timmerman-Vaughan, G.M. Overexpression of starch branching enzyme II increases short-chain branching of amylopectin and alters the physicochemical properties of starch from potato tuber. BMC Biotechnol. 2015, 15, 28. [Google Scholar] [CrossRef]
- Tanaka, N.; Fujita, N.; Nishi, A.; Satoh, H.; Hosaka, Y.; Ugaki, M.; Kawasaki, S.; Nakamura, Y. The structure of starch can be manipulated by changing the expression levels of starch branching enzyme IIb in rice endosperm. Plant Biotechnol. J. 2004, 2, 507–516. [Google Scholar] [CrossRef] [PubMed]
- Cenci, U.; Chabi, M.; Ducatez, M.; Tirtiaux, C.; Nirmal-Raj, J.; Utsumi, Y.; Kobayashi, D.; Sasaki, S.; Suzuki, E.; Nakamura, Y.; et al. Convergent Evolution of Polysaccharide Debranching Defines a Common Mechanism for Starch Accumulation in Cyanobacteria and Plants. Plant Cell 2013, 25, 3961–3975. [Google Scholar] [CrossRef]
- Cenci, U.; Nitschke, F.; Steup, M.; Minassian, B.A.; Colleoni, C.; Ball, S.G. Transition from glycogen to starch metabolism in Archaeplastida. Trends Plant Sci. 2014, 19, 18–28. [Google Scholar] [CrossRef]
- Møller, M.S.; Henriksen, A.; Svensson, B. Structure and function of α-glucan debranching enzymes. Cell Mol. Life Sci. 2016, 73, 2619–2641. [Google Scholar] [CrossRef]
- Mouille, G.; Maddelein, M.-L.; Libessart, N.; Talaga, P.; Decq, A.; Delrue, B.; Ball, S. Preamylopectin Processing: A Mandatory Step for Starch Biosynthesis in Plants. Plant Cell 1996, 8, 1353–1366. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, T.H.; Baunsgaard, L.; Blennow, A. Intermediary glucan structures formed during starch granule biosynthesis are enriched in short side chains, a dynamic pulse labeling approach. J. Biol. Chem. 2002, 277, 20249–20255. [Google Scholar] [CrossRef] [PubMed]
- Wattebled, F.; Planchot, V.; Dong, Y.; Szydlowski, N.; Pontoire, B.; Devin, A.; Ball, S.; D’Hulst, C. Further Evidence for the Mandatory Nature of Polysaccharide Debranching for the Aggregation of Semicrystalline Starch and for Overlapping Functions of Debranching Enzymes in Arabidopsis Leaves. Plant Physiol. 2008, 148, 1309–1323. [Google Scholar] [CrossRef] [PubMed]
- Dinges, J.R.; Colleoni, C.; James, M.G.; Myers, A.M. Mutational analysis of the pullulanase-type debranching enzyme of maize indicates multiple functions in starch metabolism. Plant Cell 2003, 15, 666–680. [Google Scholar] [CrossRef] [PubMed]
- Yun, M.S.; Umemoto, T.; Kawagoe, Y. Rice debranching enzyme isoamylase 3 facilitates starch metabolism and affects plastid morphogenesis. Plant Cell Physiol. 2011, 52, 1068–1082. [Google Scholar] [CrossRef]
- Li, E.; Hasjim, J.; Gilding, E.K.; Godwin, I.D.; Li, C.G.R. The role of pullulanase in starch biosynthesis, structure, and thermal properties by studying sorghum with increased pullulanase activity. Starch 2019, 71, 1900072. [Google Scholar] [CrossRef]
- Streb, S.; Delatte, T.; Umhang, M.; Eicke, S.; Schorderet, M.; Reinhardt, D.; Zeeman, S.C. Starch granule biosynthesis in Arabidopsis is abolished by removal of all debranching enzymes but restored by the subsequent removal of an endoamylase. Plant Cell 2008, 20, 3448–3466. [Google Scholar] [CrossRef]
- Fujita, N.; Toyosawa, Y.; Utsumi, Y.; Higuchi, T.; Hanashiro, I.; Ikegami, A.; Akuzawa, S.; Yoshida, M.; Mori, A.; Inomata, K.; et al. Characterization of pullulanase (PUL)-deficient mutants of rice (Oryza sativa L.) and the function of PUL on starch biosynthesis in the developing rice endosperm. J. Exp. Bot. 2009, 60, 1009–1023. [Google Scholar] [CrossRef]
- James, M.G.; Robertson, D.S.; Myers, A.M. Characterization of the maize gene sugary1, a determinant of starch composition in kernels. Plant Cell 1995, 7, 417–429. [Google Scholar]
- Morris, D.; Morris, C. Glycogen in the seed of Zea mays. J. Biol. Chem. 1939, 130, 535–544. [Google Scholar]
- Dauvillée, D.; Colleoni, C.; Mouille, G.; Morell, M.; d’Hulst, C.; Wattebled, F.; Liénard, L.; Delvallé, D.; Ral, J.-P.; Myers, A.; et al. Biochemical characterization of wild-type and mutant isoamylases of Chlamydomonas reinhardtii supports a function of the multimeric enzyme organization in amylopectin maturation. Plant Physiol. 2001, 125, 1723–1731. [Google Scholar] [CrossRef] [PubMed]
- Sim, L.; Beeren, S.R.; Findinier, J.; Dauville, D.; Ball, S.G.; Henriksen, A.; Palcic, M.M. Crystal structure of the Chlamydomonas starch debranching enzyme isoamylase ISA1 reveals insights into the mechanism of branch trimming and complex assembly. J. Biol. Chem. 2014, 289, 22991–23003. [Google Scholar] [CrossRef]
- Inouchi, N.; Glover, D.V.; Fuwa, H. Chain Length Distribution of Amylopectins of Several Single Mutants and the Normal Counterpart, and Sugary 1 Phytoglycogen in Maize (Zea mays L.). Starch Stärke 1987, 39, 259–266. [Google Scholar] [CrossRef]
- Boyer, C.D.; Liu, K.-C. Starch and water-soluble polysaccharides from sugary endosperm of sorghum. Phytochemistry 1983, 22, 2513–2515. [Google Scholar] [CrossRef]
- Blennow, A.; Engelsen, S.B. Helix-breaking news: Fighting crystalline starch energy deposits in the cell. Trends Plant Sci. 2010, 15, 236–240. [Google Scholar] [CrossRef] [PubMed]
- Baunsgaard, L.; Lütken, H.; Mikkelsen, R.; Glaring, M.A.; Pham, T.T.; Blennow, A. A novel isoform of glucan, water dikinase phosphorylates pre-phosphorylated α-glucans and is involved in starch degradation in Arabidopsis. Plant J. 2005, 41, 595–605. [Google Scholar] [CrossRef]
- Xu, X.; Dees, D.; Dechesne, A.; Huang, X.F.; Visser, R.G.F.; Trindade, L.M. Starch phosphorylation plays an important role in starch biosynthesis. Carbohydr. Polym. 2017, 157, 1628–1637. [Google Scholar] [CrossRef]
- Ritte, G.; Heydenreich, M.; Mahlow, S.; Haebel, S.; Kötting, O.; Steup, M. Phosphorylation of C6- and C3-positions of glucosyl residues in starch is catalysed by distinct dikinases. FEBS Lett. 2006, 580, 4872–4876. [Google Scholar] [CrossRef]
- Wikman, J.; Larsen, F.; Motawia, M.; Blennow, A.; Bertoft, E. Phosphate esters in amylopectin clusters of potato tuber starch. Int. J. Biol. Macromol. 2011, 48, 639–649. [Google Scholar] [CrossRef]
- Lloyd, J.R.; Kossmann, J. Transitory and storage starch metabolism: Two sides of the same coin? Curr. Opin. Biotechnol. 2015, 32, 143–148. [Google Scholar] [CrossRef]
- Critchley, J.H.; Zeeman, S.C.; Takaha, T.; Smith, A.M.; Smith, S.M. A critical role for disproportionating enzyme in starch breakdown is revealed by a knock-out mutation in Arabidopsis. Plant J. 2001, 26, 89–100. [Google Scholar] [CrossRef] [PubMed]
- Dong, X.; Zhang, D.; Liu, J.; Liu, Q.Q.; Liu, H.; Tian, L.; Jiang, L.; Qu, L.Q. Plastidial Disproportionating Enzyme Participates in Starch Synthesis in Rice Endosperm by Transferring Maltooligosyl Groups from Amylose and Amylopectin to Amylopectin. Plant Physiol 2015, 169, 2496–2512. [Google Scholar] [CrossRef] [PubMed]
- Satoh, H.; Shibahara, K.; Tokunaga, T.; Nishi, A.; Tasaki, M.; Hwang, S.-K.; Okita, T.W.; Kaneko, N.; Fujita, N.; Yoshida, M.; et al. Mutation of the plastidial α-glucan phosphorylase gene in rice affects the synthesis and structure of starch in the endosperm. Plant Cell 2008, 20, 1833–1849. [Google Scholar] [CrossRef] [PubMed]
- Higgins, J.E.; Kosar-Hashemi, B.; Li, Z.; Howitt, C.A.; Larroque, O.; Flanagan, B.; Morell, M.K.; Rahman, S. Characterization of starch phosphorylases in barley grains. J. Sci. Food Agric. 2013, 93, 2137–2145. [Google Scholar] [CrossRef] [PubMed]
- Dauvillée, D.; Chochois, V.; Steup, M.; Haebel, S.; Eckermann, N.; Ritte, G.; Ral, J.P.; Colleoni, C.; Hicks, G.; Wattebled, F.; et al. Plastidial phosphorylase is required for normal starch synthesis in Chlamydomonas reinhardtii. Plant J. 2006, 48, 274–285. [Google Scholar] [CrossRef]
- Hwang, S.K.; Nishi, A.; Satoh, H.; Okita, T.W. Rice endosperm-specific plastidial α-glucan phosphorylase is important for synthesis of short-chain malto-oligosaccharides. Arch. Biochem. Biophys. 2010, 495, 82–92. [Google Scholar] [CrossRef]
- Colleoni, C.; Dauvill e, D.; Mouille, G.; Morell, M.; Samuel, M.; Slomiany, M.C.; Li nard, L.; Wattebled, F.; d’Hulst, C.; Ball, S. Biochemical characterization of the Chlamydomonas reinhardtii alpha-1,4 glucanotransferase supports a direct function in amylopectin biosynthesis. Plant Physiol. 1999, 120, 1005–1014. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Fernie, A.R. Metabolons, Enzyme-Enzyme Assemblies that Mediate Substrate Channeling, and Their Roles in Plant Metabolism. Plant. Commun. 2020, 1, 100081. [Google Scholar] [CrossRef]
- Liu, F.; Ahmed, Z.; Lee, E.A.; Donner, E.; Liu, Q.; Ahmed, R.; Morell, M.K.; Emes, M.J.; Tetlow, I.J. Allelic variants of the amylose extender mutation of maize demonstrate phenotypic variation in starch structure resulting from modified protein-protein interactions. J. Exp. Bot. 2012, 63, 1167–1183. [Google Scholar] [CrossRef]
- Hennen-Bierwagen, T.A.; Liu, F.; Marsh, R.S.; Kim, S.; Gan, Q.; Tetlow, I.J.; Emes, M.J.; James, M.G.; Myers, A.M. Starch biosynthetic enzymes from developing maize endosperm associate in multisubunit complexes. Plant Physiol. 2008, 146, 1892–1908. [Google Scholar] [CrossRef]
- Hennen-Bierwagen, T.A.; Lin, Q.; Grimaud, F.; Planchot, V.; Keeling, P.L.; James, M.G.; Myers, A.M. Proteins from multiple metabolic pathways associate with starch biosynthetic enzymes in high molecular weight complexes: A model for regulation of carbon allocation in maize amyloplasts. Plant Physiol. 2009, 149, 1541–1559. [Google Scholar] [CrossRef] [PubMed]
- Tetlow, I.; Liu, F.; Emes, M. Protein-protein interactions during starch biosynthesis. In Starch Metabolism and Structure; Springer: New York, NY, USA, 2015; pp. 291–313. [Google Scholar]
- Hayashi, M.; Crofts, N.; Oitome, N.F.; Fujita, N. Analyses of starch biosynthetic protein complexes and starch properties from developing mutant rice seeds with minimal starch synthase activities. BMC Plant Biol. 2018, 18, 59. [Google Scholar] [CrossRef] [PubMed]


| Structural Feature | Function/Role | Enzyme/Proteins Involved |
|---|---|---|
Hilum | Initiation zone of granule growth facilitating formation of water-insoluble polyglucan | SSIII, SSIV, SSV, PTST2, PTST3, PII1/MRC (SP and DPE; provision of MOS) |
Backbone | Linear α-(1,4)-linked glucan scaffolds from which building blocks are subtended | SSIII and/or SSIII protein complexes |
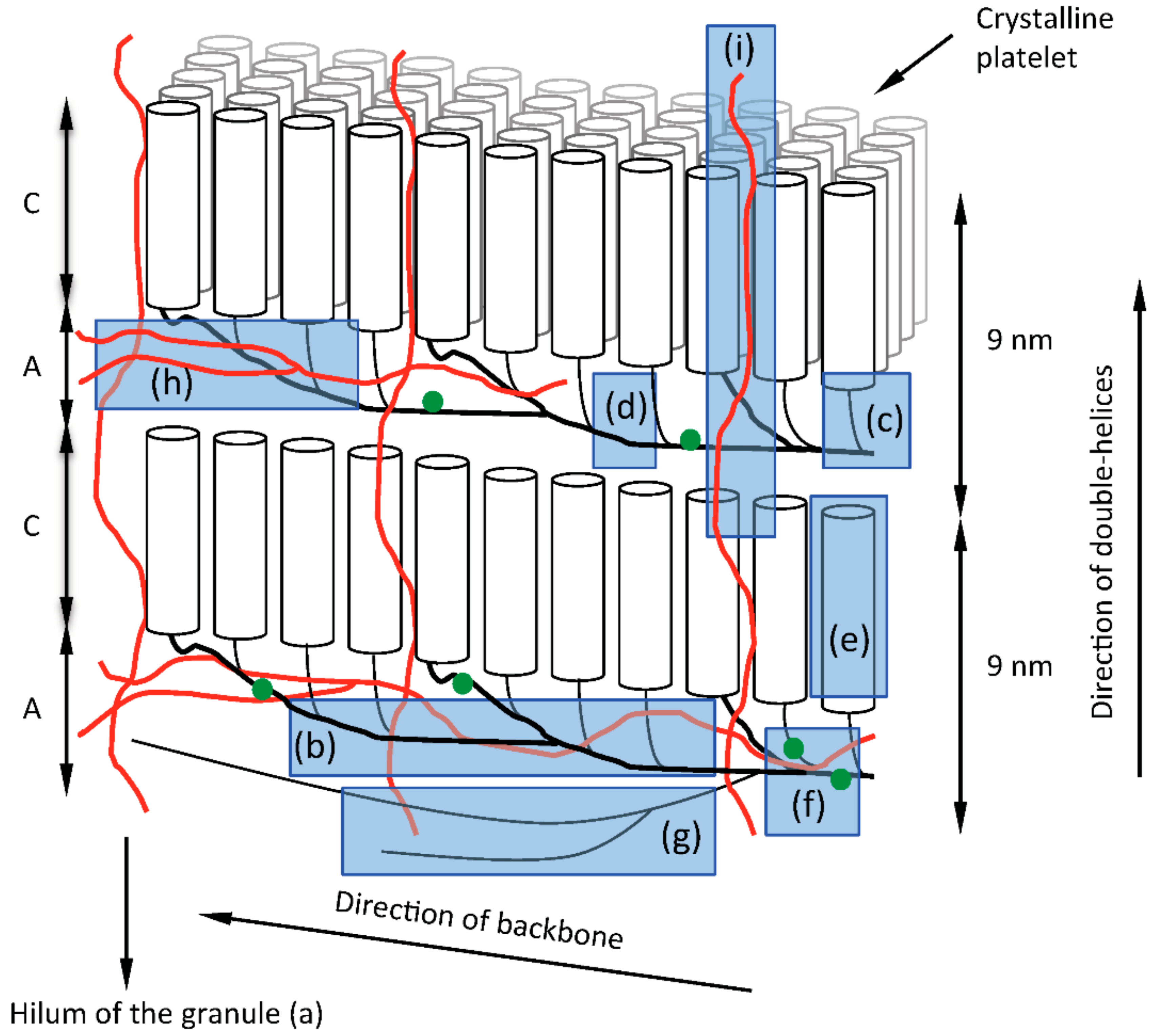
Building Blocks Short External Segments  | Major structural component of amylopectin. The framework from which short/intermediate α-glucan chains are produced which form water-excluding helical coils. Major structure responsible for the 9 nm repeat observed in coiled stacks of building blocks (blocklets?) | SSI, SSII, SBEII. The building block results from the combined actions of SS and SBE, likely SSI and/or SSII and SBEII isoforms. Further extension of the chains into short external segments is likely catalyzed by SSII. SSI, SSII and SBEII form protein complexes whose catalytic activity may be responsible for these building blocks and the subsequent short/intermediate chains pairs of which form the helical coils shown as cylinders. (DBE activity allows short/intermediate chains arising from building blocks to form water-insoluble structures via removal of selected α-(1,6)-branch linkages) |
Interblock Segments | Provide spacing between building blocks. Such spacing may influence larger scale packing of groups of building blocks possibly determining A- or B-type structural allomorphs as viewed by X-ray scattering. | SSIII and/or SSIII protein complexes (DBEactivity to reduce excess branching between building blocks) |
Superlong Chains | Packing within amorphous regions. Interacting with structurally similar amylose chains and providing structural stability. | GBSS and/or SSIII |
Linear amylose Branched amylose  | Unknown. Possibly aids packing glucan within amylopectin matrix in less dense amorphous regions. | GBSS (I and II) and PTST1. SBE isoform responsible for branching amylose unknown, possibly SBEI (SP, DPE, DBE; provision of MOS for GBSS) |
Phosphate Groups | Anionic phosphate groups cause localized mutual repulsion of glucan chains leading to structural instability, making structure prone to amylolytic action. | GWD; adds phosphate group at C6 of glucosyl residue on α-(1,4)-linked chain PWD; adds phosphate group at C3 of phosphoglucan Unknown activity responsible for phosphorylation at C2 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tetlow, I.J.; Bertoft, E. A Review of Starch Biosynthesis in Relation to the Building Block-Backbone Model. Int. J. Mol. Sci. 2020, 21, 7011. https://doi.org/10.3390/ijms21197011
Tetlow IJ, Bertoft E. A Review of Starch Biosynthesis in Relation to the Building Block-Backbone Model. International Journal of Molecular Sciences. 2020; 21(19):7011. https://doi.org/10.3390/ijms21197011
Chicago/Turabian StyleTetlow, Ian J., and Eric Bertoft. 2020. "A Review of Starch Biosynthesis in Relation to the Building Block-Backbone Model" International Journal of Molecular Sciences 21, no. 19: 7011. https://doi.org/10.3390/ijms21197011
APA StyleTetlow, I. J., & Bertoft, E. (2020). A Review of Starch Biosynthesis in Relation to the Building Block-Backbone Model. International Journal of Molecular Sciences, 21(19), 7011. https://doi.org/10.3390/ijms21197011






