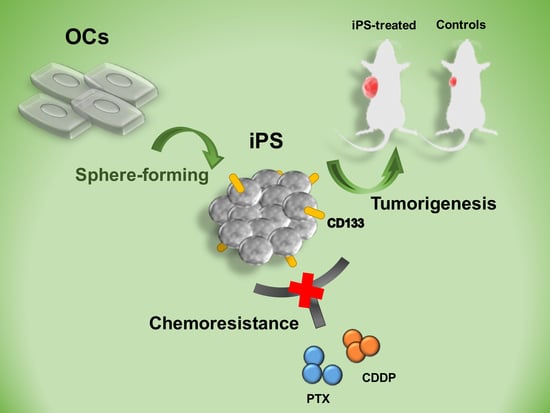
Characteristics of CD133-Sustained Chemoresistant Cancer Stem-Like Cells in Human Ovarian Carcinoma
Abstract
1. Introduction
2. Results
2.1. CD133 Expression Was Associated with the Sphere-Forming Capacity and Induced Stemness in Human OC Cells
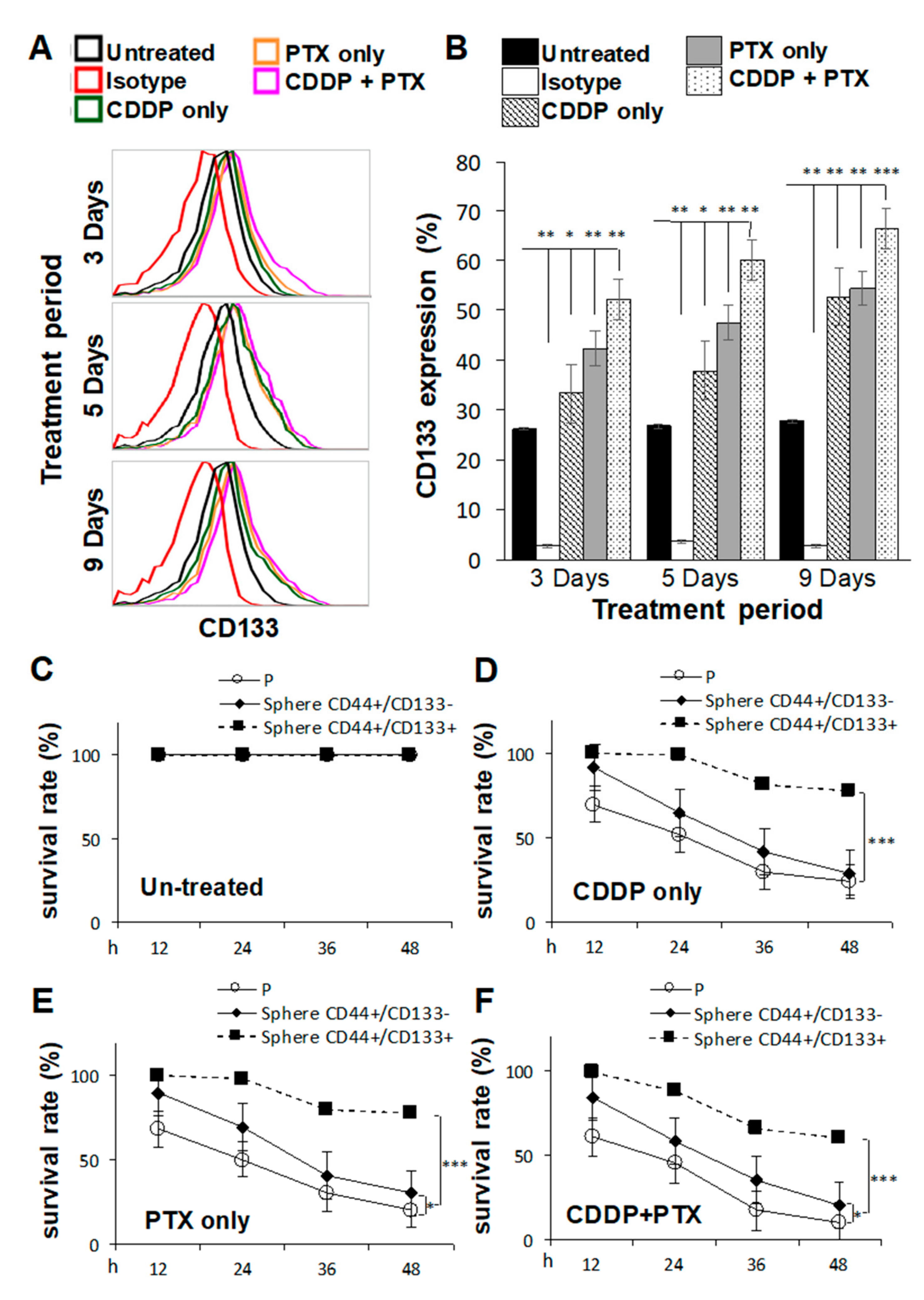
2.2. Pretreatment with Cisplatin (CDDP)- and/or Paclitaxel (PTX)-Enhanced CD133 Expression and Drug Resistance in CSC-Like SKOV3 Spheres
2.3. CD133 Promotes CSC Traits and Is Essential for the Chemoresistant Capacity
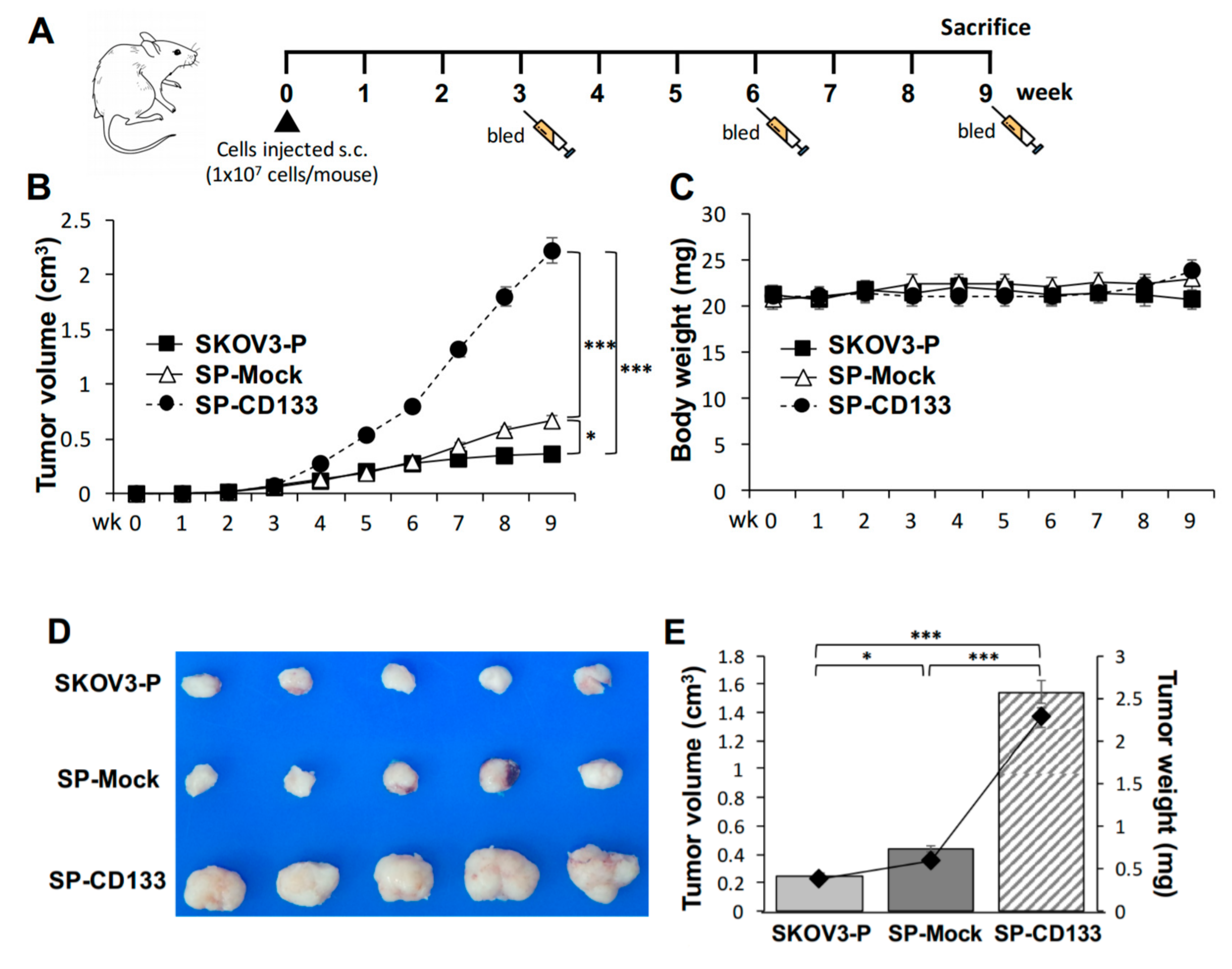
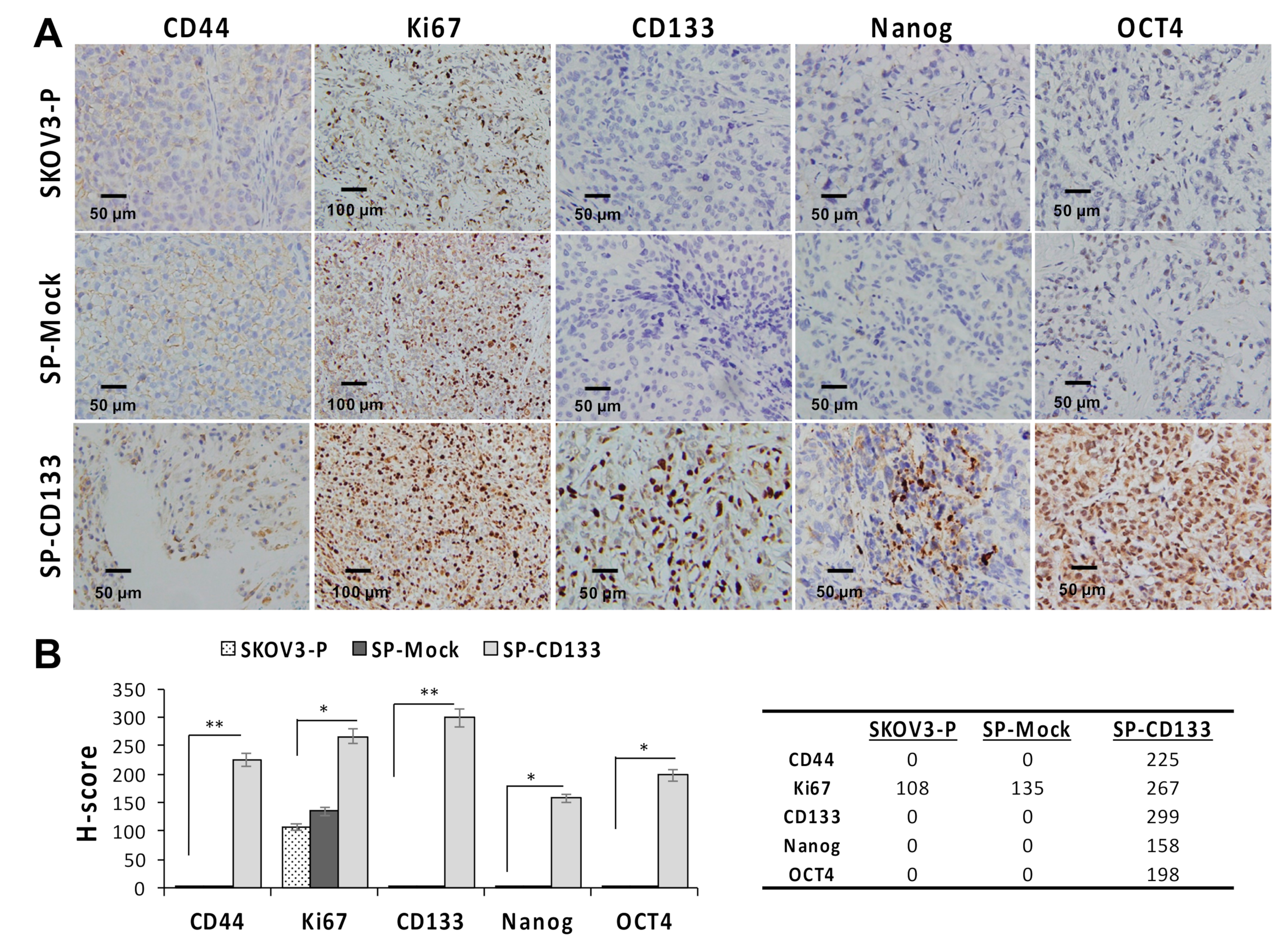
2.4. Overexpression of CD133 Enhanced the In Vivo Tumor Growth of CSC-Like SKOV3 Spheres
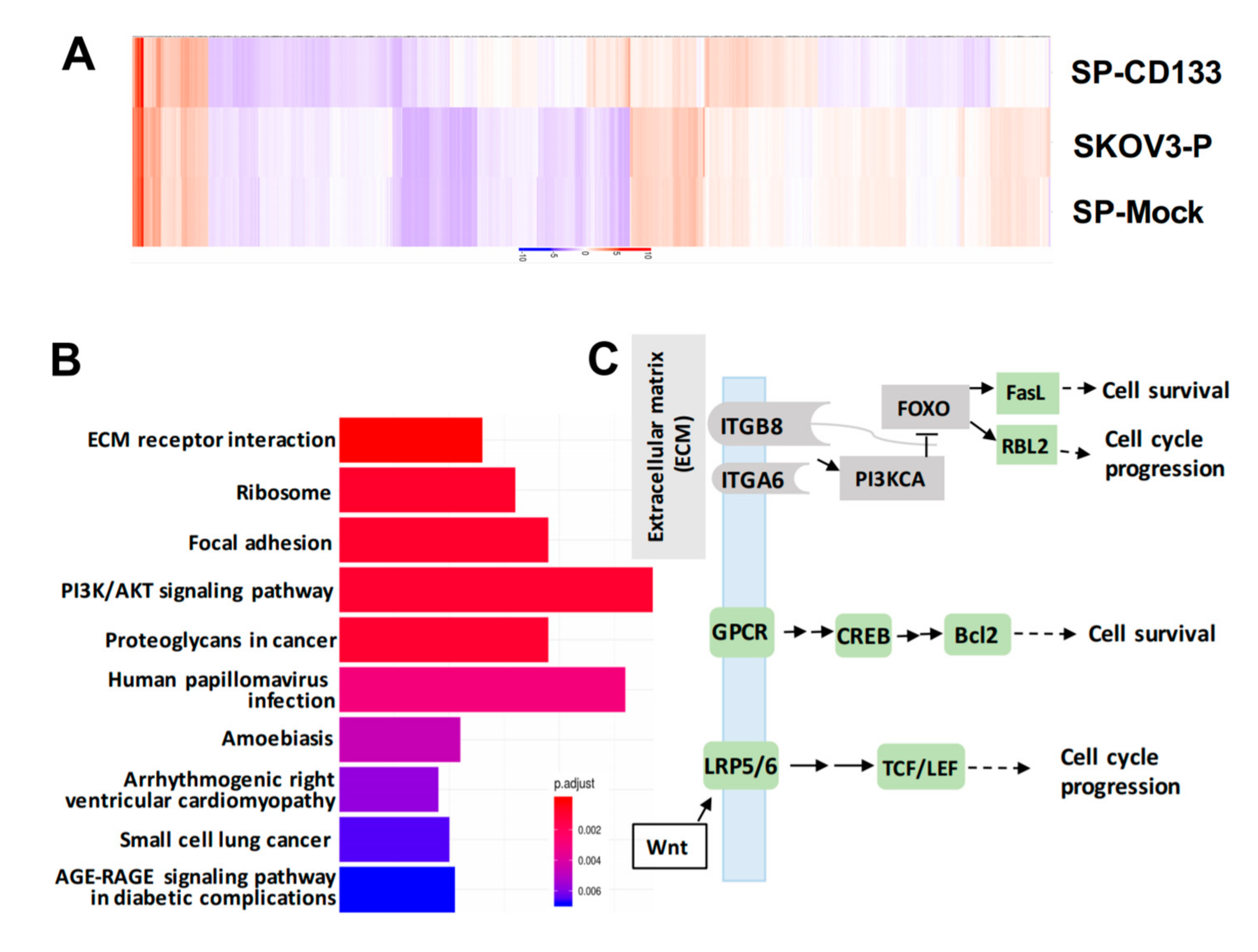
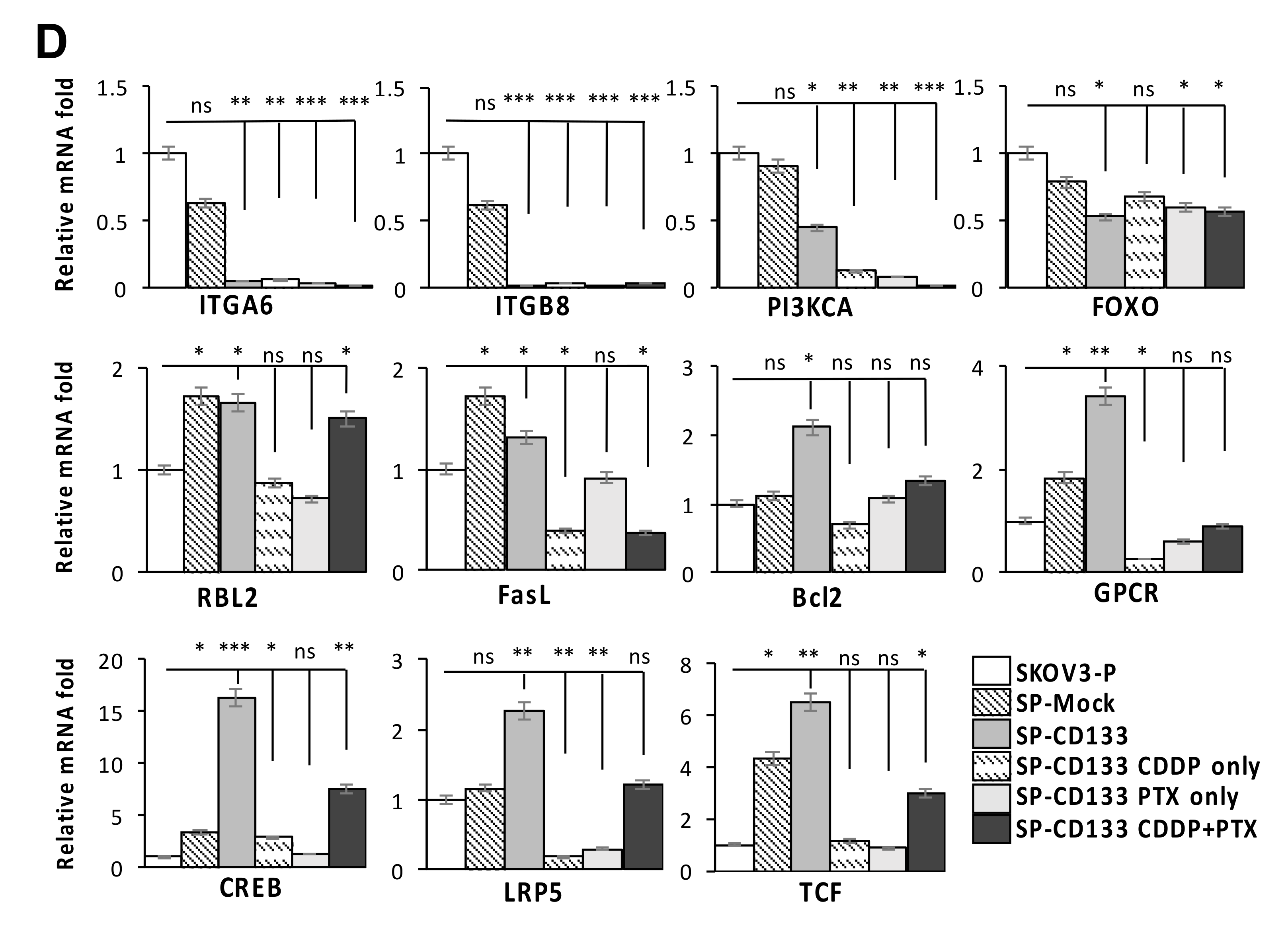
2.5. Reductions in ECM Receptor Interactions/Focal Adhesion in Combination with Induction of Cell Survival/Cell Cycle Progression Caused by CD133 Overexpression
3. Discussion
4. Materials and Methods
4.1. Cell Lines
4.2. Sphere-Formation Assay
4.3. Flow Cytometry
4.4. Cell Viability Assay
4.5. In Vitro Overexpression of CD133
4.6. Assessment of Tumor Growth Using a Xenograft Model
4.7. IHC and Immunoreactivity Scoring
4.8. RNA-Seq and Differential Expression Analysis
4.9. Real-Time Reverse-Transcription Quantitative Polymerase Chain Reaction (RT-qPCR)
4.10. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Coleman, R.L.; Monk, B.J.; Sood, A.K.; Herzog, T.J. Latest research and treatment of advanced-stage epithelial ovarian cancer. Nat. Rev. Clin. Oncol. 2013, 10, 211–224. [Google Scholar] [CrossRef] [PubMed]
- Lowe, K.A.; Chia, V.M.; Taylor, A.; O’Malley, C.; Kelsh, M.; Mohamed, M.; Mowat, F.S.; Goff, B. An international assessment of ovarian cancer incidence and mortality. Gynecol. Oncol. 2013, 130, 107–114. [Google Scholar] [CrossRef] [PubMed]
- Bowtell, D.D.; Bohm, S.; Ahmed, A.A.; Aspuria, P.J.; Bast, R.C., Jr.; Beral, V.; Berek, J.S.; Birrer, M.J.; Blagden, S.; Bookman, M.A.; et al. Rethinking ovarian cancer II: Reducing mortality from high-grade serous ovarian cancer. Nat. Rev. Cancer 2015, 15, 668–679. [Google Scholar] [CrossRef]
- Zhang, S.; Balch, C.; Chan, M.W.; Lai, H.C.; Matei, D.; Schilder, J.M.; Yan, P.S.; Huang, T.H.; Nephew, K.P. Identification and characterization of ovarian cancer-initiating cells from primary human tumors. Cancer Res. 2008, 68, 4311–4320. [Google Scholar] [CrossRef] [PubMed]
- Terraneo, N.; Jacob, F.; Dubrovska, A.; Grunberg, J. Novel Therapeutic Strategies for Ovarian Cancer Stem Cells. Front. Oncol. 2020, 10, 319. [Google Scholar] [CrossRef]
- Reya, T.; Morrison, S.J.; Clarke, M.F.; Weissman, I.L. Stem cells, cancer, and cancer stem cells. Nature 2001, 414, 105–111. [Google Scholar] [CrossRef]
- Kenda Suster, N.; Virant-Klun, I. Presence and role of stem cells in ovarian cancer. World J. Stem Cells 2019, 11, 383–397. [Google Scholar] [CrossRef]
- Pieterse, Z.; Amaya-Padilla, M.A.; Singomat, T.; Binju, M.; Madjid, B.D.; Yu, Y.; Kaur, P. Ovarian cancer stem cells and their role in drug resistance. Int. J. Biochem. Cell Biol. 2019, 106, 117–126. [Google Scholar] [CrossRef]
- Roy, L.; Cowden Dahl, K.D. Can Stemness and Chemoresistance Be Therapeutically Targeted via Signaling Pathways in Ovarian Cancer? Cancers (Basel) 2018, 10, 241. [Google Scholar] [CrossRef]
- Alvero, A.B.; Chen, R.; Fu, H.H.; Montagna, M.; Schwartz, P.E.; Rutherford, T.; Silasi, D.A.; Steffensen, K.D.; Waldstrom, M.; Visintin, I.; et al. Molecular phenotyping of human ovarian cancer stem cells unravels the mechanisms for repair and chemoresistance. Cell Cycle 2009, 8, 158–166. [Google Scholar] [CrossRef]
- Wei, X.; Dombkowski, D.; Meirelles, K.; Pieretti-Vanmarcke, R.; Szotek, P.P.; Chang, H.L.; Preffer, F.I.; Mueller, P.R.; Teixeira, J.; MacLaughlin, D.T.; et al. Mullerian inhibiting substance preferentially inhibits stem/progenitors in human ovarian cancer cell lines compared with chemotherapeutics. Proc. Natl. Acad. Sc.i USA 2010, 107, 18874–18879. [Google Scholar] [CrossRef] [PubMed]
- Gao, M.Q.; Choi, Y.P.; Kang, S.; Youn, J.H.; Cho, N.H. CD24+ cells from hierarchically organized ovarian cancer are enriched in cancer stem cells. Oncogene 2010, 29, 2672–2680. [Google Scholar] [CrossRef] [PubMed]
- Baba, T.; Convery, P.A.; Matsumura, N.; Whitaker, R.S.; Kondoh, E.; Perry, T.; Huang, Z.; Bentley, R.C.; Mori, S.; Fujii, S.; et al. Epigenetic regulation of CD133 and tumorigenicity of CD133+ ovarian cancer cells. Oncogene 2009, 28, 209–218. [Google Scholar] [CrossRef] [PubMed]
- Curley, M.D.; Therrien, V.A.; Cummings, C.L.; Sergent, P.A.; Koulouris, C.R.; Friel, A.M.; Roberts, D.J.; Seiden, M.V.; Scadden, D.T.; Rueda, B.R.; et al. CD133 expression defines a tumor initiating cell population in primary human ovarian cancer. Stem Cells 2009, 27, 2875–2883. [Google Scholar] [CrossRef]
- Zhang, J.; Guo, X.; Chang, D.Y.; Rosen, D.G.; Mercado-Uribe, I.; Liu, J. CD133 expression associated with poor prognosis in ovarian cancer. Mod. Pathol. 2012, 25, 456–464. [Google Scholar] [CrossRef]
- Landen, C.N., Jr.; Goodman, B.; Katre, A.A.; Steg, A.D.; Nick, A.M.; Stone, R.L.; Miller, L.D.; Mejia, P.V.; Jennings, N.B.; Gershenson, D.M.; et al. Targeting aldehyde dehydrogenase cancer stem cells in ovarian cancer. Mol. Cancer Ther. 2010, 9, 3186–3199. [Google Scholar] [CrossRef]
- Parte, S.C.; Batra, S.K.; Kakar, S.S. Characterization of stem cell and cancer stem cell populations in ovary and ovarian tumors. J. Ovarian Res. 2018, 11, 69. [Google Scholar] [CrossRef]
- Jiang, H.; Lin, X.; Liu, Y.; Gong, W.; Ma, X.; Yu, Y.; Xie, Y.; Sun, X.; Feng, Y.; Janzen, V.; et al. Transformation of epithelial ovarian cancer stemlike cells into mesenchymal lineage via EMT results in cellular heterogeneity and supports tumor engraftment. Mol. Med. 2012, 18, 1197–1208. [Google Scholar] [CrossRef]
- Zhou, B.B.; Zhang, H.; Damelin, M.; Geles, K.G.; Grindley, J.C.; Dirks, P.B. Tumour-initiating cells: Challenges and opportunities for anticancer drug discovery. Nat. Rev. Drug Discov. 2009, 8, 806–823. [Google Scholar] [CrossRef]
- Mohiuddin, I.S.; Wei, S.J.; Kang, M.H. Role of OCT4 in cancer stem-like cells and chemotherapy resistance. Biochim. Biophys. Acta Mol. 2020, 1866, 165432. [Google Scholar] [CrossRef]
- Steg, A.D.; Bevis, K.S.; Katre, A.A.; Ziebarth, A.; Dobbin, Z.C.; Alvarez, R.D.; Zhang, K.; Conner, M.; Landen, C.N. Stem cell pathways contribute to clinical chemoresistance in ovarian cancer. Clin. Cancer Res. 2012, 18, 869–881. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.Y.; Kim, S.; Kim, Y.T.; Lim, M.C.; Lee, B.; Jung, K.W.; Kim, J.W.; Park, S.Y.; Won, Y.J. Changes in ovarian cancer survival during the 20 years before the era of targeted therapy. BMC Cancer 2018, 18, 601. [Google Scholar] [CrossRef] [PubMed]
- Condello, S.; Sima, L.; Ivan, C.; Cardenas, H.; Schiltz, G.; Mishra, R.K.; Matei, D. Tissue Tranglutaminase Regulates Interactions between Ovarian Cancer Stem Cells and the Tumor Niche. Cancer Res. 2018, 78, 2990–3001. [Google Scholar] [CrossRef] [PubMed]
- Ruscito, I.; Cacsire Castillo-Tong, D.; Vergote, I.; Ignat, I.; Stanske, M.; Vanderstichele, A.; Ganapathi, R.N.; Glajzer, J.; Kulbe, H.; Trillsch, F.; et al. Exploring the clonal evolution of CD133/aldehyde-dehydrogenase-1 (ALDH1)-positive cancer stem-like cells from primary to recurrent high-grade serous ovarian cancer (HGSOC). A study of the Ovarian Cancer Therapy-Innovative Models Prolong Survival (OCTIPS) Consortium. Eur. J. Cancer 2017, 79, 214–225. [Google Scholar] [CrossRef] [PubMed]
- Liang, J.; Yang, B.; Cao, Q.; Wu, X. Association of Vasculogenic Mimicry Formation and CD133 Expression with Poor Prognosis in Ovarian Cancer. Gynecol. Obstet. Investig. 2016, 81, 529–536. [Google Scholar] [CrossRef]
- Zhou, Q.; Chen, A.; Song, H.; Tao, J.; Yang, H.; Zuo, M. Prognostic value of cancer stem cell marker CD133 in ovarian cancer: A meta-analysis. Int. J. Clin. Exp. Med. 2015, 8, 3080–3088. [Google Scholar]
- Wani, A.A.; Sharma, N.; Shouche, Y.S.; Bapat, S.A. Nuclear-mitochondrial genomic profiling reveals a pattern of evolution in epithelial ovarian tumor stem cells. Oncogene 2006, 25, 6336–6344. [Google Scholar] [CrossRef]
- Labidi-Galy, S.I.; Papp, E.; Hallberg, D.; Niknafs, N.; Adleff, V.; Noe, M.; Bhattacharya, R.; Novak, M.; Jones, S.; Phallen, J.; et al. High grade serous ovarian carcinomas originate in the fallopian tube. Nat. Commun. 2017, 8, 1093. [Google Scholar] [CrossRef]
- Kryczek, I.; Liu, S.; Roh, M.; Vatan, L.; Szeliga, W.; Wei, S.; Banerjee, M.; Mao, Y.; Kotarski, J.; Wicha, M.S.; et al. Expression of aldehyde dehydrogenase and CD133 defines ovarian cancer stem cells. Int. J. Cancer 2012, 130, 29–39. [Google Scholar] [CrossRef]
- Mor, G.; Alvero, A. The duplicitous origin of ovarian cancer. Rambam Maimonides Med. J. 2013, 4, e0006. [Google Scholar] [CrossRef]
- Latifi, A.; Abubaker, K.; Castrechini, N.; Ward, A.C.; Liongue, C.; Dobill, F.; Kumar, J.; Thompson, E.W.; Quinn, M.A.; Findlay, J.K.; et al. Cisplatin treatment of primary and metastatic epithelial ovarian carcinomas generates residual cells with mesenchymal stem cell-like profile. J. Cell Biochem. 2011, 112, 2850–2864. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, N.; Abubaker, K.; Findlay, J.; Quinn, M. Epithelial mesenchymal transition and cancer stem cell-like phenotypes facilitate chemoresistance in recurrent ovarian cancer. Curr. Cancer Drug Targets 2010, 10, 268–278. [Google Scholar] [CrossRef] [PubMed]
- Craveiro, V.; Yang-Hartwich, Y.; Holmberg, J.C.; Joo, W.D.; Sumi, N.J.; Pizzonia, J.; Griffin, B.; Gill, S.K.; Silasi, D.A.; Azodi, M.; et al. Phenotypic modifications in ovarian cancer stem cells following Paclitaxel treatment. Cancer Med. 2013, 2, 751–762. [Google Scholar] [CrossRef] [PubMed]
- Abubaker, K.; Latifi, A.; Luwor, R.; Nazaretian, S.; Zhu, H.; Quinn, M.A.; Thompson, E.W.; Findlay, J.K.; Ahmed, N. Short-term single treatment of chemotherapy results in the enrichment of ovarian cancer stem cell-like cells leading to an increased tumor burden. Mol. Cancer 2013, 12, 24. [Google Scholar] [CrossRef]
- Siu, M.K.; Wong, E.S.; Kong, D.S.; Chan, H.Y.; Jiang, L.; Wong, O.G.; Lam, E.W.; Chan, K.K.; Ngan, H.Y.; Le, X.F.; et al. Stem cell transcription factor NANOG controls cell migration and invasion via dysregulation of E-cadherin and FoxJ1 and contributes to adverse clinical outcome in ovarian cancers. Oncogene 2013, 32, 3500–3509. [Google Scholar] [CrossRef]
- Hu, T.; Chung, Y.M.; Guan, M.; Ma, M.; Ma, J.; Berek, J.S.; Hu, M.C. Reprogramming ovarian and breast cancer cells into non-cancerous cells by low-dose metformin or SN-38 through FOXO3 activation. Sci. Rep. 2014, 4, 5810. [Google Scholar] [CrossRef]
- Bareiss, P.M.; Paczulla, A.; Wang, H.; Schairer, R.; Wiehr, S.; Kohlhofer, U.; Rothfuss, O.C.; Fischer, A.; Perner, S.; Staebler, A.; et al. SOX2 expression associates with stem cell state in human ovarian carcinoma. Cancer Res. 2013, 73, 5544–5555. [Google Scholar] [CrossRef]
- Cooper, J.; Giancotti, F.G. Integrin Signaling in Cancer: Mechanotransduction, Stemness, Epithelial Plasticity, and Therapeutic Resistance. Cancer Cell 2019, 35, 347–367. [Google Scholar] [CrossRef]
- Tingting Zhu, R.C.; Wang, J.; Yue, H.; Lu, X.; Li, J. The prognostic value of ITGA and ITGB superfamily members in patients with high grade serous ovarian cancer. Cancer Cell Int. 2020, 20, 257. [Google Scholar] [CrossRef]
- Zhang, K.; Myllymaki, S.M.; Gao, P.; Devarajan, R.; Kytola, V.; Nykter, M.; Wei, G.H.; Manninen, A. Oncogenic K-Ras upregulates ITGA6 expression via FOSL1 to induce anoikis resistance and synergizes with alphaV-Class integrins to promote EMT. Oncogene 2017, 36, 5681–5694. [Google Scholar] [CrossRef]
- Cui, Y.; Wu, F.; Tian, D.; Wang, T.; Lu, T.; Huang, X.; Zhang, P.; Qin, L. miR-199a-3p enhances cisplatin sensitivity of ovarian cancer cells by targeting ITGB8. Oncol. Rep. 2018, 39, 1649–1657. [Google Scholar] [CrossRef] [PubMed]
- He, J.; Liu, Y.; Zhang, L.; Zhang, H. Integrin Subunit beta 8 (ITGB8) Upregulation Is an Independent Predictor of Unfavorable Survival of High-Grade Serous Ovarian Carcinoma Patients. Med. Sci. Monit. 2018, 24, 8933–8940. [Google Scholar] [CrossRef] [PubMed]
- Flesken-Nikitin, A.; Hwang, C.I.; Cheng, C.Y.; Michurina, T.V.; Enikolopov, G.; Nikitin, A.Y. Ovarian surface epithelium at the junction area contains a cancer-prone stem cell niche. Nature 2013, 495, 241–245. [Google Scholar] [CrossRef] [PubMed]
- Schindler, A.J.; Watanabe, A.; Howell, S.B. LGR5 and LGR6 in stem cell biology and ovarian cancer. Oncotarget 2018, 9, 1346–1355. [Google Scholar] [CrossRef]
- Anglesio, M.S.; Wiegand, K.C.; Melnyk, N.; Chow, C.; Salamanca, C.; Prentice, L.M.; Senz, J.; Yang, W.; Spillman, M.A.; Cochrane, D.R.; et al. Type-specific cell line models for type-specific ovarian cancer research. PLoS ONE 2013, 8, e72162. [Google Scholar] [CrossRef]
- Tsui Lien Mao, C.H.M.; Liao, Y.J.; Chen, Y.J.; Yeh, C.Y.; Liu, C.L. Ex Vivo Expanded Human Vr9Vd2 T-Cells Can Suppress Epithelial Ovarian Cancer Cell Growth. Int. J. Mol. Sci. 2019, 20, 1139. [Google Scholar] [CrossRef]
- Ishibashi, H.; Suzuki, T.; Suzuki, S.; Moriya, T.; Kaneko, C.; Takizawa, T.; Sunamori, M.; Handa, M.; Kondo, T.; Sasano, H. Sex steroid hormone receptors in human thymoma. J. Clin. Endocrinol. Metab. 2003, 88, 2309–2317. [Google Scholar] [CrossRef]


| Gene Name | Primer Sequences | Gene Bank No. |
|---|---|---|
| CD44-F CD44-R | 5′-GCTGACCTCTGCAAGGCTT-3′ 5′-GGATGTACACCCCTGTGTT-3′ | NM_000610.4 |
| Sox2-F Sox2-R | 5′-CGAGTGGAAACTTTTGTCGGA-3′ 5’-TGTGCAGCGCTCGCAG-3′ | NM_003106 |
| Oct4-F Oct4-R | 5′-GTGGAGAGCAACTCCGATG-3′ 5′-TGCTCCAGCTTCTCCTTCTC-3′ | NM_002701 |
| Nanog-F Nanog-R | 5′-ATTCAGGACAGCCCTGATTCTTC-3′ 5′-TTTTTGCGACACTCTTCTCTGC-3′ | NM_024865 |
| ITGA6-F ITGA6-R | 5′-CCATGCACGCGGATCGAGTT-3′ 5′-GGGATTCCTGCTTCGTATTA-3′ | NM_001079818.3 |
| TIGB8-F TIGB8-R | 5′-GGACGCTGCCGACTTGTCTT-3′ 5′-GGAAACCACCCTAATGTACA-3′ | NM_002214.3 |
| PIK3CA-F PIK3CA-R | 5′-CGACCATCATCAGGTGAA-3′ 5′-GCTTCTTGAGTAACACTT-3′ | NM_006218.4 |
| FOXO3-F FOXO3-R | 5′-GGAGCTGGACCCGGAGTT-3′ 5′-CGTCCTCGTCGTCTTCAT-3′ | NM_201559.3 |
| RBL2-F RBL2-R | 5′-ATGAGCGAAAGCTACACG-3′ 5′-TGCTCTGAACATTTCAGG-3′ | NM_005611.4 |
| FasL-F FasL-R | 5′-AAGAGAGATCCAGCTTGC-3′ 5′-TAGATCTGGGGATATGGG-3′ | NM_001302746.1 |
| Bcl2-F Bcl2-R GPCR-F GPCR-R | 5′-TACTAATAATAACGTGCC-3′ 5′-ATCCAGCTATTTTATTGG-3′ 5′–AATTGGAGGTTTGTTTGC-3′ 5′–ATACCCCAGTTTGACTCC-3′ | NM_000633.3 |
| NM_148963.4 | ||
| CREB-F CREB-R | 5′–AACAAATGACAGTTCAAGC-3′ 5′–GAGACTGAATAACTGATGG-3′ | NM_134442.5 |
| LRP5-F LRP5-R | 5′–AAGAAGCTGTACTGGACG-3′ 5′–AATCCGGGGCGTCTCACC-3′ | NM_001291902.2 |
| TCF-F TCF-R | 5′–TTGTGACCAGTATTAAGG-3′ 5′–AAGCCCTGGCCCCAGTCC-3′ | NM_001287182.2 |
| S26-F S26-R | 5′-CCGTGCCTCCAAGATGACAAAG-3′ 5′-GTTCGGTCCTTGCGGGCTTCAC-3′ | NC_016448.1 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, C.L.; Chen, Y.J.; Fan, M.H.; Liao, Y.J.; Mao, T.L. Characteristics of CD133-Sustained Chemoresistant Cancer Stem-Like Cells in Human Ovarian Carcinoma. Int. J. Mol. Sci. 2020, 21, 6467. https://doi.org/10.3390/ijms21186467
Liu CL, Chen YJ, Fan MH, Liao YJ, Mao TL. Characteristics of CD133-Sustained Chemoresistant Cancer Stem-Like Cells in Human Ovarian Carcinoma. International Journal of Molecular Sciences. 2020; 21(18):6467. https://doi.org/10.3390/ijms21186467
Chicago/Turabian StyleLiu, Chao Lien, Ying Jen Chen, Ming Huei Fan, Yi Jen Liao, and Tsui Lien Mao. 2020. "Characteristics of CD133-Sustained Chemoresistant Cancer Stem-Like Cells in Human Ovarian Carcinoma" International Journal of Molecular Sciences 21, no. 18: 6467. https://doi.org/10.3390/ijms21186467
APA StyleLiu, C. L., Chen, Y. J., Fan, M. H., Liao, Y. J., & Mao, T. L. (2020). Characteristics of CD133-Sustained Chemoresistant Cancer Stem-Like Cells in Human Ovarian Carcinoma. International Journal of Molecular Sciences, 21(18), 6467. https://doi.org/10.3390/ijms21186467