Potential Associations Among Alteration of Salivary miRNAs, Saliva Microbiome Structure, and Cognitive Impairments in Autistic Children
Abstract
1. Introduction
2. Results
2.1. Demographic and Neuropsychological Characteristics
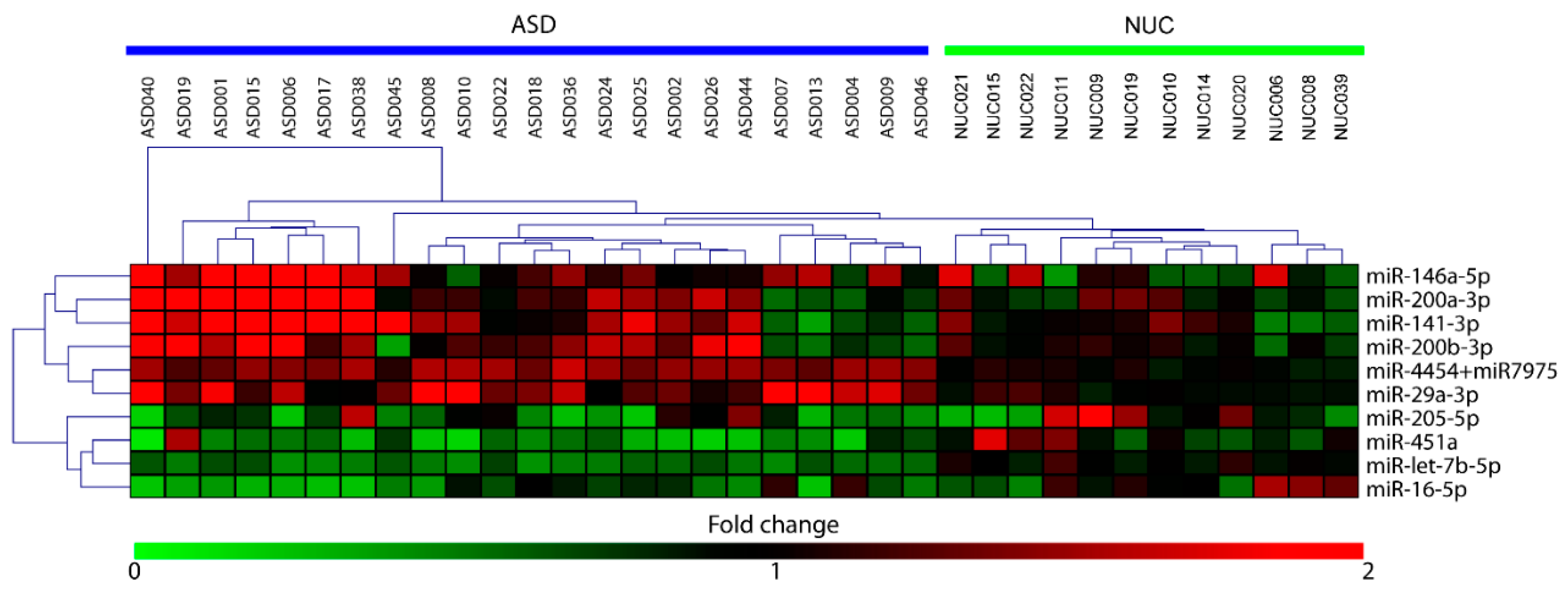
2.2. Salivary miRNA Expression Profiling
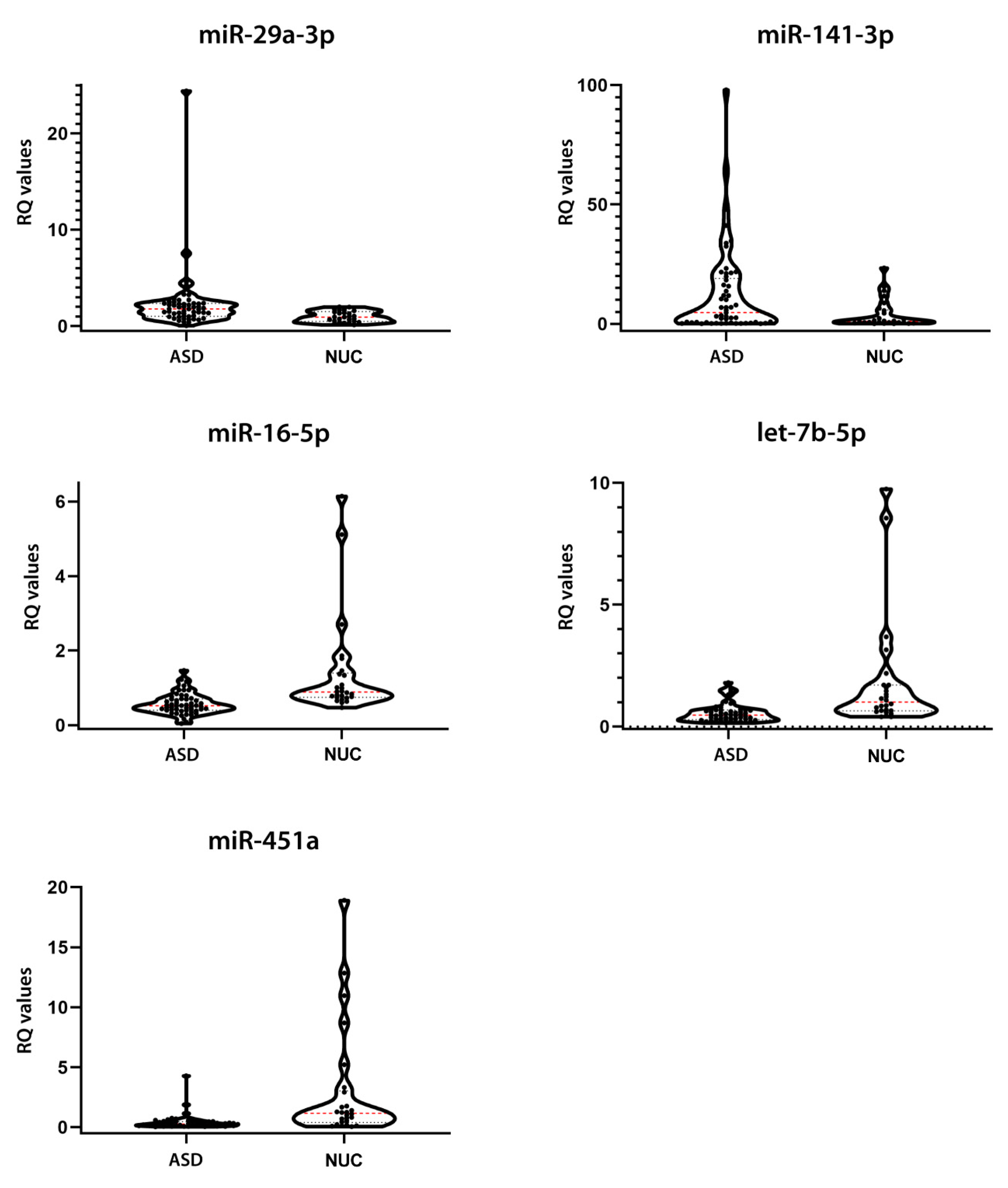
2.3. Salivary miRNA Expression Validation
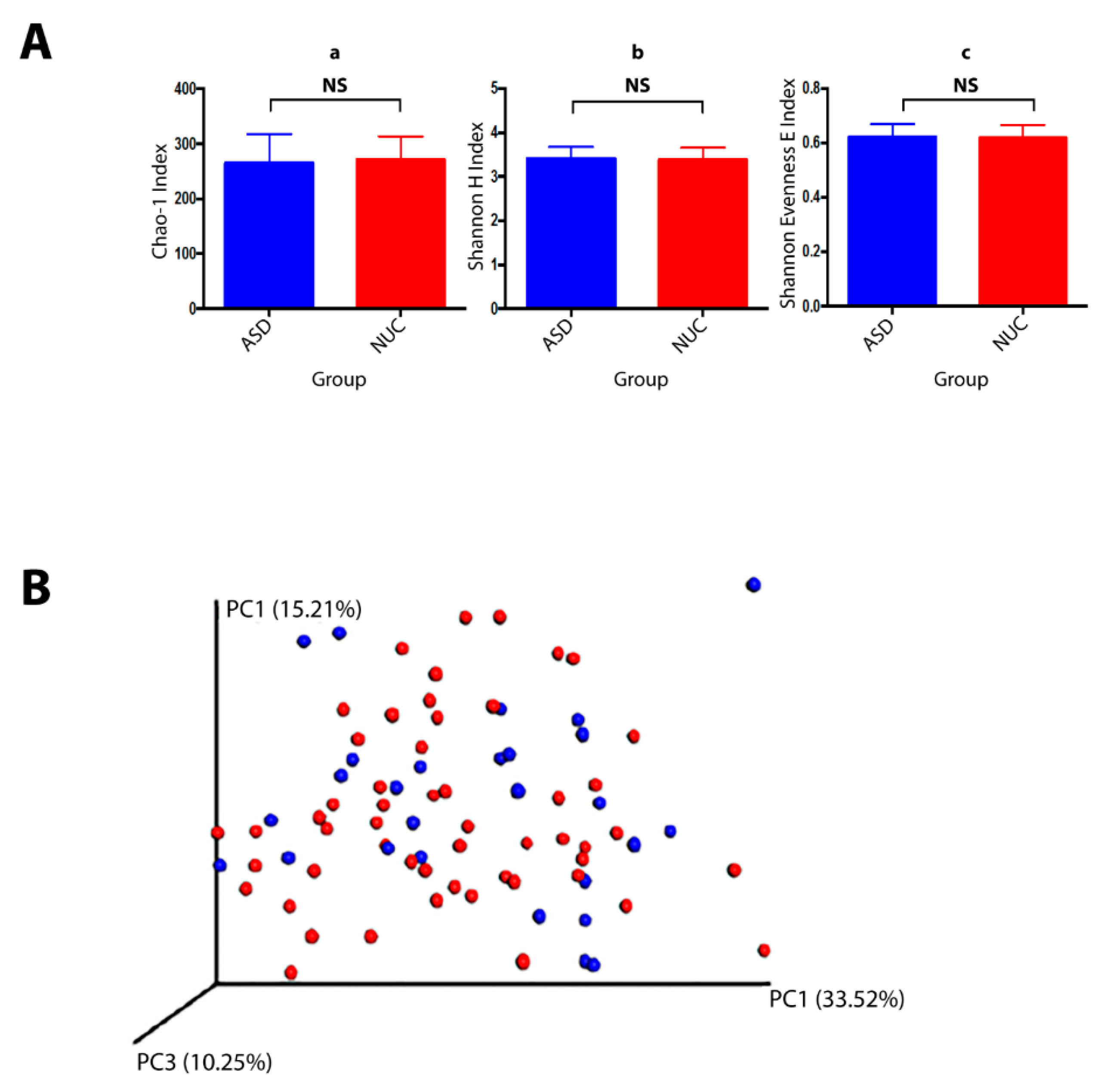
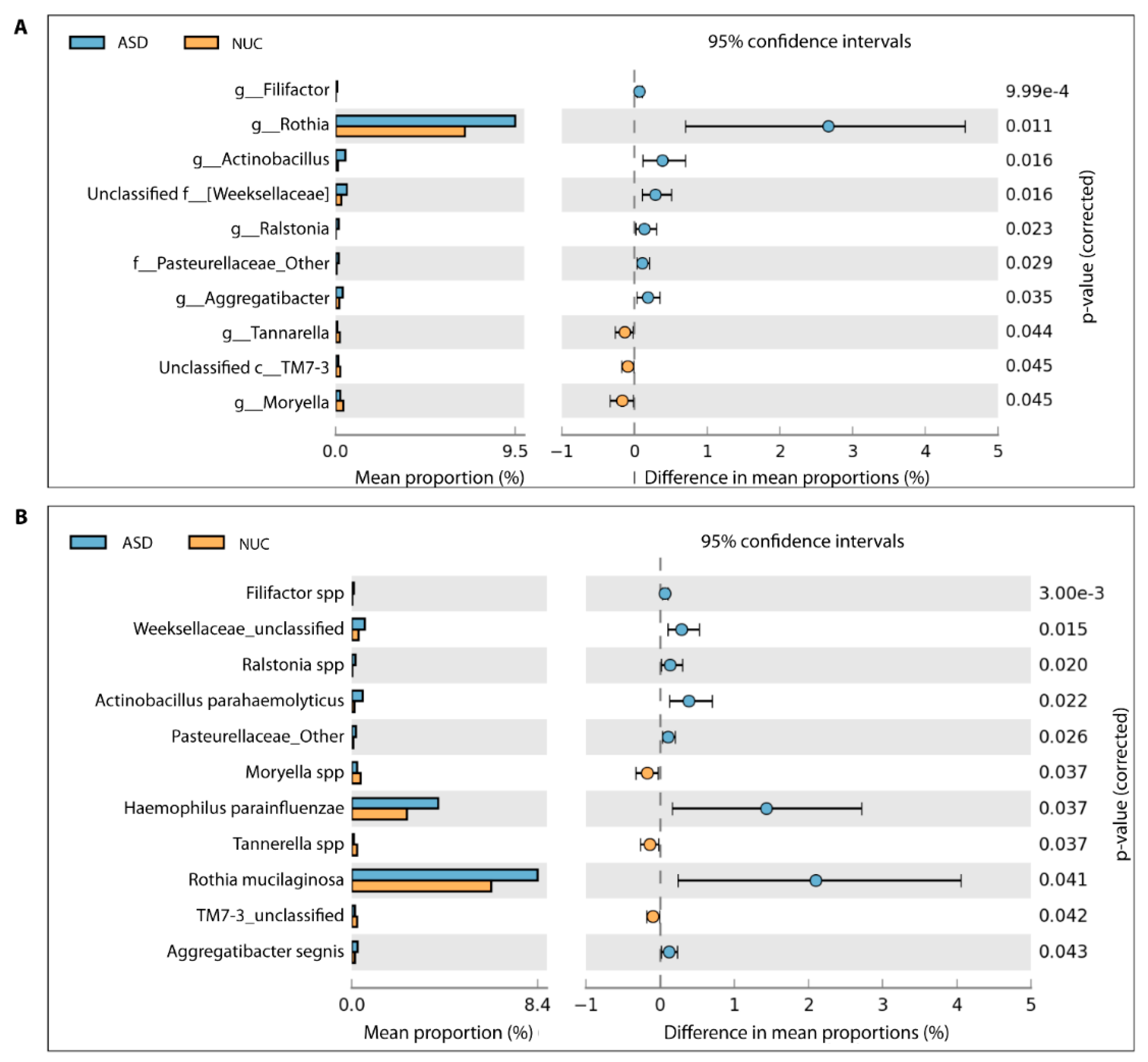
2.4. Microbial Structure of the Saliva Microbiome in Children with ASD and NUCs
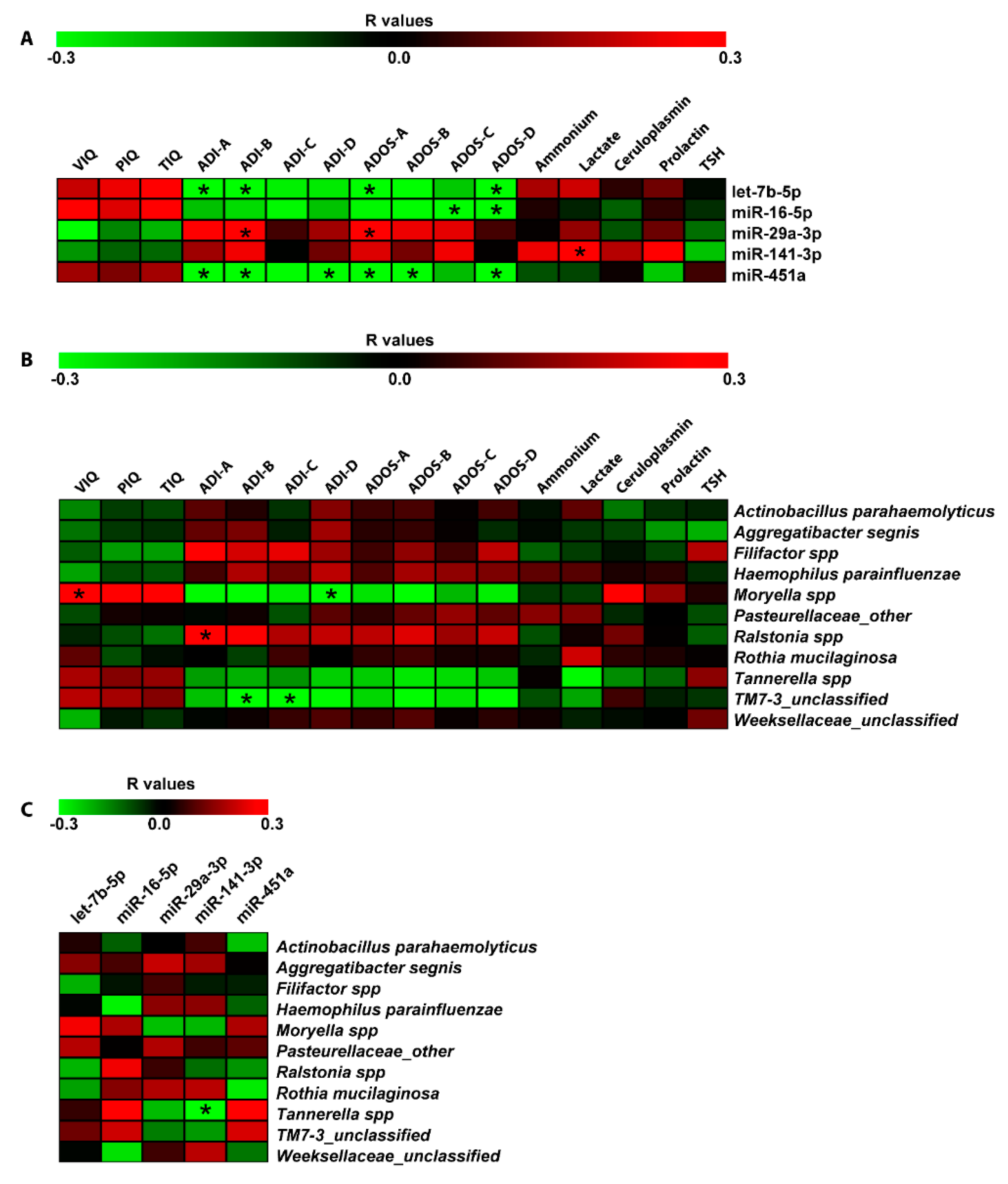
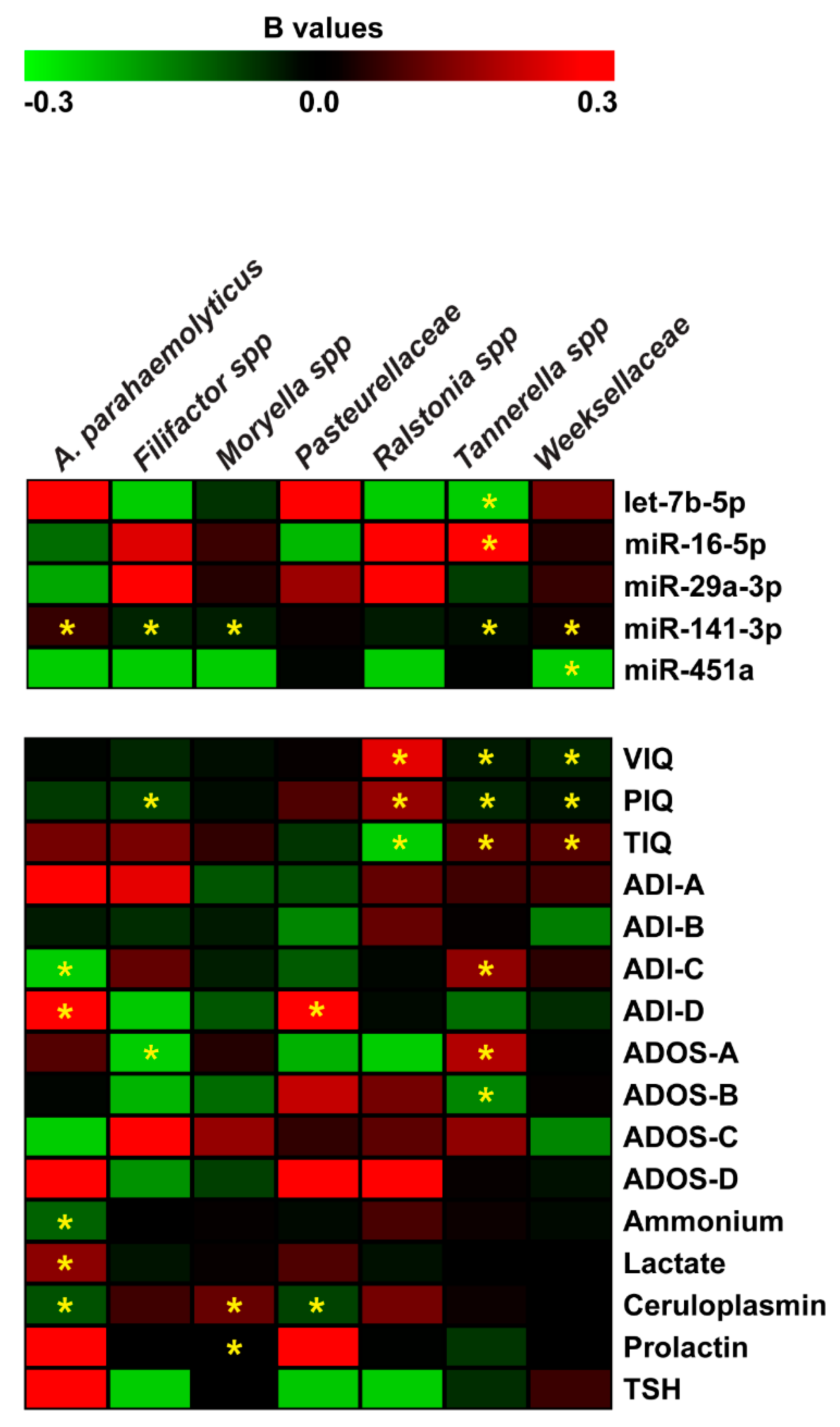
2.5. Correlation and Negative Binomial Regression Analyses Among Salivary miRNAs, Bacteria, and Neuropsychological/Hematological Parameters
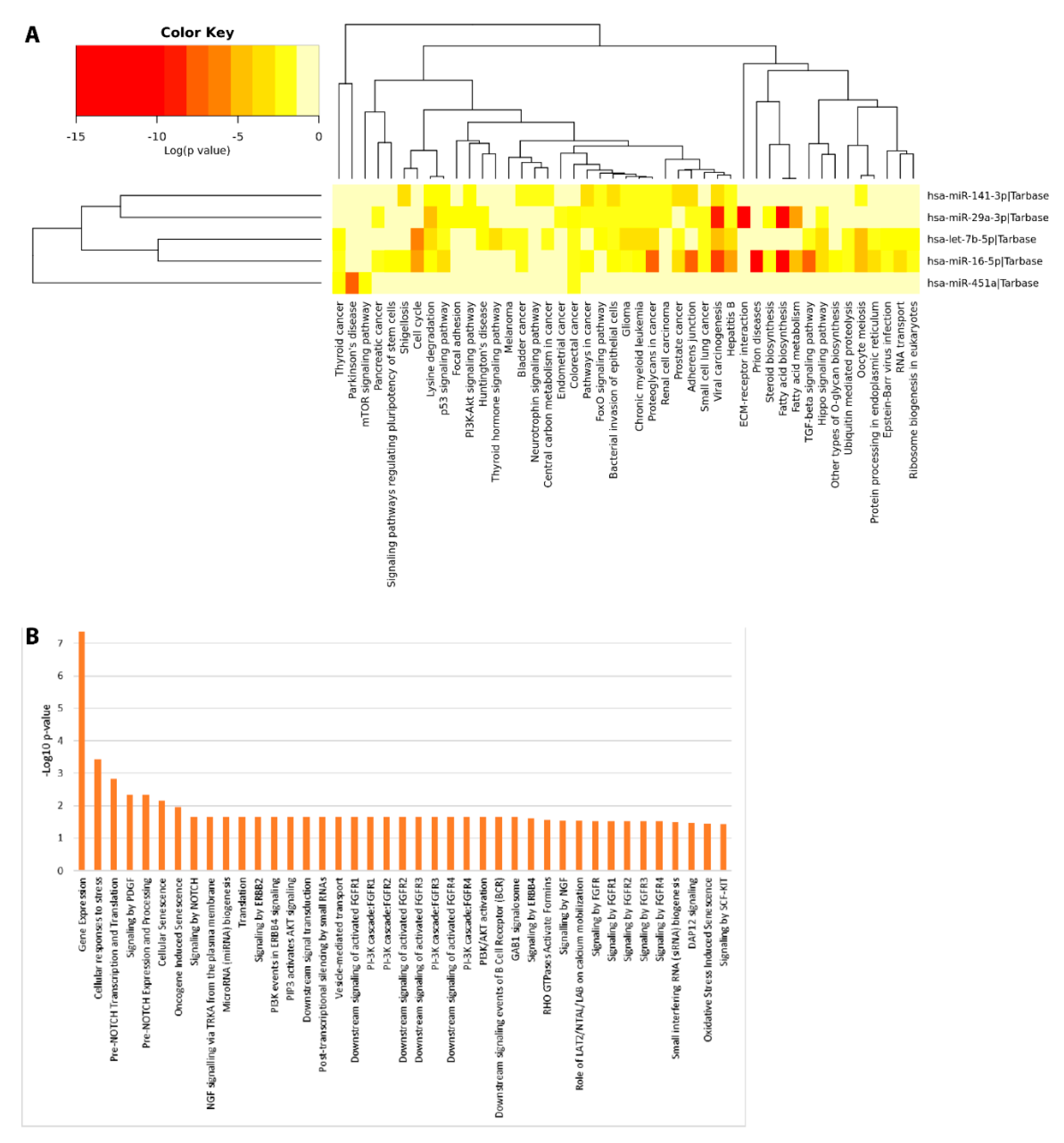
2.6. Functional Enrichment Analyses
3. Discussion
3.1. Circulating miRNAs and Microbiome Structure are Altered in Saliva of Pediatric ASD Patients
3.2. Potential Associations Among Cognitive Impairments, Salivary miRNA Expression and Microbiome Alteration in ASD Children
3.3. Potential Crosstalk between miRNAs and the Microbiome in Saliva
4. Materials and Methods
4.1. Ethics Approval and Consent to Participate
4.2. Participant Selection
4.3. Assessment
4.4. Sample Collection
4.5. RNA Extraction
4.6. MiRNA Profiling by NanoString Technology
4.7. MiRNA Data Validation by Single TaqMan Assays
4.8. DNA Extraction, 16S rRNA Gene Library Preparation, and Sequencing
4.9. Processing and Analyses of Sequencing Data
4.10. Correlation and Negative Binomial Regression Analyses
4.11. Computational Enrichment Analysis
4.12. Statistical Approach
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| ADI-R | Autism Diagnostic Interview—Revised |
| ADOS | Autism Diagnostic Observation Schedule |
| ASD | Autistic Spectrum Disorder |
| CNS | Central Nervous System |
| CNVs | Copy Number Variants |
| CSF | cerebrospinal fluid |
| df | Degree of freedom |
| DE | Differentially expressed |
| DSM-5 | Diagnostic and Statistical Manual of Mental Disorders, 5th edition |
| FC | Fold Change |
| FDR | False Discovery Rate |
| FoxO | Forkhead box O |
| GLM | Generalized linear model |
| GMN | Global median normalization |
| IQ | Intelligence Quotient |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| LR | Likelihood ratio |
| LDH | Lactate dehydrogenase |
| MeV | Multi experiment viewer |
| miRNA | microRNA |
| MLR | Multiple linear regression |
| NB | Negative binomial |
| NBR | Negative binomial regression |
| NUC | Neurologically unaffected control |
| NS | Non-significant |
| OTU | Operational taxonomic Unit |
| p | p-value |
| PCoA | Principal coordinate analysis |
| PIQ | Performance Intelligence Quotient |
| QIIME | Quantitative Insights Into Microbial Ecology |
| RQ | Relative quantification |
| rRNA | Ribosomal RNA |
| RT-PCR | Real Time-polymerase chain reactions |
| SAM | Significance of Microarrays Analysis |
| SLR | Simple linear regression |
| SMD | Standardized mean difference |
| TIQ | Total Intelligence Quotient |
| TLR-7 | Tall-like Receptor 7 |
| TSH | Thyroid-stimulating Hormone |
| VIQ | Verbal Intelligence Quotient |
| WISC-III | Wechsler Intelligence Scale for Children, III edition |
| WPSSI | Wechsler Preschool and Primary Scale of Intelligence |
| YOAD | Young-Onset Alzheimer Disease. |
References
- Fakhoury, M. Autistic spectrum disorders: A review of clinical features, theories and diagnosis. Int. J. Dev. Neurosci. 2015, 43, 70–77. [Google Scholar] [CrossRef]
- Abdolmaleky, H.M.; Zhou, J.R.; Thiagalingam, S. An update on the epigenetics of psychotic diseases and autism. Epigenomics 2015, 7, 427–449. [Google Scholar] [CrossRef] [PubMed]
- Nardone, S.; Elliott, E. The Interaction between the Immune System and Epigenetics in the Etiology of Autism Spectrum Disorders. Front. Neurosci. 2016, 10, 329. [Google Scholar] [CrossRef] [PubMed]
- Sun, W.; Poschmann, J.; Cruz-Herrera Del Rosario, R.; Parikshak, N.N.; Hajan, H.S.; Kumar, V.; Ramasamy, R.; Belgard, T.G.; Elanggovan, B.; Wong, C.C.Y.; et al. Histone Acetylome-wide Association Study of Autism Spectrum Disorder. Cell 2016, 167, 1385–1397. [Google Scholar] [CrossRef] [PubMed]
- Muhle, R.A.; Reed, H.E.; Stratigos, K.A.; Veenstra-VanderWeele, J. The Emerging Clinical Neuroscience of Autism Spectrum Disorder: A Review. JAMA Psychiatry 2018, 75, 514–523. [Google Scholar] [CrossRef] [PubMed]
- Vorstman, J.A.S.; Parr, J.R.; Moreno-De-Luca, D.; Anney, R.J.L.; Nurnberger, J.I., Jr.; Hallmayer, J.F. Autism genetics: Opportunities and challenges for clinical translation. Nat. Rev. Genet. 2017, 18, 362–376. [Google Scholar] [CrossRef]
- Miles, J.H. Autism spectrum disorders—A genetics review. Genet. Med. 2011, 13, 278–294. [Google Scholar] [CrossRef]
- Levitt, P.; Campbell, D.B. The genetic and neurobiologic compass points toward common signaling dysfunctions in autism spectrum disorders. J. Clin. Investig. 2009, 119, 747–754. [Google Scholar] [CrossRef]
- Voineagu, I.; Eapen, V. Converging Pathways in Autism Spectrum Disorders: Interplay between Synaptic Dysfunction and Immune Responses. Front. Hum. Neurosci. 2013, 7, 738. [Google Scholar] [CrossRef]
- Chen, J.; Alberts, I.; Li, X. Dysregulation of the IGF-I/PI3K/AKT/mTOR signaling pathway in autism spectrum disorders. Int. J. Dev. Neurosci. 2014, 35, 35–41. [Google Scholar] [CrossRef]
- Gokoolparsadh, A.; Sutton, G.J.; Charamko, A.; Green, N.F.; Pardy, C.J.; Voineagu, I. Searching for convergent pathways in autism spectrum disorders: Insights from human brain transcriptome studies. Cell. Mol. Life Sci. 2016, 73, 4517–4530. [Google Scholar] [CrossRef] [PubMed]
- Ansel, A.; Rosenzweig, J.P.; Zisman, P.D.; Melamed, M.; Gesundheit, B. Variation in Gene Expression in Autism Spectrum Disorders: An Extensive Review of Transcriptomic Studies. Front. Neurosci. 2016, 10, 601. [Google Scholar] [CrossRef] [PubMed]
- Adams, J.B.; Johansen, L.J.; Powell, L.D.; Quig, D.; Rubin, R.A. Gastrointestinal flora and gastrointestinal status in children with autism—Comparisons to typical children and correlation with autism severity. BMC Gastroenterol. 2011, 11, 22. [Google Scholar] [CrossRef] [PubMed]
- Williams, B.L.; Hornig, M.; Parekh, T.; Lipkin, W.I. Application of novel PCR-based methods for detection, quantitation, and phylogenetic characterization of Sutterella species in intestinal biopsy samples from children with autism and gastrointestinal disturbances. MBio 2012, 3, e00261-11. [Google Scholar] [CrossRef]
- Erny, D.; Hrabe de Angelis, A.L.; Jaitin, D.; Wieghofer, P.; Staszewski, O.; David, E.; Keren-Shaul, H.; Mahlakoiv, T.; Jakobshagen, K.; Buch, T.; et al. Host microbiota constantly control maturation and function of microglia in the CNS. Nat. Neurosci. 2015, 18, 965–977. [Google Scholar] [CrossRef]
- Desbonnet, L.; Clarke, G.; Shanahan, F.; Dinan, T.G.; Cryan, J.F. Microbiota is essential for social development in the mouse. Mol. Psychiatry 2014, 19, 146–148. [Google Scholar] [CrossRef]
- Arentsen, T.; Raith, H.; Qian, Y.; Forssberg, H.; Diaz Heijtz, R. Host microbiota modulates development of social preference in mice. Microb. Ecol. Health Dis. 2015, 26, 29719. [Google Scholar] [CrossRef]
- Crumeyrolle-Arias, M.; Jaglin, M.; Bruneau, A.; Vancassel, S.; Cardona, A.; Dauge, V.; Naudon, L.; Rabot, S. Absence of the gut microbiota enhances anxiety-like behavior and neuroendocrine response to acute stress in rats. Psychoneuroendocrinology 2014, 42, 207–217. [Google Scholar] [CrossRef]
- Li, Q.; Han, Y.; Dy, A.B.C.; Hagerman, R.J. The Gut Microbiota and Autism Spectrum Disorders. Front. Cell. Neurosci. 2017, 11, 120. [Google Scholar] [CrossRef]
- Linares, D.M.; Ross, P.; Stanton, C. Beneficial Microbes: The pharmacy in the gut. Bioengineered 2016, 7, 11–20. [Google Scholar] [CrossRef]
- Belkaid, Y.; Hand, T.W. Role of the microbiota in immunity and inflammation. Cell 2014, 157, 121–141. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, K. Influence of bacteria on epigenetic gene control. Cell. Mol. Life Sci. 2014, 71, 1045–1054. [Google Scholar] [CrossRef] [PubMed]
- Van de Wiele, T.; Van Praet, J.T.; Marzorati, M.; Drennan, M.B.; Elewaut, D. How the microbiota shapes rheumatic diseases. Nat. Rev. Rheumatol. 2016, 12, 398–411. [Google Scholar] [CrossRef]
- Clarke, G.; O’Mahony, S.M.; Dinan, T.G.; Cryan, J.F. Priming for health: Gut microbiota acquired in early life regulates physiology, brain and behaviour. Acta Paediatr. 2014, 103, 812–819. [Google Scholar] [CrossRef] [PubMed]
- Vuong, H.E.; Hsiao, E.Y. Emerging Roles for the Gut Microbiome in Autism Spectrum Disorder. Biol. Psychiatry 2017, 81, 411–423. [Google Scholar] [CrossRef] [PubMed]
- Dewhirst, F.E.; Chen, T.; Izard, J.; Paster, B.J.; Tanner, A.C.; Yu, W.H.; Lakshmanan, A.; Wade, W.G. The human oral microbiome. J. Bacteriol. 2010, 192, 5002–5017. [Google Scholar] [CrossRef]
- Krishnan, K.; Chen, T.; Paster, B.J. A practical guide to the oral microbiome and its relation to health and disease. Oral Dis. 2017, 23, 276–286. [Google Scholar] [CrossRef]
- Fabian, T.K.; Fejerdy, P.; Csermely, P. Salivary Genomics, Transcriptomics and Proteomics: The Emerging Concept of the Oral Ecosystem and their Use in the Early Diagnosis of Cancer and other Diseases. Curr. Genom. 2008, 9, 11–21. [Google Scholar] [CrossRef]
- He, J.; Li, Y.; Cao, Y.; Xue, J.; Zhou, X. The oral microbiome diversity and its relation to human diseases. Folia Microbiol. 2015, 60, 69–80. [Google Scholar] [CrossRef]
- Qin, N.; Yang, F.; Li, A.; Prifti, E.; Chen, Y.; Shao, L.; Guo, J.; Le Chatelier, E.; Yao, J.; Wu, L.; et al. Alterations of the human gut microbiome in liver cirrhosis. Nature 2014, 513, 59–64. [Google Scholar] [CrossRef]
- Farrell, J.J.; Zhang, L.; Zhou, H.; Chia, D.; Elashoff, D.; Akin, D.; Paster, B.J.; Joshipura, K.; Wong, D.T. Variations of oral microbiota are associated with pancreatic diseases including pancreatic cancer. Gut 2012, 61, 582–588. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Zhang, D.; Jia, H.; Feng, Q.; Wang, D.; Liang, D.; Wu, X.; Li, J.; Tang, L.; Li, Y.; et al. The oral and gut microbiomes are perturbed in rheumatoid arthritis and partly normalized after treatment. Nat. Med. 2015, 21, 895–905. [Google Scholar] [CrossRef] [PubMed]
- Torres, P.J.; Fletcher, E.M.; Gibbons, S.M.; Bouvet, M.; Doran, K.S.; Kelley, S.T. Characterization of the salivary microbiome in patients with pancreatic cancer. PeerJ 2015, 3, e1373. [Google Scholar] [CrossRef] [PubMed]
- Ding, T.; Schloss, P.D. Dynamics and associations of microbial community types across the human body. Nature 2014, 509, 357–360. [Google Scholar] [CrossRef] [PubMed]
- Pereira, P.A.B.; Aho, V.T.E.; Paulin, L.; Pekkonen, E.; Auvinen, P.; Scheperjans, F. Oral and nasal microbiota in Parkinson’s disease. Parkinsonism Relat. Disord. 2017, 38, 61–67. [Google Scholar] [CrossRef]
- Shoemark, D.K.; Allen, S.J. The microbiome and disease: Reviewing the links between the oral microbiome, aging, and Alzheimer’s disease. J. Alzheimer’s Dis. 2015, 43, 725–738. [Google Scholar] [CrossRef]
- Singhrao, S.K.; Harding, A.; Simmons, T.; Robinson, S.; Kesavalu, L.; Crean, S. Oral inflammation, tooth loss, risk factors, and association with progression of Alzheimer’s disease. J. Alzheimer’s Dis. 2014, 42, 723–737. [Google Scholar] [CrossRef]
- Gonzalez, A.; Hyde, E.; Sangwan, N.; Gilbert, J.A.; Viirre, E.; Knight, R. Correction for Gonzalez et al., “Migraines Are Correlated with Higher Levels of Nitrate-, Nitrite-, and Nitric Oxide-Reducing Oral Microbes in the American Gut Project Cohort”. MSystems 2017, 2, e00023-17. [Google Scholar] [CrossRef]
- Farrokhi, V.; Nemati, R.; Nichols, F.C.; Yao, X.; Anstadt, E.; Fujiwara, M.; Grady, J.; Wakefield, D.; Castro, W.; Donaldson, J.; et al. Bacterial lipodipeptide, Lipid 654, is a microbiome-associated biomarker for multiple sclerosis. Clin. Transl. Immunol. 2013, 2, e8. [Google Scholar] [CrossRef]
- Lau, H.C.; Lee, I.K.; Ko, P.W.; Lee, H.W.; Huh, J.S.; Cho, W.J.; Lim, J.O. Non-invasive screening for Alzheimer’s disease by sensing salivary sugar using Drosophila cells expressing gustatory receptor (Gr5a) immobilized on an extended gate ion-sensitive field-effect transistor (EG-ISFET) biosensor. PLoS ONE 2015, 10, e0117810. [Google Scholar] [CrossRef]
- Cabras, T.; Sanna, M.; Manconi, B.; Fanni, D.; Demelia, L.; Sorbello, O.; Iavarone, F.; Castagnola, M.; Faa, G.; Messana, I. Proteomic investigation of whole saliva in Wilson’s disease. J. Proteom. 2015, 128, 154–163. [Google Scholar] [CrossRef] [PubMed]
- Vivacqua, G.; Latorre, A.; Suppa, A.; Nardi, M.; Pietracupa, S.; Mancinelli, R.; Fabbrini, G.; Colosimo, C.; Gaudio, E.; Berardelli, A. Abnormal Salivary Total and Oligomeric Alpha-Synuclein in Parkinson’s Disease. PLoS ONE 2016, 11, e0151156. [Google Scholar] [CrossRef] [PubMed]
- Devic, I.; Hwang, H.; Edgar, J.S.; Izutsu, K.; Presland, R.; Pan, C.; Goodlett, D.R.; Wang, Y.; Armaly, J.; Tumas, V.; et al. Salivary alpha-synuclein and DJ-1: Potential biomarkers for Parkinson’s disease. Brain 2011, 134, e178. [Google Scholar] [CrossRef] [PubMed]
- Corey-Bloom, J.; Haque, A.S.; Park, S.; Nathan, A.S.; Baker, R.W.; Thomas, E.A. Salivary levels of total huntingtin are elevated in Huntington’s disease patients. Sci. Rep. 2018, 8, 7371. [Google Scholar] [CrossRef]
- Bermejo-Pareja, F.; Antequera, D.; Vargas, T.; Molina, J.A.; Carro, E. Saliva levels of Abeta1-42 as potential biomarker of Alzheimer’s disease: A pilot study. BMC Neurol. 2010, 10, 108. [Google Scholar] [CrossRef]
- Di Pietro, V.; Porto, E.; Ragusa, M.; Barbagallo, C.; Davies, D.; Forcione, M.; Logan, A.; Di Pietro, C.; Purrello, M.; Grey, M.; et al. Salivary MicroRNAs: Diagnostic Markers of Mild Traumatic Brain Injury in Contact-Sport. Front. Mol. Neurosci. 2018, 11, 290. [Google Scholar] [CrossRef]
- Hicks, S.D.; Ignacio, C.; Gentile, K.; Middleton, F.A. Salivary miRNA profiles identify children with autism spectrum disorder, correlate with adaptive behavior, and implicate ASD candidate genes involved in neurodevelopment. BMC Pediatrics 2016, 16, 52. [Google Scholar] [CrossRef]
- Griffiths, J.A.; Mazmanian, S.K. Emerging evidence linking the gut microbiome to neurologic disorders. Genome Med. 2018, 10, 98. [Google Scholar] [CrossRef]
- Bian, S.; Sun, T. Functions of noncoding RNAs in neural development and neurological diseases. Mol. Neurobiol. 2011, 44, 359–373. [Google Scholar] [CrossRef]
- Rajman, M.; Schratt, G. MicroRNAs in neural development: From master regulators to fine-tuners. Development 2017, 144, 2310–2322. [Google Scholar] [CrossRef]
- Sun, E.; Shi, Y. MicroRNAs: Small molecules with big roles in neurodevelopment and diseases. Exp. Neurol. 2015, 268, 46–53. [Google Scholar] [CrossRef] [PubMed]
- Martino, S.; di Girolamo, I.; Orlacchio, A.; Datti, A.; Orlacchio, A. MicroRNA implications across neurodevelopment and neuropathology. J. Biomed. Biotechnol. 2009, 2009, 654346. [Google Scholar] [CrossRef] [PubMed]
- Xu, B.; Karayiorgou, M.; Gogos, J.A. MicroRNAs in psychiatric and neurodevelopmental disorders. Brain Res. 2010, 1338, 78–88. [Google Scholar] [CrossRef] [PubMed]
- Rizzo, R.; Ragusa, M.; Barbagallo, C.; Sammito, M.; Gulisano, M.; Cali, P.V.; Pappalardo, C.; Barchitta, M.; Granata, M.; Condorelli, A.G.; et al. Circulating miRNAs profiles in Tourette syndrome: Molecular data and clinical implications. Mol. Brain 2015, 8, 44. [Google Scholar] [CrossRef] [PubMed]
- Cirnigliaro, M.; Barbagallo, C.; Gulisano, M.; Domini, C.N.; Barone, R.; Barbagallo, D.; Ragusa, M.; Di Pietro, C.; Rizzo, R.; Purrello, M. Expression and Regulatory Network Analysis of miR-140-3p, a New Potential Serum Biomarker for Autism Spectrum Disorder. Front. Mol. Neurosci. 2017, 10, 250. [Google Scholar] [CrossRef]
- Rajgor, D. Macro roles for microRNAs in neurodegenerative diseases. Non-Coding RNA Res. 2018, 3, 154–159. [Google Scholar] [CrossRef] [PubMed]
- Sheinerman, K.S.; Toledo, J.B.; Tsivinsky, V.G.; Irwin, D.; Grossman, M.; Weintraub, D.; Hurtig, H.I.; Chen-Plotkin, A.; Wolk, D.A.; McCluskey, L.F.; et al. Circulating brain-enriched microRNAs as novel biomarkers for detection and differentiation of neurodegenerative diseases. Alzheimer’s Res. Ther. 2017, 9, 89. [Google Scholar] [CrossRef]
- Hicks, S.D.; Middleton, F.A. A Comparative Review of microRNA Expression Patterns in Autism Spectrum Disorder. Front. Psychiatry 2016, 7, 176. [Google Scholar] [CrossRef]
- Hicks, S.D.; Rajan, A.T.; Wagner, K.E.; Barns, S.; Carpenter, R.L.; Middleton, F.A. Validation of a Salivary RNA Test for Childhood Autism Spectrum Disorder. Front. Genet. 2018, 9, 534. [Google Scholar] [CrossRef]
- Qiao, Y.; Wu, M.; Feng, Y.; Zhou, Z.; Chen, L.; Chen, F. Alterations of oral microbiota distinguish children with autism spectrum disorders from healthy controls. Sci. Rep. 2018, 8, 1597. [Google Scholar] [CrossRef]
- Hicks, S.D.; Uhlig, R.; Afshari, P.; Williams, J.; Chroneos, M.; Tierney-Aves, C.; Wagner, K.; Middleton, F.A. Oral microbiome activity in children with autism spectrum disorder. Autism Res. 2018, 11, 1286–1299. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Quinque, D.; Horz, H.P.; Li, M.; Rzhetskaya, M.; Raff, J.A.; Hayes, M.G.; Stoneking, M. Comparative analysis of the human saliva microbiome from different climate zones: Alaska, Germany, and Africa. BMC Microbiol. 2014, 14, 316. [Google Scholar] [CrossRef] [PubMed]
- Sarkar, A.; Stoneking, M.; Nandineni, M.R. Unraveling the human salivary microbiome diversity in Indian populations. PLoS ONE 2017, 12, e0184515. [Google Scholar] [CrossRef]
- Shaw, L.; Ribeiro, A.L.R.; Levine, A.P.; Pontikos, N.; Balloux, F.; Segal, A.W.; Roberts, A.P.; Smith, A.M. The Human Salivary Microbiome Is Shaped by Shared Environment Rather than Genetics: Evidence from a Large Family of Closely Related Individuals. mBio 2017, 8, e01237-17. [Google Scholar] [CrossRef] [PubMed]
- Zaura, E.; Brandt, B.W.; Prodan, A.; Teixeira de Mattos, M.J.; Imangaliyev, S.; Kool, J.; Buijs, M.J.; Jagers, F.L.; Hennequin-Hoenderdos, N.L.; Slot, D.E.; et al. On the ecosystemic network of saliva in healthy young adults. ISME J. 2017, 11, 1218–1231. [Google Scholar] [CrossRef]
- Park, N.J.; Zhou, H.; Elashoff, D.; Henson, B.S.; Kastratovic, D.A.; Abemayor, E.; Wong, D.T. Salivary microRNA: Discovery, characterization, and clinical utility for oral cancer detection. Clin. Cancer Res. 2009, 15, 5473–5477. [Google Scholar] [CrossRef]
- Ramanan, P.; Barreto, J.N.; Osmon, D.R.; Tosh, P.K. Rothia bacteremia: A 10-year experience at Mayo Clinic, Rochester, Minnesota. J. Clin. Microbiol. 2014, 52, 3184–3189. [Google Scholar] [CrossRef]
- O’Neil, C.R.; Wilson, E.; Missaghi, B. Bone and Joint Infections due to Haemophilus parainfluenzae: Case Report and Review of the Literature. Can. J. Infect. Dis. Med. Microbiol. 2016, 2016, 4503025. [Google Scholar]
- Bell, J.S.; Spencer, J.I.; Yates, R.L.; Yee, S.A.; Jacobs, B.M.; DeLuca, G.C. Invited Review: From nose to gut—the role of the microbiome in neurological disease. Neuropathol. Appl. Neurobiol. 2019, 45, 195–215. [Google Scholar] [CrossRef]
- Harding, A.; Gonder, U.; Robinson, S.J.; Crean, S.; Singhrao, S.K. Exploring the Association between Alzheimer’s Disease, Oral Health, Microbial Endocrinology and Nutrition. Front. Aging Neurosci. 2017, 9, 398. [Google Scholar] [CrossRef]
- Marchesan, J.T.; Morelli, T.; Moss, K.; Barros, S.P.; Ward, M.; Jenkins, W.; Aspiras, M.B.; Offenbacher, S. Association of Synergistetes and Cyclodipeptides with Periodontitis. J. Dent. Res. 2015, 94, 1425–1431. [Google Scholar] [CrossRef] [PubMed]
- Meuric, V.; Le Gall-David, S.; Boyer, E.; Acuna-Amador, L.; Martin, B.; Fong, S.B.; Barloy-Hubler, F.; Bonnaure-Mallet, M. Signature of Microbial Dysbiosis in Periodontitis. Appl. Environ. Microbiol. 2017, 83, e00462-17. [Google Scholar] [CrossRef] [PubMed]
- Muller, M.; Jakel, L.; Bruinsma, I.B.; Claassen, J.A.; Kuiperij, H.B.; Verbeek, M.M. MicroRNA-29a Is a Candidate Biomarker for Alzheimer’s Disease in Cell-Free Cerebrospinal Fluid. Mol. Neurobiol. 2016, 53, 2894–2899. [Google Scholar] [CrossRef] [PubMed]
- McKeever, P.M.; Schneider, R.; Taghdiri, F.; Weichert, A.; Multani, N.; Brown, R.A.; Boxer, A.L.; Karydas, A.; Miller, B.; Robertson, J.; et al. MicroRNA Expression Levels Are Altered in the Cerebrospinal Fluid of Patients with Young-Onset Alzheimer’s Disease. Mol. Neurobiol. 2018, 55, 8826–8841. [Google Scholar] [CrossRef]
- Lugli, G.; Cohen, A.M.; Bennett, D.A.; Shah, R.C.; Fields, C.J.; Hernandez, A.G.; Smalheiser, N.R. Plasma Exosomal miRNAs in Persons with and without Alzheimer Disease: Altered Expression and Prospects for Biomarkers. PLoS ONE 2015, 10, e0139233. [Google Scholar] [CrossRef]
- Margis, R.; Margis, R.; Rieder, C.R. Identification of blood microRNAs associated to Parkinsonis disease. J. Biotechnol. 2011, 152, 96–101. [Google Scholar] [CrossRef]
- Dong, H.; Wang, C.; Lu, S.; Yu, C.; Huang, L.; Feng, W.; Xu, H.; Chen, X.; Zen, K.; Yan, Q.; et al. A panel of four decreased serum microRNAs as a novel biomarker for early Parkinson’s disease. Biomarkers 2016, 21, 129–137. [Google Scholar] [CrossRef]
- Johnson, J.J.; Loeffert, A.C.; Stokes, J.; Olympia, R.P.; Bramley, H.; Hicks, S.D. Association of Salivary MicroRNA Changes With Prolonged Concussion Symptoms. JAMA Pediatrics 2018, 172, 65–73. [Google Scholar] [CrossRef]
- Huang, F.; Long, Z.; Chen, Z.; Li, J.; Hu, Z.; Qiu, R.; Zhuang, W.; Tang, B.; Xia, K.; Jiang, H. Investigation of Gene Regulatory Networks Associated with Autism Spectrum Disorder Based on MiRNA Expression in China. PLoS ONE 2015, 10, e0129052. [Google Scholar] [CrossRef]
- Sarachana, T.; Zhou, R.; Chen, G.; Manji, H.K.; Hu, V.W. Investigation of post-transcriptional gene regulatory networks associated with autism spectrum disorders by microRNA expression profiling of lymphoblastoid cell lines. Genome Med. 2010, 2, 23. [Google Scholar] [CrossRef]
- Mor, M.; Nardone, S.; Sams, D.S.; Elliott, E. Hypomethylation of miR-142 promoter and upregulation of microRNAs that target the oxytocin receptor gene in the autism prefrontal cortex. Mol. Autism 2015, 6, 46. [Google Scholar] [CrossRef] [PubMed]
- Onore, C.; Yang, H.; Van de Water, J.; Ashwood, P. Dynamic Akt/mTOR Signaling in Children with Autism Spectrum Disorder. Front. Pediatrics 2017, 5, 43. [Google Scholar] [CrossRef] [PubMed]
- de la Torre-Ubieta, L.; Won, H.; Stein, J.L.; Geschwind, D.H. Advancing the understanding of autism disease mechanisms through genetics. Nat. Med. 2016, 22, 345–361. [Google Scholar] [CrossRef]
- O’Connell, K.S.; McGregor, N.W.; Lochner, C.; Emsley, R.; Warnich, L. The genetic architecture of schizophrenia, bipolar disorder, obsessive-compulsive disorder and autism spectrum disorder. Mol. Cell. Neurosci. 2018, 88, 300–307. [Google Scholar] [CrossRef] [PubMed]
- Lathia, J.D.; Mattson, M.P.; Cheng, A. Notch: From neural development to neurological disorders. J. Neurochem. 2008, 107, 1471–1481. [Google Scholar] [CrossRef]
- Hwang, I.; Oh, H.; Santo, E.; Kim, D.Y.; Chen, J.W.; Bronson, R.T.; Locasale, J.W.; Na, Y.; Lee, J.; Reed, S.; et al. FOXO protects against age-progressive axonal degeneration. Aging Cell 2018, 17, e12701. [Google Scholar] [CrossRef] [PubMed]
- Chao, M.V.; Rajagopal, R.; Lee, F.S. Neurotrophin signalling in health and disease. Clin. Sci. 2006, 110, 167–173. [Google Scholar] [CrossRef]
- Verma, R.; Ritzel, R.M.; Harris, N.M.; Lee, J.; Kim, T.; Pandi, G.; Vemuganti, R.; McCullough, L.D. Inhibition of miR-141-3p Ameliorates the Negative Effects of Poststroke Social Isolation in Aged Mice. Stroke 2018, 49, 1701–1707. [Google Scholar] [CrossRef]
- Yang, L.; Engeland, C.G.; Cheng, B. Social isolation impairs oral palatal wound healing in sprague-dawley rats: A role for miR-29 and miR-203 via VEGF suppression. PLoS ONE 2013, 8, e72359. [Google Scholar] [CrossRef]
- Wyczechowska, D.; Lin, H.Y.; LaPlante, A.; Jeansonne, D.; Lassak, A.; Parsons, C.H.; Molina, P.E.; Peruzzi, F. A miRNA Signature for Cognitive Deficits and Alcohol Use Disorder in Persons Living with HIV/AIDS. Front. Mol. Neurosci. 2017, 10, 385. [Google Scholar] [CrossRef]
- Gidron, Y.; De Zwaan, M.; Quint, K.; Ocker, M. Influence of stress and health-behaviour on miRNA expression. Mol. Med. Rep. 2010, 3, 455–457. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Kuang, W.H.; Dong, Z.Q.; Tian, L.T.; Li, J. MicroRNA-451a, microRNA-34a-5p, and microRNA-221-3p as predictors of response to antidepressant treatment. Braz. J. Med. Biol. Res. 2018, 51, e7212. [Google Scholar] [CrossRef]
- Cassidy, S.A.; Bradley, L.; Bowen, E.; Wigham, S.; Rodgers, J. Measurement properties of tools used to assess depression in adults with and without autism spectrum conditions: A systematic review. Autism Res. 2018, 11, 738–754. [Google Scholar] [CrossRef] [PubMed]
- Khemakhem, A.M.; Frye, R.E.; El-Ansary, A.; Al-Ayadhi, L.; Bacha, A.B. Novel biomarkers of metabolic dysfunction is autism spectrum disorder: Potential for biological diagnostic markers. Metab. Brain Dis. 2017, 32, 1983–1997. [Google Scholar] [CrossRef] [PubMed]
- Ji, J.; Qin, Y.; Ren, J.; Lu, C.; Wang, R.; Dai, X.; Zhou, R.; Huang, Z.; Xu, M.; Chen, M.; et al. Mitochondria-related miR-141-3p contributes to mitochondrial dysfunction in HFD-induced obesity by inhibiting PTEN. Sci. Rep. 2015, 5, 16262. [Google Scholar] [CrossRef]
- Meltzer, S.; Bjornetro, T.; Lyckander, L.G.; Flatmark, K.; Dueland, S.; Samiappan, R.; Johansen, C.; Kalanxhi, E.; Ree, A.H.; Redalen, K.R. Corrigendum to “Circulating Exosomal miR-141-3p and miR-375 in Metastatic Progression of Rectal Cancer” [Transl Oncol 12 (8) (2019) 1038-1044]. Transl. Oncol. 2020, 13, 122–124. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Xin, H.; Fen, T. Function of microRNA141 in human breast cancer through cytotoxic CD4+ T cells regulated by MAP4K4 expression. Mol. Med. Rep. 2018, 17, 7893–7901. [Google Scholar]
- He, X.; McLean, J.S.; Edlund, A.; Yooseph, S.; Hall, A.P.; Liu, S.Y.; Dorrestein, P.C.; Esquenazi, E.; Hunter, R.C.; Cheng, G.; et al. Cultivation of a human-associated TM7 phylotype reveals a reduced genome and epibiotic parasitic lifestyle. Proc. Natl. Acad. Sci. USA 2015, 112, 244–249. [Google Scholar] [CrossRef]
- Shi, B.; Chang, M.; Martin, J.; Mitreva, M.; Lux, R.; Klokkevold, P.; Sodergren, E.; Weinstock, G.M.; Haake, S.K.; Li, H. Dynamic changes in the subgingival microbiome and their potential for diagnosis and prognosis of periodontitis. MBio 2015, 6, e01926-14. [Google Scholar] [CrossRef]
- Kamer, A.R.; Morse, D.E.; Holm-Pedersen, P.; Mortensen, E.L.; Avlund, K. Periodontal inflammation in relation to cognitive function in an older adult Danish population. J. Alzheimer’s Dis. 2012, 28, 613–624. [Google Scholar] [CrossRef]
- Bui, F.Q.; Almeida-da-Silva, C.L.C.; Huynh, B.; Trinh, A.; Liu, J.; Woodward, J.; Asadi, H.; Ojcius, D.M. Association between periodontal pathogens and systemic disease. Biomed. J. 2019, 42, 27–35. [Google Scholar] [CrossRef] [PubMed]
- Stoecklin-Wasmer, C.; Guarnieri, P.; Celenti, R.; Demmer, R.T.; Kebschull, M.; Papapanou, P.N. MicroRNAs and their target genes in gingival tissues. J. Dent. Res. 2012, 91, 934–940. [Google Scholar] [CrossRef] [PubMed]
- Moloney, G.M.; Viola, M.F.; Hoban, A.E.; Dinan, T.G.; Cryan, J.F. Faecal microRNAs: Indicators of imbalance at the host-microbe interface? Benef. Microbes 2018, 9, 175–183. [Google Scholar] [CrossRef] [PubMed]
- Yuan, C.; Burns, M.B.; Subramanian, S.; Blekhman, R. Interaction between Host MicroRNAs and the Gut Microbiota in Colorectal Cancer. MSystems 2018, 3, e00205-17. [Google Scholar] [CrossRef]
- Liu, S.; da Cunha, A.P.; Rezende, R.M.; Cialic, R.; Wei, Z.; Bry, L.; Comstock, L.E.; Gandhi, R.; Weiner, H.L. The Host Shapes the Gut Microbiota via Fecal MicroRNA. Cell Host Microbe 2016, 19, 32–43. [Google Scholar] [CrossRef] [PubMed]
- Schulte, L.N.; Eulalio, A.; Mollenkopf, H.J.; Reinhardt, R.; Vogel, J. Analysis of the host microRNA response to Salmonella uncovers the control of major cytokines by the let-7 family. EMBO J. 2011, 30, 1977–1989. [Google Scholar] [CrossRef]
- Hoeke, L.; Sharbati, J.; Pawar, K.; Keller, A.; Einspanier, R.; Sharbati, S. Intestinal Salmonella typhimurium infection leads to miR-29a induced caveolin 2 regulation. PLoS ONE 2013, 8, e67300. [Google Scholar] [CrossRef]
- Maudet, C.; Mano, M.; Sunkavalli, U.; Sharan, M.; Giacca, M.; Forstner, K.U.; Eulalio, A. Functional high-throughput screening identifies the miR-15 microRNA family as cellular restriction factors for Salmonella infection. Nat. Commun. 2014, 5, 4718. [Google Scholar] [CrossRef]
- Sharbati, J.; Lewin, A.; Kutz-Lohroff, B.; Kamal, E.; Einspanier, R.; Sharbati, S. Integrated microRNA-mRNA-analysis of human monocyte derived macrophages upon Mycobacterium avium subsp. hominissuis infection. PLoS ONE 2011, 6, e20258. [Google Scholar] [CrossRef]
- Fu, Y.; Yi, Z.; Wu, X.; Li, J.; Xu, F. Circulating microRNAs in patients with active pulmonary tuberculosis. J. Clin. Microbiol. 2011, 49, 4246–4251. [Google Scholar] [CrossRef]
- Yamada, M.; Gomez, J.C.; Chugh, P.E.; Lowell, C.A.; Dinauer, M.C.; Dittmer, D.P.; Doerschuk, C.M. Interferon-gamma production by neutrophils during bacterial pneumonia in mice. Am. J. Respir. Crit. Care Med. 2011, 183, 1391–1401. [Google Scholar] [CrossRef] [PubMed]
- Matsushima, K.; Isomoto, H.; Inoue, N.; Nakayama, T.; Hayashi, T.; Nakayama, M.; Nakao, K.; Hirayama, T.; Kohno, S. MicroRNA signatures in Helicobacter pylori-infected gastric mucosa. Int. J. Cancer 2011, 128, 361–370. [Google Scholar] [CrossRef]
- Ngounou Wetie, A.G.; Wormwood, K.L.; Russell, S.; Ryan, J.P.; Darie, C.C.; Woods, A.G. A Pilot Proteomic Analysis of Salivary Biomarkers in Autism Spectrum Disorder. Autism Res. 2015, 8, 338–350. [Google Scholar] [CrossRef] [PubMed]
- Koyama, T.; Inada, N.; Tsujii, H.; Kurita, H. Predicting children with pervasive developmental disorders using the Wechsler Intelligence Scale for Children-Third Edition. Psychiatry Clin. Neurosci. 2008, 62, 476–478. [Google Scholar] [CrossRef] [PubMed]
- Wechsler, D. WPPSI: Wechsler Preschool and Primary Scale of Intelligence; Psychological Corporation: New York, NY, USA, 1967. [Google Scholar]
- Ragusa, M.; Barbagallo, C.; Statello, L.; Caltabiano, R.; Russo, A.; Puzzo, L.; Avitabile, T.; Longo, A.; Toro, M.D.; Barbagallo, D.; et al. miRNA profiling in vitreous humor, vitreal exosomes and serum from uveal melanoma patients: Pathological and diagnostic implications. Cancer Biol. Ther. 2015, 16, 1387–1396. [Google Scholar] [CrossRef] [PubMed]
- Park, T.; Yi, S.G.; Kang, S.H.; Lee, S.; Lee, Y.S.; Simon, R. Evaluation of normalization methods for microarray data. BMC Bioinform. 2003, 4, 33. [Google Scholar] [CrossRef] [PubMed]
- Schmittgen, T.D.; Livak, K.J. Analyzing real-time PCR data by the comparative C (T) method. Nat. Protoc. 2008, 3, 1101–1108. [Google Scholar] [CrossRef]
- Klindworth, A.; Pruesse, E.; Schweer, T.; Peplies, J.; Quast, C.; Horn, M.; Glockner, F.O. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 2013, 41, e1. [Google Scholar] [CrossRef]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Pena, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef]
- Magoc, T.; Salzberg, S.L. FLASH: Fast length adjustment of short reads to improve genome assemblies. Bioinformatics 2011, 27, 2957–2963. [Google Scholar] [CrossRef]
- DeSantis, T.Z.; Hugenholtz, P.; Larsen, N.; Rojas, M.; Brodie, E.L.; Keller, K.; Huber, T.; Dalevi, D.; Hu, P.; Andersen, G.L. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl. Environ. Microbiol. 2006, 72, 5069–5072. [Google Scholar] [CrossRef] [PubMed]
- Robertson, C.E.; Harris, J.K.; Wagner, B.D.; Granger, D.; Browne, K.; Tatem, B.; Feazel, L.M.; Park, K.; Pace, N.R.; Frank, D.N. Explicet: Graphical user interface software for metadata-driven management, analysis and visualization of microbiome data. Bioinformatics 2013, 29, 3100–3101. [Google Scholar] [CrossRef] [PubMed]
- Karazsia, B.T.; van Dulmen, M.H. Regression models for count data: Illustrations using longitudinal predictors of childhood injury. J. Pediatric Psychol 2008, 33, 1076–1084. [Google Scholar] [CrossRef] [PubMed]
- Vlachos, I.S.; Zagganas, K.; Paraskevopoulou, M.D.; Georgakilas, G.; Karagkouni, D.; Vergoulis, T.; Dalamagas, T.; Hatzigeorgiou, A.G. DIANA-miRPath v3.0: Deciphering microRNA function with experimental support. Nucleic Acids Res. 2015, 43, W460–W466. [Google Scholar] [CrossRef]
- Fan, Y.; Xia, J. miRNet-Functional Analysis and Visual Exploration of miRNA-Target Interactions in a Network Context. Methods Mol. Biol. 2018, 1819, 215–233. [Google Scholar] [PubMed]
- White, J.R.; Nagarajan, N.; Pop, M. Statistical methods for detecting differentially abundant features in clinical metagenomic samples. PLoS Comput. Biol. 2009, 5, e1000352. [Google Scholar] [CrossRef]
- Parks, D.H.; Tyson, G.W.; Hugenholtz, P.; Beiko, R.G. STAMP: Statistical analysis of taxonomic and functional profiles. Bioinformatics 2014, 30, 3123–3124. [Google Scholar] [CrossRef]
- Parks, D.H.; Beiko, R.G. Identifying biologically relevant differences between metagenomic communities. Bioinformatics 2010, 26, 715–721. [Google Scholar] [CrossRef]
- Cliff, N. Dominance statistics: Ordinal analyses to answer ordinal questions. Psychol. Bull. 1993, 114, 494–509. [Google Scholar] [CrossRef]
- Cliff, N. Answering Ordinal Questions with Ordinal Data Using Ordinal Statistics. Multivar. Behav. Res. 1996, 31, 331–350. [Google Scholar] [CrossRef]
- Macbeth, G.; Razumiejczyk, E.; Ledesma, R.D.U. Cliff’s Delta Calculator: A non-parametric effect size program for two groups of observations. Univ. Psychol. 2011, 10, 545–555. [Google Scholar] [CrossRef]
- Kotrlik, J.; Williams, H.; Jabor, K. Reporting and Interpreting Effect Size in Quantitative Agricultural Education Research. J. Agric. Educ. 2011, 52, 132–142. [Google Scholar] [CrossRef]

| ASD | NUC | |
|---|---|---|
| Number of Participants | 76 | 39 |
| Sex (M:F) | 60:16 | 28:11 |
| Age (years) | 6.9 (±1.5) | 6.9 (±1.8) |
| Time of collection (h) | 09:58 (±00:28) | 10:03 (±00:30) |
| Time since last meal (h) | 2.99 (±0.12) | 2.98 (±0.13) |
| IQ | ||
| TIQ | 69.6 (±19.3) | 96.7 (±11.6) |
| VIQ | 67.8 (±19.7) | 96.2 (±13.3) |
| PIQ | 71 (±21.4) | 97.9 (±12.7) |
| ADOS | ||
| A | 4.6 (±1.9) | 0 |
| B | 7.7 (±2.4) | 0 |
| C | 2.1 (±1.2) | 0 |
| D | 2.6 (±1.3) | 0 |
| ADI-R | ||
| A | 10.9 (±3.7) | 0 |
| B | 9.4 (±2.5) | 0 |
| C | 5.9 (±2.8) | 0 |
| D | 3.2 (±1.3) | 0 |
| Prolactine | 229.8 (±89.5) | / |
| Ceruloplasmin | 27.6 (±5.6) | / |
| Lactate | 15.3 (±4.8) | / |
| Ammonium | 24.7 (±9.3) | / |
| TSH | 2.3 (±0.9) | / |
| DE miRNA ASD vs. NUC | FC | Mann–Whitney Test p-Value |
|---|---|---|
| let-7b-5p | −1.99 | 0.0002 |
| miR-16-5p | −1.68 | 0.0002 |
| miR-29a-3p | 1.43 | 0.0123 |
| miR-141-3p | 2.93 | 0.0431 |
| miR-451a | −3.58 | <0.0001 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ragusa, M.; Santagati, M.; Mirabella, F.; Lauretta, G.; Cirnigliaro, M.; Brex, D.; Barbagallo, C.; Domini, C.N.; Gulisano, M.; Barone, R.; et al. Potential Associations Among Alteration of Salivary miRNAs, Saliva Microbiome Structure, and Cognitive Impairments in Autistic Children. Int. J. Mol. Sci. 2020, 21, 6203. https://doi.org/10.3390/ijms21176203
Ragusa M, Santagati M, Mirabella F, Lauretta G, Cirnigliaro M, Brex D, Barbagallo C, Domini CN, Gulisano M, Barone R, et al. Potential Associations Among Alteration of Salivary miRNAs, Saliva Microbiome Structure, and Cognitive Impairments in Autistic Children. International Journal of Molecular Sciences. 2020; 21(17):6203. https://doi.org/10.3390/ijms21176203
Chicago/Turabian StyleRagusa, Marco, Maria Santagati, Federica Mirabella, Giovanni Lauretta, Matilde Cirnigliaro, Duilia Brex, Cristina Barbagallo, Carla Noemi Domini, Mariangela Gulisano, Rita Barone, and et al. 2020. "Potential Associations Among Alteration of Salivary miRNAs, Saliva Microbiome Structure, and Cognitive Impairments in Autistic Children" International Journal of Molecular Sciences 21, no. 17: 6203. https://doi.org/10.3390/ijms21176203
APA StyleRagusa, M., Santagati, M., Mirabella, F., Lauretta, G., Cirnigliaro, M., Brex, D., Barbagallo, C., Domini, C. N., Gulisano, M., Barone, R., Trovato, L., Oliveri, S., Mongelli, G., Spitale, A., Barbagallo, D., Di Pietro, C., Stefani, S., Rizzo, R., & Purrello, M. (2020). Potential Associations Among Alteration of Salivary miRNAs, Saliva Microbiome Structure, and Cognitive Impairments in Autistic Children. International Journal of Molecular Sciences, 21(17), 6203. https://doi.org/10.3390/ijms21176203







