Single Exon Skipping Can Address a Multi-Exon Duplication in the Dystrophin Gene
Abstract
1. Introduction
2. Results
3. Discussion
4. Materials and Methods
4.1. AO Design and Synthesis
4.2. Cell Propagation
4.3. Transfection
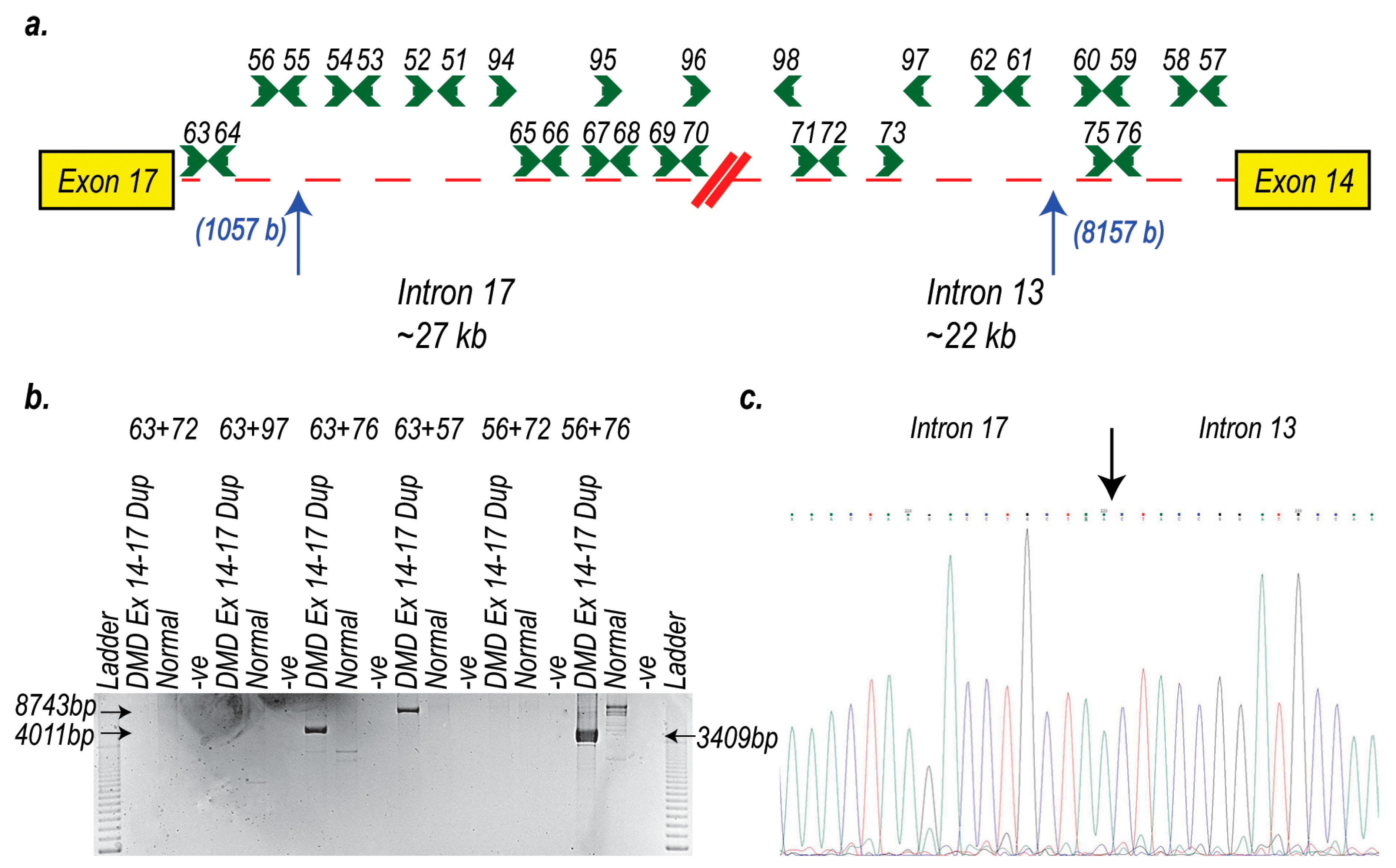
4.4. RNA Extraction and RT-PCR Assays
4.5. Gel Analysis and Imaging
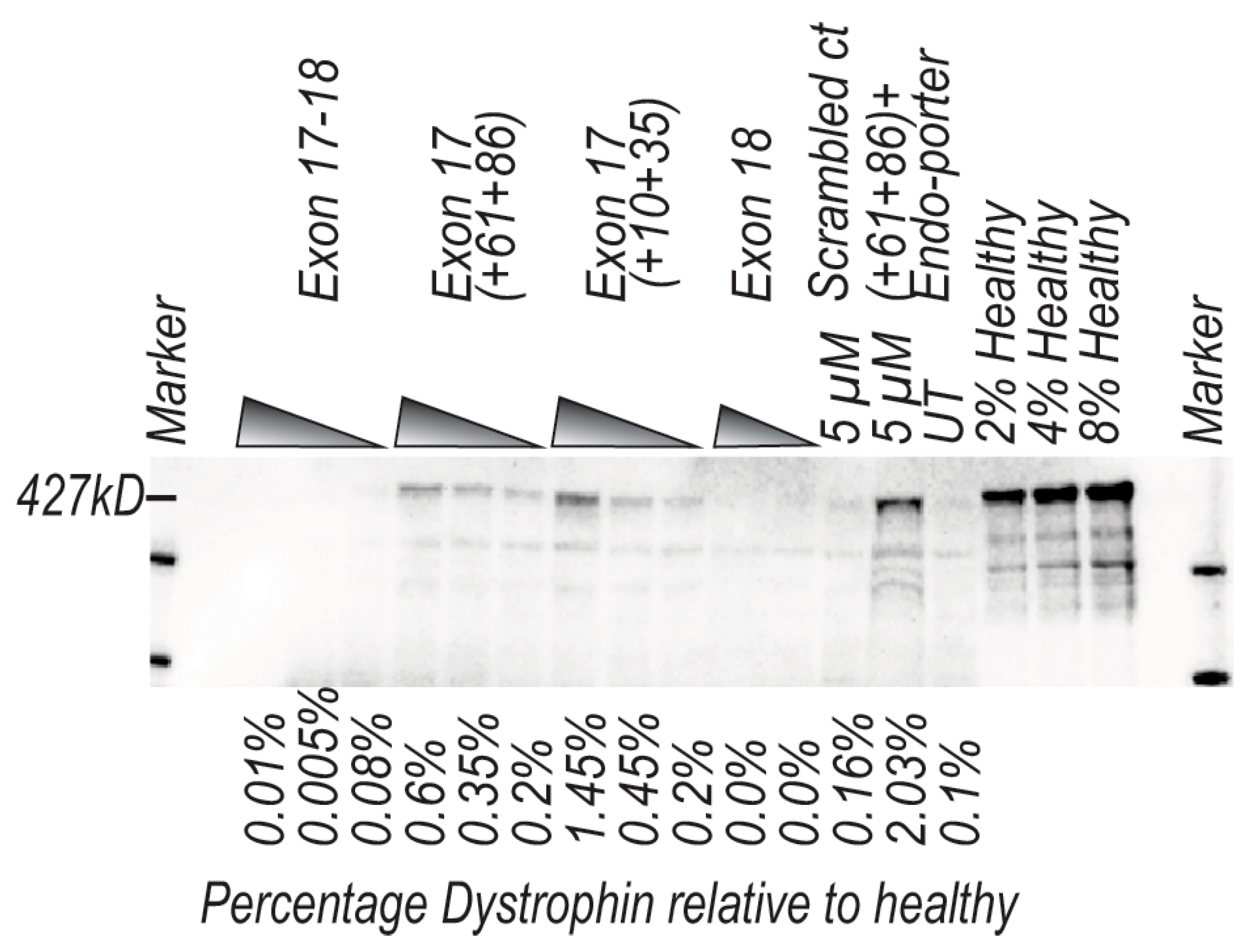
4.6. Western Blotting
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| DMD | Duchenne muscular dystrophy |
| BMD | Becker muscular dystrophy |
| AO | Antisense oligomer |
| PMO | Phosphorodiamidate morpholino oligomer |
| PPMO | Peptide conjugated phosphorodiamidate morpholino oligomer |
References
- Mendell, J.R.; Shilling, C.; Leslie, N.D.; Flanigan, K.M.; al-Dahhak, R.; Gastier-Foster, J.; Kneile, K.; Dunn, D.M.; Duval, B.; Aoyagi, A.; et al. Evidence-based path to newborn screening for Duchenne muscular dystrophy. Ann. Neurol. 2012, 71, 304–313. [Google Scholar] [CrossRef] [PubMed]
- Hoffman, E.P.; Brown, R.H., Jr.; Kunkel, L.M. Dystrophin: The protein product of the Duchenne muscular dystrophy locus. Cell 1987, 51, 919–928. [Google Scholar] [CrossRef]
- Hoffman, E.P.; Fischbeck, K.H.; Brown, R.H.; Johnson, M.; Medori, R.; Loike, J.D.; Harris, J.B.; Waterston, R.; Brooke, M.; Specht, L.; et al. Characterization of dystrophin in muscle-biopsy specimens from patients with Duchenne‘s or Becker‘s muscular dystrophy. N. Engl. J. Med. 1988, 318, 1363–1368. [Google Scholar] [CrossRef] [PubMed]
- Moser, H. Duchenne muscular dystrophy: Pathogenetic aspects and genetic prevention. Hum. Genet. 1984, 66, 17–40. [Google Scholar] [CrossRef] [PubMed]
- Koenig, M.; Beggs, A.H.; Moyer, M.; Scherpf, S.; Heindrich, K.; Bettecken, T.; Meng, G.; Muller, C.R.; Lindlof, M.; Kaariainen, H.; et al. The molecular basis for Duchenne versus Becker muscular dystrophy: Correlation of severity with type of deletion. Am. J. Hum. Genet. 1989, 45, 498–506. [Google Scholar] [PubMed]
- Koenig, M.; Monaco, A.P.; Kunkel, L.M. The complete sequence of dystrophin predicts a rod-shaped cytoskeletal protein. Cell 1988, 53, 219–228. [Google Scholar] [CrossRef]
- Monaco, A.P.; Bertelson, C.J.; Liechti-Gallati, S.; Moser, H.; Kunkel, L.M. An explanation for the phenotypic differences between patients bearing partial deletions of the DMD locus. Genomics 1988, 2, 90–95. [Google Scholar] [CrossRef]
- Aartsma-Rus, A.; Janson, A.A.; van Ommen, G.J.; van Deutekom, J.C. Antisense-induced exon skipping for duplications in Duchenne muscular dystrophy. BMC Med. Genet. 2007, 8, 1–9. [Google Scholar] [CrossRef]
- Mann, C.J.; Honeyman, K.; McClorey, G.; Fletcher, S.; Wilton, S.D. Improved antisense oligonucleotide induced exon skipping in the mdx mouse model of muscular dystrophy. J. Gene Med. 2002, 4, 644–654. [Google Scholar] [CrossRef]
- Mitrpant, C.; Fletcher, S.; Iversen, P.L.; Wilton, S.D. By-passing the nonsense mutation in the 4 CV mouse model of muscular dystrophy by induced exon skipping. J. Gene Med. 2009, 11, 46–56. [Google Scholar] [CrossRef]
- Aartsma-Rus, A.; De Winter, C.L.; Janson, A.A.; Kaman, W.E.; Van Ommen, G.J.; Den Dunnen, J.T.; Van Deutekom, J.C. Functional analysis of 114 exon-internal AONs for targeted DMD exon skipping: Indication for steric hindrance of SR protein binding sites. Oligonucleotides 2005, 15, 284–297. [Google Scholar] [CrossRef] [PubMed]
- Aartsma-Rus, A.; Janson, A.A.; Kaman, W.E.; Bremmer-Bout, M.; den Dunnen, J.T.; Baas, F.; van Ommen, G.J.; van Deutekom, J.C. Therapeutic antisense-induced exon skipping in cultured muscle cells from six different DMD patients. Hum. Mol. Genet. 2003, 12, 907–914. [Google Scholar] [CrossRef] [PubMed]
- Greer, K.L.; Lochmuller, H.; Flanigan, K.; Fletcher, S.; Wilton, S.D. Targeted exon skipping to correct exon duplications in the dystrophin gene. Mol. Nucleic Acids 2014, 3, e155. [Google Scholar] [CrossRef]
- Aartsma-Rus, A.; Van Deutekom, J.C.; Fokkema, I.F.; Van Ommen, G.J.; Den Dunnen, J.T. Entries in the Leiden Duchenne muscular dystrophy mutation database: An overview of mutation types and paradoxical cases that confirm the reading-frame rule. Muscle Nerve 2006, 34, 135–144. [Google Scholar] [CrossRef] [PubMed]
- Forrest, S.; Meloni, P.L.; Muntoni, F.; Kim, J.; Fletcher, S.; Wilton, S.D. Personalized exon skipping strategies to address clustered non-deletion dystrophin mutations. Neuromuscul Disord 2010, 20, 810–816. [Google Scholar] [CrossRef][Green Version]
- Wilton, S.D.; Fall, A.M.; Harding, P.L.; McClorey, G.; Coleman, C.; Fletcher, S. Antisense oligonucleotide-induced exon skipping across the human dystrophin gene transcript. Mol. Ther. 2007, 15, 1288–1296. [Google Scholar] [CrossRef]
- Moulton, H.M.; Moulton, J.D. Morpholinos and their peptide conjugates: Therapeutic promise and challenge for Duchenne muscular dystrophy. Biochim. Biophys. Acta 2010, 1798, 2296–2303. [Google Scholar] [CrossRef]
- Amantana, A.; Moulton, H.M.; Cate, M.L.; Reddy, M.T.; Whitehead, T.; Hassinger, J.N.; Youngblood, D.S.; Iversen, P.L. Pharmacokinetics, biodistribution, stability and toxicity of a cell-penetrating peptide-morpholino oligomer conjugate. Bioconjug. Chem. 2007, 18, 1325–1331. [Google Scholar] [CrossRef]
- Greco, S.; Cardinali, B.; Falcone, G.; Martelli, F. Circular RNAs in Muscle Function and Disease. Int. J. Mol. Sci. 2018, 19, 3454. [Google Scholar] [CrossRef]
- Suzuki, H.; Aoki, Y.; Kameyama, T.; Saito, T.; Masuda, S.; Tanihata, J.; Nagata, T.; Mayeda, A.; Takeda, S.; Tsukahara, T. Endogenous Multiple Exon Skipping and Back-Splicing at the DMD Mutation Hotspot. Int. J. Mol. Sci. 2016, 17, 1722. [Google Scholar] [CrossRef]
- Vulin, A.; Wein, N.; Simmons, T.R.; Rutherford, A.M.; Findlay, A.R.; Yurkoski, J.A.; Kaminoh, Y.; Flanigan, K.M. The first exon duplication mouse model of Duchenne muscular dystrophy: A tool for therapeutic development. Neuromuscul. Disord. 2015, 25, 827–834. [Google Scholar] [CrossRef]
- Greer, K.; Fletcher, S.; Wilton, S.D. Skipping of Duplicated Dystrophin Exons: In Vitro Induction and Assessment. Methods Mol. Biol. 2018, 1828, 219–228. [Google Scholar] [PubMed]
- Greer, K.; Mizzi, K.; Rice, E.; Kuster, L.; Barrero, R.A.; Bellgard, M.I.; Lynch, B.J.; Foley, A.R.; E., O.R.; Wilton, S.D.; et al. Pseudoexon activation increases phenotype severity in a Becker muscular dystrophy patient. Mol. Genet. Genom. Med. 2015, 3, 320–326. [Google Scholar] [CrossRef] [PubMed]
- Kunkel, T.A.; Bebenek, K. DNA replication fidelity. Annu Rev. Biochem 2000, 69, 497–529. [Google Scholar] [CrossRef] [PubMed]
- Panda, A.C.; Gorospe, M. Detection and Analysis of Circular RNAs by RT-PCR. Bio Protoc 2018, 8, e2775. [Google Scholar] [CrossRef]
- Suzuki, H.; Zuo, Y.; Wang, J.; Zhang, M.Q.; Malhotra, A.; Mayeda, A. Characterization of RNase R-digested cellular RNA source that consists of lariat and circular RNAs from pre-mRNA splicing. Nucleic Acids Res. 2006, 34, e63. [Google Scholar] [CrossRef] [PubMed]
- Cartegni, L.; Wang, J.; Zhu, Z.; Zhang, M.Q.; Krainer, A.R. ESEfinder: A web resource to identify exonic splicing enhancers. Nucleic Acids Res. 2003, 31, 3568–3571. [Google Scholar] [CrossRef]
- Rando, T.A.; Blau, H.M. Primary mouse myoblast purification, characterization, and transplantation for cell-mediated gene therapy. J. Cell Biol. 1994, 125, 1275–1287. [Google Scholar] [CrossRef]
- Harding, P.L.; Fall, A.M.; Honeyman, K.; Fletcher, S.; Wilton, S.D. The influence of antisense oligonucleotide length on dystrophin exon skipping. Mol 2007, 15, 157–166. [Google Scholar] [CrossRef]
- Adams, A.M.; Harding, P.L.; Iversen, P.L.; Coleman, C.; Fletcher, S.; Wilton, S.D. Antisense oligonucleotide induced exon skipping and the dystrophin gene transcript: cocktails and chemistries. Bmc Mol. Biol. 2007, 8, 1–8. [Google Scholar] [CrossRef]
- Toh, Z.Y.; Thandar Aung-Htut, M.; Pinniger, G.; Adams, A.M.; Krishnaswarmy, S.; Wong, B.L.; Fletcher, S.; Wilton, S.D. Deletion of Dystrophin In-Frame Exon 5 Leads to a Severe Phenotype: Guidance for Exon Skipping Strategies. PLoS ONE 2016, 11, e0145620. [Google Scholar] [CrossRef] [PubMed]
- Cooper, S.T.; Lo, H.P.; North, K.N. Single section Western blot: improving the molecular diagnosis of the muscular dystrophies. Neurology 2003, 61, 93–97. [Google Scholar] [CrossRef] [PubMed]
- Nicholson, L.V.; Davison, K.; Falkous, G.; Harwood, C.; O’Donnell, E.; Slater, C.R.; Harris, J.B. Dystrophin in skeletal muscle. I. Western blot analysis using a monoclonal antibody. J. Neurol. Sci. 1989, 94, 125–136. [Google Scholar] [CrossRef]
- Nicholson, L.V.; Johnson, M.A.; Davison, K.; O’Donnell, E.; Falkous, G.; Barron, M.; Harris, J.B. Dystrophin or a “related protein” in Duchenne muscular dystrophy? Acta Neurol. Scand. 1992, 86, 8–14. [Google Scholar] [CrossRef]
| Nomenclature | Sequence (5′-3′) |
|---|---|
| H17A(+10+35) | AGU GAU GGC UGA GUG GUG GUG ACA GC |
| H17A(+61+86) | UGU UCC CUU GUG GUC ACC GUA GUU AC |
| H18A(+24+53) | CAG CUU CUG AGC GAG UAA UCC AGC UGU GAA |
| Scrambled Sequence | GGA UGU CCU GAG UCU AGA CCC UCC G |
| Primer Location | Primer Sequence |
|---|---|
| Exon 13 Forward | TGC TTC AAG AAG ATC TAG AAC AAG AAC |
| Exon 13 Forward | CAC GCA ACT GCT GCT TTG GAA G |
| Exon 16 Forward | GCA AAC TGT ATT CAC TCA AAC |
| Exon 15 Reverse | GTG AAT CTT GTT CAC TGC ATC |
| Exon 19 Reverse | ACG TTC CAC CAG GGC CTG AG |
| Exon 20 Reverse | GAT ACT CCA GCC AGT TAA GTC T |
| Exon 21 Reverse | GGC CAC AAA GTC TGC ATC CAG |
| Exon 25 Reverse | GTC TCA AGT CTC GAA GCA AAC |
| Intron 17 Forward | TGG TGT TTT CTG GGG TGA AA |
| Intron 17 Forward | CCA GGG TAG AGA AGT TTG AA |
| Intron 13 Reverse | ACC ACT ATA TAA AAT GGC TCC |
| Intron 13 Reverse | GAG GGA GGC AGA AAA CAA CC |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Greer, K.; Johnsen, R.; Nevo, Y.; Fellig, Y.; Fletcher, S.; Wilton, S.D. Single Exon Skipping Can Address a Multi-Exon Duplication in the Dystrophin Gene. Int. J. Mol. Sci. 2020, 21, 4511. https://doi.org/10.3390/ijms21124511
Greer K, Johnsen R, Nevo Y, Fellig Y, Fletcher S, Wilton SD. Single Exon Skipping Can Address a Multi-Exon Duplication in the Dystrophin Gene. International Journal of Molecular Sciences. 2020; 21(12):4511. https://doi.org/10.3390/ijms21124511
Chicago/Turabian StyleGreer, Kane, Russell Johnsen, Yoram Nevo, Yakov Fellig, Susan Fletcher, and Steve D. Wilton. 2020. "Single Exon Skipping Can Address a Multi-Exon Duplication in the Dystrophin Gene" International Journal of Molecular Sciences 21, no. 12: 4511. https://doi.org/10.3390/ijms21124511
APA StyleGreer, K., Johnsen, R., Nevo, Y., Fellig, Y., Fletcher, S., & Wilton, S. D. (2020). Single Exon Skipping Can Address a Multi-Exon Duplication in the Dystrophin Gene. International Journal of Molecular Sciences, 21(12), 4511. https://doi.org/10.3390/ijms21124511





