Regulation of Autophagy by Protein Kinase C-ε in Breast Cancer Cells
Abstract
1. Introduction
2. Results
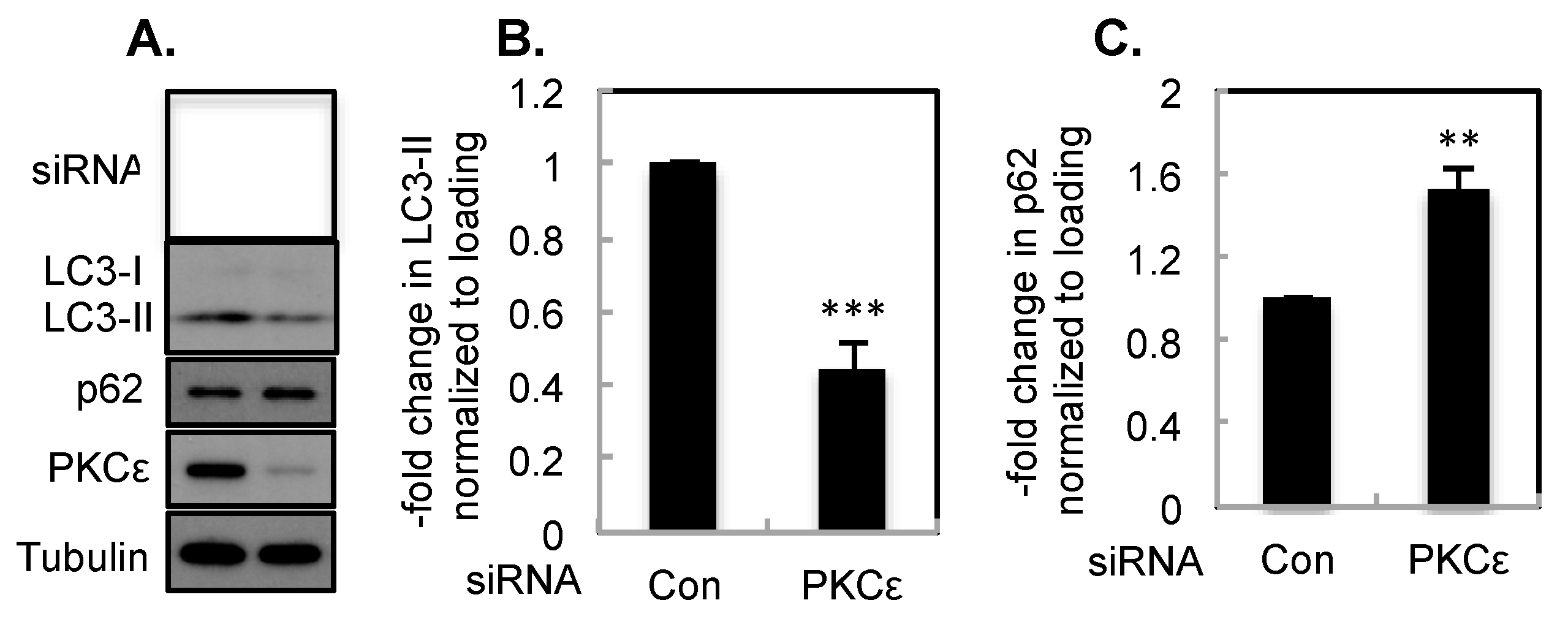
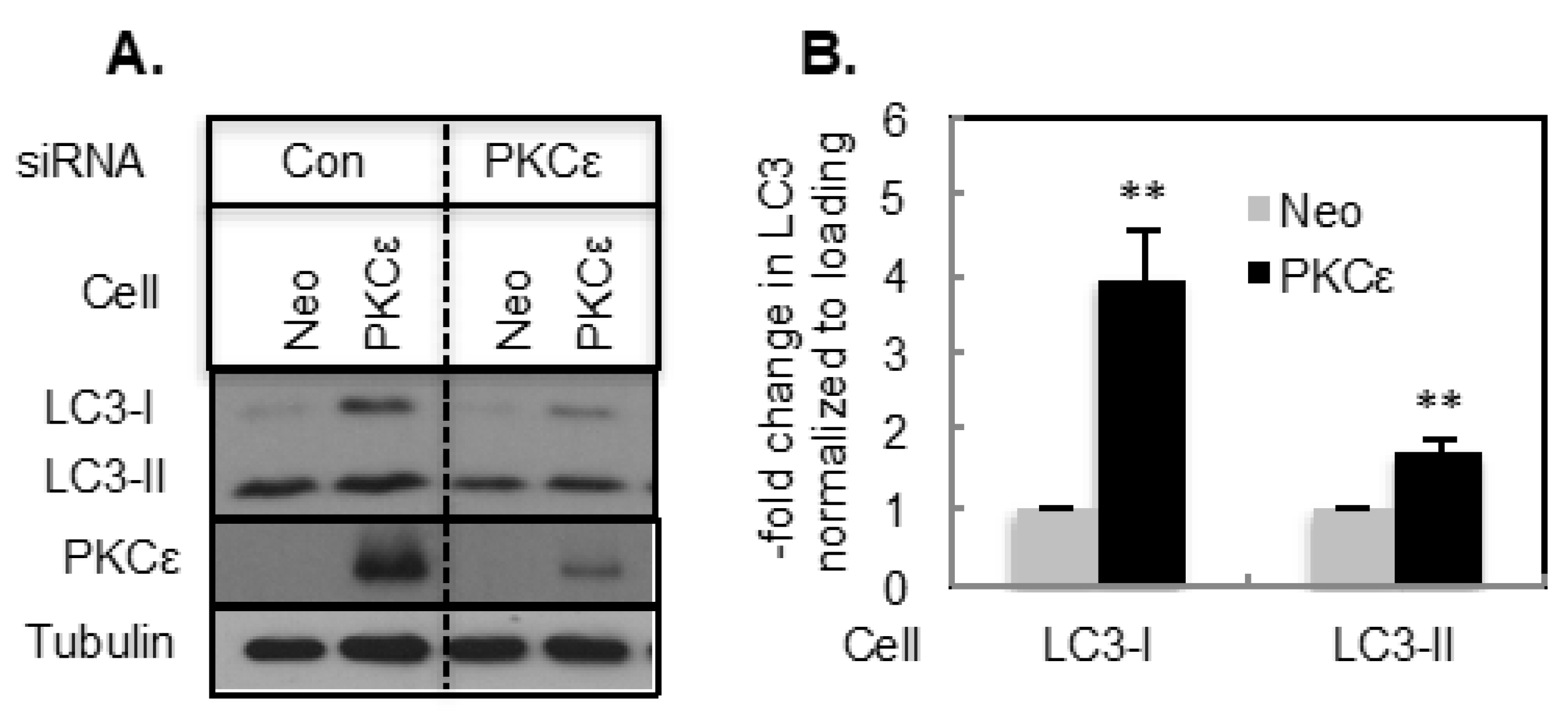
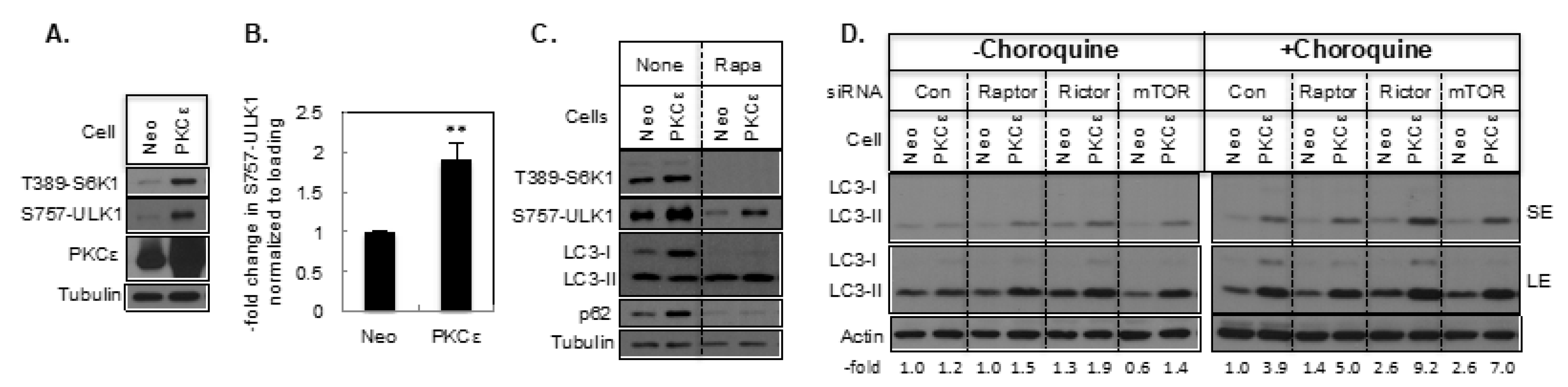
2.1. The Effect of PKCε on Basal Autophagy
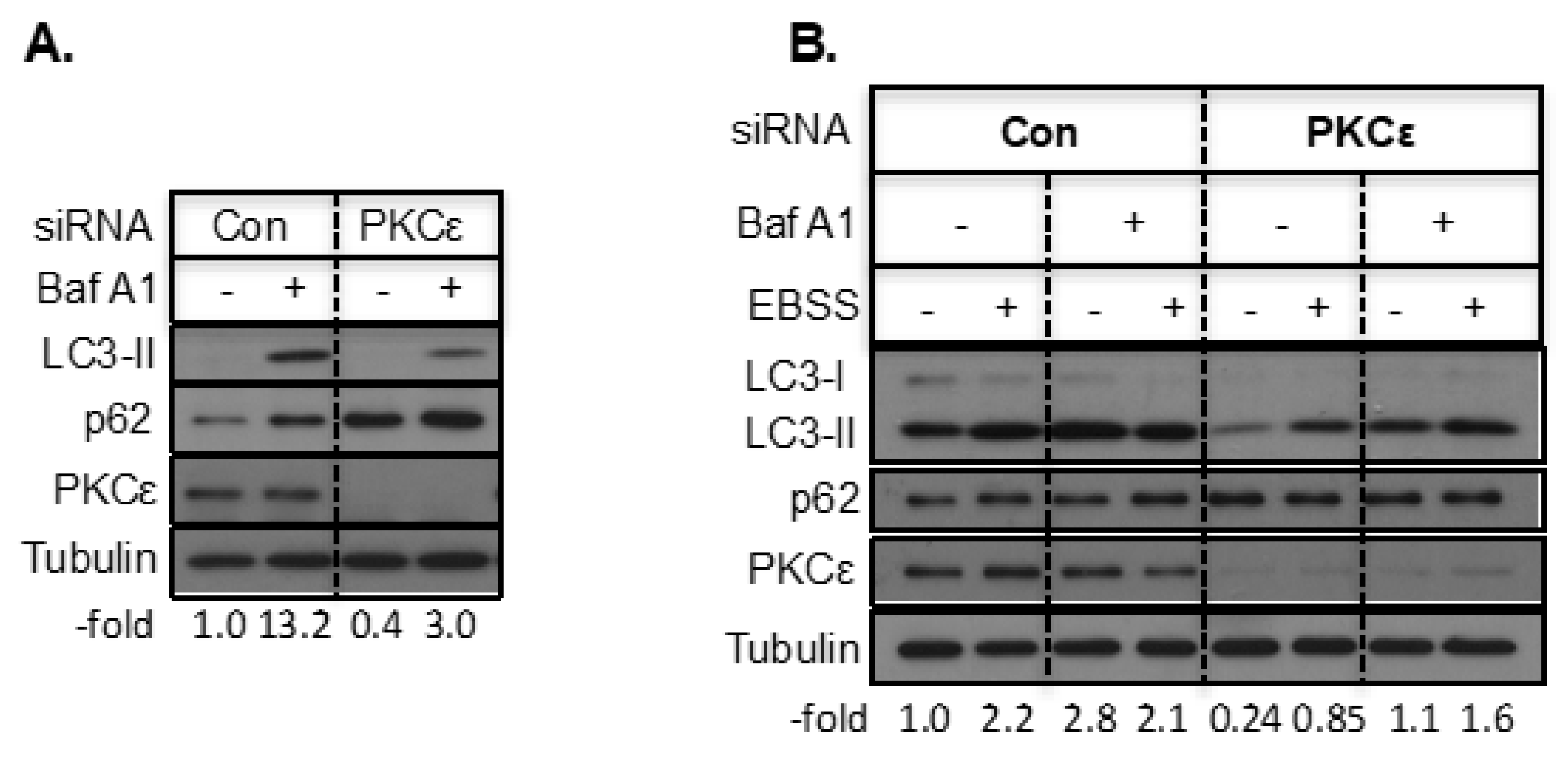
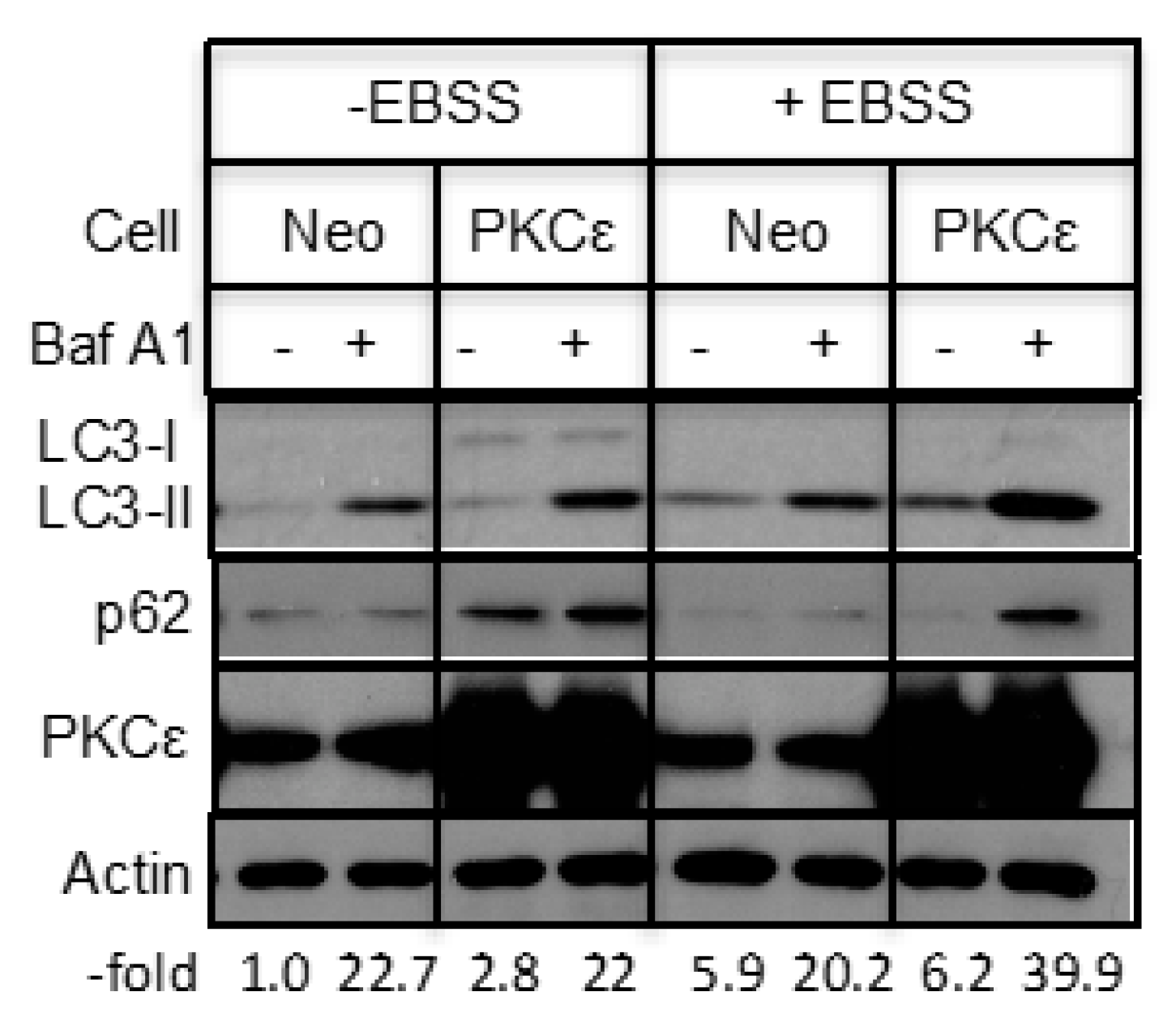
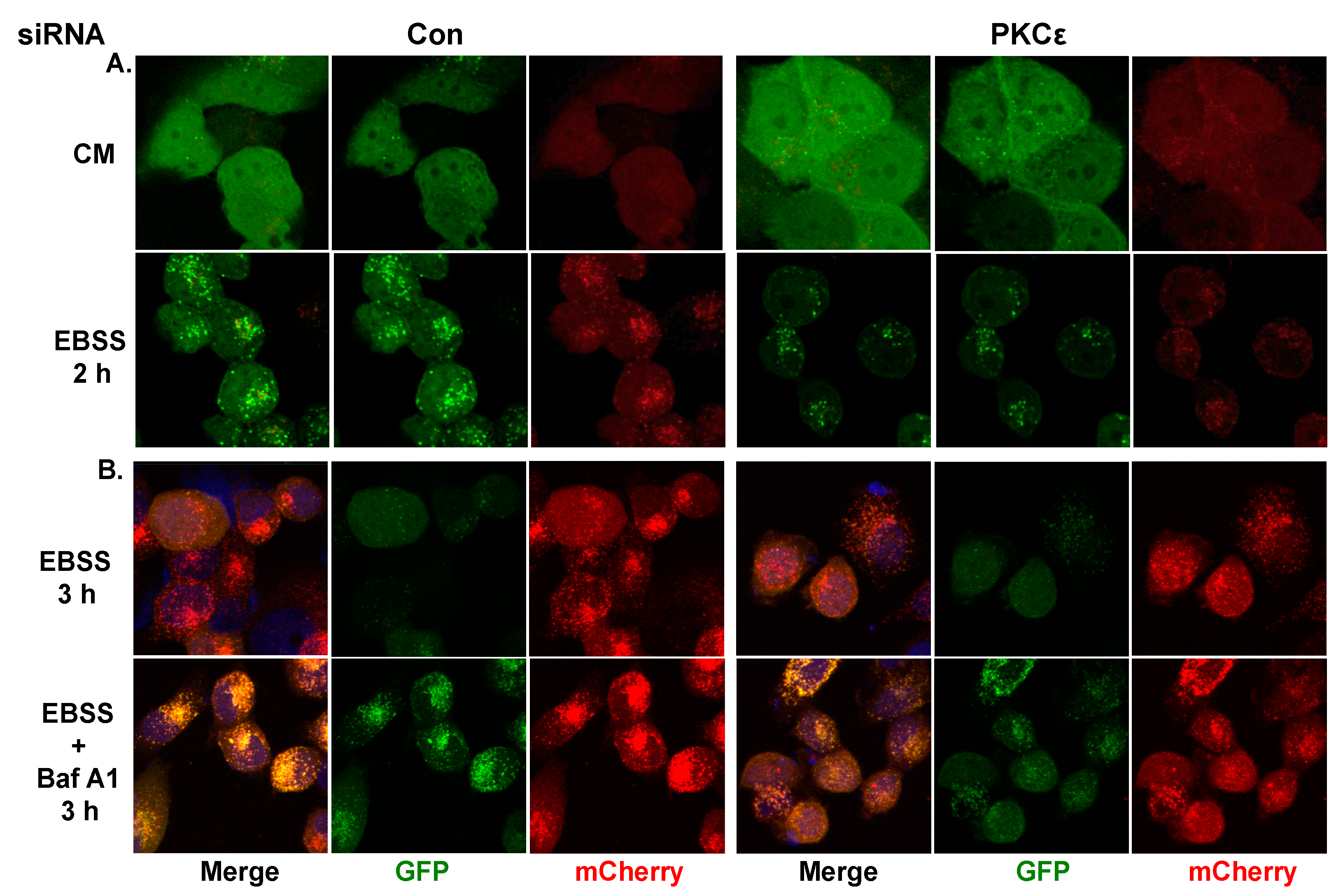
2.2. The Effect of PKCε on Starvation-Induced Autophagy
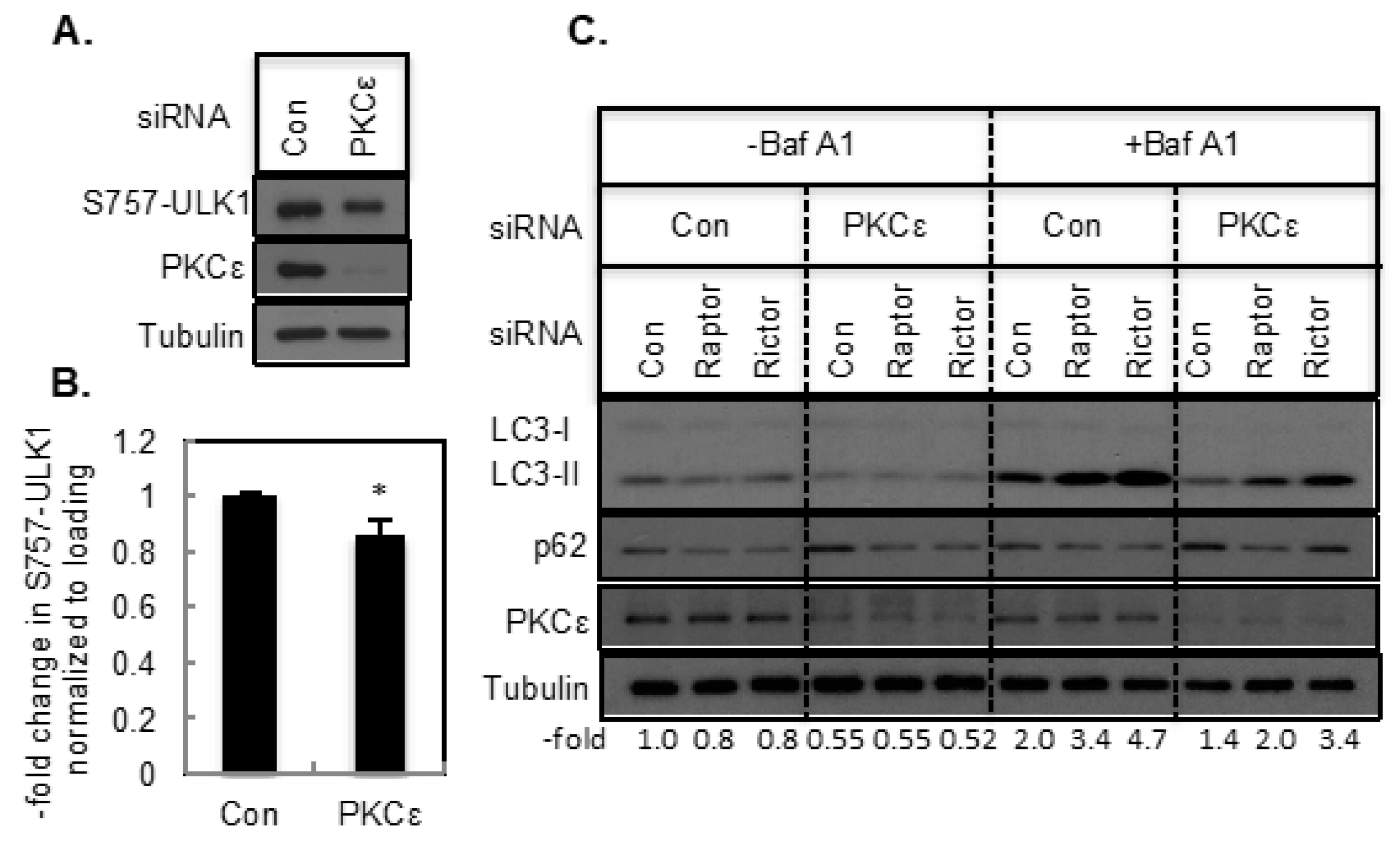
2.3. The Effect of PKCε on Autophagy Mediated by the mTOR Signaling
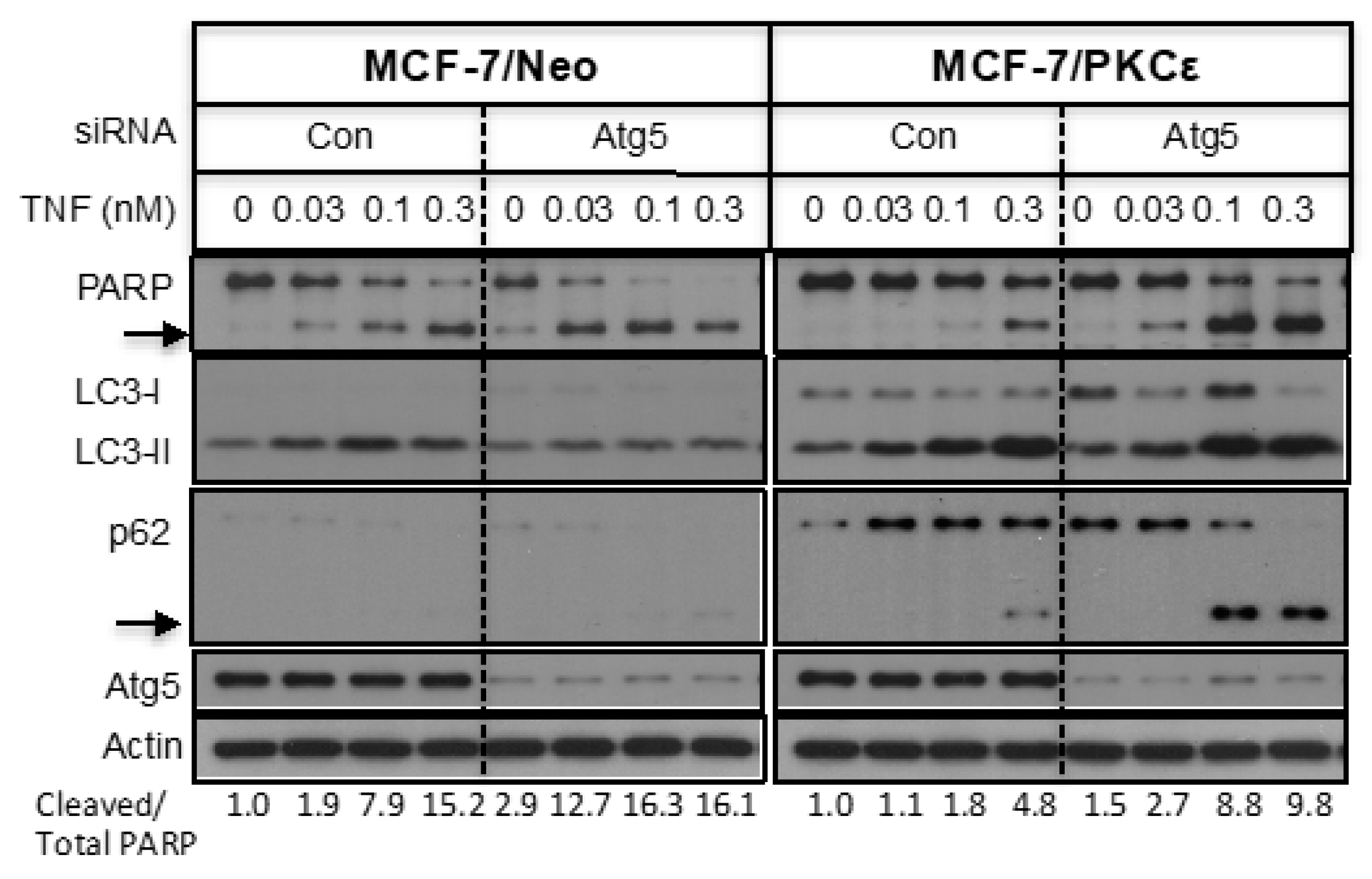
2.4. The Effect of PKCε-Mediated Autophagy on Apoptosis
3. Discussion
4. Materials and Methods
4.1. Reagents
4.2. Cell Culture and Transfection
4.3. Western Blot Analysis
4.4. Fluorescence Microscopy
4.5. Statistical Analyses
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| Atg | Autophagy-related |
| Baf A1 | Bafilomycin A1 |
| EBSS | Earle’s balanced salt solution |
| MAP1LC3/LC3 | Microtubule-associated protein 1 light chain 3 |
| mTOR | Mechanistic target of rapamycin |
| PARP | Poly (ADP-ribose) polymerase |
| PKC | Protein kinase C |
| Raptor | Regulatory-associated protein of mTOR |
| Rictor | Rapamycin-insensitive companion of mTOR |
| SDS-PAGE | Sodium dodecyl sulfate-polyacrylamide gel electrophoresis |
| TNF | Tumor necrosis factor-α |
| ULK1 | Unc-51-like kinase-1 |
References
- Nishizuka, Y. The role of protein kinase C in cell surface signal transduction and tumour promotion. Nature 1984, 308, 693–698. [Google Scholar] [CrossRef] [PubMed]
- Basu, A. The potential of protein kinase C as a target for anticancer treatment. Pharmacol. Ther. 1993, 59, 257–280. [Google Scholar] [CrossRef]
- Basu, A.; Sivaprasad, U. Protein kinase Cepsilon makes the life and death decision. Cell. Signal. 2007, 19, 1633–1642. [Google Scholar] [CrossRef]
- Gorin, M.A.; Pan, Q. Protein kinase C epsilon: An oncogene and emerging tumor biomarker. Mol. Cancer 2009, 8, 9. [Google Scholar] [CrossRef] [PubMed]
- Jain, K.; Basu, A. The Multifunctional Protein Kinase C-epsilon in Cancer Development and Progression. Cancers 2014, 6, 860–878. [Google Scholar] [CrossRef]
- Garg, R.; Benedetti, L.G.; Abera, M.B.; Wang, H.; Abba, M.; Kazanietz, M.G. Protein kinase C and cancer: What we know and what we do not. Oncogene 2014, 33, 5225–5237. [Google Scholar] [CrossRef] [PubMed]
- Lu, D.; Huang, J.; Basu, A. Protein kinase Cepsilon activates protein kinase B/Akt via DNA-PK to protect against tumor necrosis factor-alpha-induced cell death. J. Biol. Chem. 2006, 281, 22799–22807. [Google Scholar] [CrossRef]
- Lu, D.; Sivaprasad, U.; Huang, J.; Shankar, E.; Morrow, S.; Basu, A. Protein kinase C-epsilon protects MCF-7 cells from TNF-mediated cell death by inhibiting Bax translocation. Apoptosis 2007, 12, 1893–1900. [Google Scholar] [CrossRef]
- Shankar, E.; Sivaprasad, U.; Basu, A. Protein kinase C epsilon confers resistance of MCF-7 cells to TRAIL by Akt-dependent activation of Hdm2 and downregulation of p53. Oncogene 2008, 27, 3957–3966. [Google Scholar] [CrossRef]
- Steinberg, R.; Harari, O.A.; Lidington, E.A.; Boyle, J.J.; Nohadani, M.; Samarel, A.M.; Ohba, M.; Haskard, D.O.; Mason, J.C. A protein kinase Cepsilon-anti-apoptotic kinase signaling complex protects human vascular endothelial cells against apoptosis through induction of Bcl-2. J. Biol. Chem. 2007, 282, 32288–32297. [Google Scholar] [CrossRef]
- Kaleli, H.N.; Ozer, E.; Kaya, V.O.; Kutlu, O. Protein Kinase C Isozymes and Autophagy during Neurodegenerative Disease Progression. Cells 2020, 9, 553. [Google Scholar] [CrossRef] [PubMed]
- Toton, E.; Romaniuk, A.; Konieczna, N.; Hofmann, J.; Barciszewski, J.; Rybczynska, M. Impact of PKCepsilon downregulation on autophagy in glioblastoma cells. BMC Cancer 2018, 18, 185. [Google Scholar] [CrossRef] [PubMed]
- Yoshimori, T. Autophagy: A regulated bulk degradation process inside cells. Biochem. Biophys. Res. Commun. 2004, 313, 453–458. [Google Scholar] [CrossRef] [PubMed]
- Dossou, A.S.; Basu, A. The Emerging Roles of mTORC1 in Macromanaging Autophagy. Cancers 2019, 11, 1422. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.Y.; White, E. Role of autophagy in cancer prevention. Cancer Prev. Res. (Phila.) 2011, 4, 973–983. [Google Scholar] [CrossRef] [PubMed]
- Galluzzi, L.; Vitale, I.; Aaronson, S.A.; Abrams, J.M.; Adam, D.; Agostinis, P.; Alnemri, E.S.; Altucci, L.; Amelio, I.; Andrews, D.W.; et al. Molecular mechanisms of cell death: Recommendations of the Nomenclature Committee on Cell Death 2018. Cell Death Differ. 2018, 25, 486–541. [Google Scholar] [CrossRef]
- Saxton, R.A.; Sabatini, D.M. mTOR Signaling in Growth, Metabolism, and Disease. Cell 2017, 168, 960–976. [Google Scholar] [CrossRef]
- Rabanal-Ruiz, Y.; Otten, E.G.; Korolchuk, V.I. mTORC1 as the main gateway to autophagy. Essays Biochem. 2017, 61, 565–584. [Google Scholar] [CrossRef]
- Manning, B.D.; Toker, A. AKT/PKB Signaling: Navigating the Network. Cell 2017, 169, 381–405. [Google Scholar] [CrossRef]
- Sridharan, S.; Basu, A. Distinct Roles of mTOR Targets S6K1 and S6K2 in Breast Cancer. Int. J. Mol. Sci. 2020, 21, 1199. [Google Scholar] [CrossRef]
- Basu, A.; Lu, D.; Sun, B.; Moor, A.N.; Akkaraju, G.R.; Huang, J. Proteolytic activation of protein kinase C-epsilon by caspase-mediated processing and transduction of antiapoptotic signals. J. Biol. Chem. 2002, 277, 41850–41856. [Google Scholar] [CrossRef] [PubMed]
- Jain, K.; Paranandi, K.S.; Sridharan, S.; Basu, A. Autophagy in breast cancer and its implications for therapy. Am. J. Cancer Res. 2013, 3, 251–265. [Google Scholar]
- Klionsky, D.J.; Abdelmohsen, K.; Abe, A.; Abedin, M.J.; Abeliovich, H.; Acevedo Arozena, A.; Adachi, H.; Adams, C.M.; Adams, P.D.; Adeli, K.; et al. Guidelines for the use and interpretation of assays for monitoring autophagy (3rd edition). Autophagy 2016, 12, 1–222. [Google Scholar] [CrossRef] [PubMed]
- Yoshii, S.R.; Mizushima, N. Monitoring and Measuring Autophagy. Int. J. Mol. Sci. 2017, 18, 1865. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Kundu, M.; Viollet, B.; Guan, K.L. AMPK and mTOR regulate autophagy through direct phosphorylation of Ulk1. Nat. Cell Biol. 2011, 13, 132–141. [Google Scholar] [CrossRef] [PubMed]
- Shang, L.; Chen, S.; Du, F.; Li, S.; Zhao, L.; Wang, X. Nutrient starvation elicits an acute autophagic response mediated by Ulk1 dephosphorylation and its subsequent dissociation from AMPK. Proc. Natl. Acad. Sci. USA 2011, 108, 4788–4793. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Garrido, J.; Sancho-Shimizu, V.; Shenoy, A.R. Regulated proteolysis of p62/SQSTM1 enables differential control of autophagy and nutrient sensing. Sci. Signal. 2018, 11, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Norman, J.M.; Cohen, G.M.; Bampton, E.T. The in vitro cleavage of the hAtg proteins by cell death proteases. Autophagy 2010, 6, 1042–1056. [Google Scholar] [CrossRef]
- Jain, K.; Basu, A. Protein Kinase C-epsilon Promotes EMT in Breast Cancer. Breast Cancer (Auckl.) 2014, 8, 61–67. [Google Scholar] [CrossRef]
- Tian, T.; Li, X.; Zhang, J. mTOR Signaling in Cancer and mTOR Inhibitors in Solid Tumor Targeting Therapy. Int. J. Mol. Sci. 2019, 20, 755. [Google Scholar] [CrossRef]
- Li, L.; Friedrichsen, H.J.; Andrews, S.; Picaud, S.; Volpon, L.; Ngeow, K.; Berridge, G.; Fischer, R.; Borden, K.L.B.; Filippakopoulos, P.; et al. A TFEB nuclear export signal integrates amino acid supply and glucose availability. Nat. Commun. 2018, 9, 2685. [Google Scholar] [CrossRef] [PubMed]
- Ozturk, D.G.; Kocak, M.; Akcay, A.; Kinoglu, K.; Kara, E.; Buyuk, Y.; Kazan, H.; Gozuacik, D.A.-O.h.o.o. MITF-MIR211 axis is a novel autophagy amplifier system during cellular stress. Autophagy 2019, 15, 375–390. [Google Scholar] [CrossRef] [PubMed]
- Ikenoue, T.; Inoki, K.; Yang, Q.; Zhou, X.; Guan, K.L. Essential function of TORC2 in PKC and Akt turn motif phosphorylation, maturation and signalling. EMBO J. 2008, 27, 1919–1931. [Google Scholar] [CrossRef]
- Sarbassov, D.D.; Guertin, D.A.; Ali, S.M.; Sabatini, D.M. Phosphorylation and regulation of Akt/PKB by the rictor-mTOR complex. Science 2005, 307, 1098–1101. [Google Scholar] [CrossRef]
- Liu, P.; Gan, W.; Inuzuka, H.; Lazorchak, A.S.; Gao, D.; Arojo, O.; Liu, D.; Wan, L.; Zhai, B.; Yu, Y.; et al. Sin1 phosphorylation impairs mTORC2 complex integrity and inhibits downstream Akt signalling to suppress tumorigenesis. Nat. Cell Biol. 2013, 15, 1340–1350. [Google Scholar] [CrossRef] [PubMed]
- Sridharan, S.; Jain, K.; Basu, A. Regulation of autophagy by kinases. Cancers 2011, 3, 2630–2654. [Google Scholar] [CrossRef] [PubMed]
- Alers, S.; Loffler, A.S.; Wesselborg, S.; Stork, B. Role of AMPK-mTOR-Ulk1/2 in the regulation of autophagy: Cross talk, shortcuts, and feedbacks. Mol. Cell. Biol. 2012, 32, 2–11. [Google Scholar] [CrossRef] [PubMed]
- Kazyken, D.; Magnuson, B.; Bodur, C.; Acosta-Jaquez, H.A.; Zhang, D.; Tong, X.; Barnes, T.M.; Steinl, G.K.; Patterson, N.E.; Altheim, C.H.; et al. AMPK directly activates mTORC2 to promote cell survival during acute energetic stress. Sci. Signal. 2019, 12, 1–16. [Google Scholar] [CrossRef] [PubMed]
© 2020 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Basu, A. Regulation of Autophagy by Protein Kinase C-ε in Breast Cancer Cells. Int. J. Mol. Sci. 2020, 21, 4247. https://doi.org/10.3390/ijms21124247
Basu A. Regulation of Autophagy by Protein Kinase C-ε in Breast Cancer Cells. International Journal of Molecular Sciences. 2020; 21(12):4247. https://doi.org/10.3390/ijms21124247
Chicago/Turabian StyleBasu, Alakananda. 2020. "Regulation of Autophagy by Protein Kinase C-ε in Breast Cancer Cells" International Journal of Molecular Sciences 21, no. 12: 4247. https://doi.org/10.3390/ijms21124247
APA StyleBasu, A. (2020). Regulation of Autophagy by Protein Kinase C-ε in Breast Cancer Cells. International Journal of Molecular Sciences, 21(12), 4247. https://doi.org/10.3390/ijms21124247




