Sperm Differentiation: The Role of Trafficking of Proteins
Abstract
1. Introduction
2. Evolution of Mammalian Sperm
3. General Mechanisms During Sperm Differentiation
3.1. Acrosome Development
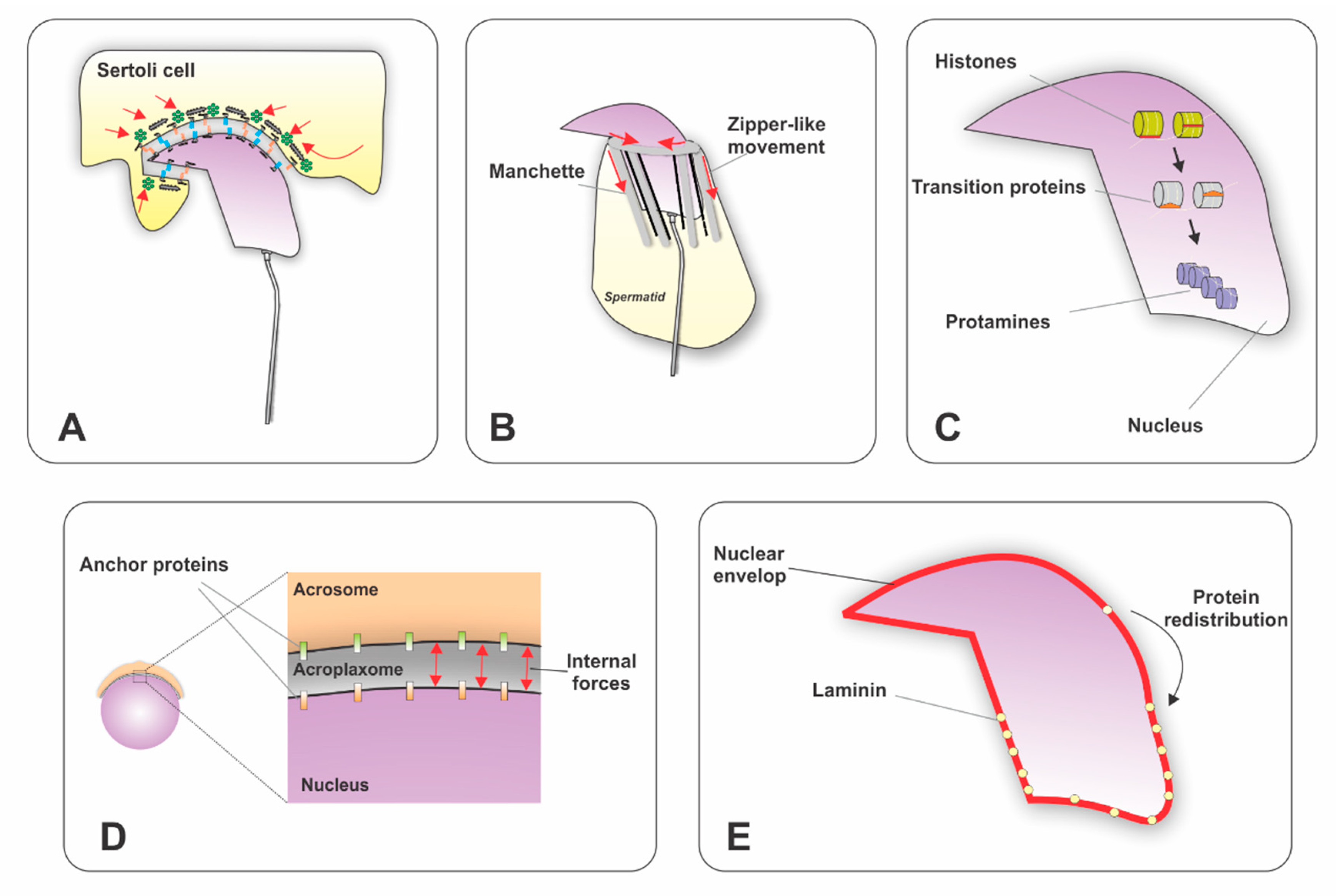
3.2. Nuclear Elongation
3.3. Flagellar Formation
4. Trafficking of Proteins During Sperm Differentiation
4.1. Components of The Delivery System
4.1.1. Tubulin
4.1.2. Actin
4.1.3. Motor proteins
4.1.4. Small GTPases (The RAB family)
4.1.5. Vesicle Coats
4.1.6. Intraflagellar Transport (IFT) Proteins
4.2. Trafficking
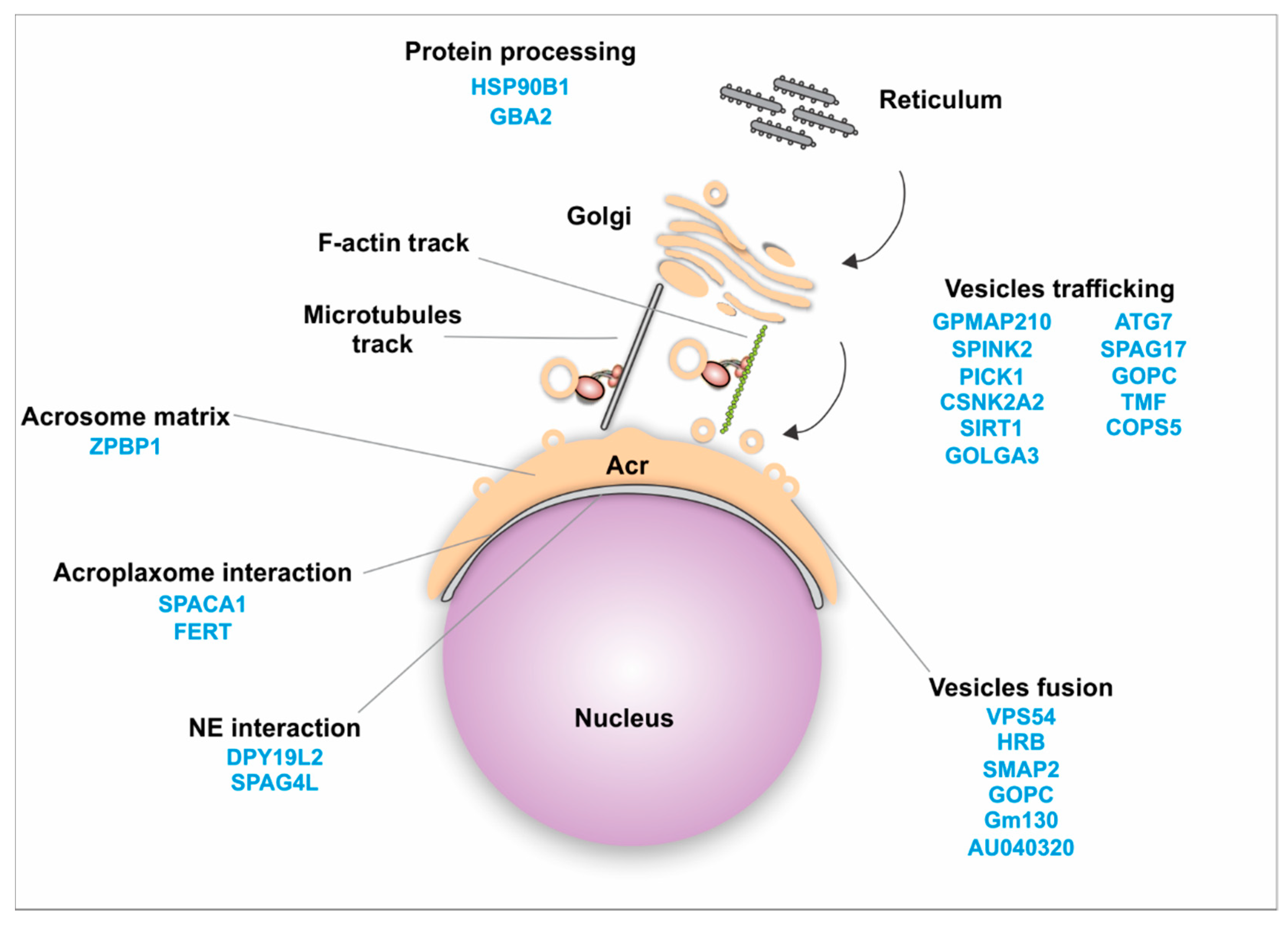
4.2.1. Golgi Transport
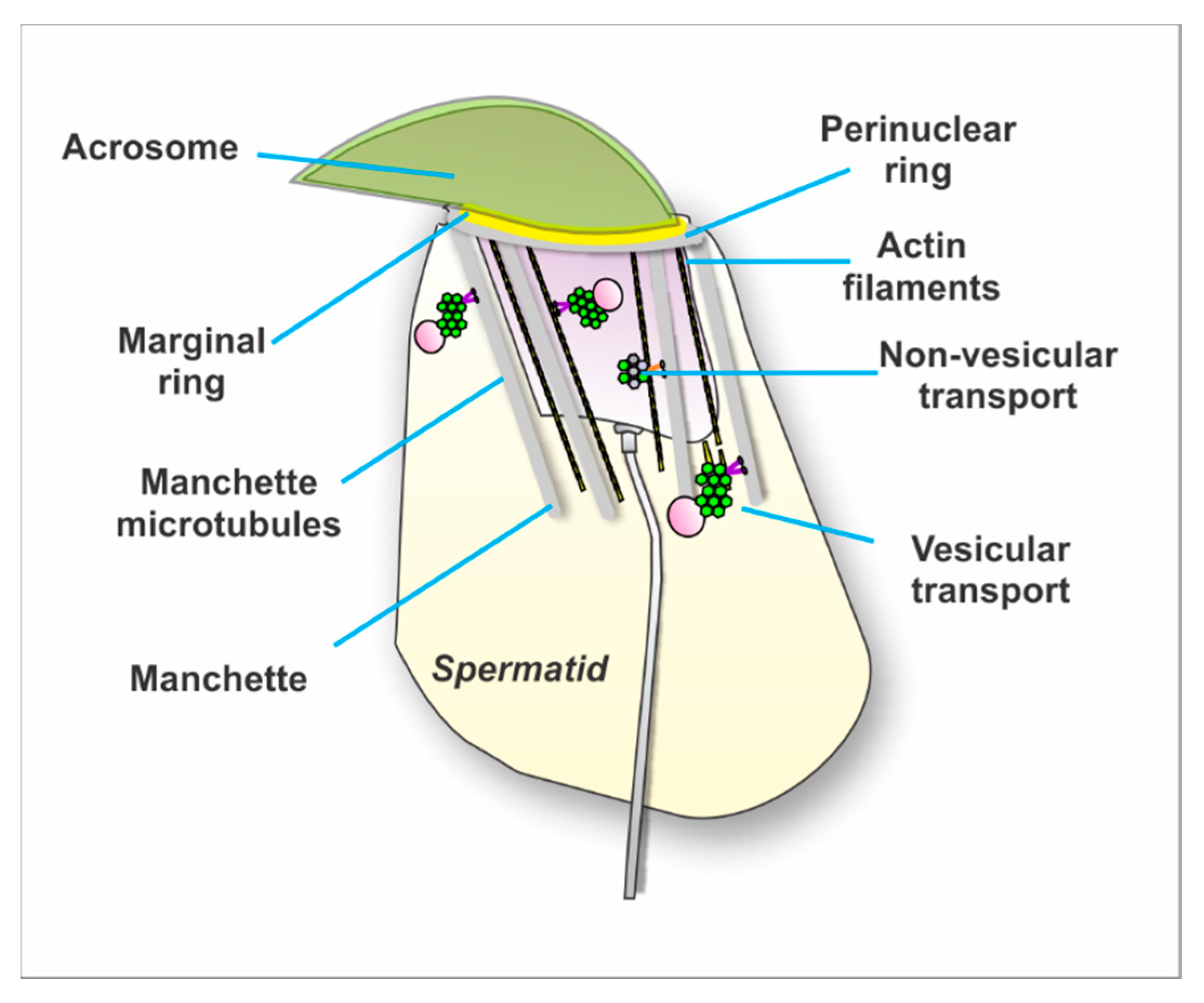
4.2.2. Intramanchette Transport (IMT)
4.3. Protein Interactomes
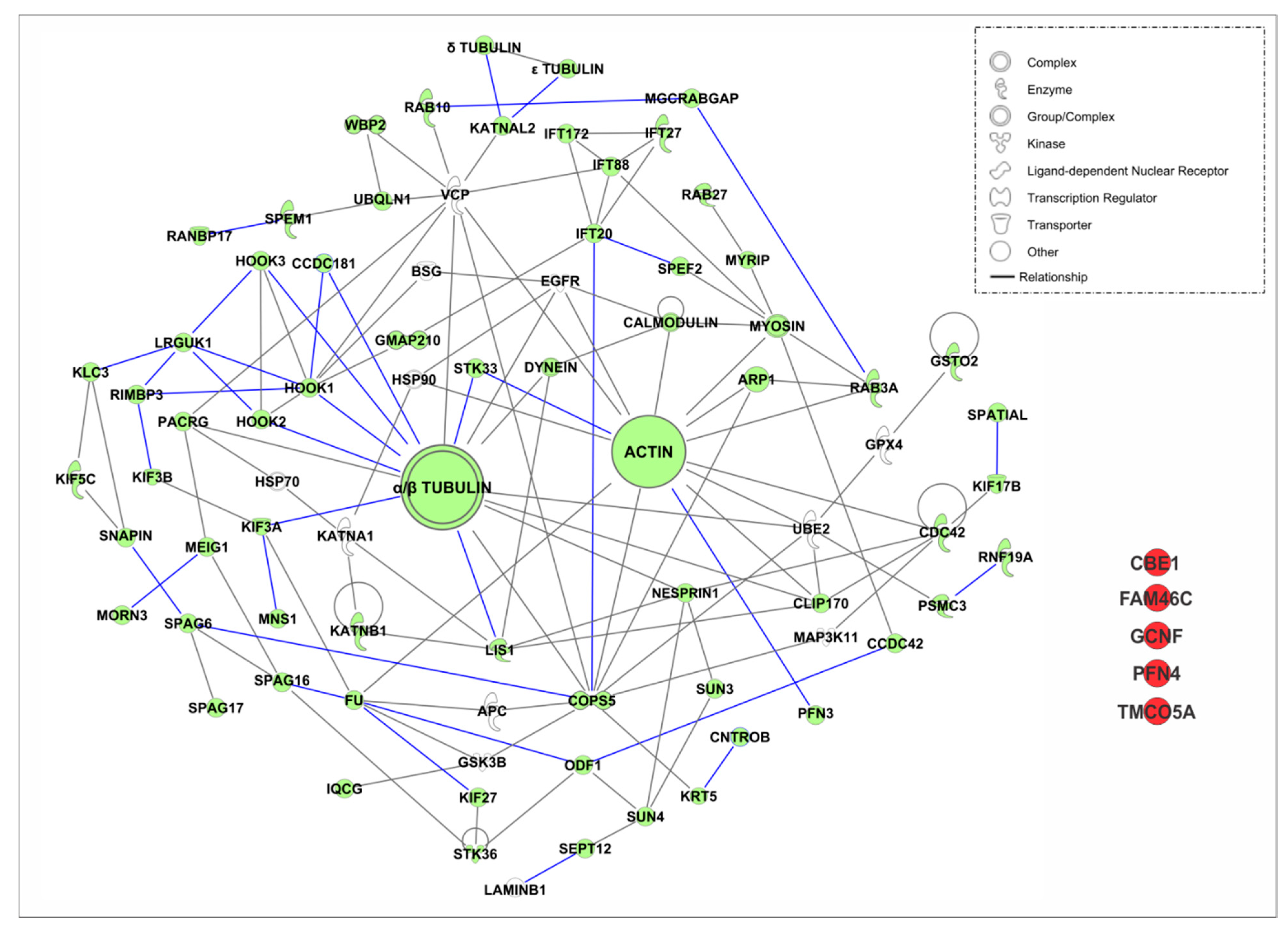
4.3.1. Intramanchette Transport Interactome
4.3.2. Golgi Transport Interactome
4.4. Novel Technology to Study Trafficking of Proteins (Super-Resolution Microscopy)
5. Conclusions and Future Directions
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Russell, L.D.; Ettlin, R.A.; Sinha Hikim, A.P.; Clegg, E.D. Histological and Histopathological Evaluation of the Testis. Cache River Press: St. Louis, MO, USA, 1990. [Google Scholar]
- Auger, J. Spermatozoa and Sperm Structure. In Encyclopedia of Reproduction, 2nd ed.; Skinner, M., Ed.; Academic Press: Amsterdam, The Netherlands, 2018; pp. 62–67. [Google Scholar]
- Cohen, J. Reproduction; Butterworth & Co.: London, UK, 1977. [Google Scholar]
- Birkhead, T.R.; Hosken, D.J.; Pitnick, S.B.T. Biology of Spermatozoa Sperm Biology; Academic Press: Amsterdam, The Netherlands, 2009. [Google Scholar]
- Temple-Smith, P.; Ravichandran, A.; Horta, F. Sperm: Comparative Vertebrate. In Encyclopedia of Reproduction, 2nd ed.; Skinner, M., Ed.; Academic Press: Amsterdam, The Netherlands, 2018; pp. 210–220. [Google Scholar]
- Brinster, R.L.; Nagano, M. Spermatogonial stem cell transplantation, cryopreservation and culture. Semin. Cell Dev. Biol. 1998, 9, 401–409. [Google Scholar] [CrossRef] [PubMed]
- Johnson, L. Efficiency of spermatogenesis. Microsc. Res. Tech. 1995, 32, 385–422. [Google Scholar] [CrossRef] [PubMed]
- DelBarco-Trillo, J.; Tourmente, M.; Roldan, E.R.S. Metabolic Rate Limits the Effect of Sperm Competition on Mammalian Spermatogenesis. PLoS ONE 2013, 8, e76510. [Google Scholar] [CrossRef] [PubMed]
- Giannakara, A.; Schärer, L.; Ramm, S.A. Sperm competition-induced plasticity in the speed of spermatogenesis. BMC Evol. Biol. 2016, 16, 60. [Google Scholar] [CrossRef]
- Firman, R.C.; Garcia-Gonzalez, F.; Simmons, L.W.; André, G.I. A competitive environment influences sperm production, but not testes tissue composition, in house mice. J. Evol. Biol. 2018, 31, 1647–1654. [Google Scholar] [CrossRef]
- Roldan, E.R.S. Sperm competition and the evolution of sperm form and function in mammals. Reprod. Domest. Anim. 2019, 54, 14–21. [Google Scholar] [CrossRef]
- Bedford, J.M. Puzzles of mammalian fertilization—And beyond. Int. J. Dev. Biol. 2008, 52, 415–426. [Google Scholar] [CrossRef]
- Bedford, J.M. Singular features of fertilization and their impact on the male reproductive system in eutherian mammals. Reproduction 2014, 147, R43–R52. [Google Scholar] [CrossRef]
- Malo, A.F.; Gomendio, M.; Garde, J.; Lang-Lenton, B.; Soler, A.J.; Roldan, E.R. Sperm design and sperm function. Biol. Lett. 2006, 2, 246–249. [Google Scholar] [CrossRef]
- Tourmente, M.; Gomendio, M.; Roldan, E.R.S. Sperm competition and the evolution of sperm design in mammals. BMC Evol. Biol. 2011, 11, 12. [Google Scholar] [CrossRef]
- Nielsen, R.; Bustamante, C.; Clark, A.G.; Glanowski, S.; Sackton, T.B.; Hubisz, M.J.; Fledel-Alon, A.; Tanenbaum, D.M.; Civello, D.; White, T.J.; et al. A Scan for Positively Selected Genes in the Genomes of Humans and Chimpanzees. PLOS Biol. 2005, 3, e170. [Google Scholar] [CrossRef] [PubMed]
- Torgerson, D.G.; Kulathinal, R.J.; Singh, R.S. Mammalian Sperm Proteins Are Rapidly Evolving: Evidence of Positive Selection in Functionally Diverse Genes. Mol. Biol. Evol. 2002, 19, 1973–1980. [Google Scholar] [CrossRef] [PubMed]
- Vicens, A.; Lüke, L.; Roldan, E.R.S. Proteins involved in motility and sperm-egg interaction evolve more rapidly in mouse spermatozoa. PLoS ONE 2014, 9, e91302. [Google Scholar] [CrossRef] [PubMed]
- Green, R.E.; Krause, J.; Briggs, A.W.; Maricic, T.; Stenzel, U.; Kircher, M.; Patterson, N.; Li, H.; Zhai, W.; Fritz, M.H.-Y.; et al. A Draft Sequence of the Neandertal Genome. Science 2010, 328, 710–722. [Google Scholar] [CrossRef]
- Zhang, Z.; Jones, B.; Tang, W.; Moss, S.B.; Wei, Z.; Ho, C.K.M.; Pollack, M.; Horowitz, E.; Bennett, J.; Baker, M.E.; et al. Dissecting the axoneme interactome: The mammalian orthologue of Chlamydomonas PF6 interacts with sperm-associated antigen 6, the mammalian orthologue of Chlamydomonas PF16. Mol. Cell. Proteom. 2005, 4, 914–923. [Google Scholar] [CrossRef]
- Kazarian, E.; Son, H.; Sapao, P.; Li, W.; Zhang, Z.; Strauss, J.F.; Teves, M.E. SPAG17 Is Required for Male Germ Cell Differentiation and Fertility. Int. J. Mol. Sci. 2018, 19, 1252. [Google Scholar] [CrossRef]
- Sapiro, R.; Tarantino, L.M.; Velazquez, F.; Kiriakidou, M.; Hecht, N.B.; Bucan, M.; Strauss, J.F. Sperm antigen 6 is the murine homologue of the Chlamydomonas reinhardtii central apparatus protein encoded by the PF16 locus. Boil. Reprod. 2000, 62, 511–518. [Google Scholar] [CrossRef]
- Williamson, S.H.; Hubisz, M.J.; Clark, A.G.; Payseur, B.A.; Bustamante, C.D.; Nielsen, R. Localizing recent adaptive evolution in the human genome. PLoS Genet. 2007, 3, e90. [Google Scholar] [CrossRef]
- Hu, M.; Ayub, Q.; Guerra-Assuncao, J.A.; Long, Q.; Ning, Z.; Huang, N.; Romero, I.G.; Mamanova, L.; Akan, P.; Liu, X.; et al. Exploration of signals of positive selection derived from genotype-based human genome scans using re-sequencing data. Qual. Life Res. 2011, 131, 665–674. [Google Scholar] [CrossRef]
- Qiu, H.; Gołas, A.; Grzmil, P.; Wojnowski, L. Lineage-Specific Duplications of Muroidea Faim and Spag6 Genes and Atypical Accelerated Evolution of the Parental Spag6 Gene. J. Mol. Evol. 2013, 77, 119–129. [Google Scholar] [CrossRef]
- Liu, Y.; Zhang, L.; Li, W.; Huang, Q.; Yuan, S.; Li, Y.; Liu, J.; Zhang, S.; Pin, G.; Song, S.; et al. The sperm associated antigen 6 interactome and its role in spermatogenesis. Reproduction 2019, 158, 183–199. [Google Scholar] [CrossRef] [PubMed]
- Sigg, M.A.; Menchen, T.; Lee, C.; Johnson, J.; Jungnickel, M.K.; Choksi, S.; Garcia, G.; Busengdal, H.; Dougherty, G.W.; Pennekamp, P.; et al. Evolutionary Proteomics Uncovers Ancient Associations of Cilia with Signaling Pathways. Dev. Cell 2017, 43, 744–762.e11. [Google Scholar] [CrossRef] [PubMed]
- Kierszenbaum, A.L.; Rivkin, E.; Tres, L.L.; Yoder, B.K.; Haycraft, C.J.; Bornens, M.; Ríos, R.M. GMAP210 and IFT88 are present in the spermatid golgi apparatus and participate in the development of the acrosome-acroplaxome complex, head-tail coupling apparatus and tail. Dev. Dyn. 2011, 240, 723–736. [Google Scholar] [CrossRef] [PubMed]
- San Agustin, J.T.; Pazour, G.J.; Witman, G.B. Intraflagellar transport is essential for mammalian spermiogenesis but is absent in mature sperm. Mol. Biol. Cell 2015, 26, 4358–4372. [Google Scholar] [CrossRef]
- Zhang, Z.; Li, W.; Zhang, Y.; Zhang, L.; Teves, M.E.; Liu, H.; Strauss, J.F.; Pazour, G.J.; Foster, J.A.; Hess, R.; et al. Intraflagellar transport protein IFT20 is essential for male fertility and spermiogenesis in mice. Mol. Boil. Cell 2016, 27, 3705–3716. [Google Scholar] [CrossRef]
- Liu, H.; Li, W.; Zhang, Y.; Zhang, Z.; Shang, X.; Zhang, L.; Zhang, S.; Li, Y.; Somoza, A.V.; Delpi, B.; et al. IFT25, an intraflagellar transporter protein dispensable for ciliogenesis in somatic cells, is essential for sperm flagella formation. Boil. Reprod. 2017, 96, 993–1006. [Google Scholar] [CrossRef]
- Zhang, Y.; Liu, H.; Li, W.; Zhang, Z.; Shang, X.; Zhang, D.; Li, Y.; Zhang, S.; Liu, J.; Hess, R.; et al. Intraflagellar transporter protein (IFT27), an IFT25 binding partner, is essential for male fertility and spermiogenesis in mice. Dev. Boil. 2017, 432, 125–139. [Google Scholar] [CrossRef]
- Shi, L.; Zhou, T.; Huang, Q.; Zhang, S.; Li, W.; Zhang, L.; Hess, R.; Pazour, G.J.; Zhang, Z. Intraflagellar transport protein 74 is essential for spermatogenesis and male fertility in mice. Boil. Reprod. 2019, 101, 188–199. [Google Scholar] [CrossRef]
- Zhang, Y.; Liu, H.; Li, W.; Zhang, Z.; Zhang, S.; E Teves, M.; Stevens, C.; A Foster, J.; E Campbell, G.; Windle, J.J.; et al. Intraflagellar transporter protein 140 (IFT140), a component of IFT-A complex, is essential for male fertility and spermiogenesis in mice. Cytoskeleton 2018, 75, 70–84. [Google Scholar] [CrossRef]
- Zhang, S.; Liu, Y.; Huang, Q.; Yuan, S.; Liu, H.; Shi, L.; Yap, Y.T.; Li, W.; Zhen, J.; Zhang, L.; et al. Murine germ cell-specific disruption of Ift172 causes defects in spermiogenesis and male fertility. Reproduction 2020, 159, 409–421. [Google Scholar] [CrossRef]
- Kierszenbaum, A.L.; Rivkin, E.; Tres, L.L. Acroplaxome, an F-actin-keratin-containing plate, anchors the acrosome to the nucleus during shaping of the spermatid head. Mol. Biol. Cell 2003, 14, 4628–4640. [Google Scholar] [CrossRef] [PubMed]
- Khawar, M.B.; Gao, H.; Li, W. Mechanism of Acrosome Biogenesis in Mammals. Front. Cell Dev. Biol. 2019, 7, 195. [Google Scholar] [CrossRef] [PubMed]
- Han, F.; Liu, C.; Zhang, L.; Zhou, Y.; Qin, Y.; Wang, Y.; Chen, M.; Duo, S.; Cui, X.; Bao, S.; et al. Globozoospermia and lack of acrosome formation in GM130-deficient mice. Cell Death Dis. 2017, 8, e2532. [Google Scholar] [CrossRef] [PubMed]
- Guidi, L.G.; Holloway, Z.G.; Arnoult, C.; Ray, P.; Monaco, A.P.; Molnár, Z.; Velayos-Baeza, A. AU040320 deficiency leads to disruption of acrosome biogenesis and infertility in homozygous mutant mice. Sci. Rep. 2018, 8, 10379. [Google Scholar] [CrossRef] [PubMed]
- Tardif, S.; Guyonnet, B.; Cormier, N.; Cornwall, G.A. Alteration in the processing of the ACRBP/sp32 protein and sperm head/acrosome malformations in proprotein convertase 4 (PCSK4) null mice. Mol. Hum. Reprod. 2012, 18, 298–307. [Google Scholar] [CrossRef]
- Wang, H.; Wan, H.; Li, X.; Liu, W.; Chen, Q.; Wang, Y.; Yang, L.; Tang, H.; Zhang, X.; Duan, E.; et al. Atg7 is required for acrosome biogenesis during spermatogenesis in mice. Cell Res. 2014, 24, 852–869. [Google Scholar] [CrossRef]
- Pereira, C.D.; Serrano, J.B.; Martins, F.; da Cruz e Silva, O.A.B.; Rebelo, S. Nuclear envelope dynamics during mammalian spermatogenesis: New insights on male fertility. Biol. Rev. 2019, 94, 1195–1219. [Google Scholar] [CrossRef]
- Lie, P.P.Y.; Mruk, D.D.; Lee, W.M.; Cheng, C.Y. Cytoskeletal dynamics and spermatogenesis. Philos. Trans. R. Soc. Lond. B. Biol. Sci. 2010, 365, 1581–1592. [Google Scholar] [CrossRef]
- Kierszenbaum, A.L.; Rivkin, E.; Tres, L.L. The actin-based motor myosin Va is a component of the acroplaxome, an acrosome-nuclear envelope junctional plate, and of manchette-associated vesicles. Cytogenet. Genome Res. 2003, 103, 337–344. [Google Scholar] [CrossRef]
- Kierszenbaum, A.L.; Rivkin, E.; Tres, L.L. Molecular biology of sperm head shaping. Soc. Reprod. Fertil. Suppl. 2007, 65, 33–43. [Google Scholar]
- Kierszenbaum, A.L.; Rivkin, E.; Talmor-Cohen, A.; Shalgi, R.; Tres, L.L. Expression of full-length and truncated Fyn tyrosine kinase transcripts and encoded proteins during spermatogenesis and localization during acrosome biogenesis and fertilization. Mol. Reprod. Dev. 2009, 76, 832–843. [Google Scholar] [CrossRef] [PubMed]
- Kierszenbaum, A.L.; Rivkin, E.; Tres, L.L. Expression of Fer testis (FerT) tyrosine kinase transcript variants and distribution sites of FerT during the development of the acrosome-acroplaxome-manchette complex in rat spermatids. Dev. Dyn. 2008, 237, 3882–3891. [Google Scholar] [CrossRef] [PubMed]
- Behnen, M.; Murk, K.; Kursula, P.; Cappallo-Obermann, H.; Rothkegel, M.; Kierszenbaum, A.L.; Kirchhoff, C. Testis-expressed profilins 3 and 4 show distinct functional characteristics and localize in the acroplaxome-manchette complex in spermatids. BMC Cell Boil. 2009, 10, 34. [Google Scholar] [CrossRef] [PubMed]
- Schütz, W.; Alsheimer, M.; Öllinger, R.; Benavente, R. Nuclear envelope remodeling during mouse spermiogenesis: Postmeiotic expression and redistribution of germline lamin B3. Exp. Cell Res. 2005, 307, 285–291. [Google Scholar] [CrossRef] [PubMed]
- Göb, E.; Schmitt, J.; Benavente, R.; Alsheimer, M. Mammalian sperm head formation involves different polarization of two novel LINC complexes. PLoS ONE 2010, 5, e12072. [Google Scholar] [CrossRef] [PubMed]
- Kracklauer, M.P.; Link, J.; Alsheimer, M. LINCing the Nuclear Envelope to Gametogenesis. In Current Topics in Developmental Biology; Academic Press: Cambridge, MA, USA, 2013; Volume 102, pp. 127–157. [Google Scholar]
- Kierszenbaum, A.L.; Rivkin, E.; Tres, L.L. Cytoskeletal track selection during cargo transport in spermatids is relevant to male fertility. Spermatogenesis 2011, 1, 221–230. [Google Scholar] [CrossRef] [PubMed]
- Kierszenbaum, A.L.; Tres, L.L. The acrosome-acroplaxome-manchette complex and the shaping of the spermatid head. Arch. Histol. Cytol. 2004, 67, 271–284. [Google Scholar] [CrossRef]
- Lehti, M.S.; Sironen, A. Formation and function of the manchette and flagellum during spermatogenesis. Reproduction 2016, 151, R43–R54. [Google Scholar] [CrossRef]
- Hao, S.-L.; Ni, F.-D.; Yang, W.-X. The dynamics and regulation of chromatin remodeling during spermiogenesis. Gene 2019, 706, 201–210. [Google Scholar] [CrossRef]
- Inaba, K. Molecular Architecture of the Sperm Flagella: Molecules for Motility and Signaling. Zoolog. Sci. 2003, 20, 1043–1056. [Google Scholar] [CrossRef]
- Sun, X.; Kovacs, T.; Hu, Y.-J.; Yang, W.-X. The role of actin and myosin during spermatogenesis. Mol. Biol. Rep. 2011, 38, 3993–4001. [Google Scholar] [CrossRef] [PubMed]
- Terman, J.R.; Kashina, A. Post-translational modification and regulation of actin. Curr. Opin. Cell Biol. 2013, 25, 30–38. [Google Scholar] [CrossRef] [PubMed]
- Yagi, A.; Paranko, J. Localization of actin, α-actinin, and tropomyosin in bovine spermatozoa and epididymal epithelium. Anat. Rec. 1992, 233, 61–74. [Google Scholar] [CrossRef] [PubMed]
- Maier, B.; Medrano, S.; Sleight, S.B.; Visconti, P.E.; Scrable, H. Developmental Association of the Synaptic Activity-Regulated Protein Arc with the Mouse Acrosomal Organelle and the Sperm Tail. Biol. Reprod. 2003, 68, 67–76. [Google Scholar] [CrossRef]
- Hara, Y.; Yamagata, K.; Oguchi, K.; Baba, T. Nuclear localization of profilin III-ArpM1 complex in mouse spermiogenesis. FEBS Lett. 2008, 582, 2998–3004. [Google Scholar] [CrossRef]
- Young, J.S.; Guttman, J.A.; Vaid, K.S.; Shahinian, H.; Vogl, A.W. Cortactin (CTTN), N-WASP (WASL), and Clathrin (CLTC) Are Present at Podosome-Like Tubulobulbar Complexes in the Rat Testis. Biol. Reprod. 2009, 80, 153–161. [Google Scholar] [CrossRef][Green Version]
- Paranko, J.; Longo, F.; Potts, J.; Krohne, G.; Franke, W.W. Widespread occurrence of calicin, a basic cytoskeletal protein of sperm cells, in diverse mammalian species. Differentiation 1988, 38, 21–27. [Google Scholar] [CrossRef]
- Sosnik, J.; Buffone, M.G.; Visconti, P.E. Analysis of CAPZA3 localization reveals temporally discrete events during the acrosome reaction. J. Cell. Physiol. 2010, 224, 575–580. [Google Scholar] [CrossRef]
- Geyer, C.B.; Inselman, A.L.; Sunman, J.A.; Bornstein, S.; Handel, M.A.; Eddy, E.M. A missense mutation in the Capza3 gene and disruption of F-actin organization in spermatids of repro32 infertile male mice. Dev. Boil. 2009, 330, 142–152. [Google Scholar] [CrossRef]
- Chapin, R.E.; Wine, R.N.; Harris, M.W.; Borchers, C.H.; Haseman, J.K. Structure and control of a cell-cell adhesion complex associated with spermiation in rat seminiferous epithelium. J. Androl. 2001, 22, 1030–1052. [Google Scholar] [CrossRef]
- Takahashi, H.; Koshimizu, U.; Miyazaki J, i.c.h.i.; Nakamura, T. Impaired spermatogenic ability of testicular germ cells in mice deficient in the LIM-kinase 2 gene. Dev. Biol. 2002, 241, 259–272. [Google Scholar] [CrossRef] [PubMed]
- Mironova, E.; Millette, C.F. Expression of the diaphanous-related formin proteins mDia1 and mDia2 in the rat testis. Dev. Dyn. 2008, 237, 2170–2176. [Google Scholar] [CrossRef] [PubMed]
- Oikonomopoulou, I.; Patel, H.; Watson, P.F.; Chantler, P.D. Relocation of myosin and actin, kinesin and tubulin in the acrosome reaction of bovine spermatozoa. Reprod. Fertil. Dev. 2009, 21, 364–377. [Google Scholar] [CrossRef] [PubMed]
- Obermann, H.; Raabe, I.; Balvers, M.; Brunswig, B.; Schulze, W.; Kirchhoff, C. Novel testis-expressed profilin IV associated with acrosome biogenesis and spermatid elongation. Mol. Hum. Reprod. 2005, 11, 53–64. [Google Scholar] [CrossRef] [PubMed]
- Lui, W.; Lee, W.M.; Cheng, C.Y. Sertoli-Germ Cell Adherens Junction Dynamics in the Testis Are Regulated by RhoB GTPase via the ROCK/LIMK Signaling Pathway. Biol. Reprod. 2003, 68, 2189–2206. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, H. Novel Actin-Like Proteins T-ACTIN 1 and T-ACTIN 2 Are Differentially Expressed in the Cytoplasm and Nucleus of Mouse Haploid Germ Cells. Boil. Reprod. 2003, 69, 475–482. [Google Scholar] [CrossRef]
- Tubb, B.; Mulholland, D.J.; Vogl, W.; Lan, Z.-J.; Niederberger, C.; Cooney, A.; Bryan, J. Testis Fascin (FSCN3): A Novel Paralog of the Actin-Bundling Protein Fascin Expressed Specifically in the Elongate Spermatid Head. Exp. Cell Res. 2002, 275, 92–109. [Google Scholar] [CrossRef]
- Rawe, V.Y.; Ramalho-Santos, J.; Payne, C.; Chemes, H.E.; Schatten, G. WAVE1, an A-kinase anchoring protein, during mammalian spermatogenesis. Hum. Reprod. 2004, 19, 2594–2604. [Google Scholar] [CrossRef]
- Gennerich, A.; Vale, R.D. Walking the walk: How kinesin and dynein coordinate their steps. Curr. Opin. Cell Biol. 2009, 21, 59–67. [Google Scholar] [CrossRef]
- Ma, D.-D.; Wang, D.-H.; Yang, W.-X. Kinesins in spermatogenesis. Biol. Reprod. 2017, 96, 267–276. [Google Scholar] [CrossRef]
- Yang, W.-X.; Sperry, A.O. C-Terminal Kinesin Motor KIFC1 Participates in Acrosome Biogenesis and Vesicle Transport. Biol. Reprod. 2003, 69, 1719–1729. [Google Scholar] [CrossRef] [PubMed]
- Yu, K.; Hou, L.; Zhu, J.Q.; Ying, X.P.; Yang, W.X. KIFC1 participates in acrosomal biogenesis, with discussion of its importance for the perforatorium in the chinese mitten crab eriocheir sinensis. Cell Tissue Res. 2009, 337, 113–123. [Google Scholar] [CrossRef] [PubMed]
- Zou, Y.; Millette, C.F.; Sperry, A.O. KRP3A and KRP3B: Candidate Motors in Spermatid Maturation in the Seminiferous Epithelium. Biol. Reprod. 2002, 66, 843–855. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Mannowetz, N.; Kartarius, S.; Wennemuth, G.; Montenarh, M. Protein kinase CK2 and new binding partners during spermatogenesis. Cell. Mol. Life Sci. 2010, 67, 3905–3913. [Google Scholar] [CrossRef]
- Navolanic, P.M.; Sperry, A.O. Identification of Isoforms of a Mitotic Motor in Mammalian Spermatogenesis. Biol. Reprod. 2000, 62, 1360–1369. [Google Scholar] [CrossRef]
- Wang, W.; Zhu, J.Q.; Yu, H.M.; Tan, F.Q.; Yang, W.X. KIFC1-like motor protein associates with the cephalopod manchette and participates in sperm nuclear morphogenesis in Octopus tankahkeei. PLoS ONE 2010, 5, e15616. [Google Scholar] [CrossRef]
- Yang, W.-X.; Jefferson, H.; Sperry, A.O. The Molecular Motor KIFC1 Associates with a Complex Containing Nucleoporin NUP62 That Is Regulated During Development and by the Small GTPase RAN. Biol. Reprod. 2006, 74, 684–690. [Google Scholar] [CrossRef]
- Saade, M.; Irla, M.; Govin, J.; Victorero, G.; Samson, M.; Nguyen, C. Dynamic distribution of Spatial during mouse spermatogenesis and its interaction with the kinesin KIF17b. Exp. Cell Res. 2007, 313, 614–626. [Google Scholar] [CrossRef]
- Lehti, M.S.; Kotaja, N.; Sironen, A. KIF3A is essential for sperm tail formation and manchette function. Mol. Cell. Endocrinol. 2013, 377, 44–55. [Google Scholar] [CrossRef]
- Miller, M.G.; Mulholland, D.J.; Vogl, A.W. Rat Testis Motor Proteins Associated with Spermatid Translocation (Dynein) and Spermatid Flagella (Kinesin-II). Biol. Reprod. 1999, 60, 1047–1056. [Google Scholar] [CrossRef]
- Fawcett, D.W. The mammalian spermatozoon. In Developmental Biology; Academic Press: Cambridge, MA, USA, 1975; pp. 394–436. [Google Scholar]
- Vaid, K.S.; Guttman, J.A.; Singaraja, R.R.; Vogl, A.W. A Kinesin Is Present at Unique Sertoli/Spermatid Adherens Junctions in Rat and Mouse Testes. Biol. Reprod. 2007, 77, 1037–1048. [Google Scholar] [CrossRef] [PubMed]
- Sperry, A.O.; Zhao, L.P. Kinesin-related proteins in the mammalian testes: Candidate motors for meiosis and morphogenesis. Mol. Biol. Cell 1996, 7, 289–305. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Kotaja, N.; Macho, B.; Sassone-Corsi, P. Microtubule-independent and protein kinase A-mediated function of kinesin KIF17b controls the intracellular transport of activator of CREM in testis (ACT). J. Biol. Chem. 2005, 280, 31739–31745. [Google Scholar] [CrossRef] [PubMed]
- Chennathukuzhi, V.; Morales, C.R.; El-Alfy, M.; Hecht, N.B. The kinesin KIF17b and RNA-binding protein TB-RBP transport specific cAMP-responsive element modulator-regulated mRNAs in male germ cells. Proc. Natl. Acad. Sci. USA 2003, 100, 15566–15571. [Google Scholar] [CrossRef] [PubMed]
- Kotaja, N.; Lin, H.; Parvinen, M.; Sassone-Corsi, P. Interplay of PIWI/Argonaute protein MIWI and kinesin KIF17b in chromatoid bodies of male germ cells. J. Cell Sci. 2006, 119, 2819–2825. [Google Scholar] [CrossRef]
- Yoshida, T.; Ioshii, S.O.; Imanaka-Yoshida, K.; Izutsu, K. Association of cytoplasmic dynein with manchette microtubules and spermatid nuclear envelope during spermiogenesis in rats. J. Cell Sci. 1994, 107, 625–633. [Google Scholar]
- Li, Y.-R.; Yang, W.-X. Myosin superfamily: The multi-functional and irreplaceable factors in spermatogenesis and testicular tumors. Gene 2016, 576, 195–207. [Google Scholar] [CrossRef]
- Schwartz, J.; Li, G.; Yang, Q.; Suri, V.; Ross, J.; Alexander, E.A. Role of SNAREs and H+-ATPase in the targeting of proton pump-coated vesicles to collecting duct cell apical membrane. Kidney Int. 2007, 72, 1310–1315. [Google Scholar] [CrossRef]
- Nakamura, Y.; Asano, A.; Hosaka, Y.; Takeuchi, T.; Iwanaga, T.; Yamano, Y. Expression and intracellular localization of TBC1D9, a Rab GTPase-accelerating protein, in mouse testes. Exp. Anim. 2015, 64, 415–424. [Google Scholar] [CrossRef]
- Belmonte, S.A.; Mayorga, L.S.; Tomes, C.N. The Molecules of Sperm Exocytosis In Sperm Acrosome Biogenesis and Function During Fertilization; Springer: Cham, Switzerland, 2016; pp. 71–92. [Google Scholar]
- Hayasaka, S.; Terada, Y.; Suzuki, K.; Murakawa, H.; Tachibana, I.; Sankai, T.; Murakami, T.; Yaegashi, N.; Okamura, K. Intramanchette transport during primate spermiogenesis: Expression of dynein, myosin Va, motor recruiter myosin Va, VIIa-Rab27a/b interacting protein, and Rab27b in the manchette during human and monkey spermiogenesis. Asian J. Androl. 2008, 10, 561–568. [Google Scholar] [CrossRef]
- Lin, Y.-H.; Ke, C.-C.; Wang, Y.-Y.; Chen, M.-F.; Chen, T.-M.; Ku, W.-C.; Chiang, H.-S.; Yeh, C.-H. RAB10 Interacts with the Male Germ Cell-Specific GTPase-Activating Protein during Mammalian Spermiogenesis. Int. J. Mol. Sci. 2017, 18, 97. [Google Scholar] [CrossRef] [PubMed]
- Lau, A.S.; Mruk, D.D. Rab8B GTPase and Junction Dynamics in the Testis. Endocrinology 2003, 144, 1549–1563. [Google Scholar] [CrossRef] [PubMed]
- Mettlen, M.; Chen, P.-H.; Srinivasan, S.; Danuser, G.; Schmid, S.L. Regulation of Clathrin-Mediated Endocytosis. Annu. Rev. Biochem. 2018, 87, 871–896. [Google Scholar] [CrossRef] [PubMed]
- Kaksonen, M.; Roux, A. Mechanisms of clathrin-mediated endocytosis. Nat. Rev. Mol. Cell Biol. 2018, 19, 313–326. [Google Scholar] [CrossRef] [PubMed]
- Moreno, R.D.; Ramalho-Santos, J.; Sutovsky, P.; Chan, E.K.L.; Schatten, G. Vesicular Traffic and Golgi Apparatus Dynamics During Mammalian Spermatogenesis: Implications for Acrosome Architecture. Biol. Reprod. 2000, 63, 89–98. [Google Scholar] [CrossRef]
- Ramalho-Santos, J.; Moreno, R.D.; Wessel, G.M.; Chan, E.K.L.; Schatten, G. Membrane Trafficking Machinery Components Associated with the Mammalian Acrosome during Spermiogenesis. Exp. Cell Res. 2001, 267, 45–60. [Google Scholar] [CrossRef]
- Martínez-Menárguez, J.A.; Avilés, M.; Madrid, J.F.; Castells, M.T.; Ballesta, J. Glycosylation in Golgi apparatus of early spermatids of rat. A high resolution lectin cytochemical study. Eur. J. Cell Biol. 1993, 61, 21–33. [Google Scholar]
- Lehti, M.S.; Sironen, A. Formation and function of sperm tail structures in association with sperm motility defects. Biol. Reprod. 2017, 97, 522–536. [Google Scholar] [CrossRef]
- Bertiaux, E.; Bastin, P. Dealing with several flagella in the same cell. Cell. Microbiol. 2020, 22, e13162. [Google Scholar] [CrossRef]
- Yang, H.; Huang, K. Dissecting the Vesicular Trafficking Function of IFT Subunits. Front. Cell Dev. Biol. 2020, 7, 352. [Google Scholar] [CrossRef]
- Funaki, T.; Kon, S.; Tanabe, K.; Natsume, W.; Sato, S.; Shimizu, T.; Yoshida, N.; Wong, W.F.; Ogura, A.; Ogawa, T.; et al. The Arf GAP SMAP2 is necessary for organized vesicle budding from the trans-Golgi network and subsequent acrosome formation in spermiogenesis. Mol. Boil. Cell 2013, 24, 2633–2644. [Google Scholar] [CrossRef] [PubMed]
- Arvan, P.; Castle, D. Regulated secretory cells go to the BAR for a bud. Nat. Rev. Endocrinol. 2013, 9, 443–444. [Google Scholar] [CrossRef] [PubMed]
- Xiao, N.; Kam, C.; Shen, C.; Jin, W.; Wang, J.; Lee, K.M.; Jiang, L.; Xia, J. PICK1 deficiency causes male infertility in mice by disrupting acrosome formation. J. Clin. Investig. 2009, 119, 802–812. [Google Scholar] [CrossRef] [PubMed]
- Yao, R.; Ito, C.; Natsume, Y.; Sugitani, Y.; Yamanaka, H.; Kuretake, S.; Yanagida, K.; Sato, A.; Toshimori, K.; Noda, T. Lack of acrosome formation in mice lacking a Golgi protein, GOPC. Proc. Natl. Acad. Sci. USA 2002, 99, 11211–11216. [Google Scholar] [CrossRef]
- Banu, Y.; Matsuda, M.; Yoshihara, M.; Kondo, M.; Sutou, S.; Matsukuma, S. Golgi matrix protein gene, Golga3/Mea2, rearranged and re-expressed in pachytene spermatocytes restores spermatogenesis in the mouse. Mol. Reprod. Dev. 2002, 61, 288–301. [Google Scholar] [CrossRef]
- Hicks, S.W.; Machamer, C.E. Isoform-specific Interaction of Golgin-160 with the Golgi-associated Protein PIST. J. Biol. Chem. 2005, 280, 28944–28951. [Google Scholar] [CrossRef]
- Bentson, L.F.; Agbor, V.A.; Agbor, L.N.; Lopez, A.C.; Nfonsam, L.E.; Bornstein, S.S.; Handel, M.A.; Linder, C.C. New point mutation in Golga3 causes multiple defects in spermatogenesis. Andrology 2013, 1, 440–450. [Google Scholar] [CrossRef]
- Bel, S.; Elkis, Y.; Lerer-Goldstein, T.; Nyska, A.; Shpungin, S.; Nir, U. Loss of TMF/ARA160 Protein Renders Colonic Mucus Refractory to Bacterial Colonization and Diminishes Intestinal Susceptibility to Acute Colitis. J. Boil. Chem. 2012, 287, 25631–25639. [Google Scholar] [CrossRef]
- Miller, V.J.; Sharma, P.; Kudlyk, T.A.; Frost, L.; Rofe, A.P.; Watson, I.J.; Duden, R.; Lowe, M.; Lupashin, V.; Ungar, D. Molecular Insights into Vesicle Tethering at the Golgi by the Conserved Oligomeric Golgi (COG) Complex and the Golgin TATA Element Modulatory Factor (TMF). J. Boil. Chem. 2012, 288, 4229–4240. [Google Scholar] [CrossRef]
- Lerer-Goldshtein, T.; Bel, S.; Shpungin, S.; Pery, E.; Motro, B.; Goldstein, R.S.; Bar-Sheshet, S.I.; Breitbart, H.; Nir, U. TMF/ARA160: A key regulator of sperm development. Dev. Boil. 2010, 348, 12–21. [Google Scholar] [CrossRef]
- Clarke, G.N.; Yanagimachi, R. Actin in mammalian sperm heads. J. Exp. Zool. 1978, 205, 125–132. [Google Scholar] [CrossRef] [PubMed]
- Hall, E.A.; Keighren, M.; Ford, M.J.; Davey, T.; Jarman, A.P.; Smith, L.B.; Jackson, I.J.; Mill, P. Acute versus chronic loss of mammalian Azi1/Cep131 results in distinct ciliary phenotypes. PLoS Genet. 2013, 9, e1003928. [Google Scholar] [CrossRef] [PubMed]
- Schwarz, T.; Prieler, B.; Schmid, J.A.; Grzmil, P.; Neesen, J. Ccdc181 is a microtubule-binding protein that interacts with Hook1 in haploid male germ cells and localizes to the sperm tail and motile cilia. Eur. J. Cell Biol. 2017, 96, 276–288. [Google Scholar] [CrossRef] [PubMed]
- Huang, Q.; Liu, H.; Zeng, J.; Li, W.; Zhang, S.; Zhang, L.; Song, S.; Zhou, T.; Sutoysky, M.; Sutoysky, P.; et al. COP9 signalosome complex subunit 5, an IFT20 binding partner, is essential to maintain male germ cell survival and acrosome biogenesis. Boil. Reprod. 2019, 233–247. [Google Scholar] [CrossRef]
- Pierre, V.; Martinez, G.; Coutton, C.; Delaroche, J.; Yassine, S.; Novella, C.; Pernet-Gallay, K.; Hennebicq, S.; Ray, P.; Arnoult, C. Absence of Dpy19l2, a new inner nuclear membrane protein, causes globozoospermia in mice by preventing the anchoring of the acrosome to the nucleus. Development 2012, 139, 2955–2965. [Google Scholar] [CrossRef]
- Nozawa, Y.I.; Yao, E.; Gacayan, R.; Xu, S.-M.; Chuang, P.-T. Mammalian Fused is essential for sperm head shaping and periaxonemal structure formation during spermatogenesis. Dev. Biol. 2014, 388, 170–180. [Google Scholar] [CrossRef]
- Miyazaki, T.; Mori, M.; Yoshida, C.A.; Ito, C.; Yamatoya, K.; Moriishi, T.; Kawai, Y.; Komori, H.; Kawane, T.; Izumi, S.-I.; et al. Galnt3 deficiency disrupts acrosome formation and leads to oligoasthenoteratozoospermia. Histochem. Cell Boil. 2012, 139, 339–354. [Google Scholar] [CrossRef]
- Bizkarguenaga, M.; Gomez-Santos, L.; Madrid, J.F.; Sáez, J.F.; Alonso, E. Increase of germ cell nuclear factor expression in globozoospermic Gopc−/− knockout mice. Andrology 2019, 7, 319–328. [Google Scholar] [CrossRef]
- Follit, J.A.; San Agustin, J.T.; Xu, F.; Jonassen, J.A.; Samtani, R.; Lo, C.W.; Pazour, G.J. The Golgin GMAP210/TRIP11 anchors IFT20 to the Golgi complex. PLoS Genet. 2008, 4, e1000315. [Google Scholar] [CrossRef]
- Hou, C.-C.; Yang, W.-X. Acroframosome-dependent KIFC1 facilitates acrosome formation during spermatogenesis in the caridean shrimp Exopalaemon modestus. PLoS ONE 2013, 8, e76065. [Google Scholar] [CrossRef]
- Liu, C.; Song, Z.; Wang, L.; Yu, H.; Liu, W.; Shang, Y.; Xu, Z.; Zhao, H.; Gao, F.; Wen, J.; et al. Sirt1 regulates acrosome biogenesis by modulating autophagic flux during spermiogenesis in mice. Development 2017, 144, 441–451. [Google Scholar] [CrossRef] [PubMed]
- Kang-Decker, N.; Mantchev, G.T.; Juneja, S.C.; McNiven, M.A.; van Deursen, J.M.A. Lack of Acrosome Formation in Hrb-Deficient Mice. Science 2001, 294, 1531–1533. [Google Scholar] [CrossRef] [PubMed]
- Audouard, C.; Christians, E. Hsp90b1 knockout targeted to male germline: A mouse model for globozoospermia. Fertil. Steril. 2011, 95, 1475–1477.e4. [Google Scholar] [CrossRef] [PubMed]
- Sironen, A.; Hansen, J.; Thomsen, B.; Andersson, M.; Vilkki, J.; Toppari, J.; Kotaja, N. Expression of SPEF2 During Mouse Spermatogenesis and Identification of IFT20 as an Interacting Protein. Boil. Reprod. 2010, 82, 580–590. [Google Scholar] [CrossRef]
- Smith, L.B.; Milne, L.; Nelson, N.; Eddie, S.; Brown, P.; Atanassova, N.; O’Bryan, M.K.; O’Donnell, L.; Rhodes, D.; Wells, S.; et al. KATNAL1 Regulation of Sertoli Cell Microtubule Dynamics Is Essential for Spermiogenesis and Male Fertility. PLoS Genet. 2012, 8, e1002697. [Google Scholar] [CrossRef]
- Liu, Y.; DeBoer, K.; De Kretser, D.M.; O’Donnell, L.; O’Connor, A.; Merriner, D.J.; Okuda, H.; Whittle, B.; Jans, D.A.; Efthymiadis, A.; et al. LRGUK-1 Is Required for Basal Body and Manchette Function during Spermatogenesis and Male Fertility. PLoS Genet. 2015, 11, e1005090. [Google Scholar] [CrossRef]
- Lin, Y.-H.; Lin, Y.-M.; Kuo, Y.-C.; Wang, Y.-Y.; Kuo, P.-L. Identification and characterization of a novel Rab GTPase-activating protein in spermatids. Int. J. Androl. 2011, 34, e358–e367. [Google Scholar] [CrossRef]
- Zhang, L.; Shang, X.-J.; Li, H.-F.; Shi, Y.-Q.; Li, W.; E Teves, M.; Wang, Z.-Q.; Jiang, G.-F.; Song, S.-Z.; Zhang, Z. Characterization of membrane occupation and recognition nexus repeat containing 3, meiosis expressed gene 1 binding partner, in mouse male germ cells. Asian J. Androl. 2014, 17, 86–93. [Google Scholar] [CrossRef]
- Rivkin, E.; Kierszenbaum, A.L.; Gil, M.; Tres, L.L. Rnf19a, a ubiquitin protein ligase, and Psmc3, a component of the 26S proteasome, Tether to the acrosome membranes and the head–tail coupling apparatus during rat spermatid development. Dev. Dyn. 2009, 238, 1851–1861. [Google Scholar] [CrossRef]
- Kierszenbaum, A.L.; Tres, L.L.; Rivkin, E.; Kang-Decker, N.; van Deursen, J.M.A. The Acroplaxome Is the Docking Site of Golgi-Derived Myosin Va/Rab27a/b-Containing Proacrosomal Vesicles in Wild-Type and Hrb Mutant Mouse Spermatids. Biol. Reprod. 2004, 70, 1400–1410. [Google Scholar] [CrossRef]
- Zhou, J.; Du, Y.R.; Qin, W.H.; Hu, Y.G.; Huang, Y.N.; Bao, L.; Han, D.; Mansouri, A.; Xu, G.L. RIM-BP3 is a manchette-associated protein essential for spermiogenesis. Development 2009, 136, 373–382. [Google Scholar] [CrossRef] [PubMed]
- Fujihara, Y.; Satouh, Y.; Inoue, N.; Isotani, A.; Ikawa, M.; Okabe, M. SPACA1-deficient male mice are infertile with abnormally shaped sperm heads reminiscent of globozoospermia. Development 2012, 139, 3583–3589. [Google Scholar] [CrossRef] [PubMed]
- Ito, C.; Toshimori, K. Acrosome markers of human sperm. Anat. Sci. Int. 2016, 91, 128–142. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Shen, X.; Jones, B.H.; Xu, B.; Herr, J.C.; Strauss, J.F., III. Phosphorylation of mouse sperm axoneme central apparatus protein SPAG16L by a testis-specific kinase, TSSK2. Biol. Reprod. 2008, 79, 75–83. [Google Scholar] [CrossRef][Green Version]
- Elkis, Y.; Bel, S.; Rahimi, R.; Lerer-Goldstein, T.; Levin-Zaidman, S.; Babushkin, T.; Shpungin, S.; Nir, U. TMF/ARA160 Governs the Dynamic Spatial Orientation of the Golgi Apparatus during Sperm Development. PLoS ONE 2015, 10, e0145277. [Google Scholar] [CrossRef]
- Dunleavy, J.; O’Bryan, M.; Stanton, P.; O’Donnell, L. The cytoskeleton in spermatogenesis. Reproduction 2019, 157, R53–R72. [Google Scholar] [CrossRef]
- Paiardi, C.; Pasini, M.E.; Gioria, M.; Berruti, G. Failure of acrosome formation and globozoospermia in the wobbler mouse, a Vps54 spontaneous recessive mutant. Spermatogenesis 2011, 1, 52–62. [Google Scholar] [CrossRef]
- Lin, Y.-N.; Roy, A.; Yan, W.; Burns, K.H.; Matzuk, M.M. Loss of zona pellucida binding proteins in the acrosomal matrix disrupts acrosome biogenesis and sperm morphogenesis. Mol. Cell. Biol. 2007, 27, 6794–6805. [Google Scholar] [CrossRef]
- Vadnais, M.L.; Aghajanian, H.K.; Lin, A.; Gerton, G.L. Signaling in sperm: Toward a molecular understanding of the acquisition of sperm motility in the mouse epididymis. Biol. Reprod. 2013, 89, 127. [Google Scholar] [CrossRef]
- Peters, J.-M. The anaphase promoting complex/cyclosome: A machine designed to destroy. Nat. Rev. Mol. Cell Biol. 2006, 7, 644–656. [Google Scholar] [CrossRef]
- Yeste, M.; Morató, R.; Rodríguez-Gil, J.E.; Bonet, S.; Prieto-Martínez, N. Aquaporins in the male reproductive tract and sperm: Functional implications and cryobiology. Reprod. Domest. Anim. 2017, 52, 12–27. [Google Scholar] [CrossRef] [PubMed]
- Kawahara, C.; Yokota, S.; Fujita, H. DDX6 localizes to nuage structures and the annulus of mammalian spermatogenic cells. Histochem. Cell Biol. 2014, 141, 111–121. [Google Scholar] [CrossRef] [PubMed]
- Mohri, H.; Inaba, K.; Ishijima, S.; Baba, S.A. Tubulin-dynein system in flagellar and ciliary movement. Proc. Jpn. Acad. Ser. B. Phys. Biol. Sci. 2012, 88, 397–415. [Google Scholar] [CrossRef] [PubMed]
- Lim, C.H.; Hamazaki, T.; Braun, E.L.; Wade, J.; Terada, N. Evolutionary genomics implies a specific function of Ant4 in mammalian and anole lizard male germ cells. PLoS ONE 2011, 6, e23122. [Google Scholar] [CrossRef]
- Zhu, Z.; Li, R.; Wang, L.; Zheng, Y.; Hoque, S.A.M.; Lv, Y.; Zeng, W. Glycogen Synthase Kinase-3 Regulates Sperm Motility and Acrosome Reaction via Affecting Energy Metabolism in Goats. Front. Physiol. 2019, 10, 968. [Google Scholar] [CrossRef]
- Revelli, A.; Ghigo, D.; Moffa, F.; Massobrio, M.; Tur-Kaspa, I. Guanylate Cyclase Activity and Sperm Function. Endocr. Rev. 2002, 23, 484–494. [Google Scholar] [CrossRef]
- Grad, I.; Cederroth, C.R.; Walicki, J.; Grey, C.; Barluenga, S.; Winssinger, N.; De Massy, B.; Nef, S.; Picard, D. The Molecular Chaperone Hsp90α Is Required for Meiotic Progression of Spermatocytes beyond Pachytene in the Mouse. PLoS ONE 2010, 5, e15770. [Google Scholar] [CrossRef]
- Myrick, D.A.; Christopher, M.A.; Scott, A.M.; Simon, A.K.; Donlin-Asp, P.G.; Kelly, W.G.; Katz, D.J. KDM1A/LSD1 regulates the differentiation and maintenance of spermatogonia in mice. PLoS ONE 2017, 12, e0177473. [Google Scholar] [CrossRef]
- Mills, R.D.; Mulhern, T.D.; Liu, F.; Culvenor, J.G.; Cheng, H.-C. Prediction of the Repeat Domain Structures and Impact of Parkinsonism-Associated Variations on Structure and Function of all Functional Domains of Leucine-Rich Repeat Kinase 2 (LRRK2). Hum. Mutat. 2014, 35, 395–412. [Google Scholar] [CrossRef]
- Ratliff, M.; Hill-Harfe, K.L.; Gleason, E.J.; Ling, H.; Kroft, T.L.; L’Hernault, S.W. MIB-1 Is Required for Spermatogenesis and Facilitates LIN-12 and GLP-1 Activity in Caenorhabditis elegans. Genetics 2018, 209, 173–193. [Google Scholar] [CrossRef]
- Rojansky, R.; Cha, M.-Y.; Chan, D.C. Elimination of paternal mitochondria in mouse embryos occurs through autophagic degradation dependent on PARKIN and MUL1. Elife 2016, 5, e17896. [Google Scholar] [CrossRef] [PubMed]
- Coene, K.; Mans, R.A.; Boldt, K.; Gloeckner, C.J.; Van Reeuwijk, J.; Bolat, E.; Roosing, S.; Letteboer, S.J.; Peters, T.A.; Cremers, F.P.; et al. The ciliopathy-associated protein homologs RPGRIP1 and RPGRIP1L are linked to cilium integrity through interaction with Nek4 serine/threonine kinase. Hum. Mol. Genet. 2011, 20, 3592–3605. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Pinheiro, A.; Silva, M.; Graça, M.I.P.D.S.; Silva, J.; Sá, R.; Sousa, M.; Barros, A.; De Almeida, I.T.; Rivera, I. Pyruvate dehydrogenase complex: MRNA and protein expression patterns of E1α subunit genes in human spermatogenesis. Gene 2012, 506, 173–178. [Google Scholar] [CrossRef] [PubMed]
- Neill, A.T.; Moy, G.W.; Vacquier, V.D. Polycystin-2 associates with the polycystin-1 homolog, suREJ3, and localizes to the acrosomal region of sea urchin spermatozoa. Mol. Reprod. Dev. 2004, 67, 472–477. [Google Scholar] [CrossRef]
- Silva, J.V.; Fardilha, M.J.F.; Fardilha, M. Phosphoprotein Phosphatase 1 Complexes in Spermatogenesis. Curr. Mol. Pharmacol. 2014, 7, 136–146. [Google Scholar] [CrossRef]
- Wang, C.; Fok, K.L.; Cai, Z.; Chen, H.; Chan, H.C. CD147 regulates extrinsic apoptosis in spermatocytes by modulating NFκB signaling pathways. Oncotarget 2017, 8, 3132–3143. [Google Scholar] [CrossRef][Green Version]
- Manku, G.; Wing, S.S.; Culty, M. Expression of the Ubiquitin Proteasome System in Neonatal Rat Gonocytes and Spermatogonia: Role in Gonocyte Differentiation1. Biol. Reprod. 2012, 44, 1–18. [Google Scholar] [CrossRef]
- Evans, C.S.; Holzbaur, E.L.F. Autophagy and mitophagy in ALS. Neurobiol. Dis. 2019, 122, 35–40. [Google Scholar] [CrossRef]
- Yu, Y.; Vanhorne, J.; Oko, R. The origin and assembly of a zona pellucida binding protein, IAM38, during spermiogenesis. Microsc. Res. Tech. 2009, 72, 558–565. [Google Scholar] [CrossRef]
- Kierszenbaum, A.L. Tyrosine protein kinases and spermatogenesis: Truncation matters. Mol. Reprod. Dev. 2006, 73, 399–403. [Google Scholar] [CrossRef]
- Wei, Y.-L.; Yang, W.-X. The acroframosome-acroplaxome-manchette axis may function in sperm head shaping and male fertility. Gene 2018, 660, 28–40. [Google Scholar] [CrossRef] [PubMed]
- Coutton, C.; Escoffier, J.; Martinez, G.; Arnoult, C.; Ray, P.F. Teratozoospermia: Spotlight on the main genetic actors in the human. Hum. Reprod. Update 2015, 21, 455–485. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Shi, Y.; Ma, S.; Huang, Q.; Yap, Y.T.; Shi, L.; Zhang, S.; Zhou, T.; Li, W.; Hu, B.; et al. Abnormal fertility, acrosome formation, IFT20 expression and localization in the conditional Gmap210 knockout mice. Am. J. Physiol. Physiol. 2019, C174–C190. [Google Scholar] [CrossRef] [PubMed]
- Miyamoto, Y.; Yoneda, Y.; Oka, M. Protein Transport Between the Nucleus and Cytoplasm. Nucl. Archit. Dyn. 2018, 387–403. [Google Scholar] [CrossRef]
- Kierszenbaum, A.L. Intramanchette transport (IMT): Managing the making of the spermatid head, centrosome, and tail. Mol. Reprod. Dev. 2002, 63, 1–4. [Google Scholar] [CrossRef] [PubMed]
- Bao, J.; Zhang, J.; Zheng, H.; Xu, C.; Yan, W. UBQLN1 interacts with SPEM1 and participates in spermiogenesis. Mol. Cell. Endocrinol. 2010, 327, 89–97. [Google Scholar] [CrossRef] [PubMed]
- Bao, J.; Wu, Q.; Song, R.; Jie, Z.; Zheng, H.; Xu, C.; Yan, W. RANBP17 is localized to the XY body of spermatocytes and interacts with SPEM1 on the manchette of elongating spermatids. Mol. Cell. Endocrinol. 2010, 333, 134–142. [Google Scholar] [CrossRef]
- Tapia Contreras, C.; Hoyer-Fender, S. CCDC42 Localizes to Manchette, HTCA and Tail and Interacts With ODF1 and ODF2 in the Formation of the Male Germ Cell Cytoskeleton. Front. cell Dev. Biol. 2019, 7, 151. [Google Scholar] [CrossRef]
- Pasek, R.; Malarkey, E.; Berbari, N.F.; Sharma, N.; Kesterson, R.A.; Tres, L.L.; Kierszenbaum, A.L.; Yoder, B.K. Coiled-coil domain containing 42 (Ccdc42) is necessary for proper sperm development and male fertility in the mouse. Dev. Boil. 2016, 412, 208–218. [Google Scholar] [CrossRef]
- Mochida, K.; Tres, L.L.; Kierszenbaum, A.L. Isolation of the Rat Spermatid Manchette and Its Perinuclear Ring. Dev Biol. 1998, 200, 46–56. [Google Scholar] [CrossRef]
- Fouquet, J.; Kann, M.; Soues, S.; Melki, R. ARP1 in Golgi organisation and attachment of manchette microtubules to the nucleus during mammalian spermatogenesis. J. Cell Sci. 2000, 113, 877–886. [Google Scholar] [PubMed]
- Li, R.-K.; Tan, J.-L.; Chen, L.-T.; Feng, J.-S.; Liang, W.-X.; Guo, X.; Liu, P.; Chen, Z.; Sha, J.-H.; Wang, Y.-F.; et al. Iqcg Is Essential for Sperm Flagellum Formation in Mice. PLoS ONE 2014, 9, e98053. [Google Scholar] [CrossRef] [PubMed]
- Pleuger, C.; Lehti, M.S.; Cooper, M.; O’Connor, A.E.; Merriner, D.J.; Smyth, I.M.; Cottle, D.; Fietz, D.; Bergmann, M.; O’Bryan, M.K. CBE1 Is a Manchette- and Mitochondria-Associated Protein With a Potential Role in Somatic Cell Proliferation. Endocrinology 2019, 160, 2573–2586. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.-Y.; Wang, Y.-Y.; Chen, Y.-L.; Chen, M.-F.; Chiang, H.-S.; Kuo, P.-L.; Lin, Y.-H. CDC42 Negatively Regulates Testis-Specific SEPT12 Polymerization. Int. J. Mol. Sci. 2018, 19, 2627. [Google Scholar] [CrossRef]
- Akhmanova, A.; Mausset-Bonnefont, A.L.; van Cappellen, W.; Keijzer, N.; Hoogenraad, C.C.; Stepanova, T.; Drabek, K.; van der Wees, J.; Mommaas, M.; Onderwater, J.; et al. The microtubule plus-end-tracking protein CLIP-170 associates with the spermatid manchette and is essential for spermatogenesis. Genes Dev. 2005, 19, 2501–2515. [Google Scholar] [CrossRef]
- Liška, F.; Gosele, C.; Rivkin, E.; Tres, L.; Cardoso, M.C.; Domaing, P.; Krejčí, E.; Šnajdr, P.; Lee-Kirsch, M.A.; de Rooij, D.G.; et al. Rat hd mutation reveals an essential role of centrobin in spermatid head shaping and assembly of the head-tail coupling apparatus. Biol. Reprod. 2009, 81, 1196–1205. [Google Scholar] [CrossRef]
- Zheng, C.; Ouyang, Y.-C.; Jiang, B.; Lin, X.; Chen, J.; Dong, M.-Z.; Zhuang, X.; Yuan, S.; Sun, Q.-Y.; Han, C. Non-canonical RNA polyadenylation polymerase FAM46C is essential for fastening sperm head and flagellum in mice. Boil. Reprod. 2019, 100, 1673–1685. [Google Scholar] [CrossRef]
- Hamilton, L.E.; Acteau, G.; Xu, W.; Sutovsky, P.; Oko, R. The developmental origin and compartmentalization of glutathione-s-transferase omega 2 isoforms in the perinuclear theca of eutherian spermatozoa. Biol. Reprod. 2017, 97, 612–621. [Google Scholar] [CrossRef]
- Walenta, J.H.; Didier, A.J.; Liu, X.; Krämer, H. The Golgi-Associated Hook3 Protein Is a Member of a Novel Family of Microtubule-Binding Proteins. J. Cell Biol. 2001, 152, 923–934. [Google Scholar] [CrossRef]
- Okuda, H.; DeBoer, K.; O’Connor, A.; Merriner, D.J.; Jamsai, D.; O’Bryan, M.K. LRGUK1 is part of a multiprotein complex required for manchette function and male fertility. FASEB J. 2016, 31, 1141–1152. [Google Scholar] [CrossRef]
- Lehti, M.S.; Zhang, F.-P.; Kotaja, N.; Sironen, A. SPEF2 functions in microtubule-mediated transport in elongating spermatids to ensure proper male germ cell differentiation. Development 2017, 144, 2683–2693. [Google Scholar] [CrossRef] [PubMed]
- Harris, T.P.; Schimenti, K.J.; Munroe, R.J.; Schimenti, J.C. IQ motif-containing G (Iqcg) is required for mouse spermiogenesis. G3 2014, 4, 367–372. [Google Scholar] [CrossRef] [PubMed]
- Dunleavy, J.; Okuda, H.; O’Connor, A.; Merriner, D.J.; O’Donnell, L.; Jamsai, D.; Bergmann, M.; O’Bryan, M.K. Katanin-like 2 (KATNAL2) functions in multiple aspects of haploid male germ cell development in the mouse. PLoS Genet. 2017, 13, e1007078. [Google Scholar] [CrossRef] [PubMed]
- O’Donnell, L.; Rhodes, D.; Smith, S.J.; Merriner, D.J.; Clark, B.J.; Borg, C.; Whittle, B.; O’Connor, A.; Smith, L.B.; McNally, F.J.; et al. An Essential Role for Katanin p80 and Microtubule Severing in Male Gamete Production. PLoS Genet. 2012, 8, e1002698. [Google Scholar] [CrossRef]
- Nayernia, K.; Vauti, F.; Meinhardt, A.; Cadenas, C.; Schweyer, S.; Meyer, B.I.; Schwandt, I.; Chowdhury, K.; Engel, W.; Arnold, H.H. Inactivation of a Testis-specific Lis1 Transcript in Mice Prevents Spermatid Differentiation and Causes Male Infertility. J. Boil. Chem. 2003, 278, 48377–48385. [Google Scholar] [CrossRef]
- Li, W.; Tang, W.; Teves, M.E.; Zhang, Z.; Zhang, L.; Li, H.; Archer, K.J.; Peterson, D.L.; Williams, D.C., Jr.; Strauss, J.F., III; et al. A MEIG1/PACRG complex in the manchette is essential for building the sperm flagella. Development 2015, 142, 921–930. [Google Scholar] [CrossRef]
- Zhang, Z.; Shen, X.; Gude, D.R.; Wilkinson, B.M.; Justice, M.J.; Flickinger, C.J.; Herr, J.C.; Eddy, E.M.; Strauss, J.F. MEIG1 is essential for spermiogenesis in mice. Proc. Natl. Acad. Sci. USA 2009, 106, 17055–17060. [Google Scholar] [CrossRef]
- Lin, Y.-H.; Lin, Y.-M.; Wang, Y.-Y.; Yu, I.-S.; Lin, Y.-W.; Wang, Y.-H.; Wu, C.-M.; Pan, H.-A.; Chao, S.-C.; Yen, P.H.; et al. The Expression Level of Septin12 Is Critical for Spermiogenesis. Am. J. Pathol. 2009, 174, 1857–1868. [Google Scholar] [CrossRef]
- Yeh, C.-H.; Kuo, P.L.; Wang, Y.Y.; Wu, Y.Y.; Chen, M.F.; Lin, D.Y.; Lai, T.H.; Chiang, H.S.; Lin, Y.H. SEPT12/SPAG4/LAMINB1 complexes are required for maintaining the integrity of the nuclear envelope in postmeiotic male germ cells. PLoS ONE 2015, 10, e0120722. [Google Scholar] [CrossRef]
- Yeh, C.H.; Wang, Y.Y.; Wee, S.K.; Chen, M.F.; Chiang, H.S.; Kuo, P.L.; Lin, Y.H. Testis-Specific SEPT12 Expression Affects SUN Protein Localization and is Involved in Mammalian Spermiogenesis. Int. J. Mol. Sci. 2019, 20, 1163. [Google Scholar] [CrossRef]
- Sironen, A.; Kotaja, N.; Mulhern, H.; Wyatt, T.A.; Sisson, J.H.; Pavlik, J.A.; Miiluniemi, M.; Fleming, M.D.; Lee, L. Loss of SPEF2 function in mice results in spermatogenesis defects and primary ciliary dyskinesia. Boil. Reprod. 2011, 85, 690–701. [Google Scholar] [CrossRef] [PubMed]
- Martins, L.R.; Bung, R.K.; Koch, S.; Richter, K.; Schwarzmüller, L.; Terhardt, D.; Kurtulmus, B.; Niehrs, C.; Rouhi, A.; Lohmann, I.; et al. Stk33 is required for spermatid differentiation and male fertility in mice. Dev. Boil. 2018, 433, 84–93. [Google Scholar] [CrossRef] [PubMed]
- Pasch, E.; Link, J.; Beck, C.; Scheuerle, S.; Alsheimer, M. The LINC complex component Sun4 plays a crucial role in sperm head formation and fertility. Biol. Open 2015, 4, 1792–1802. [Google Scholar] [CrossRef] [PubMed]
- Yang, K.; Adham, I.M.; Meinhardt, A.; Hoyer-Fender, S. Ultra-structure of the sperm head-to-tail linkage complex in the absence of the spermatid-specific LINC component SPAG4. Histochem. Cell Biol. 2018, 150, 49–59. [Google Scholar] [CrossRef]
- Kaneko, T.; Minohara, T.; Shima, S.; Yoshida, K.; Fukuda, A.; Iwamori, N.; Inai, T.; Iida, H. A membrane protein, TMCO5A, has a close relationship with manchette microtubules in rat spermatids during spermiogenesis. Mol. Reprod. Dev. 2019, 86, 330–341. [Google Scholar] [CrossRef]
- Yamase, K.; Tanigawa, Y.; Yamamoto, Y.; Tanaka, H.; Komiya, T. Mouse TMCO5 is localized to the manchette microtubules involved in vesicle transfer in the elongating spermatids. PLoS ONE 2019, 14, e0220917. [Google Scholar] [CrossRef]
- Hamilton, L.; Suzuki, J.; Acteau, G.; Shi, M.; Xu, W.; Meinsohn, M.-C.; Sutovsky, P.; Oko, R. WBP2 shares a common location in mouse spermatozoa with WBP2NL/PAWP and like its descendent is a candidate mouse oocyte-activating factor. Boil. Reprod. 2018, 99, 1171–1183. [Google Scholar] [CrossRef]
- Fok, K.L.; Chen, H.; Ruan, Y.C.; Chan, H.C. Novel regulators of spermatogenesis. Semin. Cell Dev. Biol. 2014, 29, 31–42. [Google Scholar] [CrossRef]
- Breitbart, H.; Etkovitz, N. Role and regulation of EGFR in actin remodeling in sperm capacitation and the acrosome reaction. Asian J. Androl. 2011, 13, 106–110. [Google Scholar] [CrossRef]
- Puglisi, R.; Maccari, I.; Pipolo, S.; Mangia, F.; Boitani, C. The nuclear form of glutathione peroxidase 4 colocalizes and directly interacts with protamines in the nuclear matrix during mouse sperm chromatin assembly. Spermatogenesis 2014, 4, e28460. [Google Scholar] [CrossRef]
- Kaidanovich-Beilin, O.; Woodgett, J.R. GSK-3: Functional Insights from Cell Biology and Animal Models. Front. Mol. Neurosci. 2011, 4, 40. [Google Scholar] [CrossRef] [PubMed]
- Shen, H.; Fan, X.; Zhang, Z.; Xi, H.; Ji, R.; Liu, Y.; Yue, M.; Li, Q.; He, J. Effects of elevated ambient temperature and local testicular heating on the expressions of heat shock protein 70 and androgen receptor in boar testes. Acta Histochem. 2019, 121, 297–302. [Google Scholar] [CrossRef] [PubMed]
- Gibbs, G.M.; Bianco, D.M.; Jamsai, D.; Herlihy, A.; Ristevski, S.; Aitken, R.; De Kretser, D.M.; O’Bryan, M.K. Cysteine-Rich Secretory Protein 2 Binds to Mitogen-Activated Protein Kinase Kinase Kinase 11 in Mouse Sperm. Boil. Reprod. 2007, 77, 108–114. [Google Scholar] [CrossRef] [PubMed]
- Kierszenbaum, A.L. Sperm axoneme: A tale of tubulin posttranslation diversity. Mol. Reprod. Dev. 2002, 62, 1–3. [Google Scholar] [CrossRef] [PubMed]
- Krämer, A.; Green, J.; Pollard, J., Jr.; Tugendreich, S. Causal analysis approaches in Ingenuity Pathway Analysis. Bioinformatics 2014, 30, 523–530. [Google Scholar] [CrossRef] [PubMed]
- Pleuger, C.; Fietz, D.; Hartmann, K.; Schuppe, H.-C.; Weidner, W.; Kliesch, S.; Baker, M.; O’Bryan, M.K.; Bergmann, M. Expression of ciliated bronchial epithelium 1 during human spermatogenesis. Fertil. Steril. 2017, 108, 47–54. [Google Scholar] [CrossRef] [PubMed]
- Müller, J.M.M.; Deinhardt, K.; Rosewell, I.; Warren, G.; Shima, D.T. Targeted deletion of p97 (VCP/CDC48) in mouse results in early embryonic lethality. Biochem. Biophys. Res. Commun. 2007, 354, 459–465. [Google Scholar] [CrossRef]
- Heilemann, M. Fluorescence microscopy beyond the diffraction limit. J. Biotechnol. 2010, 149, 243–251. [Google Scholar] [CrossRef]
- Sigal, Y.M.; Zhou, R.; Zhuang, X. Visualizing and discovering cellular structures with super-resolution microscopy. Science 2018, 361, 880–887. [Google Scholar] [CrossRef]
- Krapf, D.; Metzler, R. Strange interfacial molecular dynamics. Phys. Today 2019, 72, 48–54. [Google Scholar] [CrossRef]
- Manzo, C.; Garcia-Parajo, M.F. A review of progress in single particle tracking: From methods to biophysical insights. Reports Prog. Phys. 2015, 78, 124601. [Google Scholar] [CrossRef] [PubMed]
- Xu, K.; Zhong, G.; Zhuang, X. Actin, spectrin, and associated proteins form a periodic cytoskeletal structure in axons. Science 2013, 339, 452–456. [Google Scholar] [CrossRef] [PubMed]
- He, J.; Zhou, R.; Wu, Z.; Carrasco, M.A.; Kurshan, P.T.; Farley, J.E.; Simon, D.J.; Wang, G.; Han, B.; Hao, J.; et al. Prevalent presence of periodic actin-spectrin-based membrane skeleton in a broad range of neuronal cell types and animal species. Proc. Natl. Acad. Sci. USA 2016, 113, 6029–6034. [Google Scholar] [CrossRef] [PubMed]
- Sadegh, S.; Higgins, J.L.; Mannion, P.C.; Tamkun, M.M.; Krapf, D. Plasma Membrane is Compartmentalized by a Self-Similar Cortical Actin Meshwork. Phys. Rev. X 2017, 7, 11031. [Google Scholar] [CrossRef] [PubMed]
- Gervasi, M.G. The actin cytoskeleton of the mouse sperm flagellum is organized in a helical structure. J. Cell Sci. 2018, 131, jcs215897. [Google Scholar] [CrossRef]
- Ehmann, N.; Sauer, M.; Kittel, R.J. Super-resolution microscopy of the synaptic active zone. Front. Cell. Neurosci. 2015, 9, 7. [Google Scholar] [CrossRef]
- Arnspang, E.C.; Sengupta, P.; Mortensen, K.I.; Jensen, H.H.; Hahn, U.; Jensen, E.B.V.; Lippincott-Schwartz, J.; Nejsum, L.N. Regulation of Plasma Membrane Nanodomains of the Water Channel Aquaporin-3 Revealed by Fixed and Live Photoactivated Localization Microscopy. Nano Lett. 2018, 19, 699–707. [Google Scholar] [CrossRef]
- Akin, E.J.; Solé, L.; Johnson, B.; El Beheiry, M.; Masson, J.-B.; Krapf, D.; Tamkun, M.M. Single-Molecule Imaging of Na v 1.6 on the Surface of Hippocampal Neurons Reveals Somatic Nanoclusters. Biophys. J. 2016, 111, 1235–1247. [Google Scholar] [CrossRef]
- Martínez-Muñoz, L.; Rodríguez-Frade, J.M.; Barroso, R.; Sorzano C Ó, S.; Torreño-Pina, J.A.; Santiago, C.A.; Manzo, C.; Lucas, P.; García-Cuesta, E.M.; Gutierrez, E.; et al. Separating Actin-Dependent Chemokine Receptor Nanoclustering from Dimerization Indicates a Role for Clustering in CXCR4 Signaling and Function. Mol. Cell 2018, 70, 873. [Google Scholar] [CrossRef]
- Romarowski, A.; Félix, Á.G.V.; Rodríguez, P.T.; Gervasi, M.G.; Xu, X.; Luque, G.; Contreras-Jiménez, G.; Sánchez-Cárdenas, C.; Ramírez-Gómez, H.V.; Krapf, D.; et al. Super-resolution imaging of live sperm reveals dynamic changes of the actin cytoskeleton during acrosomal exocytosis. J. Cell Sci. 2018, 131, jcs218958. [Google Scholar] [CrossRef]
- Chung, J.-J.; Shim, S.-H.; Everley, R.A.; Gygi, S.P.; Zhuang, X.; E Clapham, D. Structurally distinct Ca(2+) signaling domains of sperm flagella orchestrate tyrosine phosphorylation and motility. Cell 2014, 157, 808–822. [Google Scholar] [CrossRef] [PubMed]
- Chung, J.-J.; Miki, K.; Kim, D.; Shim, S.-H.; Shi, H.; Hwang, J.Y.; Cai, X.; Iseri, Y.; Zhuang, X.; E Clapham, D. CatSperζ regulates the structural continuity of sperm Ca2+ signaling domains and is required for normal fertility. eLife 2017, 6. [Google Scholar] [CrossRef] [PubMed]
- Hwang, J.Y.; Mannowetz, N.; Zhang, Y.; Everley, R.A.; Gygi, S.P.; Bewersdorf, J.; Lishko, P.V.; Chung, J.-J. Dual Sensing of Physiologic pH and Calcium by EFCAB9 Regulates Sperm Motility. Cell 2019, 177, 1480–1494. [Google Scholar] [CrossRef] [PubMed]
- Stival, C.; Ritagliati, C.; Xu, X.; Gervasi, M.G.; Luque, G.; Graf, C.B.; De La Vega-Beltrán, J.L.; Torres, N.I.; Darszon, A.; Krapf, D.; et al. Disruption of protein kinase A localization induces acrosomal exocytosis in capacitated mouse sperm. J. Boil. Chem. 2018, 293, 9435–9447. [Google Scholar] [CrossRef]
- Alvau, A.; Battistone, M.A.; Gervasi, M.G.; Navarrete, F.A.; Xu, X.; Sánchez-Cárdenas, C.; De La Vega-Beltran, J.L.; Da Ros, V.G.; Greer, P.A.; Darszon, A.; et al. The tyrosine kinase FER is responsible for the capacitation-associated increase in tyrosine phosphorylation in murine sperm. Development 2016, 143, 2325–2333. [Google Scholar] [CrossRef]
- Frolikova, M.; Valasková, E.; Černý, J.; Lumeau, A.; Sebkova, N.; Palenikova, V.; Sanches-Hernandez, N.; Pohlová, A.; Postlerova, P.; Komrskova, K.; et al. Addressing the Compartmentalization of Specific Integrin Heterodimers in Mouse Sperm. Int. J. Mol. Sci. 2019, 20, 1004. [Google Scholar] [CrossRef]
- Frolikova, M.; Sebkova, N.; Ded, L.; Dvorakova-Hortova, K. Characterization of CD46 and β1 integrin dynamics during sperm acrosome reaction. Sci. Rep. 2016, 6, 33714. [Google Scholar] [CrossRef]
- Sharif, M.; Silva, E.; Shah, S.T.A.; Miller, D.J. Redistribution of soluble N-ethylmaleimide-sensitive-factor attachment protein receptors in mouse sperm membranes prior to the acrosome reaction. Biol. Reprod. 2017, 96, 352–365. [Google Scholar] [CrossRef][Green Version]
- Miller, M.R.; Kenny, S.J.; Mannowetz, N.; Mansell, S.A.; Wojcik, M.; Mendoza, S.; Zucker, R.S.; Xu, K.; Lishko, P.V. Asymmetrically Positioned Flagellar Control Units Regulate Human Sperm Rotation. Cell Rep. 2018, 290122. [Google Scholar] [CrossRef]
- Iyengar, P.V.; Hirota, T.; Hirose, S.; Nakamura, N. Membrane-Associated RING-CH 10 (MARCH10) is a Microtubule-Associated E3 Ubiquitin Ligase of the Spermatid Flagella. J. Biol. Chem. 2011, 286, 39082–39090. [Google Scholar] [CrossRef]
- Cesario, M.M.; Bartles, J.R. Compartmentalization, processing and redistribution of the plasma membrane protein CE9 on rodent spermatozoa. Relationship of the annulus to domain boundaries in the plasma membrane of the tail. J. Cell Sci. 1994, 107, 561–570. [Google Scholar] [PubMed]
- Krapf, D. Compartmentalization of the plasma membrane. Curr. Opin. Cell Biol. 2018, 53, 15–21. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.V.; Buchwalter, R.A.; Kao, L.-R.; Megraw, T.L. A Splice Variant of Centrosomin Converts Mitochondria to Microtubule-Organizing Centers. Curr. Biol. 2017, 27, 1928–1940.e6. [Google Scholar] [CrossRef] [PubMed]
- Vieillard, J.; Paschaki, M.; Duteyrat, J.-L.; Augière, C.; Cortier, E.; Lapart, J.-A.; Thomas, J.; Durand, B. Transition zone assembly and its contribution to axoneme formation in Drosophila male germ cells. J. Cell Boil. 2016, 214, 875–889. [Google Scholar] [CrossRef]
- Ito, C.; Yamatoya, K.; Toshimori, K. Analysis of the complexity of the sperm acrosomal membrane by super-resolution stimulated emission depletion microscopy compared with transmission electron microscopy. Microscopy 2014, 64, 279–287. [Google Scholar] [CrossRef]

| Protein | Function | Specie | References |
|---|---|---|---|
| α-ACTININ | Structure and stabilization of the cytoskeleton | Bovine | [59] |
| ARC | Possible role in the acrosome formation and the sperm acrosome reaction | Mouse | [60] |
| ARPM1 | Germ cell morphogenesis | Mouse | [61] |
| ARP2/3 | Actin polymerization | Mouse, rat, guinea pig | [45] |
| ARP3 | Actin nucleation and branching | Rat | [62] |
| CALICIN | Stabilization of the cytoskeleton | Human, mouse, rat, boar, guinea pig, bull | [63] |
| CAPZA3 | Maintains polymerized actin during spermiogenesis | Mouse | [64,65] |
| CDC42 | Actin reorganization | Rat | [66] |
| CORTACTIN | Arp2/3 complex activation; formation of actin network | Rat | [62] |
| LIMK2 | Cofilin inactivation; germ cell development | Mouse | [67] |
| MDIA1/2 | Actin nucleation | Rat | [68] |
| MYOSIN | Molecular motors | Human, mouse, rat, bovine | [69] |
| N-WASP | Arp2/3 complex activation | Rat | [62] |
| PROFILIN III | Actin monomer binding; germ cell morphogenesis | Mouse | [61] |
| PROFILIN IV | Actin monomer binding; germ cell morphogenesis | Human, mouse, rat | [70] |
| RAC1 | Actin reorganization | Rat | [66] |
| RHOB | Actin reorganization; regulation of Sertoli–germ cell adhesion | Rat | [71] |
| T-ACTIN 1/2 | Germ cell morphogenesis/sperm function | Mouse | [72] |
| TESTIS FASCIN | Actin monomer binding; germ cell morphogenesis | Human, mouse | [73] |
| WAVE1 | Arp2/3 complex activation; germ cell morphogenesis/sperm function | Human, mouse, bull, baboon | [74] |
| Stages | Kinesin Type | Functions | Species | Reference |
|---|---|---|---|---|
| Acrosome biogenesis | KIFC1 | Transport vehicles; tether acrosome to the nucleus | Rat and crustaceans | [77,78] |
| KRP3A, KRP3B | Transport vehicles like Golgi | Rat and bull | [79] | |
| KIF5C | Redistribution of proteins related to acrosome formation | Mouse | [80] | |
| Nuclear shaping | KIFC5A | Interact with manchette and promote nuclear shaping | Mouse | [81] |
| KIFC1 | Link the nucleus to the manchette; promote nucleus condensation and elongation | Mouse and Octopus tankahkeei | [82,83] | |
| KIF17B | Promote nuclear shaping in a manchette-dependent way | Mouse | [84] | |
| KIF3A | Manchette organization and sperm head shaping | Mouse | [85] | |
| Tail formation | KIF17B | Intraflagellar transport | Rat | [86] |
| KIF3A | Axoneme formation | Mouse | [85] | |
| KLC3 | Transport mitochondria to midpiece | Rat | [87] | |
| Spermatid maturation | KIF20 | Spermatid translocation | Rat and mouse | [88] |
| KRP3 | Spermatid translocation | Rat | [89] | |
| Spermatid transcription | KIF17B | ACT subcellular distribution | Monkey | [90] |
| KIF17B | Transport CREM mRNA | Mouse | [91] | |
| KIF17B | Chromatoid body movement, RNA metabolism | Mouse | [92] |
| Protein | Type of Interaction Confirmation | Reference |
|---|---|---|
| ACTIN | IPA | [119] |
| ARP2/3 | IPA | [45] |
| ATG7 | IPA | [41] |
| AU040320 | ― | [39] |
| CEP131 | IPA | [120] |
| CCDC181 | IPA | [121] |
| CLATHRIN | IPA | [103] |
| COPS5 | Co-IP, Y2H, and IPA | [122] |
| CORTACTIN | IPA | [47] |
| CSNK2A2 | IPA | [80] |
| DYP19L2 | IPA | [123] |
| FERT | IPA | [47] |
| FU | IPA | [124] |
| GALNT3 | IPA | [125] |
| GCNF | IPA | [126] |
| GOLGA3 | IPA | [113,114,115] |
| GMAP210 | IPA | [28,127] |
| GM130 | IPA | [128] |
| GOPC | IPA | [129] |
| HRB | IPA | [130] |
| HSP90B1 | IPA | [131] |
| IFT20 | Co-IP, Y2H, and IPA | [30,132] |
| IFT74 | IPA | [33] |
| IFT88 | IPA | [28] |
| KATNB1 | IPA | [133] |
| LRGUK1 | ― | [134] |
| MGCRABGAP | Co-IP | [99,135] |
| MNS1 | IPA | [85] |
| MORN3 | IPA | [136] |
| MYOSIN | IPA | [44] |
| PICK1 | IPA | [111] |
| PSMC3 | IPA | [137] |
| RAB3A | Co-IP | [135] |
| RAB27 | IPA | [138] |
| RIMBP3 | IPA | [139] |
| RNF19A | IPA | [137] |
| SIRT1 | IPA | [129] |
| SMAP2 | IPA | [109,112] |
| SNAPIN | Y2H | [26] |
| SPACA1 | IPA | [140,141] |
| SPAG6 | Co-IP, Y2H, and IPA | [26] |
| SPAG16 | Co-IP and IPA | [142] |
| SPAG17 | IPA | [21] |
| SPEF2 | Co-IP, Y2H, and IPA | [132] |
| SPINK2 | Y2H | [26] |
| STK36 | IPA | [124] |
| TMF | IPA | [143] |
| α/β-TUBULIN | IPA | [144] |
| VPS54 | IPA | [145] |
| ZPBP | ― | [146] |
| New Proteins Detected by IPA Software | ||
| AKT1 | IPA | [147] |
| APC | IPA | [148] |
| AQP1 | IPA | [149] |
| DDX6 | IPA | [150] |
| DYNC1H1 | IPA | [151] |
| GNB5 | IPA | [152] |
| GSK3B | IPA | [153] |
| GUK1 | IPA | [154] |
| HSP90 | IPA | [155] |
| KDM1A | IPA | [156] |
| KIF5C | IPA | [151] |
| KIFC3 | IPA | [151] |
| LRRK2 | IPA | [157] |
| MIB1 | IPA | [158] |
| MUL1 | IPA | [159] |
| NEK4 | IPA | [160] |
| PDHA1 | IPA | [161] |
| PKD2 | IPA | [162] |
| PPP1CA | IPA | [163] |
| SLC25A6 | IPA | [152] |
| TRAF2 | IPA | [164] |
| UBE2 | IPA | [165] |
| VCP | IPA | [166] |
| Protein | Type of Interaction Confirmation | Reference |
|---|---|---|
| ACTIN | Co-IP, IP, Y2H, and IPA | [178] |
| ARP1 | IPA | [179] |
| CALMODULIN | IPA | [180] |
| CBE1 | ― | [181] |
| CCDC42 | Co-IP and IPA | [176,177] |
| CCDC181 | Y2H | [121] |
| CDC42 | IPA | [182] |
| CLIP-170 | IPA | [183] |
| CNTROB | Co-IP | [184] |
| COPS5 | Co-IP, Y2H, and IPA | [122] |
| DYNEIN | IPA | [98,179] |
| FAM46C | ― | [185] |
| FU | IP and IPA | [124] |
| GCNF | ― | [126] |
| GMAP210 | IPA | [127] |
| GSTO2 | IPA | [186] |
| HOOK1 | Co-IP, IP, MBA, Y2H, and IPA | [121,134,139,187] |
| HOOK2 | Co-IP, IP, MBA, Y2H, and IPA | [134,187,188] |
| HOOK3 | Co-IP, IP, MBA, Y2H, and IPA | [187,188] |
| IFT20 | Co-IP, IP, Y2H, and IPA | [30,122,127,132,189] |
| IFT27 | IPA | [32] |
| IFT88 | IPA | [28] |
| IFT172 | IPA | [35] |
| IQCG | IPA | [180,190] |
| KATNAL2 | Co-IP and IPA | [191] |
| KATNB1 | IPA | [192] |
| KIF3A | Co-IP, MBA, and IPA | [85] |
| KIF3B | Y2H and IPA | [85,139] |
| KIF17B | Co-IP and IPA | [84] |
| KIF5C | IPA | [81] |
| KIF27 | IP | [124] |
| KLC3 | Co-IP, IP, Y2H, and IPA | [188] |
| KRT5 | Co-IP and IPA | [28,184] |
| LIS1 | TBA and IPA | [193] |
| LRGUK1 | Co-IP, IP, and Y2H | [134,188] |
| MGCRABGAP | Co-IP | [99,135] |
| MEIG1 | Co-IP, Y2H, and IPA | [194,195] |
| MNS1 | Co-IP | [85] |
| MORN3 | Co-IP and Y2H | [136] |
| MYRIP | IPA | [98] |
| MYOSIN | IPA | [44,98] |
| NESPRIN1 | IPA | [50] |
| ODF1 | Co-IP, IP, and IPA | [124] |
| PACRG | IPA | [194,195] |
| PFN3 | IP and Y2H | [48] |
| PFN4 | ― | [48] |
| PSMC3 | Co-IP | [137] |
| RAB3A | Co-IP and IPA | [135] |
| RAB10 | Co-IP and IPA | [99] |
| RANBP17 | Co-IP and Y2H | [175] |
| RAB27 | IPA | [98] |
| RIMBP3 | Co-IP and Y2H | [139,188] |
| RNF19A | Co-IP | [137] |
| SEPT12 | Co-IP and IPA | [182,196,197,198] |
| SNAPIN | Y2H and IPA | [26] |
| SPAG6 | Y2H and IPA | [20,22,25,26] |
| SPAG16 | Co-IP and IPA | [142,194] |
| SPAG17 | IPA | [21] |
| SPATIAL | Co-IP | [84] |
| SPEF2 | Co-IP, Y2H, and IPA | [132,199] |
| SPEM1 | Co-IP and Y2H | [91,92,120,135,136] |
| STK33 | Co-IP and IP | [200] |
| STK36 | IPA | [124] |
| SUN3 | IPA | [50] |
| SUN4 | IPA | [197,201,202] |
| TMCO5A | ― | [203,204] |
| α/β-TUBULIN | IPA | [178] |
| δ-TUBULIN | Co-IP and IPA | [191] |
| ε-TUBULIN | Co-IP and IPA | [191] |
| γ-TUBULIN | Co-IP and IPA | [191] |
| UBQLN1 | Co-IP and Y2H | [174] |
| WBP2 | IPA | [205] |
| New Proteins Detected by IPA Software | ||
| APC | IPA | [148] |
| BSG | IPA | [206] |
| EGFR | IPA | [207] |
| GPX4 | IPA | [208] |
| GSK3B | IPA | [209] |
| HSP70 | IPA | [210] |
| HSP90 | IPA | [155] |
| KATNA1 | IPA | [133] |
| LAMINB1 | Co-IP | [197] |
| MAP3K11 | IPA | [211] |
| UBE2 | IPA | [165] |
| VCP | IPA | [166] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Teves, M.E.; Roldan, E.R.S.; Krapf, D.; Strauss III, J.F.; Bhagat, V.; Sapao, P. Sperm Differentiation: The Role of Trafficking of Proteins. Int. J. Mol. Sci. 2020, 21, 3702. https://doi.org/10.3390/ijms21103702
Teves ME, Roldan ERS, Krapf D, Strauss III JF, Bhagat V, Sapao P. Sperm Differentiation: The Role of Trafficking of Proteins. International Journal of Molecular Sciences. 2020; 21(10):3702. https://doi.org/10.3390/ijms21103702
Chicago/Turabian StyleTeves, Maria E., Eduardo R. S. Roldan, Diego Krapf, Jerome F. Strauss III, Virali Bhagat, and Paulene Sapao. 2020. "Sperm Differentiation: The Role of Trafficking of Proteins" International Journal of Molecular Sciences 21, no. 10: 3702. https://doi.org/10.3390/ijms21103702
APA StyleTeves, M. E., Roldan, E. R. S., Krapf, D., Strauss III, J. F., Bhagat, V., & Sapao, P. (2020). Sperm Differentiation: The Role of Trafficking of Proteins. International Journal of Molecular Sciences, 21(10), 3702. https://doi.org/10.3390/ijms21103702








