Sushi Repeat Containing Protein X-linked 2 Is a Downstream Signal of LEM Domain Containing 1 and Acts as a Tumor-Promoting Factor in Oral Squamous Cell Carcinoma
Abstract
1. Introduction
2. Results
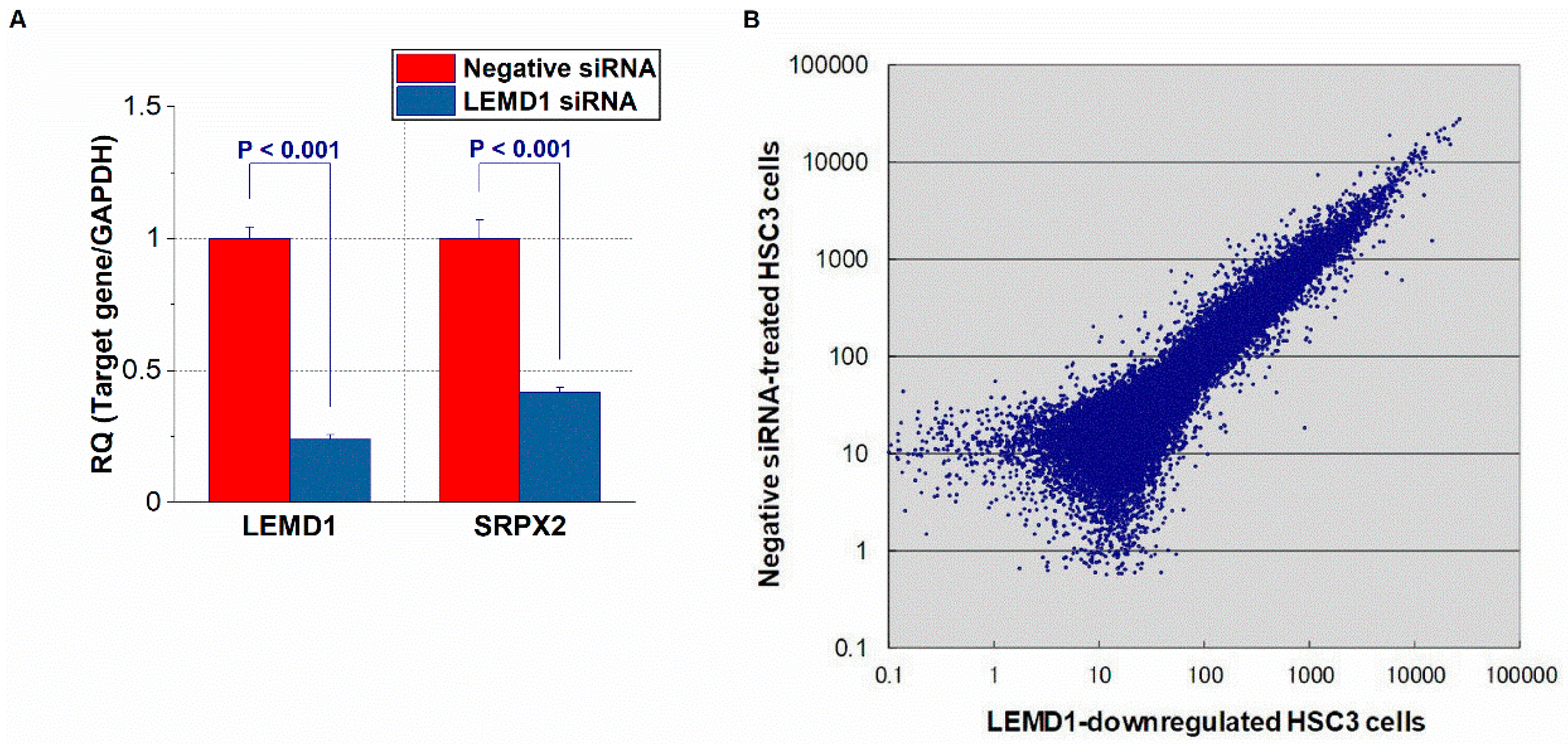
2.1. Identification of LEMD1-Related Upregulated or Downregulated Genes in OSCC Cells via a Microarray Analysis
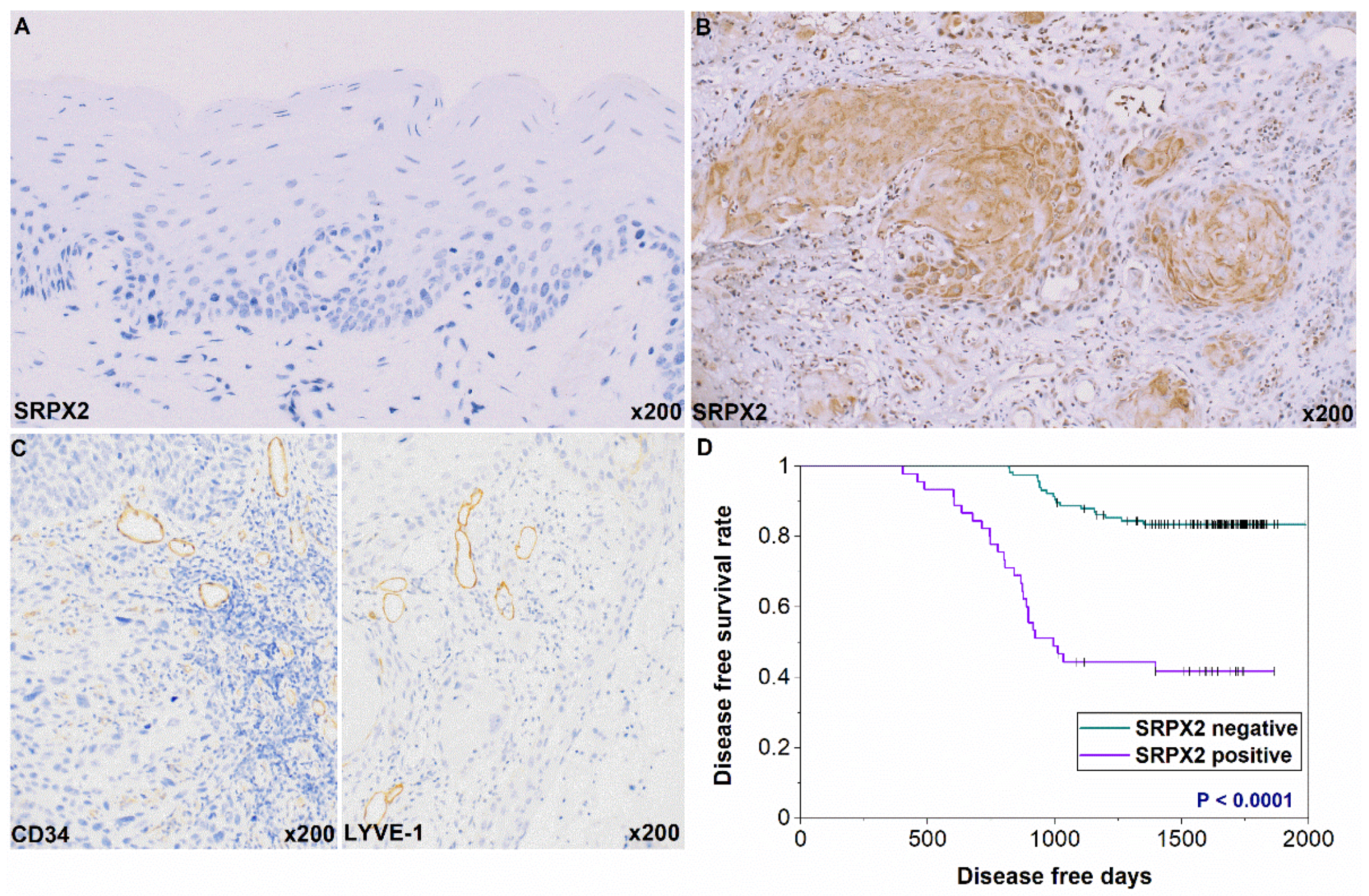
2.2. Expression and Prognosis of SRPX2 in OSCC Specimens by Immunohistochemistry
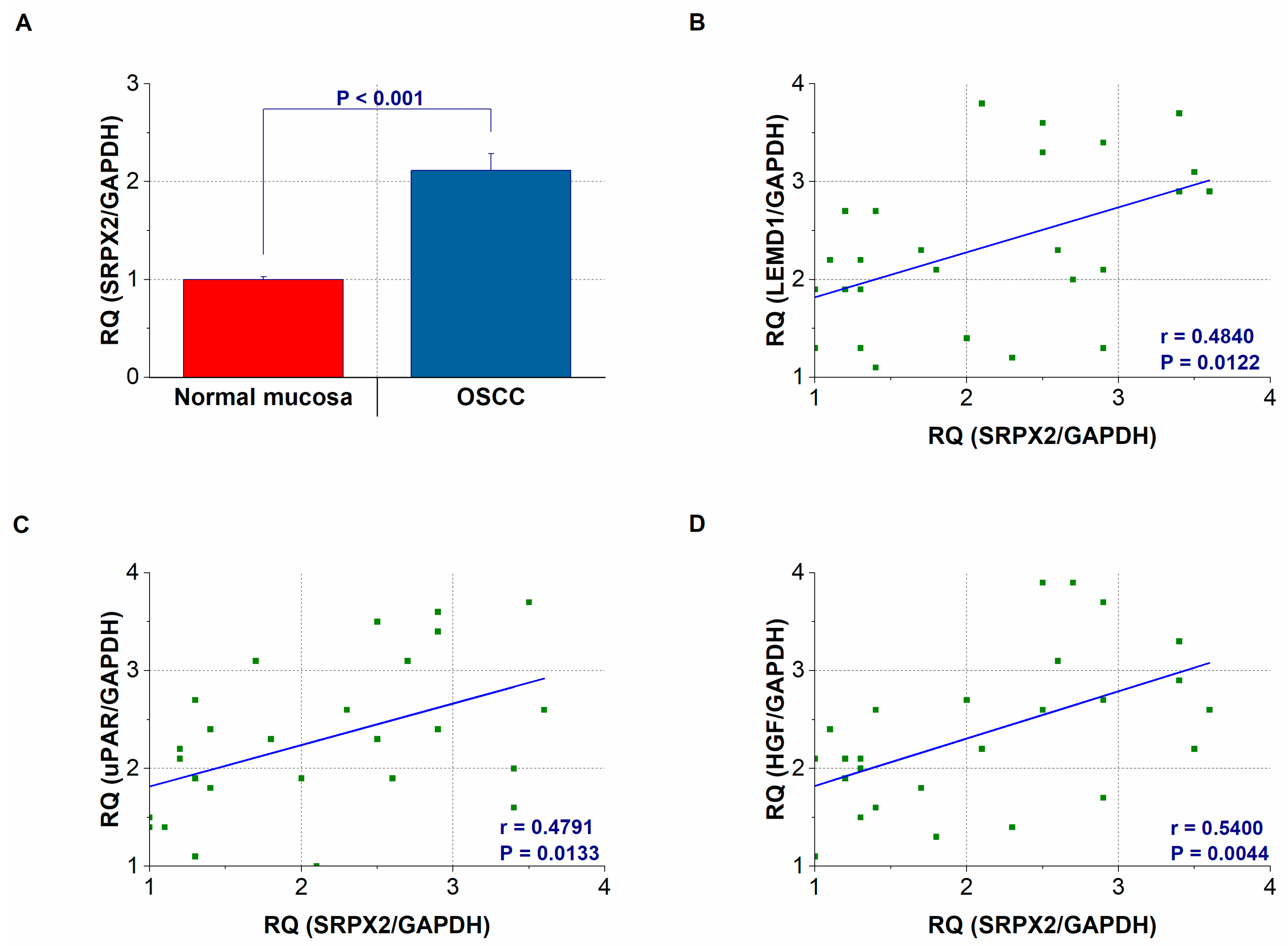
2.3. Gene Expression and Concentration of SRPX2 in OSCC Specimens
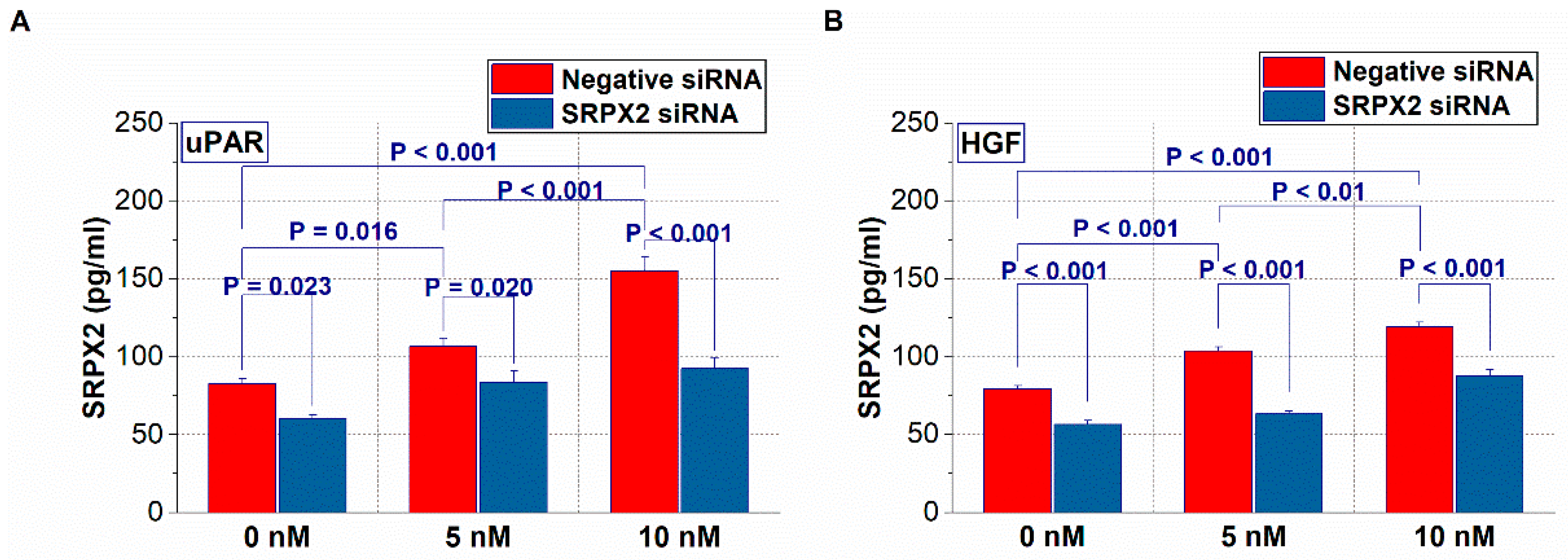
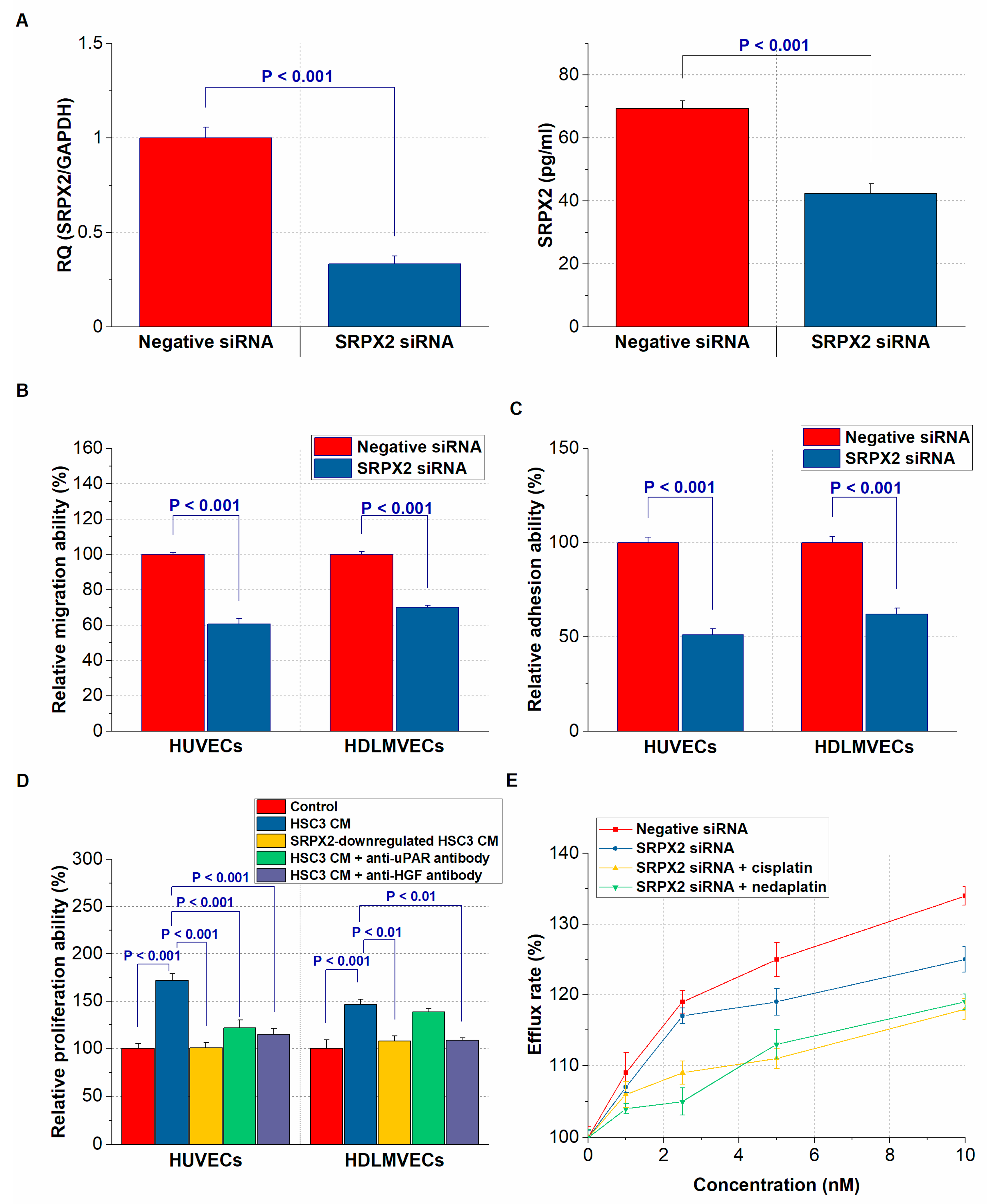
2.4. Expression of SRPX2 and the Effects of uPAR and HGF on the Secretion of SRPX2 in OSCC Cells
2.5. Effect of SRPX2 on Angiogenesis and Lymphangiogenesis of OSCC Cells
2.6. Effect of SRPX2 on the Drug Resistance of OSCC Cells
3. Discussion
4. Materials and Methods
4.1. Cell Culture
4.2. Microarray Analysis
4.3. Tumor Specimens
4.4. Immunohistochemistry
4.5. RNA Extraction and Quantitative Reverse-Transcription Polymerase Chain Reaction
4.6. ELISA for SRPX2, uPAR, and HGF
4.7. Small Interfering RNA
4.8. Treatment with Recombinant Protein and SRPX2-Conditioned Medium
4.9. Cell Proliferation of Endothelial Cells and Assays of Interactions between OSCCs and Endothelial Cells
4.10. Anticancer Resistance Assays
4.11. Statistical Analysis
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| OSCC | oral squamous cell carcinoma |
| LEMD 1 | LEM domain containing 1 |
| SRPX2 | sushi repeat containing protein X-linked 2 |
| uPAR | urokinase plasminogen activator receptor |
| HGF | hepatocyte growth factor |
| MVD | microvessel density |
| LVD | lymphovessel density |
References
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [PubMed]
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer statistics, 2019. CA A Cancer J. Clin. 2019, 69, 7–34. [Google Scholar]
- Shimomura, H.; Sasahira, T.; Nakashima, C.; Kurihara-Shimomura, M.; Kirita, T. Non-SMC condensin I complex subunit H (NCAPH) is associated with lymphangiogenesis and drug resistance in oral squamous cell carcinoma. J. Clin. Med. 2019, 9, 72. [Google Scholar] [CrossRef] [PubMed]
- Sasahira, T.; Bosserhoff, A.K.; Kirita, T. The importance of melanoma inhibitory activity gene family in the tumor progression of oral cancer. Pathol. Int. 2018, 68, 278–286. [Google Scholar] [CrossRef] [PubMed]
- Yuki, D.; Lin, Y.M.; Fujii, Y.; Nakamura, Y.; Furukawa, Y. Isolation of LEM domain-containing 1, a novel testis-specific gene expressed in colorectal cancers. Oncol. Rep. 2004, 12, 275–280. [Google Scholar] [CrossRef] [PubMed]
- Ghafouri-Fard, S.; Ousati Ashtiani, Z.; Sabah Golian, B.; Hasheminasab, S.M.; Modarressi, M.H. Expression of two testis-specific genes, SPATA19 and LEMD1, in prostate cancer. Arch. Med. Res. 2010, 41, 195–200. [Google Scholar] [CrossRef] [PubMed]
- Matsuyama, H.; Suzuki, H.I.; Nishimori, H.; Noguchi, M.; Yao, T.; Komatsu, N.; Mano, H.; Sugimoto, K.; Miyazono, K. miR-135b mediates NPM-ALK-driven oncogenicity and renders IL-17-producing immunophenotype to anaplastic large cell lymphoma. Blood 2011, 118, 6881–6892. [Google Scholar] [CrossRef] [PubMed]
- Takeda, R.; Hirohashi, Y.; Shen, M.; Wang, L.; Ogawa, T.; Murai, A.; Yamamoto, E.; Kubo, T.; Nakatsugawa, M.; Kanaseki, T.; et al. Identification and functional analysis of variants of a cancer/testis antigen LEMD1 in colorectal cancer stem-like cells. Biochem. Biophys. Res. Commun. 2017, 485, 651–657. [Google Scholar] [CrossRef]
- Li, Q.; Ge, Y.; Chen, X.; Wang, L.; Xia, Y.; Xu, Z.; Li, Z.; Wang, W.; Yang, L.; Zhang, D.; et al. LEM domain containing 1 promotes proliferation via activating the PI3K/Akt signaling pathway in gastric cancer. J. Cell Biochem. 2019, 120, 15190–15201. [Google Scholar] [CrossRef]
- Sasahira, T.; Kurihara, M.; Nakashima, C.; Kirita, T.; Kuniyasu, H. LEM domain containing 1 promotes oral squamous cell carcinoma invasion and endothelial transmigration. Br. J. Cancer 2016, 115, 52–58. [Google Scholar] [CrossRef]
- Tanaka, K.; Arao, T.; Tamura, D.; Aomatsu, K.; Furuta, K.; Matsumoto, K.; Kaneda, H.; Kudo, K.; Fujita, Y.; Kimura, H.; et al. SRPX2 is a novel chondroitin sulfate proteoglycan that is overexpressed in gastrointestinal cancer. PLoS ONE 2012, 7, e27922. [Google Scholar] [CrossRef] [PubMed]
- Sia, G.M.; Clem, R.L.; Huganir, R.L. The human language-associated gene SRPX2 regulates synapse formation and vocalization in mice. Science 2013, 342, 987–991. [Google Scholar] [CrossRef] [PubMed]
- Yamada, T.; Oshima, T.; Yoshihara, K.; Sato, T.; Nozaki, A.; Shiozawa, M.; Ota, M.; Yoshikawa, T.; Akaike, M.; Numata, K.; et al. Impact of overexpression of sushi repeat-containing protein X-linked 2 gene on outcomes of gastric cancer. J. Surg. Oncol. 2014, 109, 836–840. [Google Scholar] [CrossRef] [PubMed]
- Oster, B.; Linnet, L.; Christensen, L.L.; Thorsen, K.; Ongen, H.; Dermitzakis, E.T.; Sandoval, J.; Moran, S.; Esteller, M.; Hansen, T.F.; et al. Non-CpG island promoter hypomethylation and miR-149 regulate the expression of SRPX2 in colorectal cancer. Int. J. Cancer 2013, 132, 2303–2315. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Xu, J.; Zhang, R. SRPX2 regulates colon cancer cell metabolism by miR-192/215 via PI3K-Akt. Am. J. Transl. Res. 2018, 10, 483–490. [Google Scholar]
- Gao, Z.; Zhang, J.; Bi, M.; Han, X.; Han, Z.; Wang, H.; Ou, Y. SRPX2 promotes cell migration and invasion via FAK dependent pathway in pancreatic cancer. Int. J. Clin. Exp. Pathol. 2015, 8, 4791–4798. [Google Scholar]
- Li, H.; Zhang, S.R.; Xu, H.X.; Wang, W.Q.; Li, S.; Li, T.J.; Ni, Q.X.; Yu, X.J.; Liu, L.; Wu, C.T. SRPX2 and RAB31 are effective prognostic biomarkers in pancreatic cancer. J. Cancer 2019, 10, 2670–2678. [Google Scholar] [CrossRef]
- Tang, H.; Zhao, J.; Zhang, L.; Zhao, J.; Zhuang, Y.; Liang, P. SRPX2 enhances the epithelial-mesenchymal transition and temozolomide resistance in glioblastoma cells. Cell Mol. Neurobiol. 2016, 36, 1067–1076. [Google Scholar] [CrossRef]
- Wu, Z.; Wang, C.; Chen, Y.; Sun, Z.; Yan, W. SRPX2 promotes cell proliferation and invasion in osteosarcoma through regulating hippo Ssgnaling pathway. Onco Targets Ther. 2020, 13, 1737–1749. [Google Scholar] [CrossRef]
- Lin, X.; Chang, W.; Wang, Y.; Tian, M.; Yu, Z. SRPX2, an independent prognostic marker, promotes cell migration and invasion in hepatocellular carcinoma. Biomed. Pharmacother. 2017, 93, 398–405. [Google Scholar] [CrossRef]
- Zhang, M.; Li, X.; Fan, Z.; Zhao, J.; Liu, S.; Zhang, M.; Li, H.; Goscinski, M.A.; Fan, H.; Suo, Z. High SRPX2 protein expression predicts unfavorable clinical outcome in patients with prostate cancer. Onco Targets Ther. 2018, 11, 3149–3157. [Google Scholar] [CrossRef] [PubMed]
- He, F.; Wang, H.; Li, Y.; Liu, W.; Gao, X.; Chen, D.; Wang, Q.; Shi, G. SRPX2 knockdown inhibits cell proliferation and metastasis and promotes chemosensitivity in esophageal squamous cell carcinoma. Biomed. Pharmacother. 2019, 109, 671–678. [Google Scholar] [CrossRef] [PubMed]
- Miljkovic-Licina, M.; Hammel, P.; Garrido-Urbani, S.; Bradfield, P.F.; Szepetowski, P.; Imhof, B.A. Sushi repeat protein X-linked 2, a novel mediator of angiogenesis. Faseb J. 2009, 23, 4105–4116. [Google Scholar] [CrossRef] [PubMed]
- Liu, K.; Fan, J.; Wu, J. Sushi repeat-containing protein X-linked 2 promotes angiogenesis through the urokinase-type plasminogen activator receptor dependent integrin alphavbeta3/focal adhesion kinase pathways. Drug Discov. Ther. 2017, 11, 212–217. [Google Scholar] [CrossRef] [PubMed]
- Zhang, N.; Xie, F.; Gao, W.; Yu, S.; Qiu, L.; Lin, W.; Sun, Y.; Jia, T. Expression of hepatocyte growth factor and c-Met in non-small-cell lung cancer and association with lymphangiogenesis. Mol. Med. Rep. 2015, 11, 2797–2804. [Google Scholar] [CrossRef] [PubMed]
- Gjerstorff, M.F.; Andersen, M.H.; Ditzel, H.J. Oncogenic cancer/testis antigens: Prime candidates for immunotherapy. Oncotarget 2015, 6, 15772–15787. [Google Scholar] [CrossRef]
- Sasahira, T.; Kirita, T. Hallmarks of cancer-related newly prognostic factors of oral squamous cell carcinoma. Int. J. Mol. Sci. 2018, 19, 2413. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef]
- Allred, D.C.; Harvey, J.M.; Berardo, M.; Clark, G.M. Prognostic and predictive factors in breast cancer by immunohistochemical analysis. Mod. Pathol. 1998, 11, 155–168. [Google Scholar]
- Wang, N.; Lu, K.; Qu, H.; Wang, H.; Chen, Y.; Shan, T.; Ge, X.; Wei, Y.; Zhou, P.; Xia, J. CircRBM33 regulates IL-6 to promote gastric cancer progression through targeting miR-149. Biomed. Pharmacother. 2020, 125, 109876. [Google Scholar] [CrossRef]
- Marton, I.J.; Horvath, J.; Labiscsak, P.; Markus, B.; Dezso, B.; Szabo, A.; Tar, I.; Piffko, J.; Jakus, P.; Barabas, J.; et al. Salivary IL-6 mRNA is a robust biomarker in oral squamous cell carcinoma. J. Clin. Med. 2019, 8, 1958. [Google Scholar] [CrossRef] [PubMed]
- Sasahira, T.; Kirita, T.; Kurihara, M.; Yamamoto, K.; Bhawal, U.K.; Bosserhoff, A.K.; Kuniyasu, H. MIA-dependent angiogenesis and lymphangiogenesis are closely associated with progression, nodal metastasis and poor prognosis in tongue squamous cell carcinoma. Eur. J. Cancer 2010, 46, 2285–2294. [Google Scholar] [CrossRef] [PubMed]
- Jain, R.K.; Munn, L.L.; Fukumura, D. Dissecting tumour pathophysiology using intravital microscopy. Nat. Rev. Cancer 2002, 2, 266–276. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Kang, G.; Wang, T.; Huang, H. Tumor angiogenesis and anti-angiogenic gene therapy for cancer. Oncol. Lett. 2018, 16, 687–702. [Google Scholar] [CrossRef] [PubMed]
- O’Sullivan, B. Head and neck tumours. In UICC TNM Classification of Malignant Tumours, 8th ed.; Brierley, J.D., Gospodarowicz, M.K., Wittekind, C., Eds.; Wiley: Chichester, UK, 2017; pp. 17–54. [Google Scholar]
- Sloan, P.; Gale, N.; Hunter, K.; Lingen, M.; Nylander, K.; Reibel, J.; Salo, T.; Zain, R. Malignant surface epithelial tumors. In WHO Classification of Head and Neck Tumors, 4th ed.; El-Naggar, A., Chan, J.K.C., Grandis, J.R., Takata, T., Slootweg, P.J., Eds.; IARC Press: Lyom, Sweden, 2017; pp. 109–111. [Google Scholar]
- Sasahira, T.; Ueda, N.; Yamamoto, K.; Kurihara, M.; Matsushima, S.; Bhawal, U.K.; Kirita, T.; Kuniyasu, H. Prox1 and FOXC2 act as regulators of lymphangiogenesis and angiogenesis in oral squamous cell carcinoma. PLoS ONE 2014, 9, e92534. [Google Scholar] [CrossRef]
| Downregulated | Upregulated | ||
|---|---|---|---|
| Gene | Fold (LEMD1 Knockdown/Control) | Gene | Fold (LEMD1 Knockdown/Control) |
| CEACAM6 | 0.054 | MRPL2 | 12.481 |
| ESM1 | 0.066 | GSX2 | 10.285 |
| PHLDA1 | 0.101 | KRT5 | 9.615 |
| BIRC3 | 0.113 | BIRC6 | 8.474 |
| CALB2 | 0.115 | GSTA4 | 8.446 |
| DHRS9 | 0.121 | FOS | 7.613 |
| KPNA7 | 0.130 | AKR1B10 | 7.591 |
| SOX9 | 0.135 | BOLA3 | 7.575 |
| SERPINE2 | 0.140 | AHSG | 7.131 |
| SRPX2 | 0.154 | DST | 6.435 |
| SRPX2 Expression | |||
|---|---|---|---|
| Parameters | − (%) | + (%) | p Value * |
| Gender | |||
| Male | 61 (68.5) | 28 (31.5) | |
| Female | 55 (76.4) | 17 (23.6) | 0.2938 |
| Age | |||
| <65 | 54 (75) | 18 (25) | |
| >65 | 62 (69.7) | 27 (30.3) | 0.4844 |
| Site | |||
| Tongue | 60 (66.7) | 30 (33.3) | |
| Gingiva | 42 (80.8) | 10 (19.2) | |
| Buccal mucosa | 7 (70) | 3 (30) | |
| Other | 7 (77.8) | 2 (22.2) | 0.3305 |
| Histology | |||
| Well | 57 (75) | 19 (25) | |
| Moderately, Poorly | 59 (69.4) | 26 (30.6) | 0.4839 |
| T classification | |||
| T1-T3 | 81 (74.3) | 28 (25.7) | |
| T4 | 35 (67.3) | 17 (32.7) | 0.3554 |
| Clinical stage | |||
| I-II | 79 (75.2) | 26 (24.8) | |
| IV | 37 (66.1) | 19 (33.9) | 0.2688 |
| Nodal metastasis | |||
| Negative | 91 (77.1) | 27 (22.9) | |
| Positive | 25 (58.1) | 18 (41.9) | 0.0280 |
| MVD | 25.128 ± 14.741 | 33.109 ± 23.841 | 0.0111 |
| LVD | 20.245 ± 13.894 | 27.136 ± 19.583 | 0.0129 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sasahira, T.; Kurihara-Shimomura, M.; Nishiguchi, Y.; Shimomura, H.; Kirita, T. Sushi Repeat Containing Protein X-linked 2 Is a Downstream Signal of LEM Domain Containing 1 and Acts as a Tumor-Promoting Factor in Oral Squamous Cell Carcinoma. Int. J. Mol. Sci. 2020, 21, 3655. https://doi.org/10.3390/ijms21103655
Sasahira T, Kurihara-Shimomura M, Nishiguchi Y, Shimomura H, Kirita T. Sushi Repeat Containing Protein X-linked 2 Is a Downstream Signal of LEM Domain Containing 1 and Acts as a Tumor-Promoting Factor in Oral Squamous Cell Carcinoma. International Journal of Molecular Sciences. 2020; 21(10):3655. https://doi.org/10.3390/ijms21103655
Chicago/Turabian StyleSasahira, Tomonori, Miyako Kurihara-Shimomura, Yukiko Nishiguchi, Hiroyuki Shimomura, and Tadaaki Kirita. 2020. "Sushi Repeat Containing Protein X-linked 2 Is a Downstream Signal of LEM Domain Containing 1 and Acts as a Tumor-Promoting Factor in Oral Squamous Cell Carcinoma" International Journal of Molecular Sciences 21, no. 10: 3655. https://doi.org/10.3390/ijms21103655
APA StyleSasahira, T., Kurihara-Shimomura, M., Nishiguchi, Y., Shimomura, H., & Kirita, T. (2020). Sushi Repeat Containing Protein X-linked 2 Is a Downstream Signal of LEM Domain Containing 1 and Acts as a Tumor-Promoting Factor in Oral Squamous Cell Carcinoma. International Journal of Molecular Sciences, 21(10), 3655. https://doi.org/10.3390/ijms21103655





