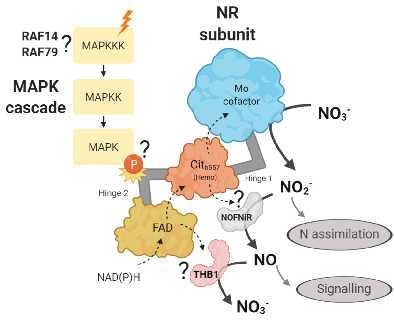
Identification of the MAPK Cascade and its Relationship with Nitrogen Metabolism in the Green Alga Chlamydomonas reinhardtii
Abstract
1. Introduction
2. Results
2.1. Chlamydomonas MAPK Family
2.2. Chlamydomonas MAPK Gene Expression
2.3. Study of a Chlamydomonas Mapk8 Mutant
2.4. Chlamydomonas MAPKK Family
2.5. Chlamydomonas MAPKK Gene Expression
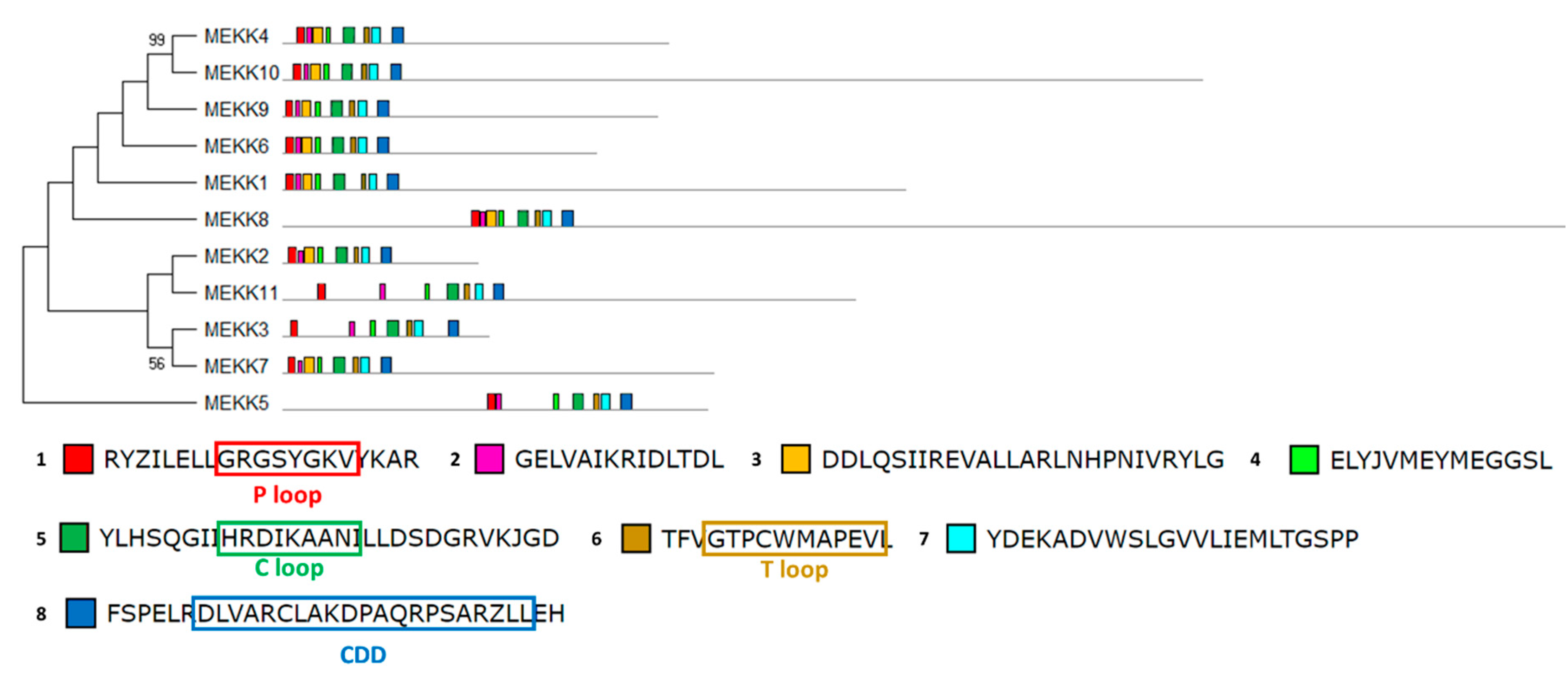
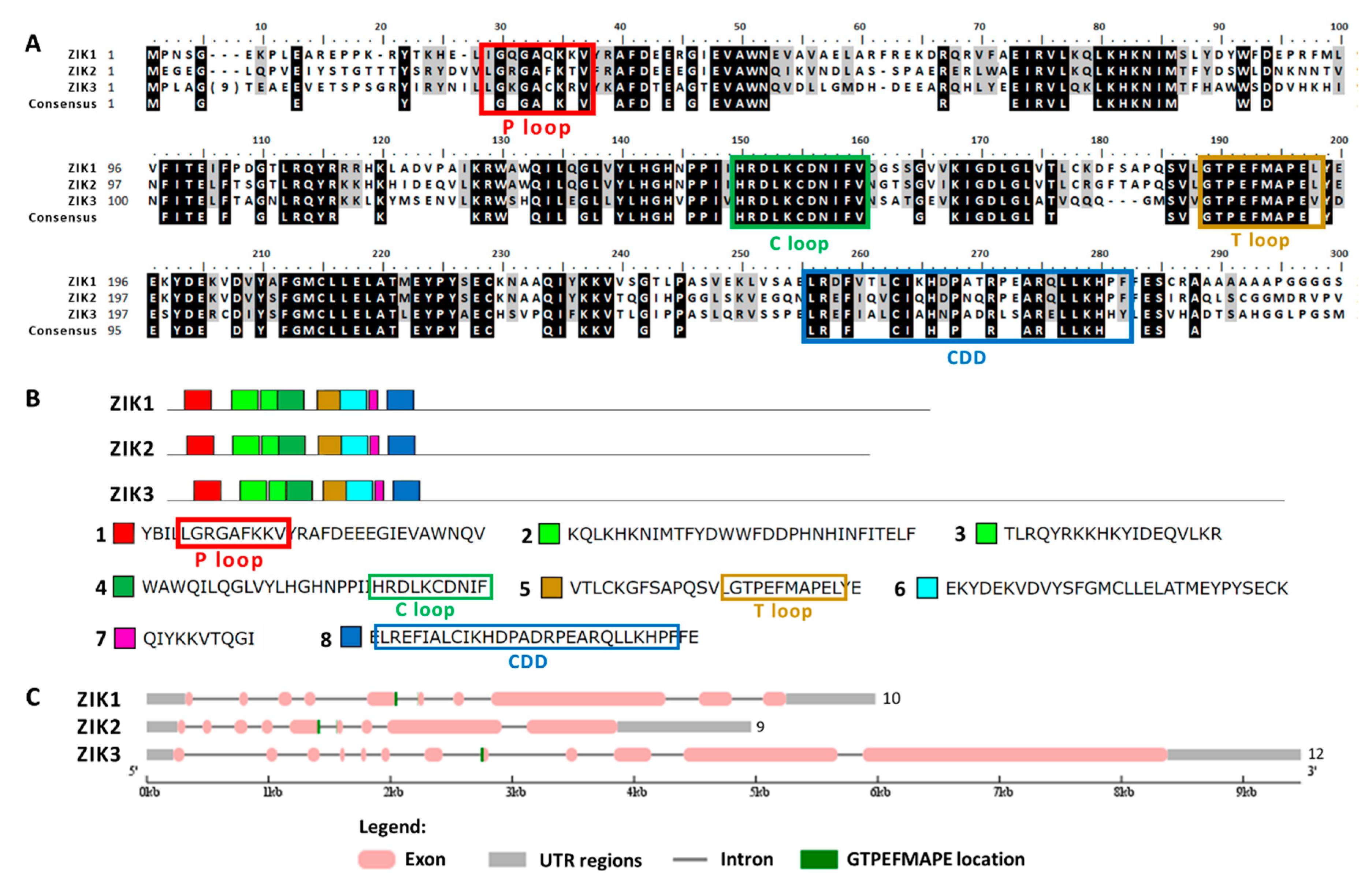
2.6. Chlamydomonas MAPKKK Family
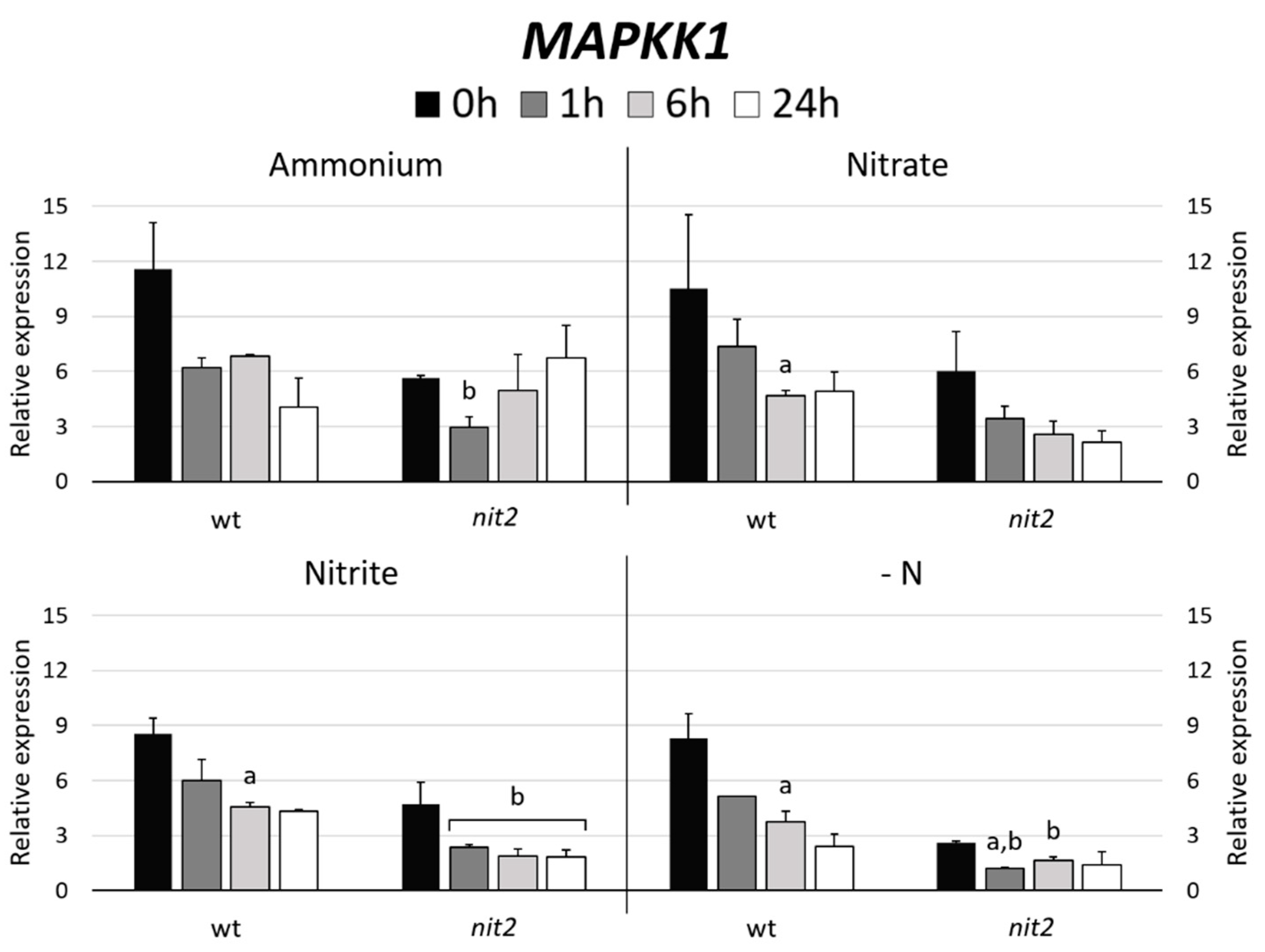
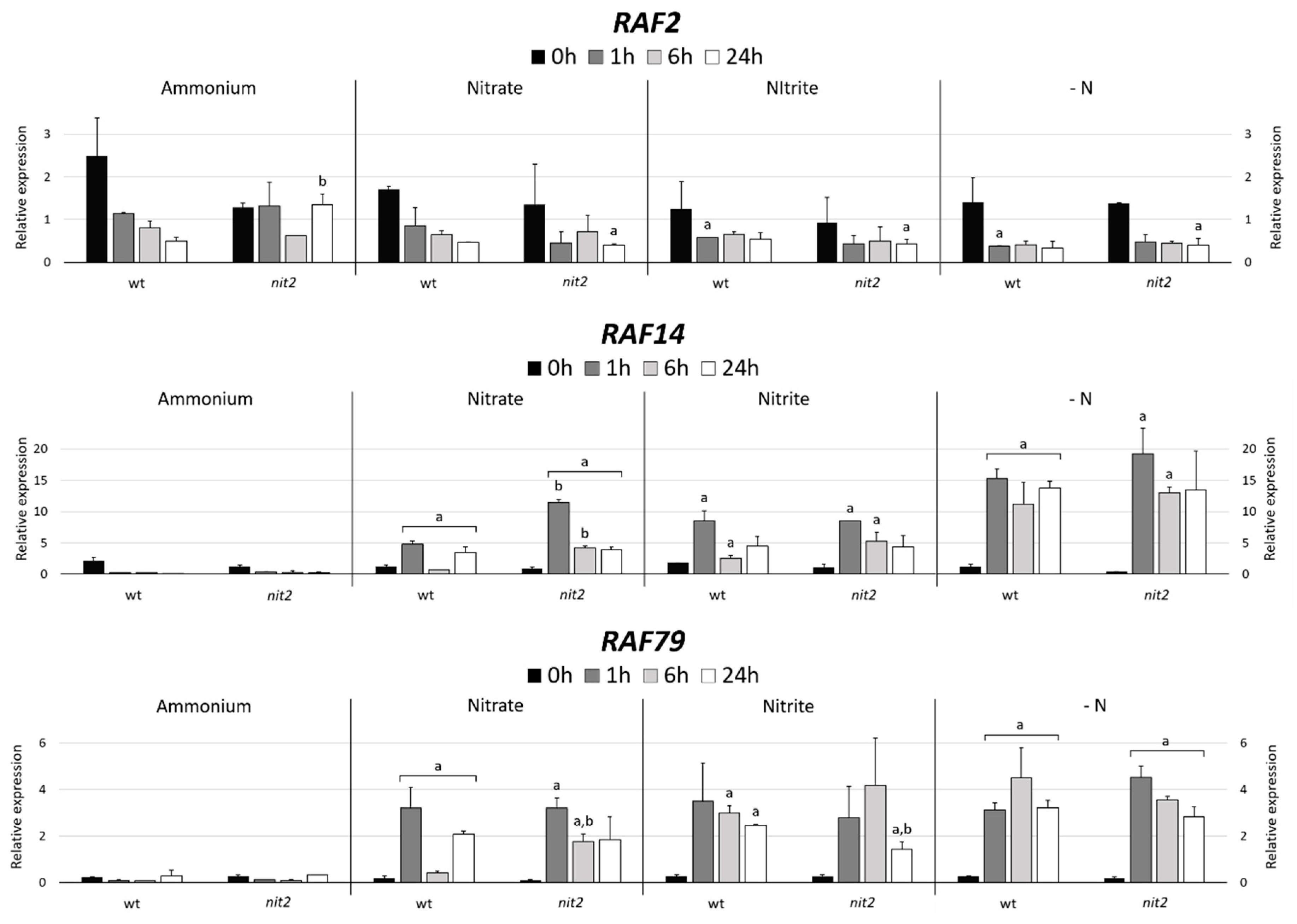
2.7. Gene Expression of MAPKKK Candidates to be Involved in the N Assimilation
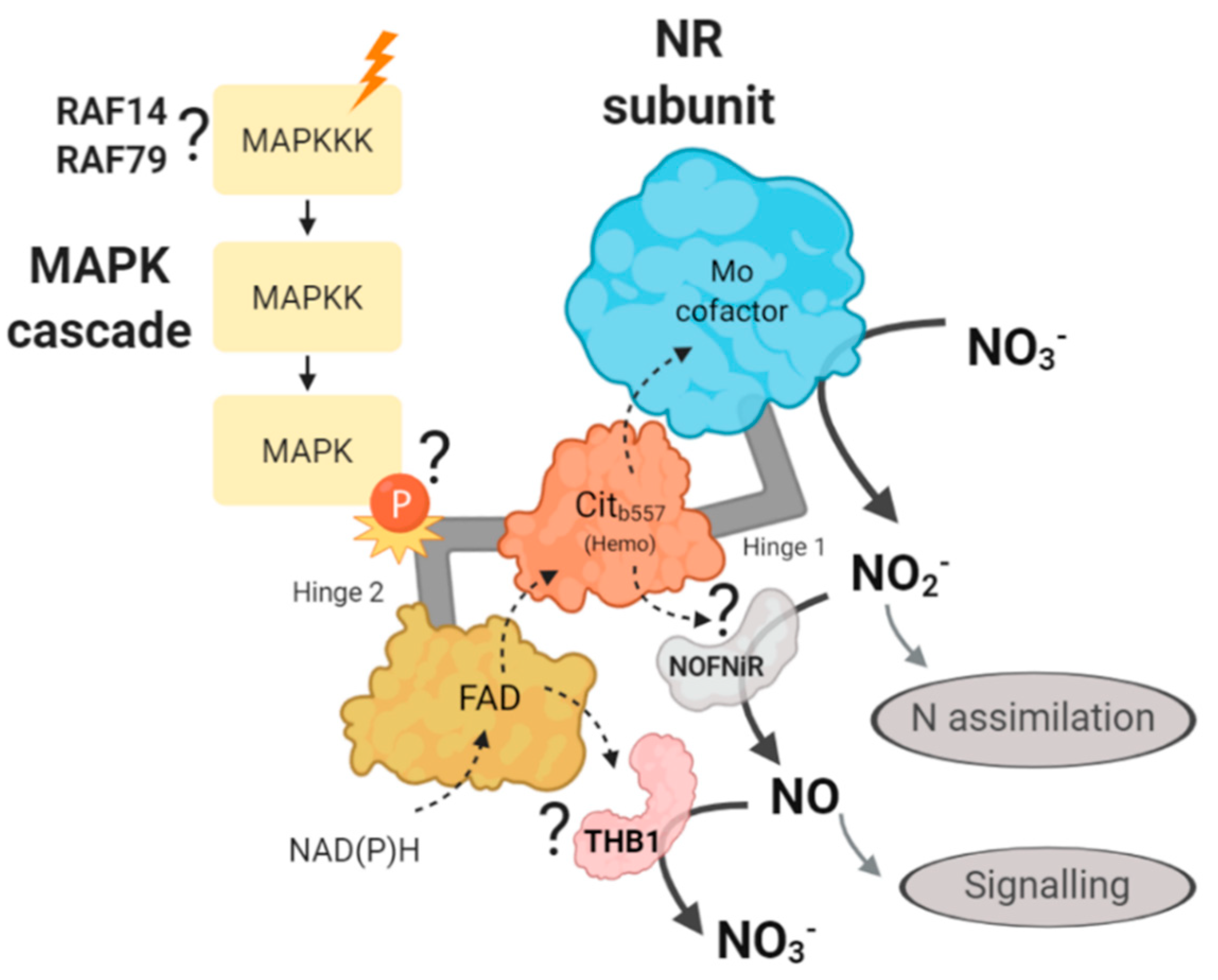
3. Discussion
4. Materials and Methods
4.1. Chlamydomonas Strains and Growth Conditions
4.2. Chlamydomonas Genetic Cross
4.3. Nitrate Reductase Activity
4.4. Quantification of Chlamydomonas Growth
4.5. Quantification of Gene Expression by RT-qPCR
4.6. MAPK Cascade Identification
4.7. Phylogenetic and Domain Analysis
4.8. Additional Bioinformatics Tools
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Harris, E.H. Chlamydomonas as a model organism. Annu. Rev. Plant Biol. 2001, 52, 363–406. [Google Scholar] [CrossRef]
- Merchant, S.S.; Prochnik, S.E.; Vallon, O.; Harris, E.H.; Karpowicz, S.J.; Witman, G.B.; Terry, A.; Salamov, A.; Fritz-Laylin, L.K.; Maréchal-Drouard, L.; et al. The Chlamydomonas genome reveals the evolution of key animal and plant functions. Science 2007, 318, 245–250. [Google Scholar] [CrossRef]
- Li, X.; Patena, W.; Fauser, F.; Jinkerson, R.E.; Saroussi, S.; Meyer, M.T.; Ivanova, N.; Robertson, J.M.; Yue, R.; Zhang, R.; et al. A genome-wide algal mutant library and functional screen identifies genes required for eukaryotic photosynthesis. Nat. Genet. 2019, 51, 627–635. [Google Scholar] [CrossRef]
- Satoh, K. Protein-pigments and the photosystem II reaction center: A glimpse into the history of research and reminiscences. Photosynth. Res. 2008, 98, 33–42. [Google Scholar] [CrossRef] [PubMed]
- Sasso, S.; Stibor, H.; Mittag, M.; Grossman, A.R. From molecular manipulation of domesticated. eLife 2018, 7, e39233. [Google Scholar] [CrossRef] [PubMed]
- Aksoy, M.; Pootakham, W.; Pollock, S.V.; Moseley, J.L.; González-Ballester, D.; Grossman, A.R. Tiered regulation of sulfur deprivation responses in Chlamydomonas reinhardtii and identification of an associated regulatory factor. Plant Physiol. 2013, 162, 195–211. [Google Scholar] [CrossRef] [PubMed]
- Melis, A. Photosynthetic H2 metabolism in Chlamydomonas reinhardtii (unicellular green algae). Planta 2007, 226, 1075–1086. [Google Scholar] [CrossRef]
- Fernandez, E.; Galvan, A. Inorganic nitrogen assimilation in Chlamydomonas. J. Exp. Bot. 2007, 58, 2279–2287. [Google Scholar] [CrossRef]
- Stein, L.Y.; Klotz, M.G. The nitrogen cycle. Curr. Biol. 2016, 26, R94–R98. [Google Scholar] [CrossRef]
- Hachiya, T.; Sakakibara, H. Interactions between nitrate and ammonium in their uptake, allocation, assimilation, and signaling in plants. J. Exp. Bot. 2017, 68, 2501–2512. [Google Scholar] [CrossRef]
- Hirsch, R.E.; Sussman, M.R. Improving nutrient capture from soil by the genetic manipulation of crop plants. Trends Biotechnol. 1999, 17, 356–361. [Google Scholar] [CrossRef]
- Cherfas, J. The Fringe of the Ocean—Under Siege from Land: The ecology of the ocean margins, crucial to human life, is being disrupted by our activities and perhaps by global change. Science 1990, 248, 163–165. [Google Scholar] [CrossRef]
- Kant, S. Understanding nitrate uptake, signaling and remobilisation for improving plant nitrogen use efficiency. Semin. Cell Dev. Biol. 2018, 74, 89–96. [Google Scholar] [CrossRef] [PubMed]
- Krapp, A. Plant nitrogen assimilation and its regulation: A complex puzzle with missing pieces. Curr. Opin. Plant Biol. 2015, 25, 115–122. [Google Scholar] [CrossRef] [PubMed]
- Sanz-Luque, E.; Chamizo-Ampudia, A.; Llamas, A.; Galvan, A.; Fernandez, E. Understanding nitrate assimilation and its regulation in microalgae. Front. Plant Sci. 2015, 6, 899. [Google Scholar] [CrossRef]
- Wang, Y.-Y.; Hsu, P.-K.; Tsay, Y.-F. Uptake, allocation and signaling of nitrate. Trends Plant Sci. 2012, 17, 458–467. [Google Scholar] [CrossRef]
- Rexach, J.; Fernández, E.; Galván, A. The Chlamydomonas reinhardtii Nar1 gene encodes a chloroplast membrane protein involved in nitrite transport. Plant Cell 2000, 12, 1441–1453. [Google Scholar] [CrossRef]
- Camargo, A.; Llamas, A.; Schnell, R.A.; Higuera, J.J.; Gonzalez-Ballester, D.; Lefebvre, P.A.; Fernandez, E.; Galvan, A. Nitrate Signaling by the Regulatory Gene NIT2 in Chlamydomonas. Plant Cell 2007, 19, 3491–3503. [Google Scholar] [CrossRef]
- Campbell, W.H.; Kinghorn, K.R. Functional domains of assimilatory nitrate reductases and nitrite reductases. Trends Biochem. Sci. 1990, 15, 315–319. [Google Scholar] [CrossRef]
- Domingos, P.; Prado, A.M.; Wong, A.; Gehring, C.; Feijo, J.A. Nitric Oxide: A Multitasked Signaling Gas in Plants. Mol. Plant 2015, 8, 506–520. [Google Scholar] [CrossRef]
- De Montaigu, A.; Sanz-Luque, E.; Galván, A.; Fernández, E. A soluble guanylate cyclase mediates negative signaling by ammonium on expression of nitrate reductase in Chlamydomonas. Plant Cell 2010, 22, 1532–1548. [Google Scholar] [CrossRef] [PubMed]
- De Mia, M.; Lemaire, S.D.; Choquet, Y.; Wollman, F.A. Nitric Oxide Remodels the Photosynthetic Apparatus upon S-Starvation in. Plant Physiol. 2019, 179, 718–731. [Google Scholar] [CrossRef] [PubMed]
- Wei, L.; Derrien, B.; Gautier, A.; Houille-Vernes, L.; Boulouis, A.; Saint-Marcoux, D.; Malnoe, A.; Rappaport, F.; de Vitry, C.; Vallon, O.; et al. Nitric Oxide-Triggered Remodeling of Chloroplast Bioenergetics and Thylakoid Proteins upon Nitrogen Starvation in Chlamydomonas reinhardtii. Plant Cell 2014, 26, 353–372. [Google Scholar] [CrossRef] [PubMed]
- Yordanova, Z.P.; Woltering, E.J.; Kapchina-Toteva, V.M.; Iakimova, E.T. Mastoparan-induced programmed cell death in the unicellular alga Chlamydomonas reinhardtii. Ann. Bot. 2013, 111, 191–205. [Google Scholar] [CrossRef] [PubMed]
- Hemschemeier, A.; Duner, M.; Casero, D.; Merchant, S.S.; Winkler, M.; Happe, T. Hypoxic survival requires a 2-on-2 hemoglobin in a process involving nitric oxide. Proc. Natl. Acad. Sci. USA 2013, 110, 10854–10859. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Tian, D.; Kong, X.; Chen, Q.; Abd Allah, E.F.; Hu, X.; Jia, A. The role of nitric oxide signalling in response to salt stress in Chlamydomonas reinhardtii. Planta 2016, 244, 651–669. [Google Scholar] [CrossRef]
- Dean, J.V.; Harper, J.E. The Conversion of Nitrite to Nitrogen Oxide(s) by the Constitutive NAD(P)H-Nitrate Reductase Enzyme from Soybean. Plant Physiol. 1988, 88, 389–395. [Google Scholar] [CrossRef]
- Sakihama, Y.; Nakamura, S.; Yamasaki, H. Nitric oxide production mediated by nitrate reductase in the green alga Chlamydomonas reinhardtii: An alternative NO production pathway in photosynthetic organisms. Plant Cell Physiol. 2002, 43, 290–297. [Google Scholar] [CrossRef]
- Gupta, K.J.; Igamberdiev, A.U.; Manjunatha, G.; Segu, S.; Moran, J.F.; Neelawarne, B.; Bauwe, H.; Kaiser, W.M. The emerging roles of nitric oxide (NO) in plant mitochondria. Plant Sci. 2011, 181, 520–526. [Google Scholar] [CrossRef]
- Rockel, P.; Strube, F.; Rockel, A.; Wildt, J.; Kaiser, W.M. Regulation of nitric oxide (NO) production by plant nitrate reductase in vivo and in vitro. J. Exp. Bot. 2002, 53, 103–110. [Google Scholar] [CrossRef]
- Chamizo-Ampudia, A.; Sanz-Luque, E.; Llamas, Á.; Ocaña-Calahorro, F.; Mariscal, V.; Carreras, A.; Barroso, J.B.; Galván, A.; Fernández, E. A dual system formed by the ARC and NR molybdoenzymes mediates nitrite-dependent NO production in Chlamydomonas. Plant Cell Environ. 2016, 39, 2097–2107. [Google Scholar] [CrossRef]
- Chamizo-Ampudia, A.; Sanz-Luque, E.; Llamas, A.; Galvan, A.; Fernandez, E. Nitrate Reductase Regulates Plant Nitric Oxide Homeostasis. Trends Plant Sci. 2017, 22, 163–174. [Google Scholar] [CrossRef]
- Rodriguez, M.C.; Petersen, M.; Mundy, J. Mitogen-activated protein kinase signaling in plants. Annu. Rev. Plant Biol. 2010, 61, 621–649. [Google Scholar] [CrossRef]
- Komis, G.; Šamajová, O.; Ovečka, M.; Šamaj, J. Cell and Developmental Biology of Plant Mitogen-Activated Protein Kinases. Annu. Rev. Plant Biol. 2018, 69, 237–265. [Google Scholar] [CrossRef]
- Neupane, S.; Schweitzer, S.E.; Neupane, A.; Andersen, E.J.; Fennell, A.; Zhou, R.; Nepal, M.P. Identification and Characterization of Mitogen-Activated Protein Kinase (MAPK) Genes in Sunflower. Plants 2019, 8, 28. [Google Scholar] [CrossRef]
- Jagodzik, P.; Tajdel-Zielinska, M.; Ciesla, A.; Marczak, M.; Ludwikow, A. Mitogen-Activated Protein Kinase Cascades in Plant Hormone Signaling. Front. Plant Sci. 2018, 9, 1387. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Gong, M.; Guo, J.; Xin, H.; Gao, Y.; Liu, C.; Dai, D.; Tang, L. Genome-wide Identification of Jatropha curcas MAPK, MAPKK, and MAPKKK Gene Families and Their Expression Profile Under Cold Stress. Sci. Rep. 2018, 8, 16163. [Google Scholar] [CrossRef] [PubMed]
- Jiang, M.; Wen, F.; Cao, J.; Li, P.; She, J.; Chu, Z. Genome-wide exploration of the molecular evolution and regulatory network of mitogen-activated protein kinase cascades upon multiple stresses in Brachypodiumdistachyon. BMC Genom. 2015, 16, 228. [Google Scholar] [CrossRef] [PubMed]
- Colcombet, J.; Sözen, C.; Hirt, H. Convergence of Multiple MAP3Ks on MKK3 Identifies a Set of Novel Stress MAPK Modules. Front. Plant Sci. 2016, 7, 1941. [Google Scholar] [CrossRef]
- Chardin, C.; Schenk, S.T.; Hirt, H.; Colcombet, J.; Krapp, A. Review: Mitogen-Activated Protein Kinases in nutritional signaling in Arabidopsis. Plant Sci. 2017, 260, 101–108. [Google Scholar] [CrossRef]
- Wang, P.; Du, Y.; Li, Y.; Ren, D.; Song, C.P. Hydrogen peroxide-mediated activation of MAP kinase 6 modulates nitric oxide biosynthesis and signal transduction in Arabidopsis. Plant Cell 2010, 22, 2981–2998. [Google Scholar] [CrossRef] [PubMed]
- Werth, E.G.; McConnell, E.W.; Gilbert, T.S.; CousoLianez, I.; Perez, C.A.; Manley, C.K.; Graves, L.M.; Umen, J.G.; Hicks, L.M. Probing the global kinome and phosphoproteome in Chlamydomonas reinhardtii via sequential enrichment and quantitative proteomics. Plant J. 2017, 89, 416–426. [Google Scholar] [CrossRef] [PubMed]
- Kalapos, B.; Hlavová, M.; Nádai, T.V.; Galiba, G.; Bišová, K.; Dóczi, R. Early Evolution of the Mitogen-Activated Protein Kinase Family in the Plant Kingdom. Sci. Rep. 2019, 9, 4094. [Google Scholar] [CrossRef] [PubMed]
- Fei, X.; Yu, J.; Li, Y.; Deng, X. CrMAPK3 regulates the expression of iron-deficiency-responsive genes in Chlamydomonas reinhardtii. BMC Biochem. 2017, 18, 6. [Google Scholar] [CrossRef] [PubMed]
- Dóczi, R.; Okrész, L.; Romero, A.E.; Paccanaro, A.; Bögre, L. Exploring the evolutionary path of plant MAPK networks. Trends Plant Sci. 2012, 17, 518–525. [Google Scholar] [CrossRef] [PubMed]
- Yang, A.; Suh, W.I.; Kang, N.K.; Lee, B.; Chang, Y.K. MAPK/ERK and JNK pathways regulate lipid synthesis and cell growth of Chlamydomonas reinhardtii under osmotic stress, respectively. Sci. Rep. 2018, 8, 13857. [Google Scholar] [CrossRef]
- Mohanta, T.K.; Arora, P.K.; Mohanta, N.; Parida, P.; Bae, H. Identification of new members of the MAPK gene family in plants shows diverse conserved domains and novel activation loop variants. BMC Genom. 2015, 16, 58. [Google Scholar] [CrossRef]
- Bigeard, J.; Hirt, H. Nuclear Signaling of Plant MAPKs. Front. Plant Sci. 2018, 9, 469. [Google Scholar] [CrossRef]
- Nagy, S.K.; Darula, Z.; Kállai, B.M.; Bögre, L.; Bánhegyi, G.; Medzihradszky, K.F.; Horváth, G.V.; Mészáros, T. Activation of AtMPK9 through autophosphorylation that makes it independent of the canonical MAPK cascades. Biochem. J. 2015, 467, 167–175. [Google Scholar] [CrossRef]
- Zhan, H.; Yue, H.; Zhao, X.; Wang, M.; Song, W.; Nie, X. Genome-Wide Identification and Analysis of MAPK and MAPKK Gene Families in Bread Wheat (Triticumaestivum L.). Genes 2017, 8, 284. [Google Scholar] [CrossRef]
- Hamel, L.P.; Nicole, M.C.; Sritubtim, S.; Morency, M.J.; Ellis, M.; Ehlting, J.; Beaudoin, N.; Barbazuk, B.; Klessig, D.; Lee, J.; et al. Ancient signals: Comparative genomics of plant MAPK and MAPKK gene families. Trends Plant Sci. 2006, 11, 192–198. [Google Scholar] [CrossRef] [PubMed]
- Jonak, C.; Okrész, L.; Bögre, L.; Hirt, H. Complexity, cross talk and integration of plant MAP kinase signalling. Curr. Opin. Plant Biol. 2002, 5, 415–424. [Google Scholar] [CrossRef]
- LópezGarcía de Lomana, A.; Schäuble, S.; Valenzuela, J.; Imam, S.; Carter, W.; Bilgin, D.D.; Yohn, C.B.; Turkarslan, S.; Reiss, D.J.; Orellana, M.V.; et al. Transcriptional program for nitrogen starvation-induced lipid accumulation in Chlamydomonas reinhardtii. Biotechnol. Biofuels 2015, 8, 207. [Google Scholar] [CrossRef]
- Zones, J.M.; Blaby, I.K.; Merchant, S.S.; Umen, J.G. High-Resolution Profiling of a Synchronized Diurnal Transcriptome from Chlamydomonas reinhardtii Reveals Continuous Cell and Metabolic Differentiation. Plant Cell 2015, 27, 2743–2769. [Google Scholar] [CrossRef] [PubMed]
- Choi, Y.E.; Rhee, J.K.; Kim, H.S.; Ahn, J.W.; Hwang, H.; Yang, J.W. Chemical Genetics Approach Reveals Importance of cAMP and MAP Kinase Signaling to Lipid and Carotenoid Biosynthesis in Microalgae. J. Microbiol. Biotechnol. 2015, 25, 637–647. [Google Scholar] [CrossRef] [PubMed]
- Roustan, V.; Bakhtiari, S.; Roustan, P.J.; Weckwerth, W. Quantitative in vivo phosphoproteomics reveals reversible signaling processes during nitrogen starvation and recovery in the biofuel model organism. Biotechnol. Biofuels 2017, 10, 280. [Google Scholar] [CrossRef]
- Marchive, C.; Roudier, F.; Castaings, L.; Bréhaut, V.; Blondet, E.; Colot, V.; Meyer, C.; Krapp, A. Nuclear retention of the transcription factor NLP7 orchestrates the early response to nitrate in plants. Nat. Commun. 2013, 4, 1713. [Google Scholar] [CrossRef]
- Forde, B.G.; Cutler, S.R.; Zaman, N.; Krysan, P.J. Glutamate signalling via a MEKK1 kinase-dependent pathway induces changes in Arabidopsis root architecture. Plant J. 2013, 75, 1–10. [Google Scholar] [CrossRef]
- Asai, T.; Tena, G.; Plotnikova, J.; Willmann, M.R.; Chiu, W.L.; Gomez-Gomez, L.; Boller, T.; Ausubel, F.M.; Sheen, J. MAP kinase signalling cascade in Arabidopsis innate immunity. Nature 2002, 415, 977–983. [Google Scholar] [CrossRef]
- Popescu, S.C.; Popescu, G.V.; Bachan, S.; Zhang, Z.; Gerstein, M.; Snyder, M.; Dinesh-Kumar, S.P. MAPK target networks in Arabidopsis thaliana revealed using functional protein microarrays. Genes Dev. 2009, 23, 80–92. [Google Scholar] [CrossRef]
- Ho, C.H.; Lin, S.H.; Hu, H.C.; Tsay, Y.F. CHL1 functions as a nitrate sensor in plants. Cell 2009, 138, 1184–1194. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.; Jun, Y.S.; Cha, O.K.; Sheen, J. Mitogen-activated protein kinases MPK3 and MPK6 are required for stem cell maintenance in the Arabidopsis shoot apical meristem. Plant Cell Rep. 2019, 38, 311–319. [Google Scholar] [CrossRef] [PubMed]
- Camps, M.; Nichols, A.; Arkinstall, S. Dual specificity phosphatases: A gene family for control of MAP kinase function. FASEB J. 2000, 14, 6–16. [Google Scholar] [CrossRef] [PubMed]
- Komis, G.; Illés, P.; Beck, M.; Šamaj, J. Microtubules and mitogen-activated protein kinase signalling. Curr. Opin. Plant Biol. 2011, 14, 650–657. [Google Scholar] [CrossRef] [PubMed]
- Lambeck, I.C.; Fischer-Schrader, K.; Niks, D.; Roeper, J.; Chi, J.C.; Hille, R.; Schwarz, G. Molecular mechanism of 14-3-3 protein-mediated inhibition of plant nitrate reductase. J. Biol. Chem. 2012, 287, 4562–4571. [Google Scholar] [CrossRef]
- Wang, P.; Du, Y.; Song, C.P. Phosphorylation by MPK6: A conserved transcriptional modification mediates nitrate reductase activation and NO production? Plant Signal. Behav. 2011, 6, 889–891. [Google Scholar] [CrossRef]
- Levine, R.P.; Ebersold, W.T. The genetics and cytology of Chlamydomonas. Annu. Rev. Microbiol. 1960, 14, 197–216. [Google Scholar] [CrossRef]
- Arnon, D.I. Copper enzumes in isolated chloroplasts. Polyphenoloxidase in Beta vulgaris. Plant Physiol. 1949, 24, 1–15. [Google Scholar] [CrossRef]
- Schloss, J.A.; Silflow, C.D.; Rosenbaum, J.L. mRNA abundance changes during flagellar regeneration in Chlamydomonas reinhardtii. Mol. Cell. Biol. 1984, 4, 424–434. [Google Scholar] [CrossRef]
- Arvidsson, S.; Kwasniewski, M.; Riaño-Pachón, D.M.; Mueller-Roeber, B. QuantPrime—A flexible tool for reliable high-throughput primer design for quantitative PCR. BMC Bioinform. 2008, 9, 465. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez-Ballester, D.; Camargo, A.; Fernandez, E. Ammonium transporter genes in Chlamydomonas: The nitrate-specific regulatory gene Nit2 is involved in Amt1;1 expression. Plant Mol. Biol. 2004, 56, 863–878. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef] [PubMed]
- Jones, D.T.; Taylor, W.R.; Thornton, J.M. The rapid generation of mutation data matrices from protein sequences. Comput. Appl. Biosci. 1992, 8, 275–282. [Google Scholar] [CrossRef] [PubMed]
- Zuckerkandl, E.; Pauling, L. Molecules as documents of evolutionary history. J. Theor. Biol. 1965, 8, 357–366. [Google Scholar] [CrossRef]
- Felsenstein, J. Confidence limits on phylogenies: An approach using the bootstrap. Evolution 1985, 39, 783–791. [Google Scholar] [CrossRef] [PubMed]







© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gomez-Osuna, A.; Calatrava, V.; Galvan, A.; Fernandez, E.; Llamas, A. Identification of the MAPK Cascade and its Relationship with Nitrogen Metabolism in the Green Alga Chlamydomonas reinhardtii. Int. J. Mol. Sci. 2020, 21, 3417. https://doi.org/10.3390/ijms21103417
Gomez-Osuna A, Calatrava V, Galvan A, Fernandez E, Llamas A. Identification of the MAPK Cascade and its Relationship with Nitrogen Metabolism in the Green Alga Chlamydomonas reinhardtii. International Journal of Molecular Sciences. 2020; 21(10):3417. https://doi.org/10.3390/ijms21103417
Chicago/Turabian StyleGomez-Osuna, Aitor, Victoria Calatrava, Aurora Galvan, Emilio Fernandez, and Angel Llamas. 2020. "Identification of the MAPK Cascade and its Relationship with Nitrogen Metabolism in the Green Alga Chlamydomonas reinhardtii" International Journal of Molecular Sciences 21, no. 10: 3417. https://doi.org/10.3390/ijms21103417
APA StyleGomez-Osuna, A., Calatrava, V., Galvan, A., Fernandez, E., & Llamas, A. (2020). Identification of the MAPK Cascade and its Relationship with Nitrogen Metabolism in the Green Alga Chlamydomonas reinhardtii. International Journal of Molecular Sciences, 21(10), 3417. https://doi.org/10.3390/ijms21103417