Derivation of Cell-Engineered Nanovesicles from Human Induced Pluripotent Stem Cells and Their Protective Effect on the Senescence of Dermal Fibroblasts
Abstract
1. Introduction
2. Results
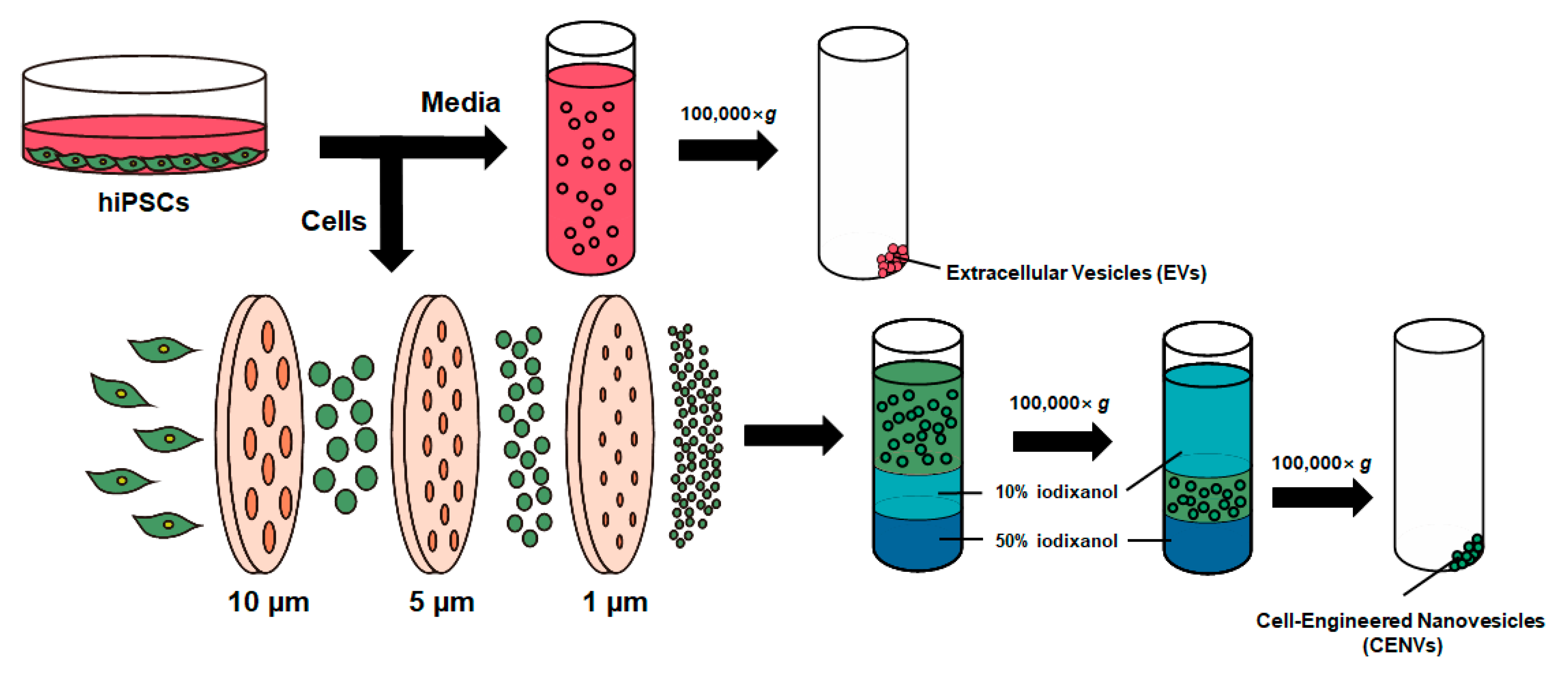
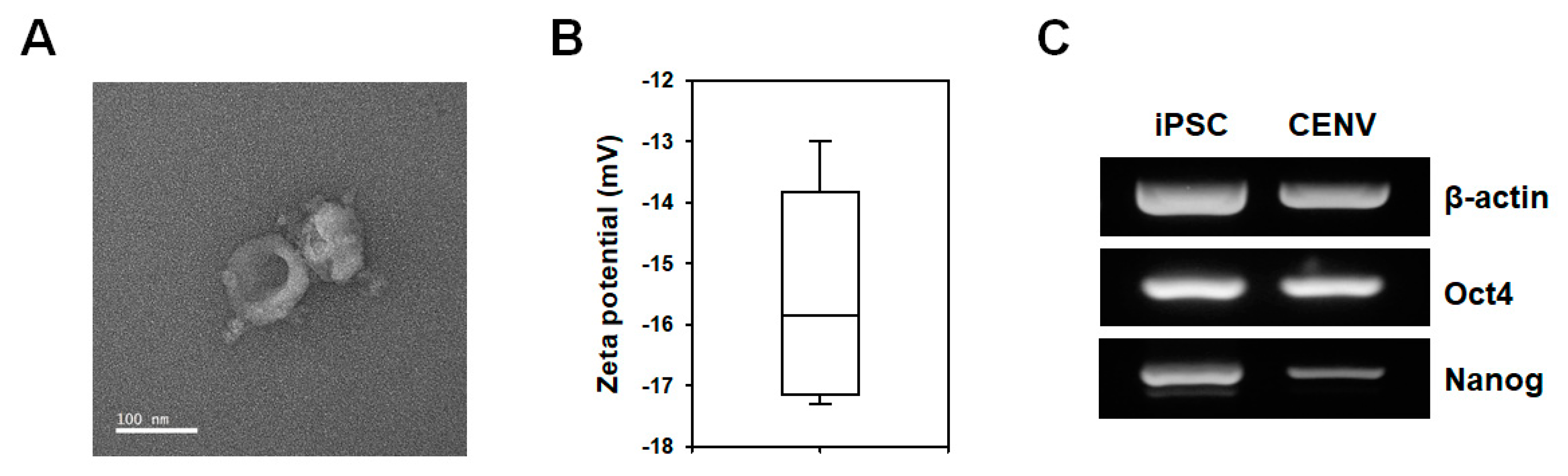
2.1. Characterization of iPSC-CENVs
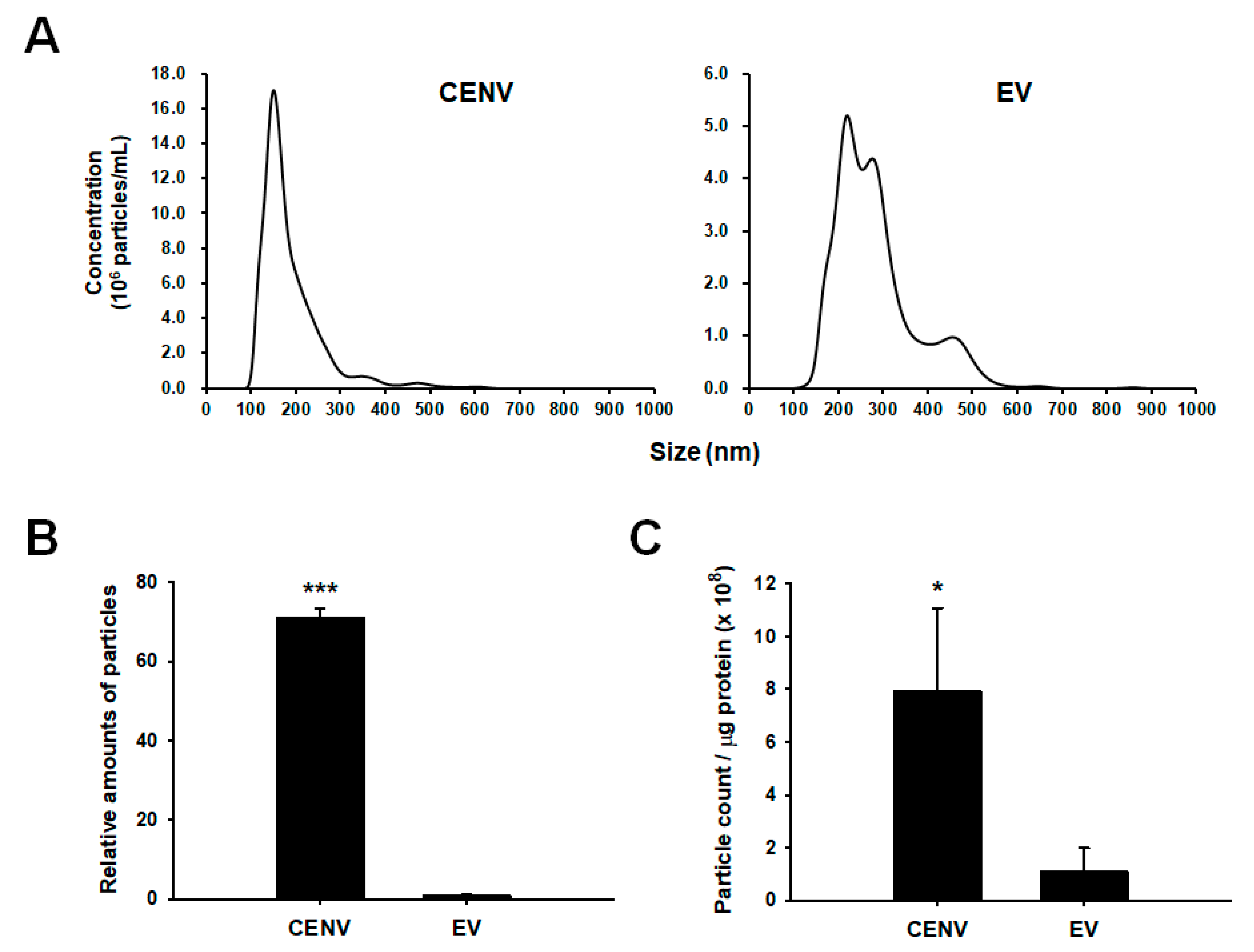
2.2. Comparison of Productivity and Purity between iPSC-CENV and iPSC-EV Sample
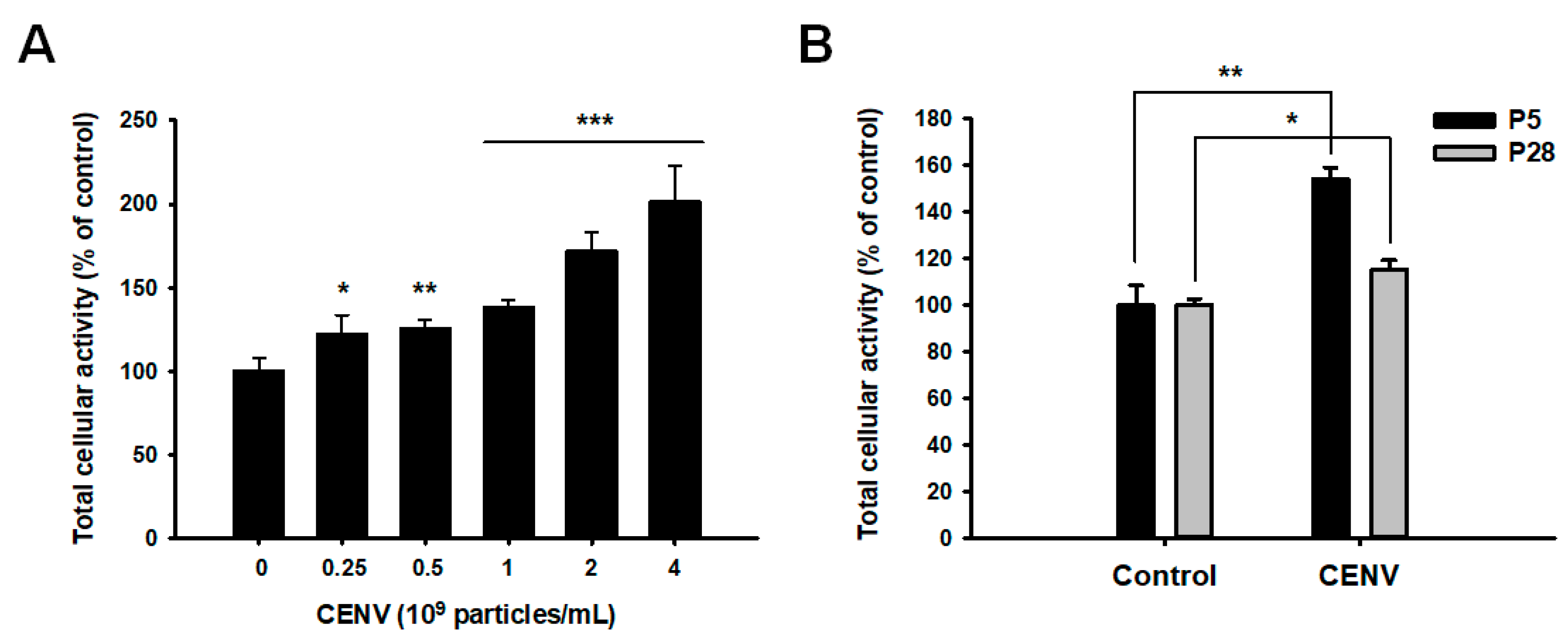
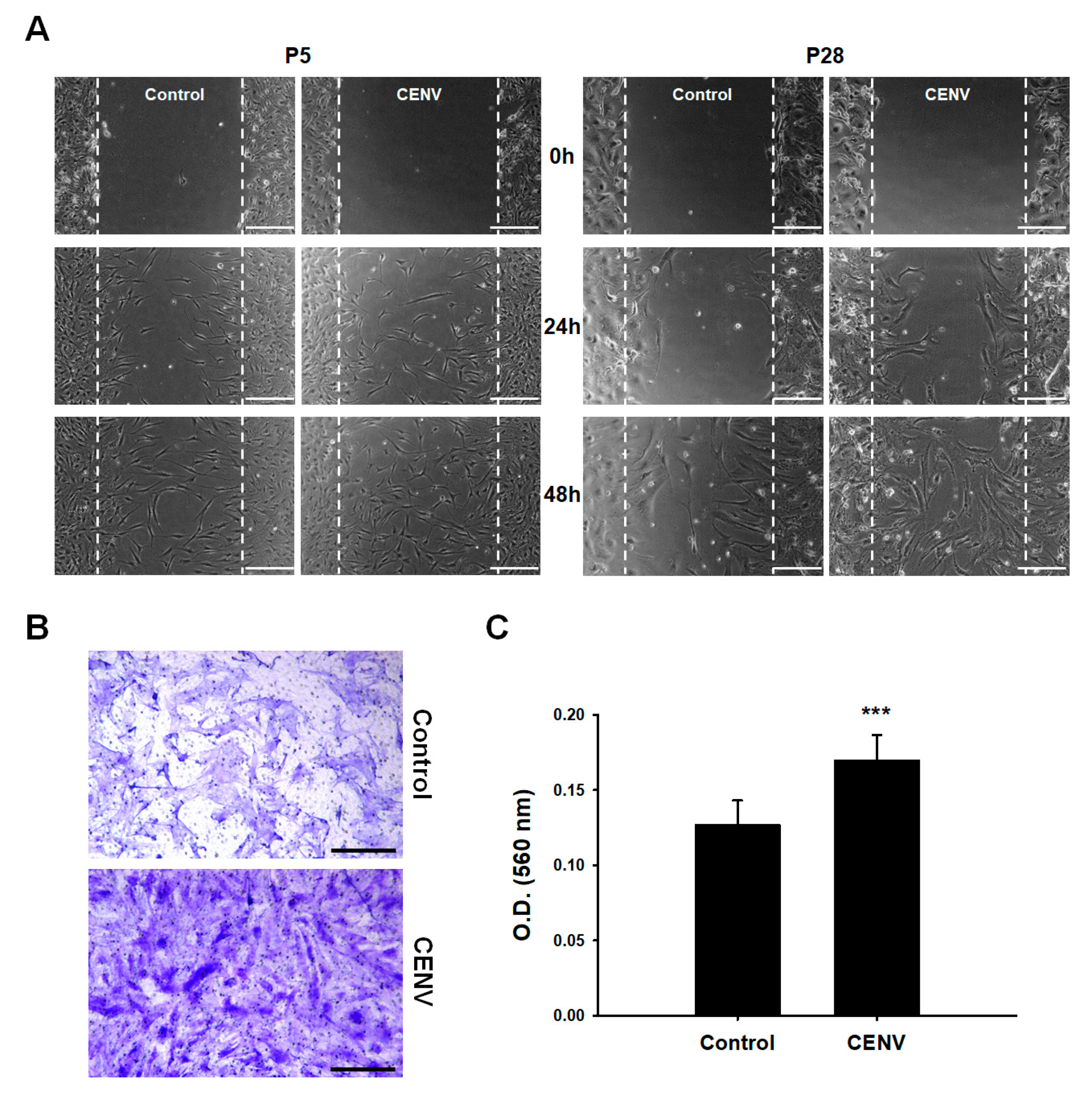
2.3. iPSC-CENV Promoted the Proliferation and Migration of HDFs
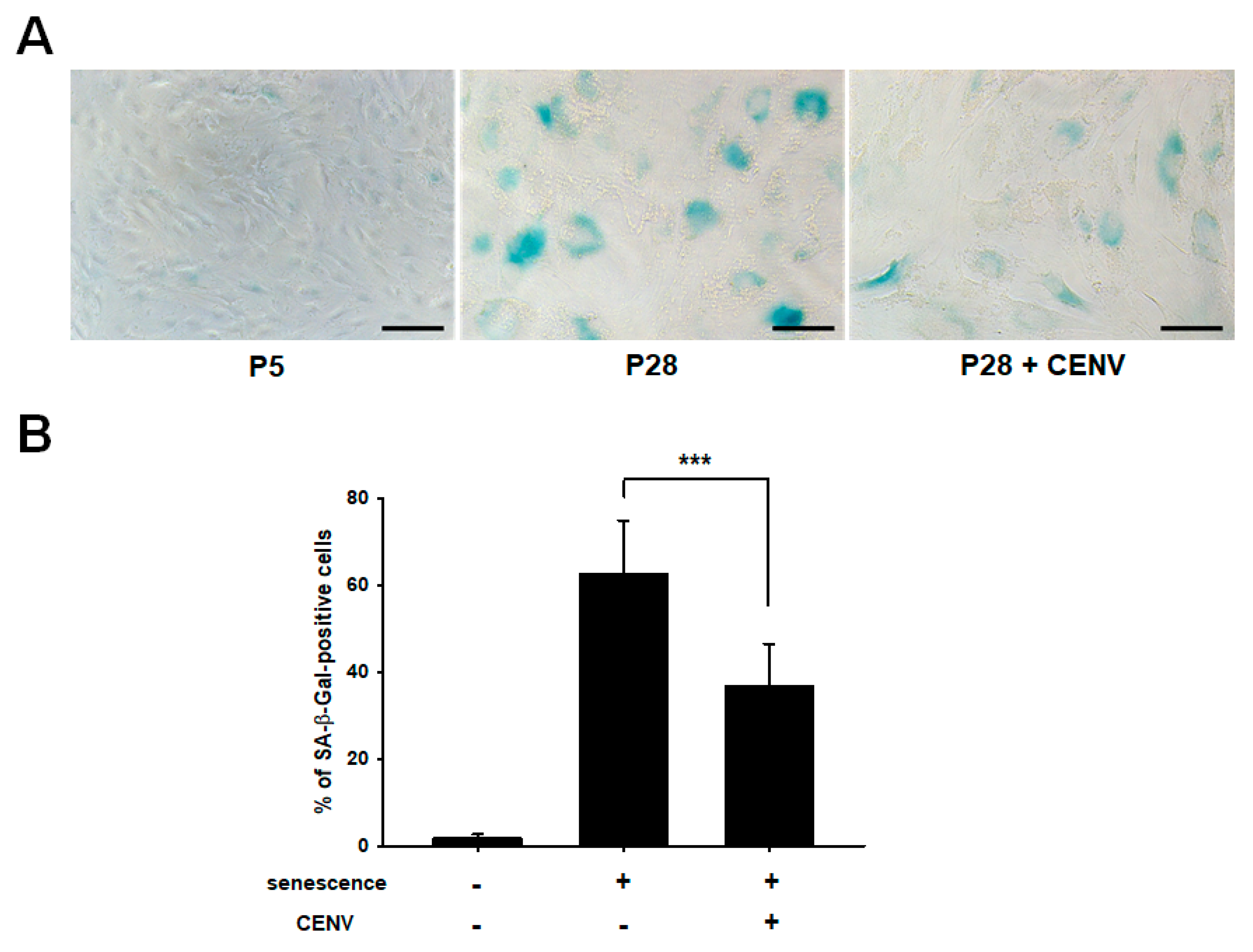
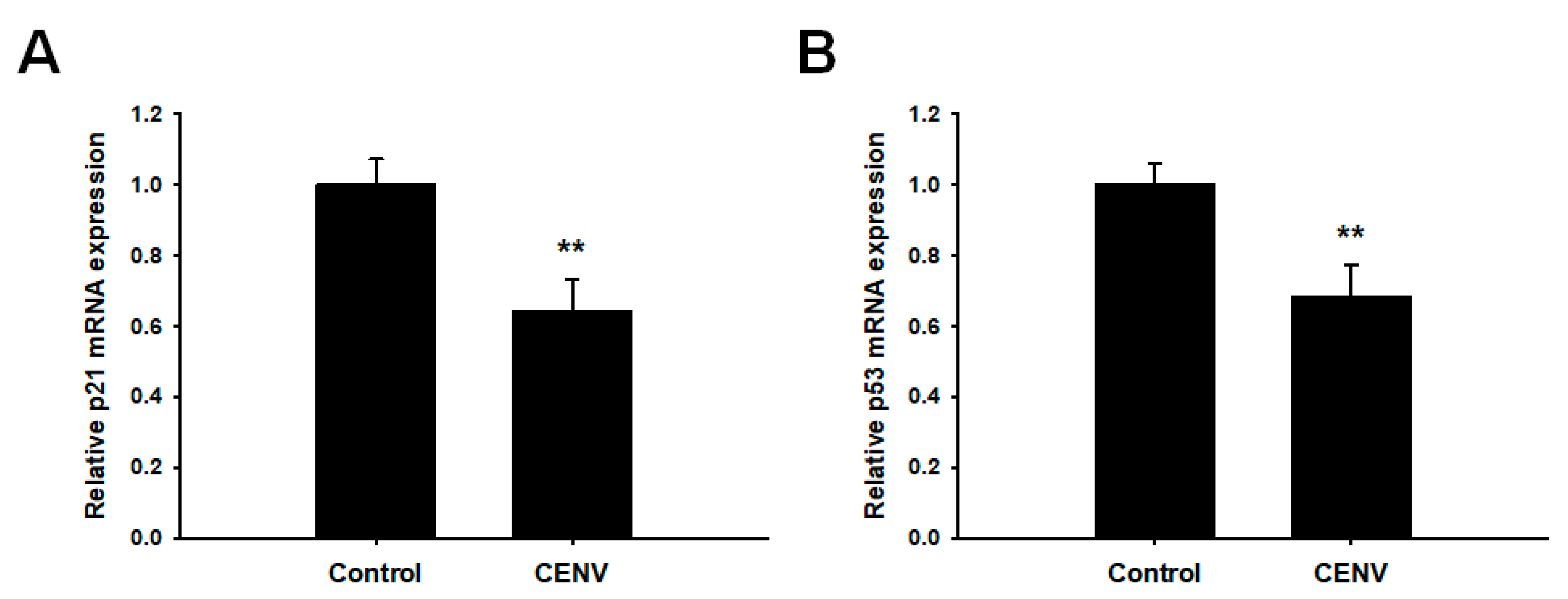
2.4. iPSC-CENV Reduced the Expression of Senescent-Related Genes
3. Discussion
4. Materials and Methods
4.1. Cell Culture
4.2. Preparation of iPSC-Derived EV and CENV
4.3. Characterization of iPSC-CENV
4.4. RNA Isolation and Reverse Transcriptase PCR (RT-PCR)
4.5. WST-8 Assay
4.6. Cell Migration Assay
4.7. SA-β-Gal Staining
4.8. Statistical Analysis
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| iPSC | Induced pluripotent stem cell |
| CENV | Cell-engineered nanovesicle |
| EV | Extracellular vesicle |
| ROS | Reactive oxygen species |
| HDF | Human dermal fibroblast |
| qPCR | Quantitative real-time polymerase chain reaction |
| RT-PCR | Reverse transcriptase polymerase chain reaction |
| ECM | Extracellular matrix |
| PSC | Pluripotent stem cell |
| CDK | Cyclin-dependent kinase |
| hUCMSC | Human umbilical cord mesenchymal stem cell |
| hPlaMSC | Human placenta mesenchymal stem cell |
| mESC | Murine embryonic stem cell |
| SA- β-Gal | Senescence-associated β-galactosidase |
| NTA | Nanoparticle tracking analysis |
| TEM | Transmission electron microscopy |
| DLS | Dynamic light scattering |
| FBS | Fetal bovine serum |
| DMEM | Dulbecco’s modified Eagle’s medium |
| PBS | Phosphate buffered saline |
| WST-8 | Water-soluble tetrazolium-8 |
| MMP | Matrix metalloproteinase |
References
- Farage, M.; Miller, K.; Elsner, P.; Maibach, H. Intrinsic and extrinsic factors in skin ageing: A review. Int. J. Cosmet. Sci. 2008, 30, 87–95. [Google Scholar] [CrossRef] [PubMed]
- Gosain, A.; DiPietro, L.A. Aging and wound healing. World J. Surg. 2004, 28, 321–326. [Google Scholar] [CrossRef] [PubMed]
- Fulop, T.; Khalil, A.; Larbi, A. The role of elastin peptides in modulating the immune response in aging and age-related diseases. Pathol. Biol. 2012, 60, 28–33. [Google Scholar] [CrossRef] [PubMed]
- López-Otín, C.; Blasco, M.A.; Partridge, L.; Serrano, M.; Kroemer, G. The hallmarks of aging. Cell 2013, 153, 1194–1217. [Google Scholar] [CrossRef]
- Tigges, J.; Krutmann, J.; Fritsche, E.; Haendeler, J.; Schaal, H.; Fischer, J.W.; Kalfalah, F.; Reinke, H.; Reifenberger, G.; Stuehler, K.; et al. The hallmarks of fibroblast ageing. Mech. Ageing Dev. 2014, 138, 26–44. [Google Scholar] [CrossRef]
- Powers, E.T.; Morimoto, R.I.; Dillin, A.; Kelly, J.W.; Balch, W.E. Biological and chemical approaches to diseases of proteostasis deficiency. Annu. Rev. Biochem. 2009, 78, 959–991. [Google Scholar] [CrossRef]
- Varani, J. Fibroblast aging: Intrinsic and extrinsic factors. Drug Discov. Today Ther. Strateg. 2010, 7, 65–70. [Google Scholar] [CrossRef]
- Brennan, M.; Bhatti, H.; Nerusu, K.C.; Bhagavathula, N.; Kang, S.; Fisher, G.J.; Varani, J.; Voorhees, J.J. Matrix metalloproteinase-1 is the major collagenolytic enzyme responsible for collagen damage in UV-irradiated human skin. Photochem. Photobiol. 2003, 78, 43–48. [Google Scholar] [CrossRef]
- Green, D.R.; Galluzzi, L.; Kroemer, G. Mitochondria and the autophagy–inflammation–cell death axis in organismal aging. Science 2011, 333, 1109–1112. [Google Scholar] [CrossRef]
- Blackburn, E.H.; Greider, C.W.; Szostak, J.W. Telomeres and telomerase: The path from maize, Tetrahymena and yeast to human cancer and aging. Nat. Med. 2006, 12, 1133–1138. [Google Scholar] [CrossRef]
- Campisi, J.; di Fagagna, F.D. Cellular senescence: When bad things happen to good cells. Nat. Rev. Mol. Cell Biol. 2007, 8, 729–740. [Google Scholar] [CrossRef] [PubMed]
- Telgenhoff, D.; Shroot, B. Cellular senescence mechanisms in chronic wound healing. Cell Death Differ. 2005, 12, 695–698. [Google Scholar] [CrossRef] [PubMed]
- Sikora, E.; Arendt, T.; Bennett, M.; Narita, M. Impact of cellular senescence signature on ageing research. Ageing Res. Rev. 2011, 10, 146–152. [Google Scholar] [CrossRef] [PubMed]
- Collado, M.; Serrano, M. Senescence in tumours: Evidence from mice and humans. Nat. Rev. Cancer 2010, 10, 51–57. [Google Scholar] [CrossRef] [PubMed]
- Rufini, A.; Tucci, P.; Celardo, I.; Melino, G. Senescence and aging: The critical roles of p53. Oncogene 2013, 32, 5129–5143. [Google Scholar] [CrossRef] [PubMed]
- Conboy, I.M.; Rando, T.A. Heterochronic parabiosis for the study of the effects of aging on stem cells and their niches. Cell Cycle 2012, 11, 2260–2267. [Google Scholar] [CrossRef]
- Kippin, T.E.; Martens, D.J.; van der Kooy, D. p21 loss compromises the relative quiescence of forebrain stem cell proliferation leading to exhaustion of their proliferation capacity. Genes Dev. 2005, 19, 756–767. [Google Scholar] [CrossRef]
- Chakkalakal, J.V.; Jones, K.M.; Basson, M.A.; Brack, A.S. The aged niche disrupts muscle stem cell quiescence. Nature 2012, 490, 355–360. [Google Scholar] [CrossRef]
- Lai, R.C.; Arslan, F.; Lee, M.M.; Sze, N.S.K.; Choo, A.; Chen, T.S.; Salto-Tellez, M.; Timmers, L.; Lee, C.N.; El Oakley, R.M.; et al. Exosome secreted by MSC reduces myocardial ischemia/reperfusion injury. Stem Cell Res. 2010, 4, 214–222. [Google Scholar] [CrossRef]
- Chen, L.; Wang, Y.; Pan, Y.; Zhang, L.; Shen, C.; Qin, G.; Ashraf, M.; Weintraub, N.; Ma, G.; Tang, Y. Cardiac progenitor-derived exosomes protect ischemic myocardium from acute ischemia/reperfusion injury. Biochem. Biophys. Res. Commun. 2013, 431, 566–571. [Google Scholar] [CrossRef]
- Sahoo, S.; Losordo, D.W. Exosomes and cardiac repair after myocardial infarction. Circ. Res. 2014, 114, 333–344. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Guan, J.; Niu, X.; Hu, G.; Guo, S.; Li, Q.; Xie, Z.; Zhang, C.; Wang, Y. Exosomes released from human induced pluripotent stem cells-derived MSCs facilitate cutaneous wound healing by promoting collagen synthesis and angiogenesis. J. Transl. Med. 2015, 13, 49. [Google Scholar] [CrossRef] [PubMed]
- Zhang, B.; Wang, M.; Gong, A.; Zhang, X.; Wu, X.; Zhu, Y.; Shi, H.; Wu, L.; Zhu, W.; Qian, H.; et al. HucMSC-exosome mediated-Wnt4 signaling is required for cutaneous wound healing. Stem Cells 2015, 33, 2158–2168. [Google Scholar] [CrossRef] [PubMed]
- Hu, L.; Wang, J.; Zhou, X.; Xiong, Z.; Zhao, J.; Yu, R.; Huang, F.; Zhang, H.; Chen, L. Exosomes derived from human adipose mensenchymal stem cells accelerates cutaneous wound healing via optimizing the characteristics of fibroblasts. Sci. Rep. 2016, 6, 32993. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Chen, C.; Hu, B.; Niu, X.; Liu, X.; Zhang, G.; Zhang, C.; Li, Q.; Wang, Y. Exosomes derived from human endothelial progenitor cells accelerate cutaneous wound healing by promoting angiogenesis through Erk1/2 signaling. Int. J. Biol. Sci. 2016, 12, 1472–1487. [Google Scholar] [CrossRef] [PubMed]
- Vlassov, A.V.; Magdaleno, S.; Setterquist, R.; Conrad, R. Exosomes: Current knowledge of their composition, biological functions, and diagnostic and therapeutic potentials. Biochim. Biophys. Acta Gen. Subj. 2012, 1820, 940–948. [Google Scholar] [CrossRef] [PubMed]
- Kowal, J.; Tkach, M.; Théry, C. Biogenesis and secretion of exosomes. Curr. Opin. Cell Biol. 2014, 29, 116–125. [Google Scholar] [CrossRef]
- Simons, M.; Raposo, G. Exosomes–vesicular carriers for intercellular communication. Curr. Opin. Cell Biol. 2009, 21, 575–581. [Google Scholar] [CrossRef]
- Oh, M.; Lee, J.; Kim, Y.J.; Rhee, W.J.; Park, J.H. Exosomes derived from human induced pluripotent stem cells ameliorate the aging of skin fibroblasts. Int. J. Mol. Sci. 2018, 19, 1715. [Google Scholar] [CrossRef]
- Rosler, E.S.; Fisk, G.J.; Ares, X.; Irving, J.; Miura, T.; Rao, M.S.; Carpenter, M.K. Long-term culture of human embryonic stem cells in feeder-free conditions. Dev. Dyn. 2004, 229, 259–274. [Google Scholar] [CrossRef]
- Chen, G.; Gulbranson, D.R.; Hou, Z.; Bolin, J.M.; Ruotti, V.; Probasco, M.D.; Smuga-Otto, K.; Howden, S.E.; Diol, N.R.; Propson, N.E.; et al. Chemically defined conditions for human iPSC derivation and culture. Nat. Methods 2011, 8, 424–429. [Google Scholar] [CrossRef] [PubMed]
- Han, C.; Jeong, D.; Kim, B.; Jo, W.; Kang, H.; Cho, S.; Kim, K.H.; Park, J. Mesenchymal stem cell engineered nanovesicles for accelerated skin wound closure. ACS Biomater. Sci. Eng. 2019, 5, 1534–1543. [Google Scholar] [CrossRef]
- Jo, W.; Jeong, D.; Kim, J.; Park, J. Self-Renewal of Bone Marrow Stem Cells by Nanovesicles Engineered from Embryonic Stem Cells. Adv. Healthc. Mater. 2016, 5, 3148–3156. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.S.; Kim, J.Y.; Cho, R.; Shin, D.M.; Lee, S.W.; Oh, Y.M. Adipose stem cell-derived nanovesicles inhibit emphysema primarily via an FGF2-dependent pathway. Exp. Mol. Med. 2017, 49, e284. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.Y.; Kumar, H.; Jo, M.J.; Kim, J.; Yoon, J.K.; Lee, J.R.; Kang, M.; Choo, Y.W.; Song, S.Y.; Kwon, S.P.; et al. Therapeutic efficacy-potentiated and diseased organ-targeting nanovesicles derived from mesenchymal stem cells for spinal cord injury treatment. Nano Lett. 2018, 18, 4965–4975. [Google Scholar] [CrossRef]
- Goh, W.J.; Zou, S.; Ong, W.Y.; Torta, F.; Alexandra, A.F.; Schiffelers, R.M.; Storm, G.; Wang, J.W.; Czarny, B.; Pastorin, G. Bioinspired cell-derived nanovesicles versus exosomes as drug delivery systems: A cost-effective alternative. Sci. Rep. 2017, 7, 14322. [Google Scholar] [CrossRef]
- Jang, S.C.; Kim, O.Y.; Yoon, C.M.; Choi, D.S.; Roh, T.Y.; Park, J.; Nilsson, J.; Lotvall, J.; Kim, Y.K.; Gho, Y.S. Bioinspired exosome-mimetic nanovesicles for targeted delivery of chemotherapeutics to malignant tumors. ACS Nano 2013, 7, 7698–7710. [Google Scholar] [CrossRef]
- Liu, S.; Mahairaki, V.; Bai, H.; Ding, Z.; Li, J.; Witwer, K.W.; Cheng, L. Highly purified human extracellular vesicles produced by stem cells alleviate aging cellular phenotypes of senescent human cells. Stem Cells 2019, 37, 779–790. [Google Scholar] [CrossRef]
- Farnebo, M.; Bykov, V.J.; Wiman, K.G. The p53 tumor suppressor: A master regulator of diverse cellular processes and therapeutic target in cancer. Biochem. Biophys. Res. Commun. 2010, 396, 85–89. [Google Scholar] [CrossRef]
- Chen, Z.; Trotman, L.C.; Shaffer, D.; Lin, H.K.; Dotan, Z.A.; Niki, M.; Koutcher, J.A.; Scher, H.I.; Ludwig, T.; Gerald, W.; et al. Crucial role of p53-dependent cellular senescence in suppression of Pten-deficient tumorigenesis. Nature 2005, 436, 725–730. [Google Scholar] [CrossRef]
- Kim, Y.Y.; Jee, H.J.; Um, J.H.; Kim, Y.M.; Bae, S.S.; Yun, J. Cooperation between p21 and Akt is required for p53-dependent cellular senescence. Aging Cell 2017, 16, 1094–1103. [Google Scholar] [CrossRef] [PubMed]
- Zou, M.; Baitei, E.; Al-Rijjal, R.; Parhar, R.; Al-Mohanna, F.; Kimura, S.; Pritchard, C.; Binessa, H.; Alzahrani, A.; Al-Khalaf, H.; et al. TSH overcomes BrafV600E-induced senescence to promote tumor progression via downregulation of p53 expression in papillary thyroid cancer. Oncogene 2016, 35, 1909–1918. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.Y.; Wang, H.J.; Tan, Y.Z. Wnt/β-catenin signaling induces the aging of mesenchymal stem cells through the DNA damage response and the p53/p21 pathway. PLoS ONE 2011, 6, e21397. [Google Scholar] [CrossRef] [PubMed]
- Campisi, J. Senescent cells, tumor suppression, and organismal aging: Good citizens, bad neighbors. Cell 2005, 120, 513–522. [Google Scholar] [CrossRef]
- Coppé, J.P.; Desprez, P.Y.; Krtolica, A.; Campisi, J. The senescence-associated secretory phenotype: The dark side of tumor suppression. Annu. Rev. Pathol. Mech. Dis. 2010, 5, 99–118. [Google Scholar] [CrossRef]
- Struewing, I.T.; Durham, S.N.; Barnett, C.D.; Mao, C.D. Enhanced endothelial cell senescence by lithium-induced matrix metalloproteinase-1 expression. J. Biol. Chem. 2009, 284, 17595–17606. [Google Scholar] [CrossRef]
- Zhu, F.; Li, Y.; Zhang, J.; Piao, C.; Liu, T.; Li, H.H.; Du, J. Senescent cardiac fibroblast is critical for cardiac fibrosis after myocardial infarction. PLoS ONE 2013, 8, e74535. [Google Scholar] [CrossRef]
- Fang, S.; Xu, C.; Zhang, Y.; Xue, C.; Yang, C.; Bi, H.; Qian, X.; Wu, M.; Ji, K.; Zhao, Y. Umbilical cord-derived mesenchymal stem cell-derived exosomal micrornas suppress myofibroblast differentiation by inhibiting the transforming growth factor-β/SMAD2 pathway during wound healing. Stem Cells Transl. Med. 2016, 5, 1425–1439. [Google Scholar] [CrossRef]
- Tooi, M.; Komaki, M.; Morioka, C.; Honda, I.; Iwasaki, K.; Yokoyama, N.; Ayame, H.; Izumi, Y.; Morita, I. Placenta mesenchymal stem cell derived exosomes confer plasticity on fibroblasts. J. Cell. Biochem. 2016, 117, 1658–1670. [Google Scholar] [CrossRef]
- Alvarez-Erviti, L.; Seow, Y.; Yin, H.; Betts, C.; Lakhal, S.; Wood, M.J. Delivery of siRNA to the mouse brain by systemic injection of targeted exosomes. Nat. Biotechnol. 2011, 29, 341–345. [Google Scholar] [CrossRef]
- Bjørge, I.; Kim, S.; Mano, J.; Kalionis, B.; Chrzanowski, W. Extracellular vesicles, exosomes and shedding vesicles in regenerative medicine–a new paradigm for tissue repair. Biomater. Sci. 2018, 6, 60–78. [Google Scholar] [CrossRef] [PubMed]
- Jeong, D.; Jo, W.; Yoon, J.; Kim, J.; Gianchandani, S.; Gho, Y.S.; Park, J. Nanovesicles engineered from ES cells for enhanced cell proliferation. Biomaterials 2014, 35, 9302–9310. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Kaslan, M.; Lee, S.H.; Yao, J.; Gao, Z. Progress in exosome isolation techniques. Theranostics 2017, 7, 789–804. [Google Scholar] [CrossRef] [PubMed]
- Greening, D.W.; Xu, R.; Ji, H.; Tauro, B.J.; Simpson, R.J. A protocol for exosome isolation and characterization: Evaluation of ultracentrifugation, density-gradient separation, and immunoaffinity capture methods. In Proteomic Profiling: Methods in Molecular Biology; Posch, A., Ed.; Humana Press: New York, NY, USA, 2015; Volume 1295, pp. 179–209. [Google Scholar]
- Patel, G.K.; Khan, M.A.; Zubair, H.; Srivastava, S.K.; Khushman, M.D.; Singh, S.; Singh, A.P. Comparative analysis of exosome isolation methods using culture supernatant for optimum yield, purity and downstream applications. Sci. Rep. 2019, 9, 5335. [Google Scholar] [CrossRef]
- Van Deun, J.; Mestdagh, P.; Sormunen, R.; Cocquyt, V.; Vermaelen, K.; Vandesompele, J.; Bracke, M.; De Wever, O.; Hendrix, A. The impact of disparate isolation methods for extracellular vesicles on downstream RNA profiling. J. Extracell. Vesicles 2014, 3, 24858. [Google Scholar] [CrossRef]
- Jo, W.; Kim, J.; Yoon, J.; Jeong, D.; Cho, S.; Jeong, H.; Yoon, Y.J.; Kim, S.C.; Gho, Y.S.; Park, J. Large-scale generation of cell-derived nanovesicles. Nanoscale 2014, 6, 12056–12064. [Google Scholar] [CrossRef]
| Gene | Primer | Sequence (5′-3′) |
|---|---|---|
| β-actin | Sense | GTG GGG CGC CCC AGG CAC CA |
| Antisense | CTC CTT AAT GTC ACG CAC GAT TT | |
| Oct4 | Sense | TGT ACT CCT CGG TCC CTT TC |
| Antisense | TCC AGG TTT TCT TTC CTA GC | |
| Nanog | Sense | CAA AGG CAA ACA ACC CAC TT |
| Antisense | ATT GTT CCA GGT CTG GTT GC | |
| p53 | Sense | GCC CAA CAA CAC CAG CTC CT |
| Antisense | CCT GGG CAT CCT TGA GTT CC | |
| p21 | Sense | GAC ACC ACT GGA GGG TGA CT |
| Antisense | CAG GTC CAC ATG GTC TTC CT |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lee, H.; Cha, H.; Park, J.H. Derivation of Cell-Engineered Nanovesicles from Human Induced Pluripotent Stem Cells and Their Protective Effect on the Senescence of Dermal Fibroblasts. Int. J. Mol. Sci. 2020, 21, 343. https://doi.org/10.3390/ijms21010343
Lee H, Cha H, Park JH. Derivation of Cell-Engineered Nanovesicles from Human Induced Pluripotent Stem Cells and Their Protective Effect on the Senescence of Dermal Fibroblasts. International Journal of Molecular Sciences. 2020; 21(1):343. https://doi.org/10.3390/ijms21010343
Chicago/Turabian StyleLee, Hyelim, Hyeonjin Cha, and Ju Hyun Park. 2020. "Derivation of Cell-Engineered Nanovesicles from Human Induced Pluripotent Stem Cells and Their Protective Effect on the Senescence of Dermal Fibroblasts" International Journal of Molecular Sciences 21, no. 1: 343. https://doi.org/10.3390/ijms21010343
APA StyleLee, H., Cha, H., & Park, J. H. (2020). Derivation of Cell-Engineered Nanovesicles from Human Induced Pluripotent Stem Cells and Their Protective Effect on the Senescence of Dermal Fibroblasts. International Journal of Molecular Sciences, 21(1), 343. https://doi.org/10.3390/ijms21010343





