A Symphony of Signals: Intercellular and Intracellular Signaling Mechanisms Underlying Circadian Timekeeping in Mice and Flies
Abstract
1. Introduction
2. Neuromodulation of Entrainment of the Mammalian Central Pacemaker
2.1. Transmission of Photic Information to the SCN: The Role of Neurotransmitters and Neuropeptides
2.1.1. Glutamate
2.1.2. Pituitary Adenylate Cyclase-Activating Polypeptide (PACAP)
2.1.3. Gamma-Aminobutyic Acid (GABA)
2.1.4. Serotonin (5-hydroxy-tryptamine)
2.1.5. Acetylcholine
2.1.6. Glycine
2.1.7. Vasoactive Intestinal Peptide (VIP)
2.1.8. Gastrin-Releasing Peptide (GRP)
2.2. Protein Kinases Implicated in the Regulation of Photic Entrainment of the SCN
2.2.1. Extracellular Signal-Regulated Kinases (ERK), Downstream Effector Kinases and Upstream Regulators
2.2.2. Mammalian Target of Rapamycin (mTOR)
2.2.3. c-Jun NH2-Terminal Kinases (JNK)
2.2.4. p38 MAPK
2.2.5. Ca2+/Calmodulin-Dependent Protein Kinase II (CaMKII)
2.2.6. cAMP-Activated Protein Kinase (PKA)
2.2.7. Protein Kinase C (PKC)
2.2.8. cGMP-Dependent Protein Kinase (PKG)
2.2.9. Salt-Inducible Kinases (SIKs)
2.2.10. Casein Kinase 1 (CK1)
2.2.11. Glycogen Synthase Kinase 3 (GSK3)
2.2.12. G-protein Coupled Receptor Kinase 2 (GRK2)
3. Neuromodulation of Circadian Timekeeping and Synchrony within the SCN
3.1. The Role of Neurotransmitters and Neuropeptides
3.1.1. GABA
3.1.2. VIP
3.1.3. Arginine Vasopressin (AVP)
3.1.4. GRP
3.1.5. Prokineticin 2
3.2. Protein Kinases that Modulate Circadian Period
3.2.1. CK1
3.2.2. Casein Kinase 2 (CK2)
3.2.3. GSK3
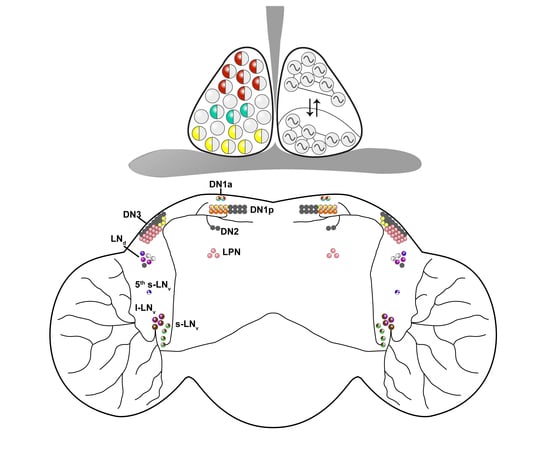
4. Neuromodulation in the Drosophila Pacemaker
4.1. Neuropeptides
4.1.1. Pigment Dispersing Factor (PDF)
4.1.2. Neuropeptide F (NPF) and Small NPF (sNPF)
4.1.3. Ion Transport Peptide (ITP)
4.1.4. Diuretic Hormone 31 (DH31)
4.1.5. CChamide 1
4.1.6. Allatostatin-C (AstC)
4.1.7. IPNamide
4.2. Neurotransmitters
4.2.1. Glutamate
4.2.2. GABA
4.2.3. Serotonin
4.2.4. Acetylcholine
4.2.5. Glycine
5. Protein Kinases Implicated in Circadian Timekeeping in Drosophila
5.1. Doubletime (DBT)
5.2. CK2
5.3. Shaggy/GSK3
5.4. NEMO/NLK
5.5. PKA
5.6. p38 MAPK
5.7. Ribosomal S6 kinase (RSK)
5.8. Ras/MAPK
5.9. TOR
5.10. AMPK/SIK3
5.11. GPRK2
6. Concluding Remarks
Author Contributions
Funding
Conflicts of Interest
References
- Partch, C.L.; Green, C.B.; Takahashi, J.S. Molecular architecture of the mammalian circadian clock. Trends Cell Biol. 2014, 24, 90–99. [Google Scholar] [CrossRef] [PubMed]
- Ralph, M.R.; Foster, R.G.; Davis, F.C.; Menaker, M. Transplanted suprachiasmatic nucleus determines circadian period. Science 1990, 247, 975–978. [Google Scholar] [CrossRef]
- Albrecht, U. Timing to Perfection: The Biology of Central and Peripheral Circadian Clocks. Neuron 2012, 74, 246–260. [Google Scholar] [CrossRef]
- Wagner, S.; Castel, M.; Gainer, H.; Yarom, Y. GABA in the mammalian suprachiasmatic nucleus and its role in diurnal rhythmicity. Nature 1997, 387, 598–603. [Google Scholar] [CrossRef]
- Abrahamson, E.E.; Moore, R.Y. Suprachiasmatic nucleus in the mouse: Retinal innervation, intrinsic organization and efferent projections. Brain Res. 2001, 916, 172–191. [Google Scholar] [CrossRef]
- Antle, M.C.; Silver, R. Orchestrating time: Arrangements of the brain circadian clock. Trends Neurosci. 2005, 28, 145–151. [Google Scholar] [CrossRef] [PubMed]
- Hendrickson, A.E.; Wagoner, N.; Cowan, W.M. An autoradiographic and electron microscopic study of retino-hypothalamic connections. Z. Zellforsch. Mikrosk. Anat. 1972, 135, 1–26. [Google Scholar] [CrossRef]
- Moore, R.Y.; Lenn, N.J. A retinohypothalamic projection in the rat. J. Comp. Neurol. 1972, 146, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Card, J.P.; Brecha, N.; Karten, H.J.; Moore, R.Y. Immunocytochemical localization of vasoactive intestinal polypeptide-containing cells and processes in the suprachiasmatic nucleus of the rat: Light and electron microscopic analysis. J. Neurosci. 1981, 1, 1289–1303. [Google Scholar] [CrossRef] [PubMed]
- Mikkelsen, J.D.; Larsen, P.J.; O’Hare, M.M.; Wiegand, S.J. Gastrin releasing peptide in the rat suprachiasmatic nucleus: An immunohistochemical, chromatographic and radioimmunological study. Neuroscience 1991, 40, 55–66. [Google Scholar] [CrossRef]
- Mendoza-Viveros, L.; Bouchard-Cannon, P.; Hegazi, S.; Cheng, A.H.; Pastore, S.; Cheng, H.-Y.M. Molecular modulators of the circadian clock: Lessons from flies and mice. Cell. Mol. Life Sci. 2017, 74, 1035–1059. [Google Scholar] [CrossRef] [PubMed]
- Yao, Z.; Shafer, O.T. The Drosophila Circadian Clock Is a Variably Coupled Network of Multiple Peptidergic Units. Science 2014, 343, 1516–1520. [Google Scholar] [CrossRef] [PubMed]
- Yao, Z.; Bennett, A.J.; Clem, J.L.; Shafer, O.T. The Drosophila Clock Neuron Network Features Diverse Coupling Modes and Requires Network-wide Coherence for Robust Circadian Rhythms. Cell Rep. 2016, 17, 2873–2881. [Google Scholar] [CrossRef] [PubMed]
- Yoshii, T.; Rieger, D.; Helfrich-Förster, C. Two clocks in the brain: An update of the morning and evening oscillator model in Drosophila. Prog. Brain Res. 2012, 199, 59–82. [Google Scholar] [PubMed]
- Nitabach, M.N.; Taghert, P.H. Organization of the Drosophila Circadian Control Circuit. Curr. Biol. 2008, 18, R84–R93. [Google Scholar] [CrossRef] [PubMed]
- Hamblen-Coyle, M.J.; Wheeler, D.A.; Rutila, J.E.; Rosbash, M.; Hall, J.C. Behavior of period-altered circadian rhythm mutants ofDrosophila in light: Dark cycles (Diptera: Drosophilidae). J. Insect Behav. 1992, 5, 417–446. [Google Scholar] [CrossRef]
- Wheeler, D.A.; Hamblen-Coyle, M.J.; Dushay, M.S.; Hall, J.C. Behavior in Light-Dark Cycles of Drosophila Mutants That Are Arrhythmic, Blind, or Both. J. Biol. Rhythms 1993, 8, 67–94. [Google Scholar] [CrossRef] [PubMed]
- Grima, B.; Chélot, E.; Xia, R.; Rouyer, F. Morning and evening peaks of activity rely on different clock neurons of the Drosophila brain. Nature 2004, 431, 869–873. [Google Scholar] [CrossRef]
- Stoleru, D.; Peng, Y.; Agosto, J.; Rosbash, M. Coupled oscillators control morning and evening locomotor behaviour of Drosophila. Nature 2004, 431, 862–868. [Google Scholar] [CrossRef]
- Rieger, D.; Shafer, O.T.; Tomioka, K.; Helfrich-Förster, C. Functional Analysis of Circadian Pacemaker Neurons in Drosophila melanogaster. J. Neurosci. 2006, 26, 2531–2543. [Google Scholar] [CrossRef]
- Shang, Y.; Griffith, L.C.; Rosbash, M. Light-arousal and circadian photoreception circuits intersect at the large PDF cells of the Drosophila brain. Proc. Natl. Acad. Sci. USA 2008, 105, 19587–19594. [Google Scholar] [CrossRef] [PubMed]
- Shafer, O.T.; Taghert, P.H. RNA-Interference Knockdown of Drosophila Pigment Dispersing Factor in Neuronal Subsets: The Anatomical Basis of a Neuropeptide’s Circadian Functions. PLoS ONE 2009, 4, e8298. [Google Scholar] [CrossRef]
- Cusumano, P.; Klarsfeld, A.; Chélot, E.; Picot, M.; Richier, B.; Rouyer, F. PDF-modulated visual inputs and cryptochrome define diurnal behavior in Drosophila. Nat. Neurosci. 2009, 12, 1431–1437. [Google Scholar] [CrossRef] [PubMed]
- Sheeba, V.; Fogle, K.J.; Holmes, T.C. Persistence of morning anticipation behavior and high amplitude morning startle response following functional loss of small ventral lateral neurons in Drosophila. PLoS ONE 2010, 5, e11628. [Google Scholar] [CrossRef]
- Schlichting, M.; Menegazzi, P.; Lelito, K.R.; Yao, Z.; Buhl, E.; Dalla Benetta, E.; Bahle, A.; Denike, J.; Hodge, J.J.; Helfrich-Förster, C.; et al. A Neural Network Underlying Circadian Entrainment and Photoperiodic Adjustment of Sleep and Activity in Drosophila. J. Neurosci. 2016, 36, 9084–9096. [Google Scholar] [CrossRef]
- Guo, F.; Yu, J.; Jung, H.J.; Abruzzi, K.C.; Luo, W.; Griffith, L.C.; Rosbash, M. Circadian neuron feedback controls the Drosophila sleep–activity profile. Nature 2016, 536, 292–297. [Google Scholar] [CrossRef]
- Stoleru, D.; Peng, Y.; Nawathean, P.; Rosbash, M. A resetting signal between Drosophila pacemakers synchronizes morning and evening activity. Nature 2005, 438, 238–242. [Google Scholar] [CrossRef] [PubMed]
- Helfrich-Förster, C. Robust circadian rhythmicity of Drosophila melanogaster requires the presence of lateral neurons: A brain-behavioral study of disconnected mutants. J. Comp. Physiol. A Sensory Neural Behav. Physiol. 1998, 182, 435–453. [Google Scholar] [CrossRef]
- Renn, S.C.; Park, J.H.; Rosbash, M.; Hall, J.C.; Taghert, P.H. A pdf neuropeptide gene mutation and ablation of PDF neurons each cause severe abnormalities of behavioral circadian rhythms in Drosophila. Cell 1999, 99, 791–802. [Google Scholar] [CrossRef]
- Peng, Y.; Stoleru, D.; Levine, J.D.; Hall, J.C.; Rosbash, M. Drosophila Free-Running Rhythms Require Intercellular Communication. PLoS Biol. 2003, 1, e13. [Google Scholar] [CrossRef]
- Lin, Y. The Neuropeptide Pigment-Dispersing Factor Coordinates Pacemaker Interactions in the Drosophila Circadian System. J. Neurosci. 2004, 24, 7951–7957. [Google Scholar] [CrossRef] [PubMed]
- Yoshii, T.; Wülbeck, C.; Sehadova, H.; Veleri, S.; Bichler, D.; Stanewsky, R.; Helfrich-Förster, C. The neuropeptide pigment-dispersing factor adjusts period and phase of Drosophila’s clock. J. Neurosci. 2009, 29, 2597–2610. [Google Scholar] [CrossRef]
- Parisky, K.M.; Agosto, J.; Pulver, S.R.; Shang, Y.; Kuklin, E.; Hodge, J.J.; Kang, K.; Liu, X.; Garrity, P.A.; Rosbash, M.; Griffith, L. PDF Cells Are a GABA-Responsive Wake-Promoting Component of the Drosophila Sleep Circuit. Neuron 2008, 60, 672–682. [Google Scholar] [CrossRef]
- Sheeba, V.; Gu, H.; Sharma, V.K.; O’Dowd, D.K.; Holmes, T.C. Circadian- and light-dependent regulation of resting membrane potential and spontaneous action potential firing of Drosophila circadian pacemaker neurons. J. Neurophysiol. 2008, 99, 976–988. [Google Scholar] [CrossRef]
- Chung, B.Y.; Kilman, V.L.; Keath, J.R.; Pitman, J.L.; Allada, R. The GABAA Receptor RDL Acts in Peptidergic PDF Neurons to Promote Sleep in Drosophila. Curr. Biol. 2009, 19, 386–390. [Google Scholar] [CrossRef] [PubMed]
- Murad, A.; Emery-Le, M.; Emery, P. A subset of dorsal neurons modulates circadian behavior and light responses in Drosophila. Neuron 2007, 53, 689–701. [Google Scholar] [CrossRef] [PubMed]
- Stoleru, D.; Nawathean, P.; de la Paz Fernández, M.; Menet, J.S.; Ceriani, M.F.; Rosbash, M. The Drosophila Circadian Network Is a Seasonal Timer. Cell 2007, 129, 207–219. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Liu, Y.; Bilodeau-Wentworth, D.; Hardin, P.E.; Emery, P. Light and Temperature Control the Contribution of Specific DN1 Neurons to Drosophila Circadian Behavior. Curr. Biol. 2010, 20, 600–605. [Google Scholar] [CrossRef]
- Kaneko, H.; Head, L.M.; Ling, J.; Tang, X.; Liu, Y.; Hardin, P.E.; Emery, P.; Hamada, F.N. Circadian Rhythm of Temperature Preference and Its Neural Control in Drosophila. Curr. Biol. 2012, 22, 1851–1857. [Google Scholar] [CrossRef]
- Cavanaugh, D.J.; Geratowski, J.D.; Wooltorton, J.R.A.; Spaethling, J.M.; Hector, C.E.; Zheng, X.; Johnson, E.C.; Eberwine, J.H.; Sehgal, A. Identification of a Circadian Output Circuit for Rest:Activity Rhythms in Drosophila. Cell 2014, 157, 689–701. [Google Scholar] [CrossRef]
- Díaz, M.M.; Schlichting, M.; Abruzzi, K.C.; Long, X.; Rosbash, M. Allatostatin-C/AstC-R2 Is a Novel Pathway to Modulate the Circadian Activity Pattern in Drosophila. Curr. Biol. 2019, 29, 13–22.e3. [Google Scholar] [CrossRef] [PubMed]
- Ni, J.D.; Gurav, A.S.; Liu, W.; Ogunmowo, T.H.; Hackbart, H.; Elsheikh, A.; Verdegaal, A.A.; Montell, C. Differential regulation of the Drosophila sleep homeostat by circadian and arousal inputs. Elife 2019, 8, e40487. [Google Scholar] [CrossRef] [PubMed]
- Moore, R.Y.; Speh, J.C.; Card, J.P. The retinohypothalamic tract originates from a distinct subset of retinal ganglion cells. J. Comp. Neurol. 1995, 352, 351–366. [Google Scholar] [CrossRef] [PubMed]
- Castel, M.; Belenky, M.; Cohen, S.; Ottersen, O.P.; Storm-Mathisen, J. Glutamate-like immunoreactivity in retinal terminals of the mouse suprachiasmatic nucleus. Eur. J. Neurosci. 1993, 5, 368–381. [Google Scholar] [CrossRef]
- De Vries, M.J.; Nunes Cardozo, B.; van der Want, J.; de Wolf, A.; Meijer, J.H. Glutamate immunoreactivity in terminals of the retinohypothalamic tract of the brown Norwegian rat. Brain Res. 1993, 612, 231–237. [Google Scholar] [CrossRef]
- Harrington, M.E.; Hoque, S.; Hall, A.; Golombek, D.; Biello, S. Pituitary adenylate cyclase activating peptide phase shifts circadian rhythms in a manner similar to light. J. Neurosci. 1999, 19, 6637–6642. [Google Scholar] [CrossRef]
- Chen, D.; Buchanan, G.F.; Ding, J.M.; Hannibal, J.; Gillette, M.U. Pituitary adenylyl cyclase-activating peptide: A pivotal modulator of glutamatergic regulation of the suprachiasmatic circadian clock. Proc. Natl. Acad. Sci. USA 1999, 96, 13468–13473. [Google Scholar] [CrossRef]
- Hannibal, J.; Møller, M.; Ottersen, O.P.; Fahrenkrug, J. PACAP and glutamate are co-stored in the retinohypothalamic tract. J. Comp. Neurol. 2000, 418, 147–155. [Google Scholar] [CrossRef]
- Nielsen, H.S.; Hannibal, J.; Knudsen, S.M.; Fahrenkrug, J. Pituitary adenylate cyclase-activating polypeptide induces period1 and period2 gene expression in the rat suprachiasmatic nucleus during late night. Neuroscience 2001, 103, 433–441. [Google Scholar] [CrossRef]
- Jin, X.; Shearman, L.P.; Weaver, D.R.; Zylka, M.J.; de Vries, G.J.; Reppert, S.M. A molecular mechanism regulating rhythmic output from the suprachiasmatic circadian clock. Cell 1999, 96, 57–68. [Google Scholar] [CrossRef]
- Yoshii, T.; Hermann-Luibl, C.; Helfrich-Förster, C. Circadian light-input pathways in Drosophila. Commun. Integr. Biol. 2016, 9, e1102805. [Google Scholar] [CrossRef]
- Li, M.-T.; Cao, L.-H.; Xiao, N.; Tang, M.; Deng, B.; Yang, T.; Yoshii, T.; Luo, D.-G. Hub-organized parallel circuits of central circadian pacemaker neurons for visual photoentrainment in Drosophila. Nat. Commun. 2018, 9, 4247. [Google Scholar] [CrossRef]
- Schlichting, M.; Menegazzi, P.; Rosbash, M.; Helfrich-Förster, C. A distinct visual pathway mediates high light intensity adaptation of the circadian clock in Drosophila. J. Neurosci. 2019, 39, 1621–1630. [Google Scholar] [CrossRef]
- Alejevski, F.; Saint-Charles, A.; Michard-Vanhée, C.; Martin, B.; Galant, S.; Vasiliauskas, D.; Rouyer, F. The HisCl1 histamine receptor acts in photoreceptors to synchronize Drosophila behavioral rhythms with light-dark cycles. Nat. Commun. 2019, 10, 252. [Google Scholar] [CrossRef]
- Rieger, D.; Stanewsky, R.; Helfrich-Förster, C. Cryptochrome, Compound Eyes, Hofbauer-Buchner Eyelets, and Ocelli Play Different Roles in the Entrainment and Masking Pathway of the Locomotor Activity Rhythm in the Fruit Fly Drosophila Melanogaster. J. Biol. Rhythms 2003, 18, 377–391. [Google Scholar] [CrossRef]
- Benito, J.; Houl, J.H.; Roman, G.W.; Hardin, P.E. The Blue-Light Photoreceptor CRYPTOCHROME Is Expressed in a Subset of Circadian Oscillator Neurons in the Drosophila CNS. J. Biol. Rhythms 2008, 23, 296–307. [Google Scholar] [CrossRef]
- Yoshii, T.; Todo, T.; Wülbeck, C.; Stanewsky, R.; Helfrich-Förster, C. Cryptochrome is present in the compound eyes and a subset ofDrosophila’s clock neurons. J. Comp. Neurol. 2008, 508, 952–966. [Google Scholar] [CrossRef]
- Stanewsky, R.; Kaneko, M.; Emery, P.; Beretta, B.; Wager-Smith, K.; Kay, S.A.; Rosbash, M.; Hall, J.C. The cryb mutation identifies cryptochrome as a circadian photoreceptor in Drosophila. Cell 1998, 95, 681–692. [Google Scholar] [CrossRef]
- Koh, K. JETLAG Resets the Drosophila Circadian Clock by Promoting Light-Induced Degradation of TIMELESS. Science 2006, 312, 1809–1812. [Google Scholar] [CrossRef]
- Yoo, S.-H.; Yamazaki, S.; Lowrey, P.L.; Shimomura, K.; Ko, C.H.; Buhr, E.D.; Siepka, S.M.; Hong, H.-K.; Oh, W.J.; Yoo, O.J.; et al. PERIOD2::LUCIFERASE real-time reporting of circadian dynamics reveals persistent circadian oscillations in mouse peripheral tissues. Proc. Natl. Acad. Sci. USA 2004, 101, 5339–5346. [Google Scholar] [CrossRef]
- Schibler, U.; Gotic, I.; Saini, C.; Gos, P.; Curie, T.; Emmenegger, Y.; Sinturel, F.; Gosselin, P.; Gerber, A.; Fleury-Olela, F.; et al. Clock-Talk: Interactions between Central and Peripheral Circadian Oscillators in Mammals. Cold Spring Harb. Symp. Quant. Biol. 2015, 80, 223–232. [Google Scholar] [CrossRef]
- Damiola, F.; Le Minh, N.; Preitner, N.; Kornmann, B.; Fleury-Olela, F.; Schibler, U. Restricted feeding uncouples circadian oscillators in peripheral tissues from the central pacemaker in the suprachiasmatic nucleus. Genes Dev. 2000, 14, 2950–2961. [Google Scholar] [CrossRef]
- Stokkan, K.-A.; Yamazaki, S.; Tei, H.; Sakaki, Y.; Menaker, M. Entrainment of the Circadian Clock in the Liver by Feeding. Science 2001, 291, 490–493. [Google Scholar] [CrossRef]
- Honma, S.; Honma, K.; Shirakawa, T.; Hiroshige, T. Rhythms in behaviors, body temperature and plasma corticosterone in SCN lesioned rats given methamphetamine. Physiol. Behav. 1988, 44, 247–255. [Google Scholar] [CrossRef]
- Levine, J.D.; Funes, P.; Dowse, H.B.; Hall, J.C. Advanced analysis of a cryptochrome mutation’s effects on the robustness and phase of molecular cycles in isolated peripheral tissues of Drosophila. BMC Neurosci. 2002, 3, 5. [Google Scholar]
- Ito, C.; Tomioka, K. Heterogeneity of the Peripheral Circadian Systems in Drosophila melanogaster: A Review. Front. Physiol. 2016, 7, 8. [Google Scholar] [CrossRef]
- Plautz, J.D.; Kaneko, M.; Hall, J.C.; Kay, S.A. Independent photoreceptive circadian clocks throughout Drosophila. Science 1997, 278, 1632–1635. [Google Scholar] [CrossRef] [PubMed]
- Xu, K.; DiAngelo, J.R.; Hughes, M.E.; Hogenesch, J.B.; Sehgal, A. The circadian clock interacts with metabolic physiology to influence reproductive fitness. Cell Metab. 2011, 13, 639–654. [Google Scholar] [CrossRef]
- Giebultowicz, J.M.; Hege, D.M. Circadian clock in Malpighian tubules. Nature 1997, 386, 664. [Google Scholar] [CrossRef] [PubMed]
- Myers, E.M.; Yu, J.; Sehgal, A. Circadian control of eclosion: Interaction between a central and peripheral clock in Drosophila melanogaster. Curr. Biol. 2003, 13, 526–533. [Google Scholar] [CrossRef]
- Krupp, J.J.; Billeter, J.-C.; Wong, A.; Choi, C.; Nitabach, M.N.; Levine, J.D. Pigment-dispersing factor modulates pheromone production in clock cells that influence mating in drosophila. Neuron 2013, 79, 54–68. [Google Scholar] [CrossRef]
- Gannon, R.L.; Rea, M.A. In situ hybridization of antisense mRNA oligonucleotides for AMPA, NMDA and metabotropic glutamate receptor subtypes in the rat suprachiasmatic nucleus at different phases of the circadian cycle. Brain Res. Mol. Brain Res. 1994, 23, 338–344. [Google Scholar] [CrossRef]
- Ito, C.; Wakamori, M.; Akaike, N. Dual effect of glycine on isolated rat suprachiasmatic neurons. Am. J. Physiol. 1991, 260, C213–C218. [Google Scholar] [CrossRef]
- van den Pol, A.N.; Hermans-Borgmeyer, I.; Hofer, M.; Ghosh, P.; Heinemann, S. Ionotropic glutamate-receptor gene expression in hypothalamus: Localization of AMPA, kainate, and NMDA receptor RNA with in situ hybridization. J. Comp. Neurol. 1994, 343, 428–444. [Google Scholar] [CrossRef]
- Bos, N.P.A.; Mirmiran, M. Effects of excitatory and inhibitory amino acids on neuronal discharges in the cultured suprachiasmatic nucleus. Brain Res. Bull. 1993, 31, 67–72. [Google Scholar] [CrossRef]
- Liou, S.Y.; Shibata, S.; Iwasaki, K.; Ueki, S. Optic nerve stimulation-induced increase of release of 3H-glutamate and 3H-aspartate but not 3H-GABA from the suprachiasmatic nucleus in slices of rat hypothalamus. Brain Res. Bull. 1986, 16, 527–531. [Google Scholar] [CrossRef]
- Cahill, G.M.; Menaker, M. Effects of excitatory amino acid receptor antagonists and agonists on suprachiasmatic nucleus responses to retinohypothalamic tract volleys. Brain Res. 1989, 479, 76–82. [Google Scholar] [CrossRef]
- Shibata, S.; Watanabe, A.; Hamada, T.; Ono, M.; Watanabe, S. N-methyl-D-aspartate induces phase shifts in circadian rhythm of neuronal activity of rat SCN in vitro. Am. J. Physiol. 1994, 267, R360–R364. [Google Scholar] [CrossRef] [PubMed]
- Colwell, C.S.; Menaker, M. NMDA as well as non-NMDA receptor antagonists can prevent the phase-shifting effects of light on the circadian system of the golden hamster. J. Biol. Rhythms 1992, 7, 125–136. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, P.K.; Baskaran, N.; van den Pol, A.N. Developmentally regulated gene expression of all eight metabotropic glutamate receptors in hypothalamic suprachiasmatic and arcuate nuclei—A PCR analysis. Brain Res. Dev. Brain Res. 1997, 102, 1–12. [Google Scholar] [CrossRef]
- Scott, G.; Rusak, B. Activation of hamster suprachiasmatic neurons in vitro via metabotropic glutamate receptors. Neuroscience 1996, 71, 533–541. [Google Scholar] [CrossRef]
- Chen, G.; van den Pol, A.N. Coexpression of Multiple Metabotropic Glutamate Receptors in Axon Terminals of Single Suprachiasmatic Nucleus Neurons. J. Neurophysiol. 1998, 80, 1932–1938. [Google Scholar] [CrossRef]
- Brancaccio, M.; Patton, A.P.; Chesham, J.E.; Maywood, E.S.; Hastings, M.H. Astrocytes Control Circadian Timekeeping in the Suprachiasmatic Nucleus via Glutamatergic Signaling. Neuron 2017, 93, 1420–1435.e5. [Google Scholar] [CrossRef] [PubMed]
- Colwell, C.S. NMDA-evoked calcium transients and currents in the suprachiasmatic nucleus: Gating by the circadian system. Eur. J. Neurosci. 2001, 13, 1420–1428. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.Y.; Choi, H.J.; Kim, J.S.; Kim, Y.S.; Jeong, D.U.; Shin, H.C.; Kim, M.J.; Han, H.; Hong, S.K.; Kim, Y.I. Voltage-gated calcium channels play crucial roles in the glutamate-induced phase shifts of the rat suprachiasmatic circadian clock. Eur. J. Neurosci. 2005, 21, 1215–1222. [Google Scholar] [CrossRef] [PubMed]
- Ding, J.M.; Buchanan, G.F.; Tischkau, S.A.; Chen, D.; Kuriashkina, L.; Faiman, L.E.; Alster, J.M.; McPherson, P.S.; Campbell, K.P.; Gillette, M.U. A neuronal ryanodine receptor mediates light-induced phase delays of the circadian clock. Nature 1998, 394, 381–384. [Google Scholar] [CrossRef] [PubMed]
- Tischkau, S.A.; Gallman, E.A.; Buchanan, G.F.; Gillette, M.U. Differential cAMP gating of glutamatergic signaling regulates long-term state changes in the suprachiasmatic circadian clock. J. Neurosci. 2000, 20, 7830–7837. [Google Scholar] [CrossRef] [PubMed]
- Tischkau, S.A.; Weber, E.T.; Abbott, S.M.; Mitchell, J.W.; Gillette, M.U. Circadian clock-controlled regulation of cGMP-protein kinase G in the nocturnal domain. J. Neurosci. 2003, 23, 7543–7550. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Ding, J.M.; Chen, D.; Weber, E.T.; Faiman, L.E.; Rea, M.A.; Gillette, M.U. Resetting the biological clock: Mediation of nocturnal circadian shifts by glutamate and NO. Science 1994, 266, 1713–1717. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, A.; Hamada, T.; Shibata, S.; Watanabe, S. Effects of nitric oxide synthase inhibitors on N-methyl-D-aspartate-induced phase delay of circadian rhythm of neuronal activity in the rat suprachiasmatic nucleus in vitro. Brain Res. 1994, 646, 161–164. [Google Scholar] [CrossRef]
- Kim, Y.I.; Choi, H.-J.; Colwell, C.S. Brain-derived neurotrophic factor regulation of N-methyl-D-aspartate receptor-mediated synaptic currents in suprachiasmatic nucleus neurons. J. Neurosci. Res. 2006, 84, 1512–1520. [Google Scholar] [CrossRef]
- Michel, S.; Clark, J.P.; Ding, J.M.; Colwell, C.S. Brain-derived neurotrophic factor and neurotrophin receptors modulate glutamate-induced phase shifts of the suprachiasmatic nucleus. Eur. J. Neurosci. 2006, 24, 1109–1116. [Google Scholar] [CrossRef] [PubMed]
- Kopp, M.D.; Schomerus, C.; Dehghani, F.; Korf, H.W.; Meissl, H. Pituitary adenylate cyclase-activating polypeptide and melatonin in the suprachiasmatic nucleus: Effects on the calcium signal transduction cascade. J. Neurosci. 1999, 19, 206–219. [Google Scholar] [CrossRef]
- Dziema, H.; Obrietan, K. PACAP Potentiates L-Type Calcium Channel Conductance in Suprachiasmatic Nucleus Neurons by Activating the MAPK Pathway. J. Neurophysiol. 2002, 88, 1374–1386. [Google Scholar] [CrossRef] [PubMed]
- Kopp, M.D.A.; Meissl, H.; Dehghani, F.; Korf, H.W. The pituitary adenylate cyclase-activating polypeptide modulates glutamatergic calcium signalling: Investigations on rat suprachiasmatic nucleus neurons. J. Neurochem. 2001, 79, 161–171. [Google Scholar] [CrossRef] [PubMed]
- Hannibal, J.; Mikkelsen, J.D.; Clausen, H.; Holst, J.J.; Wulff, B.S.; Fahrenkrug, J. Gene expression of pituitary adenylate cyclase activating polypeptide (PACAP) in the rat hypothalamus. Regul. Pept. 1995, 55, 133–148. [Google Scholar] [CrossRef]
- Hannibal, J.; Ding, J.M.; Chen, D.; Fahrenkrug, J.; Larsen, P.J.; Gillette, M.U.; Mikkelsen, J.D. Pituitary Adenylate Cyclase-Activating Peptide (PACAP) in the Retinohypothalamic Tract: A Potential Daytime Regulator of the Biological Clock. J. Neurosci. 1997, 17, 2637–2644. [Google Scholar] [CrossRef]
- Spengler, D.; Waeber, C.; Pantaloni, C.; Holsboer, F.; Bockaert, J.; Seeburg, P.H.; Journot, L. Differential signal transduction by five splice variants of the PACAP receptor. Nature 1993, 365, 170–175. [Google Scholar]
- Harmar, A.J.; Arimura, A.; Gozes, I.; Journot, L.; Laburthe, M.; Pisegna, J.R.; Rawlings, S.R.; Robberecht, P.; Said, S.I.; Sreedharan, S.P.; et al. International Union of Pharmacology. XVIII. Nomenclature of receptors for vasoactive intestinal peptide and pituitary adenylate cyclase-activating polypeptide. Pharmacol. Rev. 1998, 50, 265–270. [Google Scholar]
- Von Gall, C.; Duffield, G.E.; Hastings, M.H.; Kopp, M.D.; Dehghani, F.; Korf, H.W.; Stehle, J.H. CREB in the mouse SCN: A molecular interface coding the phase-adjusting stimuli light, glutamate, PACAP, and melatonin for clockwork access. J. Neurosci. 1998, 18, 10389–10397. [Google Scholar] [CrossRef]
- Hannibal, J.; Jamen, F.; Nielsen, H.S.; Journot, L.; Brabet, P.; Fahrenkrug, J. Dissociation between Light-Induced Phase Shift of the Circadian Rhythm and Clock Gene Expression in Mice Lacking the Pituitary Adenylate Cyclase Activating Polypeptide Type 1 Receptor. J. Neurosci. 2001, 21, 4883–4890. [Google Scholar] [CrossRef]
- Hannibal, J.; Brabet, P.; Fahrenkrug, J. Mice lacking the PACAP type I receptor have impaired photic entrainment and negative masking. Am. J. Physiol. Integr. Comp. Physiol. 2008, 295, R2050–R2058. [Google Scholar] [CrossRef]
- Kawaguchi, C.; Tanaka, K.; Isojima, Y.; Shintani, N.; Hashimoto, H.; Baba, A.; Nagai, K. Changes in light-induced phase shift of circadian rhythm in mice lacking PACAP. Biochem. Biophys. Res. Commun. 2003, 310, 169–175. [Google Scholar] [CrossRef]
- Colwell, C.S.; Michel, S.; Itri, J.; Rodriguez, W.; Tam, J.; Lelièvre, V.; Hu, Z.; Waschek, J.A. Selective deficits in the circadian light response in mice lacking PACAP. Am. J. Physiol. Integr. Comp. Physiol. 2004, 287, R1194–R1201. [Google Scholar] [CrossRef]
- Moore, R.Y.; Speh, J.C. GABA is the principal neurotransmitter of the circadian system. Neurosci. Lett. 1993, 150, 112–116. [Google Scholar] [CrossRef]
- Gao, B.; Fritschy, J.M.; Moore, R.Y. GABAA-receptor subunit composition in the circadian timing system. Brain Res. 1995, 700, 142–156. [Google Scholar] [CrossRef]
- Buijs, R.M.; Hou, Y.-X.; Shinn, S.; Renaud, L.P. Ultrastructural evidence for intra- and extranuclear projections of GABAergic neurons of the suprachiasmatic nucleus. J. Comp. Neurol. 1994, 340, 381–391. [Google Scholar] [CrossRef] [PubMed]
- Gillespie, C.F.; Mintz, E.M.; Marvel, C.L.; Huhman, K.L.; Albers, H.E. GABA(A) and GABA(B) agonists and antagonists alter the phase-shifting effects of light when microinjected into the suprachiasmatic region. Brain Res. 1997, 759, 181–189. [Google Scholar] [CrossRef]
- Jiang, Z.G.; Allen, C.N.; North, R.A. Presynaptic inhibition by baclofen of retinohypothalamic excitatory synaptic transmission in rat suprachiasmatic nucleus. Neuroscience 1995, 64, 813–819. [Google Scholar] [CrossRef]
- Liou, S.Y.; Shibata, S.; Albers, H.E.; Ueki, S. Effects of GABA and anxiolytics on the single unit discharge of suprachiasmatic neurons in rat hypothalamic slices. Brain Res. Bull. 1990, 25, 103–107. [Google Scholar] [CrossRef]
- Tominaga, K.; Shibata, S.; Hamada, T.; Watanabe, S. GABAA receptor agonist muscimol can reset the phase of neural activity rhythm in the rat suprachiasmatic nucleus in vitro. Neurosci. Lett. 1994, 166, 81–84. [Google Scholar] [CrossRef]
- Strecker, G.J.; Wuarin, J.P.; Dudek, F.E. GABAA-mediated local synaptic pathways connect neurons in the rat suprachiasmatic nucleus. J. Neurophysiol. 1997, 78, 2217–2220. [Google Scholar] [CrossRef] [PubMed]
- Wagner, S.; Sagiv, N.; Yarom, Y. GABA-induced current and circadian regulation of chloride in neurones of the rat suprachiasmatic nucleus. J. Physiol. 2001, 537, 853–869. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Reppert, S.M. GABA synchronizes clock cells within the suprachiasmatic circadian clock. Neuron 2000, 25, 123–128. [Google Scholar] [CrossRef]
- Liou, S.Y.; Albers, H.E. Single unit response of neurons within the hamster suprachiasmatic nucleus to GABA and low chloride perfusate during the day and night. Brain Res. Bull. 1990, 25, 93–98. [Google Scholar] [CrossRef]
- Aton, S.J.; Huettner, J.E.; Straume, M.; Herzog, E.D. GABA and Gi/o differentially control circadian rhythms and synchrony in clock neurons. Proc. Natl. Acad. Sci. USA 2006, 103, 19188–19193. [Google Scholar] [CrossRef] [PubMed]
- Bergeron, H.E.; Danielson, B.; Biggs, K.R.; Prosser, R.A. TTX blocks baclofen-induced phase shifts of the mammalian circadian pacemaker in vitro. Brain Res. 1999, 841, 193–196. [Google Scholar] [CrossRef]
- Mintz, E.M.; Jasnow, A.M.; Gillespie, C.F.; Huhman, K.L.; Albers, H.E. GABA interacts with photic signaling in the suprachiasmatic nucleus to regulate circadian phase shifts. Neuroscience 2002, 109, 773–778. [Google Scholar] [CrossRef]
- Smith, R.D.; Inouye, S.; Turek, F.W. Central administration of muscimol phase-shifts the mammalian circadian clock. J. Comp. Physiol. A. 1989, 164, 805–814. [Google Scholar] [CrossRef]
- Gillespie, C.F.; Van Der Beek, E.M.; Mintz, E.M.; Mickley, N.C.; Jasnow, A.M.; Huhman, K.L.; Albers, H.E. GABAergic regulation of light-induced c-Fos immunoreactivity within the suprachiasmatic nucleus. J. Comp. Neurol. 1999, 411, 683–692. [Google Scholar] [CrossRef]
- Hummer, D.L.; Ehlen, J.C.; Larkin, T.E.; McNeill, J.K.; Pamplin, J.R.; Walker, C.A.; Walker, P.V.; Dhanraj, D.R.; Albers, H.E. Sustained activation of GABAA receptors in the suprachiasmatic nucleus mediates light-induced phase delays of the circadian clock: A novel function of ionotropic receptors. Eur. J. Neurosci. 2015, 42, 1830–1838. [Google Scholar] [CrossRef]
- De Jeu, M.; Pennartz, C. Circadian modulation of GABA function in the rat suprachiasmatic nucleus: Excitatory effects during the night phase. J. Neurophysiol. 2002, 87, 834–844. [Google Scholar] [CrossRef][Green Version]
- Belenky, M.A.; Sollars, P.J.; Mount, D.B.; Alper, S.L.; Yarom, Y.; Pickard, G.E. Cell-type specific distribution of chloride transporters in the rat suprachiasmatic nucleus. Neuroscience 2010, 165, 1519–1537. [Google Scholar] [CrossRef]
- Albus, H.; Vansteensel, M.J.; Michel, S.; Block, G.D.; Meijer, J.H. A GABAergic Mechanism Is Necessary for Coupling Dissociable Ventral and Dorsal Regional Oscillators within the Circadian Clock. Curr. Biol. 2005, 15, 886–893. [Google Scholar] [CrossRef]
- McNeill, J.K.; Walton, J.C.; Albers, H.E. Functional Significance of the Excitatory Effects of GABA in the Suprachiasmatic Nucleus. J. Biol. Rhythms 2018, 33, 376–387. [Google Scholar] [CrossRef]
- Olde Engberink, A.H.O.; Meijer, J.H.; Michel, S. Chloride cotransporter KCC2 is essential for GABAergic inhibition in the SCN. Neuropharmacology 2018, 138, 80–86. [Google Scholar] [CrossRef]
- Morin, L.P.; Allen, C.N. The circadian visual system, 2005. Brain Res. Rev. 2006, 51, 1–60. [Google Scholar] [CrossRef]
- Sumner, B.E.H.; Rosie, R.; Fink, G. Relative density of 5-hydroxytryptamine receptor subtype mRNAs in female rat neuroendocrine brain determined by in situ hybridization histochemistry. Mol. Cell. Neurosci. 1992, 3, 215–223. [Google Scholar] [CrossRef]
- Belenky, M.A.; Pickard, G.E. Subcellular Distribution of 5-HT 1B and 5-HT 7 Receptors in the Mouse. J. Comp. Neurol. 2001, 388, 371–388. [Google Scholar] [CrossRef]
- Takeuchi, K.; Mohammad, S.; Ozaki, T.; Morioka, E.; Kawaguchi, K.; Kim, J.; Jeong, B.; Hong, J.H.; Lee, K.J.; Ikeda, M. Serotonin-2C receptor involved serotonin-induced Ca2+ mobilisations in neuronal progenitors and neurons in rat suprachiasmatic nucleus. Sci. Rep. 2014, 4, 4106. [Google Scholar] [CrossRef]
- Dudley, T.E.; DiNardo, L.A.; Glass, J.D. Endogenous regulation of serotonin release in the hamster suprachiasmatic nucleus. J. Neurosci. 1998, 18, 5045–5052. [Google Scholar] [CrossRef]
- Cuesta, M.; Clesse, D.; Pévet, P.; Challet, E. New light on the serotonergic paradox in the rat circadian system. J. Neurochem. 2009, 110, 231–243. [Google Scholar] [CrossRef]
- Mason, R. Circadian variation in sensitivity of suprachiasmatic and lateral geniculate neurones to 5-hydroxytryptamine in the rat. J. Physiol. 1986, 377, 1–13. [Google Scholar] [CrossRef]
- Nishino, H.; Koizumi, K. Responses of neurons in the suprachiasmatic nuclei of the hypothalamus to putative transmitters. Brain Res. 1977, 120, 167–172. [Google Scholar] [CrossRef]
- Shibata, S.; Liou, S.Y.; Ueki, S. Different effects of amino acids, acetylcholine and monoamines on neuronal activity of suprachiasmatic nucleus in rat pups and adults. Neurosci. Lett. 1983, 39, 187–192. [Google Scholar] [CrossRef]
- Miller, J.D.; Fuller, C.A. The response of suprachiasmatic neurons of the rat hypothalamus to photic and serotonergic stimulation. Brain Res. 1990, 515, 155–162. [Google Scholar] [CrossRef]
- Ying, S.W.; Rusak, B. Effects of serotonergic agonists on firing rates of photically responsive cells in the hamster suprachiasmatic nucleus. Brain Res. 1994, 651, 37–46. [Google Scholar] [CrossRef]
- Kawahara, F.; Saito, H.; Katsuki, H. Inhibition by 5-HT7 receptor stimulation of GABAA receptor-activated current in cultured rat suprachiasmatic neurones. J. Physiol. 1994, 478 Pt 1, 67–73. [Google Scholar] [CrossRef]
- Pickard, G.E.; Smith, B.N.; Belenky, M.; Rea, M.A.; Dudek, F.E.; Sollars, P.J. 5-HT1B receptor-mediated presynaptic inhibition of retinal input to the suprachiasmatic nucleus. J. Neurosci. 1999, 19, 4034–4045. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Z.G.; Teshima, K.; Yang, Y.; Yoshioka, T.; Allen, C.N. Pre- and postsynaptic actions of serotonin on rat suprachiasmatic nucleus neurons. Brain Res. 2000, 866, 247–256. [Google Scholar] [CrossRef]
- Bramley, J.R.; Sollars, P.J.; Pickard, G.E.; Dudek, F.E. 5-HT 1B Receptor-Mediated Presynaptic Inhibition of GABA Release in the Suprachiasmatic Nucleus. J. Neurophysiol. 2005, 93, 3157–3164. [Google Scholar] [CrossRef] [PubMed]
- Ying, S.W.; Rusak, B. 5-HT7 receptors mediate serotonergic effects on light-sensitive suprachiasmatic nucleus neurons. Brain Res. 1997, 755, 246–254. [Google Scholar] [CrossRef]
- Liou, S.Y.; Shibata, S.; Ueki, S. Effect of monoamines on field potentials in the suprachiasmatic nucleus of slices of hypothalamus of the rat evoked by stimulation of the optic nerve. Neuropharmacology 1986, 25, 1009–1014. [Google Scholar] [CrossRef]
- Rea, M.A.; Glass, J.D.; Colwell, C.S. Serotonin modulates photic responses in the hamster suprachiasmatic nuclei. J. Neurosci. 1994, 14, 3635–3642. [Google Scholar] [CrossRef] [PubMed]
- Pickard, G.E.; Weber, E.T.; Scott, P.A.; Riberdy, A.F.; Rea, M.A. 5HT1B receptor agonists inhibit light-induced phase shifts of behavioral circadian rhythms and expression of the immediate-early gene c-fos in the suprachiasmatic nucleus. J. Neurosci. 1996, 16, 8208–8220. [Google Scholar] [CrossRef] [PubMed]
- Sollars, P.J.; Simpson, A.M.; Ogilvie, M.D.; Pickard, G.E. Light-induced Fos expression is attenuated in the suprachiasmatic nucleus of serotonin 1B receptor knockout mice. Neurosci. Lett. 2006, 401, 209–213. [Google Scholar] [CrossRef]
- Sollars, P.J.; Ogilvie, M.D.; Simpson, A.M.; Pickard, G.E. Photic entrainment is altered in the 5-HT1B receptor knockout mouse. J. Biol. Rhythms 2006, 21, 21–32. [Google Scholar] [CrossRef] [PubMed]
- Smith, V.M.; Sterniczuk, R.; Phillips, C.I.; Antle, M.C. Altered photic and non-photic phase shifts in 5-HT(1A) receptor knockout mice. Neuroscience 2008, 157, 513–523. [Google Scholar] [CrossRef]
- Paulus, E.V.; Mintz, E.M. Photic and nonphotic responses of the circadian clock in serotonin-deficient Pet-1 knockout mice. Chronobiol. Int. 2013, 30, 1251–1260. [Google Scholar] [CrossRef]
- Paulus, E.V.; Mintz, E.M. Developmental disruption of the serotonin system alters circadian rhythms. Physiol. Behav. 2012, 105, 257–263. [Google Scholar] [CrossRef]
- Sprouse, J.; Li, X.; Stock, J.; McNeish, J.; Reynolds, L. Circadian rhythm phenotype of 5-HT7 receptor knockout mice: 5-HT and 8-OH-DPAT-induced phase advances of SCN neuronal firing. J. Biol. Rhythms 2005, 20, 122–131. [Google Scholar] [CrossRef]
- Prosser, R.A.; Lee, H.-M.; Wehner, A. Serotonergic pre-treatments block in vitro serotonergic phase shifts of the mouse suprachiasmatic nucleus circadian clock. Neuroscience 2006, 142, 547–555. [Google Scholar] [CrossRef]
- Horikawa, K.; Shibata, S. Phase-resetting response to (+)8-OH-DPAT, a serotonin 1A/7 receptor agonist, in the mouse in vivo. Neurosci. Lett. 2004, 368, 130–134. [Google Scholar] [CrossRef] [PubMed]
- Cutrera, R.; Saboureau, M.; Pévet, P. Phase-shifting effect of 8-OH-DPAT, a 5-HT1A/5-HT7 receptor agonist, on locomotor activity in golden hamster in constant darkness. Neurosci. Lett. 1996, 210, 1–4. [Google Scholar] [CrossRef]
- Horikawa, K.; Yokota, S.; Fuji, K.; Akiyama, M.; Moriya, T.; Okamura, H.; Shibata, S. Nonphotic entrainment by 5-HT1A/7 receptor agonists accompanied by reduced Per1 and Per2 mRNA levels in the suprachiasmatic nuclei. J. Neurosci. 2000, 20, 5867–5873. [Google Scholar] [CrossRef] [PubMed]
- Prosser, R.A.; Miller, J.D.; Heller, H.C. A serotonin agonist phase-shifts the circadian clock in the suprachiasmatic nuclei in vitro. Brain Res. 1990, 534, 336–339. [Google Scholar] [CrossRef]
- Kohler, M.; Kalkowski, A.; Wollnik, F. Serotonin agonist quipazine induces photic-like phase shifts of the circadian activity rhythm and c-Fos expression in the rat suprachiasmatic nucleus. J. Biol. Rhythms 1999, 14, 131–140. [Google Scholar] [CrossRef] [PubMed]
- Shibata, S.; Tsuneyoshi, A.; Hamada, T.; Tominaga, K.; Watanabe, S. Phase-resetting effect of 8-OH-DPAT, a serotonin1A receptor agonist, on the circadian rhythm of firing rate in the rat suprachiasmatic nuclei in vitro. Brain Res. 1992, 582, 353–356. [Google Scholar] [CrossRef]
- Prosser, R.A.; Heller, H.C.; Miller, J.D. Serotonergic phase shifts of the mammalian circadian clock: Effects of tetrodotoxin and high Mg2+. Brain Res. 1992, 573, 336–340. [Google Scholar] [CrossRef]
- Prosser, R.A.; Heller, H.C.; Miller, J.D. Serotonergic phase advances of the mammalian circadian clock involve protein kinase A and K+ channel opening. Brain Res. 1994, 644, 67–73. [Google Scholar] [CrossRef]
- Starkey, S.J. Melatonin and 5-hydroxytryptamine phase-advance the rat circadian clock by activation of nitric oxide synthesis. Neurosci. Lett. 1996, 211, 199–202. [Google Scholar] [CrossRef]
- Kennaway, D.J.; Moyer, R.W. Serotonin 5-HT2c agonists mimic the effect of light pulses on circadian rhythms. Brain Res. 1998, 806, 257–270. [Google Scholar] [CrossRef]
- Antle, M.C.; Ogilvie, M.D.; Pickard, G.E.; Mistlberger, R.E. Response of the mouse circadian system to serotonin 1A/2/7 agonists in vivo: Surprisingly little. J. Biol. Rhythms 2003, 18, 145–158. [Google Scholar] [CrossRef] [PubMed]
- Prosser, R.A. Serotonin phase-shifts the mouse suprachiasmatic circadian clock in vitro. Brain Res. 2003, 966, 110–115. [Google Scholar] [CrossRef]
- Sprouse, J.; Reynolds, L.; Li, X.; Braselton, J.; Schmidt, A. 8-OH-DPAT as a 5-HT7 agonist: Phase shifts of the circadian biological clock through increases in cAMP production. Neuropharmacology 2004, 46, 52–62. [Google Scholar] [CrossRef]
- Hanin, I.; Massarelli, R.; Costa, E. Acetylcholine concentrations in rat brain: Diurnal oscillation. Science 1970, 170, 341–342. [Google Scholar] [CrossRef]
- Mohan, C.; Radha, E. Circadian rhythms in the central cholinergic system in aging animals. Adv. Exp. Med. Biol. 1978, 108, 275–299. [Google Scholar]
- Murakami, N.; Takahashi, K.; Kawashima, K. Effect of light on the acetylcholine concentrations of the suprachiasmatic nucleus in the rat. Brain Res. 1984, 311, 358–360. [Google Scholar] [CrossRef][Green Version]
- Van Der Zee, E.A.; Streefland, C.; Strosberg, A.D.; Schröder, H.; Luiten, P.G. Colocalization of muscarinic and nicotinic receptors in cholinoceptive neurons of the suprachiasmatic region in young and aged rats. Brain Res. 1991, 542, 348–352. [Google Scholar] [CrossRef]
- Gannon, R.L.; Garcia, D.A.; Millan, M.J. Effects of systemically applied nAChRα7 agonists and antagonists on light-induced phase shifts of hamster circadian activity rhythms. Eur. Neuropsychopharmacol. 2014, 24, 964–973. [Google Scholar] [CrossRef]
- Wada, E.; Wada, K.; Boulter, J.; Deneris, E.; Heinemann, S.; Patrick, J.; Swanson, L.W. Distribution of alpha 2, alpha 3, alpha 4, and beta 2 neuronal nicotinic receptor subunit mRNAs in the central nervous system: A hybridization histochemical study in the rat. J. Comp. Neurol. 1989, 284, 314–335. [Google Scholar] [CrossRef]
- Yang, J.-J.; Wang, Y.-T.; Cheng, P.-C.; Kuo, Y.-J.; Huang, R.-C. Cholinergic modulation of neuronal excitability in the rat suprachiasmatic nucleus. J. Neurophysiol. 2010, 103, 1397–1409. [Google Scholar] [CrossRef]
- Earnest, D.J.; Turek, F.W. Neurochemical basis for the photic control of circadian rhythms and seasonal reproductive cycles: Role for acetylcholine. Proc. Natl. Acad. Sci. USA 1985, 82, 4277–4281. [Google Scholar] [CrossRef]
- O’Hara, B.F.; Edgar, D.M.; Cao, V.H.; Wiler, S.W.; Heller, H.C.; Kilduff, T.S.; Miller, J.D. Nicotine and nicotinic receptors in the circadian system. Psychoneuroendocrinology 1998, 23, 161–173. [Google Scholar] [CrossRef]
- Zatz, M.; Herkenham, M.A. Intraventricular carbachol mimics the phase-shifting effect of light on the circadian rhythm of wheel-running activity. Brain Res. 1981, 212, 234–238. [Google Scholar] [CrossRef]
- Trachsel, L.; Heller, H.C.; Miller, J.D. Nicotine phase-advances the circadian neuronal activity rhythm in rat suprachiasmatic nuclei explants. Neuroscience 1995, 65, 797–803. [Google Scholar] [CrossRef]
- Liu, C.; Gillette, M.U. Cholinergic regulation of the suprachiasmatic nucleus circadian rhythm via a muscarinic mechanism at night. J. Neurosci. 1996, 16, 744–751. [Google Scholar] [CrossRef] [PubMed]
- Pauly, J.R.; Horseman, N.D. Anticholinergic agents do not block light-induced circadian phase shifts. Brain Res. 1985, 348, 163–167. [Google Scholar] [CrossRef]
- Buchanan, G.F.; Gillette, M.U. New light on an old paradox: Site-dependent effects of carbachol on circadian rhythms. Exp. Neurol. 2005, 193, 489–496. [Google Scholar] [CrossRef]
- Basu, P.; Wensel, A.L.; McKibbon, R.; Lefebvre, N.; Antle, M.C. Activation of M1/4 receptors phase advances the hamster circadian clock during the day. Neurosci. Lett. 2016, 621, 22–27. [Google Scholar] [CrossRef]
- Liu, C.; Ding, J.M.; Faiman, L.E.; Gillette, M.U. Coupling of muscarinic cholinergic receptors and cGMP in nocturnal regulation of the suprachiasmatic circadian clock. J. Neurosci. 1997, 17, 659–666. [Google Scholar] [CrossRef] [PubMed]
- Ikeda, M.; Sugiyama, T.; Suzuki, K.; Moriya, T.; Shibata, S.; Katsuki, M.; Allen, C.N.; Yoshioka, T. PLC beta 4-independent Ca2+ rise via muscarinic receptors in the mouse suprachiasmatic nucleus. Neuroreport 2000, 11, 907–912. [Google Scholar] [CrossRef]
- Betz, H.; Laube, B. Glycine receptors: Recent insights into their structural organization and functional diversity. J. Neurochem. 2006, 97, 1600–1610. [Google Scholar] [CrossRef]
- Johnson, J.W.; Ascher, P. Glycine potentiates the NMDA response in cultured mouse brain neurons. Nature 1987, 325, 529–531. [Google Scholar] [CrossRef]
- Mahr, S. Anatomische Grundlagen der Übertragung circadianer Signale zwischen N. suprachiasmaticus und anderen Kerngebieten des Hypothalamus. Ph.D. Thesis, Johann Wolfgang Goethe-Universität, Frankfurt, Germany, 2008. [Google Scholar]
- Shinohara, K.; Honma, S.; Katsuno, Y.; Abe, H.; Honma, K. Circadian release of amino acids in the suprachiasmatic nucleus in vitro. Neuroreport 1998, 9, 137–140. [Google Scholar] [CrossRef] [PubMed]
- Mordel, J.; Karnas, D.; Inyushkin, A.; Challet, E.; Pévet, P.; Meissl, H. Activation of glycine receptor phase-shifts the circadian rhythm in neuronal activity in the mouse suprachiasmatic nucleus. J. Physiol. 2011, 589, 2287–2300. [Google Scholar] [CrossRef]
- Kawai, N.; Sakai, N.; Okuro, M.; Karakawa, S.; Tsuneyoshi, Y.; Kawasaki, N.; Takeda, T.; Bannai, M.; Nishino, S. The sleep-promoting and hypothermic effects of glycine are mediated by NMDA receptors in the suprachiasmatic nucleus. Neuropsychopharmacology 2015, 40, 1405–1416. [Google Scholar] [CrossRef]
- Aïoun, J.; Chambille, I.; Peytevin, J.; Martinet, L. Neurons containing gastrin-releasing peptide and vasoactive intestinal polypeptide are involved in the reception of the photic signal in the suprachiasmatic nucleus of the Syrian hamster: An immunocytochemical ultrastructural study. Cell Tissue Res. 1998, 291, 239–253. [Google Scholar] [CrossRef]
- Nielsen, H.S.; Hannibal, J.; Fahrenkrug, J. Vasoactive intestinal polypeptide induces per1 and per2 gene expression in the rat suprachiasmatic nucleus late at night. Eur. J. Neurosci. 2002, 15, 570–574. [Google Scholar] [CrossRef]
- Meyer-Spasche, A.; Piggins, H.D. Vasoactive intestinal polypeptide phase-advances the rat suprachiasmatic nuclei circadian pacemaker in vitro via protein kinase A and mitogen-activated protein kinase. Neurosci. Lett. 2004, 358, 91–94. [Google Scholar] [CrossRef]
- Harmar, A.J.; Marston, H.M.; Shen, S.; Spratt, C.; West, K.M.; Sheward, W.J.; Morrison, C.F.; Dorin, J.R.; Piggins, H.D.; Reubi, J.C.; et al. The VPAC(2) receptor is essential for circadian function in the mouse suprachiasmatic nuclei. Cell 2002, 109, 497–508. [Google Scholar] [CrossRef]
- Hughes, A.T.; Fahey, B.; Cutler, D.J.; Coogan, A.N.; Piggins, H.D. Aberrant gating of photic input to the suprachiasmatic circadian pacemaker of mice lacking the VPAC2 receptor. J. Neurosci. 2004, 24, 3522–3526. [Google Scholar] [CrossRef] [PubMed]
- Aton, S.J.; Colwell, C.S.; Harmar, A.J.; Waschek, J.; Herzog, E.D. Vasoactive intestinal polypeptide mediates circadian rhythmicity and synchrony in mammalian clock neurons. Nat. Neurosci. 2005, 8, 476–483. [Google Scholar] [CrossRef]
- Rea, M.A. VIP-stimulated cyclic AMP accumulation in the suprachiasmatic hypothalamus. Brain Res. Bull. 1990, 25, 843–847. [Google Scholar] [CrossRef]
- Irwin, R.P.; Allen, C.N. Neuropeptide-mediated calcium signaling in the suprachiasmatic nucleus network. Eur. J. Neurosci. 2010, 32, 1497–1506. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, M.; Hayashi, S.; Tamada, Y.; Ikeda, T.; Hisa, Y.; Takamatsu, T.; Ibata, Y. Direct retinal projections to GRP neurons in the suprachiasmatic nucleus of the rat. Neuroreport 1997, 8, 2187–2191. [Google Scholar] [CrossRef]
- McArthur, A.J.; Coogan, A.N.; Ajpru, S.; Sugden, D.; Biello, S.M.; Piggins, H.D. Gastrin-Releasing Peptide Phase-Shifts Suprachiasmatic Nuclei Neuronal Rhythms In Vitro. J. Neurosci. 2000, 20, 5496–5502. [Google Scholar] [CrossRef] [PubMed]
- Piggins, H.D.; Antle, M.C.; Rusak, B.; Coogan, A.N.; Piggins, H.D. Neuropeptides phase shift the mammalian circadian pacemaker. J. Neurosci. 1995, 15, 5612–5622. [Google Scholar] [CrossRef]
- Gamble, K.L.; Allen, G.C.; Zhou, T.; McMahon, D.G. Gastrin-releasing peptide mediates light-like resetting of the suprachiasmatic nucleus circadian pacemaker through cAMP response element-binding protein and Per1 activation. J. Neurosci. 2007, 27, 12078–12087. [Google Scholar] [CrossRef]
- Aida, R.; Moriya, T.; Araki, M.; Akiyama, M.; Wada, K.; Wada, E.; Shibata, S. Gastrin-releasing peptide mediates photic entrainable signals to dorsal subsets of suprachiasmatic nucleus via induction of Period gene in mice. Mol. Pharmacol. 2002, 61, 26–34. [Google Scholar] [CrossRef]
- Obrietan, K.; Impey, S.; Storm, D.R. Light and circadian rhythmicity regulate MAP kinase activation in the suprachiasmatic nuclei. Nat. Neurosci. 1998, 1, 693–700. [Google Scholar] [CrossRef]
- Coogan, A.N.; Piggins, H.D. Circadian and photic regulation of phosphorylation of ERK1/2 and Elk-1 in the suprachiasmatic nuclei of the Syrian hamster. J. Neurosci. 2003, 23, 3085–3093. [Google Scholar] [CrossRef]
- Ginty, D.D.; Kornhauser, J.M.; Thompson, M.A.; Bading, H.; Mayo, K.E.; Takahashi, J.S.; Greenberg, M.E. Regulation of CREB phosphorylation in the suprachiasmatic nucleus by light and a circadian clock. Science 1993, 260, 238–241. [Google Scholar] [CrossRef] [PubMed]
- Gau, D.; Lemberger, T.; von Gall, C.; Kretz, O.; Le Minh, N.; Gass, P.; Schmid, W.; Schibler, U.; Korf, H.W.; Schütz, G. Phosphorylation of CREB Ser142 regulates light-induced phase shifts of the circadian clock. Neuron 2002, 34, 245–253. [Google Scholar] [CrossRef]
- Obrietan, K.; Impey, S.; Smith, D.; Athos, J.; Storm, D.R. Circadian regulation of cAMP response element-mediated gene expression in the suprachiasmatic nuclei. J. Biol. Chem. 1999, 274, 17748–17756. [Google Scholar] [CrossRef]
- Travnickova-Bendova, Z.; Cermakian, N.; Reppert, S.M.; Sassone-Corsi, P. Bimodal regulation of mPeriod promoters by CREB-dependent signaling and CLOCK/BMAL1 activity. Proc. Natl. Acad. Sci. USA 2002, 99, 7728–7733. [Google Scholar] [CrossRef]
- Butcher, G.Q.; Doner, J.; Dziema, H.; Collamore, M.; Burgoon, P.W.; Obrietan, K. The p42/44 mitogen-activated protein kinase pathway couples photic input to circadian clock entrainment. J. Biol. Chem. 2002, 277, 29519–29525. [Google Scholar] [CrossRef]
- Butcher, G.Q.; Lee, B.; Hsieh, F.; Obrietan, K. Light- and clock-dependent regulation of ribosomal S6 kinase activity in the suprachiasmatic nucleus. Eur. J. Neurosci. 2004, 19, 907–915. [Google Scholar] [CrossRef] [PubMed]
- Butcher, G.Q.; Lee, B.; Cheng, H.-Y.M.; Obrietan, K. Light stimulates MSK1 activation in the suprachiasmatic nucleus via a PACAP-ERK/MAP kinase-dependent mechanism. J. Neurosci. 2005, 25, 5305–5313. [Google Scholar] [CrossRef] [PubMed]
- Cao, R.; Butcher, G.Q.; Karelina, K.; Arthur, J.S.; Obrietan, K. Mitogen- and stress-activated protein kinase 1 modulates photic entrainment of the suprachiasmatic circadian clock. Eur. J. Neurosci. 2013, 37, 130–140. [Google Scholar] [CrossRef] [PubMed]
- Cheng, H.-Y.M.; Obrietan, K.; Cain, S.W.; Lee, B.Y.; Agostino, P.V.; Joza, N.A.; Harrington, M.E.; Ralph, M.R.; Penninger, J.M. Dexras1 potentiates photic and suppresses nonphotic responses of the circadian clock. Neuron 2004, 43, 715–728. [Google Scholar] [CrossRef]
- Cheng, H.-Y.M.; Dziema, H.; Papp, J.; Mathur, D.P.; Koletar, M.; Ralph, M.R.; Penninger, J.M.; Obrietan, K. The molecular gatekeeper Dexras1 sculpts the photic responsiveness of the mammalian circadian clock. J. Neurosci. 2006, 26, 12984–12995. [Google Scholar] [CrossRef] [PubMed]
- Antoun, G.; Bouchard-Cannon, P.; Cheng, H.-Y.M. Regulation of MAPK/ERK Signaling and Photic Entrainment of the Suprachiasmatic Nucleus Circadian Clock by Raf Kinase Inhibitor Protein. J. Neurosci. 2012, 32, 4867–4877. [Google Scholar] [CrossRef]
- Yokota, S.; Yamamoto, M.; Moriya, T.; Akiyama, M.; Fukunaga, K.; Miyamoto, E.; Shibata, S. Involvement of calcium-calmodulin protein kinase but not mitogen-activated protein kinase in light-induced phase delays and Per gene expression in the suprachiasmatic nucleus of the hamster. J. Neurochem. 2001, 77, 618–627. [Google Scholar] [CrossRef]
- Yeung, K.; Seitz, T.; Li, S.; Janosch, P.; McFerran, B.; Kaiser, C.; Fee, F.; Katsanakis, K.D.; Rose, D.W.; Mischak, H.; et al. Suppression of Raf-1 kinase activity and MAP kinase signalling by RKIP. Nature 1999, 401, 173–177. [Google Scholar] [CrossRef] [PubMed]
- Cao, R.; Lee, B.; Cho, H.; Saklayen, S.; Obrietan, K. Photic regulation of the mTOR signaling pathway in the suprachiasmatic circadian clock. Mol. Cell. Neurosci. 2008, 38, 312–324. [Google Scholar] [CrossRef] [PubMed]
- Cao, R.; Anderson, F.E.; Jung, Y.-J.; Dziema, H.; Obrietan, K. Circadian regulation of mammalian target of rapamycin signaling in the mouse suprachiasmatic nucleus. Neuroscience 2011, 181, 79–88. [Google Scholar] [CrossRef]
- Cao, R.; Li, A.; Cho, H.; Lee, B.; Obrietan, K. Mammalian target of rapamycin signaling modulates photic entrainment of the suprachiasmatic circadian clock. J. Neurosci. 2010, 30, 6302–6314. [Google Scholar] [CrossRef]
- Cao, R.; Robinson, B.; Xu, H.; Gkogkas, C.; Khoutorsky, A.; Alain, T.; Yanagiya, A.; Nevarko, T.; Liu, A.C.; Amir, S.; et al. Translational control of entrainment and synchrony of the suprachiasmatic circadian clock by mTOR/4E-BP1 signaling. Neuron 2013, 79, 712–724. [Google Scholar] [CrossRef]
- Pizzio, G.A.; Hainich, E.C.; Ferreyra, G.A.; Coso, O.A.; Golombek, D.A. Circadian and photic regulation of ERK, JNK and p38 in the hamster SCN. Neuroreport 2003, 14, 1417–1419. [Google Scholar] [CrossRef]
- Yoshitane, H.; Honma, S.; Imamura, K.; Nakajima, H.; Nishide, S.; Ono, D.; Kiyota, H.; Shinozaki, N.; Matsuki, H.; Wada, N.; et al. JNK regulates the photic response of the mammalian circadian clock. EMBO Rep. 2012, 13, 455–461. [Google Scholar] [CrossRef]
- Nomura, K.; Takeuchi, Y.; Yamaguchi, S.; Okamura, H.; Fukunaga, K. Involvement of calcium/calmodulin-dependent protein kinase II in the induction of mPer1. J. Neurosci. Res. 2003, 72, 384–392. [Google Scholar] [CrossRef]
- Agostino, P.V.; Ferreyra, G.A.; Murad, A.D.; Watanabe, Y.; Golombek, D.A. Diurnal, circadian and photic regulation of calcium/calmodulin-dependent kinase II and neuronal nitric oxide synthase in the hamster suprachiasmatic nuclei. Neurochem. Int. 2004, 44, 617–625. [Google Scholar] [CrossRef]
- Golombek, D.A.; Ralph, M.R. KN-62, an inhibitor of Ca2+/calmodulin kinase II, attenuates circadian responses to light. Neuroreport 1994, 5, 1638–1640. [Google Scholar] [CrossRef]
- Fukushima, T.; Shimazoe, T.; Shibata, S.; Watanabe, A.; Ono, M.; Hamada, T.; Watanabe, S. The involvement of calmodulin and Ca2+/calmodulin-dependent protein kinase II in the circadian rhythms controlled by the suprachiasmatic nucleus. Neurosci. Lett. 1997, 227, 45–48. [Google Scholar] [CrossRef]
- Lee, J.M.; Schak, K.M.; Harrington, M.E. Inhibition of protein kinase A phase delays the mammalian circadian clock. Brain Res. 1999, 835, 350–353. [Google Scholar] [CrossRef]
- Kudo, T.; Tahara, Y.; Gamble, K.L.; McMahon, D.G.; Block, G.D.; Colwell, C.S. Vasoactive intestinal peptide produces long-lasting changes in neural activity in the suprachiasmatic nucleus. J. Neurophysiol. 2013, 110, 1097–1106. [Google Scholar] [CrossRef][Green Version]
- O’Neill, J.S.; Maywood, E.S.; Chesham, J.E.; Takahashi, J.S.; Hastings, M.H. cAMP-dependent signaling as a core component of the mammalian circadian pacemaker. Science 2008, 320, 949–953. [Google Scholar] [CrossRef]
- Sterniczuk, R.; Yamakawa, G.R.; Pomeroy, T.; Antle, M.C. Phase delays to light and gastrin-releasing peptide require the protein kinase A pathway. Neurosci. Lett. 2014, 559, 24–29. [Google Scholar] [CrossRef]
- Bonsall, D.R.; Lall, G.S. Protein kinase C differentially regulates entrainment of the mammalian circadian clock. Chronobiol. Int. 2013, 30, 460–469. [Google Scholar] [CrossRef]
- Jakubcakova, V.; Oster, H.; Tamanini, F.; Cadenas, C.; Leitges, M.; van der Horst, G.T.J.; Eichele, G. Light entrainment of the mammalian circadian clock by a PRKCA-dependent posttranslational mechanism. Neuron 2007, 54, 831–843. [Google Scholar] [CrossRef]
- Lee, B.; Almad, A.; Butcher, G.Q.; Obrietan, K. Protein kinase C modulates the phase-delaying effects of light in the mammalian circadian clock. Eur. J. Neurosci. 2007, 26, 451–462. [Google Scholar] [CrossRef]
- Albrecht, U.; Zheng, B.; Larkin, D.; Sun, Z.S.; Lee, C.C. MPer1 and mper2 are essential for normal resetting of the circadian clock. J. Biol. Rhythms 2001, 16, 100–104. [Google Scholar] [CrossRef]
- Yan, L.; Silver, R. Resetting the brain clock: Time course and localization of mPER1 and mPER2 protein expression in suprachiasmatic nuclei during phase shifts. Eur. J. Neurosci. 2004, 19, 1105–1109. [Google Scholar] [CrossRef]
- Weber, E.T.; Gannon, R.L.; Rea, M.A. cGMP-dependent protein kinase inhibitor blocks light-induced phase advances of circadian rhythms in vivo. Neurosci. Lett. 1995, 197, 227–230. [Google Scholar] [CrossRef]
- Mathur, A.; Golombek, D.A.; Ralph, M.R. cGMP-dependent protein kinase inhibitors block light-induced phase advances of circadian rhythms in vivo. Am. J. Physiol. Integr. Comp. Physiol. 1996, 270, R1031–R1036. [Google Scholar] [CrossRef]
- Ferreyra, G.A.; Golombek, D.A. Rhythmicity of the cGMP-related signal transduction pathway in the mammalian circadian system. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2001, 280, R1348–R1355. [Google Scholar] [CrossRef]
- Agostino, P.V.; Plano, S.A.; Golombek, D.A. Sildenafil accelerates reentrainment of circadian rhythms after advancing light schedules. Proc. Natl. Acad. Sci. USA 2007, 104, 9834–9839. [Google Scholar] [CrossRef]
- Jagannath, A.; Butler, R.; Godinho, S.I.H.; Couch, Y.; Brown, L.A.; Vasudevan, S.R.; Flanagan, K.C.; Anthony, D.; Churchill, G.C.; Wood, M.J.A.; et al. The CRTC1-SIK1 pathway regulates entrainment of the circadian clock. Cell 2013, 154, 1100–1111. [Google Scholar] [CrossRef]
- Gao, W.-W.; Tang, H.-M.V.; Cheng, Y.; Chan, C.-P.; Chan, C.-P.; Jin, D.-Y. Suppression of gluconeogenic gene transcription by SIK1-induced ubiquitination and degradation of CRTC1. Biochim. Biophys. acta. Gene Regul. Mech. 2018, 1861, 211–223. [Google Scholar] [CrossRef] [PubMed]
- Sakamoto, K.; Norona, F.E.; Alzate-Correa, D.; Scarberry, D.; Hoyt, K.R.; Obrietan, K. Clock and Light Regulation of the CREB Coactivator CRTC1 in the Suprachiasmatic Circadian Clock. J. Neurosci. 2013, 33, 9021–9027. [Google Scholar] [CrossRef]
- Hayasaka, N.; Hirano, A.; Miyoshi, Y.; Tokuda, I.T.; Yoshitane, H.; Matsuda, J.; Fukada, Y. Salt-inducible kinase 3 regulates the mammalian circadian clock by destabilizing PER2 protein. Elife 2017, 6, e24779. [Google Scholar] [CrossRef]
- Ralph, M.R.; Menaker, M. A mutation of the circadian system in golden hamsters. Science 1988, 241, 1225–1227. [Google Scholar] [CrossRef]
- Lowrey, P.L.; Shimomura, K.; Antoch, M.P.; Yamazaki, S.; Zemenides, P.D.; Ralph, M.R.; Menaker, M.; Takahashi, J.S. Positional syntenic cloning and functional characterization of the mammalian circadian mutation tau. Science 2000, 288, 483–492. [Google Scholar] [CrossRef]
- Pilorz, V.; Cunningham, P.S.; Jackson, A.; West, A.C.; Wager, T.T.; Loudon, A.S.I.; Bechtold, D.A. A novel mechanism controlling resetting speed of the circadian clock to environmental stimuli. Curr. Biol. 2014, 24, 766–773. [Google Scholar] [CrossRef]
- Paul, J.R.; McKeown, A.S.; Davis, J.A.; Totsch, S.K.; Mintz, E.M.; Kraft, T.W.; Cowell, R.M.; Gamble, K.L. Glycogen synthase kinase 3 regulates photic signaling in the suprachiasmatic nucleus. Eur. J. Neurosci. 2017, 45, 1102–1110. [Google Scholar] [CrossRef]
- Mehta, N.; Cheng, A.H.; Chiang, C.-K.; Mendoza-Viveros, L.; Ling, H.H.; Patel, A.; Xu, B.; Figeys, D.; Cheng, H.-Y.M. GRK2 Fine-Tunes Circadian Clock Speed and Entrainment via Transcriptional and Post-translational Control of PERIOD Proteins. Cell Rep. 2015, 12, 1272–1288. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Evans, J.A.; Leise, T.L.; Castanon-Cervantes, O.; Davidson, A.J. Dynamic Interactions Mediated by Nonredundant Signaling Mechanisms Couple Circadian Clock Neurons. Neuron 2013, 80, 973–983. [Google Scholar] [CrossRef]
- Freeman, G.M.; Krock, R.M.; Aton, S.J.; Thaben, P.; Herzog, E.D. GABA networks destabilize genetic oscillations in the circadian pacemaker. Neuron 2013, 78, 799–806. [Google Scholar] [CrossRef]
- Farajnia, S.; van Westering, T.L.E.; Meijer, J.H.; Michel, S. Seasonal induction of GABAergic excitation in the central mammalian clock. Proc. Natl. Acad. Sci. USA 2014, 111, 9627–9632. [Google Scholar] [CrossRef]
- Myung, J.; Hong, S.; DeWoskin, D.; De Schutter, E.; Forger, D.B.; Takumi, T. GABA-mediated repulsive coupling between circadian clock neurons in the SCN encodes seasonal time. Proc. Natl. Acad. Sci. USA 2015, 112, E3920–E3929. [Google Scholar] [CrossRef]
- Barca-Mayo, O.; Pons-Espinal, M.; Follert, P.; Armirotti, A.; Berdondini, L.; De Pietri Tonelli, D. Astrocyte deletion of Bmal1 alters daily locomotor activity and cognitive functions via GABA signalling. Nat. Commun. 2017, 8, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Brown, T.M.; Colwell, C.S.; Waschek, J.A.; Piggins, H.D. Disrupted Neuronal Activity Rhythms in the Suprachiasmatic Nuclei of Vasoactive Intestinal Polypeptide-Deficient Mice. J. Neurophysiol. 2007, 97, 2553–2558. [Google Scholar] [CrossRef]
- Hughes, A.T.L.; Croft, C.L.; Samuels, R.E.; Myung, J.; Takumi, T.; Piggins, H.D. Constant light enhances synchrony among circadian clock cells and promotes behavioral rhythms in VPAC2-signaling deficient mice. Sci. Rep. 2015, 5, 14044. [Google Scholar] [CrossRef]
- An, S.; Harang, R.; Meeker, K.; Granados-Fuentes, D.; Tsai, C.A.; Mazuski, C.; Kim, J.; Doyle, F.J.; Petzold, L.R.; Herzog, E.D. A neuropeptide speeds circadian entrainment by reducing intercellular synchrony. Proc. Natl. Acad. Sci. USA 2013, 110, E4355–E4361. [Google Scholar] [CrossRef]
- Ananthasubramaniam, B.; Herzog, E.D.; Herzel, H. Timing of Neuropeptide Coupling Determines Synchrony and Entrainment in the Mammalian Circadian Clock. PLoS Comput. Biol. 2014, 10, e1003565. [Google Scholar] [CrossRef]
- Liu, A.C.; Welsh, D.K.; Ko, C.H.; Tran, H.G.; Zhang, E.E.; Priest, A.A.; Buhr, E.D.; Singer, O.; Meeker, K.; Verma, I.M.; et al. Intercellular coupling confers robustness against mutations in the SCN circadian clock network. Cell 2007, 129, 605–616. [Google Scholar] [CrossRef] [PubMed]
- Maywood, E.S.; Chesham, J.E.; O’Brien, J.A.; Hastings, M.H. A diversity of paracrine signals sustains molecular circadian cycling in suprachiasmatic nucleus circuits. Proc. Natl. Acad. Sci. USA 2011, 108, 14306–14311. [Google Scholar] [CrossRef]
- Yamaguchi, Y.; Suzuki, T.; Mizoro, Y.; Kori, H.; Okada, K.; Chen, Y.; Fustin, J.-M.; Yamazaki, F.; Mizuguchi, N.; Zhang, J.; et al. Mice genetically deficient in vasopressin V1a and V1b receptors are resistant to jet lag. Science 2013, 342, 85–90. [Google Scholar] [CrossRef]
- Mieda, M.; Ono, D.; Hasegawa, E.; Okamoto, H.; Honma, K.-I.; Honma, S.; Sakurai, T. Cellular clocks in AVP neurons of the SCN are critical for interneuronal coupling regulating circadian behavior rhythm. Neuron 2015, 85, 1103–1116. [Google Scholar] [CrossRef]
- Mieda, M.; Okamoto, H.; Sakurai, T. Manipulating the Cellular Circadian Period of Arginine Vasopressin Neurons Alters the Behavioral Circadian Period. Curr. Biol. 2016, 26, 2535–2542. [Google Scholar] [CrossRef]
- Maywood, E.S.; Reddy, A.B.; Wong, G.K.Y.; O’Neill, J.S.; O’Brien, J.A.; McMahon, D.G.; Harmar, A.J.; Okamura, H.; Hastings, M.H. Synchronization and maintenance of timekeeping in suprachiasmatic circadian clock cells by neuropeptidergic signaling. Curr. Biol. 2006, 16, 599–605. [Google Scholar] [CrossRef]
- Cheng, M.Y.; Bullock, C.M.; Li, C.; Lee, A.G.; Bermak, J.C.; Belluzzi, J.; Weaver, D.R.; Leslie, F.M.; Zhou, Q.-Y. Prokineticin 2 transmits the behavioural circadian rhythm of the suprachiasmatic nucleus. Nature 2002, 417, 405–410. [Google Scholar] [CrossRef]
- Li, J.-D.; Hu, W.-P.; Boehmer, L.; Cheng, M.Y.; Lee, A.G.; Jilek, A.; Siegel, J.M.; Zhou, Q.-Y. Attenuated Circadian Rhythms in Mice Lacking the Prokineticin 2 Gene. J. Neurosci. 2006, 26, 11615–11623. [Google Scholar] [CrossRef]
- Prosser, H.M.; Bradley, A.; Chesham, J.E.; Ebling, F.J.P.; Hastings, M.H.; Maywood, E.S. Prokineticin receptor 2 (Prokr2) is essential for the regulation of circadian behavior by the suprachiasmatic nuclei. Proc. Natl. Acad. Sci. USA 2007, 104, 648–653. [Google Scholar] [CrossRef]
- Li, X.; Zhang, C.; Zhou, Q.-Y. Overexpression of Prokineticin 2 in Transgenic Mice Leads to Reduced Circadian Behavioral Rhythmicity and Altered Molecular Rhythms in the Suprachiasmatic Clock. J. Circadian Rhythms 2018, 16, 13. [Google Scholar] [CrossRef]
- Meng, Q.-J.; Logunova, L.; Maywood, E.S.; Gallego, M.; Lebiecki, J.; Brown, T.M.; Sládek, M.; Semikhodskii, A.S.; Glossop, N.R.J.; Piggins, H.D.; et al. Setting clock speed in mammals: The CK1 epsilon tau mutation in mice accelerates circadian pacemakers by selectively destabilizing PERIOD proteins. Neuron 2008, 58, 78–88. [Google Scholar] [CrossRef]
- Etchegaray, J.-P.; Yu, E.A.; Indic, P.; Dallmann, R.; Weaver, D.R. Casein kinase 1 delta (CK1delta) regulates period length of the mouse suprachiasmatic circadian clock in vitro. PLoS ONE 2010, 5, e10303. [Google Scholar] [CrossRef]
- Ishida, Y.; Yagita, K.; Fukuyama, T.; Nishimura, M.; Nagano, M.; Shigeyoshi, Y.; Yamaguchi, S.; Komori, T.; Okamura, H. Constitutive expression and delayed light response of casein kinase Iepsilon and Idelta mRNAs in the mouse suprachiasmatic nucleus. J. Neurosci. Res. 2001, 64, 612–616. [Google Scholar] [CrossRef]
- Camacho, F.; Cilio, M.; Guo, Y.; Virshup, D.M.; Patel, K.; Khorkova, O.; Styren, S.; Morse, B.; Yao, Z.; Keesler, G.A. Human casein kinase Idelta phosphorylation of human circadian clock proteins period 1 and 2. FEBS Lett. 2001, 489, 159–165. [Google Scholar] [CrossRef]
- Meng, Q.-J.; Maywood, E.S.; Bechtold, D.A.; Lu, W.-Q.; Li, J.; Gibbs, J.E.; Dupré, S.M.; Chesham, J.E.; Rajamohan, F.; Knafels, J.; et al. Entrainment of disrupted circadian behavior through inhibition of casein kinase 1 (CK1) enzymes. Proc. Natl. Acad. Sci. USA 2010, 107, 15240–15245. [Google Scholar] [CrossRef]
- Narasimamurthy, R.; Hunt, S.R.; Lu, Y.; Fustin, J.-M.; Okamura, H.; Partch, C.L.; Forger, D.B.; Kim, J.K.; Virshup, D.M. CK1δ/ε protein kinase primes the PER2 circadian phosphoswitch. Proc. Natl. Acad. Sci. USA 2018, 115, 5986–5991. [Google Scholar] [CrossRef]
- Maier, B.; Wendt, S.; Vanselow, J.T.; Wallach, T.; Reischl, S.; Oehmke, S.; Schlosser, A.; Kramer, A. A large-scale functional RNAi screen reveals a role for CK2 in the mammalian circadian clock. Genes Dev. 2009, 23, 708–718. [Google Scholar] [CrossRef]
- Tsuchiya, Y.; Akashi, M.; Matsuda, M.; Goto, K.; Miyata, Y.; Node, K.; Nishida, E. Involvement of the protein kinase CK2 in the regulation of mammalian circadian rhythms. Sci. Signal. 2009, 2, ra26. [Google Scholar] [CrossRef]
- Tamaru, T.; Hirayama, J.; Isojima, Y.; Nagai, K.; Norioka, S.; Takamatsu, K.; Sassone-Corsi, P. CK2alpha phosphorylates BMAL1 to regulate the mammalian clock. Nat. Struct. Mol. Biol. 2009, 16, 446–448. [Google Scholar] [CrossRef]
- Iwahana, E.; Akiyama, M.; Miyakawa, K.; Uchida, A.; Kasahara, J.; Fukunaga, K.; Hamada, T.; Shibata, S. Effect of lithium on the circadian rhythms of locomotor activity and glycogen synthase kinase-3 protein expression in the mouse suprachiasmatic nuclei. Eur. J. Neurosci. 2004, 19, 2281–2287. [Google Scholar] [CrossRef]
- Iitaka, C.; Miyazaki, K.; Akaike, T.; Ishida, N. A Role for Glycogen Synthase Kinase-3β in the Mammalian Circadian Clock. J. Biol. Chem. 2005, 280, 29397–29402. [Google Scholar] [CrossRef]
- Besing, R.C.; Paul, J.R.; Hablitz, L.M.; Rogers, C.O.; Johnson, R.L.; Young, M.E.; Gamble, K.L. Circadian rhythmicity of active GSK3 isoforms modulates molecular clock gene rhythms in the suprachiasmatic nucleus. J. Biol. Rhythms 2015, 30, 155–160. [Google Scholar] [CrossRef]
- Harada, Y.; Sakai, M.; Kurabayashi, N.; Hirota, T.; Fukada, Y. Ser-557-phosphorylated mCRY2 is degraded upon synergistic phosphorylation by glycogen synthase kinase-3 beta. J. Biol. Chem. 2005, 280, 31714–31721. [Google Scholar] [CrossRef]
- Kurabayashi, N.; Hirota, T.; Harada, Y.; Sakai, M.; Fukada, Y. Phosphorylation of mCRY2 at Ser557 in the hypothalamic suprachiasmatic nucleus of the mouse. Chronobiol. Int. 2006, 23, 129–134. [Google Scholar] [CrossRef]
- Yin, L.; Wang, J.; Klein, P.S.; Lazar, M.A. Nuclear receptor Rev-erbalpha is a critical lithium-sensitive component of the circadian clock. Science 2006, 311, 1002–1005. [Google Scholar] [CrossRef]
- Spengler, M.L.; Kuropatwinski, K.K.; Schumer, M.; Antoch, M.P. A serine cluster mediates BMAL1-dependent CLOCK phosphorylation and degradation. Cell Cycle 2009, 8, 4138–4146. [Google Scholar] [CrossRef]
- Paul, J.R.; Johnson, R.L.; Jope, R.S.; Gamble, K.L. Disruption of circadian rhythmicity and suprachiasmatic action potential frequency in a mouse model with constitutive activation of glycogen synthase kinase 3. Neuroscience 2012, 226, 1–9. [Google Scholar] [CrossRef]
- Lavoie, J.; Hébert, M.; Beaulieu, J.-M. Glycogen synthase kinase-3β haploinsufficiency lengthens the circadian locomotor activity period in mice. Behav. Brain Res. 2013, 253, 262–265. [Google Scholar] [CrossRef]
- Paul, J.R.; DeWoskin, D.; McMeekin, L.J.; Cowell, R.M.; Forger, D.B.; Gamble, K.L. Regulation of persistent sodium currents by glycogen synthase kinase 3 encodes daily rhythms of neuronal excitability. Nat. Commun. 2016, 7, 13470. [Google Scholar] [CrossRef]
- Helfrich-Förster, C. The period clock gene is expressed in central nervous system neurons which also produce a neuropeptide that reveals the projections of circadian pacemaker cells within the brain of Drosophila melanogaster. Proc. Natl. Acad. Sci. USA 1995, 92, 612–616. [Google Scholar] [CrossRef]
- Li, Y.; Guo, F.; Shen, J.; Rosbash, M. PDF and cAMP enhance PER stability in Drosophila clock neurons. Proc. Natl. Acad. Sci. USA 2014, 111, E1284–E1290. [Google Scholar] [CrossRef]
- Seluzicki, A.; Flourakis, M.; Kula-Eversole, E.; Zhang, L.; Kilman, V.; Allada, R. Dual PDF Signaling Pathways Reset Clocks Via TIMELESS and Acutely Excite Target Neurons to Control Circadian Behavior. PLoS Biol. 2014, 12, e1001810. [Google Scholar] [CrossRef]
- Park, J.H.; Helfrich-Förster, C.; Lee, G.; Liu, L.; Rosbash, M.; Hall, J.C. Differential regulation of circadian pacemaker output by separate clock genes in Drosophila. Proc. Natl. Acad. Sci. USA 2000, 97, 3608–3613. [Google Scholar] [CrossRef]
- King, A.N.; Barber, A.F.; Smith, A.E.; Dreyer, A.P.; Sitaraman, D.; Nitabach, M.N.; Cavanaugh, D.J.; Sehgal, A. A Peptidergic Circuit Links the Circadian Clock to Locomotor Activity. Curr. Biol. 2017, 27, 1915–1927.e5. [Google Scholar] [CrossRef] [PubMed]
- Hyun, S.; Lee, Y.; Hong, S.-T.; Bang, S.; Paik, D.; Kang, J.; Shin, J.; Lee, J.; Jeon, K.; Hwang, S.; et al. Drosophila GPCR Han is a receptor for the circadian clock neuropeptide PDF. Neuron 2005, 48, 267–278. [Google Scholar] [CrossRef]
- Lear, B.C.; Merrill, C.E.; Lin, J.-M.; Schroeder, A.; Zhang, L.; Allada, R. A G protein-coupled receptor, groom-of-PDF, is required for PDF neuron action in circadian behavior. Neuron 2005, 48, 221–227. [Google Scholar] [CrossRef] [PubMed]
- Mertens, I.; Vandingenen, A.; Johnson, E.C.; Shafer, O.T.; Li, W.; Trigg, J.S.; De Loof, A.; Schoofs, L.; Taghert, P.H. PDF receptor signaling in Drosophila contributes to both circadian and geotactic behaviors. Neuron 2005, 48, 213–219. [Google Scholar] [CrossRef]
- Im, S.H.; Taghert, P.H. PDF receptor expression reveals direct interactions between circadian oscillators in Drosophila. J. Comp. Neurol. 2010, 518, 1925–1945. [Google Scholar] [CrossRef]
- Shafer, O.T.; Kim, D.J.; Dunbar-Yaffe, R.; Nikolaev, V.O.; Lohse, M.J.; Taghert, P.H. Widespread receptivity to neuropeptide PDF throughout the neuronal circadian clock network of Drosophila revealed by real-time cyclic AMP imaging. Neuron 2008, 58, 223–237. [Google Scholar] [CrossRef]
- Klose, M.; Duvall, L.; Li, W.; Liang, X.; Ren, C.; Steinbach, J.H.; Taghert, P.H. Functional PDF Signaling in the Drosophila Circadian Neural Circuit Is Gated by Ral A-Dependent Modulation. Neuron 2016, 90, 781–794. [Google Scholar] [CrossRef]
- Liang, X.; Holy, T.E.; Taghert, P.H. Synchronous Drosophila circadian pacemakers display nonsynchronous Ca2+ rhythms in vivo. Science 2016, 351, 976–981. [Google Scholar] [CrossRef] [PubMed]
- Liang, X.; Holy, T.E.; Taghert, P.H. A Series of Suppressive Signals within the Drosophila Circadian Neural Circuit Generates Sequential Daily Outputs. Neuron 2017, 94, 1173–1189.e4. [Google Scholar] [CrossRef] [PubMed]
- Duvall, L.B.; Taghert, P.H. The Circadian Neuropeptide PDF Signals Preferentially through a Specific Adenylate Cyclase Isoform AC3 in M Pacemakers of Drosophila. PLoS Biol. 2012, 10, e1001337. [Google Scholar] [CrossRef]
- Duvall, L.B.; Taghert, P.H. E and M circadian pacemaker neurons use different PDF receptor signalosome components in drosophila. J. Biol. Rhythms 2013, 28, 239–248. [Google Scholar] [CrossRef]
- Choi, C.; Cao, G.; Tanenhaus, A.K.; McCarthy, E.V.; Jung, M.; Schleyer, W.; Shang, Y.; Rosbash, M.; Yin, J.C.P.; Nitabach, M.N. Autoreceptor control of peptide/neurotransmitter corelease from PDF neurons determines allocation of circadian activity in drosophila. Cell Rep. 2012, 2, 332–344. [Google Scholar] [CrossRef]
- Garczynski, S.F.; Brown, M.R.; Shen, P.; Murray, T.F.; Crim, J.W. Characterization of a functional neuropeptide F receptor from Drosophila melanogaster. Peptides 2002, 23, 773–780. [Google Scholar] [CrossRef]
- Mertens, I.; Meeusen, T.; Huybrechts, R.; De Loof, A.; Schoofs, L. Characterization of the short neuropeptide F receptor from Drosophila melanogaster. Biochem. Biophys. Res. Commun. 2002, 297, 1140–1148. [Google Scholar] [CrossRef]
- Vanden Broeck, J. Neuropeptides and their precursors in the fruitfly, Drosophila melanogaster. Peptides 2001, 22, 241–254. [Google Scholar] [CrossRef]
- Nässel, D.R.; Enell, L.E.; Santos, J.G.; Wegener, C.; Johard, H.A.D. A large population of diverse neurons in the Drosophila central nervous system expresses short neuropeptide F, suggesting multiple distributed peptide functions. BMC Neurosci. 2008, 9, 90. [Google Scholar] [CrossRef] [PubMed]
- Nässel, D.R.; Wegener, C. A comparative review of short and long neuropeptide F signaling in invertebrates: Any similarities to vertebrate neuropeptide Y signaling? Peptides 2011, 32, 1335–1355. [Google Scholar] [CrossRef] [PubMed]
- Lee, G.; Bahn, J.H.; Park, J.H. Sex- and clock-controlled expression of the neuropeptide F gene in Drosophila. Proc. Natl. Acad. Sci. USA 2006, 103, 12580–12585. [Google Scholar] [CrossRef] [PubMed]
- He, C.; Cong, X.; Zhang, R.; Wu, D.; An, C.; Zhao, Z. Regulation of circadian locomotor rhythm by neuropeptide Y-like system in Drosophila melanogaster. Insect Mol. Biol. 2013, 22, 376–388. [Google Scholar] [CrossRef]
- Kula-Eversole, E.; Nagoshi, E.; Shang, Y.; Rodriguez, J.; Allada, R.; Rosbash, M. Surprising gene expression patterns within and between PDF-containing circadian neurons in Drosophila. Proc. Natl. Acad. Sci. USA 2010, 107, 13497–13502. [Google Scholar] [CrossRef]
- Abruzzi, K.C.; Zadina, A.; Luo, W.; Wiyanto, E.; Rahman, R.; Guo, F.; Shafer, O.; Rosbash, M. RNA-seq analysis of Drosophila clock and non-clock neurons reveals neuron-specific cycling and novel candidate neuropeptides. PLOS Genet. 2017, 13, e1006613. [Google Scholar] [CrossRef]
- Harrington, M.E.; Nance, D.M.; Rusak, B. Double-labeling of neuropeptide Y-immunoreactive neurons which project from the geniculate to the suprachiasmatic nuclei. Brain Res. 1987, 410, 275–282. [Google Scholar] [CrossRef]
- Harrington, M.E.; Rusak, B. Lesions of the thalamic intergeniculate leaflet alter hamster circadian rhythms. J. Biol. Rhythms 1986, 1, 309–325. [Google Scholar] [CrossRef]
- Janik, D.; Mrosovsky, N. Intergeniculate leaflet lesions and behaviorally-induced shifts of circadian rhythms. Brain Res. 1994, 651, 174–182. [Google Scholar] [CrossRef]
- Johard, H.A.D.; Yoishii, T.; Dircksen, H.; Cusumano, P.; Rouyer, F.; Helfrich-Förster, C.; Nässel, D.R. Peptidergic clock neurons in Drosophila: Ion transport peptide and short neuropeptide F in subsets of dorsal and ventral lateral neurons. J. Comp. Neurol. 2009, 516, 59–73. [Google Scholar] [CrossRef] [PubMed]
- Hermann, C.; Yoshii, T.; Dusik, V.; Helfrich-Förster, C. Neuropeptide F immunoreactive clock neurons modify evening locomotor activity and free-running period in Drosophila melanogaster. J. Comp. Neurol. 2012, 520, 970–987. [Google Scholar] [CrossRef]
- Chung, B.Y.; Ro, J.; Hutter, S.A.; Miller, K.M.; Guduguntla, L.S.; Kondo, S.; Pletcher, S.D. Drosophila Neuropeptide F Signaling Independently Regulates Feeding and Sleep-Wake Behavior. Cell Rep. 2017, 19, 2441–2450. [Google Scholar] [CrossRef] [PubMed]
- He, C.; Yang, Y.; Zhang, M.; Price, J.L.; Zhao, Z. Regulation of sleep by neuropeptide Y-like system in Drosophila melanogaster. PLoS ONE 2013, 8, e74237. [Google Scholar] [CrossRef]
- Vecsey, C.G.; Pírez, N.; Griffith, L.C. The Drosophila neuropeptides PDF and sNPF have opposing electrophysiological and molecular effects on central neurons. J. Neurophysiol. 2014, 111, 1033–1045. [Google Scholar] [CrossRef] [PubMed]
- Besing, R.C.; Hablitz, L.M.; Paul, J.R.; Johnson, R.L.; Prosser, R.A.; Gamble, K.L. Neuropeptide Y-induced phase shifts of PER2::LUC rhythms are mediated by long-term suppression of neuronal excitability in a phase-specific manner. Chronobiol. Int. 2012, 29, 91–102. [Google Scholar] [CrossRef] [PubMed]
- Shigeri, Y.; Fujimoto, M. Y2 receptors for neuropeptide Y are coupled to three intracellular signal transduction pathways in a human neuroblastoma cell line. J. Biol. Chem. 1994, 269, 8842–8848. [Google Scholar]
- Shang, Y.; Donelson, N.C.; Vecsey, C.G.; Guo, F.; Rosbash, M.; Griffith, L.C. Short neuropeptide F is a sleep-promoting inhibitory modulator. Neuron 2013, 80, 171–183. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.; Shi, W.; Li, L.; Zheng, Z.; Li, T.; Bai, W.; Zhao, Z. Regulation of sleep by the short neuropeptide F (sNPF) in Drosophila melanogaster. Insect Biochem. Mol. Biol. 2013, 43, 809–819. [Google Scholar] [CrossRef] [PubMed]
- Hermann-Luibl, C.; Yoshii, T.; Senthilan, P.R.; Dircksen, H.; Helfrich-Förster, C. The ion transport peptide is a new functional clock neuropeptide in the fruit fly Drosophila melanogaster. J. Neurosci. 2014, 34, 9522–9536. [Google Scholar] [CrossRef] [PubMed]
- Kunst, M.; Hughes, M.E.; Raccuglia, D.; Felix, M.; Li, M.; Barnett, G.; Duah, J.; Nitabach, M.N. Calcitonin gene-related peptide neurons mediate sleep-specific circadian output in Drosophila. Curr. Biol. 2014, 24, 2652–2664. [Google Scholar] [CrossRef]
- Goda, T.; Tang, X.; Umezaki, Y.; Chu, M.L.; Kunst, M.; Nitabach, M.N.; Hamada, F.N. Drosophila DH31 Neuropeptide and PDF Receptor Regulate Night-Onset Temperature Preference. J. Neurosci. 2016, 36, 11739–11754. [Google Scholar] [CrossRef] [PubMed]
- Goda, T.; Umezaki, Y.; Alwattari, F.; Seo, H.W.; Hamada, F.N. Neuropeptides PDF and DH31 hierarchically regulate free-running rhythmicity in Drosophila circadian locomotor activity. Sci. Rep. 2019, 9, 838. [Google Scholar] [CrossRef]
- Goda, T.; Doi, M.; Umezaki, Y.; Murai, I.; Shimatani, H.; Chu, M.L.; Nguyen, V.H.; Okamura, H.; Hamada, F.N. Calcitonin receptors are ancient modulators for rhythms of preferential temperature in insects and body temperature in mammals. Genes Dev. 2018, 32, 140–155. [Google Scholar] [CrossRef]
- Hansen, K.K.; Hauser, F.; Williamson, M.; Weber, S.B.; Grimmelikhuijzen, C.J.P. The Drosophila genes CG14593 and CG30106 code for G-protein-coupled receptors specifically activated by the neuropeptides CCHamide-1 and CCHamide-2. Biochem. Biophys. Res. Commun. 2011, 404, 184–189. [Google Scholar] [CrossRef]
- Ida, T.; Takahashi, T.; Tominaga, H.; Sato, T.; Sano, H.; Kume, K.; Ozaki, M.; Hiraguchi, T.; Shiotani, H.; Terajima, S.; et al. Isolation of the bioactive peptides CCHamide-1 and CCHamide-2 from Drosophila and their putative role in appetite regulation as ligands for G protein-coupled receptors. Front. Endocrinol. (Lausanne). 2012, 3, 177. [Google Scholar] [CrossRef] [PubMed]
- Fujiwara, Y.; Hermann-Luibl, C.; Katsura, M.; Sekiguchi, M.; Ida, T.; Helfrich-Förster, C.; Yoshii, T. The CCHamide1 Neuropeptide Expressed in the Anterior Dorsal Neuron 1 Conveys a Circadian Signal to the Ventral Lateral Neurons in Drosophila melanogaster. Front. Physiol. 2018, 9, 1276. [Google Scholar] [CrossRef] [PubMed]
- Birgül, N.; Weise, C.; Kreienkamp, H.J.; Richter, D. Reverse physiology in drosophila: Identification of a novel allatostatin-like neuropeptide and its cognate receptor structurally related to the mammalian somatostatin/galanin/opioid receptor family. EMBO J. 1999, 18, 5892. [Google Scholar] [CrossRef]
- Dulcis, D.; Jamshidi, P.; Leutgeb, S.; Spitzer, N.C. Neurotransmitter Switching in the Adult Brain Regulates Behavior. Science 2013, 340, 449–453. [Google Scholar] [CrossRef]
- Deats, S.P.; Adidharma, W.; Yan, L. Hypothalamic dopaminergic neurons in an animal model of seasonal affective disorder. Neurosci. Lett. 2015, 602, 17–21. [Google Scholar] [CrossRef]
- Dumbell, R.A.; Scherbarth, F.; Diedrich, V.; Schmid, H.A.; Steinlechner, S.; Barrett, P. Somatostatin Agonist Pasireotide Promotes a Physiological State Resembling Short-Day Acclimation in the Photoperiodic Male Siberian Hamster (Phodopus sungorus). J. Neuroendocrinol. 2015, 27, 588–599. [Google Scholar] [CrossRef]
- Tanaka, M.; Okamura, H.; Matsuda, T.; Shigeyoshi, Y.; Hisa, Y.; Chihara, K.; Ibata, Y. Somatostatin neurons form a distinct peptidergic neuronal group in the rat suprachiasmatic nucleus: A double labeling in situ hybridization study. Neurosci. Lett. 1996, 215, 119–122. [Google Scholar] [CrossRef]
- Takeuchi, J.; Nagasaki, H.; Shinohara, K.; Inouye, S.T. A circadian rhythm of somatostatin messenger RNA levels, but not of vasoactive intestinal polypeptide/peptide histidine isoleucine messenger RNA levels in rat suprachiasmatic nucleus. Mol. Cell. Neurosci. 1992, 3, 29–35. [Google Scholar] [CrossRef]
- Hamada, T.; Shibata, S.; Tsuneyoshi, A.; Tominaga, K.; Watanabe, S. Effect of somatostatin on circadian rhythms of firing and 2-deoxyglucose uptake in rat suprachiasmatic slices. Am. J. Physiol. 1993, 265, R1199–R1204. [Google Scholar] [CrossRef] [PubMed]
- Fukuhara, C.; Hamada, T.; Shibata, S.; Watanabe, S.; Aoki, K.; Inouye, S.I. Phase advances of circadian rhythms in somatostatin depleted rats: Effects of cysteamine on rhythms of locomotor activity and electrical discharge of the suprachiasmatic nucleus. J. Comp. Physiol. A. 1994, 175, 677–685. [Google Scholar] [CrossRef] [PubMed]
- Shafer, O.T.; Helfrich-Förster, C.; Renn, S.C.P.; Taghert, P.H. Reevaluation of Drosophila melanogaster’s neuronal circadian pacemakers reveals new neuronal classes. J. Comp. Neurol. 2006, 498, 180–193. [Google Scholar] [CrossRef]
- Verleyen, P.; Baggerman, G.; Wiehart, U.; Schoeters, E.; Van Lommel, A.; De Loof, A.; Schoofs, L. Expression of a novel neuropeptide, NVGTLARDFQLPIPNamide, in the larval and adult brain of Drosophila melanogaster. J. Neurochem. 2004, 88, 311–319. [Google Scholar] [CrossRef] [PubMed]
- Baggerman, G.; Cerstiaens, A.; De Loof, A.; Schoofs, L. Peptidomics of the Larval Drosophila melanogaster Central Nervous System. J. Biol. Chem. 2002, 277, 40368–40374. [Google Scholar] [CrossRef]
- Hamasaka, Y.; Rieger, D.; Parmentier, M.; Grau, Y.; Helfrich-Förster, C.; Nässel, D.R. Glutamate and its metabotropic receptor in Drosophila clock neuron circuits. J. Comp. Neurol. 2007, 505, 32–45. [Google Scholar] [CrossRef] [PubMed]
- Collins, B.; Kane, E.A.; Reeves, D.C.; Akabas, M.H.; Blau, J. Balance of Activity between LNvs and Glutamatergic Dorsal Clock Neurons Promotes Robust Circadian Rhythms in Drosophila. Neuron 2012, 74, 706–718. [Google Scholar] [CrossRef] [PubMed]
- Collins, B.; Kaplan, H.S.; Cavey, M.; Lelito, K.R.; Bahle, A.H.; Zhu, Z.; Macara, A.M.; Roman, G.; Shafer, O.T.; Blau, J. Differentially Timed Extracellular Signals Synchronize Pacemaker Neuron Clocks. PLoS Biol. 2014, 12, e1001959. [Google Scholar] [CrossRef] [PubMed]
- Hamasaka, Y.; Wegener, C.; Nässel, D.R. GABA modulates Drosophila circadian clock neurons via GABAB receptors and decreases in calcium. J. Neurobiol. 2005, 65, 225–240. [Google Scholar] [CrossRef]
- Dahdal, D.; Reeves, D.C.; Ruben, M.; Akabas, M.H.; Blau, J. Drosophila Pacemaker Neurons Require G Protein Signaling and GABAergic Inputs to Generate Twenty-Four Hour Behavioral Rhythms. Neuron 2010, 68, 964–977. [Google Scholar] [CrossRef]
- Lelito, K.R.; Shafer, O.T. Reciprocal cholinergic and GABAergic modulation of the small ventrolateral pacemaker neurons of Drosophila’s circadian clock neuron network. J. Neurophysiol. 2012, 107, 2096–2108. [Google Scholar] [CrossRef]
- Gmeiner, F.; Kołodziejczyk, A.; Yoshii, T.; Rieger, D.; Nässel, D.R.; Helfrich-Förster, C. GABA(B) receptors play an essential role in maintaining sleep during the second half of the night in Drosophila melanogaster. J. Exp. Biol. 2013, 216, 3837–3843. [Google Scholar] [CrossRef]
- Buhl, E.; Bradlaugh, A.; Ogueta, M.; Chen, K.-F.; Stanewsky, R.; Hodge, J.J.L. Quasimodo mediates daily and acute light effects on Drosophila clock neuron excitability. Proc. Natl. Acad. Sci. USA 2016, 113, 13486–13491. [Google Scholar] [CrossRef]
- Li, Q.; Li, Y.; Wang, X.; Qi, J.; Jin, X.; Tong, H.; Zhou, Z.; Zhang, Z.C.; Han, J. Fbxl4 Serves as a Clock Output Molecule that Regulates Sleep through Promotion of Rhythmic Degradation of the GABAA Receptor. Curr. Biol. 2017, 27, 3616–3625.e5. [Google Scholar] [CrossRef]
- Liu, S.; Lamaze, A.; Liu, Q.; Tabuchi, M.; Yang, Y.; Fowler, M.; Bharadwaj, R.; Zhang, J.; Bedont, J.; Blackshaw, S.; et al. WIDE AWAKE mediates the circadian timing of sleep onset. Neuron 2014, 82, 151–166. [Google Scholar] [CrossRef]
- Hamasaka, Y.; Nässel, D.R. Mapping of serotonin, dopamine, and histamine in relation to different clock neurons in the brain of Drosophila. J. Comp. Neurol. 2006, 494, 314–330. [Google Scholar] [CrossRef]
- Yuan, Q.; Lin, F.; Zheng, X.; Sehgal, A. Serotonin modulates circadian entrainment in Drosophila. Neuron 2005, 47, 115–127. [Google Scholar] [CrossRef]
- Nichols, C.D. 5-HT2 receptors in Drosophila are expressed in the brain and modulate aspects of circadian behaviors. Dev. Neurobiol. 2007, 67, 752–763. [Google Scholar] [CrossRef]
- McCarthy, E.V.; Wu, Y.; Decarvalho, T.; Brandt, C.; Cao, G.; Nitabach, M.N. Synchronized bilateral synaptic inputs to Drosophila melanogaster neuropeptidergic rest/arousal neurons. J. Neurosci. 2011, 31, 8181–8193. [Google Scholar] [CrossRef]
- Wegener, C.; Hamasaka, Y.; Nässel, D.R. Acetylcholine increases intracellular Ca2+ via nicotinic receptors in cultured PDF-containing clock neurons of Drosophila. J. Neurophysiol. 2004, 91, 912–923. [Google Scholar] [CrossRef]
- Keene, A.C.; Mazzoni, E.O.; Zhen, J.; Younger, M.A.; Yamaguchi, S.; Blau, J.; Desplan, C.; Sprecher, S.G. Distinct Visual Pathways Mediate Drosophila Larval Light Avoidance and Circadian Clock Entrainment. J. Neurosci. 2011, 31, 6527–6534. [Google Scholar] [CrossRef]
- Helfrich-Förster, C.; Edwards, T.; Yasuyama, K.; Wisotzki, B.; Schneuwly, S.; Stanewsky, R.; Meinertzhagen, I.A.; Hofbauer, A. The extraretinal eyelet of Drosophila: Development, ultrastructure, and putative circadian function. J. Neurosci. 2002, 22, 9255–9266. [Google Scholar] [CrossRef]
- Muraro, N.I.; Ceriani, M.F. Acetylcholine from Visual Circuits Modulates the Activity of Arousal Neurons in Drosophila. J. Neurosci. 2015, 35, 16315–16327. [Google Scholar] [CrossRef]
- Fogle, K.J.; Parson, K.G.; Dahm, N.A.; Holmes, T.C. CRYPTOCHROME Is a Blue-Light Sensor That Regulates Neuronal Firing Rate. Science 2011, 331, 1409–1413. [Google Scholar] [CrossRef]
- Frenkel, L.; Muraro, N.I.; Beltrán González, A.N.; Marcora, M.S.; Bernabó, G.; Hermann-Luibl, C.; Romero, J.I.; Helfrich-Förster, C.; Castaño, E.M.; Marino-Busjle, C.; et al. Organization of Circadian Behavior Relies on Glycinergic Transmission. Cell Rep. 2017, 19, 72–85. [Google Scholar] [CrossRef] [PubMed]
- Price, J.L.; Blau, J.; Rothenfluh, A.; Abodeely, M.; Kloss, B.; Young, M.W. double-time is a novel Drosophila clock gene that regulates PERIOD protein accumulation. Cell 1998, 94, 83–95. [Google Scholar] [CrossRef]
- Kloss, B.; Price, J.L.; Saez, L.; Blau, J.; Rothenfluh, A.; Wesley, C.S.; Young, M.W. The Drosophila clock gene double-time encodes a protein closely related to human casein kinase Iepsilon. Cell 1998, 94, 97–107. [Google Scholar] [CrossRef]
- Preuss, F.; Fan, J.-Y.; Kalive, M.; Bao, S.; Schuenemann, E.; Bjes, E.S.; Price, J.L. Drosophila doubletime mutations which either shorten or lengthen the period of circadian rhythms decrease the protein kinase activity of casein kinase I. Mol. Cell. Biol. 2004, 24, 886–898. [Google Scholar] [CrossRef] [PubMed]
- Rothenfluh, A.; Abodeely, M.; Young, M.W. Short-period mutations of per affect a double-time-dependent step in the Drosophila circadian clock. Curr. Biol. 2000, 10, 1399–1402. [Google Scholar] [CrossRef]
- Suri, V.; Hall, J.C.; Rosbash, M. Two novel doubletime mutants alter circadian properties and eliminate the delay between RNA and protein in Drosophila. J. Neurosci. 2000, 20, 7547–7555. [Google Scholar] [CrossRef] [PubMed]
- Muskus, M.J.; Preuss, F.; Fan, J.-Y.; Bjes, E.S.; Price, J.L. Drosophila DBT lacking protein kinase activity produces long-period and arrhythmic circadian behavioral and molecular rhythms. Mol. Cell. Biol. 2007, 27, 8049–8064. [Google Scholar] [CrossRef] [PubMed]
- Zheng, X.; Sowcik, M.; Chen, D.; Sehgal, A. Casein kinase 1 promotes synchrony of the circadian clock network. Mol. Cell. Biol. 2014, 34, 2682–2694. [Google Scholar] [CrossRef]
- Konopka, R.J.; Benzer, S. Clock mutants of Drosophila melanogaster. Proc. Natl. Acad. Sci. USA 1971, 68, 2112–2116. [Google Scholar] [CrossRef]
- Chiu, J.C.; Ko, H.W.; Edery, I. NEMO/NLK phosphorylates PERIOD to initiate a time-delay phosphorylation circuit that sets circadian clock speed. Cell 2011, 145, 357–370. [Google Scholar] [CrossRef]
- Chiu, J.C.; Vanselow, J.T.; Kramer, A.; Edery, I. The phospho-occupancy of an atypical SLIMB-binding site on PERIOD that is phosphorylated by DOUBLETIME controls the pace of the clock. Genes Dev. 2008, 22, 1758–1772. [Google Scholar] [CrossRef]
- Ko, H.W.; Jiang, J.; Edery, I. Role for Slimb in the degradation of Drosophila Period protein phosphorylated by Doubletime. Nature 2002, 420, 673–678. [Google Scholar] [CrossRef]
- Bao, S.; Rihel, J.; Bjes, E.; Fan, J.Y.; Price, J.L. The Drosophila double-timeS mutation delays the nuclear accumulation of period protein and affects the feedback regulation of period mRNA. J. Neurosci. 2001, 21, 7117–7126. [Google Scholar] [CrossRef]
- Cyran, S.A.; Yiannoulos, G.; Buchsbaum, A.M.; Saez, L.; Young, M.W.; Blau, J. The Double-Time Protein Kinase Regulates the Subcellular Localization of the Drosophila Clock Protein Period. J. Neurosci. 2005, 25, 5430–5437. [Google Scholar] [CrossRef]
- Kivimäe, S.; Saez, L.; Young, M.W. Activating PER repressor through a DBT-directed phosphorylation switch. PLoS Biol. 2008, 6, e183. [Google Scholar] [CrossRef]
- Kloss, B.; Rothenfluh, A.; Young, M.W.; Saez, L. Phosphorylation of period is influenced by cycling physical associations of double-time, period, and timeless in the Drosophila clock. Neuron 2001, 30, 699–706. [Google Scholar] [CrossRef]
- Kim, E.Y.; Edery, I. Balance between DBT/CKIepsilon kinase and protein phosphatase activities regulate phosphorylation and stability of Drosophila CLOCK protein. Proc. Natl. Acad. Sci. USA 2006, 103, 6178–6183. [Google Scholar] [CrossRef]
- Yu, W.; Zheng, H.; Price, J.L.; Hardin, P.E. DOUBLETIME plays a noncatalytic role to mediate CLOCK phosphorylation and repress CLOCK-dependent transcription within the Drosophila circadian clock. Mol. Cell. Biol. 2009, 29, 1452–1458. [Google Scholar] [CrossRef]
- Konopka, R.J.; Smith, R.F.; Orr, D. Characterization of Andante, a new Drosophila clock mutant, and its interactions with other clock mutants. J. Neurogenet. 1991, 7, 103–114. [Google Scholar] [CrossRef]
- Akten, B.; Jauch, E.; Genova, G.K.; Kim, E.Y.; Edery, I.; Raabe, T.; Jackson, F.R. A role for CK2 in the Drosophila circadian oscillator. Nat. Neurosci. 2003, 6, 251–257. [Google Scholar] [CrossRef]
- Lin, J.-M.; Kilman, V.L.; Keegan, K.; Paddock, B.; Emery-Le, M.; Rosbash, M.; Allada, R. A role for casein kinase 2α in the Drosophila circadian clock. Nature 2002, 420, 816–820. [Google Scholar] [CrossRef]
- Smith, E.M.; Lin, J.-M.; Meissner, R.-A.; Allada, R. Dominant-negative CK2alpha induces potent effects on circadian rhythmicity. PLoS Genet. 2008, 4, e12. [Google Scholar] [CrossRef]
- Lin, J.-M.; Schroeder, A.; Allada, R. In Vivo Circadian Function of Casein Kinase 2 Phosphorylation Sites in Drosophila PERIOD. J. Neurosci. 2005, 25, 11175–11183. [Google Scholar] [CrossRef]
- Meissner, R.-A.; Kilman, V.L.; Lin, J.-M.; Allada, R. TIMELESS is an important mediator of CK2 effects on circadian clock function in vivo. J. Neurosci. 2008, 28, 9732–9740. [Google Scholar] [CrossRef]
- Szabó, A.; Papin, C.; Zorn, D.; Ponien, P.; Weber, F.; Raabe, T.; Rouyer, F. The CK2 kinase stabilizes CLOCK and represses its activity in the Drosophila circadian oscillator. PLoS Biol. 2013, 11, e1001645. [Google Scholar] [CrossRef]
- Martinek, S.; Inonog, S.; Manoukian, A.S.; Young, M.W. A role for the segment polarity gene shaggy/GSK-3 in the Drosophila circadian clock. Cell 2001, 105, 769–779. [Google Scholar] [CrossRef]
- Top, D.; Harms, E.; Syed, S.; Adams, E.L.; Saez, L. GSK-3 and CK2 Kinases Converge on Timeless to Regulate the Master Clock. Cell Rep. 2016, 16, 357–367. [Google Scholar] [CrossRef]
- Ko, H.W.; Kim, E.Y.; Chiu, J.; Vanselow, J.T.; Kramer, A.; Edery, I. A hierarchical phosphorylation cascade that regulates the timing of PERIOD nuclear entry reveals novel roles for proline-directed kinases and GSK-3beta/SGG in circadian clocks. J. Neurosci. 2010, 30, 12664–12675. [Google Scholar] [CrossRef]
- Yu, W.; Houl, J.H.; Hardin, P.E. NEMO kinase contributes to core period determination by slowing the pace of the Drosophila circadian oscillator. Curr. Biol. 2011, 21, 756–761. [Google Scholar] [CrossRef]
- Levine, J.D.; Casey, C.I.; Kalderon, D.D.; Jackson, F.R. Altered circadian pacemaker functions and cyclic AMP rhythms in the Drosophila learning mutant dunce. Neuron 1994, 13, 967–974. [Google Scholar] [CrossRef]
- Majercak, J.; Kalderon, D.; Edery, I. Drosophila melanogaster deficient in protein kinase A manifests behavior-specific arrhythmia but normal clock function. Mol. Cell. Biol. 1997, 17, 5915–5922. [Google Scholar] [CrossRef]
- Park, S.K.; Sedore, S.A.; Cronmiller, C.; Hirsh, J. Type II cAMP-dependent protein kinase-deficient Drosophila are viable but show developmental, circadian, and drug response phenotypes. J. Biol. Chem. 2000, 275, 20588–20596. [Google Scholar] [CrossRef]
- Dusik, V.; Senthilan, P.R.; Mentzel, B.; Hartlieb, H.; Wülbeck, C.; Yoshii, T.; Raabe, T.; Helfrich-Förster, C. The MAP kinase p38 is part of Drosophila melanogaster’s circadian clock. PLoS Genet. 2014, 10, e1004565. [Google Scholar] [CrossRef]
- Vrailas-Mortimer, A.D.; Ryan, S.M.; Avey, M.J.; Mortimer, N.T.; Dowse, H.; Sanyal, S. p38 MAP kinase regulates circadian rhythms in Drosophila. J. Biol. Rhythms 2014, 29, 411–426. [Google Scholar] [CrossRef]
- Akten, B.; Tangredi, M.M.; Jauch, E.; Roberts, M.A.; Ng, F.; Raabe, T.; Jackson, F.R. Ribosomal s6 kinase cooperates with casein kinase 2 to modulate the Drosophila circadian molecular oscillator. J. Neurosci. 2009, 29, 466–475. [Google Scholar] [CrossRef]
- Beck, K.; Hovhanyan, A.; Menegazzi, P.; Helfrich-Förster, C.; Raabe, T. Drosophila RSK Influences the Pace of the Circadian Clock by Negative Regulation of Protein Kinase Shaggy Activity. Front. Mol. Neurosci. 2018, 11, 122. [Google Scholar] [CrossRef]
- Tangredi, M.M.; Ng, F.S.; Jackson, F.R. The C-terminal kinase and ERK-binding domains of Drosophila S6KII (RSK) are required for phosphorylation of the protein and modulation of circadian behavior. J. Biol. Chem. 2012, 287, 16748–16758. [Google Scholar] [CrossRef]
- Williams, J.A.; Su, H.S.; Bernards, A.; Field, J.; Sehgal, A. A circadian output in Drosophila mediated by neurofibromatosis-1 and Ras/MAPK. Science 2001, 293, 2251–2256. [Google Scholar] [CrossRef]
- Weber, F.; Hung, H.-C.; Maurer, C.; Kay, S.A. Second messenger and Ras/MAPK signalling pathways regulate CLOCK/CYCLE-dependent transcription. J. Neurochem. 2006, 98, 248–257. [Google Scholar] [CrossRef]
- Zheng, X.; Sehgal, A. AKT and TOR signaling set the pace of the circadian pacemaker. Curr. Biol. 2010, 20, 1203–1208. [Google Scholar] [CrossRef]
- Cho, E.; Kwon, M.; Jung, J.; Hyun Kang, D.; Jin, S.; Choi, S.-E.; Kang, Y.; Kim, E.Y. AMP-activated protein kinase regulates circadian rhythm by affecting CLOCK in Drosophila. J. Neurosci. 2019, 39, 3537–3550. [Google Scholar] [CrossRef]
- Fujii, S.; Emery, P.; Amrein, H. SIK3-HDAC4 signaling regulates Drosophila circadian male sex drive rhythm via modulating the DN1 clock neurons. Proc. Natl. Acad. Sci. USA 2017, 114, E6669–E6677. [Google Scholar] [CrossRef]
- Tanoue, S.; Krishnan, P.; Chatterjee, A.; Hardin, P.E. G protein-coupled receptor kinase 2 is required for rhythmic olfactory responses in Drosophila. Curr. Biol. 2008, 18, 787–794. [Google Scholar] [CrossRef]
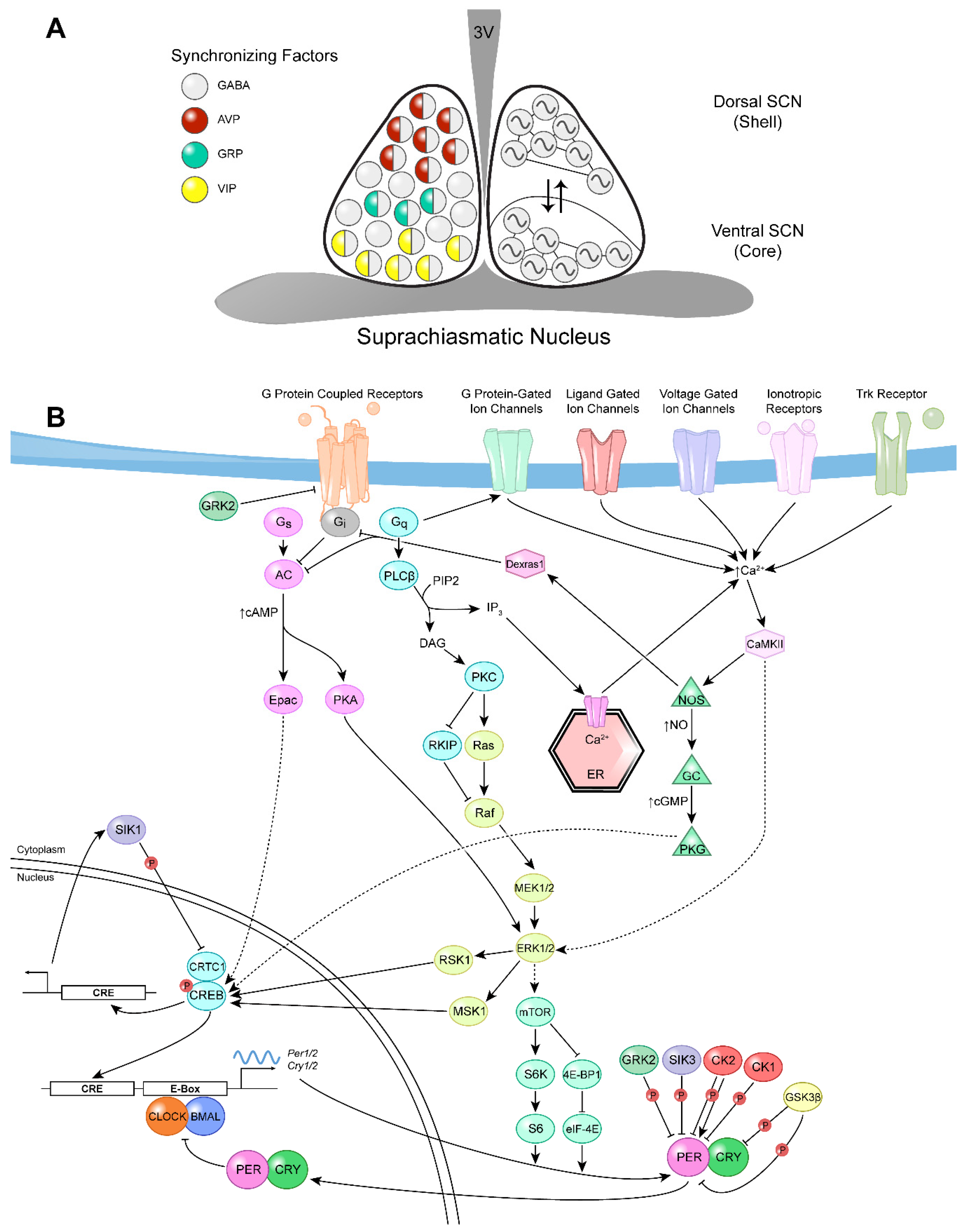
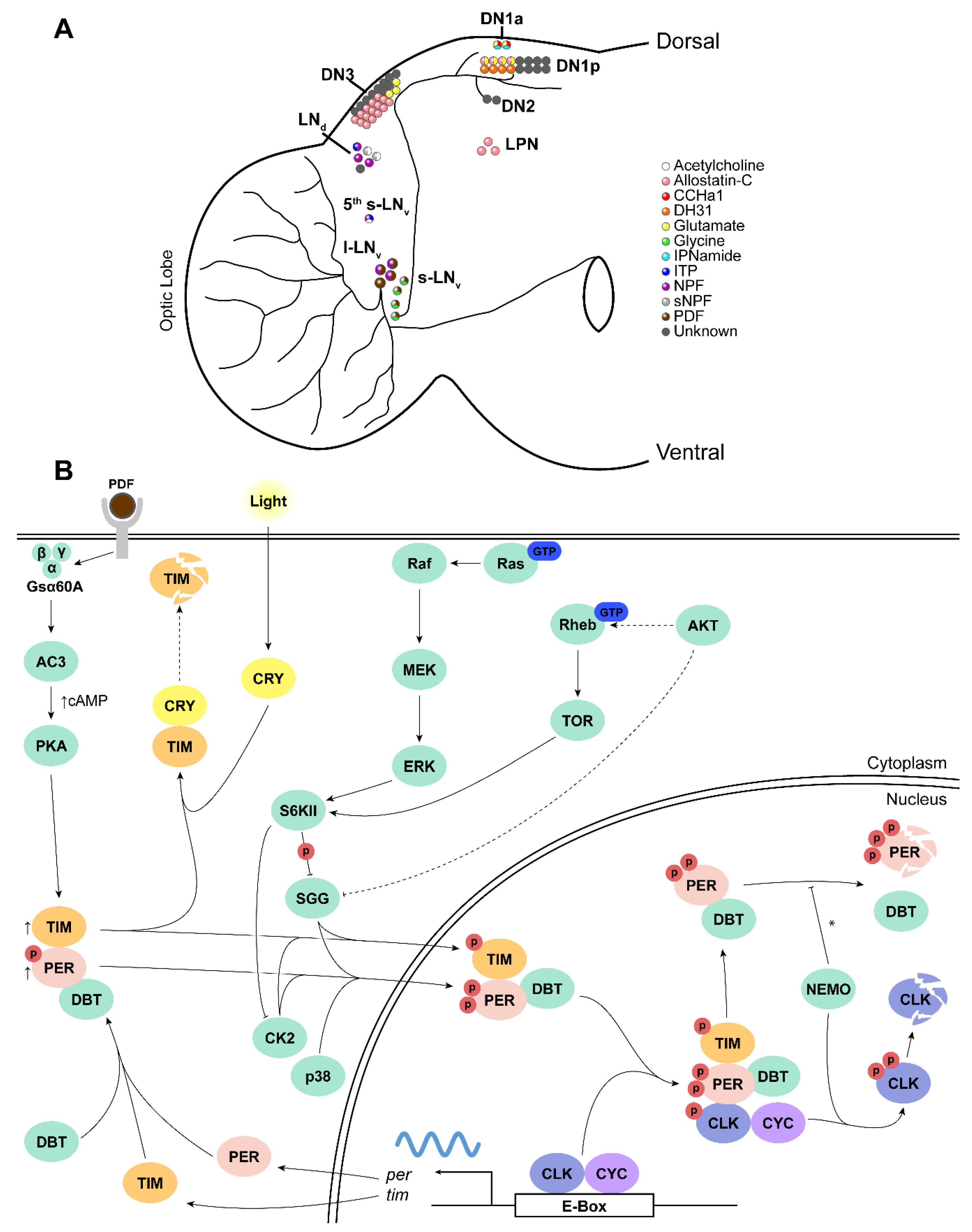
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hegazi, S.; Lowden, C.; Rios Garcia, J.; Cheng, A.H.; Obrietan, K.; Levine, J.D.; Cheng, H.-Y.M. A Symphony of Signals: Intercellular and Intracellular Signaling Mechanisms Underlying Circadian Timekeeping in Mice and Flies. Int. J. Mol. Sci. 2019, 20, 2363. https://doi.org/10.3390/ijms20092363
Hegazi S, Lowden C, Rios Garcia J, Cheng AH, Obrietan K, Levine JD, Cheng H-YM. A Symphony of Signals: Intercellular and Intracellular Signaling Mechanisms Underlying Circadian Timekeeping in Mice and Flies. International Journal of Molecular Sciences. 2019; 20(9):2363. https://doi.org/10.3390/ijms20092363
Chicago/Turabian StyleHegazi, Sara, Christopher Lowden, Julian Rios Garcia, Arthur H. Cheng, Karl Obrietan, Joel D. Levine, and Hai-Ying Mary Cheng. 2019. "A Symphony of Signals: Intercellular and Intracellular Signaling Mechanisms Underlying Circadian Timekeeping in Mice and Flies" International Journal of Molecular Sciences 20, no. 9: 2363. https://doi.org/10.3390/ijms20092363
APA StyleHegazi, S., Lowden, C., Rios Garcia, J., Cheng, A. H., Obrietan, K., Levine, J. D., & Cheng, H.-Y. M. (2019). A Symphony of Signals: Intercellular and Intracellular Signaling Mechanisms Underlying Circadian Timekeeping in Mice and Flies. International Journal of Molecular Sciences, 20(9), 2363. https://doi.org/10.3390/ijms20092363