Dynamic Chloroplast Genome Rearrangement and DNA Barcoding for Three Apiaceae Species Known as the Medicinal Herb “Bang-Poong”
Abstract
1. Introduction
2. Results
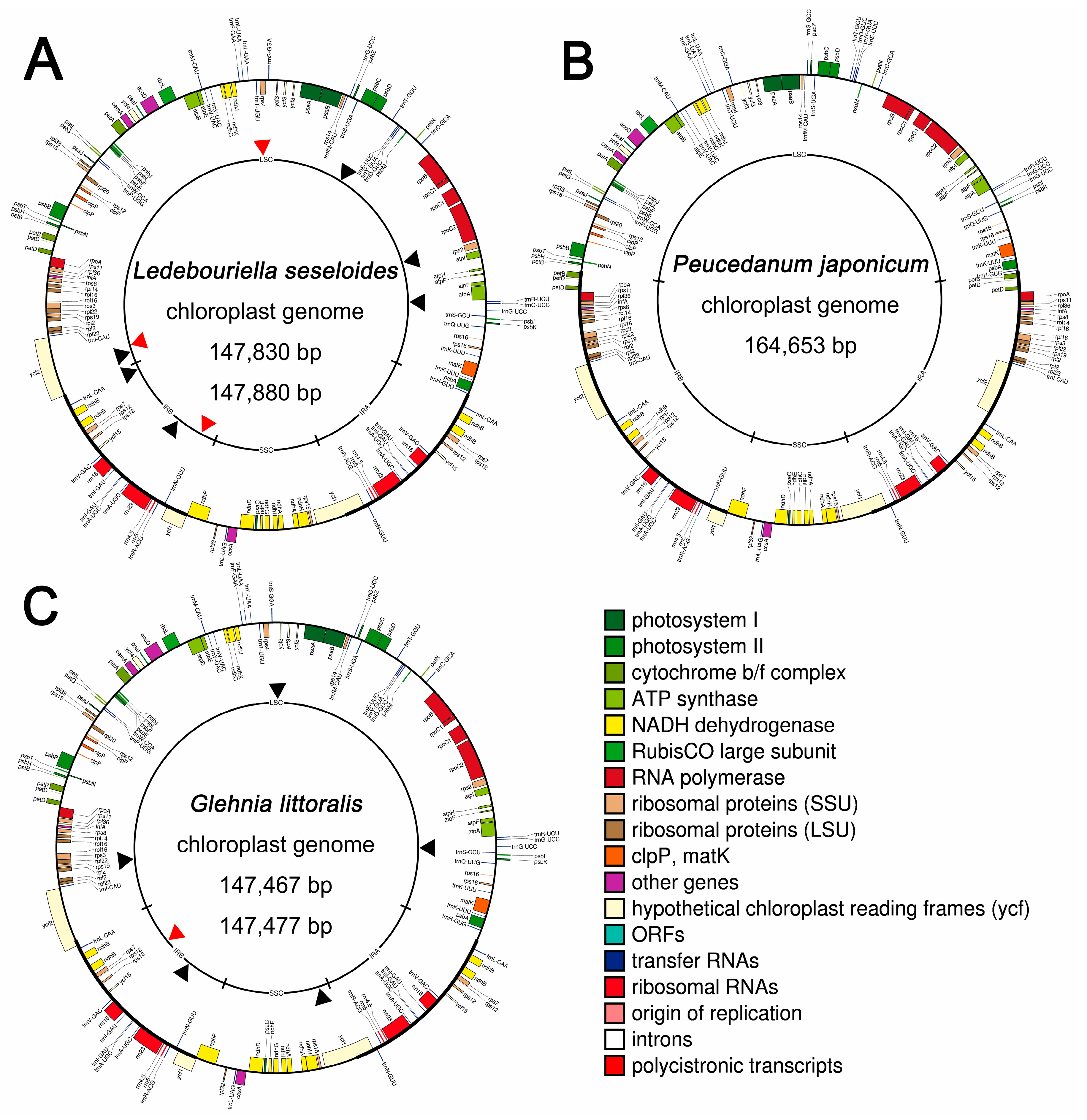
2.1. Six Complete Cp Genomes and 45S Nrdna Sequences of Three Apiaceae Species
2.2. Identification of Intraspecies Variations within Three Apiaceae Cp Genomes
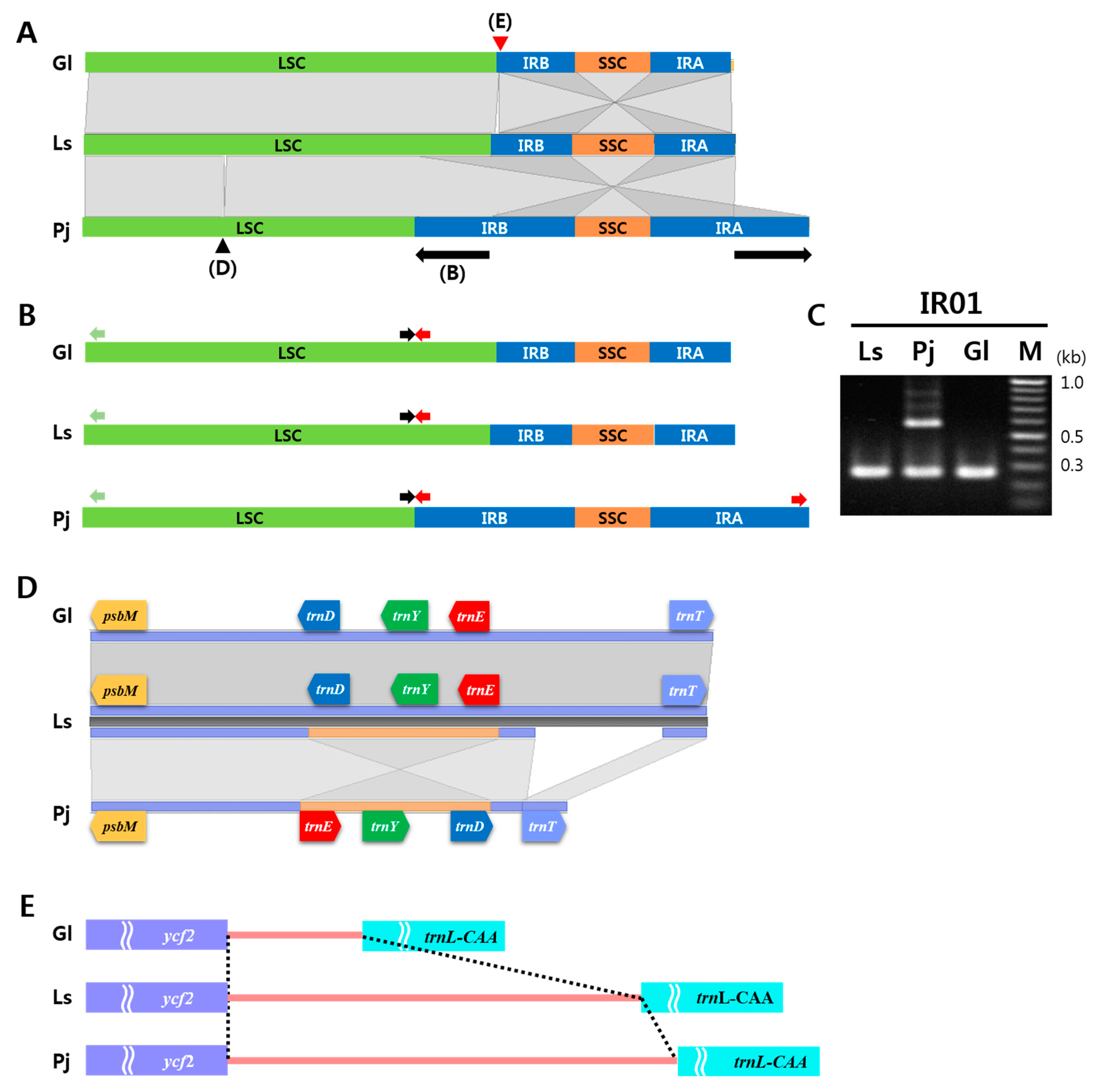
2.3. IR Expansion and Structural Variations among Three Apiaceae Species
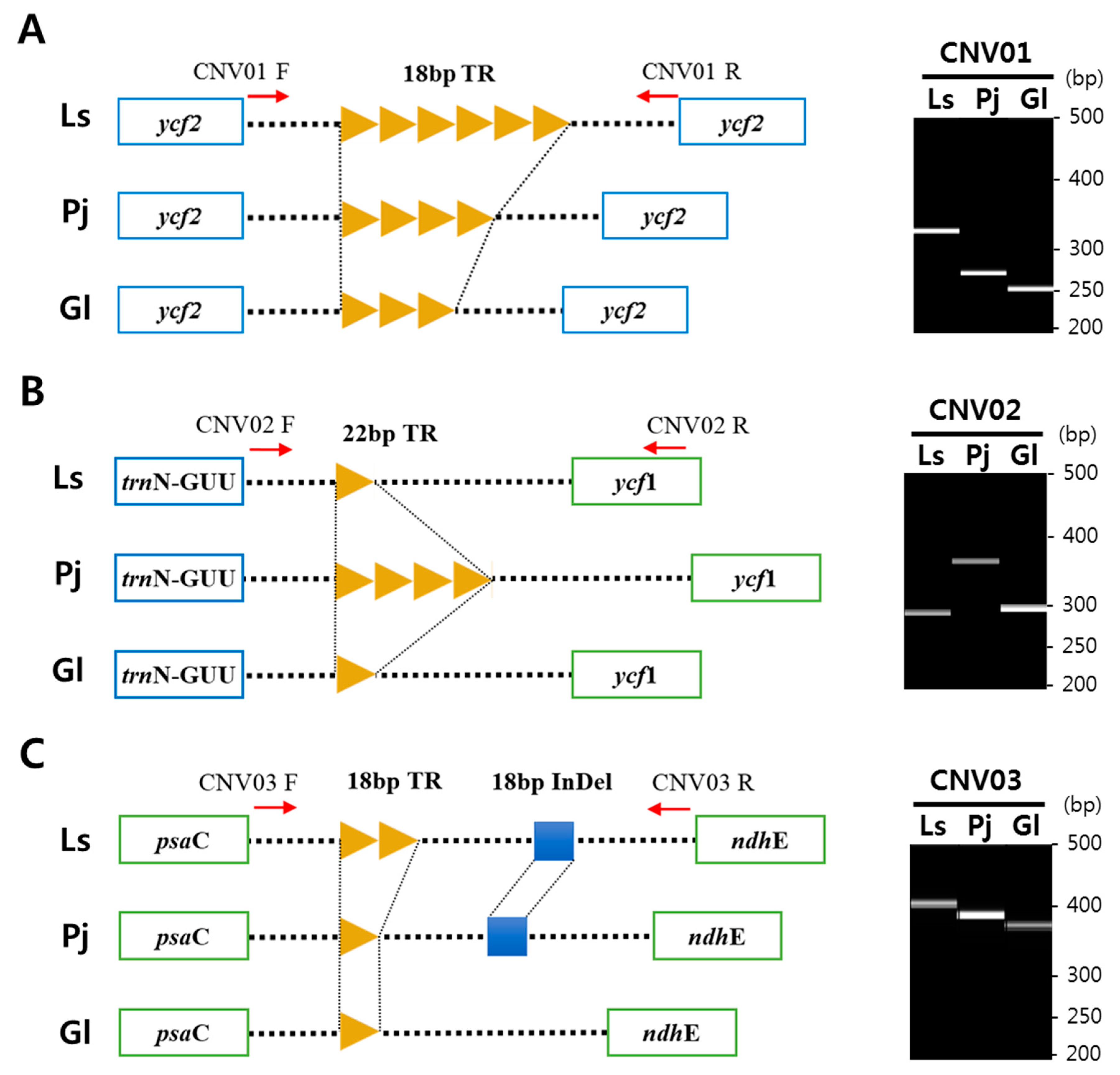
2.4. Tandem Repeats and Copy Number Variations in the Cp Genomes of Three Species
2.5. Sequence Variations of 45S DNA Sequences of Three Species
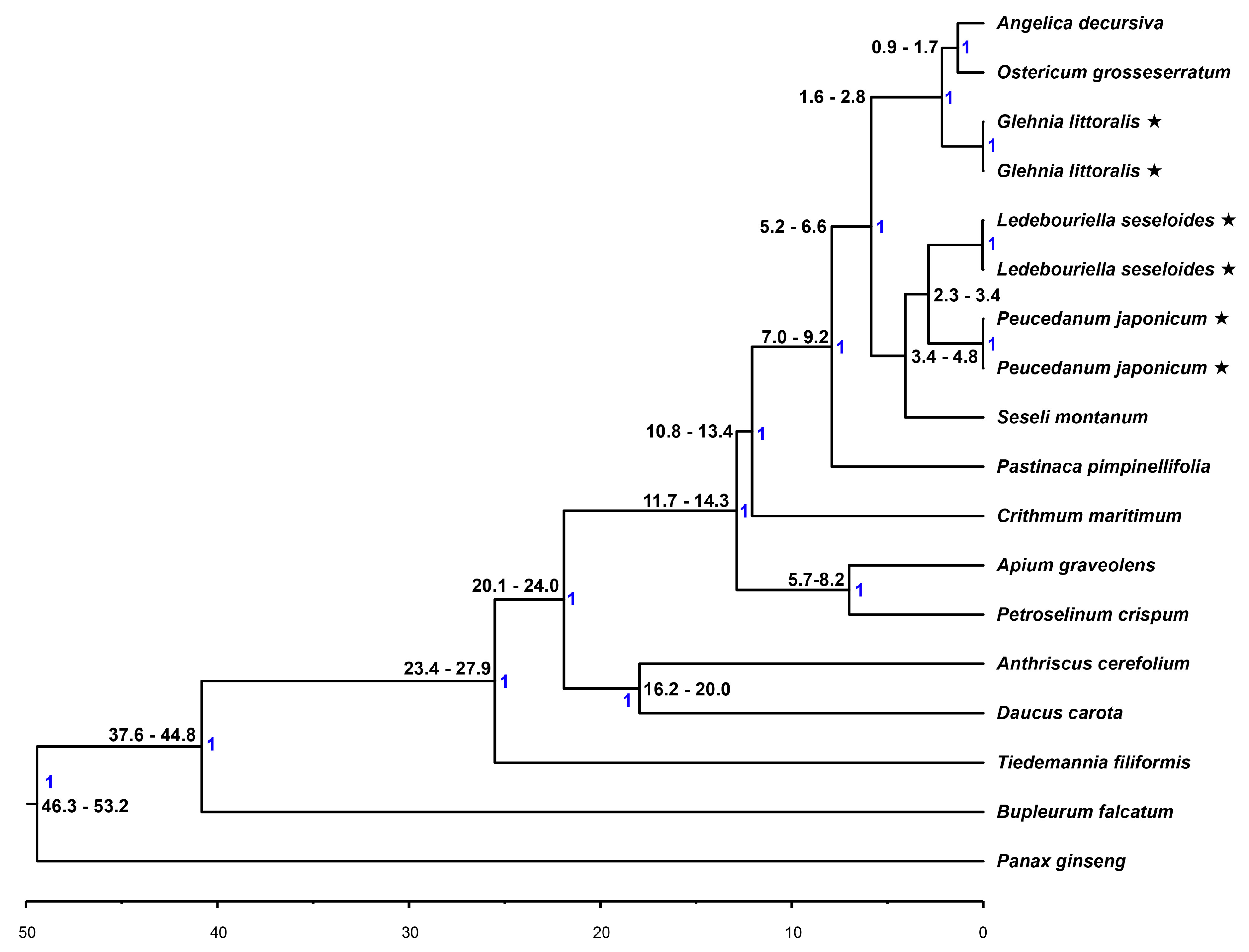
2.6. Phylogenetic Analysis and Divergence Time Estimation
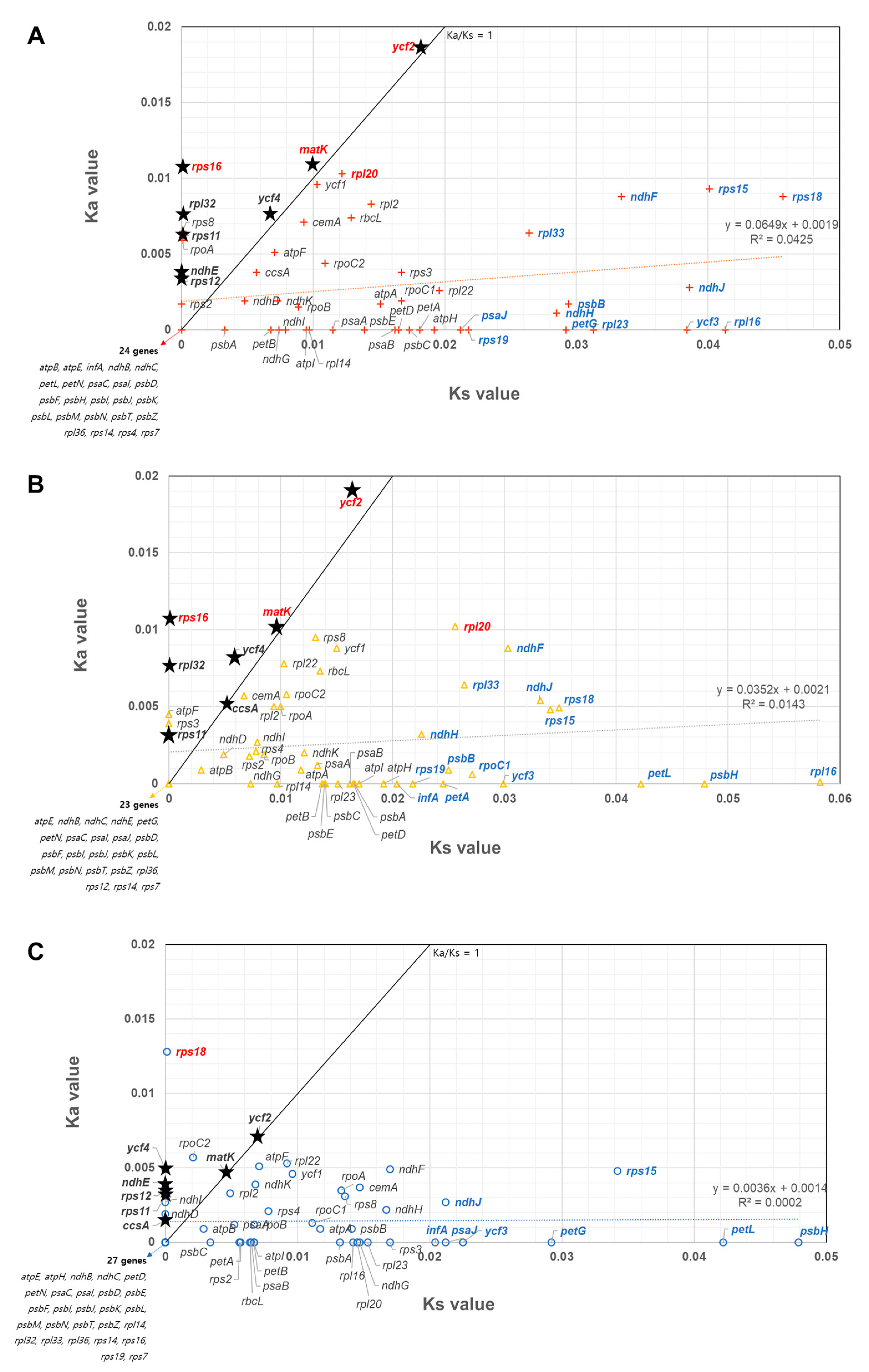
2.7. Comparison of Mutation Rate among All Cp Protein-Coding Genes in the Three Apiaceae Species
2.8. Development of Barcode Markers Derived from cp Genomes and 45S nrDNA Sequences
3. Discussion
3.1. Molecular Phylogeny of the Apiaceae Species
3.2. Intraspecies Chloroplast Variation and DNA Barcoding Markers for Species Authentication
3.3. Unique Structural Changes in the cp Genome of P. japonicum
3.4. Variation in the cp Genomes and 45S nrDNA of Three Species
3.5. Cp Gene Selection Pressure and Phylogenetic Relationship of Three Apiaceae Species
4. Materials and Methods
4.1. Plant Materials
4.2. Genomic DNA Preparation and Whole-Genome Shotgun Sequencing
4.3. Cp Genome and 45S nrDNA Assembly
4.4. Gene and Structural Annotation
4.5. Comparative Sequence Analysis at the Intraspecies and Interspecies Level
4.6. Development and Validation of DNA Barcode Markers
4.7. Calculation of Nucleotide Substitution Value
4.8. Phylogenetic Analysis
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Pimenov, M.G.; Leonov, M.V.E. The genera of the Umbelliferae: A nomenclator. In Kew Bulletin; Royal Botanic Gardens, Kew: Richmond, UK, 1993; Volume 49, pp. 592–593. [Google Scholar]
- Wu, Z.; Raven, P.H.; Hong, D. Flora of China. Volume 14: Apiaceae through Ericaceae; Science Press: Beijing, China, 2005; p. 581. [Google Scholar]
- Commission, C.P. Pharmacopoeia of the People’s Republic of China; China Medical Science and Technology Press: Beijing, China, 2015. [Google Scholar]
- Committee, J. The Japanese Pharmacopoeia, 16th ed.; The Ministry of Health, Labour and Welfare: Tokyo, Japan, 2011; p. 2319. [Google Scholar]
- Bae, K. The Medicinal Plants of Korea; Kyo-Hak Publishing: Seoul, Korea, 2000; p. 461. [Google Scholar]
- Hsu, H.; Chen, Y.; Sheu, S.; Hsu, C.; Chen, C.; Chang, H. Oriental Materia Medica: A Concise Guide; Oriental Healing Arts Institute: Long Beach, CA, USA, 1986; p. 582. [Google Scholar]
- Bell, C.R.; Constance, L. Chromosome numbers in Umbelliferae. II. Am. J. Bot. 1960, 47, 24–32. [Google Scholar] [CrossRef]
- Constance, L.; Chuang, T.I.; Bell, C.R. Chromosome numbers in Umbelliferae. V. Am. J. Bot. 1976, 63, 608–625. [Google Scholar] [CrossRef]
- Ali, M.A.; Gyulai, G.; Hidvegi, N.; Kerti, B.; Al Hemaid, F.M.; Pandey, A.K.; Lee, J. The changing epitome of species identification–DNA barcoding. Saudi J. Biol. Sci. 2014, 21, 204–231. [Google Scholar]
- Sugiura, M. The chloroplast genome. Plant Mol. Biol. 1992, 19, 149–168. [Google Scholar] [CrossRef]
- Álvarez, I.; Wendel, J.F. Ribosomal ITS sequences and plant phylogenetic inference. Mol. Phylogenet. Evol. 2003, 29, 417–434. [Google Scholar] [CrossRef]
- Hollingsworth, M.L.; Andra Clark, A.; Forrest, L.L.; Richardson, J.; Pennington, R.T.; Long, D.G.; Cowan, R.; Chase, M.W.; Gaudeul, M.; Hollingsworth, P.M. Selecting barcoding loci for plants: Evaluation of seven candidate loci with species-level sampling in three divergent groups of land plants. Mol. Ecol. Res. 2009, 9, 439–457. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.; Lee, S.-C.; Lee, J.; Yu, Y.; Yang, K.; Choi, B.-S.; Koh, H.-J.; Waminal, N.E.; Choi, H.-I.; Kim, N.-H. Complete chloroplast and ribosomal sequences for 30 accessions elucidate evolution of Oryza AA genome species. Sci. Rep. 2015, 5, 15655. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.; Lee, S.-C.; Lee, J.; Lee, H.O.; Joh, H.J.; Kim, N.-H.; Park, H.-S.; Yang, T.-J. Comprehensive survey of genetic diversity in chloroplast genomes and 45S nrDNAs within Panax ginseng species. PLoS ONE 2015, 10, e0117159. [Google Scholar] [CrossRef] [PubMed]
- Drude, O. Die naturlichen Pflanzenfamilien (Engler, A.) 3. In Umbelliferae; Verlag von Wilhelm Engelmann: Leipizig, German, 1897; Volume 3, pp. 63–150. [Google Scholar]
- Downie, S.R.; Katz-Downie, D.S.; Watson, M.F. A phylogeny of the flowering plant family Apiaceae based on chloroplast DNA rpl16 and rpoC1 intron sequences: Towards a suprageneric classification of subfamily Apioideae. Am. J. Bot. 2000, 87, 273–292. [Google Scholar] [CrossRef]
- Joh, H.J.; Kim, N.-H.; Jayakodi, M.; Jang, W.; Park, J.Y.; Kim, Y.C.; In, J.-G.; Yang, T.-J. Authentication of golden-berry P. ginseng cultivar ‘Gumpoong’ from a landrace ‘Hwangsook’based on pooling method using chloroplast-derived markers. Plant Breed Biotech. 2017, 5, 16–24. [Google Scholar] [CrossRef]
- Nguyen, V.B.; Giang, V.N.L.; Waminal, N.E.; Park, H.-S.; Kim, N.-H.; Jang, W.; Lee, J.; Yang, T.-J. Comprehensive comparative analysis of chloroplast genomes from seven Panax species and development of an authentication system based on species-unique single nucleotide polymorphism markers. J. Ginseng Res. 2018. [Google Scholar] [CrossRef]
- Palmer, J.D.; Thompson, W.F. Chloroplast DNA rearrangements are more frequent when a large inverted repeat sequence is lost. Cell 1982, 29, 537–550. [Google Scholar] [CrossRef]
- Goulding, S.E.; Olmstead, R.G.; Morden, C.W.; Wolfe, K.H. Ebb and flow of the chloroplast inverted repeat. Mol. Gen. Genet. MGG 1996, 252, 195–206. [Google Scholar] [CrossRef]
- Dugas, D.V.; Hernandez, D.; Koenen, E.J.; Schwarz, E.; Straub, S.; Hughes, C.E.; Jansen, R.K.; Nageswara-Rao, M.; Staats, M.; Trujillo, J.T.; et al. Mimosoid legume plastome evolution: IR expansion, tandem repeat expansions, and accelerated rate of evolution in clpP. Sci. Rep. 2015, 5, 16958. [Google Scholar] [CrossRef]
- Park, S.; An, B.; Park, S. Reconfiguration of the plastid genome in Lamprocapnos spectabilis: IR boundary shifting, inversion, and intraspecific variation. Sci. Rep. 2018, 8, 13568. [Google Scholar] [CrossRef]
- Dong, W.; Liu, J.; Yu, J.; Wang, L.; Zhou, S. Highly variable chloroplast markers for evaluating plant phylogeny at low taxonomic levels and for DNA barcoding. PLoS ONE 2012, 7, e35071. [Google Scholar] [CrossRef]
- Dong, W.; Xu, C.; Li, C.; Sun, J.; Zuo, Y.; Shi, S.; Cheng, T.; Guo, J.; Zhou, S. ycf1, the most promising plastid DNA barcode of land plants. Sci. Rep. 2015, 5, 8348. [Google Scholar] [CrossRef]
- Chen, S.; Yao, H.; Han, J.; Liu, C.; Song, J.; Shi, L.; Zhu, Y.; Ma, X.; Gao, T.; Pang, X. Validation of the ITS2 region as a novel DNA barcode for identifying medicinal plant species. PLoS ONE 2010, 5, e8613. [Google Scholar] [CrossRef] [PubMed]
- Kim, C.-K.; Seol, Y.-J.; Perumal, S.; Lee, J.; Waminal, N.E.; Jayakodi, M.; Lee, S.-C.; Jin, S.; Choi, B.-S.; Yu, Y. Re-exploration of U’s Triangle Brassica Species Based on Chloroplast Genomes and 45S nrDNA Sequences. Sci. Rep. 2018, 8, 7353. [Google Scholar] [CrossRef] [PubMed]
- Wolfe, K.H.; Li, W.H.; Sharp, P.M. Rates of nucleotide substitution vary greatly among plant mitochondrial, chloroplast, and nuclear DNAs. Proc. Natl. Acad. Sci. USA 1987, 84, 9054–9058. [Google Scholar] [CrossRef]
- Plunkett, G.M.; Soltis, D.E.; Soltis, P.S. Evolutionary patterns in Apiaceae: Inferences based on matK sequence data. System. Bot. 1996, 477–495. [Google Scholar] [CrossRef]
- Plunkett, G.M.; Soltis, D.E.; Soltis, P.S. Clarification of the relationship between Apiaceae and Araliaceae based on matK and rbcL sequence data. Am. J. Bot. 1997, 84, 565–580. [Google Scholar] [CrossRef]
- Downie, S.R.; Ramanath, S.; Katz-Downie, D.S.; Llanas, E. Molecular systematics of Apiaceae subfamily Apioideae: Phylogenetic analyses of nuclear ribosomal DNA internal transcribed spacer and plastid rpoC1 intron sequences. Am. J. Bot. 1998, 85, 563–591. [Google Scholar] [CrossRef]
- WANG, C.B.; MA, X.G.; HE, X.J. A taxonomic re-assessment in the Chinese Bupleurum (Apiaceae): Insights from morphology, nuclear ribosomal internal transcribed spacer, and chloroplast (trnH-psbA, matK) sequences. J. System. Evol. 2011, 49, 558–589. [Google Scholar] [CrossRef]
- Feist, M.A.E.; Downie, S.R.; Magee, A.R.; Liu, M.R. Revised generic delimitations for Oxypolis and Ptilimnium (Apiaceae) based on leaf morphology, comparative fruit anatomy, and phylogenetic analysis of nuclear rDNA ITS and cpDNA trnQ-trnK intergenic spacer sequence data. Taxon 2012, 61, 402–418. [Google Scholar] [CrossRef]
- Downie, S.R.; Katz-Downie, D.S. A molecular phylogeny of Apiaceae subfamily Apioideae: Evidence from nuclear ribosomal DNA internal transcribed spacer sequences. Am. J. Bot. 1996, 83, 234–251. [Google Scholar] [CrossRef]
- Downie, S.R.; Spalik, K.; Katz-Downie, D.S.; Reduron, J.-P. Major clades within Apiaceae subfamily Apioideae as inferred by phylogenetic analysis of nrDNA ITS sequences. Plant Div. Evol. 2010, 128, 111–136. [Google Scholar] [CrossRef]
- Liu, J.; Shi, L.; Han, J.; Li, G.; Lu, H.; Hou, J.; Zhou, X.; Meng, F.; Downie, S.R. Identification of species in the angiosperm family Apiaceae using DNA barcodes. Mol. Ecol. Res. 2014, 14, 1231–1238. [Google Scholar] [CrossRef]
- Allen, G.; Flores-Vergara, M.; Krasynanski, S.; Kumar, S.; Thompson, W. A modified protocol for rapid DNA isolation from plant tissues using cetyltrimethylammonium bromide. Nat. Protoc. 2006, 1, 2320. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.O.; Kim, K.; Lee, S.-C.; Lee, J.; Lee, J.; Kim, S.; Yang, T.-J. The complete chloroplast genome sequence of Ledebouriella seseloides (Hoffm.) H. Wolff. Mit. DNA Part A 2016, 27, 3498–3499. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.-C.; Oh Lee, H.; Kim, K.; Kim, S.; Yang, T.-J. The complete chloroplast genome sequence of the medicinal plant Glehnia littoralis F. Schmidt ex Miq.(Apiaceae). Mit. DNA Part A 2016, 27, 3674–3675. [Google Scholar] [CrossRef] [PubMed]
- Ruhlman, T.; Lee, S.-B.; Jansen, R.K.; Hostetler, J.B.; Tallon, L.J.; Town, C.D.; Daniell, H. Complete plastid genome sequence of Daucus carota: Implications for biotechnology and phylogeny of angiosperms. BMC Genom. 2006, 7, 222. [Google Scholar] [CrossRef] [PubMed]
- Kurtz, S.; Phillippy, A.; Delcher, A.L.; Smoot, M.; Shumway, M.; Antonescu, C.; Salzberg, S.L. Versatile and open software for comparing large genomes. Genome Biol. 2004, 5, R12. [Google Scholar] [CrossRef] [PubMed]
- Wyman, S.K.; Jansen, R.K.; Boore, J.L. Automatic annotation of organellar genomes with DOGMA. Bioinformatics 2004, 20, 3252–3255. [Google Scholar] [CrossRef]
- Lohse, M.; Drechsel, O.; Bock, R. OrganellarGenomeDRAW (OGDRAW): A tool for the easy generation of high-quality custom graphical maps of plastid and mitochondrial genomes. Cur. Genet. 2007, 52, 267–274. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef] [PubMed]
- Frazer, K.A.; Pachter, L.; Poliakov, A.; Rubin, E.M.; Dubchak, I. VISTA: Computational tools for comparative genomics. Nucl. Acids Res. 2004, 32, W273–W279. [Google Scholar] [CrossRef]
- Brudno, M.; Malde, S.; Poliakov, A.; Do, C.B.; Couronne, O.; Dubchak, I.; Batzoglou, S. Glocal alignment: Finding rearrangements during alignment. Bioinformatics 2003, 19 (Suppl. 1), i54–i62. [Google Scholar] [CrossRef]
- Schwartz, S.; Kent, W.J.; Smit, A.; Zhang, Z.; Baertsch, R.; Hardison, R.C.; Haussler, D.; Miller, W. Human–Mouse Alignments with BLASTZ. Genome Res. 2003, 13, 103–107. [Google Scholar] [CrossRef]
- Yang, Z. PAML: A program package for phylogenetic analysis by maximum likelihood. Comput. Appl. Biosci. 1997, 13, 555–556. [Google Scholar] [CrossRef] [PubMed]
- Bouckaert, R.; Heled, J.; Kühnert, D.; Vaughan, T.; Wu, C.-H.; Xie, D.; Suchard, M.A.; Rambaut, A.; Drummond, A.J. BEAST 2: A software platform for Bayesian evolutionary analysis. PLoS Comput. Biol. 2014, 10, e1003537. [Google Scholar] [CrossRef] [PubMed]
| Species (Common Name in Traditional Korean Medicine) | Accession | Raw Data (Gbp) | Chloroplast Genome | 45S nrDNA | ||||
|---|---|---|---|---|---|---|---|---|
| Length (bp) | Coverage (x) d | GenBank Accession No. (Reference) | Length (bp) e | Coverage (x) d | GenBank Accession No. (Reference) | |||
| Ledebouriella seseloides (Bang-poong) | Ls-01 a | 1.67 | 147,880 | 255.15 | KT153021 (Lee et al., 2015a) | 5815 | 659.94 | KX757774 (This study) |
| Ls-02 b | 5.05 | 147,830 | 177.95 | KU866529 (This study) | 5815 | 2550.62 | KX757775 (This study) | |
| Peucedanum japonicum (Sik-bang-poong) | Pj-01 a | 2.08 | 164,653 | 369.41 | KU866530 (This study) | 5815 | 943.43 | KX757776 (This study) |
| Pj-02 b | 5.18 | 164,653 | 945.25 | KU866531 (This study) | 5815 | 2418.35 | KX757777 (This study) | |
| Glehnia littoralis (Hae-bang-poong) | Gl-01 c | 1.06 | 147,467 | 146.53 | KT153022 (Lee et al., 2015b) | 5812 | 399.28 | KX757778 (This study) |
| Gl-02 b | 4.72 | 147,477 | 367.41 | KU866532 (This study) | 5812 | 1638.57 | KX757779 (This study) | |
| No. | TR Unit Sequence | Unit Length (bp) | Copy Number | Position | ||
|---|---|---|---|---|---|---|
| Ls | Pj | Gl | ||||
| 1 | AATAAGTAACTAG | 13 | 2 | 1 | 1 | trnK-UUU ~ rps16 |
| 2 | TTCTTCTATACAAA | 14 | 2 | 2 | 2 | rps16 ~ trnQ-UUG |
| 3 | AAAGATATGATTCATA | 16 | 2 | 2 | 1 | rps16 ~ trnQ-UUG |
| 4 | TAAAAAATATAAA | 13 | 2 | 1 | 1 | atpF ~ atpH |
| 5 | AGGAATCATTAAA | 13 | 2 | 1 | 1 | rps2 ~ rpoC2 |
| 6 | TATATTGTTATAAT | 14 | 1 | 1 | 2 | rpoB ~ trnC-GCA |
| 7 | TATAATATTAATAAG | 15 | 2 | nf | nf | trnE-UUC ~ trnT-GGU |
| 8 | TATATAGGAAGTATGAT | 17 | nf | 2 | 1 | trnS-UGA ~ lhbA |
| 9 | ATCTAGATAAGATTTATA | 18 | 2 | 1 | 1 | trnM-CAU ~ psbD |
| 10 | TTAAATGGTATTTATTAAT | 19 | 2 | 1 | 1 | trnS-UGA ~ lhbA |
| 11 | ATTAGCCATTACTAATAG | 18 | 1 | 2 | 1 | psaA ~ ycf3 |
| 12 | TAACCTAAGTGCAAAAATAGA | 21 | 1 | 2 | 1 | ycf3 ~ trnS-GGA |
| 13 | TATTCTATATATATATATTCTATATA | 26 | 1 | 2 | 1 | rps4 ~ trnT-UGU |
| 14 | TTTATATATATATATAT | 17 | 2 | 1 | 1 | ndhC ~ trnV-UAC |
| 15 | TTATCTTTATAATTTAA | 18 | 1 | 2 | 1 | accD ~ psaI |
| 16 | TATATGTATATTGA | 14 | 2 | 2 | 1 | psaI ~ ycf4 |
| 17 | TTATTCCAGTAAAA | 14 | 2 | 2 | 2 | cemA ~ petA |
| 18 | AAGGAAGTACTC | 12 | 1 | 2 | 1 | psbE ~ petL |
| 19 | ATTATATATAT | 11 | nf | 3 | 1 | trnW-CCA ~ trnP-UGG |
| 20 | TTATACAAGGTACTTAAATGTAAA | 24 | 2 | 1 | 1 | psaJ ~ rpl33 |
| 21 | TTCTATATAGAACATAATTAAATA | 24 | 2 | 2 | 2 | rpl33 ~ rps18 |
| 22 | TAAAGTTCCAACTAAAAAG | 19 | 2 | 1 | 2 | psbT ~ psbN |
| 23 | TCCCCTTTTT | 10 | 1 | 2 | 1 | rps11 ~ rpl36 |
| 24 | ATATTTTTAAATTG | 14 | 2 | 2 | 3 | rpl16 ~ rps3 |
| 25 | TCTTATAGAATTAGAATTGT | 20 | 2 | 1 | 1 | rpl16 ~ rps3 |
| 26 | CCTATTGCCGATACA | 15 | 2 | 2 | 2 | ycf2 |
| 27 | TAGTGACGATATTGATGC | 18 | 6/5 a | 4 | 3 | ycf2 (marker ID CNV01) |
| 28 | AAACTCTCTTCAAGAGTTATTAACACCAACCCGGTGTTC | 39 | nf | 2 | nf | ycf2 ~ trnL-CAA |
| 29 | TGGTGGAGATCAGAAAGAGAA | 21 | 2 | 1 | nf | ycf2 ~ trnL-CAA |
| 30 | TGTAATGTACTT | 12 | 2 | 1 | 1 | ycf2 ~ trnL-CAA |
| 31 | TCTTTTCTTCCGTGATGAACT | 21 | 3 | 1 | 1 | ycf15 |
| 32 | ACCGAAAGGAAAAGCGTGAA | 20 | 2 | 2 | 3 | trnI-GAU ~ trnI-GAU |
| 33 | CATTGTTCAACTCTTTGACAACACGAAAAAAC | 32 | 2 | 2 | 2 | rrn4.5 ~ rrn5 |
| 34 | AAAAAGAAATAAAT | 14 | 2 | 2 | 1 | trnR-ACG |
| 35 | TTCTATTTCTTTTCTATATATG | 22 | 1 | 4 | 1 | trnN-GUU ~ ycf1 (marker ID CNV02) |
| 36 | TATTAATATTAATAAAT | 17 | 2 | nf | nf | ycf1 ~ ndhF |
| 37 | TATATATGTATAAA | 14 | 2 | nf | nf | rpl32 ~ trnL-UAG |
| 38 | AAATATTAATCTACTTCT | 18 | 2 | 1 | 1 | psaC ~ ndhE (marker ID CNV03) |
| 39 | TTATGAATATAA | 12 | 2 | 2 | 2 | psaC ~ ndhE |
| 40 | TATTATTTTTTATTA | 15 | 2 | 1 | 1 | ndhA ~ orf188 |
| Marker ID | Primer Sequence (5′–3′) | Product Size a | Location |
|---|---|---|---|
| Ls/Pj/Gl (bp) | |||
| CNV01 | F: GGTCAAATACCTAGCGACAA | 308/272/254 | ycf2 |
| R: TTATGCAAGGAGACATTGCT | |||
| CNV02 | F: CATCCATATCCCAATTCCAT | 305/371/305 | trnN-GUU ~ ycf1 |
| R: GCTCGGAGAAGGAAGAGATA | |||
| CNV03 | F: CATTGAGTGCACCCTATACA | 404/386/368 | psaC ~ ndhE |
| R: TCGTAACAGAAAATCAACTCG | |||
| InDel01 | F: AACGAATCCTACGGTTTCTC | 266/289/269 | trnS-GCU ~ trnR-UCU |
| R: TGTCGAACAGGGATAATTTG | |||
| InDel02 | F: TCTCGCTTTTTAGTCAGTTTG | 380/379/416 | ndhF ~ rpl32 |
| R: GCCTAATGAAAAGCCTAATGA | |||
| InDel03 | F: CAGGAGGATAGCAAGTTACAA | 412/571/571 | rps12 ~ trnV-GAC |
| R: CAACGCCACTATTCTTGAAC | |||
| InDel04 | F: CTATATGTATATACAATAACGAATCA | 652/198/633 | trnE-UUC ~ trnT-GGU |
| R: GTTCAAGAATAGTGGCGTTG | |||
| IR01 | LSC P: CCTAGCTGCTGTTGAAGCTC | na/576/na | psbA |
| Control F: GACGACTGAGCCAACTTGAT | 262/262/262 | psbH ~ petB | |
| Control R: TCGAGACGTTCTTCAAACCA | |||
| nrDNA01 | Specific F: GTTAACAATTAGGGCGAGCA | na/na/291 | ITS1 |
| Control F: GCATCGATGAAGAACGTAGC | 78/78/78 | 5.8S | |
| Control R: GCGTTCAAAGACTCGATGGT |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lee, H.O.; Joh, H.J.; Kim, K.; Lee, S.-C.; Kim, N.-H.; Park, J.Y.; Park, H.-S.; Park, M.-S.; Kim, S.; Kwak, M.; et al. Dynamic Chloroplast Genome Rearrangement and DNA Barcoding for Three Apiaceae Species Known as the Medicinal Herb “Bang-Poong”. Int. J. Mol. Sci. 2019, 20, 2196. https://doi.org/10.3390/ijms20092196
Lee HO, Joh HJ, Kim K, Lee S-C, Kim N-H, Park JY, Park H-S, Park M-S, Kim S, Kwak M, et al. Dynamic Chloroplast Genome Rearrangement and DNA Barcoding for Three Apiaceae Species Known as the Medicinal Herb “Bang-Poong”. International Journal of Molecular Sciences. 2019; 20(9):2196. https://doi.org/10.3390/ijms20092196
Chicago/Turabian StyleLee, Hyun Oh, Ho Jun Joh, Kyunghee Kim, Sang-Choon Lee, Nam-Hoon Kim, Jee Young Park, Hyun-Seung Park, Mi-So Park, Soonok Kim, Myounghai Kwak, and et al. 2019. "Dynamic Chloroplast Genome Rearrangement and DNA Barcoding for Three Apiaceae Species Known as the Medicinal Herb “Bang-Poong”" International Journal of Molecular Sciences 20, no. 9: 2196. https://doi.org/10.3390/ijms20092196
APA StyleLee, H. O., Joh, H. J., Kim, K., Lee, S.-C., Kim, N.-H., Park, J. Y., Park, H.-S., Park, M.-S., Kim, S., Kwak, M., Kim, K.-y., Lee, W. K., & Yang, T.-J. (2019). Dynamic Chloroplast Genome Rearrangement and DNA Barcoding for Three Apiaceae Species Known as the Medicinal Herb “Bang-Poong”. International Journal of Molecular Sciences, 20(9), 2196. https://doi.org/10.3390/ijms20092196




