Proteomics Analysis of E. angustifolia Seedlings Inoculated with Arbuscular Mycorrhizal Fungi under Salt Stress
Abstract
1. Introduction
2. Results
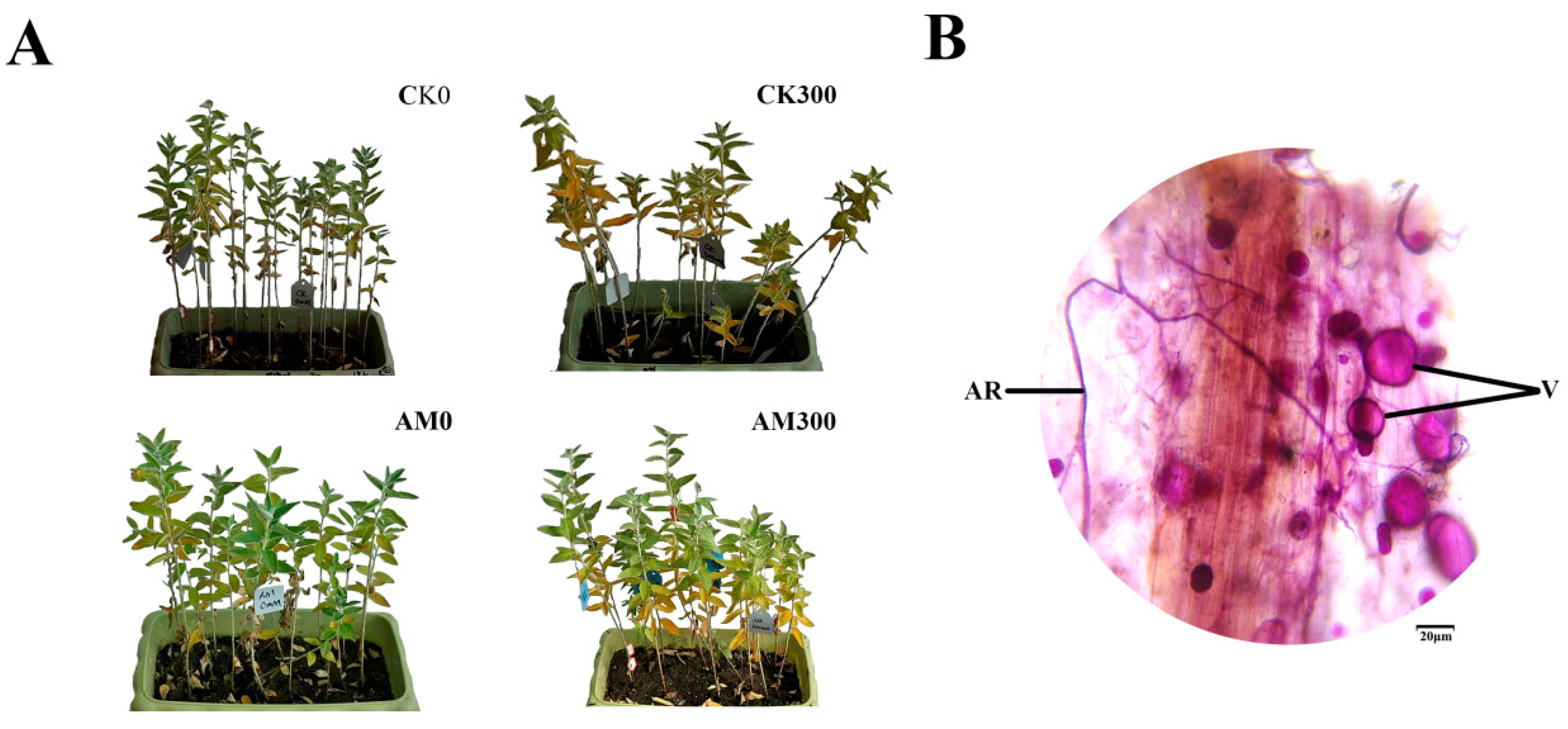
2.1. Growth of E. angustifolia under Salt Stress and Colonization of AMF in the Plant Roots
2.2. Effects of RI and CK on Height, Diameter, and Roots of E. angustifolia under Salt Stress
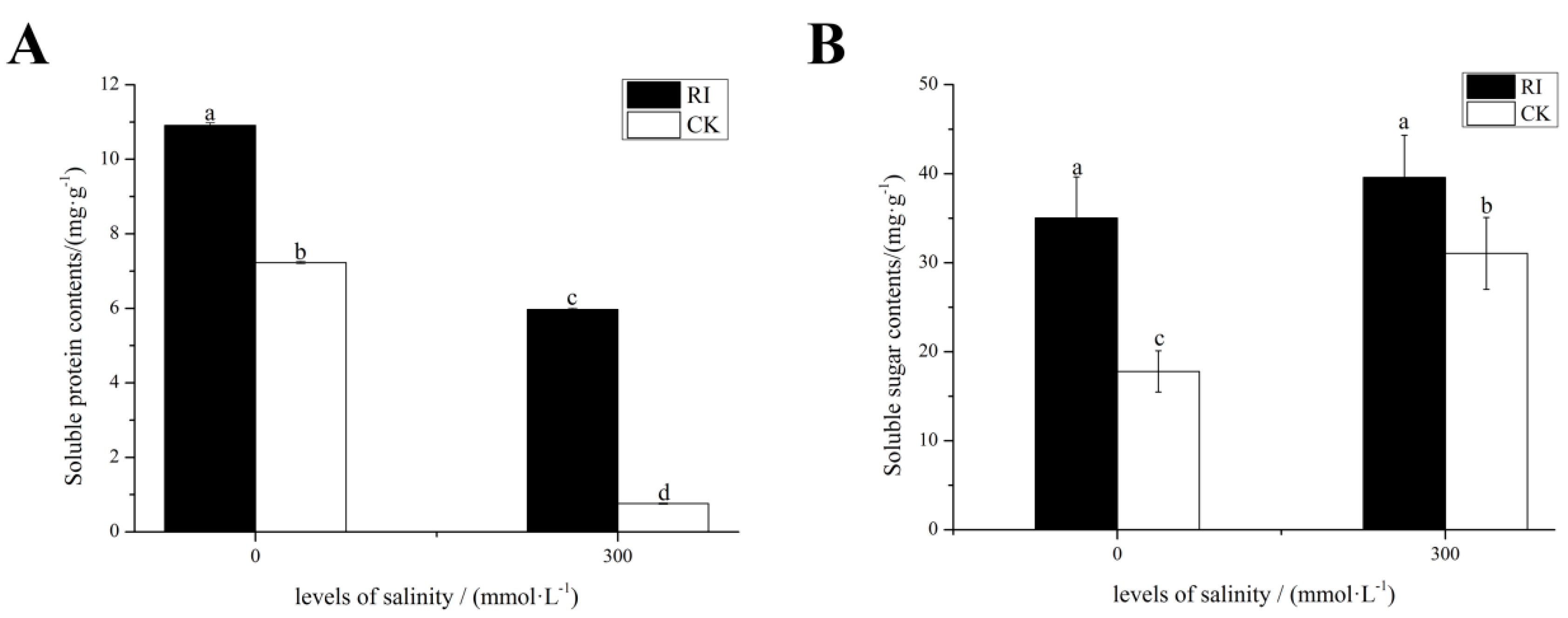
2.3. Effects of RI and CK on the Soluble Sugar Content, Soluble Protein Content in the Leaves of E. angustifolia under Salt Stress
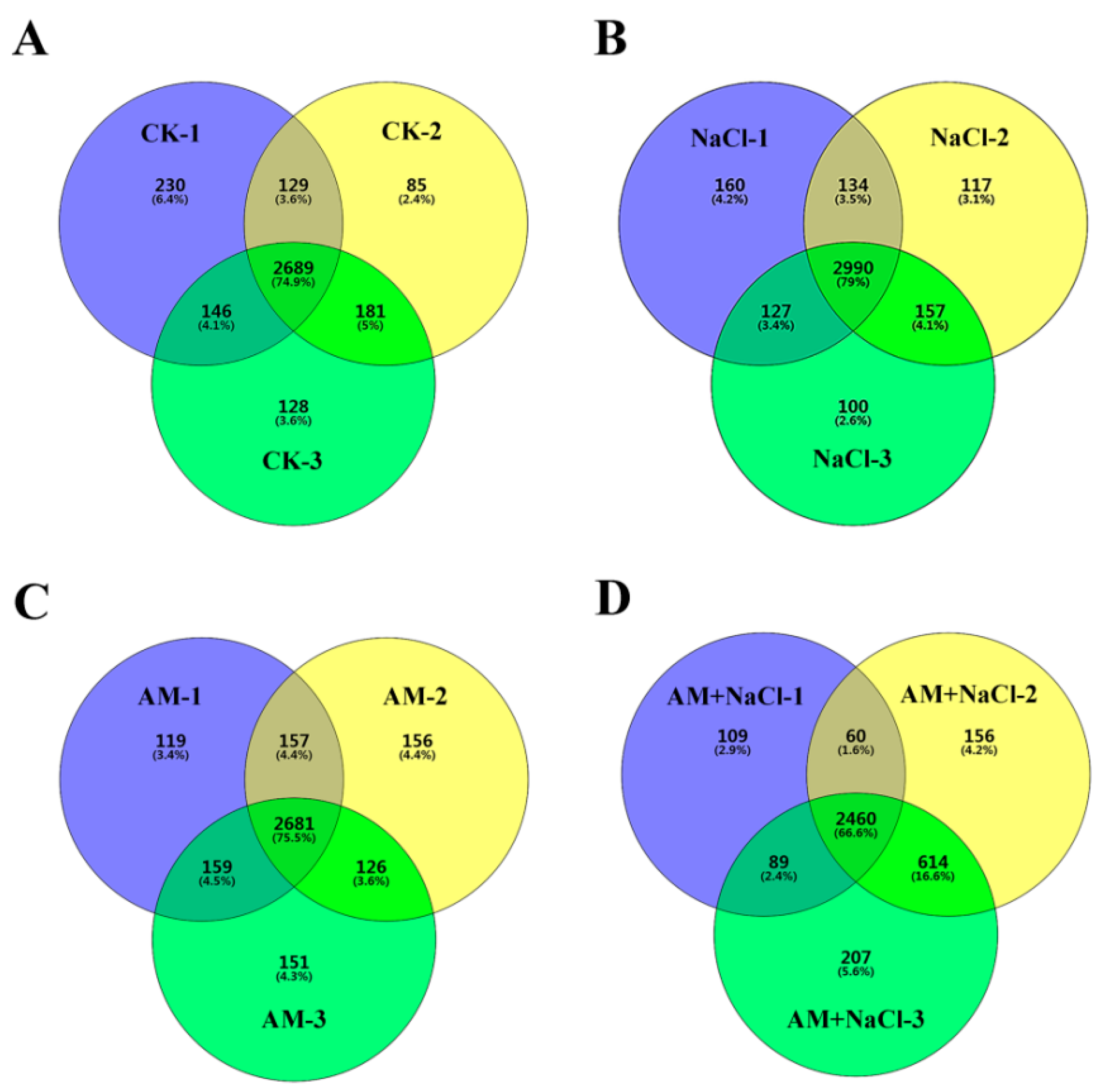
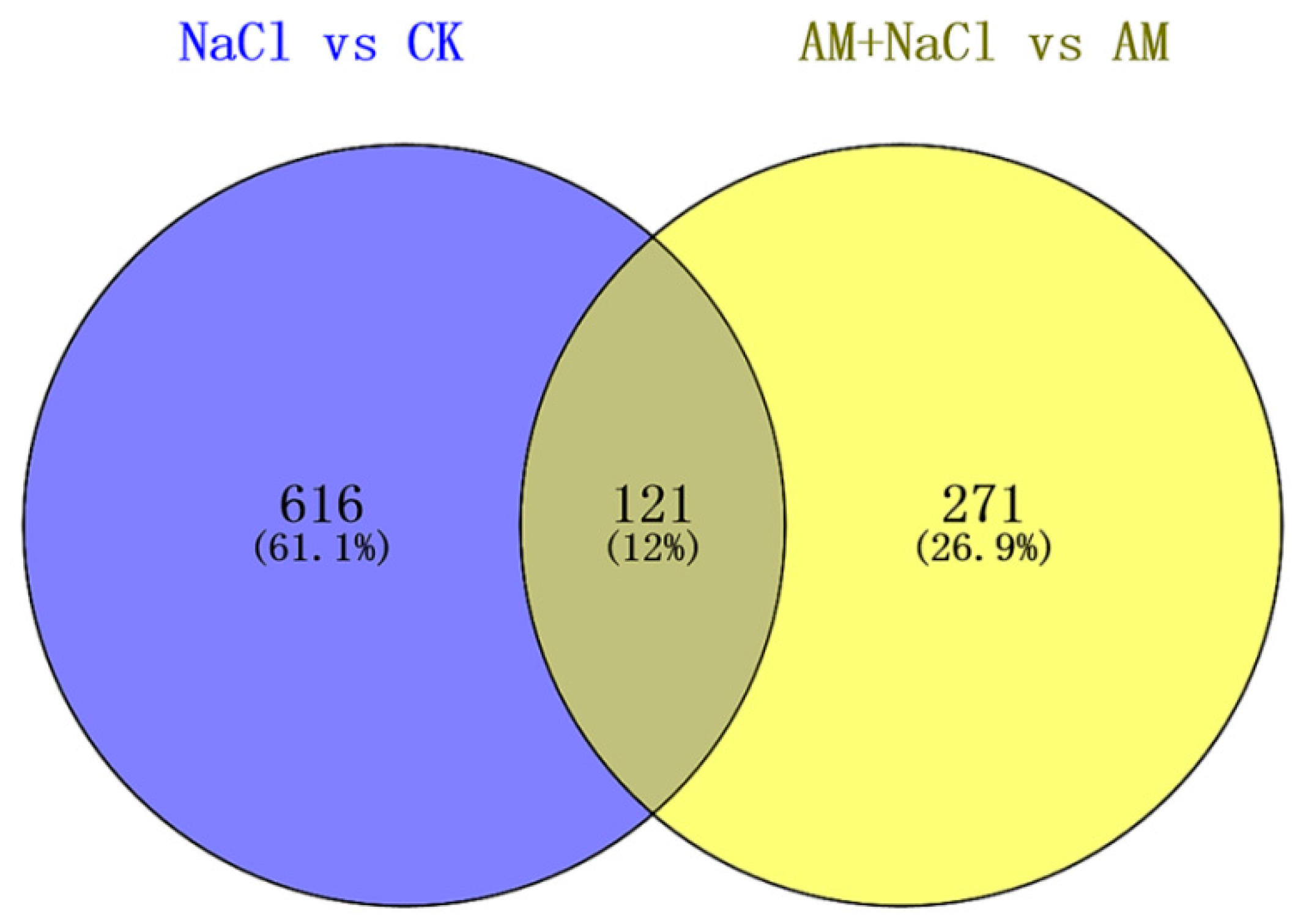
2.4. Effect of RI on Protein Abundance under Salt Stress
2.5. Functional Classification of Proteins
2.5.1. Salt Tolerance-Related Proteins Induced by Symbiosis
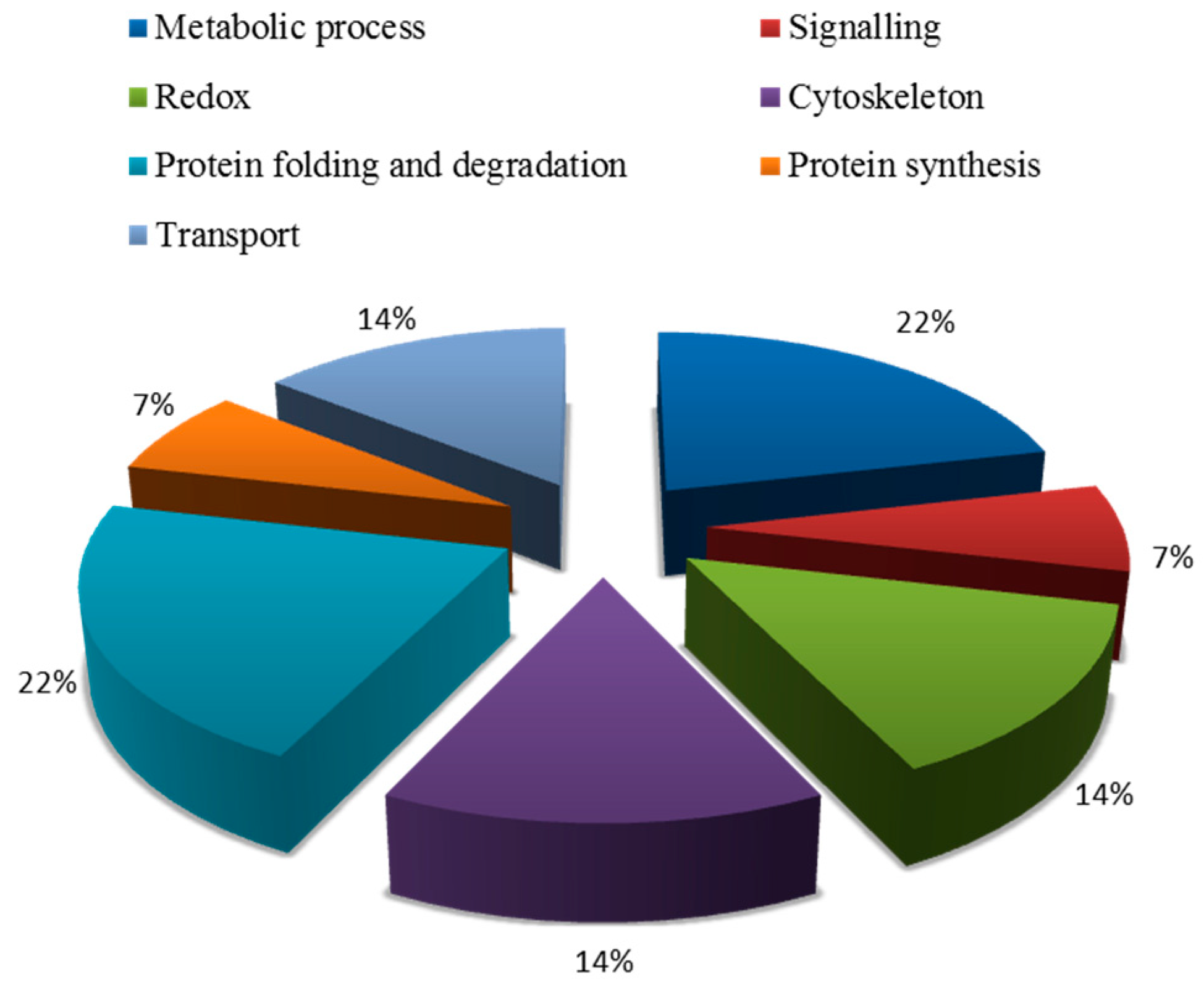
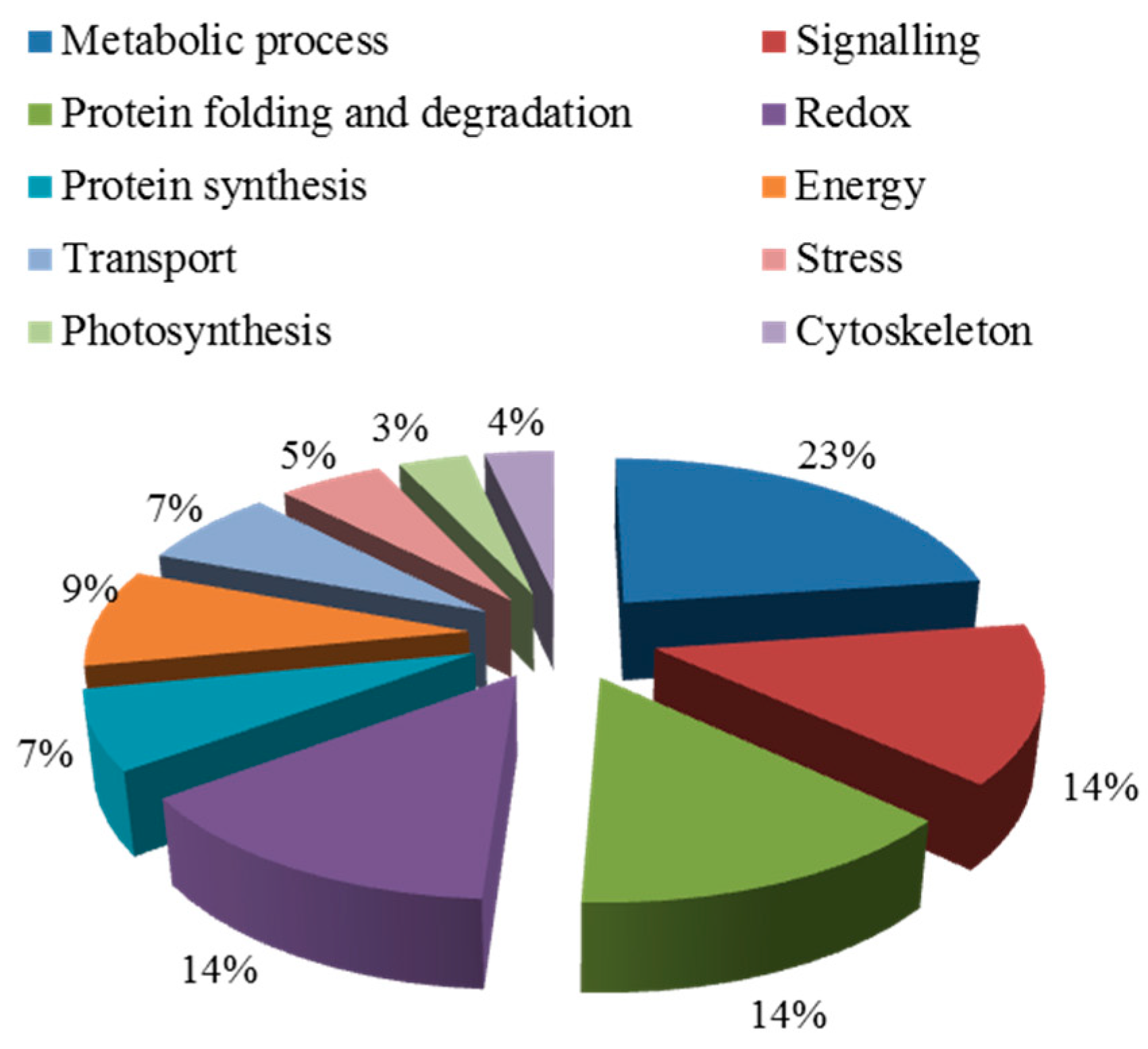
2.5.2. Functional Classification of Salt Tolerance-Related Proteins Induced by Symbiosis
2.5.3. Salt-Induced Mycorrhizal Protein
2.5.4. Functional Classification of Salt-Induced Mycorrhizal Protein
3. Discussion
3.1. Proteins Related to Metabolism
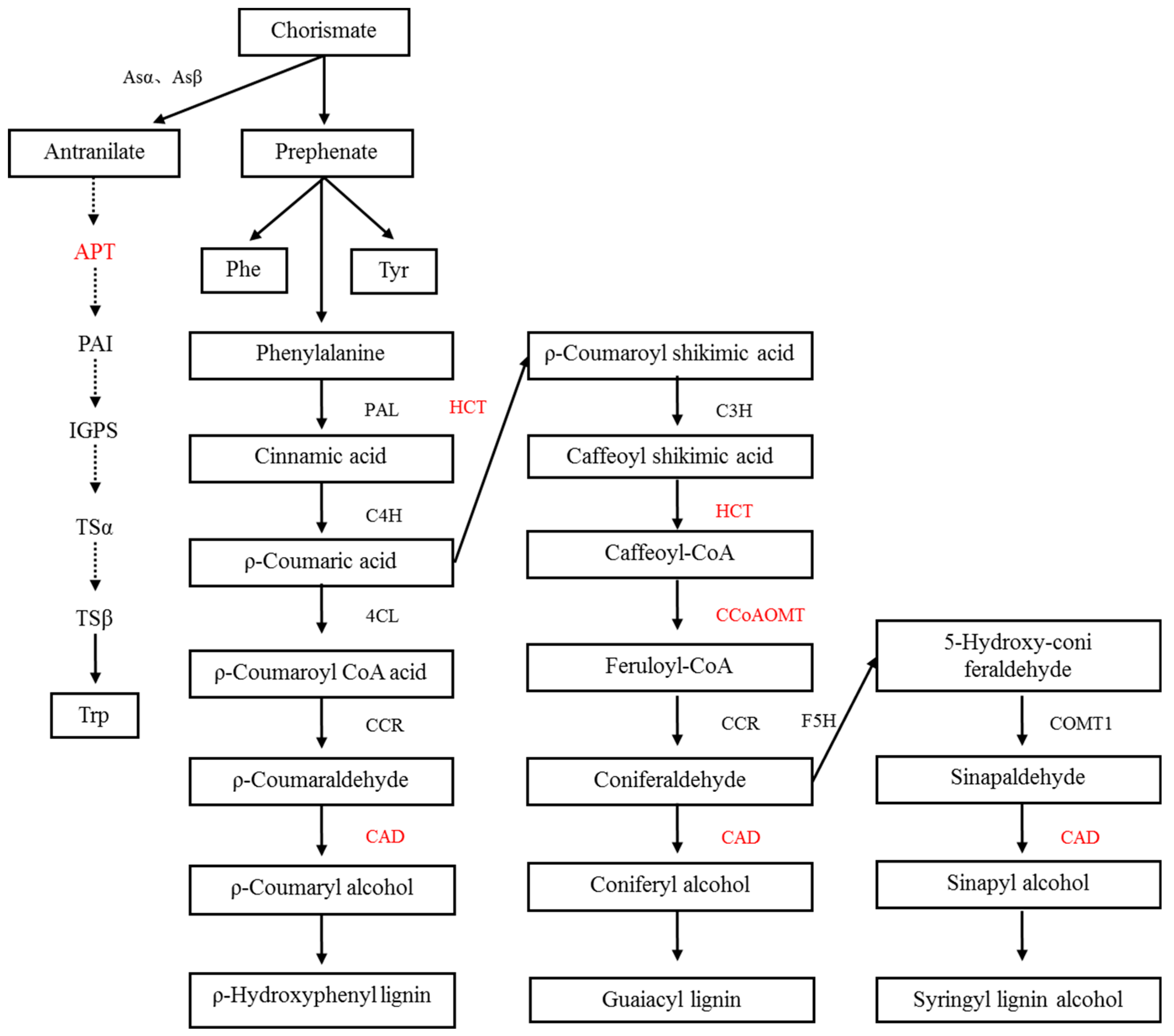
- In this study, we found a chorismate mutase (CM), which catalyzes the conversion of branched acid to prebenzoic acid. Prebenzoic acid can produce phenolic compounds through the phenylpropane metabolic pathway, including phenylalanine (Phe), tyrosine (Tyr), anthocyanin, and tannin [20,21]. Phenylpropane metabolism is indirectly generated by the shikimic acid pathway. This pathway might play an important role in plant stress defense. We found three proteins that relate to the phenylalanine metabolic pathways, including shikimate O-hydroxycinnamoyltransferase, cinnamyl alcohol dehydrogenase, and caffeoyl-CoA O-methyltransferase. Flavonoids are synthesized by the condensation of phenylpropane derivatives with malonate monoacyl coenzyme A. In addition, shikimic acid O-hydroxyacinnamate transferase and caffeoyl coenzyme A-O-methyltransferase were also involved.
- In this study, we found that the phosphoribosyltransferase (APT) was up-regulated, which was the first key enzyme in the tryptophan production reaction of o-aminobenzoic acid. Phosphorylribosyltransferase activity of o-aminobenzoic acid was enhanced, which accelerated the synthesis of tryptophan in plants under salt stress. It is well known that tryptophan is a precursor of auxin (indole acetic acid) as well as protein synthesis in plants. Auxin response was also identified in the leaves of E. angustifolia. We deduced that these two pathways synthesize auxin to maintain the growth and metabolism of mycorrhizal plants under salt stress.
3.2. Protein Synthesis, Folding, and Degradation Related Proteins
3.3. Signal Transduction-Related Proteins
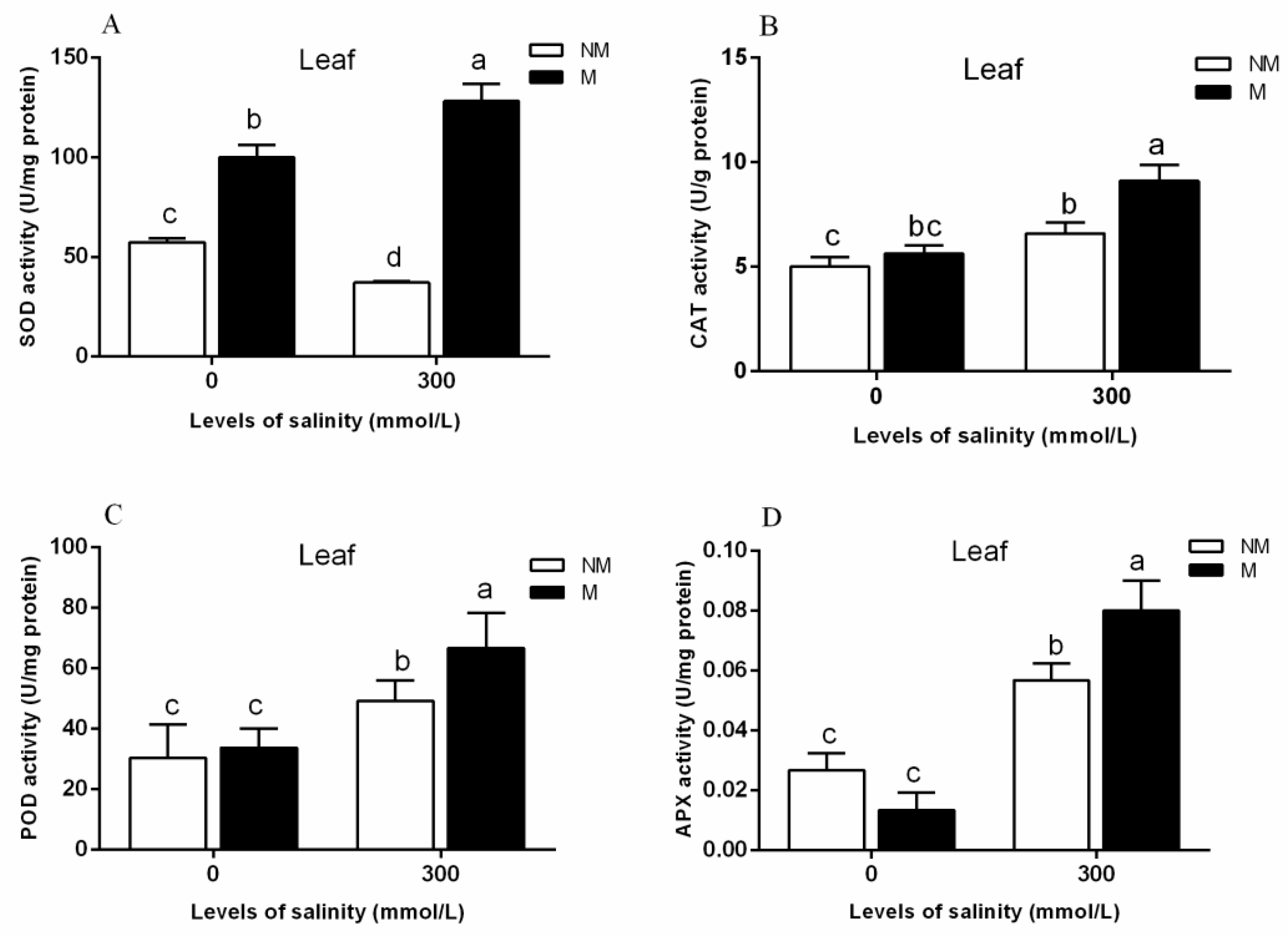
3.4. ROS Scavenging-Related Proteins
3.5. Energy-Related Protein
3.6. Photosynthesis-Related Proteins
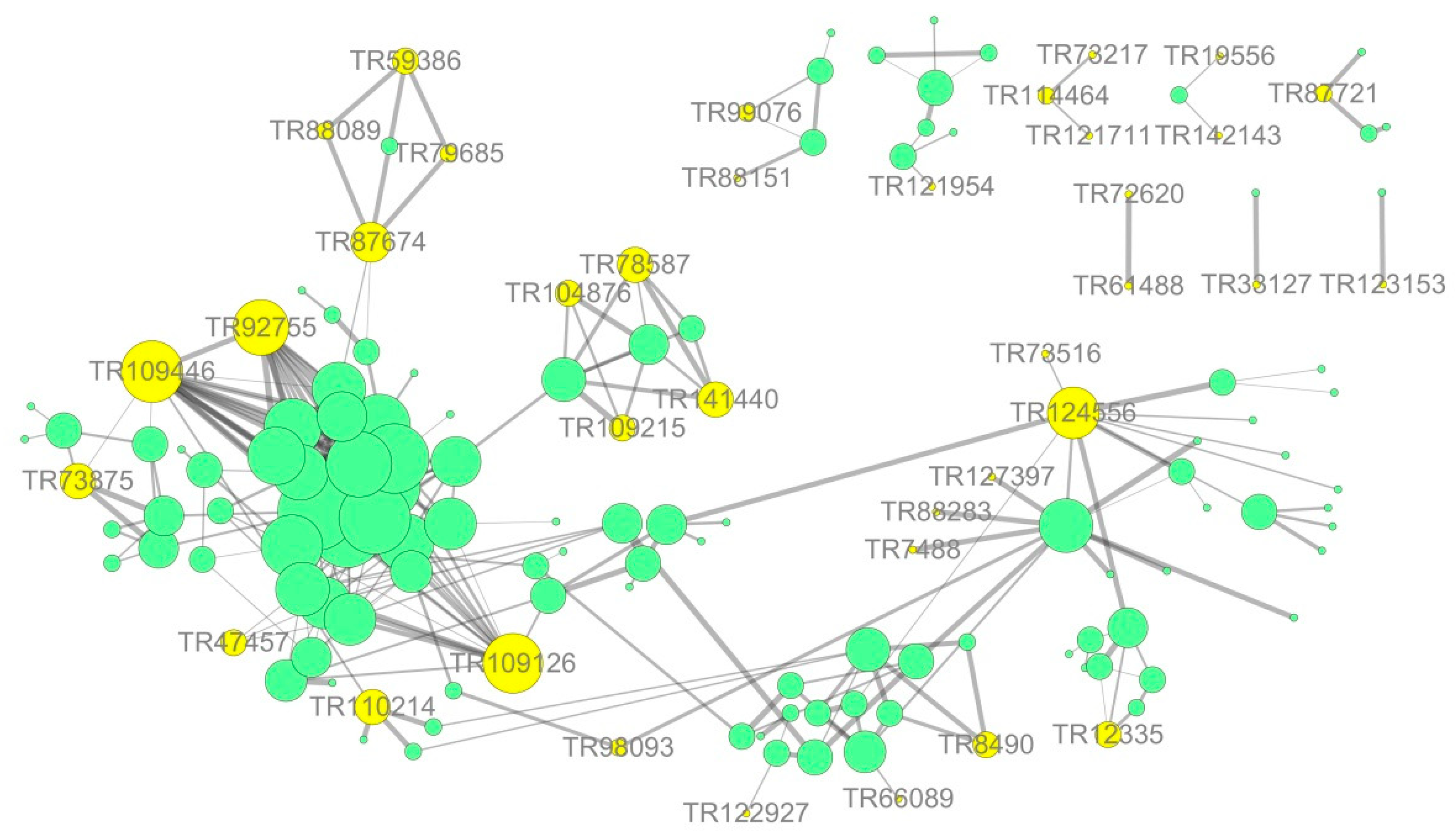
3.7. Network Interaction Predictions Based on Differential Expression
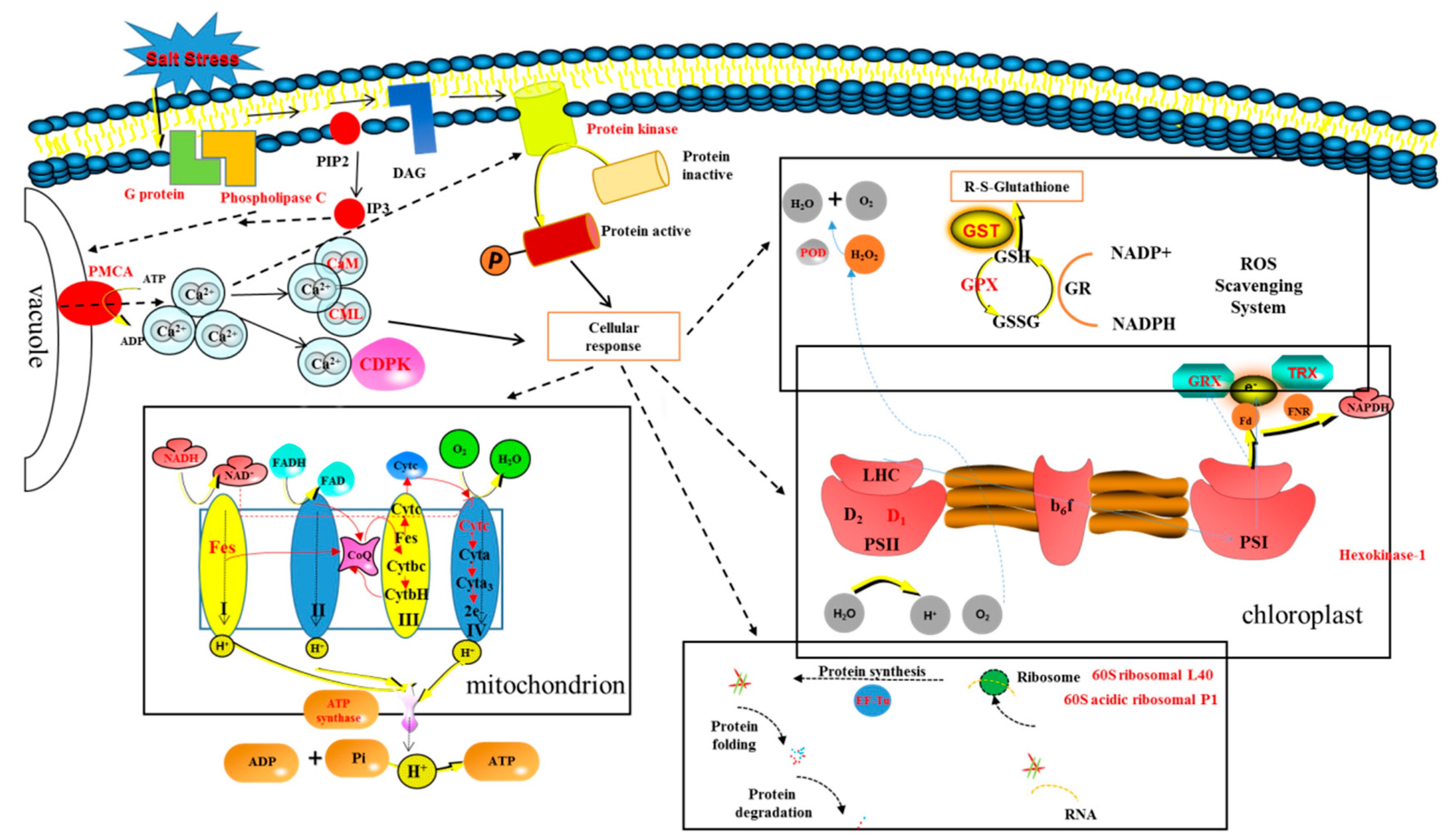
- AMF accelerates the secondary metabolism of plants, mainly phenylpropanoid metabolism (shikimate O-hydroxycinnamoyltransferase, cinnamyl alcohol dehydrogenase, and caffeoyl-CoA O-methyltransferase), reducing salt damage to plants.
- AMF enhances the signal transduction of the second messenger Ca2+ (G protein, phospholipase C, plasma membrane Ca2+ transporter ATPase (PMCA), calcium binding (CML), calcium-dependent kinases (CDPKs), and calmodulin (CaM), increasing the speed of sensing and transmitting of stress signals, allowing plants to follow up.
- Among a variety of antioxidant pathways (TRX/GRX, GST/GPX, POD), AMF enhances the antioxidant capacity of plants by increasing ROS clearance.
- AMF promotes protein biosynthesis, speeding up protein folding, and inhibiting protein degradation (ubiquitin-60S ribosomal, 60S acidic ribosomal, elongation factor mitochondrial, ubiquitin activating enzyme E1, and E3 ubiquitin- ligase).
- In the chloroplast, AMF maintains the PSII reaction centre conformation stability and speeds up photosynthetic electron transport (TRX/GRX, photosystem II D1 precursor processing PSB27- chloroplastic); in mitochondria, AMF enhances the synthesis of ATP (NADH dehydrogenase, cytochrome C oxidase, iron sulfur protein NADH dehydrogenase, and ATP synthase), providing sufficient energy for cellular activities.
4. Materials and Methods
4.1. Experimental Materials and Salinity Treatments
4.2. Measurement of AMF Colonization and Growth Parameter
4.3. Measurement of Soluble Sugar Content, Soluble Protein Content in the Leaves of E. angustifolia under Salt Stress
4.4. Extraction and Quantification of Proteins
4.5. FASP Digestion
4.6. LC-MS/MS Analysis
4.7. Database Search and Protein Quantification
4.8. Protein GO Functional Annotation and KEGG Pathway Annotation
4.9. Protein—Protein Interact Network (PPI)
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Appendix A
| TR Number | Protein Name | Unique Peptides | Sequence Coverage [%] | Mol. Weight [kDa] | AM/CK | AM+NaCl/AM | Fold Change | t Test p Value |
|---|---|---|---|---|---|---|---|---|
| Metabolic process | ||||||||
| 115007 | Auxin response 4 | 2 | 5.7 | 52.521 | under | over | ||
| 142773 | purple acid phosphatase 2-like [64,65] | 2 | 11.2 | 53.569 | under | over | ||
| 71893 | probable alkaline neutral invertase B [22] | 2 | 4 | 65.249 | under | over | ||
| Protein folding and degradation | ||||||||
| 101573 | AAA-ATPase At5g17760-like [66] | 2 | 8 | 58.706 | under | over | ||
| 12335 | Heat shock 70 kDa partial [67] | 1 | 55.3 | 22.844 | under | over | ||
| 143526 | small heat shock protein 17.3 kDa | 2 | 1.1 | 81.397 | over | over | 2.32 | 0.032 |
| Protein synthesis | ||||||||
| 109446 | ubiquitin-60S ribosomal L40 [23] | 1 | 41.4 | 14.643 | under | over | ||
| Redox | ||||||||
| 127397 | thioredoxin 1 [68,69] | 1 | 10.5 | 20.973 | under | over | ||
| 130845 | glutaredoxin 3 [37] | 2 | 24 | 13.046 | under | over | ||
| Cytoskeleton | ||||||||
| 114464 | actin-related 7 [70] | 1 | 4.7 | 39.154 | under | over | ||
| Transport | ||||||||
| 119356 | transmembrane 147 [71] | 2 | 10.5 | 25.299 | under | over | ||
| 88151 | vesicle transport v-SNARE 13 [72] | 2 | 15.8 | 24.96 | under | over | ||
| Signalling | ||||||||
| 136541 | serine/threonine-protein kinase PRP4 [73,74] | 1 | 5.1 | 28.709 | under | over | ||
| TR Number | Protein Name | Unique Peptides | Sequence Coverage [%] | Mol. Weight [kDa] | AM+NaCl/AM | Fold Change | t Test p Value |
|---|---|---|---|---|---|---|---|
| Metabolic process | |||||||
| 103310 | beta-glucosidase 40 [75] | 3 | 7.7 | 60.254 | over | ||
| 142616 | beta-glucosidase 42 [75] | 7 | 18.8 | 57.009 | over | 2.16 | 0.037 |
| 105849 | 1,2-dihydroxy-3-keto-5-methylthiopentene dioxygenase [76] | 6 | 32.2 | 23.919 | over | 2.42 | 0.014 |
| 121954 | anthocyanidin reductase [77] | 2 | 5.2 | 37.122 | over | ||
| 123153 | shikimate O-hydroxycinnamoyltransferase-like | 2 | 5.5 | 47.978 | over | ||
| 124556 | acetyl-CoA carboxylase [78] | 5 | 4.8 | 155.54 | over | ||
| 47457 | chorismate mutase chloroplastic-like [20,21] | 1 | 8.7 | 14.428 | over | ||
| 66089 | probable cinnamyl alcohol dehydrogenase 1 [20,21] | 9 | 31.5 | 38.574 | over | 2.22 | 0.006 |
| 99076 | caffeoyl-CoA O-methyltransferase [20,21] | 1 | 1.5 | 72.473 | over | ||
| 73516 | phosphoglucan phosphatase chloroplastic isoform X1 | 1 | 2 | 62.123 | over | ||
| 98093 | delta-1-pyrroline-5-carboxylate dehydrogenase mitochondrial-like [2] | 9 | 35.5 | 37.633 | over | 2.07 | 0.001 |
| 99156 | anthranilate phosphoribosyltransferase [20,21] | 2 | 3.3 | 83.928 | over | 2.06 | 0.016 |
| 120341 | UDP-glucose 4-epimerase [79] | 15 | 49.7 | 43.428 | over | 2.68 | 0.007 |
| Signalling | |||||||
| 87721 | phospholipase C | 6 | 14 | 59.562 | over | ||
| 114971 | calcium-dependent kinase 29 [30] | 1 | 6 | 51.414 | over | ||
| 142143 | calcium-transporting ATPase plasma membrane-type [28] | 1 | 11.4 | 15.308 | over | ||
| 155255 | glycine-rich 2-like [80] | 2 | 32.8 | 11.363 | over | 2.17 | 0.018 |
| 19556 | calcium-transporting ATPase plasma membrane-type [28] | 1 | 8.8 | 18.439 | over | ||
| 67342 | probable calcium-binding CML20 [29] | 3 | 15.8 | 22.997 | over | ||
| 61926 | calmodulin-7 [30] | 7 | 46.7 | 21.009 | over | 2.7 | 0.046 |
| 24540 | guanylate-binding family | 1 | 7.8 | 13.089 | over | ||
| Protein folding and degradation | |||||||
| 110214 | E3 ubiquitin- ligase UPL3 [26] | 2 | 2.3 | 196.04 | over | ||
| 122927 | peptidyl-prolyl cis-trans isomerase FKBP12 [23,25] | 2 | 21.4 | 12.076 | over | ||
| 74495 | FKBP-type peptidyl-prolyl cis-trans isomerase 5 isoform 1 [23,25] | 1 | 8.9 | 33.514 | over | ||
| 61488 | prefoldin subunit 2 [23,25] | 3 | 31.1 | 16.223 | over | ||
| 72620 | prefoldin subunit 1 [23,25] | 3 | 26.4 | 14.922 | over | ||
| 91034 | peptidyl-prolyl cis-trans isomerase CYP18-1 [23,25] | 1 | 8.8 | 17.474 | over | ||
| 93048 | peptidyl-prolyl cis-trans isomerase FKBP62-like [23,25] | 1 | 20.8 | 13.805 | over | ||
| 73875 | ubiquitin-like 1-activating enzyme E1 B [26] | 6 | 11.4 | 74.427 | over | 2.64 | 0.000 |
| Protein synthesis | |||||||
| 92755 | 60S acidic ribosomal P1 | 2 | 36.9 | 14.968 | over | 2.73 | 0.013 |
| 33127 | exportin-2 [81] | 1 | 14.1 | 11.097 | over | ||
| 106351 | nuclear pore complex NUP98A [82,83] | 1 | 3.8 | 50.09 | over | ||
| 109126 | elongation factor mitochondrial [24] | 2 | 5.4 | 44.427 | over | ||
| Redox | |||||||
| 101431 | monothiol glutaredoxin-mitochondrial | 3 | 15.6 | 19.243 | over | 2.27 | 0.010 |
| 12964 | peroxidase 16-like | 1 | 7.6 | 13.952 | over | ||
| 29124 | probable linoleate 9S-lipoxygenase 5 | 2 | 19.3 | 17.097 | over | 3.06 | 0.036 |
| 59386 | glutathione S-transferase L3-like [39,40] | 11 | 42.6 | 29.735 | over | 2.65 | 0.003 |
| 79685 | probable glutathione peroxidase 2 [45,46] | 6 | 33.2 | 21.733 | over | 2.35 | 0.017 |
| 97566 | 2,4-D inducible glutathione S-transferase [39,40] | 4 | 22.7 | 25.559 | over | ||
| 87674 | glutathione S-transferase T1 [39,40] | 3 | 16.8 | 27.036 | over | ||
| 88089 | probable glutathione peroxidase 8 [45,46] | 3 | 17 | 19.394 | over | 2.43 | 0.006 |
| Energy | |||||||
| 109215 | ATP synthase subunit mitochondrial | 4 | 60.6 | 14.69 | over | 3.36 | 0.015 |
| 141440 | NADH dehydrogenase [ubiquinone] 1 beta subcomplex subunit 7 | 5 | 40.6 | 15.264 | over | 2.59 | 0.010 |
| 28754 | NADH dehydrogenase [ubiquinone] 1 beta subcomplex subunit 7 | 1 | 7.9 | 11.862 | over | ||
| 78587 | NADH dehydrogenase [ubiquinone] iron-sulfur mitochondrial | 2 | 9.4 | 18.933 | over | 2.19 | 0.029 |
| 104876 | cytochrome c oxidase subunit mitochondrial-like | 2 | 30.3 | 11.185 | over | 2.27 | 0.050 |
| Transport | |||||||
| 88283 | aquaporin PIP2-1-like [84] | 1 | 4.9 | 30.753 | over | ||
| 7488 | aquaporin PIP1-4 [84] | 2 | 11.9 | 21.091 | over | ||
| 87290 | mechanosensitive ion channel 1 [85] | 2 | 2.5 | 85.926 | over | ||
| 88244 | vacuolar sorting-associated 2 homolog 1 [86] | 2 | 6.7 | 25.106 | over | ||
| Stress | |||||||
| 109624 | GDSL esterase lipase 1 [87] | 3 | 13.2 | 42.914 | over | ||
| 93161 | GDSL esterase lipase At3g27950-like [88] | 2 | 8.9 | 30.265 | over | 2.07 | 0.029 |
| 58645 | stress response NST1-like | 1 | 2.8 | 35.36 | over | ||
| Photosynthesis | |||||||
| 71168 | photosystem II D1 precursor processing PSB27-chloroplastic-like [54,55,56] | 2 | 12.3 | 23.507 | over | ||
| 8490 | hexokinase-3 isoform X1 [89] | 1 | 14.9 | 16.704 | over | 2.5 | 0.002 |
| Cytoskeleton | |||||||
| 73217 | profilin | 2 | 13.1 | 19.413 | over | 2.45 | 0.039 |
| 121711 | actin-depolymerizing factor 1 [90] | 2 | 26.6 | 16.042 | over | ||
References
- Sun, Y.F.; Song, F.Q.; Chang, W.; Fan, X.X. The effects of the AMF on the growth and physiology of the E. angustifolia seedlings under the stress of saline. Sci. Silvae Sin. 2016, 52, 18–27. [Google Scholar]
- Porcel, R.; Aroca, R.; Ruiz-Lozano, J.M. Salinity stress alleviation using arbuscular mycorrhizal fungi. Agron. Sustain. Dev. 2012, 32, 181–200. [Google Scholar] [CrossRef]
- Kapoor, R.; Evelin, H.; Mathur, P.; Giri, B. Plant Acclimation to Environmental Stress; Springer: New York, NY, USA, 2012; pp. 359–401. [Google Scholar]
- Li, X.L.; Feng, G. Arbuscular Mycorrhizal Ecological Physiology; Sino-Culture Press: Beijing, China, 2001. [Google Scholar]
- Guo, X.Z.; Bi, G.C. Mycorrhiza and Its Application Technology; China Forestry Publishing House: Beijing, China, 1989. [Google Scholar]
- Li, G.Z.; Chen, Z.C.; Li, X.C.; Sheng, J.D.; Huang, C.F.; Jin, J.X. Spatial distribution of arbuscular mycorrhizal fungi in the roots of Phragmites australis in Xinjiang. Pr. Sci. 2016, 33, 1267–1274. [Google Scholar]
- Ruiz-Lozano, J.M.; Porcel, R.; Azcon, C.; Aroca, R. Regulationby arbuscular mycorrhizae of the integrated physiological responseto salinity in plants: New challenges in physiological andmolecular studies. J. Exp. Bot. 2012, 5, 1–12. [Google Scholar]
- Kumar, A.; Sharma, S.; Mishra, S. Influence ofarbuscularmycorrhizal (AM) fungi and salinity on seedling growth, soluteaccumulation, and mycorrhizal dependency of Jatrophacurcas L. J. Plant Growth Regul. 2010, 29, 297–306. [Google Scholar] [CrossRef]
- Li, M.N.; Zhang, K.; Long, G.; Sun, Y.; Kang, J.M.; Zhang, T.J.; Cao, S.H. iTRAQ-based comparative proteomic analysis reveals tissue-specific and novel early-stage molecular mechanisms of salt stress response in Carex rigescens. Environ. Exp. Bot. 2017, 143, 99–114. [Google Scholar] [CrossRef]
- Ma, Q.L.; Kang, J.M.; Long, R.C.; Cui, Y.G.; Zhang, T.J.; Xiong, J.B.; Yang, Q.C.; Sun, Y. Proteomic analysis of salt and osmotic-drought stress in alfalfaseedlings. J. Integr. Agric. 2016, 15, 2266–2278. [Google Scholar] [CrossRef]
- Yu, J.J.; Chen, S.X.; Dai, S.J. Physiological and proteomic analysis of salinity tolerance in Puccinellia tenuiflora. J. Proteome Res. 2011, 10, 3852–3870. [Google Scholar] [CrossRef] [PubMed]
- Banaei-Asl, F.; Farajzadeh, D.; Bandehagh, A.; Komatsu, S. Comprehensive proteomic analysis of canola leaf inoculated with a plant growth-promoting bacterium, Pseudomonas fluorescens, under salt stress. Biochim. Biophys. Acta 2016, 1864, 1222–1236. [Google Scholar] [CrossRef] [PubMed]
- Yin, Y.Q.; Yang, R.Q.; Han, Y.B.; Gu, Z.X. Comparative proteomic and physiological analysesreveal the protective effect of exogenous calcium on the germinating soybean response to salt stress. J. Proteom. 2015, 113, 110–126. [Google Scholar] [CrossRef] [PubMed]
- Niu, X.P.; Xu, J.T.; Chen, T.; Tao, A.; Qi, J.M. Proteomic changes in kenaf (Hibiscus cannabinus L.) leaves under saltstress. Ind. Crop. Prod. 2016, 91, 255–263. [Google Scholar] [CrossRef]
- Lv, D.W.; Zhu, G.R.; Zhu, D.; Bian, Y.W.; Liang, X.N.; Cheng, Z.W.; Deng, X.; Yan, Y.M. Proteomic and phosphoproteomic analysis reveals the response and defense mechanism in leaves of diploid wheat T. monococcum under salts tress and recovery. J. Proteom. 2016, 143, 93–105. [Google Scholar] [CrossRef] [PubMed]
- Morandi, D. Occurrence of Phytoalexins and Phenolic com Pounds in endomyeorrhizalinteractions, and their Potential role in biological control. Plant Soil 1996, 185, 241–251. [Google Scholar] [CrossRef]
- Akiyama, K.; Matsuoka, H.; Hayashi, H. Isolation and identification of a phosphate deficieney-induced C-glycosylflavonoid that stimulates arbuscular mycorrhiza formation in melon roots. Mol. Plant-Microbe Interact. 2002, 15, 334–340. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Yan, X.F. Effects of arbuscular mycorrhizal fungi on plant secondary metabolism. Chin. J. Plant Ecol. 2006, 30, 514–521. [Google Scholar]
- Larose, G.; Chnevert, R.; Moutoglis, P.; Gagne, S.; Piche, Y.; Vietheilig, H. Flavonoid Ievels in roots of Medicagosativa are modulated by the developmental stage of the symbiosis and the root colonizing arbuseular mycorrhizal fungus. J. Plant Physiol. 2002, 159, 1329–1339. [Google Scholar] [CrossRef]
- Qian, X.; Yu, K.J.; Yuan, G.S.; Yang, H.Y.; Pan, Q.H.; Zhu, B.Q. Research progress in plant anthranilate synthase. J. Trop. Biol. 2015, 6, 504–511. [Google Scholar]
- Pérez, E.; Rubio, M.B.; Cardoza, R.E.; Gutiérrez, S.; Bettiol, W.; Monte, E.; Hermosa, R. The importance of chorismate mutase in the biocontrol potential of Trichoderma parareesei. Front. Microbiol. 2015, 6, 1181. [Google Scholar] [CrossRef]
- Niu, J.Q.; Wang, A.Q.; Huang, J.L.; Yang, L.T.; Li, Y.R. Cloning and expression analysis of sugarcane alkaline/neutral invertase gene SoNIN1. Acta Agron. Sin. 2014, 40, 253–263. [Google Scholar] [CrossRef]
- Li, H.Y.; Yu, P.; Zhang, Y.X.; Wu, C.; Ma, C.Q.; Yu, B.; Zhu, N.; Koh, J.; Chen, S.X. Salt stress response of membrane proteome of sugarbeet monosomic addition line M14. J. Proteom. 2015, 127, 18–33. [Google Scholar] [CrossRef]
- Suzuki, H.; Ueda, T.; Taguchi, H.; Takeuchi, N. Chaperoneproperties of mammalian mitochondrial translation elongationfactor Tu. J. Biol. Chem. 2007, 282, 4076–4084. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.J.; Chen, S.X.; Dai, S.J. Comparative proteomic analysis of Puccinellia tenuiflora leaves under Na2CO3 stress. Int. J. Mol. Sci. 2013, 14, 1740–1762. [Google Scholar] [CrossRef] [PubMed]
- Lyzenga, W.J.; Stone, S.L. Abiotic stress tolerance mediated by protein ubiquitination. J. Exp. Botary 2012, 63, 599–616. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.S.; Lan, H.Y. Signal Transduction Pathways in Response to Salt Stress in Plants. Plant Physiol. J. 2011, 47, 119–128. [Google Scholar]
- Kim, J.S.; Park, S.J.; Kwak, K.J.; Kim, Y.O.; Kim, J.K.; Song, J.; Jang, B.; Jung, S.H.; Kang, H. Cold shock domain proteins and glycine-rich RNA-binding proteins from Arabidopsis thaliana can promote the cold adaptation process in Escherichia coli. Nucleic Acids Res. 2007, 35, 505–516. [Google Scholar] [CrossRef]
- Harmon, A.C. Calcium-regulated protein kinases of plants. Gravit. Space Biol. Bull. 2003, 16, 83–90. [Google Scholar] [PubMed]
- Asano, T.; Hayashi, N.; Kikuchi, S.; Ohsugi, R. CDPK-mediated abiotic stress signaling. Plant Signal. Behav. 2012, 7, 817–821. [Google Scholar] [CrossRef] [PubMed]
- Choudhury, S.; Panda, P.; Sahoo, L.; Panda, S.K. Reactive oxygen species signalingin plants under abiotic stress. Plant Signal. Behav. 2013, 8, e23681. [Google Scholar] [CrossRef]
- Wang, L.X.; Pan, D.Z.; Li, J.; Tan, F.L.; Benning, S.H.; Liang, W.Y.; Chen, W. Proteomic analysis of changes in the Kandelia candel chloroplast proteins reveals pathways associated with salt tolerance. Plant Sci. 2015, 231, 159–172. [Google Scholar] [CrossRef] [PubMed]
- Rouhier, N.; Gelhaye, E.; Jacquot, J.P. Redox control by dithiol–disulfide exchange in plants: II. The cytosolic and mitochondrial systems. Ann. N. Y. Acad. Sci. 2002, 973, 520–528. [Google Scholar] [CrossRef] [PubMed]
- Scheibe, R.; Dietz, K.J. Reduction-oxidation network for flexible adjustment ofcellular metabolism in photoautotrophic cells. Plant Cell Environ. 2012, 35, 202–216. [Google Scholar] [CrossRef] [PubMed]
- Jacquot, J.P.; Rouhier, N.; Gelhaye, E. Redox control by dithiol–disulfide exchange in plants: I. The chloroplastic systems. Ann. N. Y. Acad. Sci. 2002, 973, 508–519. [Google Scholar] [CrossRef] [PubMed]
- Michelet, L.; Zaffagnini, M.; Morisse, S.; Sparla, F.; Pérez-Pérez, M.E.; Francia, F.; Danon, A.; Marchand, C.H.; Fermani, S.; Trost, P.; et al. Redox regulation of the Calvin–Benson cycle: Something old, something new. Front. Plant Sci. 2013, 4, 470. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.; Huang, C.; Xie, Y.; Song, F.; Zhou, X. A tomato glutaredoxin gene SLGRX1 regulates plant responses to oxidative, drought and salt stresses. Planta 2010, 232, 1499–1509. [Google Scholar] [CrossRef] [PubMed]
- Song, C.Q.; Miao, H.F.; Zhu, B. The role of plant glutathione-S-transferase in phytoremediation. Anhui Agric. Bull. 2010, 16, 1007–7731. [Google Scholar]
- Mittler, R. Oxidative stress, antioxidants and stress tolerance. Trends Plant Sci. 2002, 7, 405–410. [Google Scholar] [CrossRef]
- Margis, R.; Dunand, C.; Teixeira, F.K.; Pinheiro, M.M. Glutathione peroxidase family-an evolutionary overview. FEBS J. 2008, 275, 3959–3970. [Google Scholar] [CrossRef] [PubMed]
- Sun, G.R.; Wang, J.B.; Cao, W.Z.; Du, K.; Zhang, B. GST activity and its related indexes in the Chloroplasts of Puccinellia tenuiflora seedlings under Na2CO3 stress. Northwest Bot. Gaz. 2005, 25, 2495–2501. [Google Scholar]
- Avsian, K.O.; Gueta, D.Y.; Ben, H.G.; Eshdat, Y. Regulation of stress-induced phospholipid hydroperoxide glutathione peroxidase expression in citrus. Planta 1999, 4, 469–477. [Google Scholar] [CrossRef]
- Beeor-Tzahar, T.; Ben-Hayyim, G.; Holland, D.; Faltun, Z.; Eshdat, Y. A stress-associated citrus protein is a distinct plant phospholipid hydroperoxide glutathione peroxidase. FEBS Lett. 1995, 366, 151–155. [Google Scholar] [CrossRef]
- Kim, Y.J.; Jang, M.G.; Noh, H.Y.; Lee, H.J.; Sukweenadhi, J.; Kim, J.H.; Kim, S.Y.; Kwon, W.S.; Yang, D.C. Molecular characterization of two glutathione peroxidase genes of Panax ginseng and their expression analysis against environmental stresses. Gene 2014, 1, 33–41. [Google Scholar] [CrossRef] [PubMed]
- Roxas, V.P.; Smith, R.K., Jr.; Allen, E.R.; Allen, R.D. Overexpression of glutathione S-transferase/glutathione peroxidase enhances growth of transgenic tobacco seedlings during stress. Nat. Biotechnol. 1997, 15, 988–991. [Google Scholar] [CrossRef] [PubMed]
- Roxas, V.P.; Lodhi, S.A.; Garrett, D.K.; Mahan, J.R.; Allen, R.D. Stress Tolerance in Transgenic Tobacco Seedlings that Overexpress Glutathione S-Transferase/Glutathione Peroxidase. Plant Cell Physiol. 2000, 41, 1229–1234. [Google Scholar] [CrossRef] [PubMed]
- Chang, W.; Sui, X.; Fan, X.X.; Jia, T.T. Arbuscular Mycorrhizal Symbiosis Modulates Antioxidant Response and Ion Distribution in Salt-Stressed Elaeagnus angustifolia Seedlings. Front. Microbiol. 2018, 9, 652. [Google Scholar] [CrossRef] [PubMed]
- Talaat, N.B.; Shawky, B.T. Protective effects of arbuscular mycorrhizal fungi on wheat (Triticum aestivum L.) plants exposed to salinity. Environ. Exp. Bot. 2014, 98, 20–31. [Google Scholar] [CrossRef]
- Yue, Y.N. Effects of Arbuscular Mycorrhizal Fungi on Plant Salt Tolerance in Songnen Saline Grassland; Northeast Forestry University: Haerbin, China, 2015. [Google Scholar]
- Sheng, M.; Tang, M.; Chen, H.; Yang, B.W.; Zhang, F.F.; Huang, Y.H. Influence of arbuscular mycorrhizae on photosynthesis and water status of maize plants under salt stress. Mycorrhiza 2008, 18, 287–296. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.X.; Wang, Y.N.; Sun, S.N.; Mu, C.S.; Yan, X.F. Effects of arbuscular mycorrhizal fungi on the growth, photosynthesis and photosynthetic pigments of Leymus chinensis seedlings under salt-alkali stress and nitrogen deposition. Sci. Total Environ. 2017, 576, 234–241. [Google Scholar] [CrossRef]
- Liu, H.G. AMF to Improve the Mechanism of Salt Resistance of Lycium barbarum L.; Northwest A & F University: Yangling, China, 2016. [Google Scholar]
- Tang, J. Effects of Mycorrhizal Fungi (AMF) on Salinity Tolerance in the Cultivation of Healthy Vegetables; Sichuan Agricultural University: Yaan, China, 2016. [Google Scholar]
- Järvi, S.; Suorsa, M.; Aro, E.M. Photosystem II repair in plant chloroplasts-Regulation, assisting proteins and shared components with photosystem II biogenesis. Biochim. Biophys. Acta 2015, 1847, 900–909. [Google Scholar] [CrossRef] [PubMed]
- Greenberg, B.M.; Gaba, V.; Canaani, O.; Malkin, S.; Mattoo, A.K.; Edelman, M. Separate photosensitizers mediate degradation of the 32 kDa photosystem II reaction center protein in the visible and UV spectral regions. Proc. Natl. Acad. Sci. USA 1989, 86, 6617–6620. [Google Scholar] [CrossRef]
- Zhu, S.Q.; Xia, S.L.; Chen, Q. Advances in the regulation mechanism of photosystem IID1 protein expression. Guizhou Agric. Sci. 2012, 40, 37–42. [Google Scholar]
- Guo, X. Role of Glutathione GRX in Response to Temperature Stress in Tomato. Master’s Thesis, Zhejiang University, Hangzhou, China, 2016. [Google Scholar]
- Jia, T.T.; Chang, W.; Fan, X.X. Effects of Arbuscular mycorrhizal fungi on photosynthetic and chlorophyll fluorescence characteristics in Elaeagnus angustifolia seedlings under salt stress. Acta Ecol. Sin. 2018, 38, 1337–1347. [Google Scholar]
- Phillips, J.M.; Hayman, D.S. Improved procedures for clearing roots and staining parasitic and vesicular-arbuscular mycorrhizal fungi for rapid assessment of infection. Trans. Br. Mycol. Soc. 1970, 55, 158–161. [Google Scholar] [CrossRef]
- Wang, X.K.; Huang, S.L. Plant Physiological and Biochemical Experimental Principles and Technology, 3rd ed.; Higher Education Press: Beijing, China, 2015. [Google Scholar]
- Zhang, Z.L.; Qu, W.J. Plant Physiology Experimental Guide, 3rd ed.; Higher Education Press: Beijing, China, 2003. [Google Scholar]
- Wisniewski, J.R.; Zougman, A.; Nagaraj, N.; Mann, M. Universal sample preparation method for proteome analysis. Nat. Methods 2009, 6, 359–362. [Google Scholar] [CrossRef] [PubMed]
- Cox, J.; Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 2008, 6, 1367–1372. [Google Scholar] [CrossRef] [PubMed]
- Del Pozo, J.C.; Allona, I.; Rubio, V.; Leyva, A.; de la Peña, A.; Aragoncillo, C.; Paz-Ares, J. A type 5 acid phosphatase gene from Arabidopsis thaliana is induced by phosphate starvation and by some other types of phosphate mobilising/oxidativestress conditions. Plant J. 1999, 19, 579–589. [Google Scholar] [CrossRef] [PubMed]
- Li, W.Y.; Shao, G.; Lam, H. Ectopic expression of GmPAP3 alleviates oxidative damage caused by salinity and osmotic stresses. New Phytol. 2008, 178, 80–91. [Google Scholar] [CrossRef] [PubMed]
- Monroe, N.; Hill, C.P. Meiotic Clade AAA ATPases: Protein Polymer Disassembly Machines. J. Mol. Biol. 2016, 428 Pt B, 1897–1911. [Google Scholar] [CrossRef]
- Ren, Y.; Pan, H.; Yang, Y.; Pan, B.; Bu, W. Molecular cloning, characterization and functional analysis of a heat shock protein 70 gene in Cyclina sinensis. Fish Shellfish Immunol. 2016, 58, 663–668. [Google Scholar] [CrossRef]
- Wong, J.H.; Cai, N.; Balmer, Y.; Tanaka, C.K.; Vensel, W.H.; Hurkman, W.J.; Buchanan, B.B. Thioredoxin targets of developing wheat seeds ide ntified by complementary proteomic approaches. Phytochemistry 2004, 65, 1629–1640. [Google Scholar] [CrossRef]
- Traverso, J.A.; Vignols, F.; Cazalis, R.; Pulido, A.; Sahrawy, M.; Cejudo, F.J.; Meyer, Y.; Chueca, A. PsTRXh1 and PsTRXh2 Are Both Pea h-Type Thioredoxins with Antagonistic Behavior in Redox Imbalances. Plant Physiol. 2007, 143, 300. [Google Scholar] [CrossRef]
- Wang, J.; Zhang, S.B.; Ma, D.; Ma, G.Y. Advances in Plant Actin Research. J. Anhui Agric. Sci. 2007, 35, 2860–2863. [Google Scholar]
- Song, J.H.; Zhang, L.X. Progress on the transmembrane protein in plant. J. Biol. 2009, 26, 62–64. [Google Scholar]
- Weber, T.; Parlati, F.; McNew, J.A.; Johnston, R.J.; Westermann, B.; Söllner, T.H.; Rothman, J.E. SNAREpins are functionally resistant to dis-ruption by NSF and αSNAP. J. Cell Biol. 2000, 149, 1063–1072. [Google Scholar] [CrossRef]
- Sun, X.L.; Yu, Q.Y.; Tang, L.L.; Ji, W.; Bai, X.; Cai, H.; Liu, X.F.; Ding, X.D.; Zhu, Y.M. GsSRK, a G-type lectin S-receptor-like serine/threonine protein kinase, is apositive regulator of plant tolerance to salt stress. J. Plant Physiol. 2013, 170, 505–515. [Google Scholar] [CrossRef] [PubMed]
- Ge, R.C.; Chen, G.P.; Zhao, B.C.; Shen, Y.Z.; Huang, Z.J. Cloning and functional characterization of a wheat serine/threoninekinase gene (TaSTK) related to salt-resistance. Plant Sci. 2007, 173, 55–60. [Google Scholar] [CrossRef]
- Lee, K.H.; Piao, H.L.; Kim, H.Y.; Choi, S.M.; Jiang, F.; Hartung, W.; Hwang, I.; Kwak, J.M.; Lee, I.J.; Hwang, I. Activation of glucosidase via stress-induced polymerization rapidly increases active pools of abscisic acid. Cell 2006, 126, 1109–1120. [Google Scholar] [CrossRef]
- Ge, C.; Wan, D.; Wang, Z.; Ding, Y.; Wang, Y.; Shang, Q.; Ma, F.; Luo, S. A proteomic analysis of rice seedlings responding to 1,2,4-trichlorobenzene stress. J. Environ. Sci. 2008, 20, 309–319. [Google Scholar] [CrossRef]
- Luo, P.; Shen, Y.X.; Jin, S.J. Overexpression of Rosa rugosa anthocyanidin reductase enhances tobacco tolerance to abiotic stress through increased ROS scavenging and modulation of ABA signaling. Plant Sci. 2016, 245, 35–39. [Google Scholar] [CrossRef]
- Lee, J.Q.; Zheng, S.X.; Yu, Z.N. Acetyl-CoA carboxylase: Advances in research on key enzymes of fatty acid metabolism and gene cloning. J. Appl. Environ. Biol. 2011, 17, 753–758. [Google Scholar]
- Zhang, Q.; Hrmova, M.; Shirley, N.J.; Lahnstein, J.; Fincher, G.B. Gene expression patterns and catalytic properties of UDP-D-glucose4-epimerases from barley Hordeum vulgare L. Biochem. J. 2006, 394, 115–124. [Google Scholar] [CrossRef]
- Singh, S.; Virdi, A.S.; Jaswal, R. A temperature-responsive gene in sorghum encodes a glycine-rich protein that interacts with calmodulin. Biochimie 2017, 137, 115–123. [Google Scholar] [CrossRef]
- Gupta, A.; Kailasam, S.; Bansall, M. Insights into the Structural Dynamics of Nucleocytoplasmic Transport of tRNA by Exportin-t. Biophys. J. 2016, 110, 1264–1279. [Google Scholar] [CrossRef]
- Kohler, A.; Hurt, E. Exporting RNA from the nucleus to the cytoplasm. Nat. Rev. Mol. Cell Biol. 2007, 8, 761–773. [Google Scholar] [CrossRef]
- Mattaj, I.W.; Englmeier, L. Nucleocytoplasmic transport: The soluble phase. Annu. Rev. Biochem. 1998, 67, 265–306. [Google Scholar] [CrossRef]
- Aroca, R.; Porcel, R.; Ruiz-Lozano, J.M. How does arbuscular mycorrhizal symbiosis regulate root hydraulic properties and plasma membrane aquaporins in Phaseolus vulgaris under drought, cold or salinity stresses? New Phytol. 2007, 73, 808–816. [Google Scholar] [CrossRef]
- Geelen, D.; Leyman, B.; Batoko, H.; Di Sansebastiano, G.P.; Moore, I.; Blatt, M.R. The abscisic acid-related SNARE homolog NtSyr1contributes to secretion and growth: Evidence from competition with its cytosolic domain. Plant Cell 2002, 14, 387–406. [Google Scholar] [CrossRef]
- Iqbal, M.S.; Siddiqui, A.A.; Alam, A.; Goyal, M.; Banerjee, C.; Sarkar, S.; Mazumder, S.; De, R.; Nag, S.; Saha, S.J.; et al. Expression, purification and characterization of Plasmodium falciparum vacuolar protein sorting 29. Protein Expr. Purif. 2016, 120, 7–15. [Google Scholar] [CrossRef]
- Hong, J.K.; Choi, H.W.; Hwang, I.S.; Kim, D.S.; Kim, N.H.; Choi, D.S.; Kim, Y.J.; Hwang, B.K. Function of a novel GDSL-type pepper lipase gene, CaGLIP1, in disease sus-ceptibility and abiotic stress tolerance. Planta 2008, 227, 539–558. [Google Scholar] [CrossRef]
- Naranjo, M.; Forment, J.; Roldan, M.; Serrano, R.; Vicente, O. Overexpression of Arabidopsis thaliana LTL1, a salt-induced gene encod-ing a GDSL-motif lipase, increases salt tolerance in yeast and transgenic plants. Plant Cell Environ. 2006, 29, 1890–1900. [Google Scholar] [CrossRef]
- Li, N.N.; Qian, W.J.; Wang, L.; Cao, H.L.; Hao, X.Y.; Yang, Y.J.; Wang, X.C. Isolation and expression features of hexose kinase genes under various abiotic stresses in the tea plant (Camellia sinensis). J. Plant Physiol. 2017, 209, 95–104. [Google Scholar] [CrossRef]
- Van Gisbergen, P.A.; Bezanilla, M. Plant formins: Membrane anchors for actin polymerization. Trends Cell Biol. 2013, 23, 227–233. [Google Scholar] [CrossRef]

| Level of Salinity/(mmol/L) | Different Treatment | Height/(cm) | Diameter/(mm) | Length/(cm) | Area/(cm2) |
|---|---|---|---|---|---|
| 0 | CK | 45.50 ± 0.24c | 5.65 ± 0.17b | 985.73 ± 27.80b | 146.04 ± 5.98c |
| RI | 49.07 ± 0.54a | 6.54 ± 0.20a | 1256.7 ± 22.52a | 213.07 ± 13.04a | |
| Significance | ** | ** | ** | ** | |
| 300 | CK | 39.57 ± 0.26f | 3.99 ± 0.14e | 763.64 ± 23.34e | 93.68 ± 6.27e |
| RI | 43.17 ± 0.21de | 4.82 ± 0.11d | 896.56 ± 42.36bcd | 126.96 ± 8.03cd | |
| Significance | ** | ** | * | ** |
| Treatments | Number of Differential Proteins |
|---|---|
| NaCl vs CK variation analysis | 402 a + 335 b |
| AM vs CK variation analysis | 35 a + 152 b |
| AM + NaCl vs AM variation analysis | 166 a + 226 b |
| AM + NaCl vs NaCl variation analysis | 62 a + 189 b |
| Item | Value |
|---|---|
| Main search ppm | 6 |
| Missed cleavage | 2 |
| MS/MS tolerance ppm | 20 |
| De-Isotopic | True |
| Enzyme | Trypsin |
| Database | P16440_Unigene.fasta.transdecoder_73797_20161212.fasta |
| Fixed modification | Carbamidomethyl (C) |
| Variable modification | Oxidation(M), Acetyl (Protein N-term) |
| Decoy database pattern | reverse |
| LFQ | True |
| LFQ min. ratio count | 1 |
| Match between runs | 2min |
| Peptide FDR | 0.01 |
| Protein FDR | 0.01 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jia, T.; Wang, J.; Chang, W.; Fan, X.; Sui, X.; Song, F. Proteomics Analysis of E. angustifolia Seedlings Inoculated with Arbuscular Mycorrhizal Fungi under Salt Stress. Int. J. Mol. Sci. 2019, 20, 788. https://doi.org/10.3390/ijms20030788
Jia T, Wang J, Chang W, Fan X, Sui X, Song F. Proteomics Analysis of E. angustifolia Seedlings Inoculated with Arbuscular Mycorrhizal Fungi under Salt Stress. International Journal of Molecular Sciences. 2019; 20(3):788. https://doi.org/10.3390/ijms20030788
Chicago/Turabian StyleJia, Tingting, Jian Wang, Wei Chang, Xiaoxu Fan, Xin Sui, and Fuqiang Song. 2019. "Proteomics Analysis of E. angustifolia Seedlings Inoculated with Arbuscular Mycorrhizal Fungi under Salt Stress" International Journal of Molecular Sciences 20, no. 3: 788. https://doi.org/10.3390/ijms20030788
APA StyleJia, T., Wang, J., Chang, W., Fan, X., Sui, X., & Song, F. (2019). Proteomics Analysis of E. angustifolia Seedlings Inoculated with Arbuscular Mycorrhizal Fungi under Salt Stress. International Journal of Molecular Sciences, 20(3), 788. https://doi.org/10.3390/ijms20030788






