Citrus Taste Modification Potentials by Genetic Engineering
Abstract
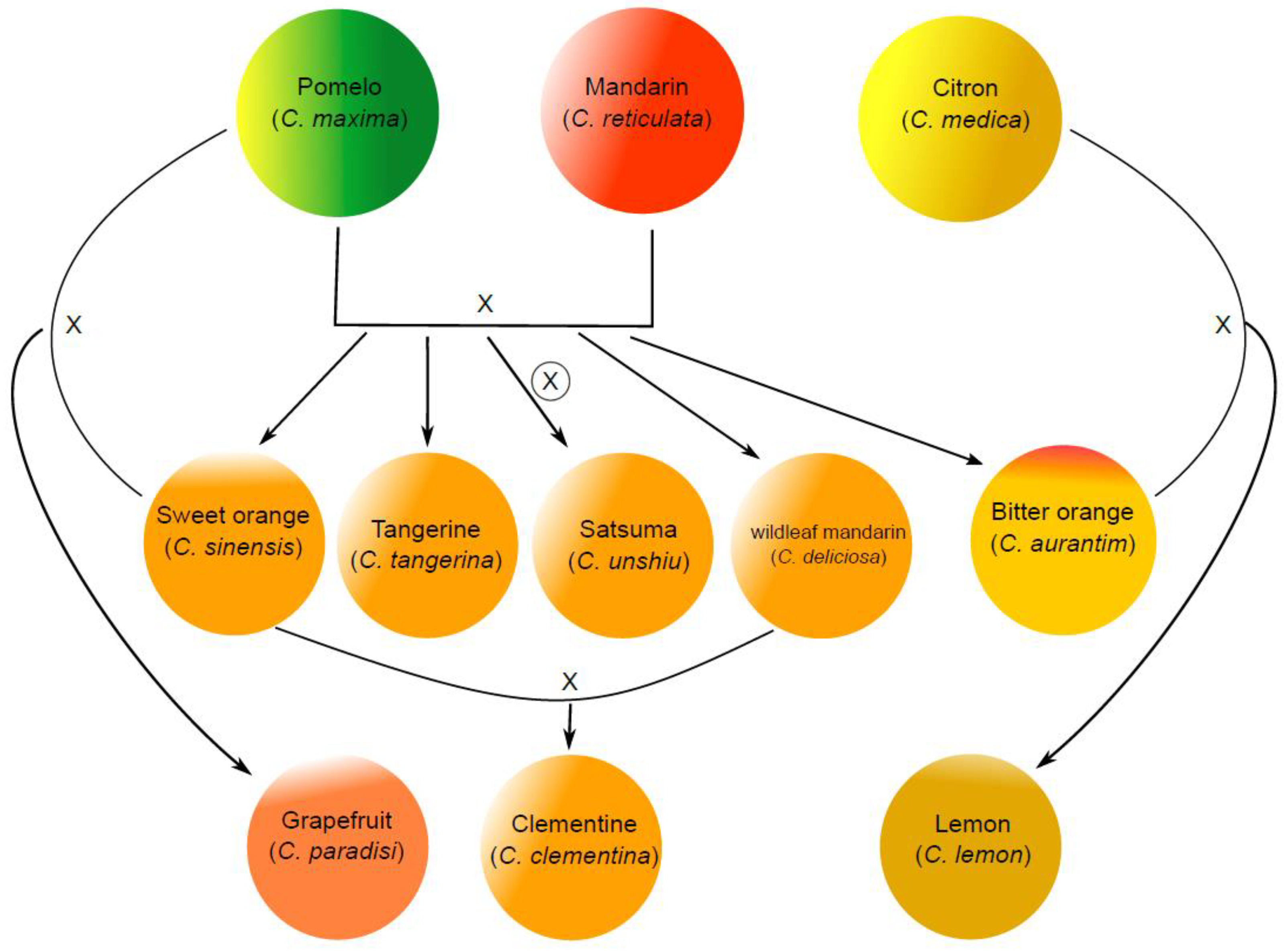
1. Introduction
2. Citrus Taste
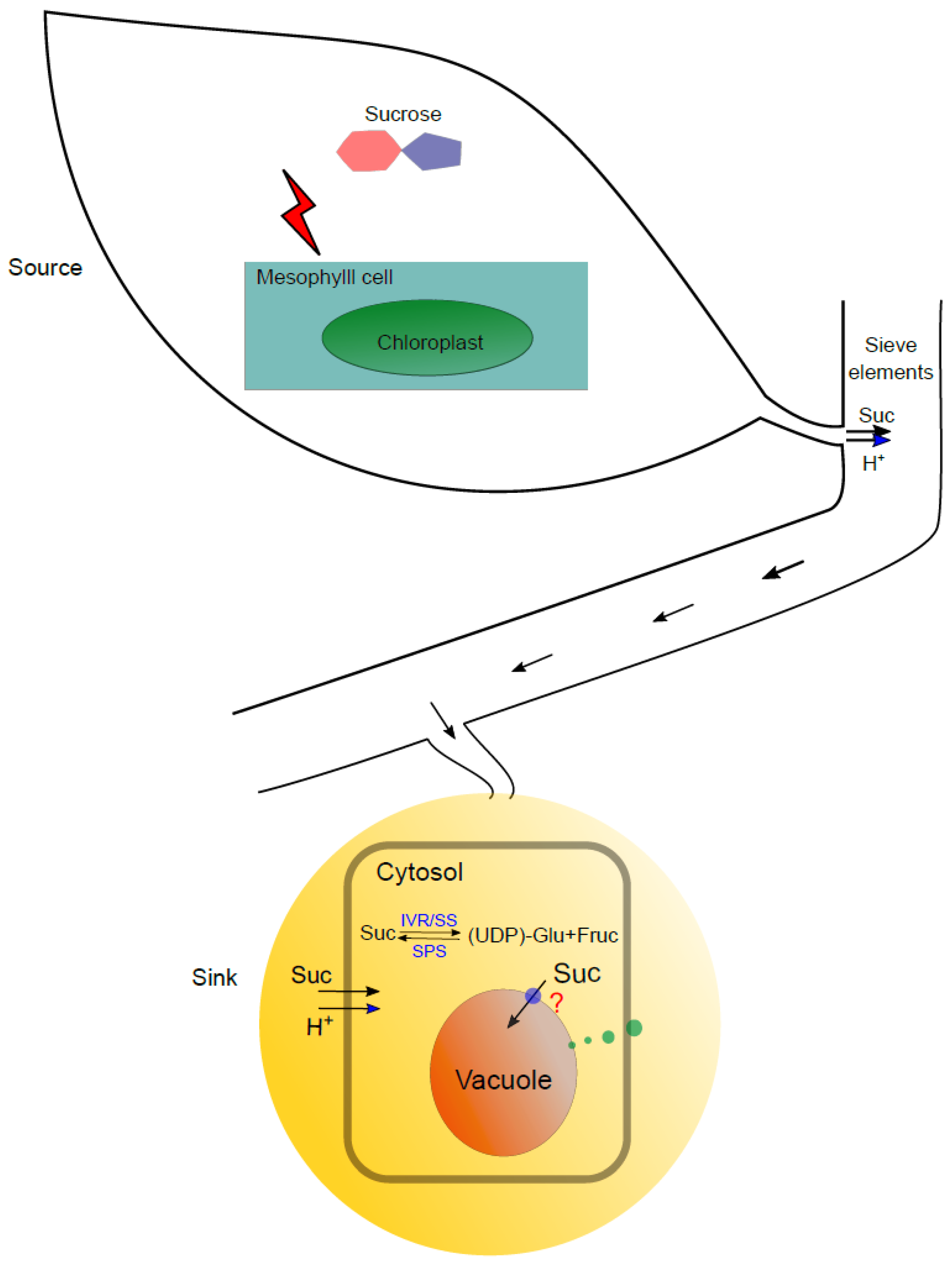
2.1. Sweet
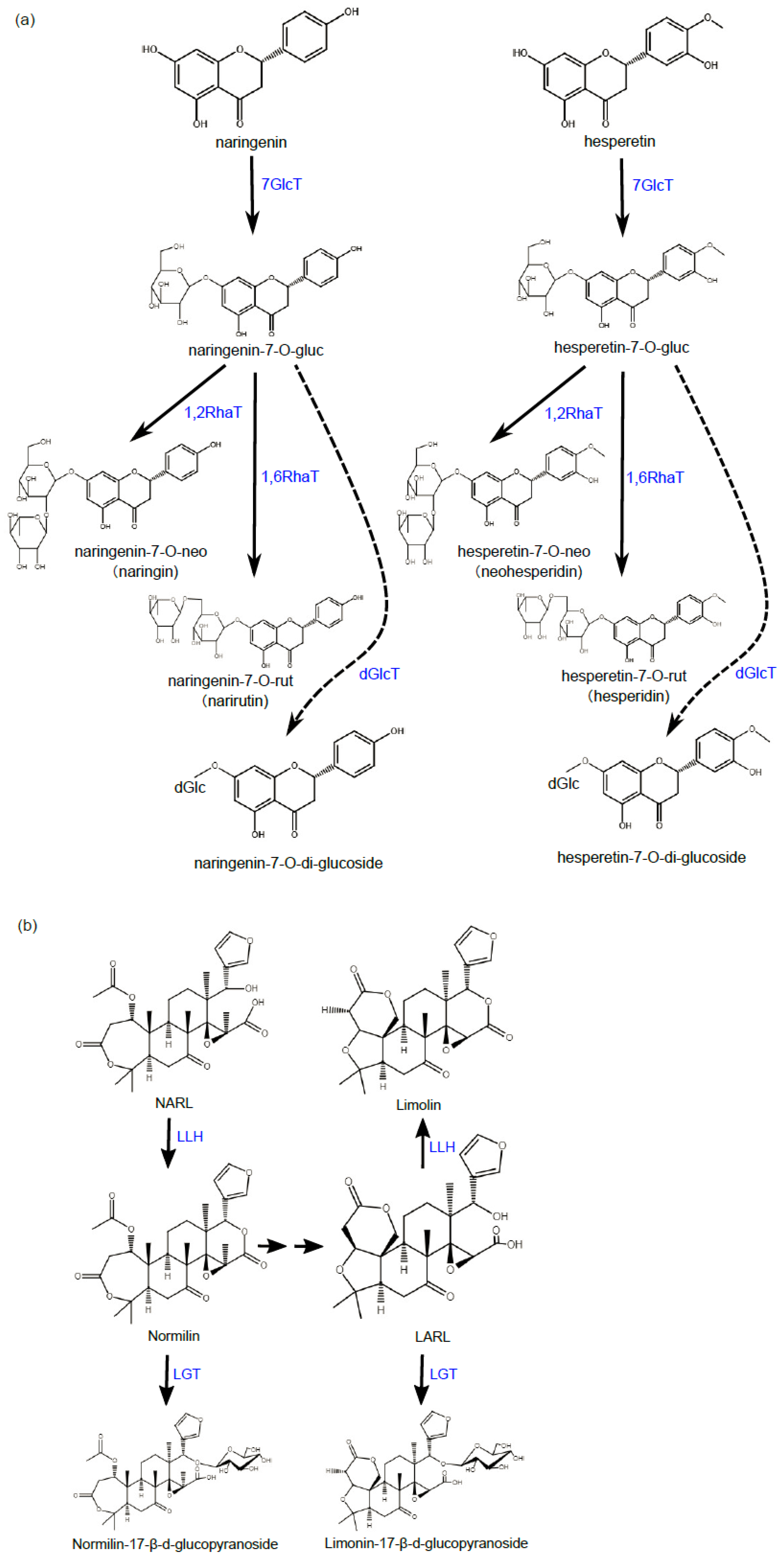
2.2. Bitter
2.2.1. Cm1,2RhaT
2.2.2. Cs1,6RhaT
2.2.3. CitdGlcT
2.2.4. CitLGTs
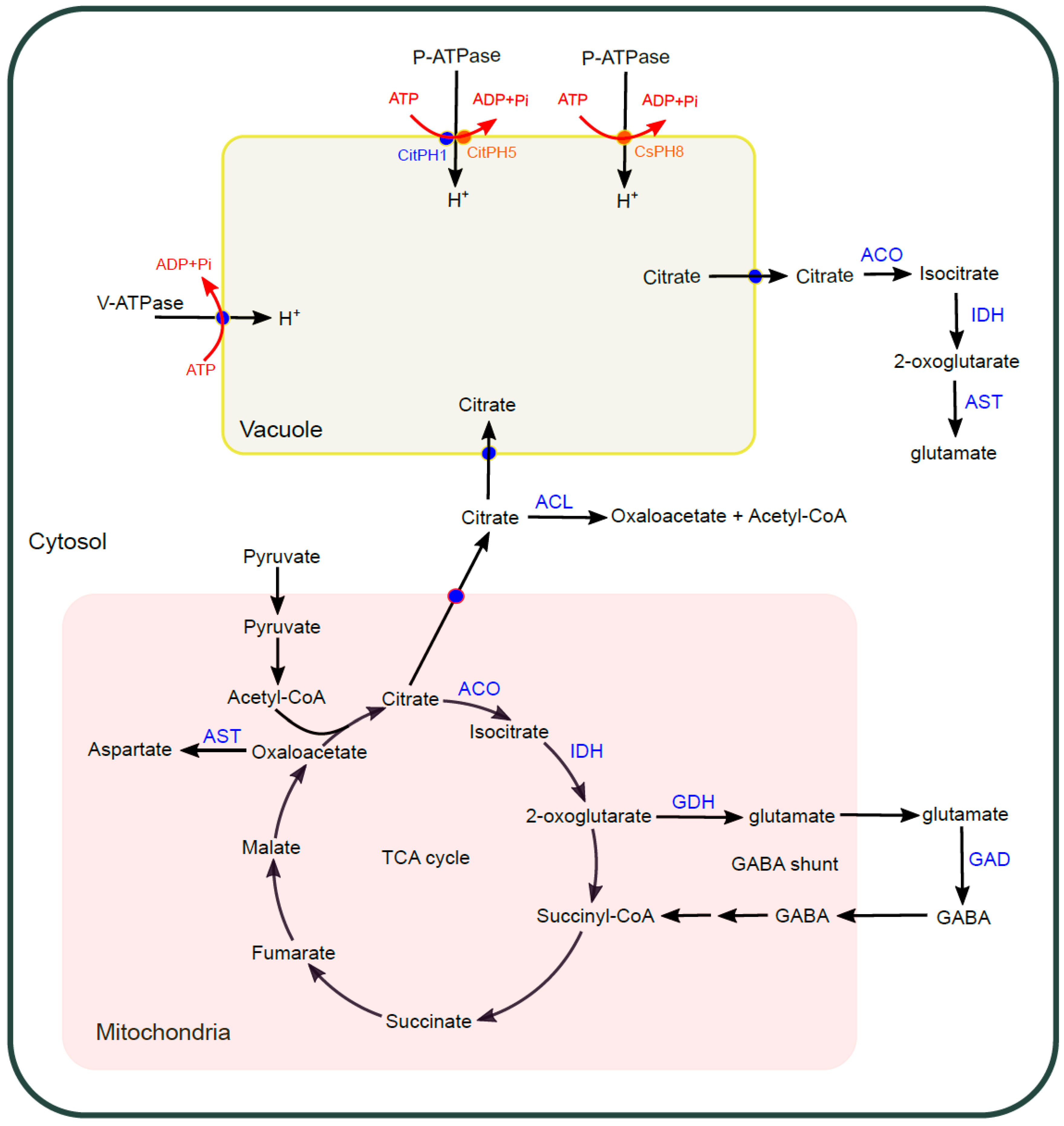
2.3. Sour
2.3.1. V-ATPase
2.3.2. P-ATPase
3. Citrus Genetic Modification Approach
4. Conclusions and Perspectives
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Frydman, A.; Liberman, R.; Huhman, D.V.; Carmeli-Weissberg, M.; Sapir-Mir, M.; Ophir, R.; L, W.S.; Eyal, Y. The molecular and enzymatic basis of bitter/non-bitter flavor of citrus fruit: Evolution of branch-forming rhamnosyltransferases under domestication. Plant J. 2013, 73, 166–178. [Google Scholar] [CrossRef] [PubMed]
- Wu, G.A.; Terol, J.; Ibanez, V.; Lopez-Garcia, A.; Perez-Roman, E.; Borreda, C.; Domingo, C.; Tadeo, F.R.; Carbonell-Caballero, J.; Alonso, R.; et al. Genomics of the origin and evolution of Citrus. Nature 2018, 554, 311–316. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Heying, E.; Tanumihardjo, S.A. History, global distribution, and nutritional importance of citrus fruits. Compr. Rev. Food Sci. Food Saf. 2012, 11, 530–545. [Google Scholar] [CrossRef]
- Shimizu, T.; Kitajima, A.; Nonaka, K.; Yoshioka, T.; Ohta, S.; Goto, S.; Toyoda, A.; Fujiyama, A.; Mochizuki, T.; Nagasaki, H.; et al. Hybrid origins of citrus varieties inferred from dna marker analysis of nuclear and organelle genomes. PLoS ONE 2016, 11, e0166969. [Google Scholar] [CrossRef]
- Bharate, S.S.; Bharate, S.B. Non-enzymatic browning in citrus juice: Chemical markers, their detection and ways to improve product quality. J. Food Sci. Technol. 2014, 51, 2271–2288. [Google Scholar] [CrossRef]
- Gyawali, R.; Kim, K.S. Anticancer phytochemicals of citrus fruits—A review. J. Anim. Res. 2014, 4, 85–95. [Google Scholar] [CrossRef]
- Klee, H.J. Improving the flavor of fresh fruits: Genomics, biochemistry, and biotechnology. New Phytol. 2010, 187, 44–56. [Google Scholar] [CrossRef]
- Kore, V.T.; Chakraborty, I. Efficacy of various techniques on biochemical characteristics and bitterness of pummelo juice. J. Food Sci. Technol. 2015, 52, 6073–6077. [Google Scholar] [CrossRef]
- Talon, M.; Gmitter, F.G., Jr. Citrus genomics. Int. J. Plant Genom. 2008, 2008, 17. [Google Scholar] [CrossRef]
- Echeverria, E.; Ismail, M. Sugars unrelated to brix changes in stored citrus fruits. HortScience 1990, 25, 710. [Google Scholar] [CrossRef]
- Yamaki, S. Metabolism and accumulation of sugars translocated to fruit and their regulation. J. Japan. Soc. Hort. Sci. 2010, 79, 1–15. [Google Scholar] [CrossRef]
- Stein, O.; Granot, D. An overview of sucrose synthases in plants. Front. Plant Sci. 2019, 10, 95. [Google Scholar] [CrossRef] [PubMed]
- Lin, Q.; Wang, C.; Dong, W.; Jiang, Q.; Wang, D.; Li, S.; Chen, M.; Liu, C.; Sun, C.; Chen, K. Transcriptome and metabolome analyses of sugar and organic acid metabolism in Ponkan (Citrus reticulata) fruit during fruit maturation. Gene 2015, 554, 64–74. [Google Scholar] [CrossRef]
- Lowell, C.A.; Tomlinson, P.T.; Koch, K.E. Sucrose-metabolizing enzymes in transport tissues and adjacent sink structures in developing citrus fruit. Plant Physiol. 1989, 90, 1394–1402. [Google Scholar] [CrossRef] [PubMed]
- Sauer, N. Molecular physiology of higher plant sucrose transporters. FEBS Lett. 2007, 581, 2309–2317. [Google Scholar] [CrossRef] [PubMed]
- Etxeberria, E.; Gonzalez, P. Sucrose transport into citrus juice cells: Evidence for an endocytic transport system. J. Am. Soc. Hort. Sci. 2005, 130, 269–274. [Google Scholar] [CrossRef]
- Komatsu, A.; Moriguchi, T.; Koyama, K.; Omura, M.; Akihama, T. Analysis of sucrose synthase genes in citrus suggests different roles and phylogenetic relationships. J. Exp. Bot. 2002, 53, 61–71. [Google Scholar] [CrossRef]
- Islam, M.Z.; Jin, L.-F.; Shi, C.-Y.; Liu, Y.-Z.; Peng, S.-A. Citrus sucrose transporter genes: Genome-wide identification and transcript analysis in ripening and ABA-injected fruits. Tree Genet. Genomes 2015, 11, 97. [Google Scholar] [CrossRef]
- Islam, M.Z.; Hu, X.M.; Jin, L.F.; Liu, Y.Z.; Peng, S.A. Genome-wide identification and expression profile analysis of citrus sucrose synthase genes: Investigation of possible roles in the regulation of sugar accumulation. PLoS ONE 2014, 9, e113623. [Google Scholar] [CrossRef]
- Omura, M.; Shimada, T. Citrus breeding, genetics and genomics in Japan. Breed. Sci. 2016, 66, 3–17. [Google Scholar] [CrossRef]
- Katz, E.; Boo, K.H.; Kim, H.Y.; Eigenheer, R.A.; Phinney, B.S.; Shulaev, V.; Negre-Zakharov, F.; Sadka, A.; Blumwald, E. Label-free shotgun proteomics and metabolite analysis reveal a significant metabolic shift during citrus fruit development. J. Exp. Bot. 2011, 62, 5367–5384. [Google Scholar] [CrossRef] [PubMed]
- Kore, V.T.; Tawade, S.S.; Devi, L.H.; Chakraborty, I. Use of pre and post harvest low cost techniques to control/minimize citrus juice bitterness. Am. Res. J. Food Nutr. 2018, 1, 11–21. [Google Scholar]
- Puri, M.; MarwahaJ, S.S.; KothariJ, R.M.; Kennedy, J.F. Biochemical basis of bitterness in citrus fruit juices and biotech approaches for debittering. Crit. Rev. Biotechnol. 1996, 16, 145–155. [Google Scholar] [CrossRef]
- Raithore, S.; Dea, S.; McCollum, G.; Manthey, J.A.; Bai, J.; Leclair, C.; Hijaz, F.; Narciso, J.A.; Baldwin, E.A.; Plotto, A. Development of delayed bitterness and effect of harvest date in stored juice from two complex citrus hybrids. J. Sci. Food Agric. 2016, 96, 422–429. [Google Scholar] [CrossRef] [PubMed]
- Bi, J.; Li, H.; Wang, H. Delayed bitterness of citrus wine is removed through the selection of fining agents and fining optimization. Front. Chem. 2019, 7. [Google Scholar] [CrossRef]
- Zaare-Nahandi, F.; Hosseinkhani, S.; Zamani, Z.; Asadi-Abkenar, A.; Omidbaigi, R. Delay expression of limonoid UDP-glucosyltransferase makes delayed bitterness in citrus. Biochem. Biophys. Res. Commun. 2008, 371, 59–62. [Google Scholar] [CrossRef]
- Frydman, A.; Weisshaus, O.; Bar-Peled, M.; Huhman, D.V.; Sumner, L.W.; Marin, F.R.; Lewinsohn, E.; Fluhr, R.; Gressel, J.; Eyal, Y. Citrus fruit bitter flavors: Isolation and functional characterization of the gene Cm1,2RhaT encoding a 1,2 rhamnosyltransferase, a key enzyme in the biosynthesis of the bitter flavonoids of citrus. Plant J. 2004, 40, 88–100. [Google Scholar] [CrossRef]
- Lewinsohn, E.; Britsch, L.; Mazur, Y.; Gressel, J. Flavanone glycoside biosynthesis in citrus: Chalcone synthase, UDP-glucose: flavanone-7-o-glucosyl-transferase and -rhamnosyl-transferase activities in cell-free extracts. Plant Physiol. 1989, 91, 1323–1328. [Google Scholar] [CrossRef]
- Stevens, Y.; Van Rymenant, E.; Grootaert, C.; Van Camp, J.; Possemiers, S.; Masclee, A.; Jonkers, D. The intestinal fate of citrus flavanones and their effects on gastrointestinal health. Nutrients 2019, 11, 1464. [Google Scholar] [CrossRef]
- Li, L.J.; Wu, Z.Y.; Yu, Y.; Zhang, L.J.; Zhu, Y.B.; Ni, H.; Chen, F. Development and characterization of an alpha-l-rhamnosidase mutant with improved thermostability and a higher efficiency for debittering orange juice. Food Chem. 2018, 245, 1070–1078. [Google Scholar] [CrossRef]
- Bar-Peled, M.; Fluhr, R.; Gressel, J. Juvenile-specific localization and accumulation of a rhamnosyltransferase and its bitter flavonoid in foliage, flowers, and young citrus fruits. Plant Physiol. 1993, 103, 1377–1384. [Google Scholar] [CrossRef] [PubMed]
- Panche, A.N.; Diwan, A.D.; Chandra, S.R. Flavonoids: An overview. J. Nutr. Sci. 2016, 5. [Google Scholar] [CrossRef] [PubMed]
- Berhow, M.A.; Tisserat, B.; Kanes, K.; Vandercook, C. Survey of phenolic compounds produced in citrus. In US Department of Agriculture/Agricultural Research Service; Bulletin, T., Ed.; US Government Printing Office: Washington, DC, USA, 1998. [Google Scholar]
- Ohashi, T.; Hasegawa, Y.; Misaki, R.; Fujiyama, K. Substrate preference of citrus naringenin rhamnosyltransferases and their application to flavonoid glycoside production in fission yeast. Appl. Microbiol. Biotechnol. 2016, 100, 687–696. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Yuan, Z.; Zhang, H.; Li, W.; Shi, M.; Peng, Z.; Li, M.; Tian, J.; Deng, X.; Cheng, Y.; et al. Cit1,2RhaT and two novel CitdGlcTs participate in flavor-related flavonoid metabolism during citrus fruit development. J. Exp. Bot. 2019, 70, 2759–2771. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.; Liu, X.; Xiong, B.; Qiu, X.; Sun, G.; Wang, X.; Zhang, X.; Dong, Z.; Wang, Z. Variation in limonin and nomilin content in citrus fruits of eight varieties determined by modified HPLC. Food Sci. Biotechnol. 2019, 28, 641–647. [Google Scholar] [CrossRef]
- Breksa, A.P.; Zukas, A.A.; Manners, G.D. Determination of limonoate and nomilinoate A-ring lactones in citrus juices by liquid chromatography-electrospray ionization mass spectrometry. J. Chromatogr. A 2005, 1064, 187–191. [Google Scholar] [CrossRef]
- Maier, V.P.; Hasegawa, S.; Bennett, R.D.; Echols, L.C. Limonin and limonoids chemistry, biochemistry, and juice bitterness. Citrus Nutr. Qual. 1980, 143, 63–82, Chapter 4. [Google Scholar]
- Kita, M.; Endo, T.; Shimada, T.; Moriguchi, T.; Hirata, Y.; Hasegawa, S.; Omura, M. Allelic structures of UDP-glucose: Limonoid glucosyltransferase affect limonoid bitterness in Citrus unshiu and C. sinensis. Euphytica 2003, 132, 87–94. [Google Scholar] [CrossRef]
- Arora, S.; Mohanpuria, P.; Sidhu, G.S.; Yadav, I.S.; Kumari, V. Cloning and characterization of limonoid glucosyltransferase from Kinnow Mandarin (Citrus reticulata Blanco). Food Technol. Biotechnol. 2018, 56, 228–237. [Google Scholar] [CrossRef]
- Endo, T.; Kita, M.; Shimada, T.; Moriguchi, T.; Hidaka, T.; Matsumoto, R.; Hasegawa, S.; Omura, M. Modification of limonoid metabolism in suspension cell culture of citrus. Plant Biotechnol. 2002, 19, 397–403. [Google Scholar] [CrossRef]
- Herman, Z.; Fong, C.H.; Ou, P.; Hasegawa, S. Limonoid glucosides in orange juices by HPLC. J. Agric. Food Chem. 1990, 38, 1860–1861. [Google Scholar] [CrossRef]
- Vikram, A.; Jayaprakasha, G.K.; Patil, B.S. Simultaneous determination of citrus limonoid aglycones and glucosides by high performance liquid chromatography. Anal. Chim. Acta 2007, 590, 180–186. [Google Scholar] [CrossRef] [PubMed]
- Ishii, T.; Ohta, H.; Nogata, Y.; Yano, M.; Hasegawa, S. Limonoids in seeds of iyo tangor (Citrus iyo hort. ex Tanaka). Food Sci. Technol. Res. 2003, 9, 162–164. [Google Scholar] [CrossRef]
- Kita, M.; Hirata, Y.; Moriguchi, T.; Endo-Inagaki, T.; Matsumoto, R.; Hasegawa, S.; Suhayda, C.G.; Omura, M. Molecular cloning and characterization of a novel gene encoding limonoid UDP-glucosyltransferase in Citrus. FEBS Lett. 2000, 469, 173–178. [Google Scholar] [CrossRef]
- Etienne, A.; Genard, M.; Lobit, P.; Mbeguie, A.M.D.; Bugaud, C. What controls fleshy fruit acidity? A review of malate and citrate accumulation in fruit cells. J. Exp. Bot. 2013, 64, 1451–1469. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.H.; Liu, J.J.; Chen, K.L.; Li, H.W.; He, J.; Guan, B.; He, L. Comparative transcriptome and proteome profiling of two Citrus sinensis cultivars during fruit development and ripening. BMC Genom. 2017, 18. [Google Scholar] [CrossRef]
- Goldenberg, L.; Yaniv, Y.; Porat, R.; Carmi, N. Mandarin fruit quality: A review. J. Sci. Food Agric. 2018, 98, 18–26. [Google Scholar] [CrossRef]
- Al-Mouei, R.; Choumane, W. Physiochemical juice characteristics of various citrus species in Syria. Int. J. Plant Soil Sci. 2014, 3, 1083–1095. [Google Scholar] [CrossRef]
- Güler, Z.; Gül, E. Volatile organic compounds in the aril juices and seeds from selected five pomegranate (Punica granatum L.) cultivars. Int. J. Food Prop. 2017, 20, 281–293. [Google Scholar] [CrossRef]
- Butelli, E.; Licciardello, C.; Ramadugu, C.; Durand-Hulak, M.; Celant, A.; Reforgiato Recupero, G.; Froelicher, Y.; Martin, C. Noemi controls production of flavonoid pigments and fruit acidity and illustrates the domestication routes of modern citrus varieties. Curr. Biol. 2019, 29, 158–164. [Google Scholar] [CrossRef]
- Terol, J.; Soler, G.; Talon, M.; Cercos, M. The aconitate hydratase family from Citrus. BMC Plant Biol. 2010, 10. [Google Scholar] [CrossRef] [PubMed]
- Cercos, M.; Soler, G.; Iglesias, D.J.; Gadea, J.; Forment, J.; Talon, M. Global analysis of gene expression during development and ripening of citrus fruit flesh. A proposed mechanism for citric acid utilization. Plant Mol. Biol. 2006, 62, 513–527. [Google Scholar] [CrossRef] [PubMed]
- Degu, A.; Hatew, B.; Nunes-Nesi, A.; Shlizerman, L.; Zur, N.; Katz, E.; Fernie, A.R.; Blumwald, E.; Sadka, A. Inhibition of aconitase in citrus fruit callus results in a metabolic shift towards amino acid biosynthesis. Planta 2011, 234, 501–513. [Google Scholar] [CrossRef] [PubMed]
- Brune, A.; Gonzalez, P.; Goren, R.; Zehavi, U.; Echeverria, E. Citrate uptake into tonoplast vesicles from acid lime (Citrus aurantifolia) juice cells. J. Membr. Biol. 1998, 166, 197–203. [Google Scholar] [CrossRef] [PubMed]
- Ratajczak, R. Structure, function and regulation of the plant vacuolar H+-translocating ATPase. Biochim. Biophys. Acta 2000, 1465, 17–36. [Google Scholar] [CrossRef]
- Li, S.J.; Yin, X.R.; Xie, X.L.; Allan, A.C.; Ge, H.; Shen, S.L.; Chen, K.S. The Citrus transcription factor, CitERF13, regulates citric acid accumulation via a protein-protein interaction with the vacuolar proton pump, CitVHA-c4. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef] [PubMed]
- Shimada, T.; Nakano, R.; Shulaev, V.; Sadka, A.; Blumwald, E. Vacuolar citrate/H+ symporter of citrus juice cells. Planta 2006, 224, 472–480. [Google Scholar] [CrossRef]
- Brune, A. Vacuolar acidification in citrus fruit: Comparison between acid lime (Citrus aurantifolia) and sweet lime (Citrus limmetioides) juice cells. J. Am. Soc. Hort. Sci. 2002, 127, 171–177. [Google Scholar] [CrossRef]
- Muller, M.; Irkens-Kiesecker, U.; Rubinstein, B.; Taiz, L. On the mechanism of hyperacidification in lemon. Comparison of the vacuolar H+-ATPase activities of fruits and epicotyls. J. Biol. Chem. 1996, 271, 1916–1924. [Google Scholar] [CrossRef]
- Faraco, M.; Spelt, C.; Bliek, M.; Verweij, W.; Hoshino, A.; Espen, L.; Prinsi, B.; Jaarsma, R.; Tarhan, E.; de Boer, A.H.; et al. Hyperacidification of vacuoles by the combined action of two different P-ATPases in the tonoplast determines flower color. Cell Rep. 2014, 6, 32–43. [Google Scholar] [CrossRef]
- Strazzer, P.; Spelt, C.E.; Li, S.; Bliek, M.; Federici, C.T.; Roose, M.L.; Koes, R.; Quattrocchio, F.M. Hyperacidification of Citrus fruits by a vacuolar proton-pumping P-ATPase complex. Nat. Commun. 2019, 10, 744. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Ye, J.; Liu, C.; Xu, Q.; Long, L.; Deng, X. The citrus CsPH4-Noemi regulatory complex is involved in proanthocyanidin biosynthesis via a positive feedback loop. J. Exp. Bot. 2019. [Google Scholar] [CrossRef] [PubMed]
- Shi, C.Y.; Hussain, S.B.; Yang, H.; Bai, Y.X.; Khan, M.A.; Liu, Y.Z. CsPH8, a P-type proton pump gene, plays a key role in the diversity of citric acid accumulation in Citrus fruits. Plant Sci. 2019, 289, 110288. [Google Scholar] [CrossRef] [PubMed]
- Shi, C.Y.; Song, R.Q.; Hu, X.M.; Liu, X.; Jin, L.F.; Liu, Y.Z. Citrus PH5-like H+-ATPase genes: Identification and transcript analysis to investigate their possible relationship with citrate accumulation in fruits. Front. Plant Sci. 2015, 6. [Google Scholar] [CrossRef]
- Zhang, S.; Liang, M.; Wang, N.; Xu, Q.; Deng, X.; Chai, L. Reproduction in woody perennial Citrus: An update on nucellar embryony and self-incompatibility. Plant Reprod. 2018, 31, 43–57. [Google Scholar] [CrossRef]
- Endo, T.; Shimada, T.; Fujii, H.; Kobayashi, Y.; Araki, T.; Omura, M. Ectopic expression of an FT homolog from citrus confers an early flowering phenotype on trifoliate orange (Poncirus trifoliata L. Raf.). Transgenic Res. 2005, 14, 703–712. [Google Scholar] [CrossRef]
- Dutt, M.; Grosser, J. Using genetically modified biotechnology to improve citrus. Citrus Ind. 2005, 10–13. Available online: https://crec.ifas.ufl.edu/extension/trade_journals/2015/2015_September_biotech.pdf (accessed on 31 October 2019).
- Velazquez, K.; Aguero, J.; Vives, M.C.; Aleza, P.; Pina, J.A.; Moreno, P.; Navarro, L.; Guerri, J. Precocious flowering of juvenile citrus induced by a viral vector based on Citrus leaf blotch virus: A new tool for genetics and breeding. Plant Biotechnol. J. 2016, 14, 1976–1985. [Google Scholar] [CrossRef]
- Iwata, H.; Minamikawa, M.F.; Kajiya-Kanegae, H.; Ishimori, M.; Hayashi, T. Genomics-assisted breeding in fruit trees. Breed. Sci. 2016, 66, 100–115. [Google Scholar] [CrossRef]
- Yu, Y.; Bai, J.; Chen, C.; Plotto, A.; Yu, Q.; Baldwin, E.A.; Gmitter, F.G., Jr. Identification of QTLs controlling aroma volatiles using a ‘Fortune’ × ‘Murcott’ (Citrus reticulata) population. BMC Genom. 2017, 18. [Google Scholar] [CrossRef]
- Rommens, C.M. All-native DNA transformation: A new approach to plant genetic engineering. Trends Plant Sci. 2004, 9, 457–464. [Google Scholar] [CrossRef] [PubMed]
- Butelli, E.; Licciardello, C.; Zhang, Y.; Liu, J.; Mackay, S.; Bailey, P.; Reforgiato-Recupero, G.; Martin, C. Retrotransposons control fruit-specific, cold-dependent accumulation of anthocyanins in blood oranges. Plant Cell 2012, 24, 1242–1255. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Acanda, Y.; Canton, M.; Zale, J. Efficient biolistic transformation of immature citrus rootstocks using phosphomannose-isomerase selection. Plants 2019, 8, 390. [Google Scholar] [CrossRef] [PubMed]
- Schaeffer, S.M.; Nakata, P.A. The expanding footprint of CRISPR/Cas9 in the plant sciences. Plant Cell Rep. 2016, 35, 1451–1468. [Google Scholar] [CrossRef] [PubMed]
- Jia, H.; Wang, N. Targeted genome editing of sweet orange using Cas9/sgRNA. PLoS ONE 2014, 9, e93806. [Google Scholar] [CrossRef]
- Jia, H.; Zou, X.; Orbovic, V.; Wang, N. Genome editing in citrus tree with CRISPR/Cas9. Methods Mol. Biol. 2019, 1917, 235–241. [Google Scholar]
- Jia, H.; Xu, J.; Orbovic, V.; Zhang, Y.; Wang, N. Editing citrus genome via SaCas9/sgRNA system. Front. Plant Sci. 2017, 8. [Google Scholar] [CrossRef]
- Peng, A.; Chen, S.; Lei, T.; Xu, L.; He, Y.; Wu, L.; Yao, L.; Zou, X. Engineering canker-resistant plants through CRISPR/Cas9-targeted editing of the susceptibility gene CsLOB1 promoter in citrus. Plant Biotechnol. J. 2017, 15, 1509–1519. [Google Scholar] [CrossRef]
- Jia, H.; Zhang, Y.; Orbovic, V.; Xu, J.; White, F.F.; Jones, J.B.; Wang, N. Genome editing of the disease susceptibility gene CsLOB1 in citrus confers resistance to citrus canker. Plant Biotechnol. J. 2017, 15, 817–823. [Google Scholar] [CrossRef]
- Zhu, C.; Zheng, X.; Huang, Y.; Ye, J.; Chen, P.; Zhang, C.; Zhao, F.; Xie, Z.; Zhang, S.; Wang, N.; et al. Genome sequencing and CRISPR/Cas9 gene editing of an early flowering Mini-Citrus (Fortunella hindsii). Plant Biotechnol. J. 2019, 17, 2199–2210. [Google Scholar] [CrossRef]
- Jia, H.; Orbovic, V.; Wang, N. CRISPR-LbCas12a-mediated modification of citrus. Plant Biotechnol. J. 2019, 17, 1928–1937. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Ke, F.; Nie, Z.; Wang, P.; Xu, J. Citrus genetic engineering for disease resistance: Past, present and future. Int. J. Mol. Sci. 2019, 20, 5256. [Google Scholar] [CrossRef]
- Yu, Y.; Bai, J.; Chen, C.; Plotto, A.; Baldwin, E.A.; Gmitter, F.G. Comparative analysis of juice volatiles in selected mandarins, mandarin relatives and other citrus genotypes. J. Sci. Food Agric. 2018, 98, 1124–1131. [Google Scholar] [CrossRef] [PubMed]
- Abouzari, A.; Mahdi Nezhad, N. The investigation of citrus fruit quality. Popular characteristic and breeding. Acta Univ. Agric. Silvic. Mendel. Brun. 2016, 64, 725–740. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, L.-J.; Tan, W.-S.; Li, W.-J.; Zhu, Y.-B.; Cheng, Y.-S.; Ni, H. Citrus Taste Modification Potentials by Genetic Engineering. Int. J. Mol. Sci. 2019, 20, 6194. https://doi.org/10.3390/ijms20246194
Li L-J, Tan W-S, Li W-J, Zhu Y-B, Cheng Y-S, Ni H. Citrus Taste Modification Potentials by Genetic Engineering. International Journal of Molecular Sciences. 2019; 20(24):6194. https://doi.org/10.3390/ijms20246194
Chicago/Turabian StyleLi, Li-Jun, Wan-Seng Tan, Wen-Jing Li, Yan-Bing Zhu, Yi-Sheng Cheng, and Hui Ni. 2019. "Citrus Taste Modification Potentials by Genetic Engineering" International Journal of Molecular Sciences 20, no. 24: 6194. https://doi.org/10.3390/ijms20246194
APA StyleLi, L.-J., Tan, W.-S., Li, W.-J., Zhu, Y.-B., Cheng, Y.-S., & Ni, H. (2019). Citrus Taste Modification Potentials by Genetic Engineering. International Journal of Molecular Sciences, 20(24), 6194. https://doi.org/10.3390/ijms20246194





