RNA Sequencing Reveals Differential Gene Expression of Cerrena Unicolor in Response to Variable Lighting Conditions
Abstract
1. Introduction
2. Results
2.1. Transcriptome Sequencing and Identification of Transcripts Coding for Fungal Photoreceptors
2.2. Analysis of Differentially Expressed Genes (DEGs)
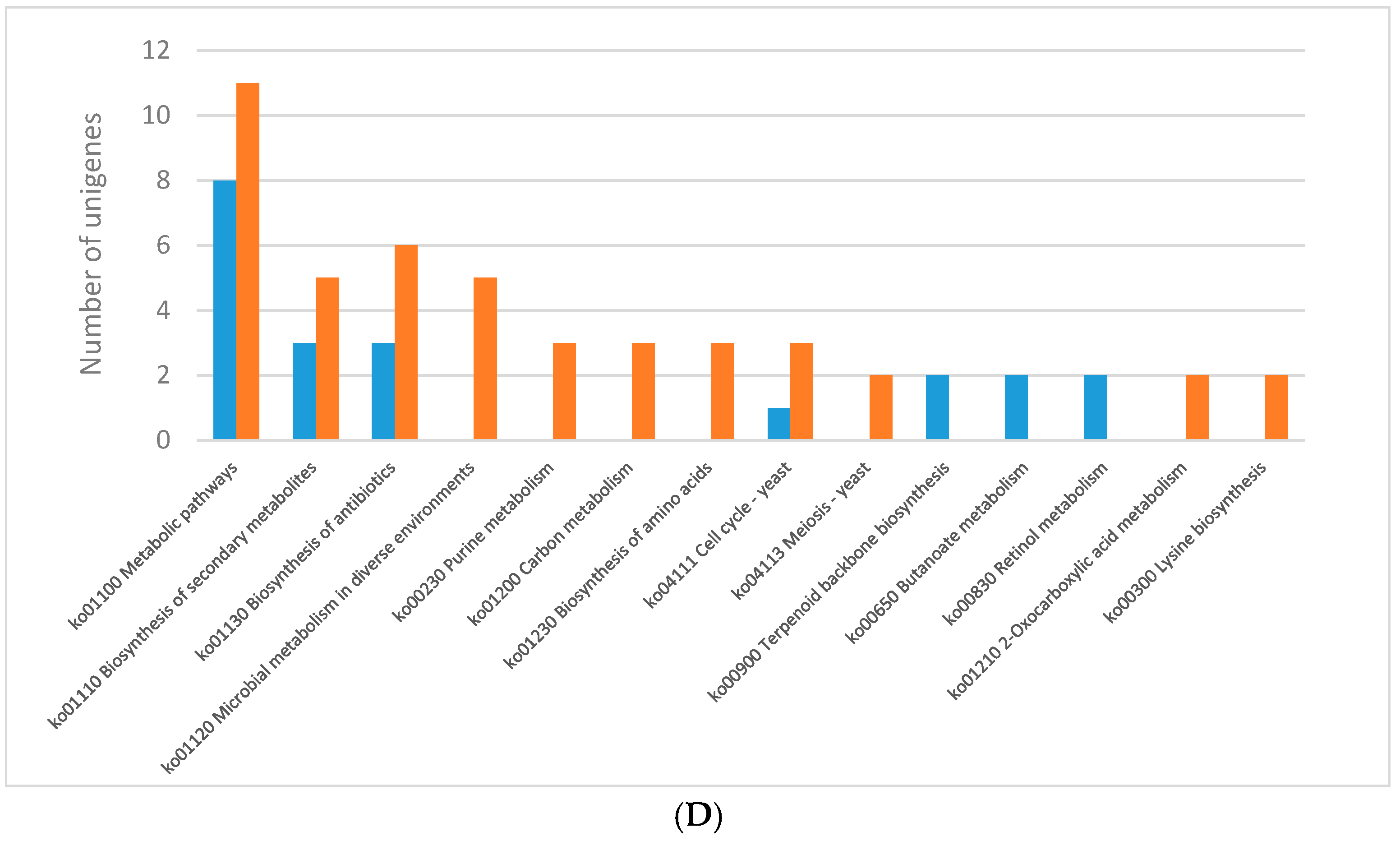
2.3. KEGG Pathway Enrichment of DEGs
2.3.1. Light Regulates Primary Metabolism of C. Unicolor
2.3.2. Effect of Light on the Expression of Wood-Degrading Enzymes
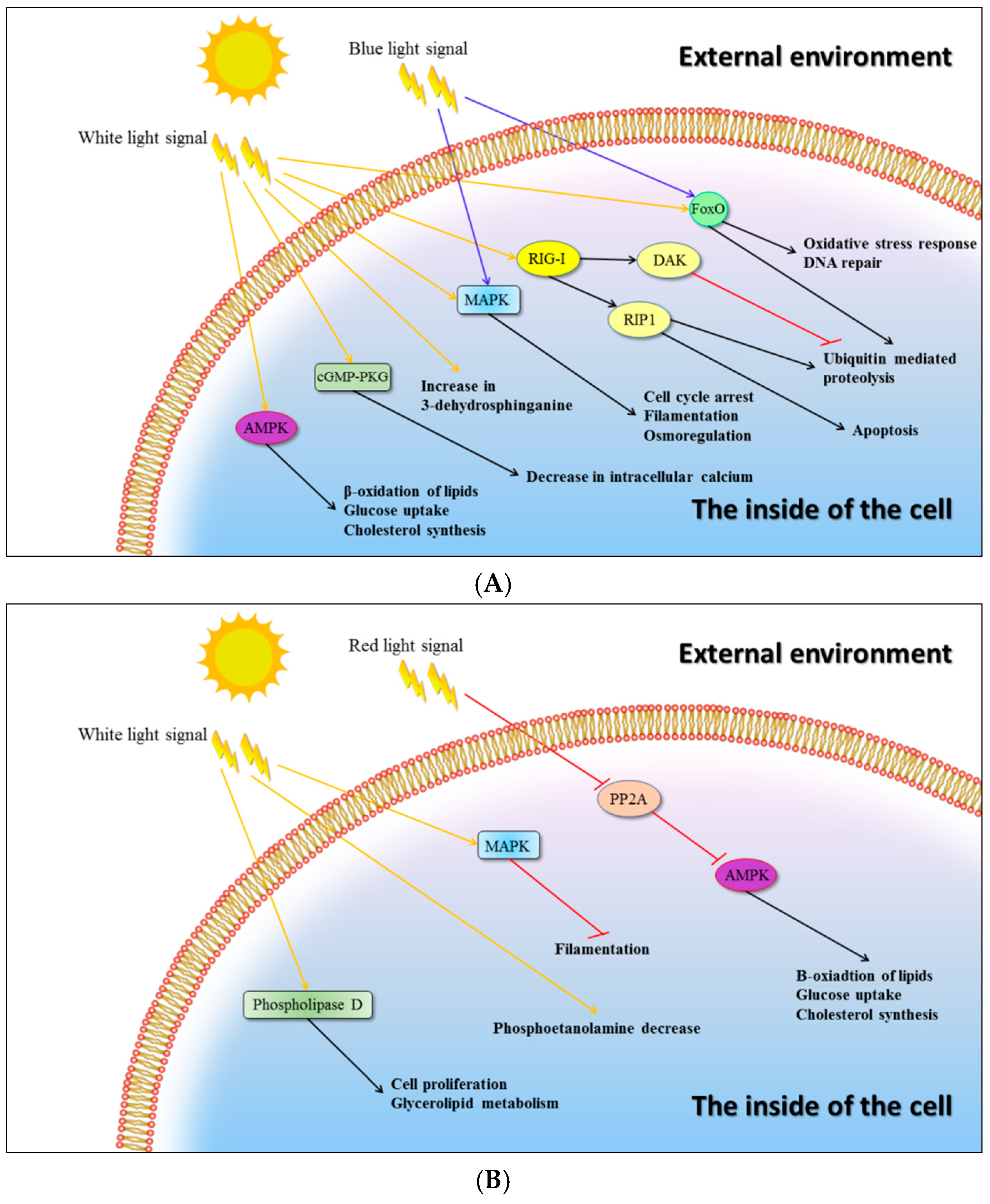
2.4. Signalling Pathways
3. Discussion
4. Materials and Methods
4.1. Medium and Growth Conditions
4.2. RNA Extraction and Sequencing
4.3. NGS Data Analysis
4.4. Nucleotide Sequence Accession Numbers
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| DEGs | Differentially Expressed Genes |
| LDA | Lignin-Degrading Auxiliary Enzyme |
| LME | Lignin-Modifying Enzyme |
Appendix A
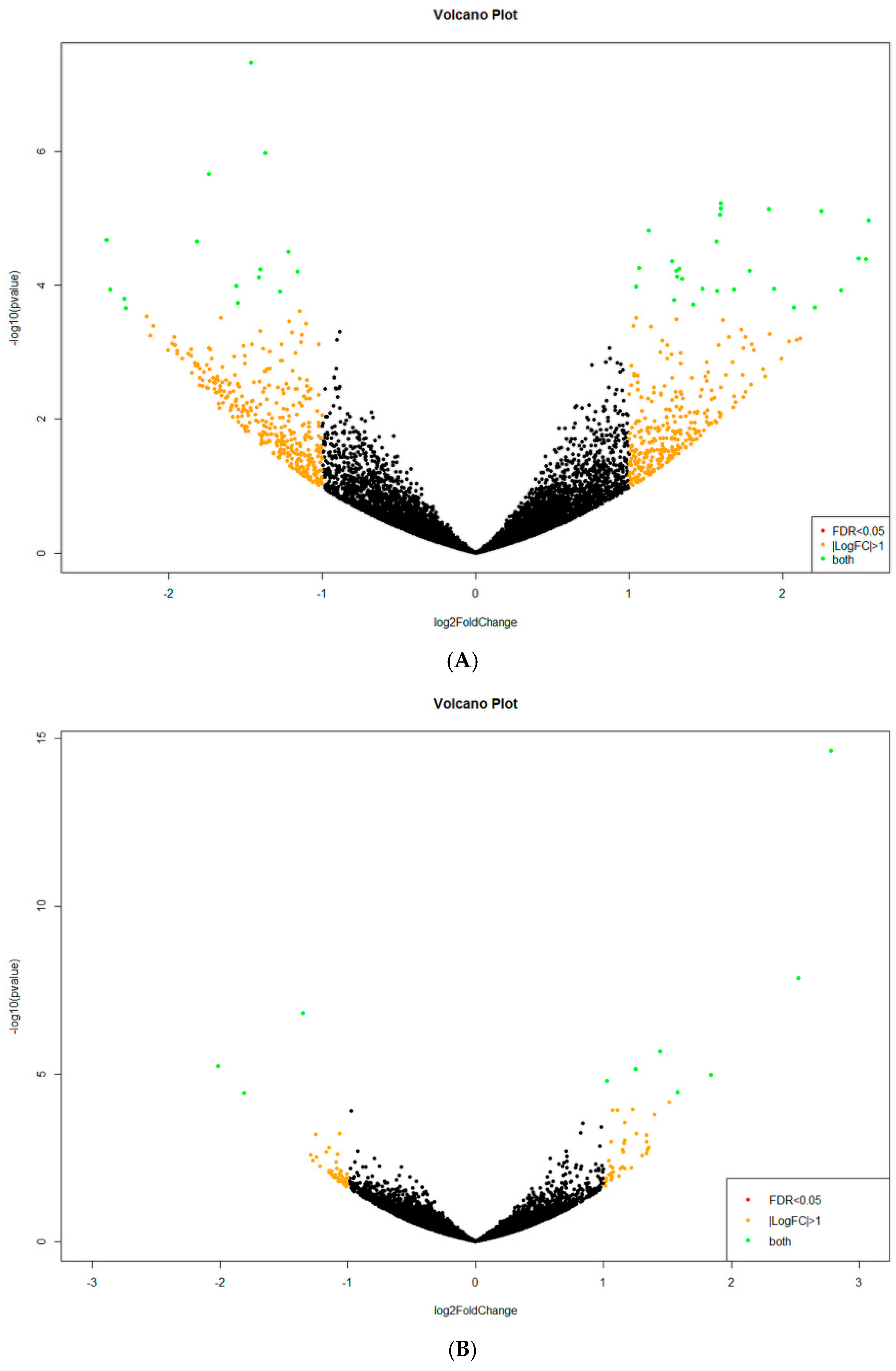
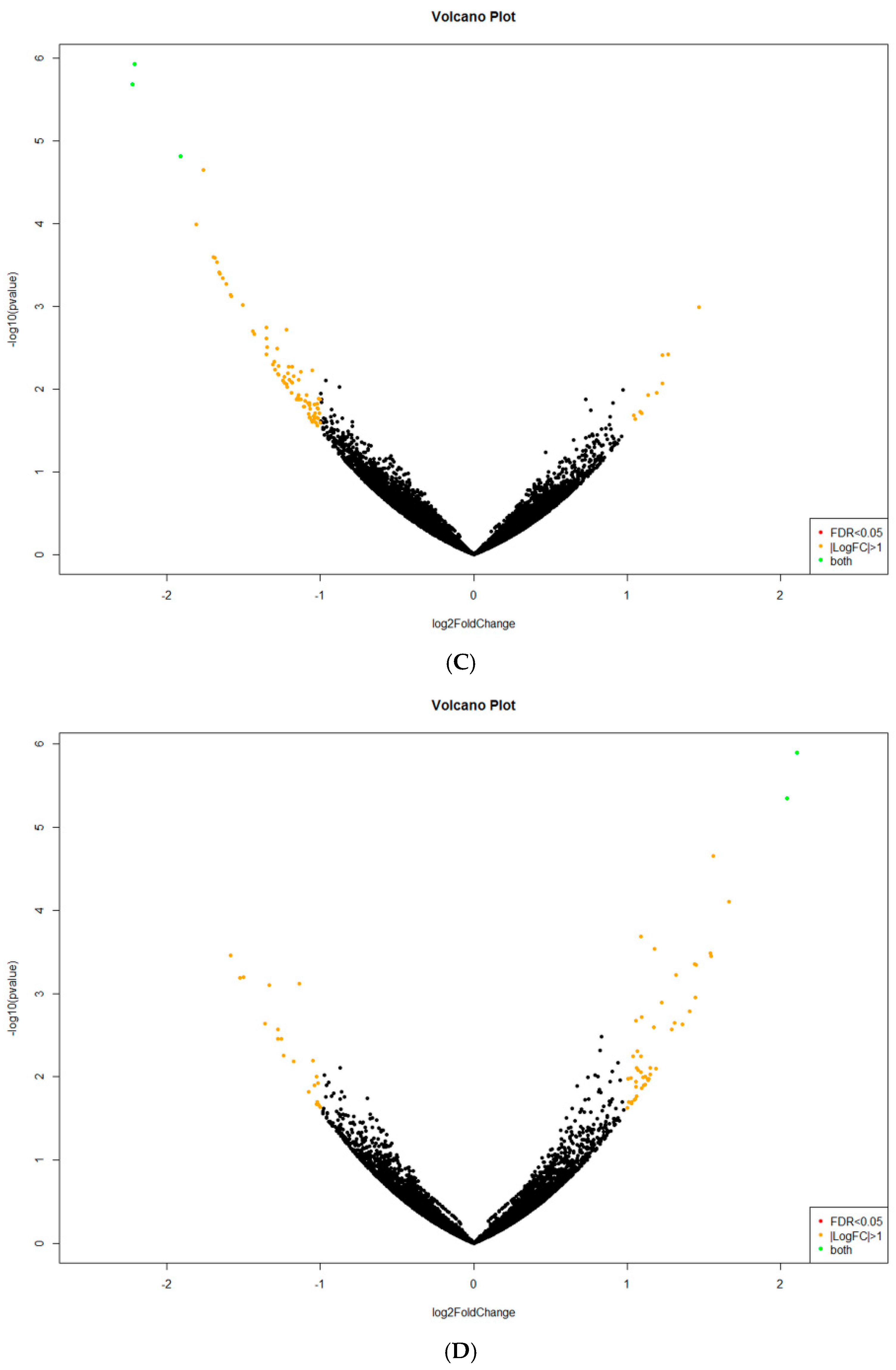
Appendix B
| Gene ID | Blast Best Hit | Signalling Pathway | Expression | |
|---|---|---|---|---|
| Log2 Fold Change | p-Value | |||
| White vs. Dark | ||||
| XLOC_005484 | Ras-related protein RABB1c | AMPK | 1.747524 | 0.002175 |
| XLOC_000988 | Dihydroxyacetone kinase 2 | RIG-I-like receptor | 1.59843 | 7.42 × 10−6 |
| XLOC_003649 | Calcium-transporting ATPase sarcoplasmic/endoplasmic reticulum type | cGMP-PKG | 1.344607 | 0.026421 |
| XLOC_010509 | Flocculation protein FLO11 | MAPK | 1.340754 | 0.009125 |
| XLOC_002816 | Transcription factor atf1 | MAPK | 1.325213 | 0.024773 |
| XLOC_008849 | Serine palmitoyltransferase 2 | Increase in 3-dehydrosphinganine | 1.145672 | 0.042199 |
| XLOC_001876 | Serine/threonine-protein phosphatase 2B catalytic subunit | cGMP-PKG/MAPK | 1.068844 | 0.03373 |
| XLOC_008348 | Serine/threonine-protein kinase STK11 | AMPK/FoxO | 0.958811 | 0.008435 |
| XLOC_002560 | Mitogen-activated protein kinase dlk-1 | MAPK | 0.942856 | 0.00767 |
| XLOC_010380 | Receptor-interacting serine/threonine-protein kinase 1 | RIG-I-like receptor | 0.786022 | 0.015669 |
| XLOC_000296 | Protein phosphatase 2C homolog 1 | MAPK | 0.734535 | 0.010766 |
| XLOC_003540 | Striatin-3 | MAPK | −0.66931 | 0.0101 |
| XLOC_008420 | DNA damage checkpoint protein rad24 | MAPK | −0.79174 | 0.041947 |
| XLOC_001763 | ADP-ribosylation factor 6 | Phospholipase D | −1.17631 | 0.003653 |
| XLOC_000832 | Sphingosine-1-phosphate lyase | Phosphoetanolamine decrease | −1.30105 | 0.018164 |
| Blue vs. Dark | ||||
| XLOC_010509 | Flocculation protein FLO11 | MAPK | 0.709652 | 0.026203 |
| XLOC_009396 | Catalase-1 | MAPK/FoxO | 0.590792 | 0.035802 |
| Red vs. Dark | ||||
| XLOC_001068 | Protein phosphatase PP2A regulatory subunit B | PP2A/AMPK | −0.79388 | 0.04389 |
References
- Anastasi, A.; Vizzini, A.; Prigione, V.; Varese, G. Wood degrading fungi: Morphology, metabolism and environmental applications. In A Textbook of Molecular Biotechnology, 0th ed.; Chauhan, A.K., Varma, A., Eds.; I.K. International: New Delhi, India, 2009; pp. 957–993. ISBN 9789380026374. [Google Scholar]
- Mester, T.; Varela, E.; Tien, M. Wood Degradation by Brown-Rot and White-Rot Fungi. In Genetics and Biotechnology; Kück, U., Ed.; Springer: Berlin/Heidelberg, Germany, 2004; pp. 355–368. ISBN 9783642076671. [Google Scholar]
- Pinzari, F.; Ceci, A.; Abu-Samra, N.; Canfora, L.; Maggi, O.; Persiani, A. Phenotype MicroArray system in the study of fungal functional diversity and catabolic versatility. Res. Microbiol. 2016, 167, 710–722. [Google Scholar] [CrossRef] [PubMed]
- Blanchette, R.A.; Nilsson, T.; Daniel, G.; Abad, A. Biological Degradation of Wood. Biol. Degrad. Wood 1990, 225, 141–174. [Google Scholar]
- Tisch, D.; Schmoll, M. Light regulation of metabolic pathways in fungi. Appl. Microbiol. Biotechnol. 2010, 85, 1259–1277. [Google Scholar] [CrossRef] [PubMed]
- Fischer, R.; Aguirre, J.; Herrera-Estrella, A.; Corrochano, L.M. The complexity of fungal vision. Microbiol. Spectr. 2016, 4. [Google Scholar] [CrossRef]
- Fuller, K.K.; Dunlap, J.C.; Loros, J.J. Fungal Light Sensing at the Bench and Beyond. Adv. Genet. 2016, 96, 1–51. [Google Scholar] [PubMed]
- Velculescu, V.E.; Zhang, L.; Zhou, W.; Vogelstein, J.; Basrai, M.A.; Bassett, D.E.; Hieter, P.; Vogelstein, B.; Kinzler, K.W. Characterization of the yeast transcriptome. Cell 1997, 88, 243–251. [Google Scholar] [CrossRef]
- Martinez, D.; Challacombe, J.; Morgenstern, I.; Hibbett, D.; Schmoll, M.; Kubicek, C.P.; Ferreira, P.; Ruiz-Duenas, F.J.; Martinez, A.T.; Kersten, P.; et al. Genome, transcriptome, and secretome analysis of wood decay fungus Postia placenta supports unique mechanisms of lignocellulose conversion. Proc. Natl. Acad. Sci. USA 2009, 106, 1954–1959. [Google Scholar] [CrossRef]
- Rohr, C.O.; Levin, L.N.; Mentaberry, A.N.; Wirth, S.A. A first insight into Pycnoporus sanguineus BAFC 2126 transcriptome. PLoS ONE 2013, 8, e81033. [Google Scholar] [CrossRef]
- Yu, G.J.; Wang, M.; Huang, J.; Yin, Y.L.; Chen, Y.J.; Jiang, S.; Jin, Y.X.; Lan, X.Q.; Wong, B.H.; Liang, Y.; et al. Deep insight into the Ganoderma lucidum by comprehensive analysis of its transcriptome. PLoS ONE 2012, 7, e44031. [Google Scholar] [CrossRef]
- Hori, C.; Ishida, T.; Igarashi, K.; Samejima, M.; Suzuki, H.; Master, E.; Ferreira, P.; Ruiz-Duenas, F.J.; Held, B.; Canessa, P.; et al. Analysis of the Phlebiopsis gigantea genome, transcriptome and secretome provides insight into its pioneer colonization strategies of wood. PLoS Genet. 2014, 10, e1004759. [Google Scholar] [CrossRef]
- Huang, Y.; Wu, X.; Jian, D.; Zhan, Y.; Fan, G. De novo transcriptome analysis of a medicinal fungi Phellinus linteus and identification of SSR markers. Biotechnol. Biotechnol. Equip. 2015, 29, 395–403. [Google Scholar] [CrossRef] [PubMed]
- Janusz, G.; Mazur, A.; Wielbo, J.; Koper, P.; Żebracki, K.; Pawlik, A.; Ciołek, B.; Paszczyński, A.; Kubik-Komar, A. Comparative transcriptomic analysis of Cerrena unicolor revealed differential expression of genes engaged in degradation of various kinds of wood. Microbiol. Res. 2018, 207, 256–268. [Google Scholar] [CrossRef] [PubMed]
- Meijueiro, M.L.; Santoyo, F.; Ramirez, L.; Pisabarro, A.G. Transcriptome characteristics of filamentous fungi deduced using high-throughput analytical technologies. Brief Funct. Genom. 2014, 13, 440–450. [Google Scholar] [CrossRef] [PubMed]
- Velez, P.; Alejandri-Ramírez, N.D.; González, M.C.; Estrada, K.J.; Sanchez-Flores, A.; Dinkova, T.D. Comparative transcriptome analysis of the cosmopolitan marine fungus Corollospora maritima under two physiological conditions. G3 Genes Genomes Genet. 2015, 5, 1805–1814. [Google Scholar] [CrossRef] [PubMed]
- Szilágyi, M.; Miskei, M.; Karányi, Z.; Lenkey, B.; Pócsi, I.; Emri, T. Transcriptome changes initiated by carbon starvation in Aspergillus nidulans. Microbiology 2013, 159, 176–190. [Google Scholar] [CrossRef] [PubMed]
- Mizerska-Dudka, M.; Jaszek, M.; Blachowicz, A.; Rejczak, T.P.; Matuszewska, A.; Osinska-Jaroszuk, M.; Stefaniuk, D.; Janusz, G.; Sulej, J.; Kandefer-Szerszen, M. Fungus Cerrena unicolor as an effective source of new antiviral, immunomodulatory, and anticancer compounds. Int. J. Biol. Macromol. 2015, 79, 459–468. [Google Scholar] [CrossRef] [PubMed]
- Belova, O.V.; Lisov, A.V.; Vinokurova, N.G.; Kostenevich, A.A.; Sapunova, L.I.; Lobanok, A.G.; Leont’evskii, A. Xylanase and cellulase of fungus Cerrena unicolor VKM F-3196: Production, properties, and applications for the saccharification of plant material. Prikl. Biokhim. Mikrobiol. 2014, 50, 171–176. [Google Scholar] [CrossRef]
- Zhang, H.; Zhang, J.; Zhang, X.; Geng, A. Purification and characterization of a novel manganese peroxidase from white-rot fungus Cerrena unicolor BBP6 and its application in dye decolorization and denim bleaching. Proc. Biochem. 2018, 66, 222–229. [Google Scholar] [CrossRef]
- Janusz, G.; Jaszek, M.; Justyna, S.; Osińska-Jaroszuk, M. Effect of different wavelengths of light on laccase, cellobiose dehydrogenase, and proteases produced by Cerrena unicolor, Pycnoporus sanguineus, and Phlebia lindtneri. Acta Biochim. Pol. 2016, 63, 223–228. [Google Scholar] [CrossRef]
- Heberle, H.; Meirelles, G.V.; da Silva, F.R.; Telles, G.P.; Minghim, R. InteractiVenn: A web-based tool for the analysis of sets through Venn diagrams. BMC Bioinform. 2015, 16, 169. [Google Scholar] [CrossRef]
- Chen, C.-H.; Ringelberg, C.S.; Gross, R.H.; Dunlap, J.C.; Loros, J.J. Genome-wide analysis of light-inducible responses reveals hierarchical light signalling in Neurospora. EMBO J. 2009, 28, 1029–1042. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.; Yang, F.; Smith, K.M.; Peterson, M.; Dekhang, R.; Zhang, Y.; Zucker, J.; Bredeweg, E.L.; Mallappa, C.; Zhou, X.; et al. Genome-wide characterization of light-regulated genes in Neurospora crassa. G3 Genes Genomes Genet. 2014, 4, 1731–1745. [Google Scholar] [CrossRef] [PubMed]
- Sharrock, R.A. The phytochrome red/far-red photoreceptor superfamily. Genome Biol. 2008, 9, 230. [Google Scholar] [CrossRef] [PubMed]
- Xie, C.; Gong, W.; Zhu, Z.; Yan, L.; Hu, Z.; Peng, Y. Comparative transcriptomics of Pleurotus eryngii reveals blue-light regulation of carbohydrate-active enzymes (CAZymes) expression at primordium differentiated into fruiting body stage. Genomics 2018, 110, 201–209. [Google Scholar] [CrossRef] [PubMed]
- Ávalos, J.; Bejarano, E.R.; Cerdá-Olmedo, E. Photoinduction of carotenoid biosynthesis. Methods Enzymol. 1993, 214, 283–294. [Google Scholar]
- Wang, F.; Liu, Q.; Zhang, J.; Liu, K.; Li, K.; Liu, G.; Dong, C. Comparative transcriptome analysis between a spontaneous albino mutant and its sibling strain of Cordyceps militaris in response to light stress. Front. Microbiol. 2018, 9, 1237. [Google Scholar] [CrossRef] [PubMed]
- Dasgupta, A.; Fuller, K.K.; Dunlap, J.C.; Loros, J.J. Seeing the world differently: Variability in the photosensory mechanisms of two model fungi. Environ. Microbiol. 2016, 18, 5–20. [Google Scholar] [CrossRef] [PubMed]
- Bayram, Ö.; Feussner, K.; Dumkow, M.; Herrfurth, C.; Feussner, I.; Braus, G.H. Changes of global gene expression and secondary metabolite accumulation during light-dependent Aspergillus nidulans development. Fungal Genet. Biol. 2016, 87, 30–53. [Google Scholar] [CrossRef]
- Tani, S.; Kawaguchi, T.; Kobayashi, T. Complex regulation of hydrolytic enzyme genes for cellulosic biomass degradation in filamentous fungi. Appl. Microbiol. Biotechnol. 2014, 98, 4829–4837. [Google Scholar] [CrossRef]
- Tang, L.H.; Jian, H.H.; Song, C.Y.; Bao, D.P.; Shang, X.D.; Wu, D.Q.; Tan, Q.; Zhang, X.H. Transcriptome analysis of candidate genes and signaling pathways associated with light-induced brown film formation in Lentinula edodes. Appl. Microbiol. Biotechnol. 2013, 97, 4977–4989. [Google Scholar] [CrossRef]
- Purschwitz, J.; Muller, S.; Kastner, C.; Schoser, M.; Haas, H.; Espeso, E.A.; Atoui, A.; Calvo, A.M.; Fischer, R. Functional and physical interaction of blue- and red-light sensors in Aspergillus nidulans. Curr. Biol. 2008, 18, 255–259. [Google Scholar] [CrossRef] [PubMed]
- Heinrich, K.; Leslie, D.J.; Jonas, K. Modulation of bacterial proliferation as a survival strategy. Adv. Appl. Microbiol. 2015, 92, 127–171. [Google Scholar] [PubMed]
- Esquivel-Naranjo, E.U.; Garcia-Esquivel, M.; Medina-Castellanos, E.; Correa-Perez, V.A.; Parra-Arriaga, J.L.; Landeros-Jaime, F.; Cervantes-Chavez, J.A.; Herrera-Estrella, A. A Trichoderma atroviride stress-activated MAPK pathway integrates stress and light signals. Mol. Microbiol. 2016, 100, 860–876. [Google Scholar] [CrossRef] [PubMed]
- Brunner, M.; Kaldi, K. Interlocked feedback loops of the circadian clock of Neurospora crassa. Mol. Microbiol. 2008, 68, 255–262. [Google Scholar] [CrossRef] [PubMed]
- Vanden Wymelenberg, A.; Gaskell, J.; Mozuch, M.; Sabat, G.; Ralph, J.; Skyba, O.; Mansfield, S.D.; Blanchette, R.A.; Martinez, D.; Grigoriev, I.; et al. Comparative transcriptome and secretome analysis of wood decay fungi Postia placenta and Phanerochaete chrysosporium. Appl. Environ. Microbiol. 2010, 76, 3599–3610. [Google Scholar] [CrossRef] [PubMed]
- Schmoll, M. Regulation of plant cell wall degradation by light in Trichoderma. Fungal Biol. Biotechnol. 2018, 5, 10. [Google Scholar] [CrossRef]
- Sakamoto, Y.; Sato, S.; Ito, M.; Ando, Y.; Nakahori, K.; Muraguchi, H. Blue light exposure and nutrient conditions influence the expression of genes involved in simultaneous hyphal knot formation in Coprinopsis cinerea. Microbiol. Res. 2018, 217, 81–90. [Google Scholar] [CrossRef]
- Hernández, C.A.; Perroni, Y.; Pérez, J.A.G.; Rivera, B.G.; Alarcón, E. Light-induced inhibition of laccase in Pycnoporus sanguineus. Folia Microbiol. 2016, 61, 137–142. [Google Scholar] [CrossRef]
- Lindeberg, G.; Holm, G. Occurrence of tyrosinase and laccase in fruit bodies and mycelia of some Hymenomycetes. Physiol. Plant. 1952, 5, 100–114. [Google Scholar] [CrossRef]
- Homer, N.; Merriman, B.; Nelson, S.F. BFAST: An alignment tool for large scale genome resequencing. PLoS ONE 2009, 4, e7767. [Google Scholar] [CrossRef]
- Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011, 17, 10–12. [Google Scholar] [CrossRef]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Meth. 2012, 9, 357–359. [Google Scholar] [CrossRef] [PubMed]
- Liao, Y.; Smyth, G.K.; Shi, W. The Subread aligner: Fast, accurate and scalable read mapping by seed-and-vote. Nucl. Acids Res. 2013, 41, e108. [Google Scholar] [CrossRef]
- Roberts, A.; Pimentel, H.; Trapnell, C.; Pachter, L. Identification of novel transcripts in annotated genomes using RNA-Seq. Bioinformatics 2011, 27, 2325–2329. [Google Scholar] [CrossRef] [PubMed]
- Le Roes-Hill, M.; Rohland, J.; Burton, S. Actinobacteria isolated from termite guts as a source of novel oxidative enzymes. Antonie Van Leeuwenhoek 2011, 100, 589–605. [Google Scholar] [CrossRef] [PubMed]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef]
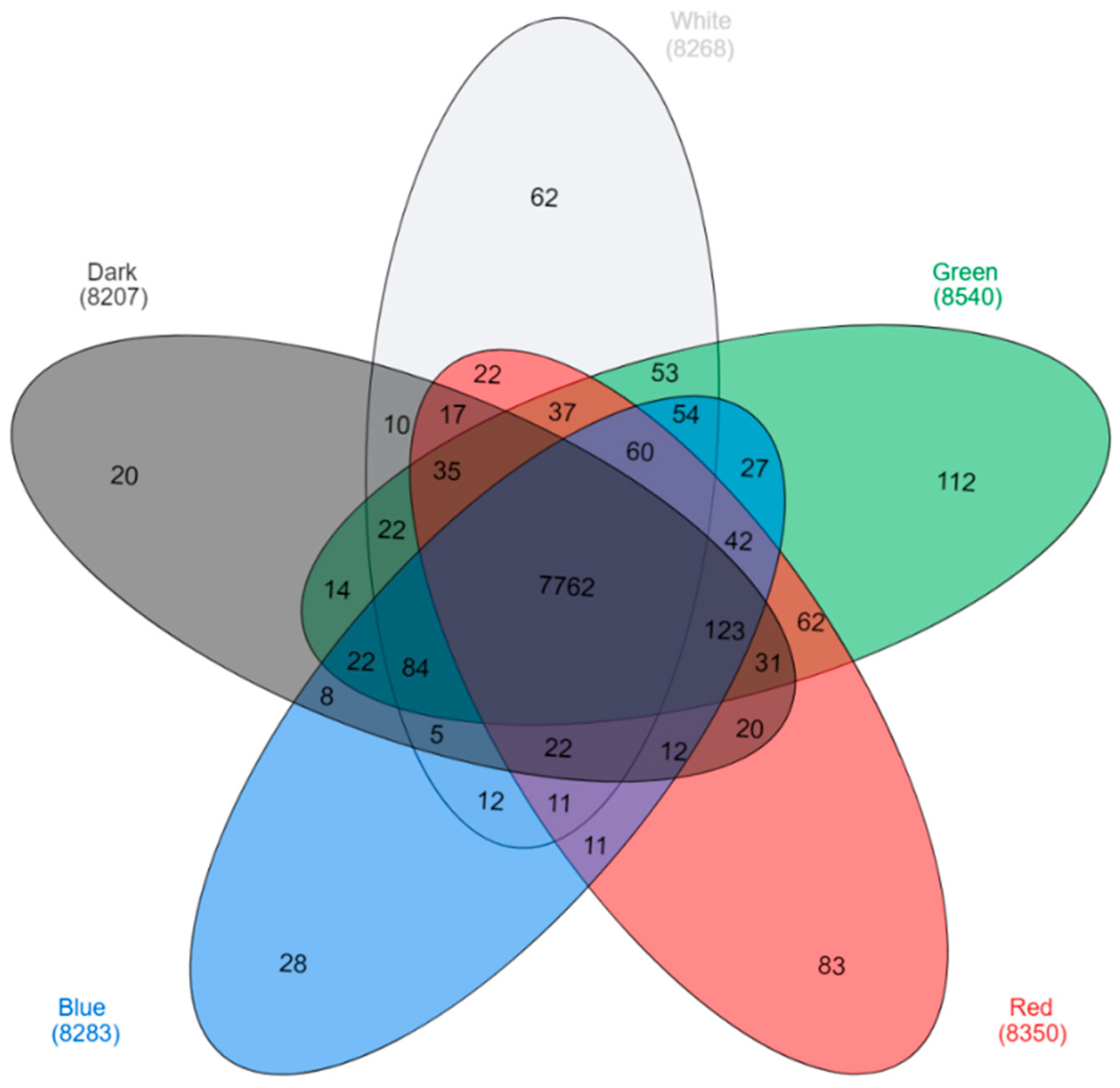
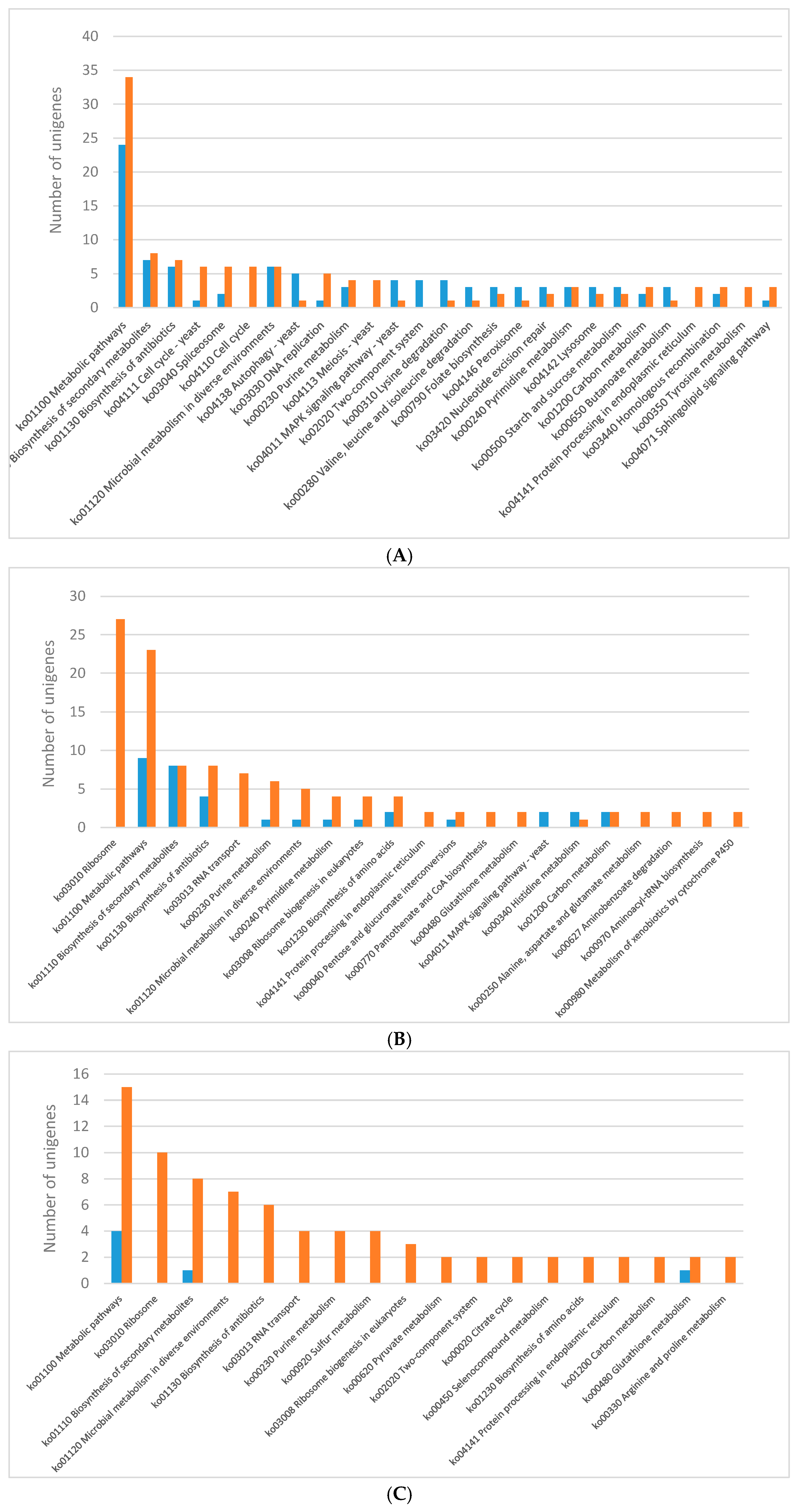
| Sample Name | Number of Total Reads | Average Number of Reads per Lighting Condition | Per cent of too Short Reads | Per cent of Reads with too Many Ns | Number of rRNA Mapped Reads | Number of Reads which Passed Filtering, Quality Control, rRNA Removal (% of Total Reads) 1 | Number of Mapped Reads 1 |
|---|---|---|---|---|---|---|---|
| W_1 | 112,287,853 | 120,864,091 | 16.4% | 69.4% | 458,752 | 15,517,301 (13.82%) | 47,884,777 |
| W_2 | 121,609,141 | 22.3% | 1.9% | 1,815,166 | 90,460,336 (74.39%) | ||
| W_3 | 128,695,279 | 13.9% | 1.2% | 3,393,460 | 105,877,886 (82.27%) | ||
| G_1 | 128,090,596 | 113,654,401 | 17.0% | 59.7% | 437,929 | 29,427,455 (22.97%) | 34,288,490 |
| G_2 | 131,301,239 | 19.2% | 2.2% | 1,151,865 | 102,046,831 (77.72%) | ||
| G_3 | 81,571,367 | 20.3% | 1.0% | 2,134,477 | 62,027,269 (76.04%) | ||
| B_1 | 120,729,656 | 128,318,401 | 17.8% | 1.0% | 2,226,961 | 95,742,071 (79.30%) | 47,208,269 |
| B_2 | 121,879,416 | 20.6% | 2.1% | 1,342,182 | 92,929,241 (76.25%) | ||
| B_2 | 142,346,130 | 36.5% | 0.9% | 1,787,916 | 87,349,092 (61.36%) | ||
| R_1 | 141,730,254 | 128,365,999 | 16.2% | 58.6% | 591,388 | 35,131,581 (24.79%) | 40,335,011 |
| R_2 | 121,611,135 | 22.3% | 1.9% | 1,519,062 | 90,665,568 (74.55%) | ||
| R_3 | 121,756,608 | 19.8% | 1.0% | 2,341,637 | 94,055,653 (77.25%) | ||
| D_1 | 139,936,639 | 123,716,947 | 13.9% | 1.1% | 3,078,557 | 115,934,413 (82.85%) | 48,645,244 |
| D_2 | 134,402,468 | 23.0% | 1.7% | 989,101 | 100,168,832 (74.53%) | ||
| D_3 | 96,811,733 | 31.3% | 1.8% | 918,966 | 63,824,504 (65.93%) |
| Gene ID | SwissProt Best Hit | Putative Function |
|---|---|---|
| XLOC_005534 | SwissProt_best_hit “White collar 2 protein (7e-12, sp|P78714|WC2_NEUCR)” | white collar protein |
| XLOC_005717 | SwissProt_best_hit “White collar 2 protein (1e-11, sp|P78714|WC2_NEUCR)” | white collar protein |
| XLOC_007605 | SwissProt_best_hit “White collar 1 protein (8e-63, sp|Q01371|WC1_NEUCR)” | white collar protein |
| XLOC_011520 | SwissProt_best_hit “White collar 1 protein (1e-10, sp|Q01371|WC1_NEUCR)” | white collar protein |
| XLOC_008457 | SwissProt_best_hit “Deoxyribodipyrimidine photo-lyase (6e-97, sp|P27526|PHR_NEUCR)” | cryptochrome |
| XLOC_003467 | SwissProt_best_hit “Deoxyribodipyrimidine photo-lyase (6e-97, sp|P27526|PHR_NEUCR” | cryptochrome |
| XLOC_010394 | SwissProt_best_hit “Light-sensor Protein kinase (2e-08, sp|P25848|PHY1_CERPU” | phytochrome |
| XLOC_011661 | Light-sensor Protein kinase (4e-17, sp|P25848|PHY1_CERPU) | phytochrome |
| XLOC_003467 | Protein FDD123 (1e-30, sp|O74631|FD123_TRAVE) | opsin |
| Compared Lightning Conditions | Number of Differentially Expressed Genes (at p < 0.05) | |
|---|---|---|
| Up-Regulated | Down-Regulated | |
| blue vs. dark | 176 | 227 |
| green vs. dark | 41 | 167 |
| red vs. dark | 132 | 95 |
| white vs. dark | 454 | 457 |
| Gene ID | Blast Best Hit | Putative Function | Expression | |
|---|---|---|---|---|
| log2 Fold Change | p-Value | |||
| White vs. Dark | ||||
| XLOC_007791 | Cytochrome P450 3A11 (2e-20, sp|Q64459|CP3AB_MOUSE) | LDA 1 | 1.527678 | 0.012165 |
| XLOC_005202 | Cytochrome P450 52A5 (7e-52, sp|Q12581|CP52X_CANMA) | LDA 1 | 1.312698 | 0.03231 |
| XLOC_004360 | Manganese peroxidase 3 (2e-75, sp|Q96TS6|PEM3_PHLRA) | LME 2 | 1.310116 | 0.015162 |
| XLOC_006405 | Beta-glucuronidase (3e-14, sp|A2QEQ6|GUS79_ASPNC) | hemicellulase | 1.18915 | 0.013524 |
| XLOC_007195 | Beta-glucosidase cel3A (3e-36, sp|G4NI45|CEL3A_MAGO7) | cellulase | 1.186582 | 0.028688 |
| XLOC_006061 | Cytochrome P450 734A1 (2e-23, sp|O48786|C734A_ARATH) | LDA 1 | 1.015716 | 0.001687 |
| XLOC_007410 | Cytochrome P450 4A10 (3e-20, sp|P08516|CP4AA_RAT) | LDA 1 | 0.996072 | 0.016423 |
| XLOC_000665 | Alcohol oxidase (6e-93, sp|P04841|ALOX_PICAN) | LDA 1 | 0.953991 | 0.014911 |
| XLOC_001403 | Cytochrome P450 67 (Fragment) (4e-60, sp|O00061|CP67_UROFA) | LDA 1 | 0.936259 | 0.012773 |
| XLOC_007076 | Xyloglucanase (0.0, sp|Q7Z9M8|XG74_HYPJQ) | hemicellulase | 0.863734 | 0.04043 |
| XLOC_002989 | Endo-1,4-beta-xylanase C (2e-132, sp|B7SIW2|XYNC_PHACH) | hemicellulase | 0.855198 | 0.017668 |
| XLOC_008979 | Laccase (1e-32, sp|O59896|LAC1_PYCCI) | LME 2 | −0.67001 | 0.031583 |
| XLOC_008955 | Laccase (2e-170, sp|Q02497|LAC1_TRAHI) | LME 2 | −0.92358 | 0.037087 |
| XLOC_007951 | Probable endo-beta-1,4-glucanase D (3e-34, sp|B0Y9G4|EGLD_ASPFC) | cellulase | −1.01988 | 0.044786 |
| XLOC_000669 | Laccase (2e-80, sp|Q01679|LAC1_PHLRA) | LME 2 | −1.172 | 0.045441 |
| XLOC_011551 | Laccase-2 (0.0, sp|Q12718|LAC2_TRAVE) | LME 2 | −1.53184 | 0.008822 |
| Blue vs. Dark | ||||
| XLOC_006977 | Probable beta-glucosidase G (2e-15, sp|B8NMR5|BGLG_ASPFN) | cellulase | 1.398314 | 0.00017 |
| XLOC_008717 | Aryl-alcohol dehydrogenase [NADP(+)] (4e-39, sp|Q01752|AAD_PHACH) | LDA 1 | 0.940399 | 0.030909 |
| XLOC_009983 | Pyranose 2-oxidase (3e-32, sp|Q6QWR1|P2OX_PHACH) | LDA 1 | 0.905345 | 0.041241 |
| XLOC_008418 | Beta-glucuronidase (2e-06, sp|A2QEQ6|GUS79_ASPNC) | hemicellulase | 0.640958 | 0.04928 |
| XLOC_003435 | Putative monooxygenase Rv1533 (1e-10, sp|O06179|Y1533_MYCTU) | LDA 1 | 0.615906 | 0.043912 |
| XLOC_001060 | Cytochrome P450 67 (Fragment) (2e-43, sp|O00061|CP67_UROFA) | LDA 1 | −0.4461 | 0.046049 |
| XLOC_006587 | Exoglucanase 1 (2e-111, sp|P13860|GUX1_PHACH) | cellulase | −0.47763 | 0.048334 |
| Green vs. Dark | ||||
| XLOC_006977 | Probable beta-glucosidase G (2e-15, sp|B8NMR5|BGLG_ASPFN) | cellulase | 0.93716 | 0.044217 |
| XLOC_006843 | Xyloglucan-specific endo-beta-1,4-glucanase A (7e-61, sp|Q5BG78|XGEA_EMENI) | hemicellulase | 0.736937 | 0.038777 |
| XLOC_008194 | Aryl-alcohol dehydrogenase [NADP(+)] (8e-126, sp|Q01752|AAD_PHACH) | LDA 1 | −0.85863 | 0.037114 |
| XLOC_010403 | Cytochrome P450 3A30 (4e-16, sp|Q9PVE8|C330_FUNHE) | LDA 1 | −1.78285 | 2.49 × 10−5 |
| Red vs. Dark | ||||
| XLOC_007791 | Cytochrome P450 3A11 (2e-20, sp|Q64459|CP3AB_MOUSE) | LDA 1 | 2.048597 | 4.47 × 10−6 |
| XLOC_006731 | Cytochrome P450 monooxygenase yanC (9e-49, sp|G3Y416|YANC_ASPNA) | LDA 1 | 1.552875 | 0.000342 |
| XLOC_000034 | Probable mannan endo-1,4-beta-mannosidase F (1e-50, sp|Q5AR04|MANF_EMENI) | hemicellulase | 1.451517 | 0.001082 |
| XLOC_006683 | Cytochrome P450 monooxygenase yanC (2e-28, sp|G3Y416|YANC_ASPNA) | LDA 1 | 1.315805 | 0.002199 |
| XLOC_002405 | Aryl-alcohol dehydrogenase [NADP(+)] (3e-171, sp|Q01752|AAD_PHACH) | LDA 1 | 1.295598 | 0.002627 |
| XLOC_001969 | Cytochrome P450 monooxygenase yanC (3e-104, sp|G3Y416|YANC_ASPNA) | LDA 1 | 0.917329 | 0.031318 |
| XLOC_010055 | Versatile peroxidase VPL1 (9e-155, sp|Q9UR19|VPL1_PLEER) | LME 2 | 0.861786 | 0.037494 |
| XLOC_007717 | Cytochrome P450 monooxygenase yanC (0.0005, sp|G3Y416|YANC_ASPNA) | LDA 1 | 0.828494 | 0.04104 |
| XLOC_004247 | Manganese peroxidase 3 (6e-50, sp|Q96TS6|PEM3_PHLRA) | LME 2 | 0.819839 | 0.014128 |
| XLOC_011551 | Laccase-2 (0.0, sp|Q12718|LAC2_TRAVE) | LME 2 | −1.27902 | 0.003542 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pawlik, A.; Mazur, A.; Wielbo, J.; Koper, P.; Żebracki, K.; Kubik-Komar, A.; Janusz, G. RNA Sequencing Reveals Differential Gene Expression of Cerrena Unicolor in Response to Variable Lighting Conditions. Int. J. Mol. Sci. 2019, 20, 290. https://doi.org/10.3390/ijms20020290
Pawlik A, Mazur A, Wielbo J, Koper P, Żebracki K, Kubik-Komar A, Janusz G. RNA Sequencing Reveals Differential Gene Expression of Cerrena Unicolor in Response to Variable Lighting Conditions. International Journal of Molecular Sciences. 2019; 20(2):290. https://doi.org/10.3390/ijms20020290
Chicago/Turabian StylePawlik, Anna, Andrzej Mazur, Jerzy Wielbo, Piotr Koper, Kamil Żebracki, Agnieszka Kubik-Komar, and Grzegorz Janusz. 2019. "RNA Sequencing Reveals Differential Gene Expression of Cerrena Unicolor in Response to Variable Lighting Conditions" International Journal of Molecular Sciences 20, no. 2: 290. https://doi.org/10.3390/ijms20020290
APA StylePawlik, A., Mazur, A., Wielbo, J., Koper, P., Żebracki, K., Kubik-Komar, A., & Janusz, G. (2019). RNA Sequencing Reveals Differential Gene Expression of Cerrena Unicolor in Response to Variable Lighting Conditions. International Journal of Molecular Sciences, 20(2), 290. https://doi.org/10.3390/ijms20020290







