Apoptotic Fragmentation of Tricellulin
Abstract
1. Introduction
2. Results
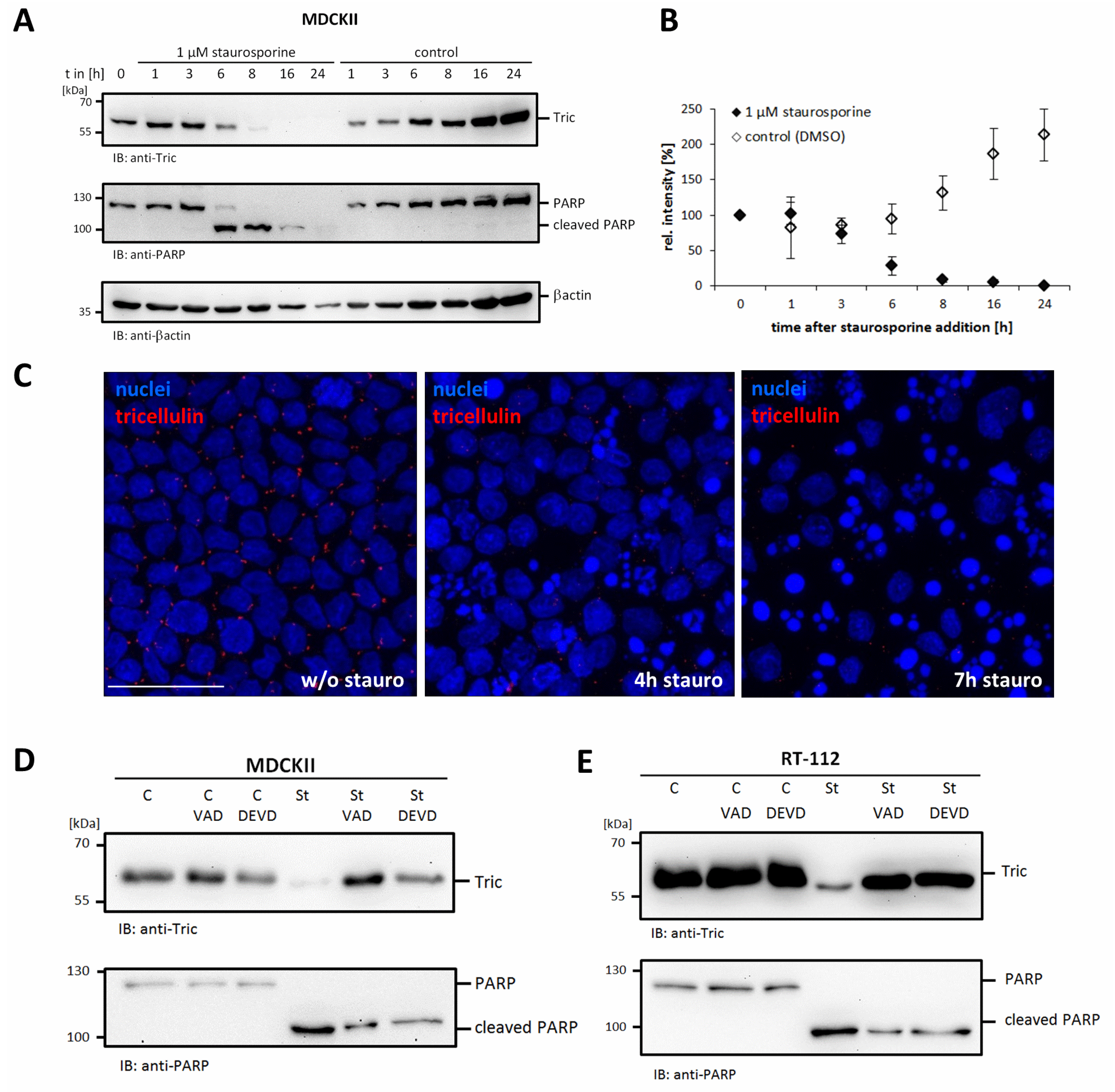
2.1. Caspase-Dependent Cleavage of Tricellulin upon Apoptotic Stimuli
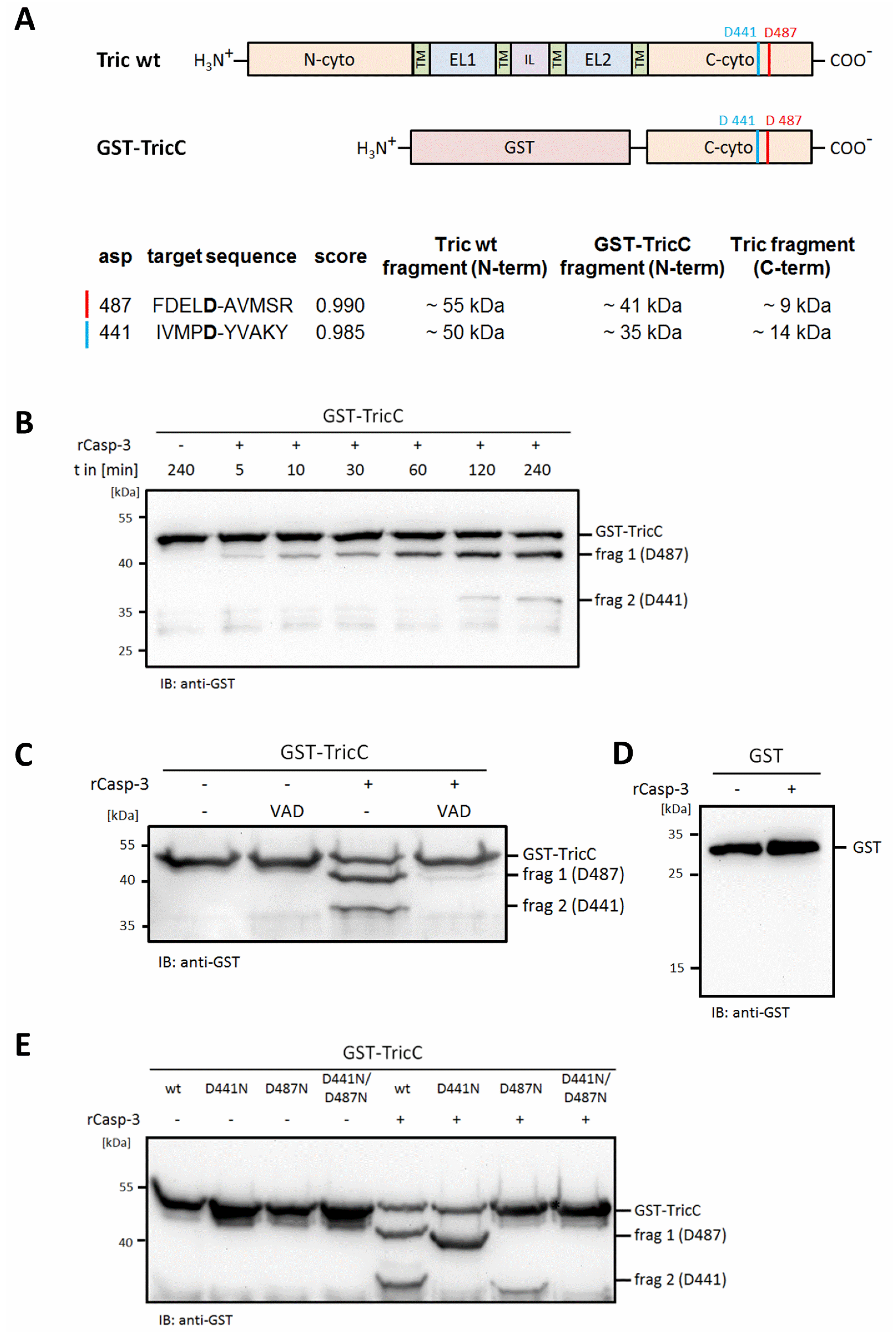
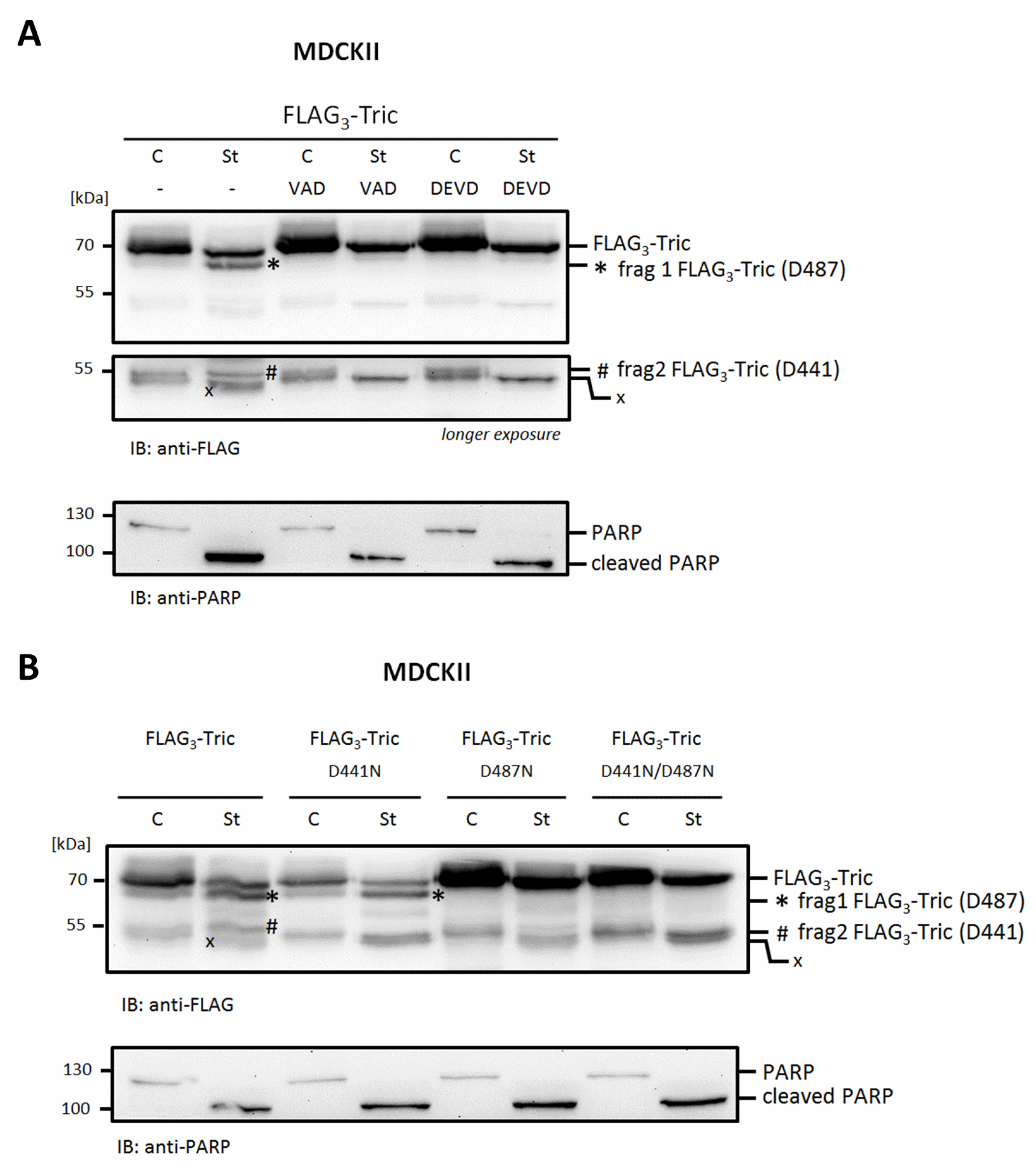
2.2. Mapping of Potential Caspase Cleavage-Sites in Tricellulin
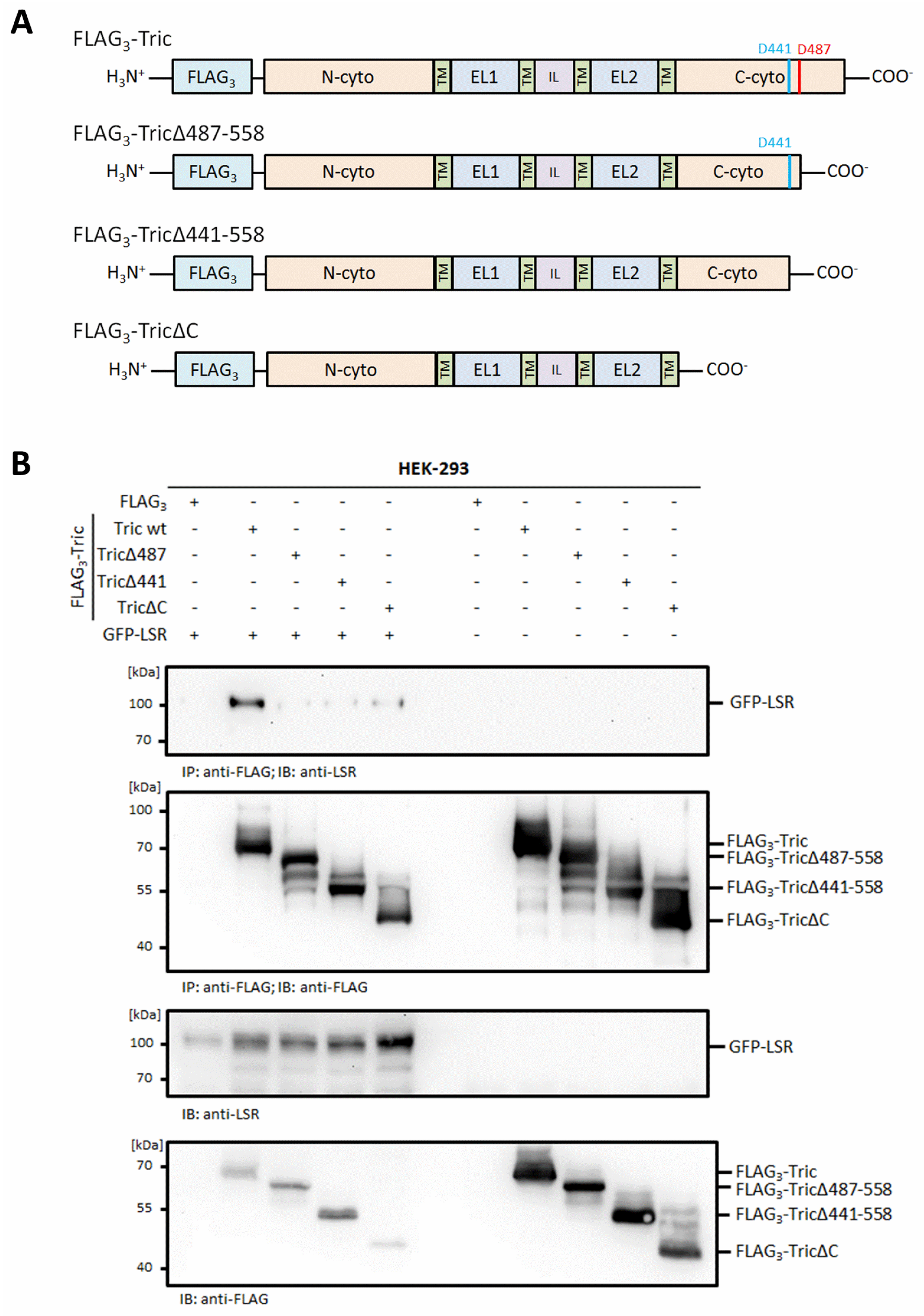
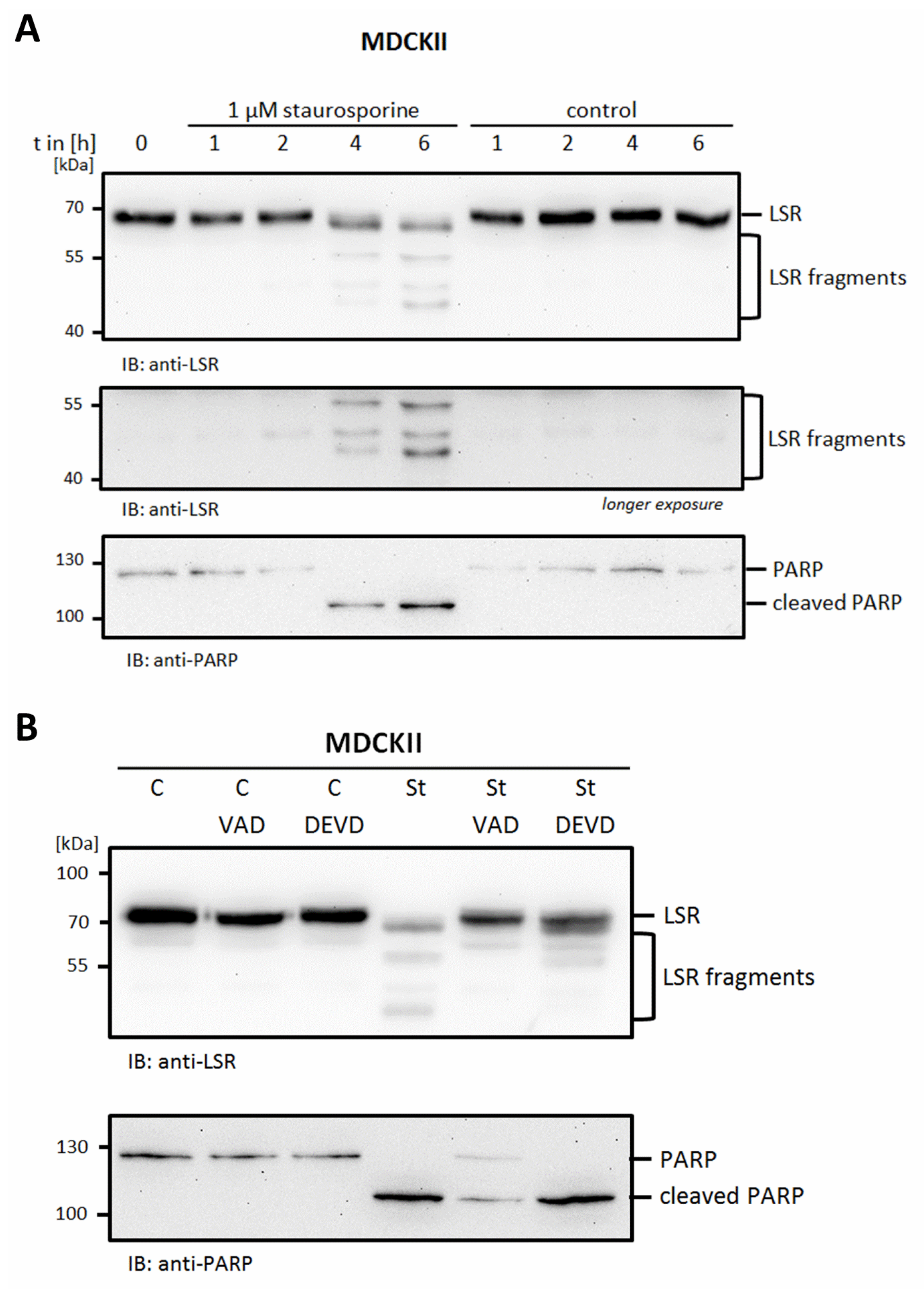
2.3. The Functional Interaction of Tricellulin and LSR Is Disrupted during Apoptosis
3. Discussion
4. Materials and Methods
4.1. Cell Culture
4.2. Reagents and Antibodies
4.3. Plasmids, Site-Directed Mutagenesis and Transient Transfections
4.4. Induction of Apoptosis and Preparation of Cell Lysates
4.5. In Vitro Caspase Cleavage
4.6. Cell Lysis and Co-Immunoprecipitation
4.7. Western Blot Analysis
4.8. Immunofluorescence Microscopy
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| bTJ | Bicellular tight junction |
| CHAPS | 3-[(3-cholamidopropyl)dimethylammonio]-1-propanesulfonate |
| DAPI | 4,6-diamidino-2-phenylindole |
| DMEM | Dulbecco’s Modified Eagle’s Medium |
| DMSO | Dimethyl sulfoxide |
| EDTA | Ethylenediaminetetraacetic acid |
| FBS | Fetal bovine serum |
| GFP | Green-fluorescent protein |
| GST | Glutathione S-transferase |
| HRP | Horseradish peroxidase |
| LSR | Lipolysis-stimulated lipoprotein receptor |
| mab | Monoclonal antibody |
| MDCK | Madin-Darby canine kidney cells |
| pab | Polyclonal antibody |
| PARP | Poly [ADP-ribose] polymerase |
| PEI | Polyethlenimine |
| RIPA | Radioimmunoprecipitation Assay |
| SBOE-PCR | Splicing by overlap extension PCR |
| SDS | Sodium dodecyl sulfate |
| SDS-PAGE | Sodium dodecyl sulfate polyacrylamide gel electrophoresis |
| TBS-T | Tris-buffered saline/Tween 20 |
| TER | Transepithelial resistance |
| TJ | Tight junction |
| tTJ | Tricellular tight junction |
| ZO-1 | Zona occludens-1 |
| Z-DEVD-FMK | N-Benzyloxycarbonyl-Asp-Glu-Val-Asp(OMe) fluoromethylketone |
| Z-VAD-FMK | N-Benzyloxycarbonyl-Val-Ala-Asp(OMe) fluoromethylketone |
References
- Barmeyer, C.; Schulzke, J.D.; Fromm, M. Claudin-related intestinal diseases. Semin. Cell Dev. Biol. 2015, 42, 30–38. [Google Scholar] [CrossRef] [PubMed]
- Chawla, L.S.; Fink, M.; Goldstein, S.L.; Opal, S.; Gomez, A.; Murray, P.; Gomez, H.; Kellum, J.A.; Workgrp, A.X. The epithelium as a target in sepsis. Shock 2016, 45, 249–258. [Google Scholar] [CrossRef] [PubMed]
- Haussner, F.; Chakraborty, S.; Halbgebauer, R.; Huber-Lang, M. Challenge to the intestinal mucosa during sepsis. Front. Immunol. 2019, 10, 891. [Google Scholar] [CrossRef] [PubMed]
- Zihni, C.; Balda, M.S.; Matter, K. Signalling at tight junctions during epithelial differentiation and microbial pathogenesis. J. Cell Sci. 2014, 127, 3401–3413. [Google Scholar] [CrossRef] [PubMed]
- Higashi, T.; Miller, A.L. Tricellular junctions: How to build junctions at the TRICkiest points of epithelial cells. Mol. Biol. Cell 2017, 28, 2023–2034. [Google Scholar] [CrossRef]
- Staehelin, L.A. Further observation on the fine structure of freeze-cleaved tight junctions. J. Cell Sci. 1973, 13, 763–786. [Google Scholar] [PubMed]
- Ikenouchi, J.; Furuse, M.; Furuse, K.; Sasaki, H.; Tsukita, S.; Tsukita, S. Tricellulin constitutes a novel barrier at tricellular contacts of epithelial cells. J. Cell Biol. 2005, 171, 939–945. [Google Scholar] [CrossRef]
- Higashi, T.; Tokuda, S.; Kitajiri, S.; Masuda, S.; Nakamura, H.; Oda, Y.; Furuse, M. Analysis of the ‘angulin’ proteins LSR, ILDR1 and ILDR2-tricellulin recruitment, epithelial barrier function and implication in deafness pathogenesis. J. Cell Sci. 2013, 126, 966–977. [Google Scholar] [CrossRef]
- Furuse, M.; Itoh, M.; Hirase, T.; Nagafuchi, A.; Yonemura, S.; Tsukita, S.; Tsukita, S. Direct association of occludin with ZO-1 and its possible involvement in the localization of occludin at tight junctions. J. Cell Biol. 1994, 127, 1617–1626. [Google Scholar] [CrossRef]
- Schuetz, A.; Radusheva, V.; Krug, S.M.; Heinemann, U. Crystal structure of the tricellulin C-terminal coiled-coil domain reveals a unique mode of dimerization. Ann. N. Y. Acad. Sci. 2017, 1405, 147–159. [Google Scholar] [CrossRef]
- Raleigh, D.R.; Marchiando, A.M.; Zhang, Y.; Shen, L.; Sasaki, H.; Wang, Y.; Long, M.; Turner, J.R. Tight junction-associated MARVEL proteins marveld3, tricellulin, and occludin have distinct but overlapping functions. Mol. Biol. Cell 2010, 21, 1200–1213. [Google Scholar] [CrossRef] [PubMed]
- Krug, S.M.; Amasheh, M.; Dittmann, I.; Christoffel, I.; Fromm, M.; Amasheh, S. Sodium caprate as an enhancer of macromolecule permeation across tricellular tight junctions of intestinal cells. Biomaterials 2013, 34, 275–282. [Google Scholar] [CrossRef] [PubMed]
- Krug, S.M.; Amasheh, S.; Richter, J.F.; Milatz, S.; Günzel, D.; Westphal, J.K.; Huber, O.; Schulzke, J.D.; Fromm, M. Tricellulin forms a barrier to macromolecules in tricellular tight junctions without affecting ion permeability. Mol. Biol. Cell 2009, 20, 3713–3724. [Google Scholar] [CrossRef] [PubMed]
- Ikenouchi, J.; Sasaki, H.; Tsukita, S.; Furuse, M.; Tsukita, S. Loss of occludin affects tricellular localization of tricellulin. Mol. Biol. Cell 2008, 19, 4687–4693. [Google Scholar] [CrossRef] [PubMed]
- Westphal, J.K.; Dörfel, M.J.; Krug, S.M.; Cording, J.D.; Piontek, J.; Blasig, I.E.; Tauber, R.; Fromm, M.; Huber, O. Tricellulin forms homomeric and heteromeric tight junctional complexes. Cell. Mol. Life Sci. 2010, 67, 2057–2068. [Google Scholar] [CrossRef] [PubMed]
- Cording, J.; Berg, J.; Kading, N.; Bellmann, C.; Tscheik, C.; Westphal, J.K.; Milatz, S.; Günzel, D.; Wolburg, H.; Piontek, J.; et al. In tight junctions, claudins regulate the interactions between occludin, tricellulin and marvelD3, which, inversely, modulate claudin oligomerization. J. Cell Sci. 2013, 126, 554–564. [Google Scholar] [CrossRef]
- Masuda, S.; Oda, Y.; Sasaki, H.; Ikenouchi, J.; Higashi, T.; Akashi, M.; Nishi, E.; Furuse, M. LSR defines cell corners for tricellular tight junction formation in epithelial cells. J. Cell Sci. 2011, 124, 548–555. [Google Scholar] [CrossRef]
- Kohno, T.; Konno, T.; Kojima, T. Role of tricellular tight junction protein lipolysis-stimulated lipoprotein receptor (LSR) in cancer cells. Int. J. Mol. Sci. 2019, 20, 3555. [Google Scholar] [CrossRef]
- Higashi, T.; Katsuno, T.; Kitajiri, S.; Furuse, M. Deficiency of angulin-2/ILDR1, a tricellular tight junction-associated membrane protein, causes deafness with cochlear hair cell degeneration in mice. PLoS ONE 2015, 10, e0120674. [Google Scholar] [CrossRef]
- Riazuddin, S.; Ahmed, Z.M.; Fanning, A.S.; Lagziel, A.; Kitajiri, S.; Ramzan, K.; Khan, S.N.; Chattaraj, P.; Friedman, P.L.; Anderson, J.M.; et al. Tricellulin is a tight-junction protein necessary for hearing. Am. J. Hum. Genet. 2006, 79, 1040–1051. [Google Scholar] [CrossRef]
- Ramzan, K.; Shaikh, R.S.; Ahmad, J.; Khan, S.N.; Riazuddin, S.; Ahmed, Z.M.; Friedman, T.B.; Wilcox, E.R.; Riazuddin, S. A new locus for nonsyndromic deafness DFNB49 maps to chromosome 5q12.3-q14.1. Hum. Genet. 2005, 116, 407–412. [Google Scholar] [CrossRef] [PubMed]
- Nayak, G.; Lee, S.I.; Yousaf, R.; Edelmann, S.E.; Trincot, C.; Van Itallie, C.M.; Sinha, G.P.; Rafeeq, M.; Jones, S.M.; Belyantseva, I.A.; et al. Tricellulin deficiency affects tight junction architecture and cochlear hair cells. J. Clin. Investig. 2013, 123, 4036–4049. [Google Scholar] [CrossRef] [PubMed]
- Kamitani, T.; Sakaguchi, H.; Tamura, A.; Miyashita, T.; Yamazaki, Y.; Tokumasu, R.; Inamoto, R.; Matsubara, A.; Mori, N.; Hisa, Y.; et al. Deletion of tricellulin causes progressive hearing loss associated with degeneration of cochlear hair cells. Sci. Rep. 2015, 5, 18402. [Google Scholar] [CrossRef] [PubMed]
- Morozko, E.L.; Nishio, A.; Ingham, N.J.; Chandra, R.; Fitzgerald, T.; Martelletti, E.; Borck, G.; Wilson, E.; Riordan, G.P.; Wangemann, P.; et al. ILDR1 null mice, a model of human deafness DFNB42, show structural aberrations of tricellular tight junctions and degeneration of auditory hair cells. Hum. Mol. Genet. 2015, 24, 609–624. [Google Scholar] [CrossRef] [PubMed]
- Sang, Q.; Li, W.; Xu, Y.; Qu, R.G.; Xu, Z.G.; Feng, R.Z.; Jin, L.; He, L.; Li, H.W.; Wang, L. ILDR1 deficiency causes degeneration of cochlear outer hair cells and disrupts the structure of the organ of Corti: A mouse model for human DFNB42. Biol. Open 2015, 4, 411–418. [Google Scholar] [CrossRef][Green Version]
- Dörfel, M.J.; Huber, O. Modulation of tight junction structure and function by kinases and phosphatases targeting occludin. J. Biomed. Biotechnol. 2012, 807356. [Google Scholar] [CrossRef] [PubMed]
- Dörfel, M.J.; Huber, O. A phosphorylation hotspot within the occludin C-terminal domain. Ann. N. Y. Acad. Sci. 2012, 1257, 38–44. [Google Scholar] [CrossRef]
- Manda, B.; Mir, H.; Gangwar, R.; Meena, A.S.; Amin, S.; Shukla, P.K.; Dalal, K.; Suzuki, T.; Rao, R. Phosphorylation hotspot in the C-terminal domain of occludin regulates the dynamics of epithelial junctional complexes. J. Cell Sci. 2018, 131. [Google Scholar] [CrossRef]
- Dörfel, M.J.; Westphal, J.K.; Huber, O. Differential phosphorylation of occludin and tricellulin by CK2 and CK1. Ann. N. Y. Acad. Sci. 2009, 1165, 69–73. [Google Scholar] [CrossRef]
- Nakatsu, D.; Kano, F.; Taguchi, Y.; Sugawara, T.; Nishizono, T.; Nishikawa, K.; Oda, Y.; Furuse, M.; Murata, M. JNK1/2-dependent phosphorylation of angulin-1/LSR is required for the exclusive localization of angulin-1/LSR and tricellulin at tricellular contacts in EpH4 epithelial sheet. Genes Cells 2014, 19, 565–581. [Google Scholar] [CrossRef]
- Jennek, S.; Mittag, S.; Reiche, J.; Westphal, J.K.; Seelk, S.; Dorfel, M.J.; Pfirrmann, T.; Friedrich, K.; Schutz, A.; Heinemann, U.; et al. Tricellulin is a target of the ubiquitin ligase Itch. Ann. N. Y. Acad. Sci. 2017, 1397, 157–168. [Google Scholar] [CrossRef] [PubMed]
- Ramirez, M.L.G.; Salvesen, G.S. A primer on caspase mechanisms. Semin. Cell Dev. Biol. 2018, 82, 79–85. [Google Scholar] [CrossRef] [PubMed]
- Songane, M.; Khair, M.; Saleh, M. An updated view on the functions of caspases in inflammation and immunity. Semin. Cell Dev. Biol. 2018, 82, 137–149. [Google Scholar] [CrossRef] [PubMed]
- Gagliardi, P.A.; Primo, L. Death for life: A path from apoptotic signaling to tissue-scale effects of apoptotic epithelial extrusion. Cell. Mol. Life Sci. 2019, 76, 3571–3581. [Google Scholar] [CrossRef] [PubMed]
- Bojarski, C.; Weiske, J.; Schöneberg, T.; Schröder, W.; Mankertz, J.; Schulzke, J.D.; Florian, P.; Fromm, M.; Tauber, R.; Huber, O. The specific fates of tight junction proteins in apoptotic epithelial cells. J. Cell Sci. 2004, 117, 2097–2107. [Google Scholar] [CrossRef] [PubMed]
- Steinhusen, U.; Weiske, J.; Badock, V.; Tauber, R.; Bommert, K.; Huber, O. Cleavage and shedding of E-cadherin after induction of apoptosis. J. Biol. Chem. 2001, 276, 4972–4980. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; van Raam, B.J.; Salvesen, G.S.; Cieplak, P. Caspase cleavage sites in the human proteome: CaspDB, a database of predicted substrates. PLoS ONE 2014, 9, e110539. [Google Scholar] [CrossRef] [PubMed]
- Iwamoto, N.; Higashi, T.; Furuse, M. Localization of angulin-1/LSR and tricellulin at tricellular contacts of brain and retinal endothelial cells in vivo. Cell. Struct. Funct. 2014, 39, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Sohet, F.; Lin, C.; Munji, R.N.; Lee, S.Y.; Ruderisch, N.; Soung, A.; Arnold, T.D.; Derugin, N.; Vexler, Z.S.; Yen, F.T.; et al. LSR/angulin-1 is a tricellular tight junction protein involved in blood-brain barrier formation. J. Cell Biol. 2015, 208, 703–711. [Google Scholar] [CrossRef] [PubMed]
- Bojarski, C.; Gitter, A.H.; Bendfeldt, K.; Mankertz, J.; Schmitz, H.; Wagner, S.; Fromm, M.; Schulzke, J.D. Permeability of human HT-29/B6 colonic epithlium as a function of apoptosis. J. Physiol. 2001, 535, 541–552. [Google Scholar] [CrossRef] [PubMed]
- Herren, B.; Levkau, B.; Raines, E.W.; Ross, R. Cleavage of β-catenin and plakoglobin and shedding of VE-cadherin during endothelial apoptosis: Evidence for a role for caspases and metalloproteinases. Mol. Biol. Cell 1998, 9, 1589–1601. [Google Scholar] [CrossRef] [PubMed]
- Steinhusen, U.; Badock, V.; Bauer, A.; Behrens, J.; Wittmann-Liebold, B.; Dörken, B.; Bommert, K. Apoptosis induced cleavage of β-catenin by caspase-3 results in proteolytic fragments with reduced transactivation potential. J. Biol. Chem. 2000, 275, 16345–16353. [Google Scholar] [CrossRef] [PubMed]
- Weiske, J.; Schöneberg, T.; Schröder, W.; Hatzfeld, M.; Tauber, R.; Huber, O. The fate of desmosomal proteins in apoptotic cells. J. Biol. Chem. 2001, 276, 41175–41181. [Google Scholar] [CrossRef] [PubMed]
- Multhaup, G.; Huber, O.; Buee, L.; Galas, M.C. Amyloid Precursor Protein (APP) Metabolites APP intracellular fragment (AICD), Aβ42, and Tau in nuclear roles. J. Biol. Chem. 2015, 290, 23515–23522. [Google Scholar] [CrossRef] [PubMed]
- Bray, S.J.; Gomez-Lamarca, M. Notch after cleavage. Curr. Opin. Cell Biol. 2018, 51, 103–109. [Google Scholar] [CrossRef]
- Takasawa, A.; Murata, M.; Takasawa, K.; Ono, Y.; Osanai, M.; Tanaka, S.; Nojima, M.; Kono, T.; Hirata, K.; Kojima, T.; et al. Nuclear localization of tricellulin promotes the oncogenic property of pancreatic cancer. Sci. Rep. 2016, 6, 33582. [Google Scholar] [CrossRef]
- Eum, S.Y.; Jaraki, D.; Bertrand, L.; Andras, I.E.; Toborek, M. Disruption of epithelial barrier by quorum-sensing N-3-(oxododecanoyl)-homoserine lactone is mediated by matrix metalloproteinases. Am. J. Physiol. Gastrointest. Liver Physiol. 2014, 306, G992–G1001. [Google Scholar] [CrossRef] [PubMed]
- Wetzel, F.; Mittag, S.; Cano-Cortina, M.; Wagner, T.; Kramer, O.H.; Niedenthal, R.; Gonzalez-Mariscal, L.; Huber, O. SUMOylation regulates the intracellular fate of ZO-2. Cell. Mol. Life Sci. 2017, 74, 373–392. [Google Scholar] [CrossRef]
- Mittag, S.; Valenta, T.; Weiske, J.; Bloch, L.; Klingel, S.; Gradl, D.; Wetzel, F.; Chen, Y.; Petersen, I.; Basler, K.; et al. A novel role for the tumour suppressor nitrilase1 modulating the Wnt/β-catenin signalling pathway. Cell Discov. 2016, 2. [Google Scholar] [CrossRef]

| Antibody | Dilution | Target | Species |
|---|---|---|---|
| anti-FLAG® M2 | WB (1:5.000); IP 2 μg | FLAG-tag | mouse mab |
| anti-GST | WB (1:20.000) | glutathione-S-transferase | rabbit pab |
| anti-LSR (D3E3N) XP® | WB (1:1.000) | LSR | rabbit mab |
| anti-PARP-1 (Ab-2) | WB (1:1.000) | PARP | mouse mab |
| anti-tricellulin (54H19L38) ABfinity™ | WB (1:1.000) IF (1 μg/mL) | tricellulin | rabbit mab |
| anti-βactin | WB (1:1000) | βactin | mouse |
| goat anti-mouse-HRP | WB (1:50.000–1:100.000) | mouse IgG | goat |
| goat anti-rabbit-HRP | WB (1:50.000–1:100.000) | rabbit IgG | goat |
| donkey anti-rabbit-Alexa594 | IF (1: 1.000) | rabbit IgG | donkey |
| Mutation | Product | Sequence Oligonucleotides |
|---|---|---|
| 1 | 5′-GCG GGT ACC GGA TCC GCC GCC ATG TCA AAT GAT GGA AGA TCC-3′ 5′-CAC ATA GTT GGG CAT CAC GAT-3′ | |
| Tric-D441N | 2 | 5′-ATC GTG ATG CCC AAC TAT GTG-3′ 5′-GCG GGT ACC GGA TCC TTA AGA ATA ACC TTG TAC ATC-3′ |
| 1 + 2 (SBOE) | 5′-GCG GGT ACC GGA TCC GCC GCC ATG TCA AAT GAT GGA AGA TCC-3′ 5′-GCG GGT ACC GGA TCC TTA AGA ATA ACC TTG TAC ATC-3′ | |
| 1 | 5′-GCG GGT ACC GGA TCC GCC GCC ATG TCA AAT GAT GGA AGA TCC-3′ 5′-CAC TGC ATT CAG CTC ATC AAA-3′ | |
| Tric-D487N | 2 | 5′-TTT GAT GAG CTG AAT GCA GTG-3′ 5′-GCG GGT ACC GGA TCC TTA AGA ATA ACC TTG TAC ATC-3′ |
| 1 + 2 (SBOE) | 5′-GCG GGT ACC GGA TCC GCC GCC ATG TCA AAT GAT GGA AGA TCC-3′ 5′-GCG GGT ACC GGA TCC TTA AGA ATA ACC TTG TAC ATC-3′ |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Janke, S.; Mittag, S.; Reiche, J.; Huber, O. Apoptotic Fragmentation of Tricellulin. Int. J. Mol. Sci. 2019, 20, 4882. https://doi.org/10.3390/ijms20194882
Janke S, Mittag S, Reiche J, Huber O. Apoptotic Fragmentation of Tricellulin. International Journal of Molecular Sciences. 2019; 20(19):4882. https://doi.org/10.3390/ijms20194882
Chicago/Turabian StyleJanke, Susanne, Sonnhild Mittag, Juliane Reiche, and Otmar Huber. 2019. "Apoptotic Fragmentation of Tricellulin" International Journal of Molecular Sciences 20, no. 19: 4882. https://doi.org/10.3390/ijms20194882
APA StyleJanke, S., Mittag, S., Reiche, J., & Huber, O. (2019). Apoptotic Fragmentation of Tricellulin. International Journal of Molecular Sciences, 20(19), 4882. https://doi.org/10.3390/ijms20194882






