Heat-Responsive Proteomics of a Heat-Sensitive Spinach Variety
Abstract
1. Introduction
2. Results
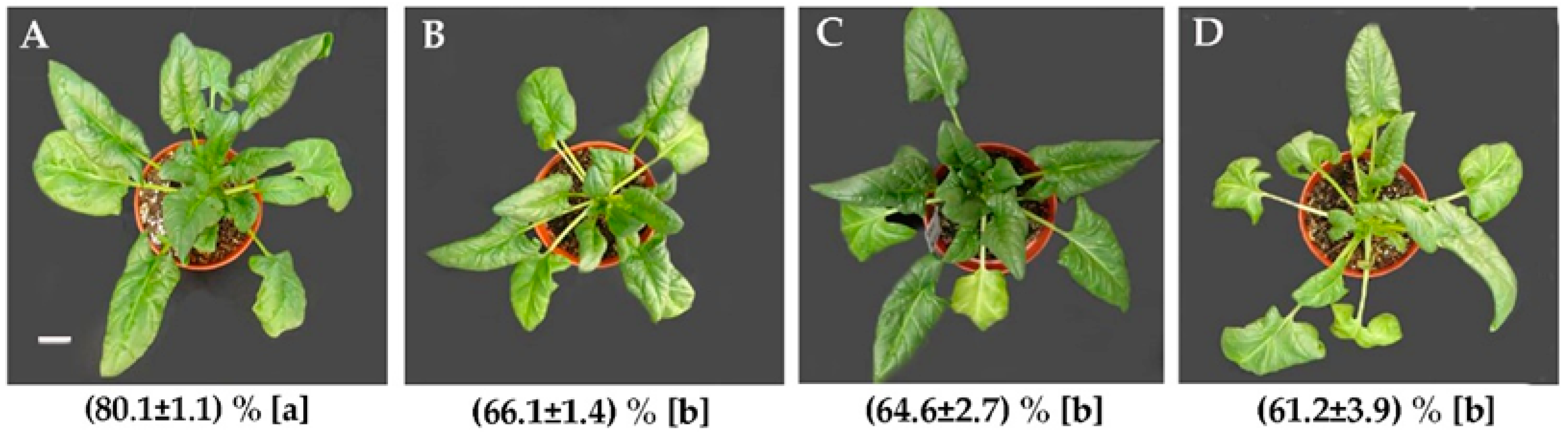
2.1. Morphology and Relative Water Content (RWC)
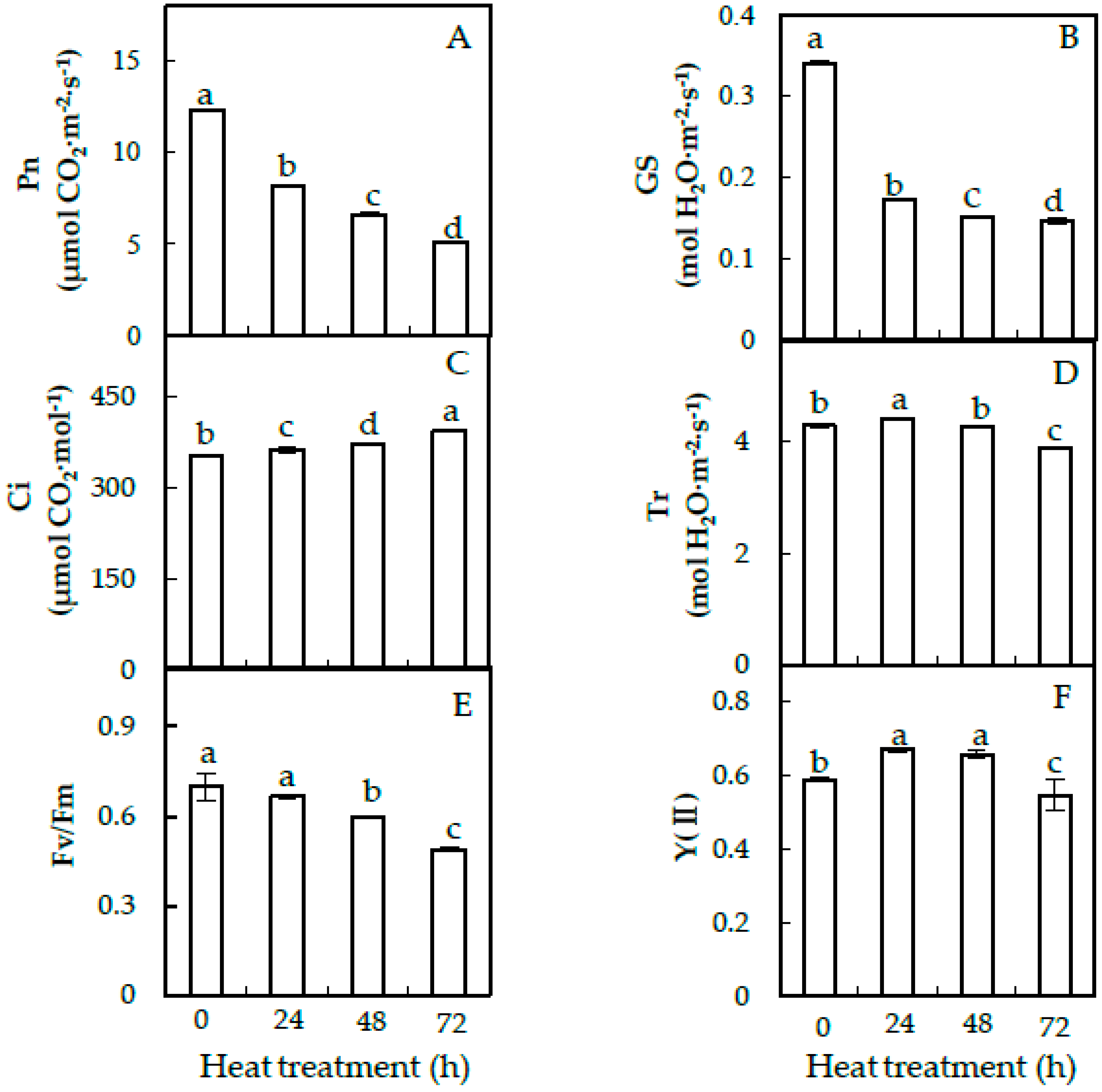
2.2. Photosynthesis and Chlorophyll Fluorescence Parameters
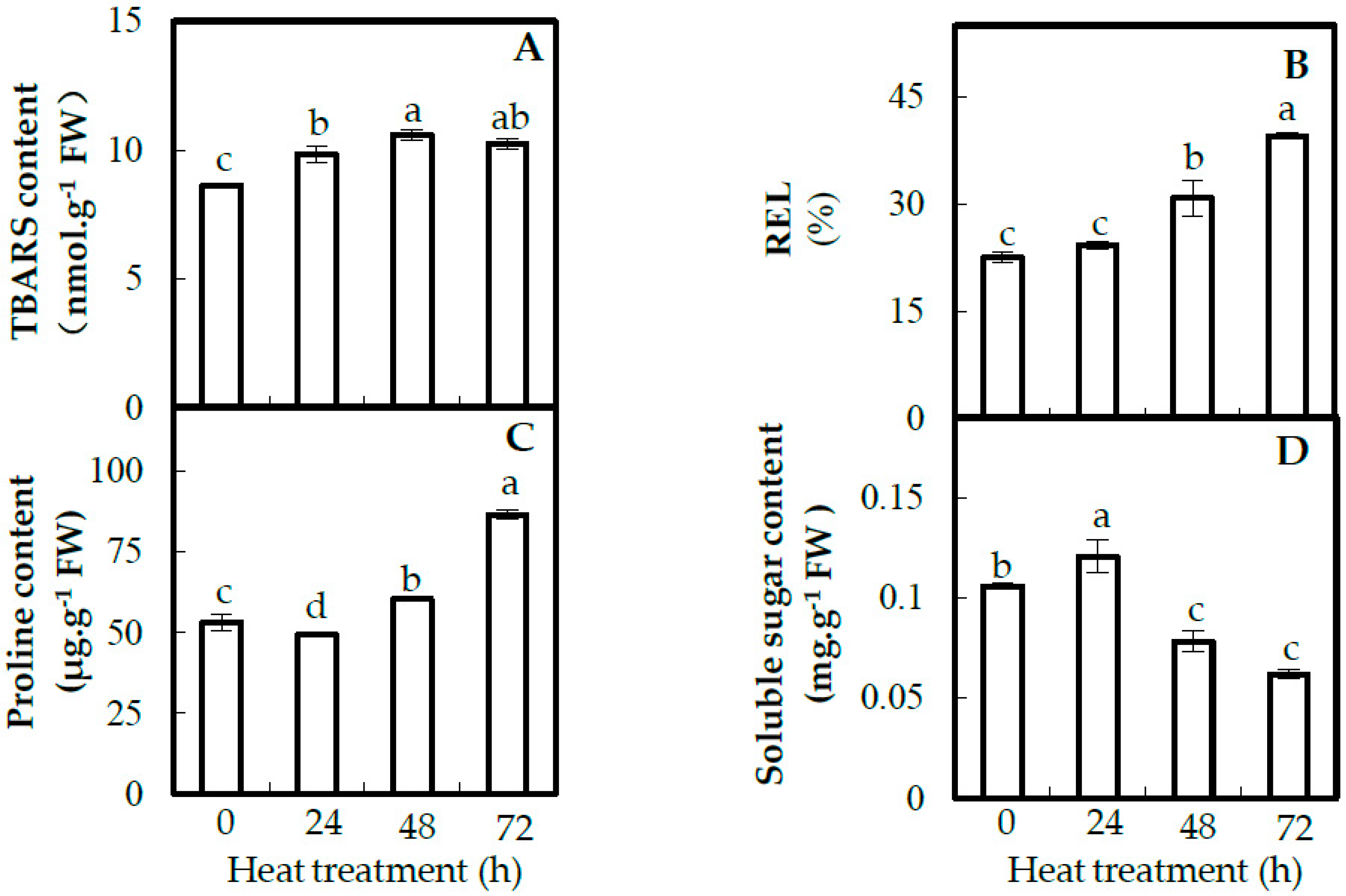
2.3. Plasma Membrane Integrity and Osmolyte Accumulation in Leaves
2.4. Activities of Antioxidant Enzymes and Antioxidant Contents in Leaves
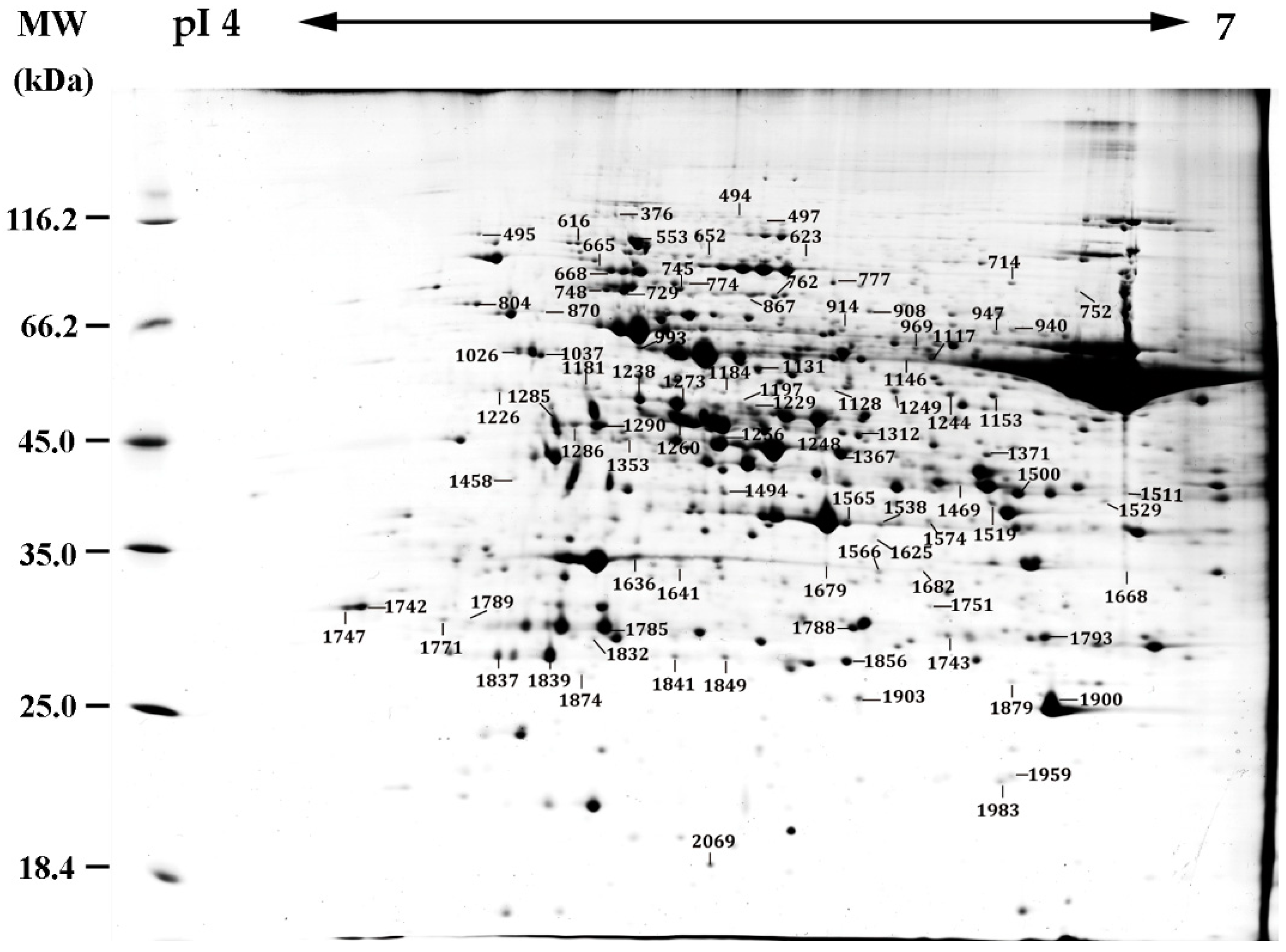
2.5. Identification of Heat-Responsive Proteins by 2DE-Based and iTRAQ-Based Proteomics
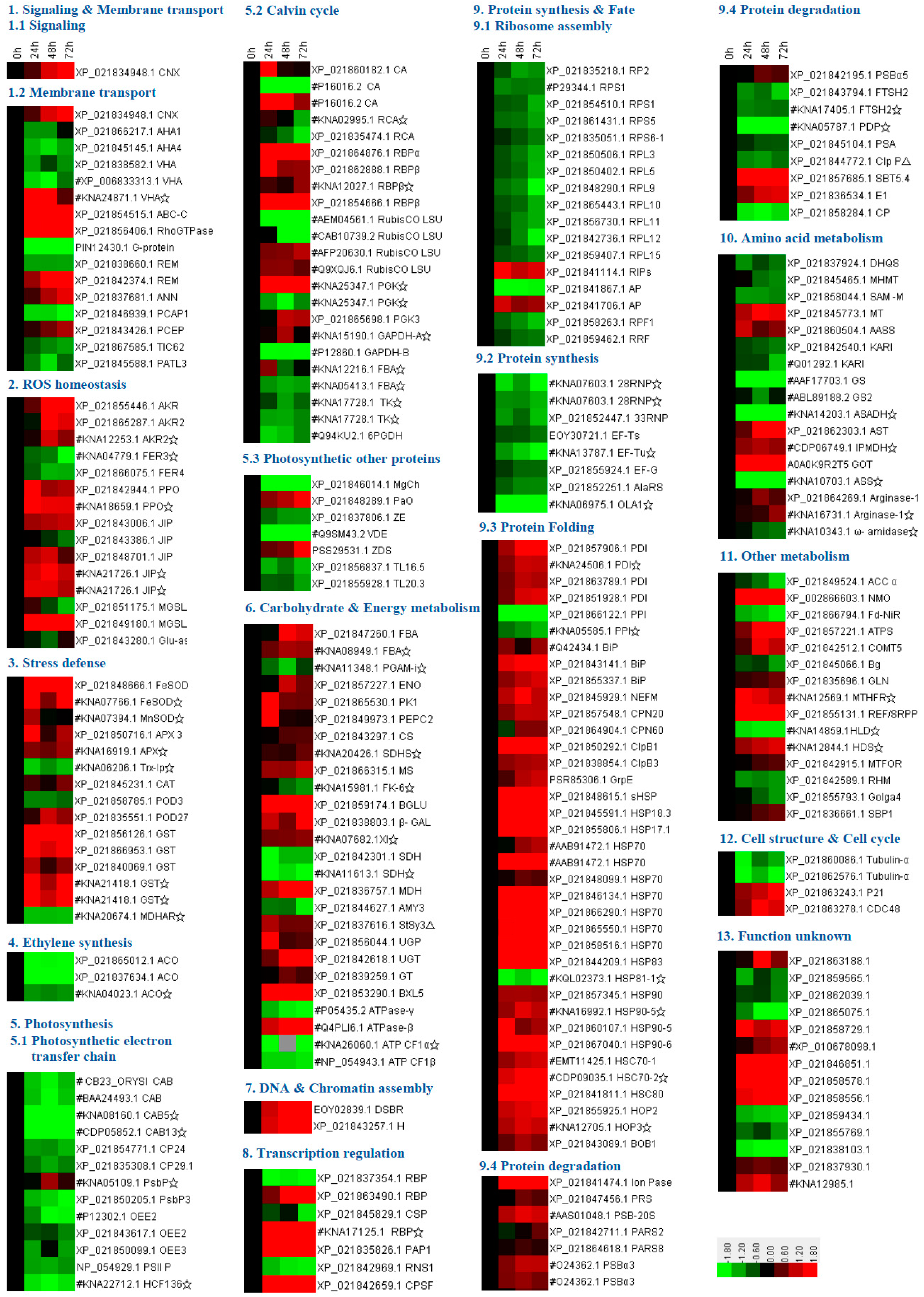
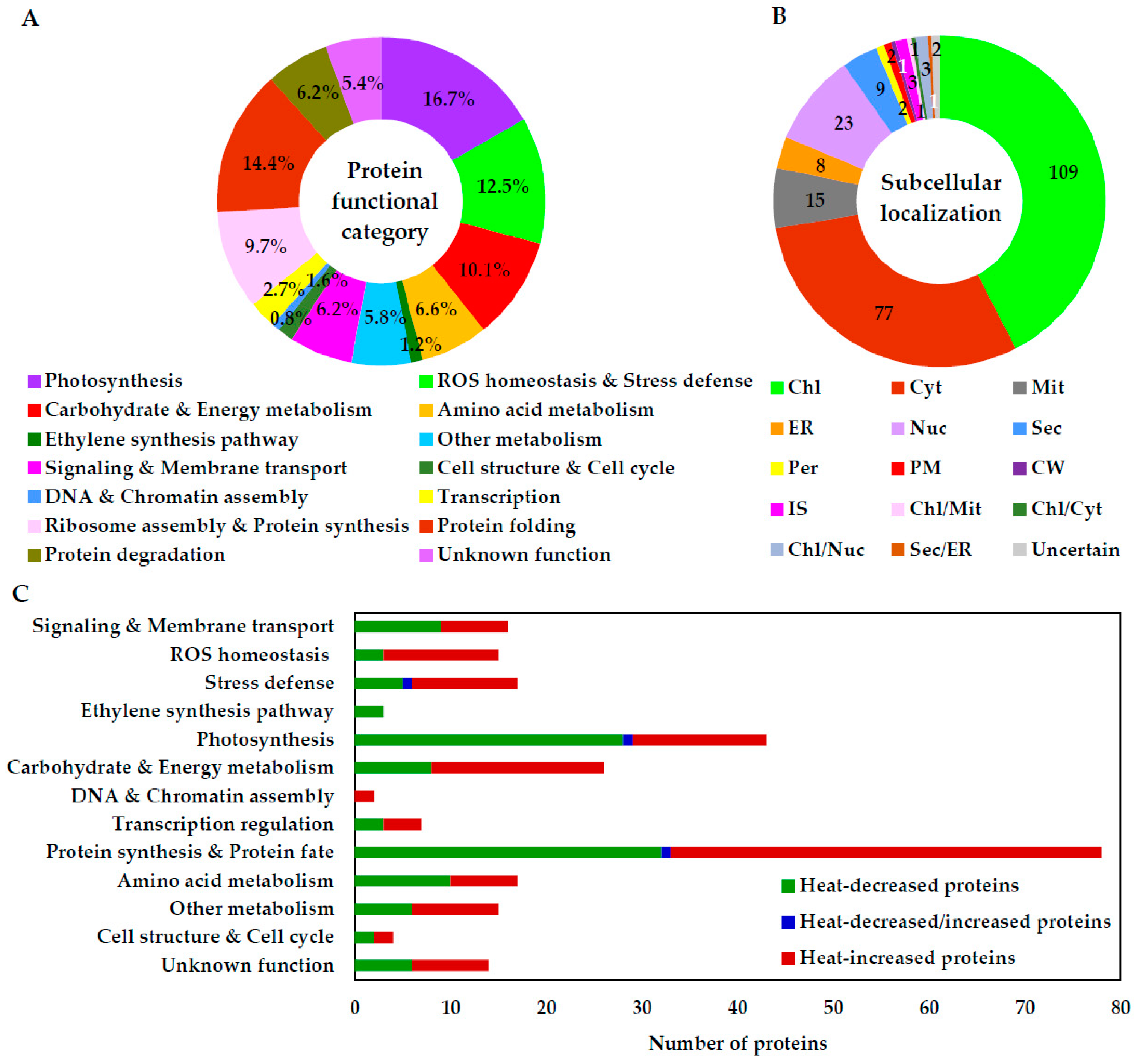
2.6. Annotation and Functional Categorization of HRPs
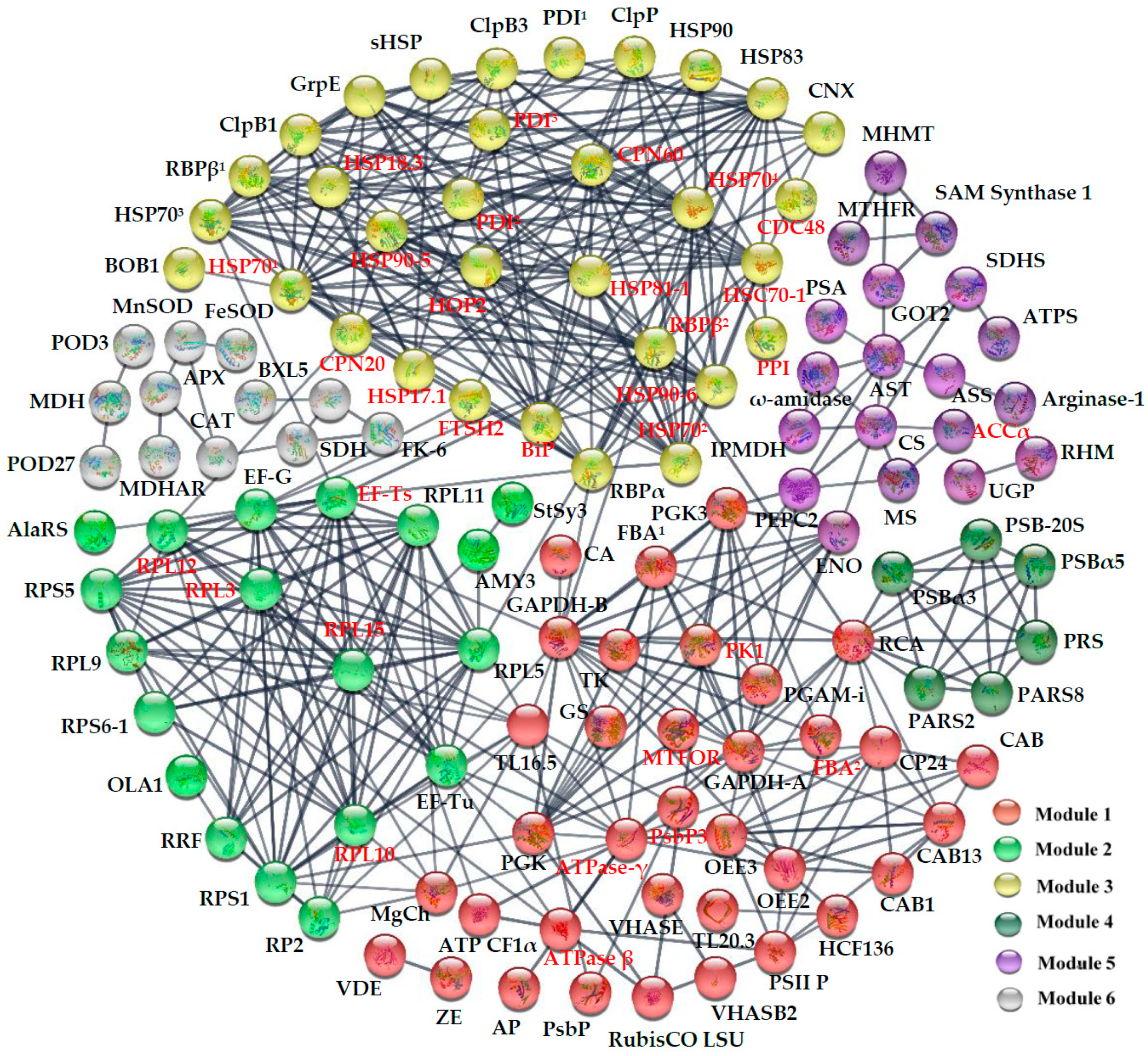
2.7. Subcellular Localization and Protein–Protein Interaction (PPI) Network of HRPs
3. Discussion
3.1. Heat-Inhibited Photosynthesis in Heat-Sensitive Spinach Variety Sp73
3.2. Heat-Altered ROS Scavenging Pathways in Spinach Sp73
3.3. Heat-Stress Signaling and Transport Pathways in Spinach Sp73
3.4. Heat Stress-Perturbed Diverse Primary and Secondary Metabolisms
3.5. Heat Stress-Induced Transcriptional Regulation and Protein Processing in Sp73
4. Materials and Methods
4.1. Plant Cultivation and Treatment
4.2. RWC Measurement
4.3. Photosynthesis and Chlorophyll Fuorescence Parameter Measurement
4.4. Determination of TBARS Content, REL, Total Soluble Sugar, and Proline Contents
4.5. Determination of ROS and Antioxidant Substance Contents, and Antioxidant Enzyme Activity Assay
4.6. Quantitative Proteomics Analyses
4.7. Protein Function Classification and Cluster Analysis
4.8. Protein Subcellular Localization Prediction
4.9. Protein-Protein Interaction Prediction
4.10. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| 2DE | Two-dimensional gel electrophoresis |
| AKR | Aldo/keto reductase |
| ANN | Annexin |
| APX | Ascorbate peroxidase |
| AsA | Ascorbate |
| ATP | Adenosine-triphosphate |
| BLAST | Basic Local Alignment Search Tool |
| CAT | Catalase |
| Ci | Intercellular CO2 |
| DHA | Dehydroascorbate |
| DHAR | Dehydroascorbate reductase |
| Fv/Fm | PSII maximum quantum yield |
| FW | Fresh weight |
| GPX | Glutathione peroxidase |
| GR | Glutathione reductase |
| Gs | Stomata conductance |
| GSH | Reduced glutathione |
| GSSG | Oxidized glutathione |
| GST | Glutathione S-transferase |
| H2O2 | Hydrogen peroxide |
| HHT | Hour of heat treatment |
| HRP | Heat stress-responsive protein |
| IEF | Isoelectric focusing |
| IPG | Immobilized pH gradient |
| iTRAQ | Isobaric tags for relative and absolute quantification |
| MALDI | Matrix-assisted laser desorption/ ionization |
| MAPK | Mitogen-activated protein kinase |
| MDHAR | Monodehydroascorbate reductase |
| MS | Mass spectrometry |
| NCBInr | National Center for Biotechnology Information non-redundant |
| O2•− | Superoxide anion radicals |
| OEC | Oxygen evolving complex |
| pI | Isoelectric point |
| Pn | Photosynthesis rate |
| POD | Peroxidase |
| PPI | Protein-protein interactions |
| PSII | Photosystem II |
| RBP | RNA-binding protein family member |
| RCA | Ribulose bisphosphate carboxylase/oxygenase activase |
| REL | Relative electrolyte leakage |
| ROS | Reactive oxygen species |
| RubisCO | Ribulose-1,5-bisphosphate carboxylase/oxygenase |
| RWC | Relative water content |
| SDS-PAGE | Sodium dodecyl sulfate polyacrylamide gel electrophoresis |
| SOD | Superoxide dismutase |
| Sp73 | Heat-sensitive spinach |
| Sp75 | Heat-tolerant spinach |
| STRING | Search tool for recurring instances of neighbouring genes |
| TBARS | Thiobarbituric acid reactive substance |
| TOF | Time of flight |
| Tr | Transpiration rate |
| Y(II) | Effective PSII quantum yield |
References
- Fahad, S.; Bajwa, A.A.; Nazir, U.; Anjum, S.A.; Farooq, A.; Zohaib, A.; Sadia, S.; Nasim, W.; Adkins, S.; Saud, S.; et al. Crop production under drought and heat stress: Plant responses and management options. Front Plant Sci. 2017, 8, 1147. [Google Scholar] [CrossRef] [PubMed]
- Lamaoui, M.; Jemo, M.; Datla, R.; Bekkaoui, F. Heat and drought stresses in crops and approaches for their mitigation. Front. Chem. 2018, 6, 26. [Google Scholar] [CrossRef] [PubMed]
- Bita, C.E.; Zenoni, S.; Vriezen, W.H.; Mariani, C.; Pezzotti, M.; Gerats, T. Temperature stress differentially modulates transcription in meiotic anthers of heat-tolerant and heat-sensitive tomato plants. BMC Genom. 2011, 12, 384. [Google Scholar] [CrossRef] [PubMed]
- Mittler, R.; Finka, A.; Goloubinoff, P. How do plants feel the heat? Trends Biochem. Sci. 2012, 37, 118–125. [Google Scholar] [CrossRef] [PubMed]
- Bokszczanin, K.L.; Fragkostefanakis, S. Perspectives on deciphering mechanisms underlying plant heat stress response and thermotolerance. Front Plant Sci. 2013, 4, 315. [Google Scholar] [CrossRef] [PubMed]
- Hasanuzzaman, M.; Nahar, K.; Alam, M.M.; Roychowdhury, R.; Fujita, M. Physiological, biochemical, and molecular mechanisms of heat stress tolerance in plants. Int. J. Mol. Sci. 2013, 14, 9643–9684. [Google Scholar] [CrossRef] [PubMed]
- McClung, C.R.; Davis, S.J. Ambient thermometers in plants: From physiological outputs towards mechanisms of thermal sensing. Curr. Biol. 2010, 20, R1086–R1092. [Google Scholar] [CrossRef] [PubMed]
- Mathur, S.; Agrawal, D.; Jajoo, A. Photosynthesis: Response to high temperature stress. J. Photochem. Photobiol. B. 2014, 137, 116–126. [Google Scholar] [CrossRef]
- Wang, X.; Xu, C.; Cai, X.; Wang, Q.; Dai, S. Heat-responsive photosynthetic and signaling pathways in plants: Insight from proteomics. Int. J. Mol. Sci. 2017, 18, 2191. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Wei, Z.; Qiao, Z.; Wu, Z.; Cheng, L.; Wang, Y. Proteomics analysis of alfalfa response to heat stress. PLoS ONE 2013, 8, e82725. [Google Scholar] [CrossRef]
- Zhao, Q.; Chen, W.; Bian, J.; Xie, H.; Li, Y.; Xu, C.; Ma, J.; Guo, S.; Chen, J.; Cai, X.; et al. Proteomics and phosphoproteomics of heat stress-responsive mechanisms in Spinach. Front Plant Sci. 2018, 9, 800. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Zhu, G.; Guo, Q.; Zhu, Z.; Wang, C.; Liu, Z. A comparative proteomic analysis of Pinellia ternata leaves exposed to heat stress. Int. J. Mol. Sci. 2013, 10, 20614–20634. [Google Scholar] [CrossRef] [PubMed]
- Rocco, M.; Arena, S.; Renzone, G.; Scippa, G.S.; Lomaglio, T.; Verrillo, F.; Scaloni, A.; Marra, M. Proteomic analysis of temperature stress-responsive proteins in Arabidopsis thaliana rosette leaves. Mol. Biosyst. 2013, 9, 1257–1267. [Google Scholar] [CrossRef] [PubMed]
- Han, F.; Chen, H.; Li, X.J.; Yang, M.F.; Liu, G.S.; Shen, S.H. A comparative proteomic analysis of rice seedlings under various high-temperature stresses. Biochim. Biophys. Acta. 2009, 1794, 1625–1634. [Google Scholar] [CrossRef] [PubMed]
- Lee, D.G.; Ahsan, N.; Lee, S.H.; Young Kang, K.; Dong Bahk, J.; Lee, I.J.; Lee, B.H. A proteomic approach in analyzing heat-responsive proteins in rice leaves. Proteomics 2007, 7, 3369–3383. [Google Scholar] [CrossRef] [PubMed]
- Scafaro, A.P.; Haynes, P.A.; Atwell, B.J. Physiological and molecular changes in Oryza meridionalis Ng., a heat-tolerant species of wild rice. J. Exp. Bot. 2010, 61, 191–202. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.; Li, R.; Wang, R.; Wang, X.; Zheng, W.; Sun, Q.; Tong, S.; Dai, S.; Xu, S. Comparative proteomic analysis of flag leaves reveals new insight into wheat heat adaptation. Front Plant Sci. 2017, 8, 1086. [Google Scholar] [CrossRef]
- Wang, X.; Dinler, B.S.; Vignjevic, M.; Jacobsen, S.; Wollenweber, B. Physiological and proteome studies of responses to heat stress during grain filling in contrasting wheat cultivars. Plant Sci. 2015, 230, 33–50. [Google Scholar] [CrossRef]
- Hu, X.; Wu, L.; Zhao, F.; Zhang, D.; Li, N.; Zhu, G.; Li, C.; Wang, W. Phosphoproteomic analysis of the response of maize leaves to drought, heat and their combination stress. Front Plant Sci. 2015, 6, 298. [Google Scholar] [CrossRef]
- Zhao, F.; Zhang, D.; Zhao, Y.; Wang, W.; Yang, H.; Tai, F.; Li, C.; Hu, X. The difference of physiological and proteomic changes in maize leaves adaptation to drought, heat, and combined both stresses. Front Plant Sci. 2016, 7, 1471. [Google Scholar] [CrossRef]
- Ahsan, N.; Donnart, T.; Nouri, M.Z.; Komatsu, S. Tissue-specific defense and thermo-adaptive mechanisms of soybean seedlings under heat stress revealed by proteomic approach. J. Proteome Res. 2010, 9, 4189–4204. [Google Scholar] [CrossRef] [PubMed]
- Das, A.; Eldakak, M.; Paudel, B.; Kim, D.W.; Hemmati, H.; Basu, C.; Rohila, J.S. Leaf proteome analysis reveals prospective drought and heat stress response mechanisms in soybean. Biomed Res. Int. 2016, 2016, 6021047. [Google Scholar] [CrossRef] [PubMed]
- Huang, W.; Ma, H.Y.; Huang, Y.; Li, Y.; Wang, G.L.; Jiang, Q.; Wang, F.; Xiong, A.S. Comparative proteomic analysis provides novel insights into chlorophyll biosynthesis in celery under temperature stress. Physiol. Plant 2017, 161, 468–485. [Google Scholar] [CrossRef] [PubMed]
- Yan, J.; Yu, L.; Xuan, J.; Lu, Y.; Lu, S.; Zhu, W. De novo transcriptome sequencing and gene expression profiling of spinach (Spinacia oleracea L.) leaves under heat stress. Sci. Rep. 2016, 6, 19473. [Google Scholar] [CrossRef] [PubMed]
- Tang, Y.; Wen, X.; Lu, Q.; Yang, Z.; Cheng, Z.; Lu, C. Heat stress induces an aggregation of the light-harvesting complex of photosystem Ⅱ in spinach plants. Plant Physiol. 2007, 143, 629–638. [Google Scholar] [CrossRef] [PubMed]
- Komayama, K.; Khatoon, M.; Takenaka, D.; Horie, J.; Yamashita, A.; Yoshioka, M.; Nakayama, Y.; Yoshida, M.; Ohira, S.; Morita, N.; et al. Quality control of photosystem II: Cleavage and aggregation of heat-damaged D1 protein in spinach thylakoids. Biochim. Biophys. Acta. 2007, 1767, 838–846. [Google Scholar] [CrossRef]
- Yamashita, A.; Nijo, N.; Pospisil, P.; Morita, N.; Takenaka, D.; Aminaka, R.; Yamamoto, Y.; Yamamoto, Y. Quality control of photosystem II: Reactive oxygen species are responsible for the damage to photosystem II under moderate heat stress. J. Biol. Chem. 2008, 283, 28380–28391. [Google Scholar] [CrossRef]
- Jianfu, J.; Liu, X.; Liu, G.-T.; Liu, C.; Li, S.; Wang, L. Integrating omics and alternative splicing reveals insights into grape response to high temperature. Plant Physiol. 2017, 173, 1502–1518. [Google Scholar]
- Wise, R.; Olson, A.J.; Schrader, S.M.; Sharkey, T. Electron transport is the functional limitation of photosynthesis in field-grown Pima cotton plants at high temperature. Plant Cell Environ. 2004, 27, 717–724. [Google Scholar] [CrossRef]
- Camejo, D.; Rodriguez, P.; Morales, M.A.; Dell’Amico, J.M.; Torrecillas, A.; Alarcon, J.J. High temperature effects on photosynthetic activity of two tomato cultivars with different heat susceptibility. J. Plant Physiol. 2005, 162, 281–289. [Google Scholar] [CrossRef]
- McDonald, G.; Paulsen, G.M. High temperature effects on photosynthesis and water relations of grain legumes. Plant Soil 1997, 196, 47–58. [Google Scholar] [CrossRef]
- As, W.; Galani, S.; Ashraf, M.; Foolad, M. Heat tolerance in plants: An overview. Environ. Exp. Bot. 2007, 61, 199–223. [Google Scholar]
- Xu, C.; Huang, B. Differential proteomic response to heat stress in thermal Agrostis scabra and heat-sensitive Agrostis stolonifera. Physiol. Plant. 2010, 139, 192–204. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Cai, J.; Jiang, D.; Liu, F.; Dai, T.; Cao, W. Pre-anthesis high-temperature acclimation alleviates damage to the flag leaf caused by post-anthesis heat stress in wheat. J. Plant Physiol. 2011, 168, 585–593. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.T.; Ma, L.; Duan, W.; Wang, B.C.; Li, J.H.; Xu, H.G.; Yan, X.Q.; Yan, B.F.; Li, S.H.; Wang, J. Differential proteomic analysis of grapevine leaves by iTRAQ reveals responses to heat stress and subsequent recovery. BMC Plant Biol. 2014, 14, 110. [Google Scholar] [CrossRef] [PubMed]
- Portis, A.R., Jr. Rubisco activase-Rubisco’s catalytic chaperone. Photosynth. Res. 2003, 75, 11–27. [Google Scholar] [CrossRef] [PubMed]
- Kurek, I.; Chang, T.K.; Bertain, S.M.; Madrigal, A.; Liu, L.; Lassner, M.W.; Zhu, G. Enhanced thermostability of Arabidopsis Rubisco activase improves photosynthesis and growth rates under moderate heat stress. Plant Cell 2007, 19, 3230–3241. [Google Scholar] [CrossRef]
- Ristic, Z.; Momcilovic, I.; Bukovnik, U.; Prasad, P.V.; Fu, J.; Deridder, B.P.; Elthon, T.E.; Mladenov, N. Rubisco activase and wheat productivity under heat-stress conditions. J. Exp. Bot. 2009, 60, 4003–4014. [Google Scholar] [CrossRef]
- Horton, P. Crop Improvement through Alteration in the Photosynthetic Membrane; ISB News Report; Virginia Tech: Blacksburg, VA, USA, 2002. [Google Scholar]
- de Pinto, M.C.; Paradiso, A.; Leonetti, P.; De Gara, L. Hydrogen peroxide, nitric oxide and cytosolic ascorbate peroxidase at the crossroad between defence and cell death. Plant J. 2006, 48, 784–795. [Google Scholar] [CrossRef]
- Mittler, R.; Vanderauwera, S.; Gollery, M.; Van Breusegem, F. Reactive oxygen gene network of plants. Trends Plant Sci. 2004, 9, 490–498. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Yang, Y.; Sun, X.; Lin, H.; Chen, J.; Ren, J.; Hu, X.; Yang, Y. Comparative physiological and proteomic analyses of poplar (Populus yunnanensis) plantlets exposed to high temperature and drought. PLoS ONE 2014, 9, e107605. [Google Scholar] [CrossRef] [PubMed]
- Kumar, R. Protection against heat stress in wheat involves change in cell membrane stability, antioxidant enzymes, osmolyte, H2O2 and transcript of heat shock protein. Plant Physiol. Bioch. 2012, 4, 83–91. [Google Scholar]
- Sairam, R.K.; Srivastava, G.; Saxena, D. Increased antioxidant activity under elevated temperatures: A mechanism of heat stress tolerance in wheat genotypes. Biol. Plant. 2000, 43, 245–251. [Google Scholar] [CrossRef]
- Sengupta, D.; Naik, D.; Reddy, A.R. Plant aldo-keto reductases (AKRs) as multi-tasking soldiers involved in diverse plant metabolic processes and stress defense: A structure-function update. J. Plant Physiol. 2015, 179, 40–55. [Google Scholar] [CrossRef] [PubMed]
- Hayat, S.; Hayat, Q.; Alyemeni, M.N.; Wani, A.S.; Pichtel, J.; Ahmad, A. Role of proline under changing environments: A review. Plant Signal Behav. 2012, 7, 1456–1466. [Google Scholar] [CrossRef]
- Wahid, A.; Close, T. Expression of dehydrins under heat stress and their relationship with water relations of sugarcane leaves. Biol. Plant. 2007, 51, 104–109. [Google Scholar] [CrossRef]
- Rivero, R.; Jm, R.; Romero, L. Oxidative metabolism in tomato plants subjected to heat stress. J. Hortic. Sci. Biotech. 2004, 79, 560–564. [Google Scholar] [CrossRef]
- Cvikrova, M.; Gemperlova, L.; Dobra, J.; Martincova, O.; Prasil, I.T.; Gubis, J.; Vankova, R. Effect of heat stress on polyamine metabolism in proline-over-producing tobacco plants. Plant Sci. 2012, 182, 49–58. [Google Scholar] [CrossRef]
- Gaxiola, R.A.; Palmgren, M.G.; Schumacher, K. Plant proton pumps. FEBS Lett. 2007, 581, 2204–2214. [Google Scholar] [CrossRef]
- Savchenko, A.; Vieille, C.; Kang, S.; Zeikus, J.G. Pyrococcus furiosus alpha-amylase is stabilized by calcium and zinc. Biochemistry 2002, 41, 6193–6201. [Google Scholar] [CrossRef] [PubMed]
- Ruan, J.; Haerdter, R.; Gerendas, J. Impact of nitrogen supply on carbon/nitrogen allocation: A case study on amino acids and catechins in green tea [Camellia sinensis (L.) O. Kuntze] plants. Plant Biol. (Stuttg.) 2010, 12, 724–734. [Google Scholar] [CrossRef] [PubMed]
- Laino, P.; Shelton, D.; Finnie, C.; De Leonardis, A.; Mastrangelo, A.; Svensson, B.; Lafiandra, D.; Masci, S. Comparative proteome analysis of metabolic proteins from seeds of durum wheat (cv. Svevo) subjected to heat stress. Proteomics 2010, 10, 2359–2368. [Google Scholar] [CrossRef] [PubMed]
- Yang, F.; Jorgensen, A.D.; Li, H.; Sondergaard, I.; Finnie, C.; Svensson, B.; Jiang, D.; Wollenweber, B.; Jacobsen, S. Implications of high-temperature events and water deficits on protein profiles in wheat (Triticum aestivum L. cv. Vinjett) grain. Proteomics 2011, 11, 1684–1695. [Google Scholar] [CrossRef] [PubMed]
- Majoul, T.; Bancel, E.; Triboi, E.; Ben Hamida, J.; Branlard, G. Proteomic analysis of the effect of heat stress on hexaploid wheat grain: Characterization of heat-responsive proteins from non-prolamins fraction. Proteomics 2004, 4, 505–513. [Google Scholar] [CrossRef] [PubMed]
- Xu, C.; Huang, B. Root proteomic responses to heat stress in two Agrostis grass species contrasting in heat tolerance. J. Exp. Bot. 2008, 59, 4183–4194. [Google Scholar] [CrossRef]
- Cai, X.; Gao, C.; Su, B.; Tan, F.; Yang, N.; Wang, G. Expression profiling and microbial ligand binding analysis of high-mobility group box-1 (HMGB1) in turbot (Scophthalmus mavximus L.). Fish Shellfish Immunol. 2018, 78, 100–108. [Google Scholar] [CrossRef] [PubMed]
- Hopfner, K.P.; Karcher, A.; Shin, D.S.; Craig, L.; Arthur, L.M.; Carney, J.P.; Tainer, J.A. Structural biology of Rad50 ATPase: ATP-driven conformational control in DNA double-strand break repair and the ABC-ATPase superfamily. Cell 2000, 101, 789–800. [Google Scholar] [CrossRef]
- Johnova, P.; Skalak, J.; Saiz-Fernandez, I.; Brzobohaty, B. Plant responses to ambient temperature fluctuations and water-limiting conditions: A proteome-wide perspective. Biochim. Biophys. Acta. 2016, 1864, 916–931. [Google Scholar] [CrossRef]
- Larkindale, J.; Hall, J.D.; Knight, M.R.; Vierling, E. Heat stress phenotypes of Arabidopsis mutants implicate multiple signaling pathwayes in the acquisition of thermotolerance. Plant Physiol. 2005, 138, 882–897. [Google Scholar] [CrossRef]
- Salvucci, M.E. Association of Rubisco activase with chaperonin-60beta: A possible mechanism for protecting photosynthesis during heat stress. J. Exp. Bot. 2008, 59, 1923–1933. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Chen, J.; Liu, Q.; Ben, C.; Todd, C.D.; Shi, J.; Yang, Y.; Hu, X. Comparative proteomic analysis of the thermotolerant plant Portulaca oleracea acclimation to combined high temperature and humidity stress. J. Proteome Res. 2012, 11, 3605–3623. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Li, G.; Huang, W.; Bi, T.; Chen, G.; Tang, Z.; Su, W.; Sun, W. Proteomic study of Carissa spinarum in response to combined heat and drought stress. Proteomics 2010, 10, 3117–3129. [Google Scholar] [CrossRef] [PubMed]
- Perez, D.E.; Hoyer, J.S.; Johnson, A.I.; Moody, Z.R.; Lopez, J.; Kaplinsky, N.J. BOBBER1 is a noncanonical Arabidopsis small heat shock protein required for both development and thermotolerance. Plant Physiol. 2009, 151, 241–252. [Google Scholar] [CrossRef]
- Yu, J.; Chen, S.; Zhao, Q.; Wang, T.; Yang, C.; Diaz, C.; Sun, G.; Dai, S. Physiological and proteomic analysis of salinity tolerance in Puccinellia tenuiflora. J. Proteome Res. 2011, 10, 3852–3870. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Chen, S.; Zhang, H.; Shi, L.; Cao, F.; Guo, L.; Xie, Y.; Wang, T.; Yan, X.; Dai, S. Desiccation tolerance mechanism in resurrection fern-ally Selaginella tamariscina revealed by physiological and proteomic analysis. J. Proteome Res. 2010, 9, 6561–6577. [Google Scholar] [CrossRef]
- Ibrahim, M.H.; Jaafar, H.Z. Primary, secondary metabolites, H2O2, malondialdehyde and photosynthetic responses of Orthosiphon stimaneus Benth. to different irradiance levels. Molecules 2012, 17, 1159–1176. [Google Scholar] [CrossRef]
- Yin, Z.; Balmant, K.; Geng, S.; Zhu, N.; Zhang, T.; Dufresne, C.; Dai, S.; Chen, S. Bicarbonate induced redox proteome changes in Arabidopsis suspension cells. Front Plant Sci. 2017, 8, 58. [Google Scholar] [CrossRef]
- Suo, J.; Zhao, Q.; Zhang, Z.; Chen, S.; Cao, J.; Liu, G.; Wei, X.; Wang, T.; Yang, C.; Dai, S. Cytological and proteomic analyses of osmunda cinnamomea germinating spores reveal characteristics of fern spore germination and rhizoid tip growth. Mol. Cell. Proteom. 2015, 14, 2510–2534. [Google Scholar] [CrossRef]
- Protein Database. Available online: http://www.spinachbase.org/ (accessed on 20 September 2018).
- PSI-BLAST and PHI-BLAST Program. Available online: http://www.ncbi.nlm.nih.gov/BLAST/ (accessed on 15 April 2018).
- KEGG Pathway Database. Available online: http://www.kegg.jp/kegg (accessed on 15 April 2018).
- UniProt Database. Available online: http://www.ebi.uniprot.org/ (accessed on 15 May 2018).
- Gene Ontology Database. Available online: http://geneontology.org (accessed on 15 May 2018).
- YLoc: Interpretable Subcellular Localization Prediction. Available online: http://abi.inf.uni-tuebingen.de/Services/YLoc/webloc.cgi (accessed on 15 August 2018).
- LocTree3: Protein Subcellular Localization Prediction System. Available online: https://rostlab.org/services/loctree3/ (accessed on 15 August 2018).
- ngLOC: A Bayesian Method for Predicting Protein Subcellular Localization. Available online: http://genome.unmc.edu/ngLOC/index.html (accessed on 15 August 2018).
- Plant-mPLoc: Predicting Subcellular Localization of Plant Proteins Including Those with Multiple Sites. Available online: http://www.csbio.sjtu.edu.cn/bioinf/plant-multi/# (accessed on 15 August 2018).
- TargetP Server. Available online: http://www.cbs.dtu.dk/services/TargetP/ (accessed on 15 August 2018).
- Web Tool of STRING 10.5. Available online: http://string-db.org (accessed on 20 September 2018).


© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, S.; Yu, J.; Li, Y.; Zhang, H.; Bao, X.; Bian, J.; Xu, C.; Wang, X.; Cai, X.; Wang, Q.; et al. Heat-Responsive Proteomics of a Heat-Sensitive Spinach Variety. Int. J. Mol. Sci. 2019, 20, 3872. https://doi.org/10.3390/ijms20163872
Li S, Yu J, Li Y, Zhang H, Bao X, Bian J, Xu C, Wang X, Cai X, Wang Q, et al. Heat-Responsive Proteomics of a Heat-Sensitive Spinach Variety. International Journal of Molecular Sciences. 2019; 20(16):3872. https://doi.org/10.3390/ijms20163872
Chicago/Turabian StyleLi, Shanshan, Juanjuan Yu, Ying Li, Heng Zhang, Xuesong Bao, Jiayi Bian, Chenxi Xu, Xiaoli Wang, Xiaofeng Cai, Quanhua Wang, and et al. 2019. "Heat-Responsive Proteomics of a Heat-Sensitive Spinach Variety" International Journal of Molecular Sciences 20, no. 16: 3872. https://doi.org/10.3390/ijms20163872
APA StyleLi, S., Yu, J., Li, Y., Zhang, H., Bao, X., Bian, J., Xu, C., Wang, X., Cai, X., Wang, Q., Wang, P., Guo, S., Miao, Y., Chen, S., Qin, Z., & Dai, S. (2019). Heat-Responsive Proteomics of a Heat-Sensitive Spinach Variety. International Journal of Molecular Sciences, 20(16), 3872. https://doi.org/10.3390/ijms20163872





