Marek’s Disease Virus Infection Induced Mitochondria Changes in Chickens
Abstract
1. Introduction
2. Results
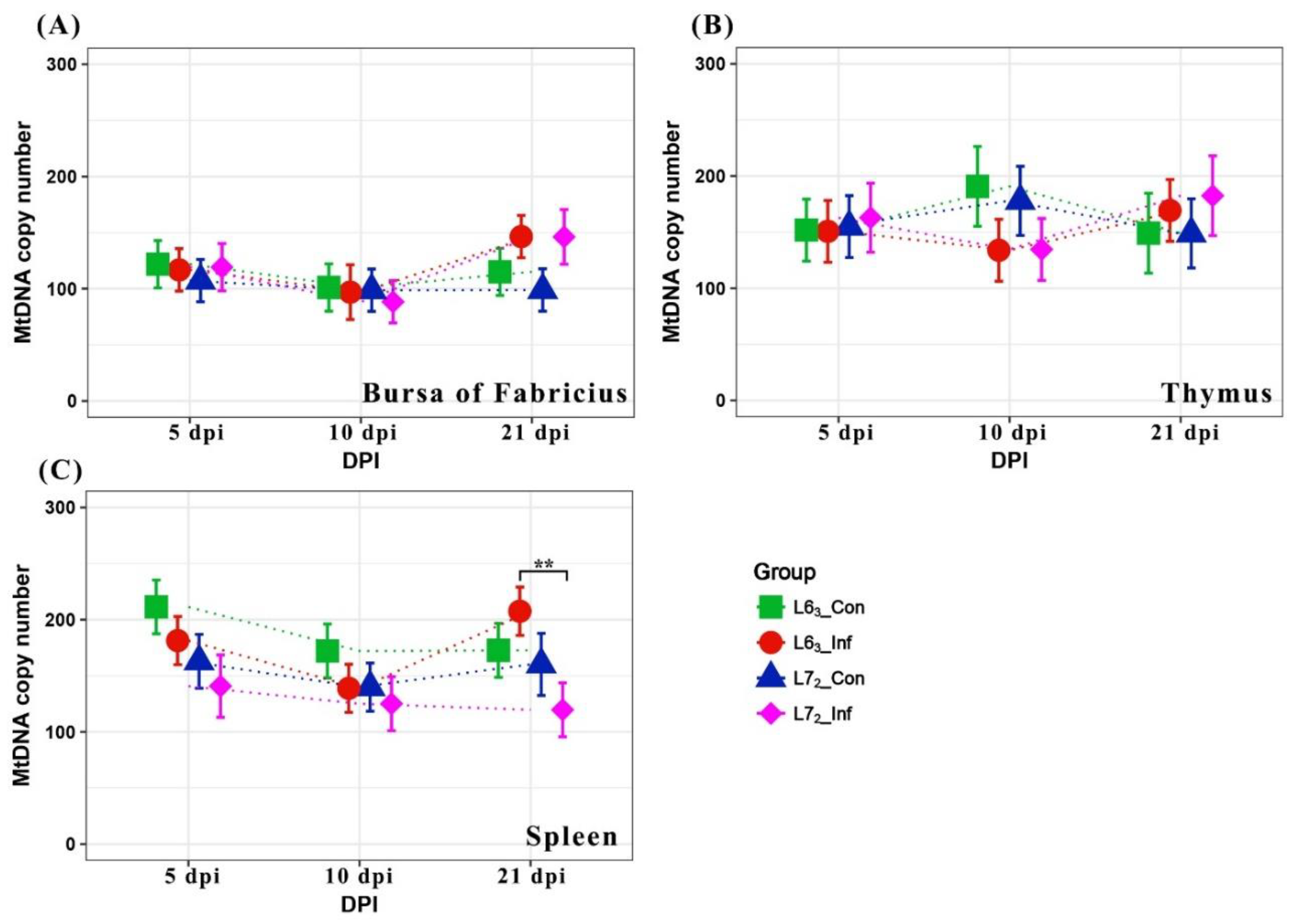
2.1. Mitochondrial DNA Copy Number Variation
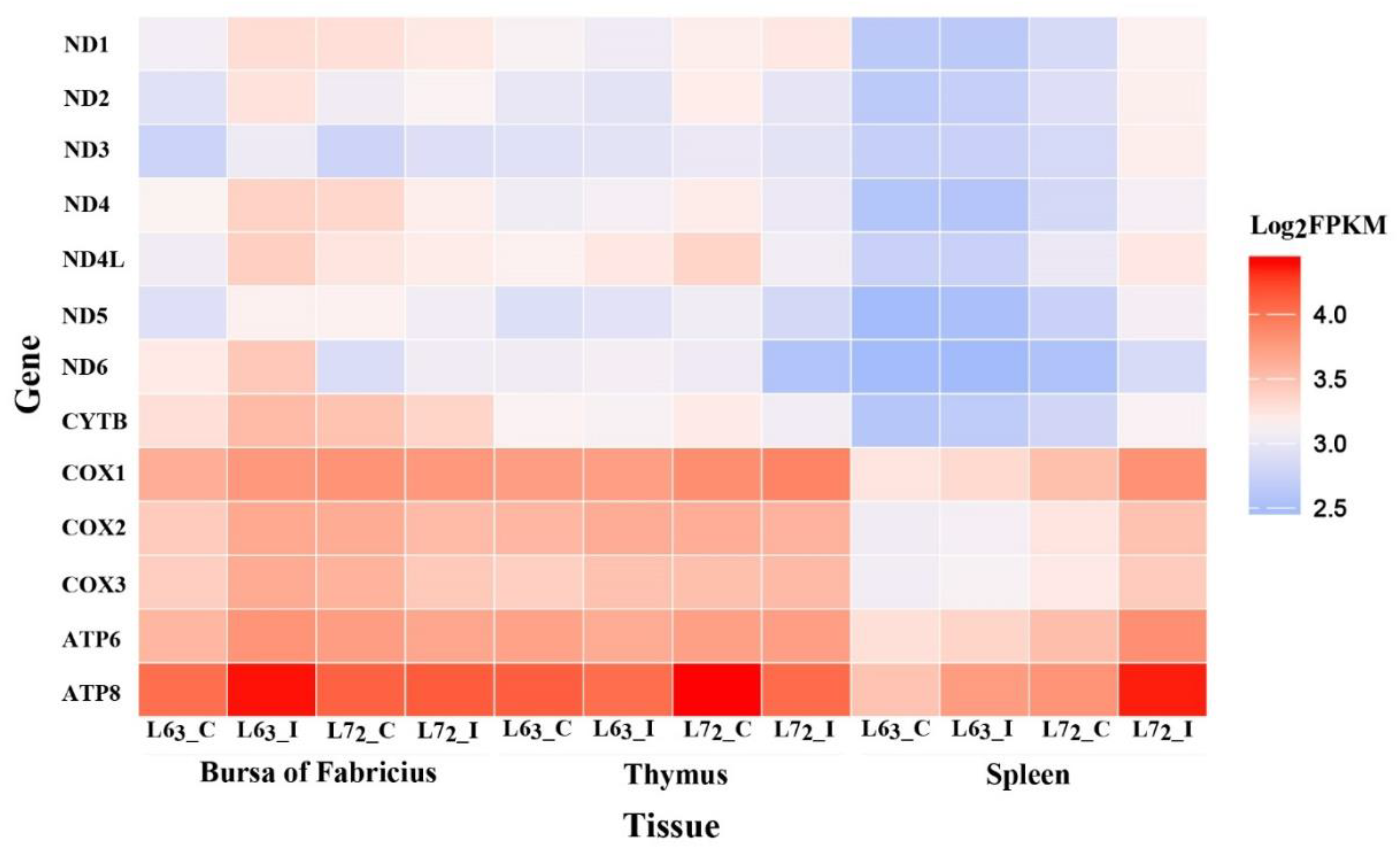
2.2. The Expressions of Mitochondrial DNA-coding Genes
2.3. Differentially Expressed MitoProteome Nuclear Genes
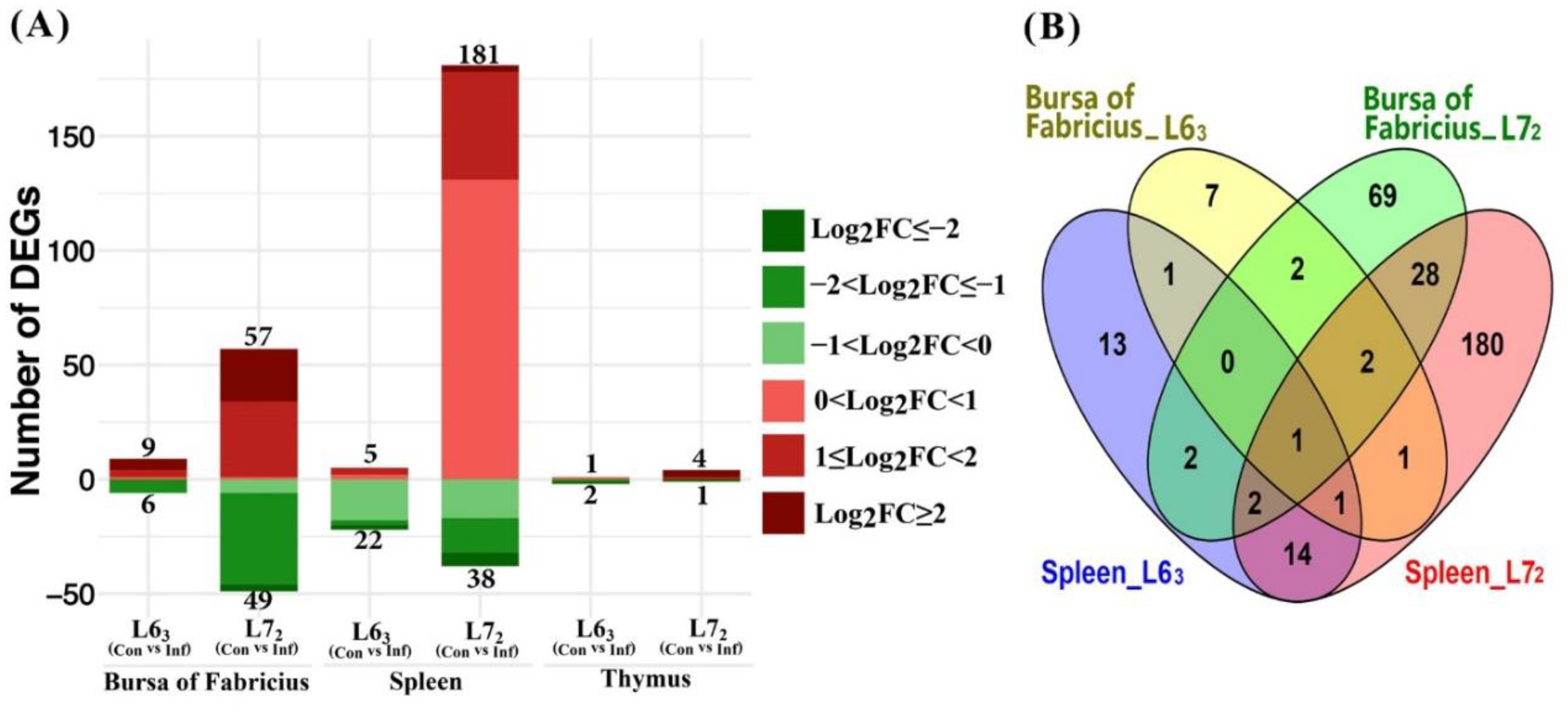
2.3.1. Mitochondria-Related Nuclear DEGs Induced by MDV
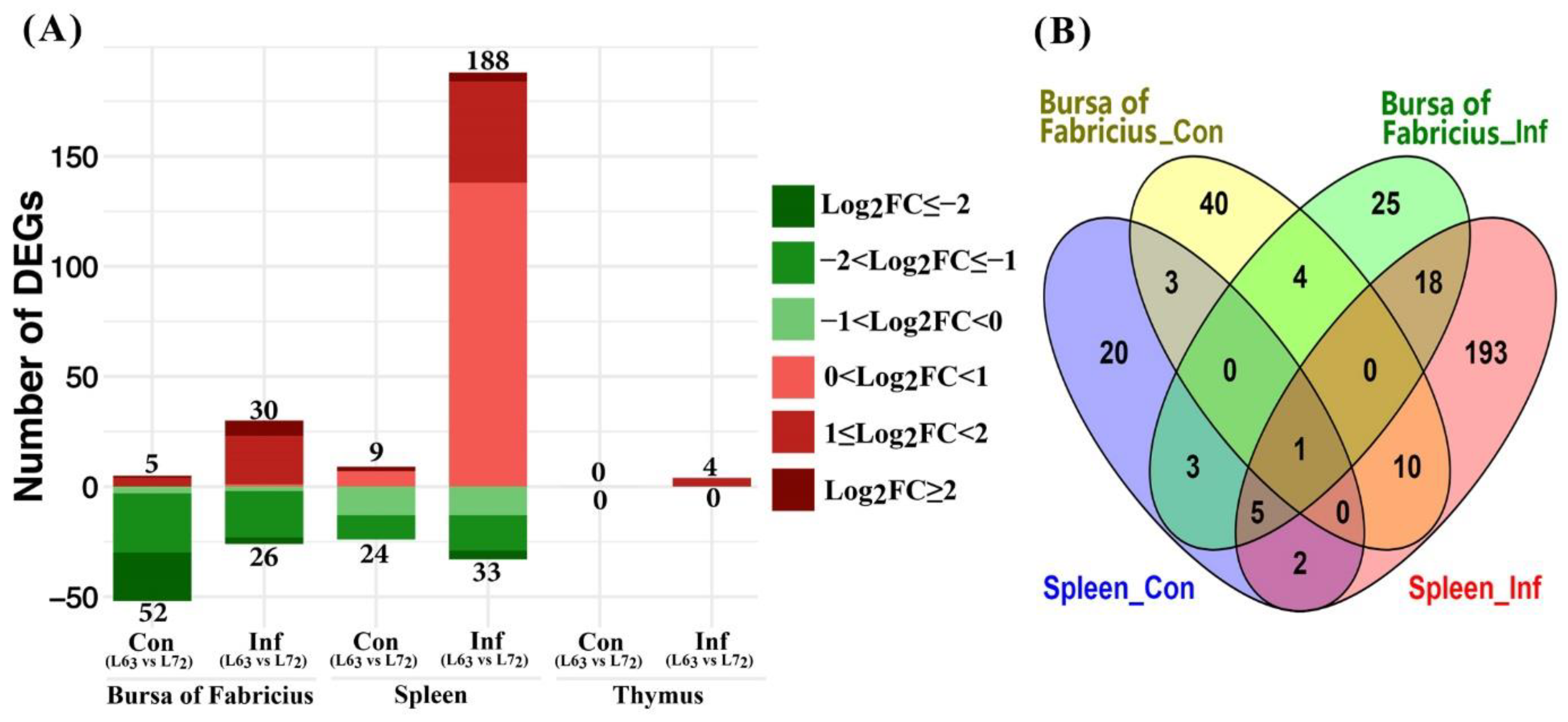
2.3.2. Mitochondria-Related Nuclear DEGs between Two Chicken Lines
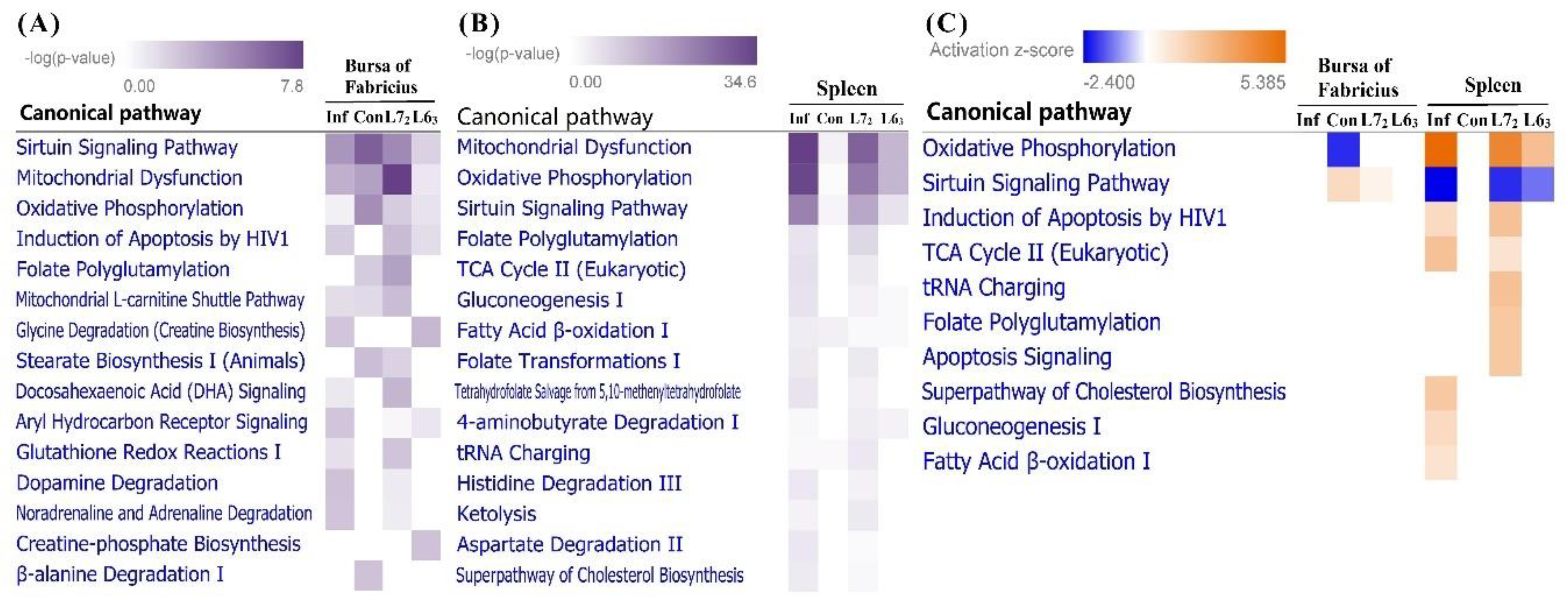
2.3.3. Canonical Pathways Prediction
2.3.4. Nuclear Genes Involving in the mtDNA Replication, Transcription and Maintenance
3. Discussion
3.1. MtDNA Content and Gene Expression
3.2. Mitochondria-Related Nuclear Genes and Pathway Analysis
3.3. The Mitochondrial DNA Replication and Transcription upon MDV Infection
4. Materials and Methods
4.1. Ethics Statement
4.2. Chickens, Treatment, and Samples
4.3. Quantification of mtDNA Copy Number
4.4. RNA Sequencing
4.5. MitoProteome Gene Differential Expression Analysis and Pathway Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Mayr, J.A. Lipid metabolism in mitochondrial membranes. J. Inherit. Metab. Dis. 2015, 38, 137–144. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.H.; Wei, Y.H. Role of mitochondrial dysfunction and dysregulation of Ca(2+) homeostasis in the pathophysiology of insulin resistance and type 2 diabetes. J. Biomed. Sci. 2017, 24, 70. [Google Scholar] [CrossRef] [PubMed]
- Monlun, M.; Hyernard, C.; Blanco, P.; Lartigue, L.; Faustin, B. Mitochondria as Molecular Platforms Integrating Multiple Innate Immune Signalings. J. Mol. Biol. 2017, 429, 1–13. [Google Scholar] [CrossRef] [PubMed]
- West, A.P.; Shadel, G.S. Mitochondrial DNA in innate immune responses and inflammatory pathology. Nat. Rev. Immunol. 2017, 17, 363–375. [Google Scholar] [CrossRef] [PubMed]
- Kwak, G.H.; Kim, T.H.; Kim, H.Y. Down-regulation of MsrB3 induces cancer cell apoptosis through reactive oxygen species production and intrinsic mitochondrial pathway activation. Biochem. Biophys. Res. Commun. 2017, 483, 468–474. [Google Scholar] [CrossRef]
- Yan, X.J.; Yu, X.; Wang, X.P.; Jiang, J.F.; Yuan, Z.Y.; Lu, X.; Lei, F.; Xing, D.M. Mitochondria play an important role in the cell proliferation suppressing activity of berberine. Sci. Rep. 2017, 7, 41712. [Google Scholar] [CrossRef] [PubMed]
- Reverter, A.; Okimoto, R.; Sapp, R.; Bottje, W.G.; Hawken, R.; Hudson, N.J. Chicken muscle mitochondrial content appears co-ordinately regulated and is associated with performance phenotypes. Biol. Open 2017, 6, 50–58. [Google Scholar] [CrossRef]
- Weikard, R.; Kuehn, C. Different mitochondrial DNA copy number in liver and mammary gland of lactating cows with divergent genetic background for milk production. 2018. [Google Scholar] [CrossRef]
- Mengel-From, J.; Thinggaard, M.; Dalgard, C.; Kyvik, K.O.; Christensen, K.; Christiansen, L. Mitochondrial DNA copy number in peripheral blood cells declines with age and is associated with general health among elderly. Hum. Genet. 2014, 133, 1149–1159. [Google Scholar] [CrossRef]
- Ashar, F.N.; Zhang, Y.; Longchamps, R.J.; Lane, J.; Moes, A.; Grove, M.L.; Mychaleckyj, J.C.; Taylor, K.D.; Coresh, J.; Rotter, J.I.; et al. Association of Mitochondrial DNA Copy Number With Cardiovascular Disease. Jama Cardiol. 2017, 2, 1247–1255. [Google Scholar] [CrossRef]
- Leavy, O. T cells: Mitochondria and T cell activation. Nat. Rev. Immunol. 2013, 13, 224. [Google Scholar] [CrossRef]
- Liu, P.S.; Ho, P.C. Mitochondria: A master regulator in macrophage and T cell immunity. Mitochondrion 2018, 41, 45–50. [Google Scholar] [CrossRef] [PubMed]
- Sandoval, H.; Kodali, S.; Wang, J. Regulation of B cell fate, survival, and function by mitochondria and autophagy. Mitochondrion 2018, 41, 58–65. [Google Scholar] [CrossRef] [PubMed]
- Anand, S.K.; Tikoo, S.K. Viruses as modulators of mitochondrial functions. Adv. Virol. 2013, 2013, 738794. [Google Scholar] [CrossRef] [PubMed]
- Saffran, H.A.; Pare, J.M.; Corcoran, J.A.; Weller, S.K.; Smiley, J.R. Herpes simplex virus eliminates host mitochondrial DNA. Embo Rep. 2007, 8, 188–193. [Google Scholar] [CrossRef] [PubMed]
- Weinberg, S.E.; Sena, L.A.; Chandel, N.S. Mitochondria in the regulation of innate and adaptive immunity. Immunity 2015, 42, 406–417. [Google Scholar] [CrossRef] [PubMed]
- Robinson, C.M.; Hunt, H.D.; Cheng, H.H.; Delany, M.E. Chromosomal integration of an avian oncogenic herpesvirus reveals telomeric preferences and evidence for lymphoma clonality. Herpesviridae 2010, 1, 5. [Google Scholar] [CrossRef] [PubMed]
- Robinson, C.M.; Cheng, H.H.; Delany, M.E. Temporal kinetics of Marek’s disease herpesvirus: Integration occurs early after infection in both B and T cells. Cytogenet. Genome Res. 2014, 144, 142–154. [Google Scholar] [CrossRef]
- McPherson, M.C.; Delany, M.E. Virus and host genomic, molecular, and cellular interactions during Marek’s disease pathogenesis and oncogenesis. Poult. Sci. 2016, 95, 412–429. [Google Scholar] [CrossRef]
- McPherson, M.C.; Cheng, H.H.; Delany, M.E. Marek’s disease herpesvirus vaccines integrate into chicken host chromosomes yet lack a virus-host phenotype associated with oncogenic transformation. Vaccine 2016, 34, 5554–5561. [Google Scholar] [CrossRef]
- Boodhoo, N.; Gurung, A.; Sharif, S.; Behboudi, S. Marek’s disease in chickens: A review with focus on immunology. Vet. Res. 2016, 47, 119. [Google Scholar] [CrossRef]
- Luo, J.; Yu, Y.; Chang, S.; Tian, F.; Zhang, H.; Song, J. DNA Methylation Fluctuation Induced by Virus Infection Differs between MD-resistant and -susceptible Chickens. Front. Genet. 2012, 3, 20. [Google Scholar] [CrossRef] [PubMed]
- Xie, Q.; Chang, S.; Dong, K.; Dunn, J.R.; Song, J.; Zhang, H. Genomic Variation between Genetic Lines of White Leghorns Differed in Resistance to Marek’s Disease. J. Clin. Epigenetics 2017, 03. [Google Scholar] [CrossRef]
- Xu, L.; He, Y.; Ding, Y.; Sun, G.; Carrillo, J.A.; Li, Y.; Ghaly, M.M.; Ma, L.; Zhang, H.; Liu, G.E.; et al. Characterization of Copy Number Variation’s Potential Role in Marek’s Disease. Int. J. Mol. Sci. 2017, 18. [Google Scholar] [CrossRef] [PubMed]
- Bacon, L.D.; Hunt, H.D.; Cheng, H.H. A review of the development of chicken lines to resolve genes determining resistance to diseases. Poult. Sci. 2000, 79, 1082–1093. [Google Scholar] [CrossRef] [PubMed]
- Han, B.; He, Y.; Zhang, L.; Ding, Y.; Lian, L.; Zhao, C.; Song, J.; Yang, N. Long intergenic non-coding RNA GALMD3 in chicken Marek’s disease. Sci. Rep. 2017, 7, 10294. [Google Scholar] [CrossRef] [PubMed]
- Chodkowski, M.; Serafinska, I.; Brzezicka, J.; Golke, A.; Slonska, A.; Krzyzowska, M.; Orlowski, P.; Baska, P.; Banbura, M.W.; Cymerys, J. Human herpesvirus type 1 and type 2 disrupt mitochondrial dynamics in human keratinocytes. 2018. [Google Scholar] [CrossRef]
- Reznik, E.; Miller, M.L.; Şenbabaoğlu, Y.; Riaz, N.; Sarungbam, J.; Tickoo, S.K.; Al-Ahmadie, H.A.; Lee, W.; Seshan, V.E.; Hakimi, A.A. Mitochondrial DNA copy number variation across human cancers. eLife 2016, 5, e10769. [Google Scholar] [CrossRef] [PubMed]
- Mesri, E.A.; Feitelson, M.A.; Munger, K. Human viral oncogenesis: A cancer hallmarks analysis. Cell Host Microbe 2014, 15, 266–282. [Google Scholar] [CrossRef]
- Calvo, S.E.; Mootha, V.K. The mitochondrial proteome and human disease. Annu. Rev. Genom. Hum. Genet. 2010, 11, 25–44. [Google Scholar] [CrossRef] [PubMed]
- Smith, J.; Sadeyen, J.R.; Paton, I.R.; Hocking, P.M.; Salmon, N.; Fife, M.; Nair, V.; Burt, D.W.; Kaiser, P. Systems analysis of immune responses in Marek’s disease virus-infected chickens identifies a gene involved in susceptibility and highlights a possible novel pathogenicity mechanism. J. Virol. 2011, 85, 11146–11158. [Google Scholar] [CrossRef]
- Gonzalez Herrera, K.N.; Lee, J.; Haigis, M.C. Intersections between mitochondrial sirtuin signaling and tumor cell metabolism. Crit. Rev. Biochem. Mol. Biol. 2015, 50, 242–255. [Google Scholar] [CrossRef]
- Viale, A.; Pettazzoni, P.; Lyssiotis, C.A.; Ying, H.; Sanchez, N.; Marchesini, M.; Carugo, A.; Green, T.; Seth, S.; Giuliani, V.; et al. Oncogene ablation-resistant pancreatic cancer cells depend on mitochondrial function. Nature 2014, 514, 628–632. [Google Scholar] [CrossRef] [PubMed]
- Sullivan, L.B.; Gui, D.Y.; Hosios, A.M.; Bush, L.N.; Freinkman, E.; Vander Heiden, M.G. Supporting Aspartate Biosynthesis Is an Essential Function of Respiration in Proliferating Cells. Cell 2015, 162, 552–563. [Google Scholar] [CrossRef] [PubMed]
- Molina, J.R.; Sun, Y.; Protopopova, M.; Gera, S.; Bandi, M.; Bristow, C.; McAfoos, T.; Morlacchi, P.; Ackroyd, J. An inhibitor of oxidative phosphorylation exploits cancer vulnerability. Nat. Med. 2018, 24, 1036–1046. [Google Scholar] [CrossRef] [PubMed]
- Cruz, P.M.; Mo, H.; McConathy, W.J.; Sabnis, N.; Lacko, A.G. The role of cholesterol metabolism and cholesterol transport in carcinogenesis: A review of scientific findings, relevant to future cancer therapeutics. Front. Pharmacol. 2013, 4, 119. [Google Scholar] [CrossRef] [PubMed]
- Bringman-Rodenbarger, L.R.; Guo, A.H.; Lyssiotis, C.A.; Lombard, D.B. Emerging Roles for SIRT5 in Metabolism and Cancer. Antioxid. Redox Signal. 2018, 28, 677–690. [Google Scholar] [CrossRef]
- Chalkiadaki, A.; Guarente, L. The multifaceted functions of sirtuins in cancer. Nat. Rev. Cancer 2015, 15, 608–624. [Google Scholar] [CrossRef] [PubMed]
- O’Callaghan, C.; Vassilopoulos, A. Sirtuins at the crossroads of stemness, aging, and cancer. Aging Cell 2017, 16, 1208–1218. [Google Scholar] [CrossRef]
- Zhao, S.; Xu, W.; Jiang, W.; Yu, W.; Lin, Y.; Zhang, T.; Yao, J.; Zhou, L.; Zeng, Y.; Li, H.; et al. Regulation of cellular metabolism by protein lysine acetylation. Sci. (New YorkN.Y.) 2010, 327, 1000–1004. [Google Scholar] [CrossRef]
- Bonda, D.J.; Lee, H.G.; Camins, A.; Pallas, M.; Casadesus, G.; Smith, M.A.; Zhu, X. The sirtuin pathway in ageing and Alzheimer disease: Mechanistic and therapeutic considerations. Lancet. Neurol. 2011, 10, 275–279. [Google Scholar] [CrossRef]
- Gong, M.; Hay, S.; Marshall, K.R.; Munro, A.W.; Scrutton, N.S. DNA binding suppresses human AIF-M2 activity and provides a connection between redox chemistry, reactive oxygen species, and apoptosis. J. Biol. Chem. 2007, 282, 30331–30340. [Google Scholar] [CrossRef]
- Gurung, A.; Kamble, N.; Kaufer, B.B.; Pathan, A.; Behboudi, S. Association of Marek’s Disease induced immunosuppression with activation of a novel regulatory T cells in chickens. Plos Pathog. 2017, 13, e1006745. [Google Scholar] [CrossRef] [PubMed]
- Pearce, S.F.; Rebelo-Guiomar, P.; D’Souza, A.R.; Powell, C.A.; Van Haute, L.; Minczuk, M. Regulation of Mammalian Mitochondrial Gene Expression: Recent Advances. Trends Biochem. Sci. 2017, 42, 625–639. [Google Scholar] [CrossRef] [PubMed]
- Korhonen, J.A.; Gaspari, M.; Falkenberg, M. TWINKLE Has 5’ -> 3’ DNA helicase activity and is specifically stimulated by mitochondrial single-stranded DNA-binding protein. J. Biol. Chem. 2003, 278, 48627–48632. [Google Scholar] [CrossRef] [PubMed]
- Milenkovic, D.; Matic, S.; Kuhl, I.; Ruzzenente, B.; Freyer, C.; Jemt, E.; Park, C.B.; Falkenberg, M.; Larsson, N.G. TWINKLE is an essential mitochondrial helicase required for synthesis of nascent D-loop strands and complete mtDNA replication. Hum. Mol. Genet. 2013, 22, 1983–1993. [Google Scholar] [CrossRef] [PubMed]
- Miralles Fuste, J.; Shi, Y.; Wanrooij, S.; Zhu, X.; Jemt, E.; Persson, O.; Sabouri, N.; Gustafsson, C.M.; Falkenberg, M. In vivo occupancy of mitochondrial single-stranded DNA binding protein supports the strand displacement mode of DNA replication. Plos Genet. 2014, 10, e1004832. [Google Scholar] [CrossRef] [PubMed]
- Ruhanen, H.; Borrie, S.; Szabadkai, G.; Tyynismaa, H.; Jones, A.W.; Kang, D.; Taanman, J.W.; Yasukawa, T. Mitochondrial single-stranded DNA binding protein is required for maintenance of mitochondrial DNA and 7S DNA but is not required for mitochondrial nucleoid organisation. Biochim. Et Biophys. Acta 2010, 1803, 931–939. [Google Scholar] [CrossRef] [PubMed]
- Kaukonen, J.; Juselius, J.K.; Tiranti, V.; Kyttala, A.; Zeviani, M.; Comi, G.P.; Keranen, S.; Peltonen, L.; Suomalainen, A. Role of adenine nucleotide translocator 1 in mtDNA maintenance. Sci. (New YorkN.Y.) 2000, 289, 782–785. [Google Scholar] [CrossRef] [PubMed]
- Kornblum, C.; Nicholls, T.J.; Haack, T.B.; Scholer, S.; Peeva, V.; Danhauser, K.; Hallmann, K.; Zsurka, G.; Rorbach, J.; Iuso, A.; et al. Loss-of-function mutations in MGME1 impair mtDNA replication and cause multisystemic mitochondrial disease. Nat. Genet. 2013, 45, 214–219. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, N.; Ronchi, D.; Comi, G.P. Genes and Pathways Involved in Adult Onset Disorders Featuring Muscle Mitochondrial DNA Instability. Int. J. Mol. Sci. 2015, 16, 18054–18076. [Google Scholar] [CrossRef]
- Thompson, K.; Majd, H.; Dallabona, C.; Reinson, K.; King, M.S.; Alston, C.L.; He, L.; Lodi, T.; Jones, S.A.; Fattal-Valevski, A.; et al. Recurrent De Novo Dominant Mutations in SLC25A4 Cause Severe Early-Onset Mitochondrial Disease and Loss of Mitochondrial DNA Copy Number. Am. J. Hum. Genet. 2016, 99, 860–876. [Google Scholar] [CrossRef]
- Jia, P.P.; Junaid, M.; Ma, Y.B.; Ahmad, F.; Jia, Y.F.; Li, W.G.; Pei, D.S. Role of human DNA2 (hDNA2) as a potential target for cancer and other diseases: A systematic review. Dna Repair 2017, 59, 9–19. [Google Scholar] [CrossRef] [PubMed]
- Hyvarinen, A.K.; Pohjoismaki, J.L.; Reyes, A.; Wanrooij, S.; Yasukawa, T.; Karhunen, P.J.; Spelbrink, J.N.; Holt, I.J.; Jacobs, H.T. The mitochondrial transcription termination factor mTERF modulates replication pausing in human mitochondrial DNA. Nucleic Acids Res. 2007, 35, 6458–6474. [Google Scholar] [CrossRef] [PubMed]
- Pellegrini, M.; Asin-Cayuela, J.; Erdjument-Bromage, H.; Tempst, P.; Larsson, N.G.; Gustafsson, C.M. MTERF2 is a nucleoid component in mammalian mitochondria. Biochim. Et Biophys. Acta 2009, 1787, 296–302. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Zhou, G.; Yu, M.; He, Y.; Tang, W.; Lai, J.; He, J.; Liu, W.; Tan, D. Cloning and functional analysis of human mTERFL encoding a novel mitochondrial transcription termination factor-like protein. Biochem. Biophys. Res. Commun. 2005, 337, 1112–1118. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.; Gao, P.; Qiu, S.; Zhang, L.; Yang, L.; Zuo, J.; Zhong, C.; Zhu, S.; Liu, W. MTERF2 contributes to MPP(+)-induced mitochondrial dysfunction and cell damage. Biochem. Biophys. Res. Commun. 2016, 471, 177–183. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y.; Posse, V.; Zhu, X.; Hyvarinen, A.K.; Jacobs, H.T.; Falkenberg, M.; Gustafsson, C.M. Mitochondrial transcription termination factor 1 directs polar replication fork pausing. Nucleic Acids Res. 2016, 44, 5732–5742. [Google Scholar] [CrossRef]
- Clemente, P.; Pajak, A.; Laine, I.; Wibom, R.; Wedell, A.; Freyer, C.; Wredenberg, A. SUV3 helicase is required for correct processing of mitochondrial transcripts. Nucleic Acids Res. 2015, 43, 7398–7413. [Google Scholar] [CrossRef]
- Wang, D.D.; Guo, X.E.; Modrek, A.S.; Chen, C.F.; Chen, P.L.; Lee, W.H. Helicase SUV3, polynucleotide phosphorylase, and mitochondrial polyadenylation polymerase form a transient complex to modulate mitochondrial mRNA polyadenylated tail lengths in response to energetic changes. J. Biol. Chem. 2014, 289, 16727–16735. [Google Scholar] [CrossRef]
- Litonin, D.; Sologub, M.; Shi, Y.; Savkina, M.; Anikin, M.; Falkenberg, M.; Gustafsson, C.M.; Temiakov, D. Human mitochondrial transcription revisited: Only TFAM and TFB2M are required for transcription of the mitochondrial genes in vitro. J. Biol. Chem. 2010, 285, 18129–18133. [Google Scholar] [CrossRef]
- Stiles, A.R.; Simon, M.T.; Stover, A.; Eftekharian, S.; Khanlou, N.; Wang, H.L.; Magaki, S.; Lee, H.; Partynski, K.; Dorrani, N.; et al. Mutations in TFAM, encoding mitochondrial transcription factor A, cause neonatal liver failure associated with mtDNA depletion. Mol. Genet. Metab. 2016, 119, 91–99. [Google Scholar] [CrossRef]
- Baixauli, F.; Acin-Perez, R.; Villarroya-Beltri, C.; Mazzeo, C.; Nunez-Andrade, N.; Gabande-Rodriguez, E.; Ledesma, M.D.; Blazquez, A.; Martin, M.A.; Falcon-Perez, J.M.; et al. Mitochondrial Respiration Controls Lysosomal Function during Inflammatory T Cell Responses. Cell Metab. 2015, 22, 485–498. [Google Scholar] [CrossRef] [PubMed]
- Hudson, G.; Chinnery, P.F. Mitochondrial DNA polymerase-gamma and human disease. Hum. Mol. Genet. 2006, 15, R244–R252. [Google Scholar] [CrossRef]
- Garcia-Gomez, S.; Reyes, A.; Martinez-Jimenez, M.I.; Chocron, E.S.; Mouron, S.; Terrados, G.; Powell, C.; Salido, E.; Mendez, J.; Holt, I.J.; et al. PrimPol, an archaic primase/polymerase operating in human cells. Mol. Cell 2013, 52, 541–553. [Google Scholar] [CrossRef] [PubMed]
- Lefai, E.; Fernandez-Moreno, M.A.; Alahari, A.; Kaguni, L.S.; Garesse, R. Differential regulation of the catalytic and accessory subunit genes of Drosophila mitochondrial DNA polymerase. J. Biol. Chem. 2000, 275, 33123–33133. [Google Scholar] [CrossRef] [PubMed]
- Varma, H.; Faust, P.L.; Iglesias, A.D.; Lagana, S.M.; Wou, K.; Hirano, M.; DiMauro, S.; Mansukani, M.M.; Hoff, K.E.; Nagy, P.L.; et al. Whole exome sequencing identifies a homozygous POLG2 missense variant in an infant with fulminant hepatic failure and mitochondrial DNA depletion. Eur. J. Med Genet. 2016, 59, 540–545. [Google Scholar] [CrossRef] [PubMed]
- Danaher, R.J.; McGarrell, B.S.; Stromberg, A.J.; Miller, C.S. Herpes simplex virus type 1 modulates cellular gene expression during quiescent infection of neuronal cells. Arch. Virol. 2008, 153, 1335–1345. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Ding, Y.; Zhan, F.; Zhang, H.; Han, B.; Hu, G.; Zhao, K.; Yang, N.; Yu, Y.; Mao, L.; et al. The conservation and signatures of lincRNAs in Marek’s disease of chicken. Sci. Rep. 2015, 5, 15184. [Google Scholar] [CrossRef] [PubMed]
- Cotter, D.; Guda, P.; Fahy, E.; Subramaniam, S. MitoProteome: Mitochondrial protein sequence database and annotation system. Nucleic Acids Res. 2004, 32, D463–D467. [Google Scholar] [CrossRef]
| Tissue | Comparison | Up-Regulated Gene | Down-Regulated Gene |
|---|---|---|---|
| Bursa of Fabricius | Con (L63 vs. L72) | ND6 | |
| Thymus | L72 (Con vs. Inf) | ND6 | |
| Spleen | L63 (Con vs. Inf) | ND1, ND2, ND4, ND5, COX1, COX2, CYTB | |
| L72 (Con vs. Inf) | ND1, ND2, ND3, ND4, ND5, ND6, COX1, COX2, CYTB, ATP6 | ||
| Con (L63 vs. L72) | ND6 | ||
| Inf (L63 vs. L72) | ND1, ND2, ND3, ND4, ND5, COX2, CYTB, ATP6 |
| Tissue | Comparison Subset | Up-Regulated Gene | Down-Regulated Gene |
|---|---|---|---|
| Bursa of Fabricius | L63 (Con vs. Inf) | SLC25A4 | POLG2 |
| L72 (Con vs. Inf) | SLC25A4 | DNA2, MGME1, TK2 | |
| Con (L63 vs. L72) | SLC25A4, MPV17 | ||
| Inf (L63 vs. L72) | SLC25A4, DNA2, TK2 | ||
| Spleen | L63 (Con vs. Inf) | POLG2 | |
| L72 (Con vs. Inf) | TWNK, SSBP1, DNA2, MGME1, SLC25A4 | ||
| Inf (L63 vs. L72) | TWNK, SSBP1, DNA2, MGME1, SLC25A4, TFAM, MTERF2, SUPV3L1 | POLG2 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chu, Q.; Ding, Y.; Cai, W.; Liu, L.; Zhang, H.; Song, J. Marek’s Disease Virus Infection Induced Mitochondria Changes in Chickens. Int. J. Mol. Sci. 2019, 20, 3150. https://doi.org/10.3390/ijms20133150
Chu Q, Ding Y, Cai W, Liu L, Zhang H, Song J. Marek’s Disease Virus Infection Induced Mitochondria Changes in Chickens. International Journal of Molecular Sciences. 2019; 20(13):3150. https://doi.org/10.3390/ijms20133150
Chicago/Turabian StyleChu, Qin, Yi Ding, Wentao Cai, Lei Liu, Huanmin Zhang, and Jiuzhou Song. 2019. "Marek’s Disease Virus Infection Induced Mitochondria Changes in Chickens" International Journal of Molecular Sciences 20, no. 13: 3150. https://doi.org/10.3390/ijms20133150
APA StyleChu, Q., Ding, Y., Cai, W., Liu, L., Zhang, H., & Song, J. (2019). Marek’s Disease Virus Infection Induced Mitochondria Changes in Chickens. International Journal of Molecular Sciences, 20(13), 3150. https://doi.org/10.3390/ijms20133150






