Myeloid-Specific Deletion of the AMPKα2 Subunit Alters Monocyte Protein Expression and Atherogenesis
Abstract
1. Introduction
2. Results
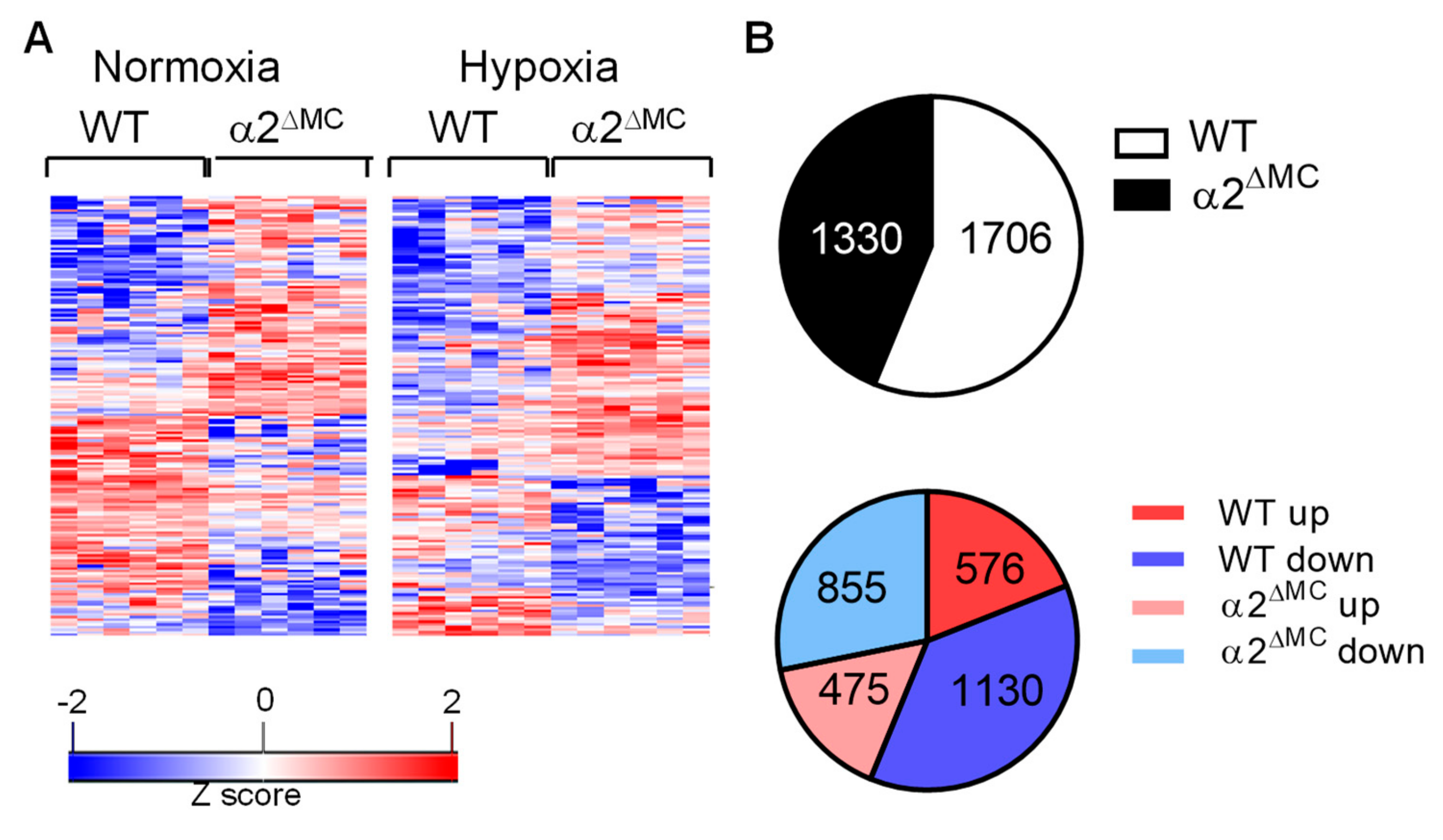
2.1. Consequences of the Myeloid-Specific Deletion of AMPKα2 on Monocyte Protein Expression
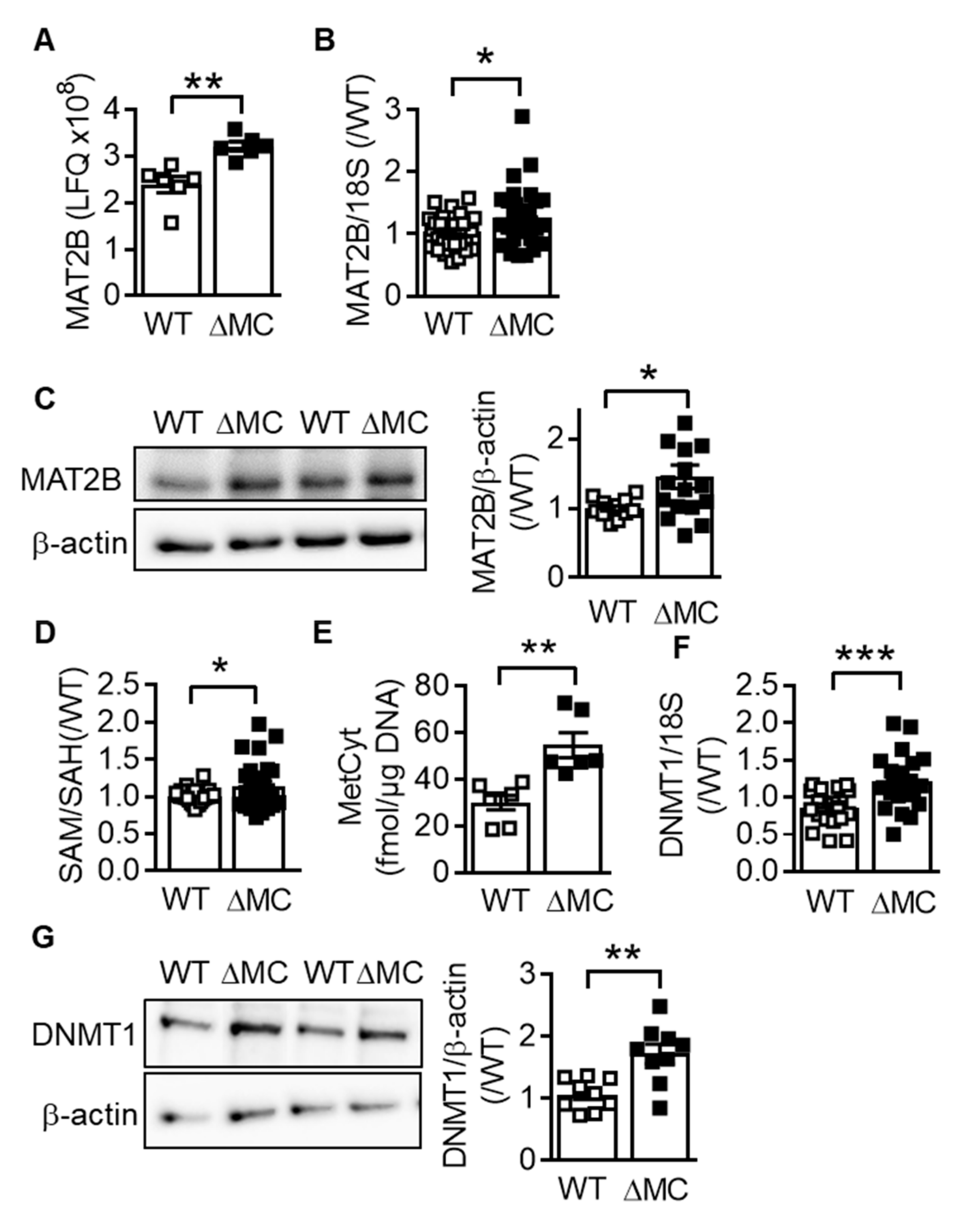
2.2. Consequences of Myeloid Cell AMPKα2 Deletion on the S-adenosyl Methionine Metabolism
2.3. Consequences of Myeloid Cell AMPKα2 Deletion on DNA Methylation
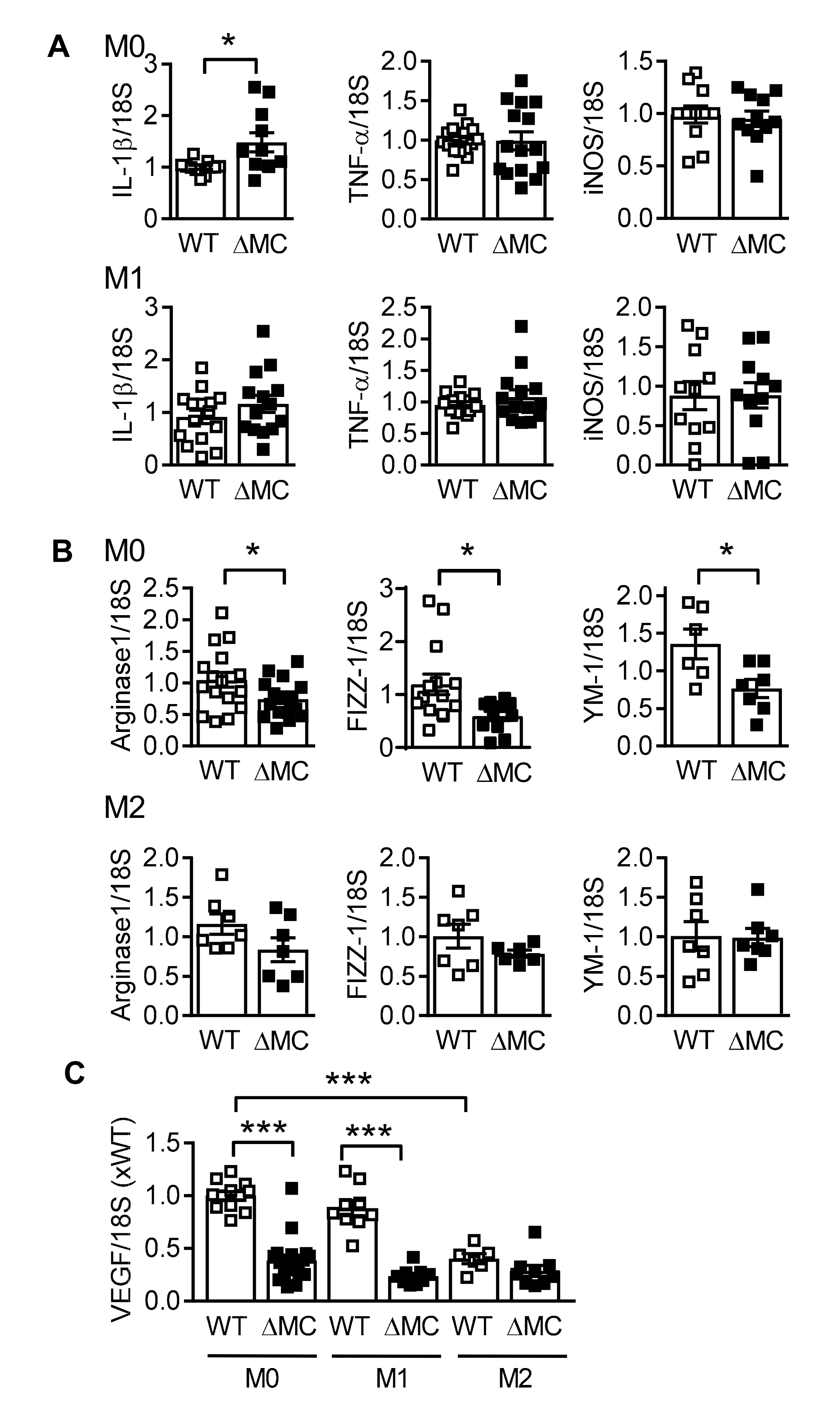
2.4. Consequence of AMPKα2 Deletion on Macrophage Polarization
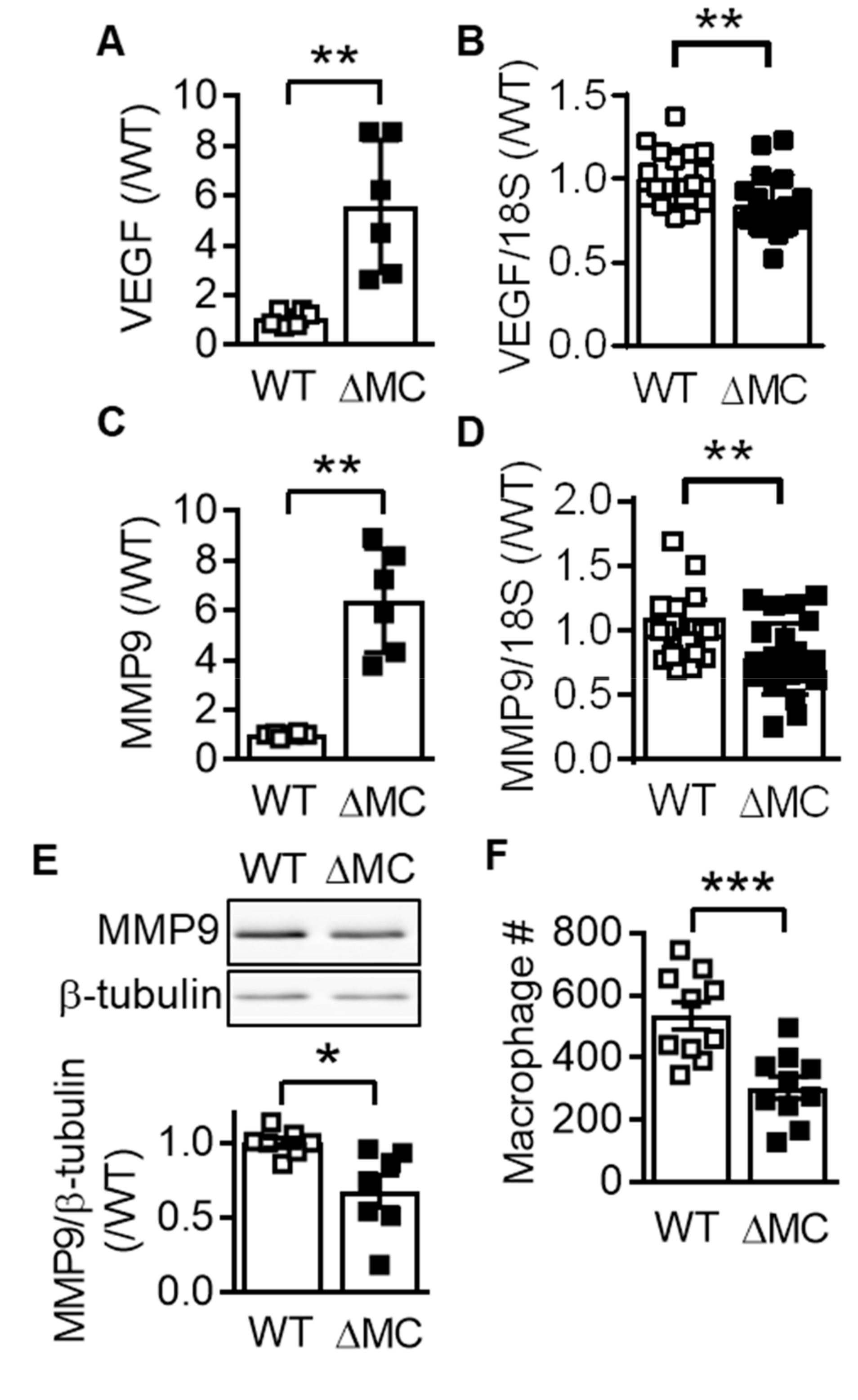
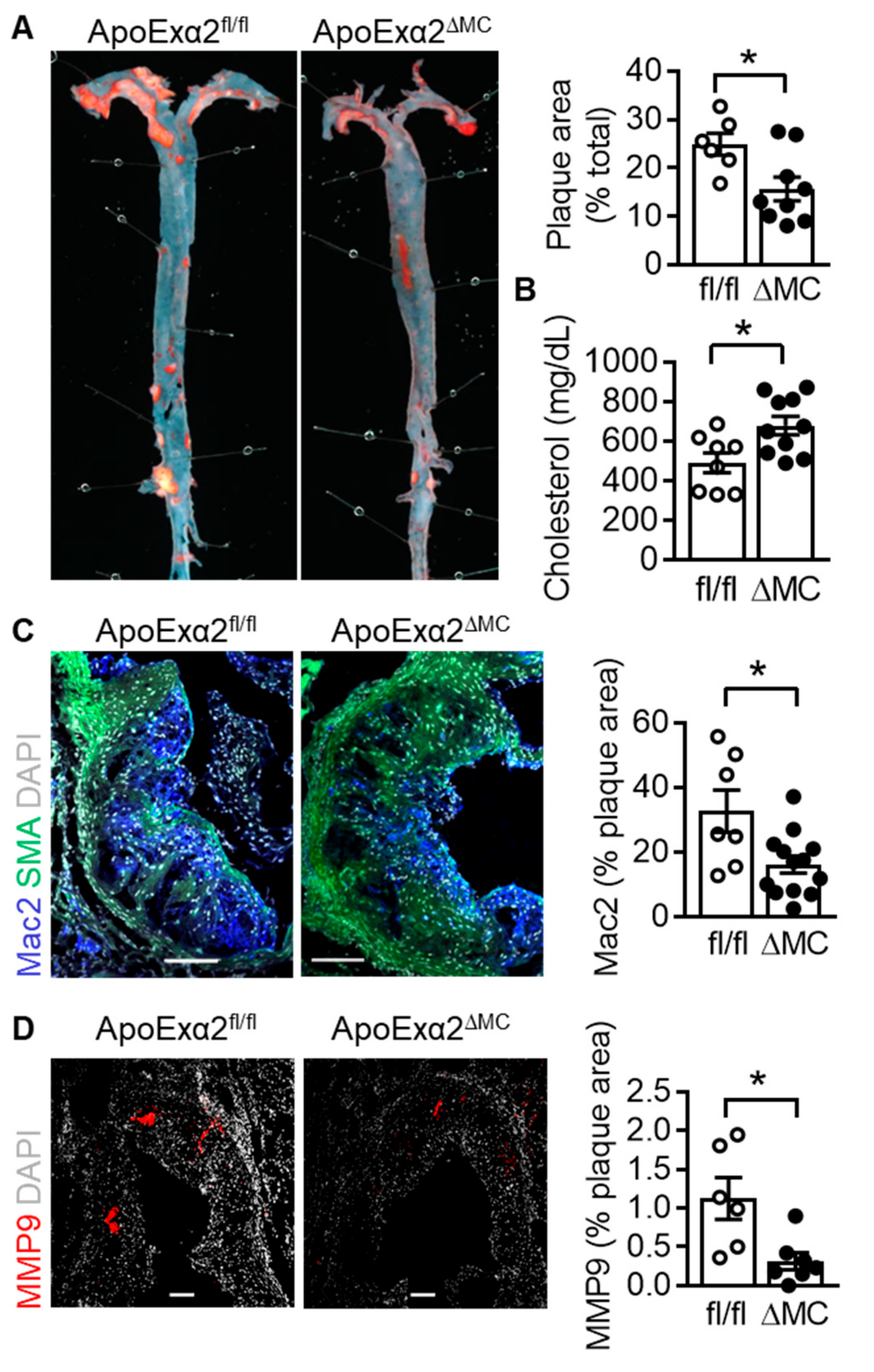
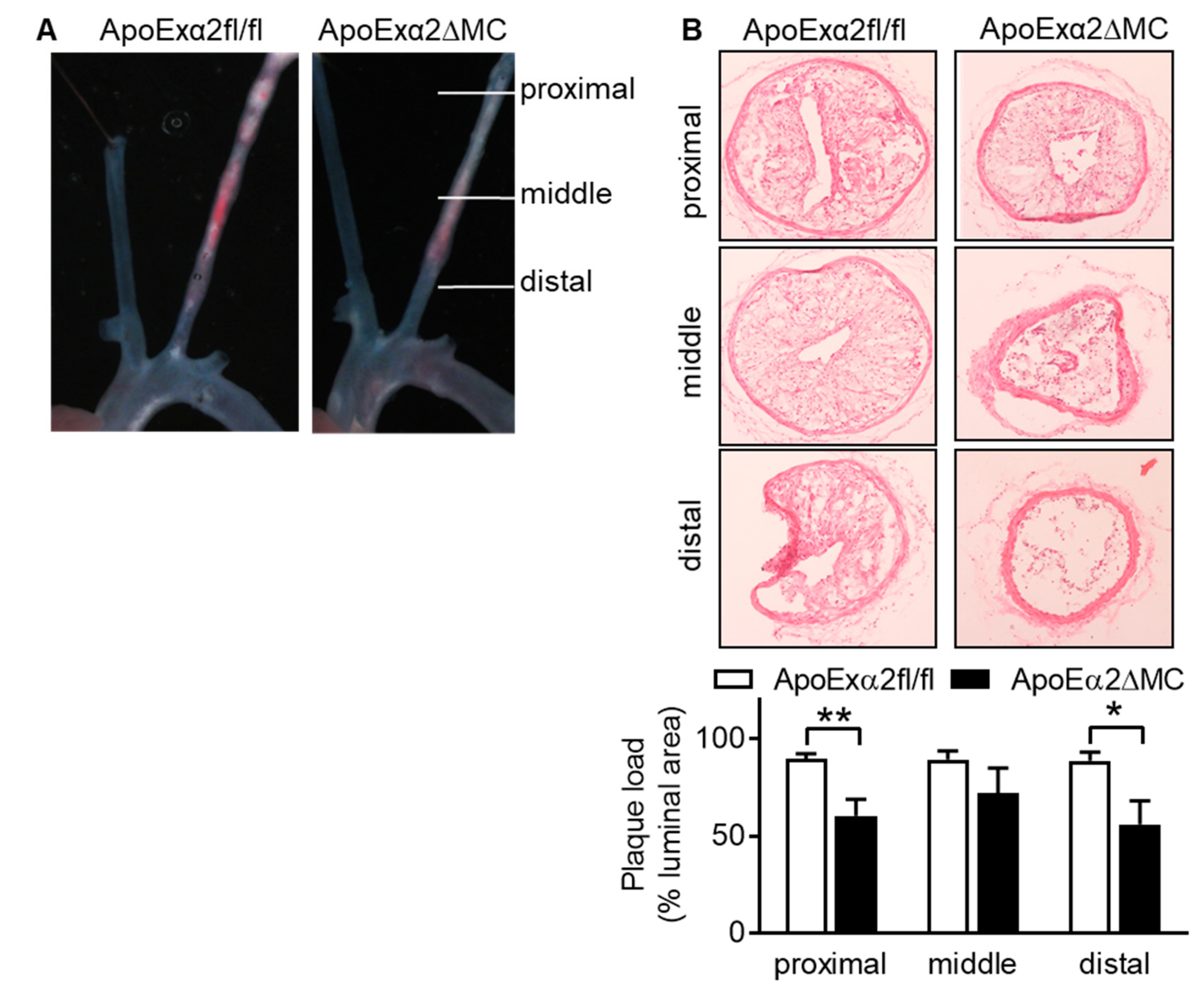
2.5. Effect of AMPKα2 Deletion in Myeloid Cells on Atherosclerotic Plaque Formation
3. Discussion
4. Materials and Methods
4.1. Materials
4.2. Animals
4.3. Partial Carotid Ligation
4.4. Western Diet
4.5. Histochemical Staining
4.6. RNA Isolation and Real Time Quantitative PCR (RT-qPCR)
4.7. DNA Methylation Quantification
4.8. Immunoblotting
4.9. Fluorescence-activated Cell Sorting
4.10. Isolation of Bone Marrow Cells
4.11. Proteomics: Sample Preparation
4.12. Proteomics: Liquid Chromatography/Mass Spectrometry (LC/MS)
4.13. Proteomics: MS Data Analysis
4.14. Metabolomics: Liquid Chromatography/Tandem Mass Spectrometry (LC-MS/MS)
4.15. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| AMPK | AMP-activated protein kinase |
| AMP | Adenosine monophosphate |
| ApoE | Apolipoprotein E |
| ATP | Adenosine triphosphate |
| CpG | Cytosine-guanine dinucleotide |
| DNMT | DNA methyltransferase |
| FACs | Fluorescence-activated cell sorting |
| FIZZ1 | Found in inflammatory zone 1 |
| IFN | Interferon |
| IL | Interleukin |
| iNOS | inducible nitric oxide synthase |
| LCA | Left carotid artery |
| LC/MS | Liquid chromatography/Mass spectrometery |
| LPS | Lipopolysaccharide |
| MAT2B | Methionine adenosyltransferase 2B |
| MMP | Matrix metalloproteinase |
| RCA | Right carotid artery |
| SAM | S-adenosyl methionine |
| SAH | S-adenosyl homocysteine |
| TNFα | Tumor necrosis factor alpha |
| VEGF | Vascular endothelial cell growth factor |
| YM1 | Chitinase 3 like protein 3 |
References
- Ross, F.A.; MacKintosh, C.; Hardie, D.G. AMP-activated protein kinase: A cellular energy sensor that comes in 12 flavours. FEBS J. 2016, 283, 2987–3001. [Google Scholar] [CrossRef] [PubMed]
- Hardie, D.G.; Lin, S.C. AMP-activated protein kinase-not just an energy sensor. F1000Research 2017, 6, 1724. [Google Scholar] [CrossRef] [PubMed]
- Foretz, M.; Guihard, S.; Leclerc, J.; Fauveau, V.; Couty, J.P.; Andris, F.; Gaudry, M.; Andreelli, F.; Vaulont, S.; Viollet, B. Maintenance of red blood cell integrity by AMP-activated protein kinase α1 catalytic subunit. FEBS Lett. 2010, 584, 3667–3671. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Dale, G.L.; Song, P.; Viollet, B.; Zou, M.H. AMPKα1 deletion shortens erythrocyte life span in mice. J. Biol. Chem. 2010, 285, 19976–19985. [Google Scholar] [CrossRef] [PubMed]
- Viollet, B.; Andreelli, F.; Jorgensen, S.B.; Perrin, C.; Geloen, A.; Flamez, D.; Mu, J.; Lenzner, C.; Baud, O.; Bennoun, M.; et al. The AMP-activated protein kinase α2 catalytic subunit controls whole-body insulin sensitivity. J. Clin. Investig. 2003, 111, 91–98. [Google Scholar] [CrossRef] [PubMed]
- Viollet, B.; Andreelli, F.; Jorgensen, S.B.; Perrin, C.; Flamez, D.; Mu, J.; Wojtaszewski, J.F.; Schuit, F.C.; Birnbaum, M.; Richter, E.; et al. Physiological role of AMP-activated protein kinase (AMPK): Insights from knockout mouse models. Biochem. Soc. Trans. 2003, 31, 216–219. [Google Scholar] [CrossRef]
- Salt, I.P.; Hardie, D.G. AMP-Activated protein kinase: An ubiquitous signaling pathway with key roles in the cardiovascular system. Circ. Res. 2017, 120, 1825. [Google Scholar] [CrossRef]
- Morrow, V.A.; Foufelle, F.; Connell, J.M.C.; Petrie, J.R.; Gould, G.W.; Salt, I.P. Direct activation of AMP-activated protein kinase stimulates nitric oxide synthesis in human aortic endothelial cells. J. Biol. Chem. 2003, 278, 31629–31639. [Google Scholar] [CrossRef]
- Stahmann, N.; Woods, A.; Spengler, K.; Heslegrave, A.; Bauer, R.; Krause, S.; Viollet, B.; Carling, D.; Heller, R. Activation of AMP-activated protein kinase by vascular endothelial growth factor mediates endothelial angiogenesis independent of nitric-oxide synthase. J. Biol. Chem. 2010, 285, 10638–10652. [Google Scholar] [CrossRef]
- Liu, C.; Liang, B.; Wang, Q.; Wu, J.; Zou, M.H. Activation of the AMP-activated protein kinase α1 alleviates endothelial cell apoptosis by increasing the expression of anti-apoptotic proteins BCL-2 and survivin. J. Biol. Chem. 2010, 285, 15346–16355. [Google Scholar] [CrossRef]
- Zippel, N.; Abdel Malik, R.; Frömel, T.; Popp, R.; Bess, E.; Strilic, B.; Wettschureck, N.; Fleming, I.; Fisslthaler, B. Transforming growth factor-β-activated kinase 1 regulates angiogenesis via AMP-activated protein kinase-α1 and redox balance in endothelial cells. Arterioscler. Thromb. Vasc. Biol. 2013, 33, 2792–2799. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Zhang, M.; Liang, B.; Xu, J.; Xie, Z.; Liu, C.; Viollet, B.; Yan, D.; Zou, M.H. AMPKa2 deletion causes aberrant expression and activation of NAD(P)H oxidase and consequent endothelial dysfunction in vivo. Role of 26S proteasomes. Circ. Res. 2010, 106, 1117–1128. [Google Scholar] [CrossRef] [PubMed]
- Bess, E.; Fisslthaler, B.; Frömel, T.; Fleming, I. Nitric oxide-induced activation of the AMP-activated protein kinase α2 subunit attenuates IκB kinase activity and inflammatory responses in endothelial cells. PLoS ONE 2011, 6, e20848. [Google Scholar] [CrossRef]
- Carroll, K.C.; Viollet, B.; Suttles, J. AMPKα1 deficiency amplifies proinflammatory myeloid APC activity and CD40 signaling. J. Leukoc. Biol. 2013, 94, 1113–1121. [Google Scholar] [CrossRef] [PubMed]
- Sag, D.; Carling, D.; Stout, R.D.; Suttles, J. Adenosine 5-monophosphate-activated protein kinase promotes macrophage polarization to an anti-inflammatory functional phenotype. J. Immunol. 2008, 181, 8633–8641. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.P.; Brown, J.R.; Sag, D.; Zhang, L.; Suttles, J. Adenosine 5-monophosphate-activated protein kinase regulates IL-10-mediated anti-inflammatory signaling pathways in macrophages. J. Immunol. 2015, 194, 584–594. [Google Scholar] [CrossRef] [PubMed]
- Kohlstedt, K.; Trouvain, C.; Namgaladze, D.; Fleming, I. Adipocyte-derived lipids increase angiotensin-converting enzyme (ACE) expression and modulate macrophage phenotype. Basic Res. Cardiol. 2011, 106, 205–215. [Google Scholar] [CrossRef]
- Abdel Malik, R.; Zippel, N.; Frömel, T.; Heidler, J.; Zukunft, S.; Walzog, B.; Ansari, N.; Pampaloni, F.; Wingert, S.; Rieger, M.A.; et al. AMP-activated protein kinase α2 in neutrophils regulates hypoxia-inducible factor-1α and a network of proteins affecting metabolism and vascular repair after ischemia. Circ. Res. 2017, 120, 99–109. [Google Scholar] [CrossRef]
- Igarashi, K.; Katoh, Y. Metabolic aspects of epigenome: Coupling of S-adenosylmethionine synthesis and gene regulation on chromatin by SAMIT module. Epigenetics Dev. Dis. 2013, 61, 105–118. [Google Scholar] [CrossRef]
- Carling, D.; Clarke, P.R.; Zammit, V.A.; Hardie, D.G. Purification and characterization of the AMP-activated protein kinase. Copurification of acetyl-CoA carboxylase kinase and 3-hydroxy-3-methylglutaryl-CoA reductase kinase activities. Eur. J. Biochem. 1989, 186, 129–136. [Google Scholar] [CrossRef]
- Fisslthaler, B.; Fleming, I.; Keserü, B.; Walsh, K.; Busse, R. Fluid shear stress and NO decrease the activity of the hydroxy-methylglutaryl coenzyme A reductase in endothelial cells via the AMP-activated protein kinase and FoxO1. Circ. Res. 2007, 100, e12–e21. [Google Scholar] [CrossRef] [PubMed]
- Nam, D.; Ni, C.W.; Rezvan, A.; Suo, J.; Budzyn, K.; Llanos, A.; Harrison, D.; Giddens, D.; Jo, H. Partial carotid ligation is a model of acutely induced disturbed flow, leading to rapid endothelial dysfunction and atherosclerosis. Am. J. Physiol. Heart Circ. Physiol. 2009, 297, H1535–H1543. [Google Scholar] [CrossRef] [PubMed]
- Cui, S.; Li, W.; Lv, X.; Wang, P.; Gao, Y.; Huang, G. Folic acid supplementation delays atherosclerotic lesion development by modulating Mcp1 and Vegf DNA methylation levels in vivo and in vitro. Int. J. Mol. Sci. 2017, 18, 990. [Google Scholar] [CrossRef]
- Marin, T.L.; Gongol, B.; Zhang, F.; Martin, M.; Johnson, D.A.; Xiao, H.; Wang, Y.; Subramaniam, S.; Chien, S.; Shyy, J.Y. AMPK promotes mitochondrial biogenesis and function by phosphorylating the epigenetic factors DNMT1, RBBP7, and HAT1. Sci. Signal 2017, 10, eaaf7478. [Google Scholar] [CrossRef] [PubMed]
- Palomer, X.; Salvado, L.; Barroso, E.; Vazquez-Carrera, M. An overview of the crosstalk between inflammatory processes and metabolic dysregulation during diabetic cardiomyopathy. Int. J. Cardiol. 2013, 168, 3160–3172. [Google Scholar] [CrossRef] [PubMed]
- Ouimet, M.; Ediriweera, H.N.; Gundra, U.M.; Sheedy, F.J.; Ramkhelawon, B.; Hutchison, S.B.; Rinehold, K.; van Solingen, S.C.; Fullerton, M.D.; Cecchini, K.; et al. MicroRNA-33-dependent regulation of macrophage metabolism directs immune cell polarization in atherosclerosis. J. Clin. Investig. 2015, 125, 4334–4348. [Google Scholar] [CrossRef] [PubMed]
- Price, N.L.; Rotllan, N.; Canfran-Duque, A.; Zhang, X.; Pati, P.; Arias, N.; Moen, J.; Mayr, M.; Ford, D.A.; Baldan, A.; et al. Genetic dissection of the impact of miR-33a and miR-33b during the progression of atherosclerosis. Cell Rep. 2017, 21, 1317–1330. [Google Scholar] [CrossRef]
- Ohki, Y.; Heissig, B.; Sato, Y.; Akiyama, H.; Zhu, Z.; Hicklin, D.J.; Shimada, K.; Ogawa, H.; Daida, H.; Hattori, K.; et al. Granulocyte colony-stimulating factor promotes neovascularization by releasing vascular endothelial growth factor from neutrophils. FASEB J. 2005, 19, 2005–2007. [Google Scholar] [CrossRef]
- Döring, Y.; Drechsler, M.; Wantha, S.; Kemmerich, K.; Lievens, D.; Vijayan, S.; Gallo, R.L.; Weber, C.; Soehnlein, O. Lack of neutrophil-derived CRAMP reduces atherosclerosis in mice. Circ. Res. 2012, 110, 1052–1056. [Google Scholar] [CrossRef]
- Wantha, S.; Alard, J.E.; Megens, R.T.A.; van der Does, A.M.; Döring, Y.; Drechsler, M.; Pham, C.T.N.; Wang, M.W.; Wang, J.M.; Gallo, R.L.; et al. Neutrophil-derived cathelicidin promotes adhesion of classical monocytes. Circ. Res. 2013, 112, 792–801. [Google Scholar] [CrossRef]
- Zhu, H.; Zhang, M.; Liu, Z.; Xing, J.; Moriasi, C.; Dai, X.; Zou, M.H. AMP-Activated protein kinase α1 in macrophages promotes collateral remodeling and arteriogenesis in mice in vivo. Arterioscler. Thromb. Vasc. Biol. 2016, 36, 1868–1878. [Google Scholar] [CrossRef] [PubMed]
- Dong, Y.; Zhang, M.; Liang, B.; Xie, Z.; Zhao, Z.; Asfa, S.; Choi, H.C.; Zou, M.H. Reduction of AMP-activated protein kinase α2 increases endoplasmic reticulum stress and atherosclerosis in vivo. Circulation 2010, 121, 792–803. [Google Scholar] [CrossRef] [PubMed]
- Ding, Y.; Zhang, M.; Zhang, W.; Lu, Q.; Cai, Z.; Song, P.; Okon, I.S.; Xiao, L.; Zou, M.H. AMP-Activated protein kinase alpha 2 deletion induces VSMC phenotypic switching and reduces features of atherosclerotic plaque stability. Circ. Res. 2016, 119, 718–730. [Google Scholar] [CrossRef] [PubMed]
- Cai, Z.; Ding, Y.; Zhang, M.; Lu, Q.; Wu, S.; Zhu, H.; Song, P.; Zou, M.H. Ablation of adenosine monophosphate-activated protein kinase α1 in vascular smooth muscle cells promotes diet-induced atherosclerotic calcification in vivo. Circ. Res. 2016, 119, 422–433. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Zhu, H.; Ding, Y.; Liu, Z.; Cai, Z.; Zou, M.H. AMP-activated protein kinase α1 promotes atherogenesis by increasing monocyte-to-macrophage differentiation. J. Biol. Chem. 2017, 292, 7888–7903. [Google Scholar] [CrossRef] [PubMed]
- Kohlstedt, K.; Trouvain, C.; Boettger, T.; Shi, L.; Fisslthaler, B.; Fleming, I. AMP-activated protein kinase regulates endothelial cell angiotensin-converting enzyme expression via p53 and the post-transcriptional regulation of microRNA-143/145. Circ. Res. 2013, 112, 1150–1158. [Google Scholar] [CrossRef]
- Gongol, B.; Sari, I.; Bryant, T.; Rosete, G.; Marin, T. AMPK: An epigenetic landscape modulator. Int. J. Mol. Sci. 2018, 19, 3238. [Google Scholar] [CrossRef]
- Fleming, I.; Fisslthaler, B.; Dixit, M.; Busse, R. Role of PECAM-1 in the shear-stress-induced activation of Akt and the endothelial nitric oxide synthase (eNOS) in endothelial cells. J. Cell Sci. 2005, 118, 4103–4111. [Google Scholar] [CrossRef]
- Galkina, E.; Kadl, A.; Sanders, J.; Varughese, D.; Sarembock, I.J.; Ley, K. Lymphocyte recruitment into the aortic wall before and during development of atherosclerosis is partially L-selectin dependent. J. Exp. Med. 2006, 203, 1273–1282. [Google Scholar] [CrossRef]
- Wisniewski, J.R.; Zougman, A.; Nagaraj, N.; Mann, M. Universal sample preparation method for proteome analysis. Nat. Methods 2009, 6, 359–362. [Google Scholar] [CrossRef]
- Rappsilber, J.; Mann, M.; Ishihama, Y. Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nat. Protoc. 2007, 2, 1896–1906. [Google Scholar] [CrossRef] [PubMed]
- Tyanova, S.; Temu, T.; Cox, J. The MaxQuant computational platform for mass spectrometry-based shotgun proteomics. Nat. Protoc. 2016, 11, 2301–2319. [Google Scholar] [CrossRef] [PubMed]
- Tyanova, S.; Temu, T.; Sinitcyn, P.; Carlson, A.; Hein, M.Y.; Geiger, T.; Mann, M.; Cox, J. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 2016, 13, 731–740. [Google Scholar] [CrossRef] [PubMed]
| Target mRNA | Forward Primer | Reverse Primer |
|---|---|---|
| 18S RNA | 5′-CTTTGGTCGCTCGCTCCTC-3′ | 5′-CTGACCGGGTTGGTTTTGAT-3′ |
| VEGF | 5′-GCACTGGACCCTGGCTTTACTGCTGTA-3′ | 5′-GAACTTGATCACTTCATGGGACTTCTGCTC-3′ |
| MMP9 | 5′-GAAGGCAAACCCTGTGTT-3′ | 5′-AGAGTACTGCTTGCCCAGGA-3′ |
| Mat2B | 5′-CAGAGGTTCCCCACACATGT-3′ | 5′-GGGGAGGTTGAAGGCATCTG-3′ |
| DNMT1 | 5′-AAGGGGGCCCTGACCGCTTC-3′ | 5′-CCGAAATGCCTGGGCTGCCG-3′ |
| IL-1β | 5′-CAGGCAGGCAGTATCACTCA-3′ | 5′-AGCTCATATGGGTCCGACAG-3′ |
| TNF-α | 5′-GGCCTTCCTACCTTCAGACC-3′ | 5′-CCGGCCTTCCAAATAAATAC-3′ |
| iNOS | 5′-GTGGTGACAAGCACATTTGG-3′ | 5′-GGCCTTCCTACCTTCAGACC-3′ |
| Arginase | 5′-GTGAAGAACCCACGGTCTGT-3′ | 5′-CTGGTTGTCAGGGGAGTGTT-3′ |
| FIZZ1 | 5′-CCCTTCTCATCTGCATCTCC-3′ | 5′-CAGTAGCAGTCATCCCAGCA-3′ |
| YM1 | 5′-CTGGAATTGGTGCCCCTACAA-3′ | 5′-TCATAACCAACCCACTCATTACC-3′ |
| Target Region | Forward Primer | Reverse Primer |
|---|---|---|
| Vegfa | 5′-CTCCTCTCCCTTCTGGAACC-3′ | 5′-GAGGGGAGGAAGAGAAGGAA-3′ |
| Mmp9 | 5′-CAATCCCTAGTCGCTGCTTC-3′ | 5′-AGGAAGGGACTCAATCAGCA-3′ |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fisslthaler, B.; Zippel, N.; Abdel Malik, R.; Delgado Lagos, F.; Zukunft, S.; Thoele, J.; Siuda, D.; Soehnlein, O.; Wittig, I.; Heidler, J.; et al. Myeloid-Specific Deletion of the AMPKα2 Subunit Alters Monocyte Protein Expression and Atherogenesis. Int. J. Mol. Sci. 2019, 20, 3005. https://doi.org/10.3390/ijms20123005
Fisslthaler B, Zippel N, Abdel Malik R, Delgado Lagos F, Zukunft S, Thoele J, Siuda D, Soehnlein O, Wittig I, Heidler J, et al. Myeloid-Specific Deletion of the AMPKα2 Subunit Alters Monocyte Protein Expression and Atherogenesis. International Journal of Molecular Sciences. 2019; 20(12):3005. https://doi.org/10.3390/ijms20123005
Chicago/Turabian StyleFisslthaler, Beate, Nina Zippel, Randa Abdel Malik, Fredy Delgado Lagos, Sven Zukunft, Janina Thoele, Daniel Siuda, Oliver Soehnlein, Ilka Wittig, Juliana Heidler, and et al. 2019. "Myeloid-Specific Deletion of the AMPKα2 Subunit Alters Monocyte Protein Expression and Atherogenesis" International Journal of Molecular Sciences 20, no. 12: 3005. https://doi.org/10.3390/ijms20123005
APA StyleFisslthaler, B., Zippel, N., Abdel Malik, R., Delgado Lagos, F., Zukunft, S., Thoele, J., Siuda, D., Soehnlein, O., Wittig, I., Heidler, J., Weigert, A., & Fleming, I. (2019). Myeloid-Specific Deletion of the AMPKα2 Subunit Alters Monocyte Protein Expression and Atherogenesis. International Journal of Molecular Sciences, 20(12), 3005. https://doi.org/10.3390/ijms20123005






