Jasmonic Acid-Induced VQ-Motif-Containing Protein OsVQ13 Influences the OsWRKY45 Signaling Pathway and Grain Size by Associating with OsMPK6 in Rice
Abstract
1. Introduction
2. Results
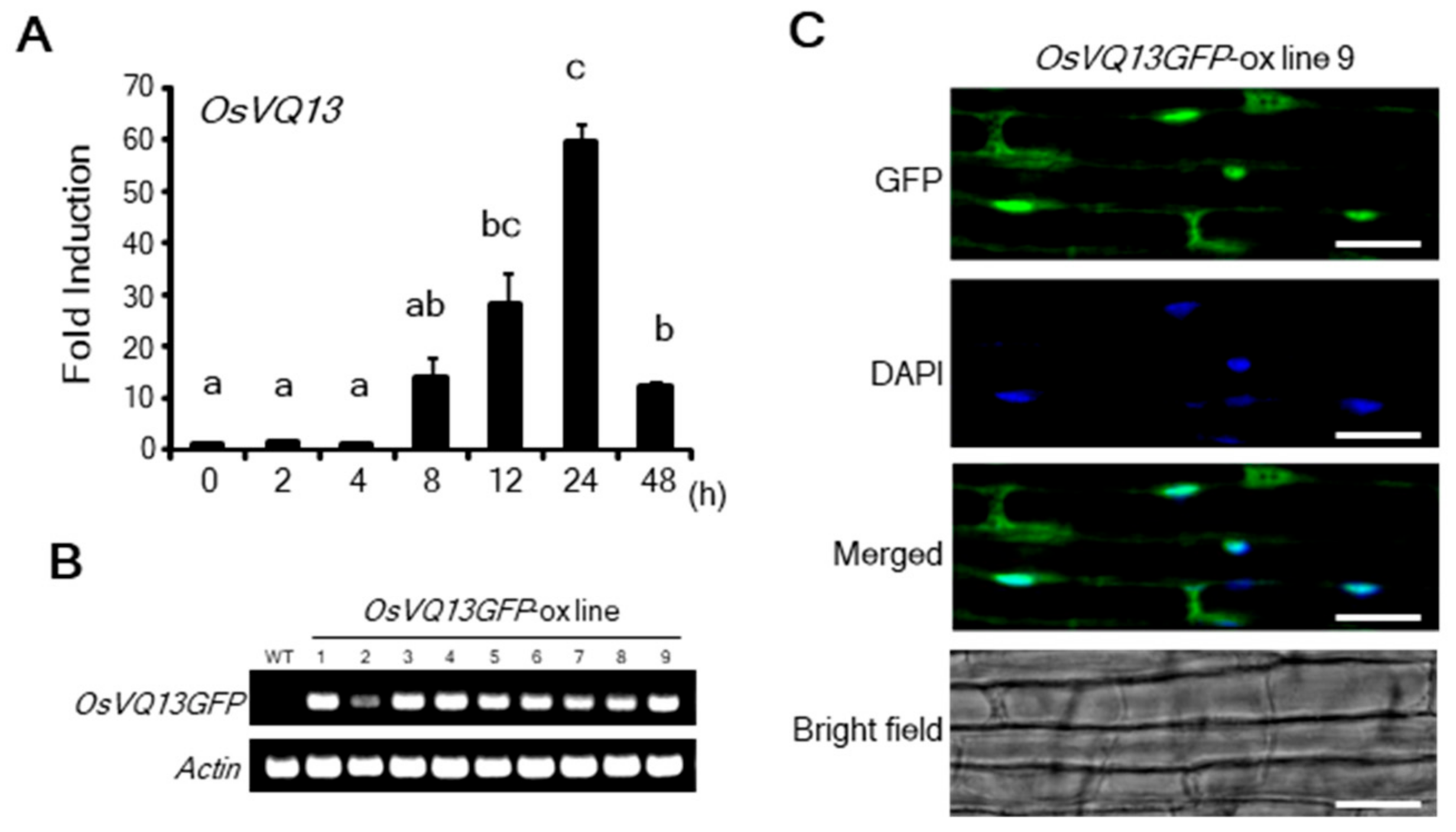
2.1. Properties of OsVQ13
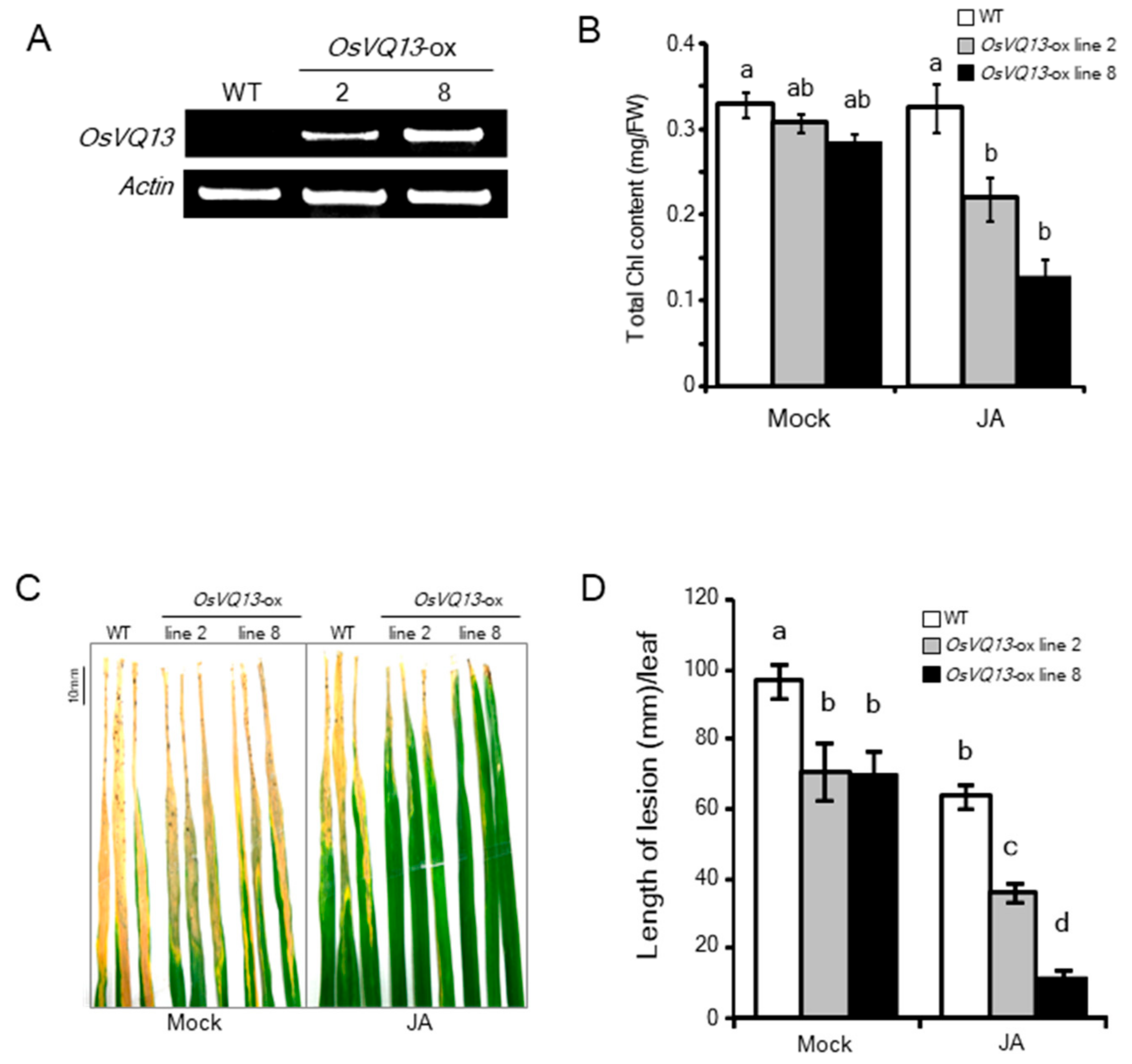
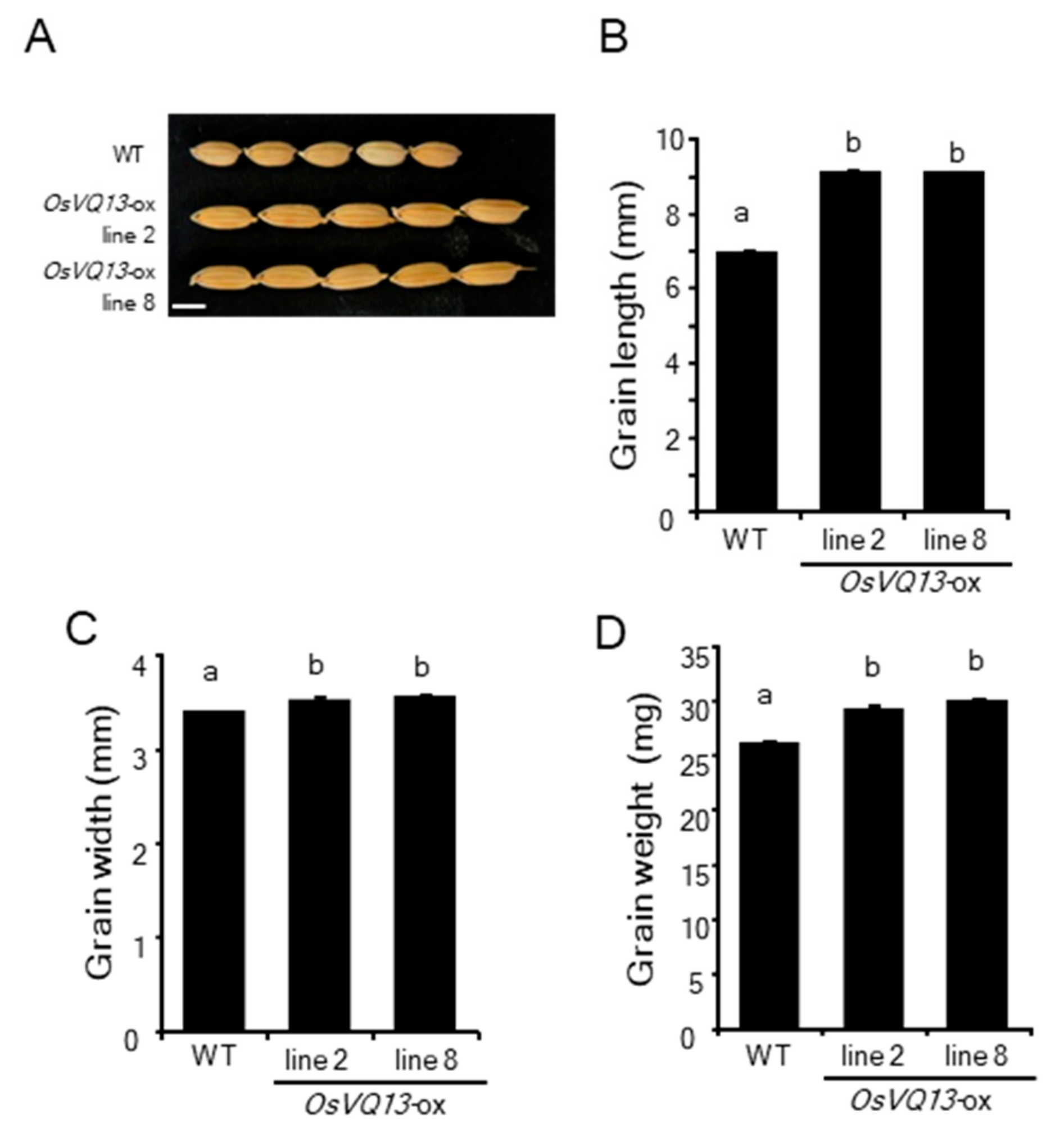
2.2. Phenotypes of OsVQ13-Overexpressing Rice Plants
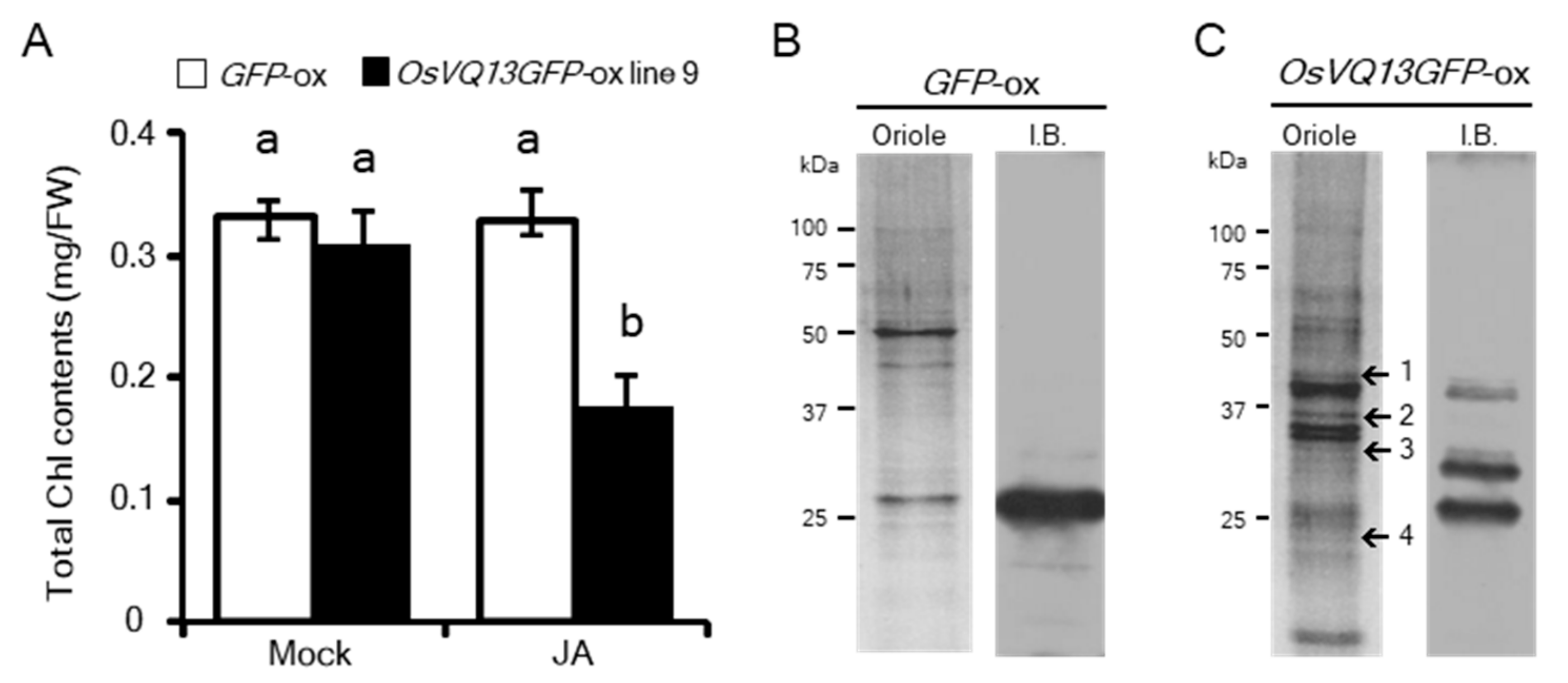
2.3. Identification of OsVQ13-Associating Proteins
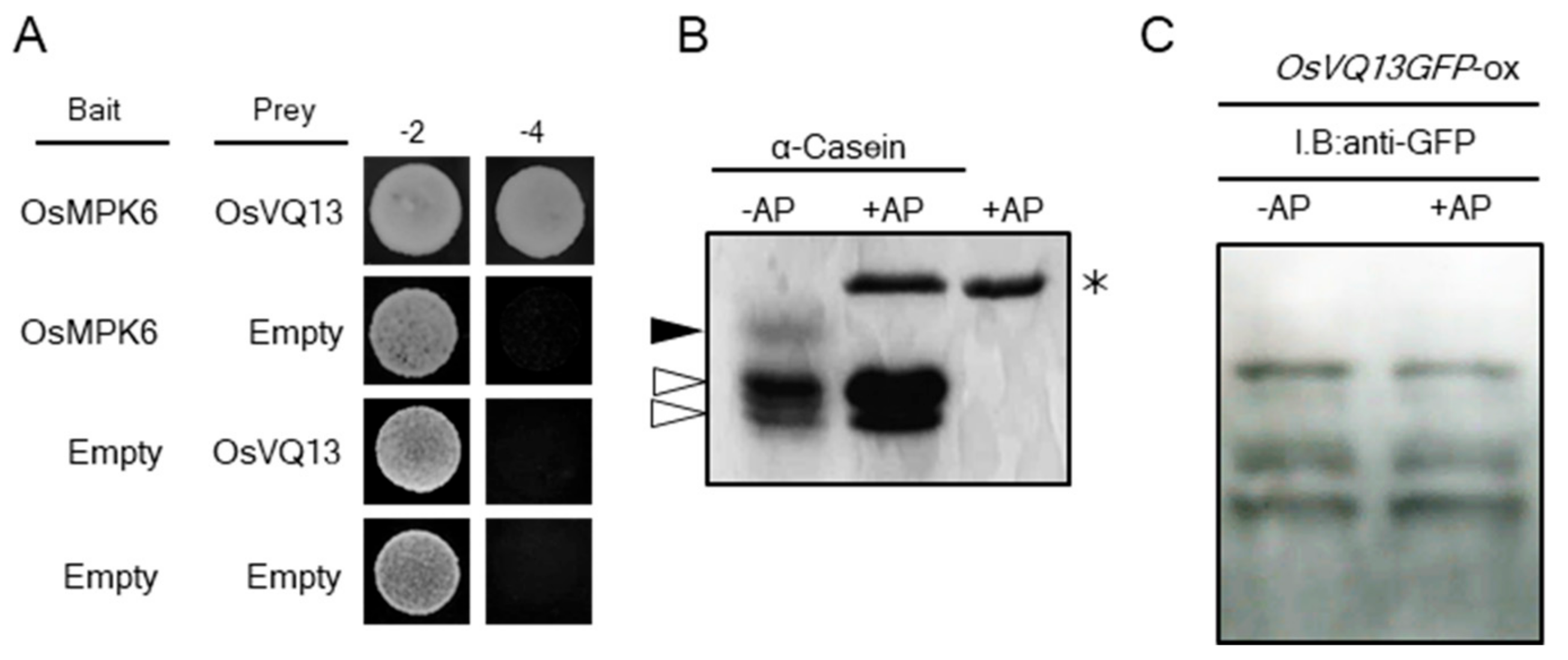
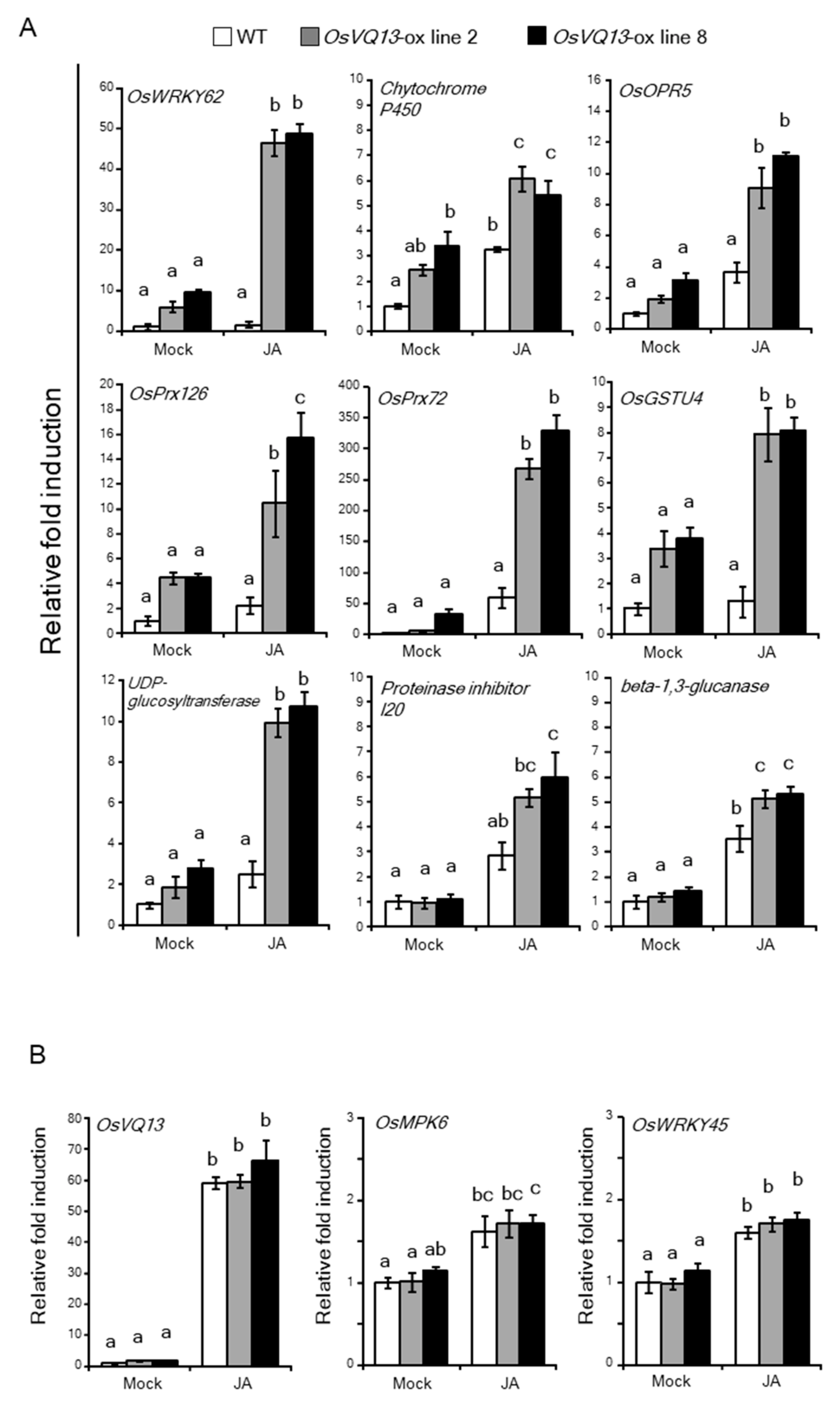
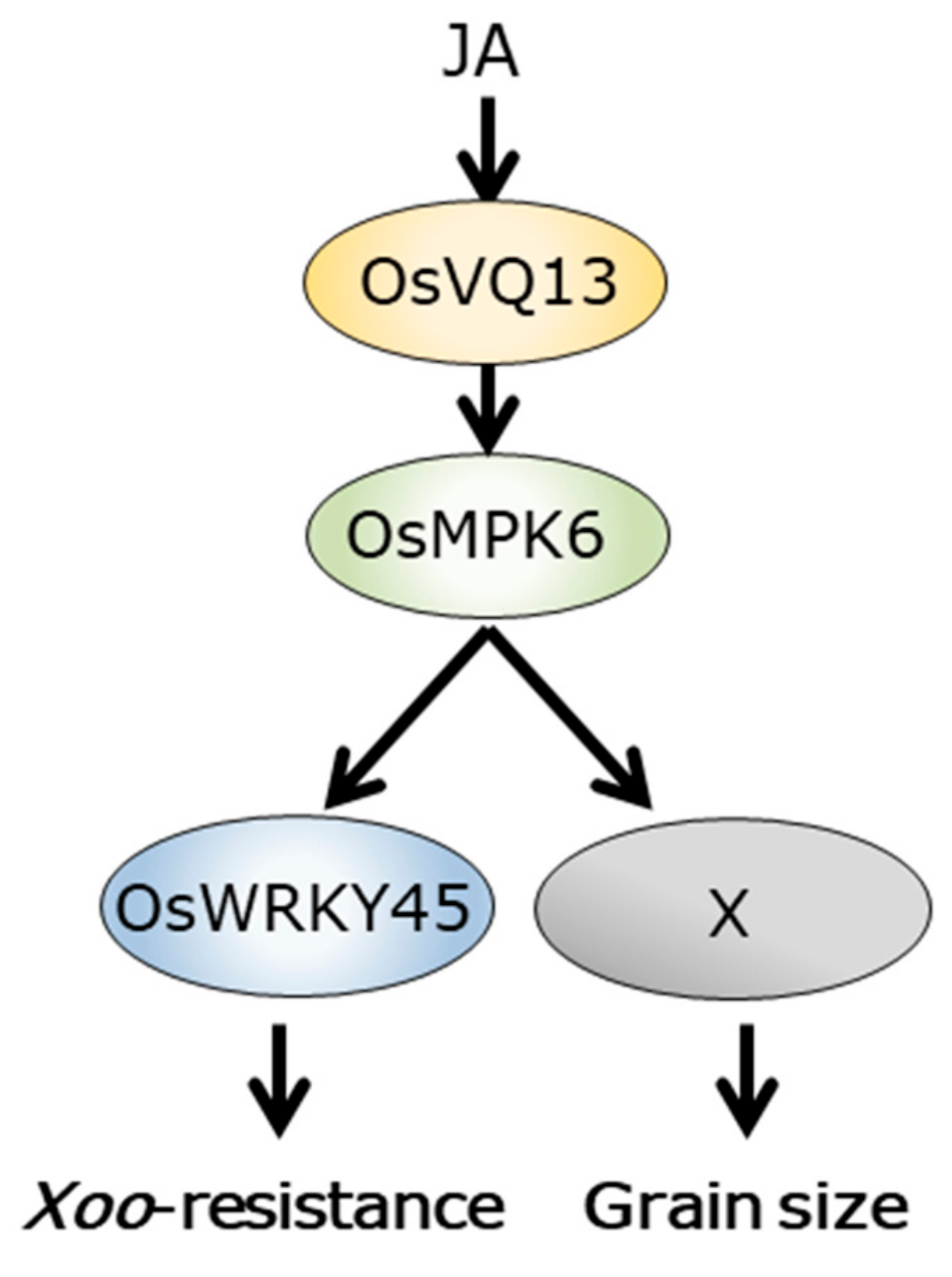
2.4. OsVQ13 Affected OsMPK6-Mediated Signaling Pathways in Rice
3. Discussion
4. Materials and Methods
4.1. Plant Materials, Jasmonate Treatment, and Bacterial Inoculation
4.2. RT-qPCR
4.3. Construction of OsVQ13-Overexpressing Vectors and Rice Transformation
4.4. Chlorophyll Content Measurement
4.5. Subcellular Localization Assay
4.6. Identification of OsVQ13-Associating Proteins
4.6.1. In Vivo Purification of the OsVQ13GFP Protein
4.6.2. Co-Immunoprecipitation and Protein Gel Blot Analysis
4.6.3. Protein Identification Using TOF MS
4.6.4. Phos-tag® SDS-PAGE
4.7. Yeast Two-Hybrid System
4.8. Additional Information
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- De Vleesschauwer, D.; Gheysen, G.; Höfte, M. Hormone defense networking in rice: Tales from a different world. Trends Plant Sci. 2013, 18, 555–565. [Google Scholar] [CrossRef]
- Shimono, M.; Sugano, S.; Nakayama, A.; Jiang, C.J.; Ono, K.; Toki, S.; Takatsuji, H. Rice WRKY45 plays a crucial role in benzothiadiazole-inducible blast resistance. Plant Cell 2007, 19, 2064–2076. [Google Scholar] [CrossRef]
- Bakshi, M.; Oelmüller, R. WRKY transcription factors: Jack of many trades in plants. Plant Signal. Behav. 2014, 9, e27700. [Google Scholar] [CrossRef]
- Nakayama, A.; Fukushima, S.; Goto, S.; Matsushita, A.; Shimono, M.; Sugano, S.; Jiang, C.J.; Akagi, A.; Yamazaki, M.; Inoue, H.; et al. Genome-wide identification of WRKY45-regulated genes that mediate benzothiadiazole-induced defense responses in rice. BMC Plant Biol. 2013, 13, 150. [Google Scholar] [CrossRef]
- Ding, Y.; Sun, T.; Ao, K.; Peng, Y.; Zhang, Y.; Li, X.; Zhang, Y. Opposite roles of salicylic acid receptors NPR1 and NPR3/NPR4 in transcriptional regulation of plant immunity. Cell 2018, 173, 1454–1467.e15. [Google Scholar] [CrossRef]
- Chern, M.; Fitzgerald, H.A.; Canlas, P.E.; Navarre, D.A.; Ronald, P.C. Overexpression of a rice NPR1 homolog leads to constitutive activation of defense response and hypersensitivity to light. Mol. Plant Microbe Interact. 2005, 18, 511–520. [Google Scholar] [CrossRef]
- Sugano, S.; Jiang, C.J.; Miyazawa, S.I.; Masumoto, C.; Yazawa, K.; Hayashi, N.; Shimono, M.; Nakayama, A.; Miyao, M.; Takatsuji, H. Role of OsNPR1 in rice defense program as revealed by genome-wide expression analysis. Plant Mol. Biol. 2010, 74, 549–562. [Google Scholar] [CrossRef]
- Liu, Q.; Ning, Y.; Zhang, Y.; Yu, N.; Zhao, C.; Zhan, X.; Wu, W.; Chen, D.; Wei, X.; Wang, G.L.; et al. OsCUL3a negatively regulates cell death and immunity by degrading OsNPR1 in rice. Plant Cell 2017, 29, 345–359. [Google Scholar] [CrossRef]
- Ranjan, A.; Vadassery, J.; Patel, H.K.; Pandey, A.; Palaparthi, R.; Mithöfer, A.; Sonti, R.V. Upregulation of jasmonate biosynthesis and jasmonate-responsive genes in rice leaves in response to a bacterial pathogen mimic. Funct. Integr. Genom. 2015, 15, 363–373. [Google Scholar] [CrossRef]
- Yamada, S.; Kano, A.; Tamaoki, D.; Miyamoto, A.; Shishido, H.; Miyoshi, S.; Taniguchi, S.; Akimitsu, K.; Gomi, K. Involvement of OsJAZ8 in jasmonate-induced resistance to bacterial blight in rice. Plant Cell Physiol. 2012, 53, 2060–2072. [Google Scholar] [CrossRef]
- Tao, Z.; Liu, H.; Qiu, D.; Zhou, Y.; Li, X.; Xu, C.; Wang, S. A pair of allelic WRKY genes play opposite roles in rice-bacteria interactions. Plant Physiol. 2009, 151, 936–948. [Google Scholar] [CrossRef]
- Deng, H.; Liu, H.; Li, X.; Xiao, J.; Wang, S. A CCCH-type zinc finger nucleic acid-binding protein quantitatively confers resistance against rice bacterial blight disease. Plant Physiol. 2012, 158, 876–889. [Google Scholar] [CrossRef]
- Uji, Y.; Taniguchi, S.; Tamaoki, D.; Shishido, H.; Akimitsu, K.; Gomi, K. Overexpression of OsMYC2 results in the up-regulation of early JA-rresponsive genes and bacterial blight resistance in rice. Plant Cell Physiol. 2016, 57, 1814–1827. [Google Scholar] [CrossRef]
- Pauwels, L.; Barbero, G.F.; Geerinck, J.; Tilleman, S.; Grunewald, W.; Pérez, A.C.; Chico, J.M.; Bossche, R.V.; Sewell, J.; Gil, E.; et al. NINJA connects the co-repressor TOPLESS to jasmonate signalling. Nature 2010, 464, 788–791. [Google Scholar] [CrossRef]
- Kashihara, K.; Onohata, T.; Okamoto, Y.; Uji, Y.; Mochizuki, S.; Akimitsu, K.; Gomi, K. Overexpression of OsNINJA1 negatively affects a part of OsMYC2-mediated abiotic and biotic responses in rice. J. Plant Physiol. 2019, 232, 180–187. [Google Scholar] [CrossRef]
- Tanaka, K.; Taniguchi, S.; Tamaoki, D.; Yoshitomi, K.; Akimitsu, K.; Gomi, K. Multiple roles of plant volatiles in jasmonate-induced defense response in rice. Plant Signal. Behav. 2014, 9, e29247. [Google Scholar] [CrossRef]
- Taniguchi, S.; Hosokawa-Shinonaga, Y.; Tamaoki, D.; Yamada, S.; Akimitsu, K.; Gomi, K. Jasmonate induction of the monoterpene linalool confers resistance to rice bacterial blight and its biosynthesis is regulated by JAZ protein in rice. Plant Cell Environ. 2014, 37, 451–461. [Google Scholar] [CrossRef]
- Taniguchi, S.; Miyoshi, S.; Tamaoki, D.; Yamada, S.; Tanaka, K.; Uji, Y.; Tanaka, S.; Akimitsu, K.; Gomi, K. Isolation of jasmonate-induced sesquiterpene synthase of rice: Product of which has an antifungal activity against Magnaporthe oryzae. J. Plant Physiol. 2014, 171, 625–632. [Google Scholar] [CrossRef]
- Yoshitomi, K.; Taniguchi, S.; Tanaka, K.; Uji, Y.; Akimitsu, K.; Gomi, K. Rice terpene synthase 24 (OsTPS24) encodes a jasmonate-responsive monoterpene synthase that produces an antibacterial γ-terpinene against rice pathogen. J. Plant Physiol. 2016, 191, 120–126. [Google Scholar] [CrossRef]
- Kiryu, M.; Hamanaka, M.; Yoshitomi, K.; Mochizuki, S.; Akimitsu, K.; Gomi, K. Rice terpene synthase 18 (OsTPS18) encodes a sesquiterpene synthase that produces an antibacterial (E)-nerolidol against a bacterial pathogen of rice. J. Gen. Plant Pathol. 2018, 84, 221–229. [Google Scholar] [CrossRef]
- Cheng, Y.; Zhou, Y.; Yang, Y.; Chi, Y.-J.; Zhou, J.; Chen, J.Y.; Wang, F.; Fan, B.; Shi, K.; Zhou, Y.H.; et al. Structural and functional analysis of VQ motif-containing proteins in Arabidopsis as interacting proteins of WRKY transcription factors. Plant Physiol. 2012, 159, 810–825. [Google Scholar] [CrossRef]
- Kim, D.Y.; Kwon, S.I.; Choi, C.; Lee, H.; Ahn, I.; Park, S.R.; Bae, S.C.; Lee, S.C.; Hwang, D.J. Expression analysis of rice VQ genes in response to biotic and abiotic stresses. Gene 2013, 529, 208–214. [Google Scholar] [CrossRef]
- Wang, X.; Zhang, H.; Sun, G.; Jin, Y.; Qiu, L. Identification of active VQ motif-containing genes and the expression patterns under low nitrogen treatment in soybean. Gene 2014, 543, 237–243. [Google Scholar] [CrossRef]
- Song, W.; Zhao, H.; Zhang, X.; Lei, L.; Lai, J. Genome-wide identification of VQ motif-containing proteins and their expression profiles under abiotic stresses in maize. Front. Plant Sci. 2016, 6. [Google Scholar] [CrossRef]
- Hu, P.; Zhou, W.; Cheng, Z.; Fan, M.; Wang, L.; Xie, D. JAV1 controls jasmonate-regulated plant defense. Mol. Cell 2013, 50, 504–515. [Google Scholar] [CrossRef]
- Andreasson, E.; Jenkins, T.; Brodersen, P.; Thorgrimsen, S.; Petersen, N.H.T.; Zhu, S.; Qiu, J.L.; Micheelsen, P.; Rocher, A.; Petersen, M.; et al. The MAP kinase substrate MKS1 is a regulator of plant defense responses. EMBO J. 2005, 24, 2579–2589. [Google Scholar] [CrossRef]
- Petersen, K.; Qiu, J.L.; Lütje, J.; Fiil, B.K.; Hansen, S.; Mundy, J.; Petersen, M. Arabidopsis MKS1 is involved in basal immunity and requires an intact N-terminal domain for proper function. PLoS ONE 2010, 5, e14364. [Google Scholar] [CrossRef]
- Qiu, J.L.; Fiil, B.K.; Petersen, K.; Nielsen, H.B.; Botanga, C.J.; Thorgrimsen, S.; Palma, K.; Suarez-Rodriguez, M.C.; Sandbech-Clausen, S.; Lichota, J.; et al. Arabidopsis MAP kinase 4 regulates gene expression through transcription factor release in the nucleus. EMBO J. 2008, 27, 2214–2221. [Google Scholar] [CrossRef]
- Birkenbihl, R.P.; Diezel, C.; Somssich, I.E. Arabidopsis WRKY33 is a key transcriptional regulator of hormonal and metabolic responses toward Botrytis cinerea infection. Plant Physiol. 2012, 159, 266–285. [Google Scholar] [CrossRef]
- Li, N.; Li, X.; Xiao, J.; Wang, S. Comprehensive analysis of VQ motif-containing gene expression in rice defense responses to three pathogens. Plant Cell Rep. 2014, 33, 1493–1505. [Google Scholar] [CrossRef]
- Jing, Y.; Lin, R. The VQ motif-containing protein family of plant-specific transcriptional regulators. Plant Physiol. 2015, 169, 371–378. [Google Scholar] [CrossRef]
- Kishi-Kaboshi, M.; Okada, K.; Kurimoto, L.; Murakami, S.; Umezawa, T.; Shibuya, N.; Yamane, H.; Miyao, A.; Takatsuji, H.; Takahashi, A.; et al. A rice fungal MAMP-responsive MAPK cascade regulates metabolic flow to antimicrobial metabolite synthesis. Plant J. 2010, 63, 599–612. [Google Scholar] [CrossRef]
- Ma, H.; Chen, J.; Zhang, Z.; Ma, L.; Yang, Z.; Zhang, Q.; Li, X.; Xiao, J.; Wang, S. MAPK kinase 10.2 promotes disease resistance and drought tolerance by activating different MAPKs in rice. Plant J. 2017, 92, 557–570. [Google Scholar] [CrossRef]
- Kinoshita, E.; Kinoshita-Kikuta, E.; Takiyama, K.; Koike, T. Phosphate-binding tag, a new tool to visualize phosphorylated proteins. Mol. Cell Proteom. 2006, 5, 749–757. [Google Scholar] [CrossRef]
- Hosokawa, T.; Saito, T.; Asada, A.; Fukunaga, K.; Hisanaga, S. Quantitative measurement of in vivo phosphorylation states of Cdk5 activator p35 by Phos-tag SDS-PAGE. Mol. Cell Proteom. 2010, 9, 1133–1143. [Google Scholar] [CrossRef]
- Ueno, Y.; Yoshida, R.; Kishi-Kaboshi, M.; Matsushita, A.; Jiang, C.J.; Goto, S.; Takahashi, A.; Hirochika, H.; Takatsuji, H. MAP kinases phosphorylate rice WRKY45. Plant Signal. Behav. 2013, 8, e24510. [Google Scholar] [CrossRef]
- Ueno, Y.; Yoshida, R.; Kishi-Kaboshi, M.; Matsushita, A.; Jiang, C.J.; Goto, S.; Takahashi, A.; Hirochika, H.; Takatsuji, H. Abiotic Stresses Antagonize the Rice Defence Pathway through the Tyrosine-Dephosphorylation of OsMPK6. PLoS Pathog. 2015, 11, e1005231. [Google Scholar] [CrossRef]
- Liu, S.; Hua, L.; Dong, S.; Chen, H.; Zhu, X.; Jiang, J.; Zhang, F.; Li, Y.; Fang, X.; Chen, F. OsMAPK6, a mitogen-activated protein kinase, influences rice grain size and biomass production. Plant J. 2015, 84, 672–681. [Google Scholar] [CrossRef]
- Xu, R.; Duan, P.; Yu, H.; Zhou, Z.; Zhang, B.; Wang, R.; Li, J.; Zhang, G.; Zhuang, S.; Lyu, J.; et al. Control of grain size and weight by the OsMKKK10-OsMKK4-OsMAPK6 signaling pathway in rice. Mol. Plant 2018, 11, 860–873. [Google Scholar] [CrossRef]
- Katou, S.; Kuroda, K.; Seo, S.; Yanagawa, Y.; Tsuge, T.; Yamazaki, M.; Miyao, A.; Hirochika, H.; Ohashi, Y. A calmodulin-binding mitogen-activated protein kinase phosphatase is induced by wounding and regulates the activities of stress-related mitogen-activated protein kinases in rice. Plant Cell Physiol. 2007, 48, 332–344. [Google Scholar] [CrossRef]
- Ueno, Y.; Matsushita, A.; Inoue, H.; Yoshida, R.; Jiang, C.J.; Takatsuji, H. WRKY45 phosphorylation at threonine 266 acts negatively on WRKY45-dependent blast resistance in rice. Plant Signal. Behav. 2017, 12, e1356968. [Google Scholar] [CrossRef]
- Tamaoki, D.; Seo, S.; Yamada, S.; Kano, A.; Miyamoto, A.; Shishido, H.; Miyoshi, S.; Taniguchi, S.; Akimitsu, K.; Gomi, K. Jasmonic acid and salicylic acid activate a common defense system in rice. Plant Signal. Behav. 2013, 8, e24260. [Google Scholar] [CrossRef]
- Lee, M.O.; Cho, K.; Kim, S.H.; Jeong, S.H.; Kim, J.A.; Jung, Y.H.; Shim, J.; Shibato, J.; Rakwal, R.; Tamogami, S.; et al. Novel rice OsSIPK is a multiple stress responsive MAPK family member showing rhythmic expression at mRNA level. Planta 2008, 227, 981–990. [Google Scholar] [CrossRef]
- Wakuta, S.; Suzuki, E.; Saburi, W.; Matsuura, H.; Nabeta, K.; Imai, R.; Matsui, H. OsJAR1 and OsJAR2 are jasmonyl-L-isoleucine synthases involved in wound- and pathogen-induced jasmonic acid signalling. Biochem. Biophys. Res. Commun. 2011, 409, 634–639. [Google Scholar] [CrossRef]
- Riemann, M.; Haga, K.; Shimizu, T.; Okada, K.; Ando, S.; Mochizuki, S.; Nishizawa, Y.; Yamanouchi, U.; Nick, P.; Yano, M.; et al. Identification of rice Allene Oxide Cyclase mutants and the function of jasmonate for defence against Magnaporthe oryzae. Plant J. 2013, 74, 226–238. [Google Scholar] [CrossRef]
- Shimizu, T.; Miyamoto, K.; Miyamoto, K.; Minami, E.; Nishizawa, Y.; Iino, M.; Nojiri, H.; Yamane, H.; Okada, K. OsJAR1 contributes mainly to biosynthesis of the stress-induced jasmonoyl-isoleucine involved in defense responses in rice. Biosci. Biotechnol. Biochem. 2013, 77, 1556–1564. [Google Scholar] [CrossRef]
- Zhang, X.; Bao, Y.; Shan, D.; Wang, Z.; Song, X.; Wang, Z.; Wang, J.; He, L.; Wu, L.; Zhang, Z.; et al. Magnaporthe oryzae induces the expression of a microRNA to suppress the immune response in rice. Plant Physiol. 2018, 177, 352–368. [Google Scholar]
- Patkar, R.N.; Benke, P.I.; Qu, Z.; Chen, Y.Y.C.; Yang, F.; Swarup, S.; Naqvi, N.I. A fungal monooxygenase-derived jasmonate attenuates host innate immunity. Nat. Chem. Biol. 2015, 11, 733–740. [Google Scholar] [CrossRef]
- Yuan, Y.; Zhong, S.; Li, Q.; Zhu, Z.; Lou, Y.; Wang, L.; Wang, J.; Wang, M.; Li, Q.; Yang, D.; et al. Functional analysis of rice NPR1-like genes reveals that OsNPR1/NH1 is the rice orthologue conferring disease resistance with enhanced herbivore susceptibility. Plant Biotechnol. J. 2007, 5, 313–324. [Google Scholar] [CrossRef]
- Lee, S.H.; Sakuraba, Y.; Lee, T.; Kim, K.W.; An, G.; Lee, H.Y.; Paek, N.C. Mutation of Oryza sativa CORONATINE INSENSITIVE 1b (OsCOI1b) delays leaf senescence. J. Integr. Plant Biol. 2015, 57, 562–576. [Google Scholar] [CrossRef]
- Takasaki, H.; Maruyama, K.; Kidokoro, S.; Ito, Y.; Fujita, Y.; Shinozaki, K.; Yamaguchi-Shinozaki, K.; Nakashima, K. The abiotic stress-responsive NAC-type transcription factor OsNAC5 regulates stress-inducible genes and stress tolerance in rice. Mol. Genet. Genom. 2010, 284, 173–183. [Google Scholar] [CrossRef]
- Song, S.Y.; Chen, Y.; Chen, J.; Dai, X.Y.; Zhang, W.H. Physiological mechanisms underlying OsNAC5-dependent tolerance of rice plants to abiotic stress. Planta 2011, 234, 331–345. [Google Scholar] [CrossRef]
- Kazan, K. Diverse roles of jasmonates and ethylene in abiotic stress tolerance. Trends Plant Sci. 2015, 20, 219–229. [Google Scholar] [CrossRef]
- Toda, Y.; Tanaka, M.; Ogawa, D.; Kurata, K.; Kurotani, K.I.; Habu, Y.; Ando, T.; Sugimoto, K.; Mitsuda, N.; Katoh, E.; et al. RICE SALT SENSITIVE3 forms a ternary complex with JAZ and class-C bHLH factors and regulates jasmonate-induced gene expression and root cell elongation. Plant Cell 2013, 25, 1709–1725. [Google Scholar] [CrossRef]
- Sakuraba, Y.; Piao, W.; Lim, J.H.; Han, S.H.; Kim, Y.S.; An, G.; Paek, N.C. Rice ONAC106 inhibits leaf senescence and increases salt tolerance and tiller angle. Plant Cell Physiol. 2015, 56, 2325–2339. [Google Scholar] [CrossRef]
- Uji, Y.; Akimitsu, K.; Gomi, K. Identification of OsMYC2-regulated senescence-associated genes in rice. Planta 2017, 245, 1241–1246. [Google Scholar] [CrossRef]
- Kauffman, H.E.; Reddy, A.P.K.; Hsieh, S.P.Y.; Merca, S.D. An improved technique for evaluating resistance of rice varieties to Xanthomonas oryzae. Plant Dis. Rep. 1973, 57, 537–541. [Google Scholar]
- Sato, H.; Imiya, Y.; Ida, S.; Ichii, M. Characterization of four molybdenum cofactor mutants of rice, Oryza sativa L. Plant Sci. 1996, 119, 39–47. [Google Scholar] [CrossRef]
- Gomi, K.; Satoh, M.; Ozawa, R.; Shinonaga, Y.; Sanada, S.; Sasaki, K.; Matsumura, M.; Ohashi, Y.; Kanno, H.; Akimitsu, K.; et al. Role of hydroperoxide lyase in white-backed planthopper (Sogatella furcifera Horváth)-induced resistance to bacterial blight in rice, Oryza sativa L. Plant J. 2010, 61, 46–57. [Google Scholar] [CrossRef]
- Nishizawa, Y.; Nishio, Z.; Nakazono, K.; Soma, M.; Nakajima, E.; Ugaki, M.; Hibi, T. Enhanced resistance to blast (Magnaporthe grisea) in transgenic Japonica rice by constitutive expression of rice chitinase. Theor. Appl. Genet. 1999, 99, 383–390. [Google Scholar] [CrossRef]
- Shen, W.J.; Forde, B.G. Efficient transformation of Agrobacterium spp. by high voltage electroporation. Nucleic Acids Res. 1989, 17, 8385. [Google Scholar]
- Hiei, Y.; Ohta, S.; Komari, T.; Kumashiro, T. Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA. Plant J. 1994, 6, 271–282. [Google Scholar] [CrossRef]
- Arnon, D.I. Copper enzymes in isolated chloroplasts. Polyphenoloxidase in Beta vulgaris. Plant Physiol. 1949, 24, 1–15. [Google Scholar] [CrossRef]
- Shevchenko, A.; Jensen, O.N.; Podtelejnikov, A.V.; Sagliocco, F.; Wilm, M.; Vorm, O.; Mortensen, P.; Shevchenko, A.; Boucherie, H.; Mann, M. Linking genome and proteome by mass spectrometry: Large-scale identification of yeast proteins from two dimensional gels. Proc. Natl. Acad. Sci. USA 1996, 93, 14440–14445. [Google Scholar] [CrossRef]
| No. | Locus No. | Protein Name | Predicted MW (kDa) | Occurrence/Total No. of Experiments |
|---|---|---|---|---|
| 1 | Os06g06090 | Mitogen-activated protein kinase 6 (OsMPK6) | 45 | 4/5 |
| 2 | Os11g08210 | NAC domain-containing protein 5 (OsNAC5) | 35 | 3/5 |
| 3 | Os10g41330 | Ethylene response factor 36 (OsERF36) | 29 | 2/5 |
| 4 | Os01g66030 | MADS-box transcription factor 2 (OsMADS2) | 24 | 2/5 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Uji, Y.; Kashihara, K.; Kiyama, H.; Mochizuki, S.; Akimitsu, K.; Gomi, K. Jasmonic Acid-Induced VQ-Motif-Containing Protein OsVQ13 Influences the OsWRKY45 Signaling Pathway and Grain Size by Associating with OsMPK6 in Rice. Int. J. Mol. Sci. 2019, 20, 2917. https://doi.org/10.3390/ijms20122917
Uji Y, Kashihara K, Kiyama H, Mochizuki S, Akimitsu K, Gomi K. Jasmonic Acid-Induced VQ-Motif-Containing Protein OsVQ13 Influences the OsWRKY45 Signaling Pathway and Grain Size by Associating with OsMPK6 in Rice. International Journal of Molecular Sciences. 2019; 20(12):2917. https://doi.org/10.3390/ijms20122917
Chicago/Turabian StyleUji, Yuya, Keita Kashihara, Haruna Kiyama, Susumu Mochizuki, Kazuya Akimitsu, and Kenji Gomi. 2019. "Jasmonic Acid-Induced VQ-Motif-Containing Protein OsVQ13 Influences the OsWRKY45 Signaling Pathway and Grain Size by Associating with OsMPK6 in Rice" International Journal of Molecular Sciences 20, no. 12: 2917. https://doi.org/10.3390/ijms20122917
APA StyleUji, Y., Kashihara, K., Kiyama, H., Mochizuki, S., Akimitsu, K., & Gomi, K. (2019). Jasmonic Acid-Induced VQ-Motif-Containing Protein OsVQ13 Influences the OsWRKY45 Signaling Pathway and Grain Size by Associating with OsMPK6 in Rice. International Journal of Molecular Sciences, 20(12), 2917. https://doi.org/10.3390/ijms20122917





