Combined Therapy with SS31 and Mitochondria Mitigates Myocardial Ischemia-Reperfusion Injury in Rats
Abstract
1. Introduction
2. Results
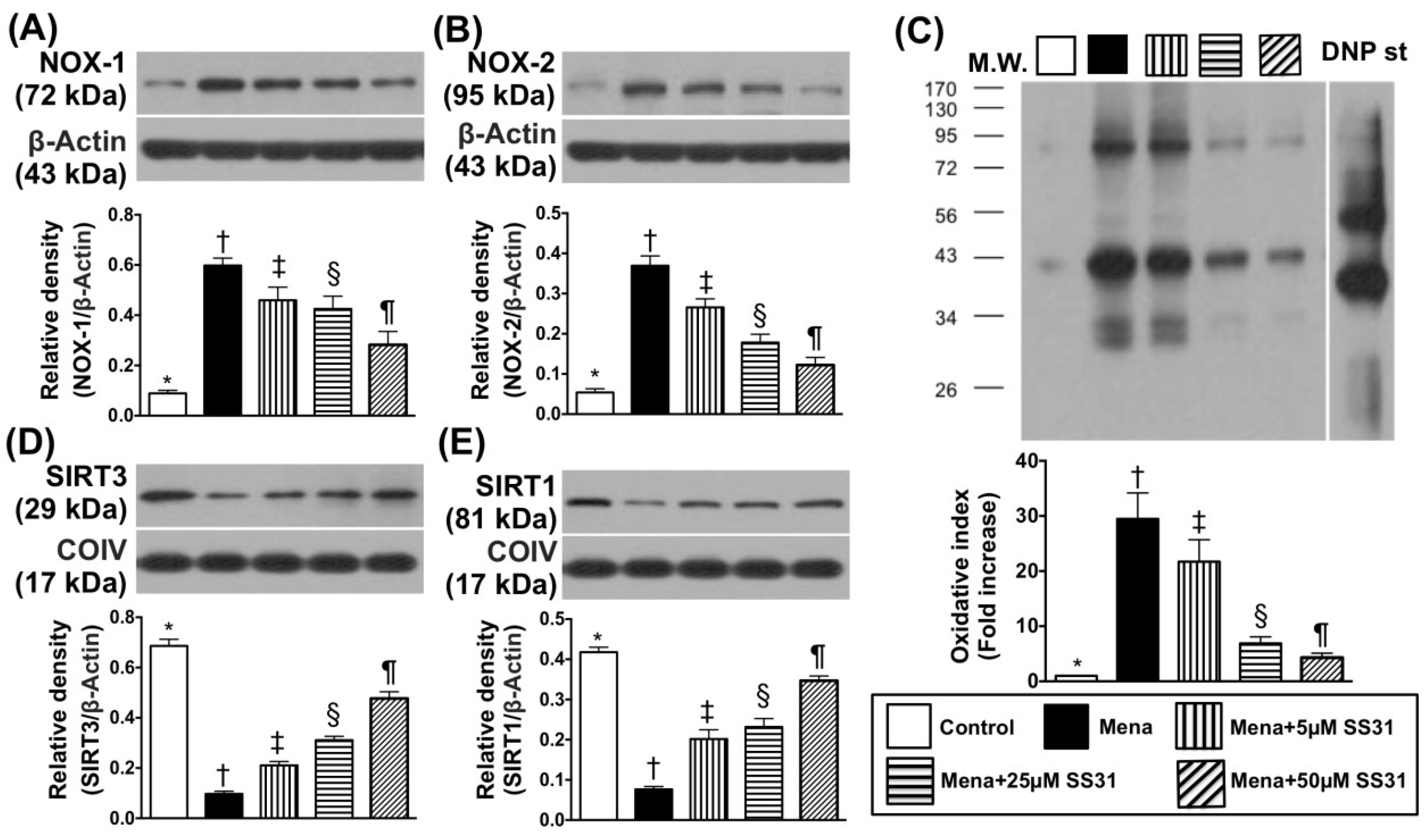
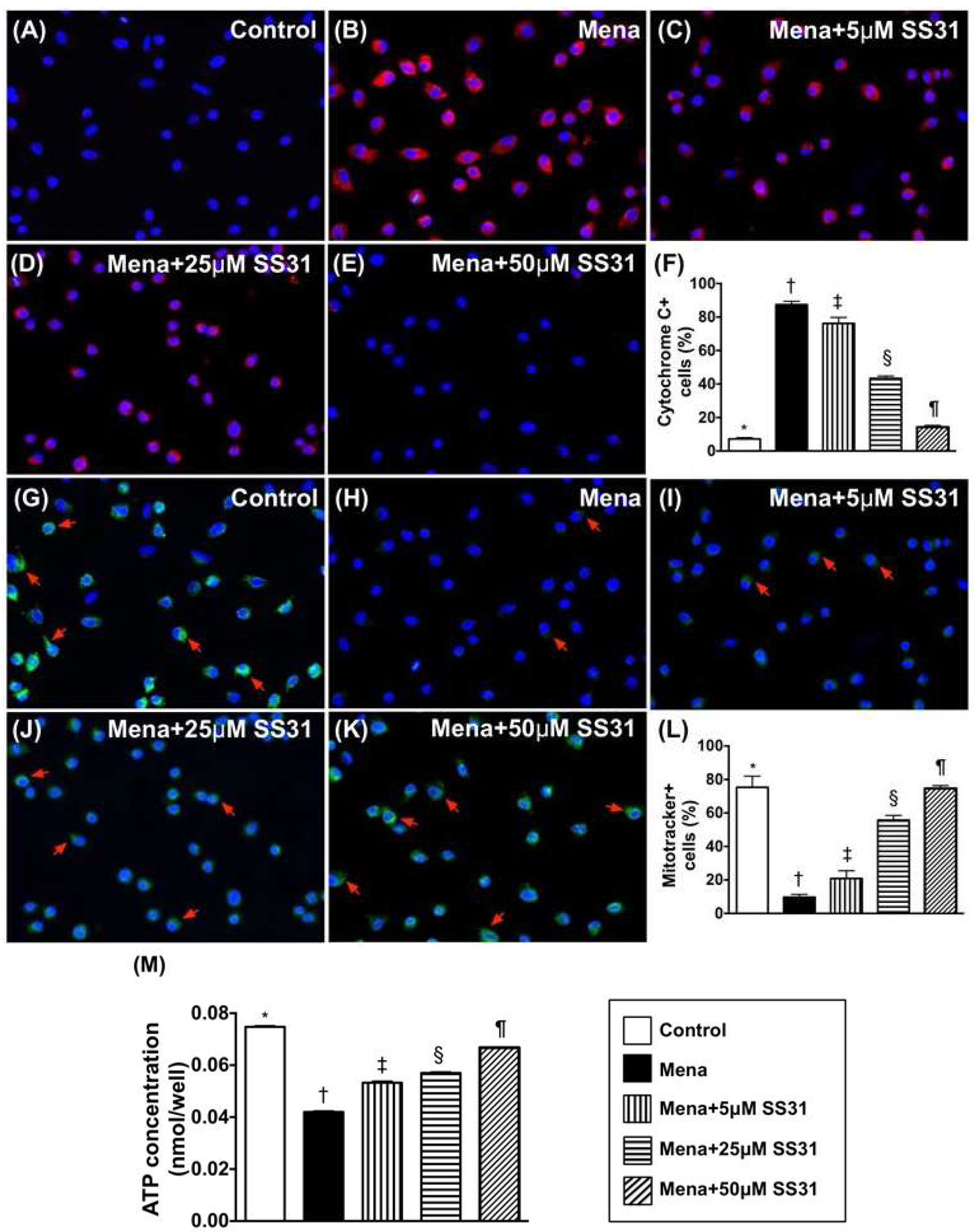
2.1. SS31 Suppresses Menadione-Induced Oxidative Stress and Preserves ATP in H9C2 Cells
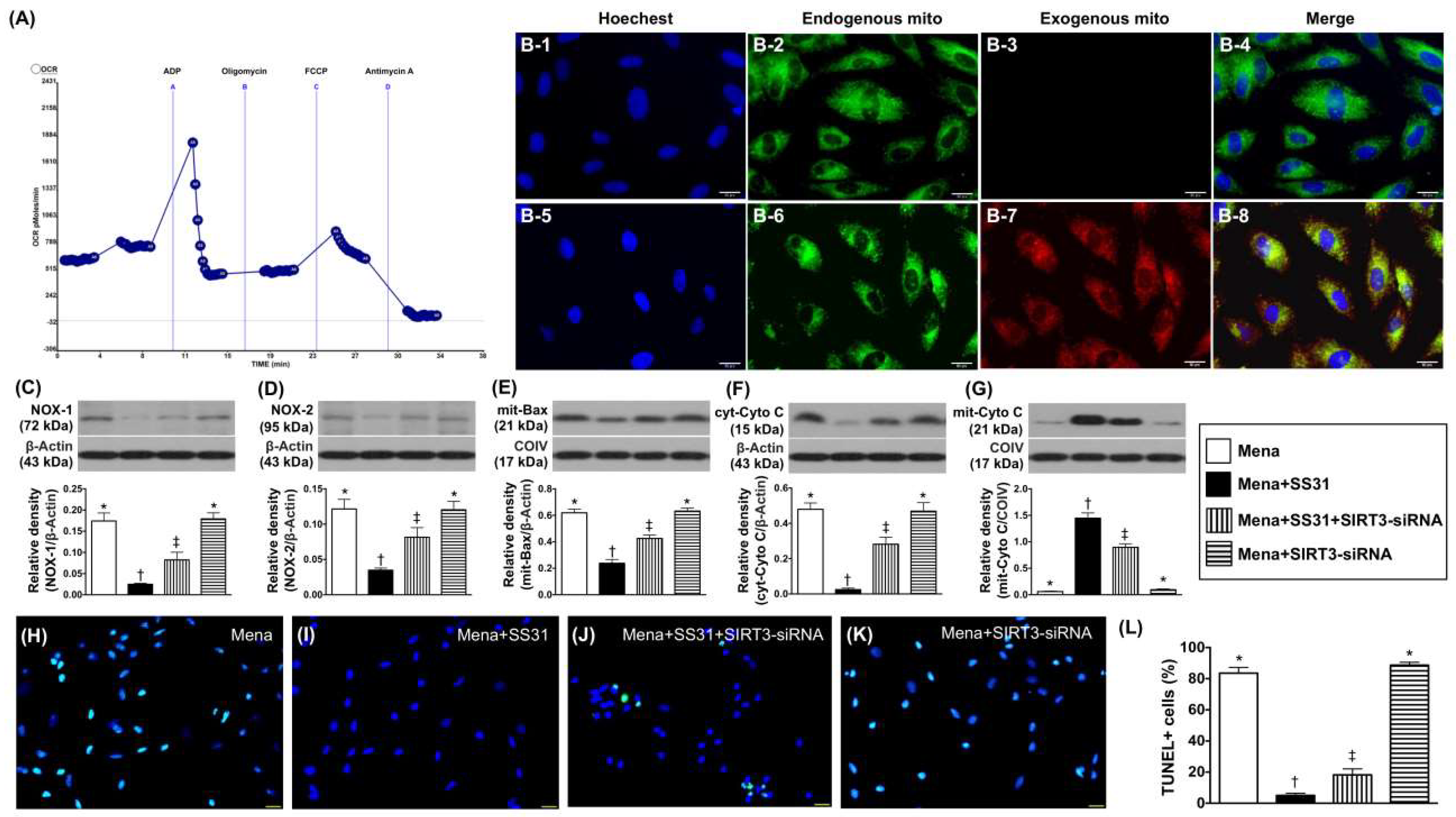
2.2. Mitochondrial Transfusion Refreshes Intracellular Mitochondria and SIRT3 Suppression Abrogates the Cyto-Protective Effects of SS31 by Increasing Oxidative Stress in Menadione-Treated H9C2 Cells
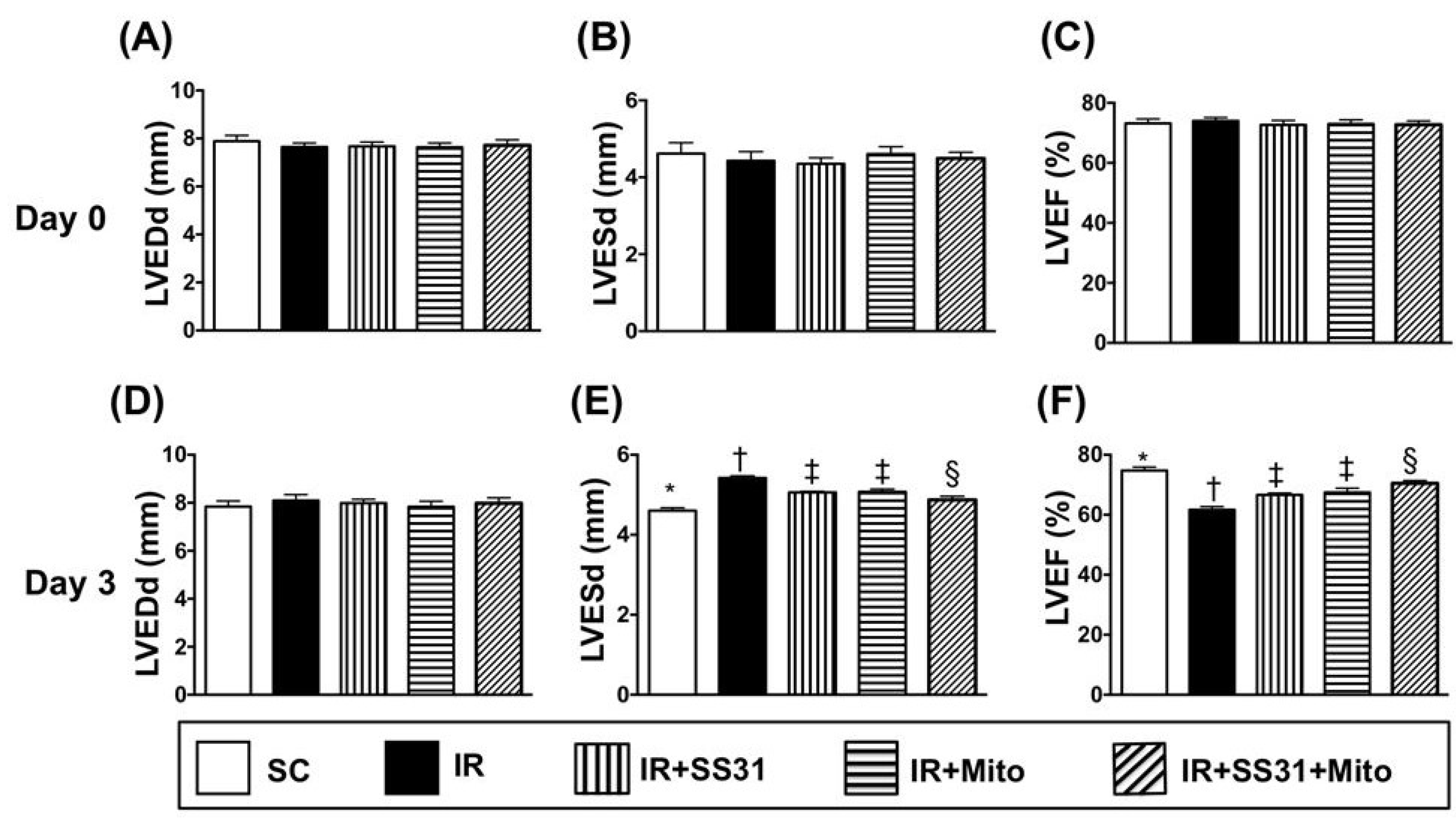
2.3. Transthoracic Echocardiographic Findings after IR Procedure
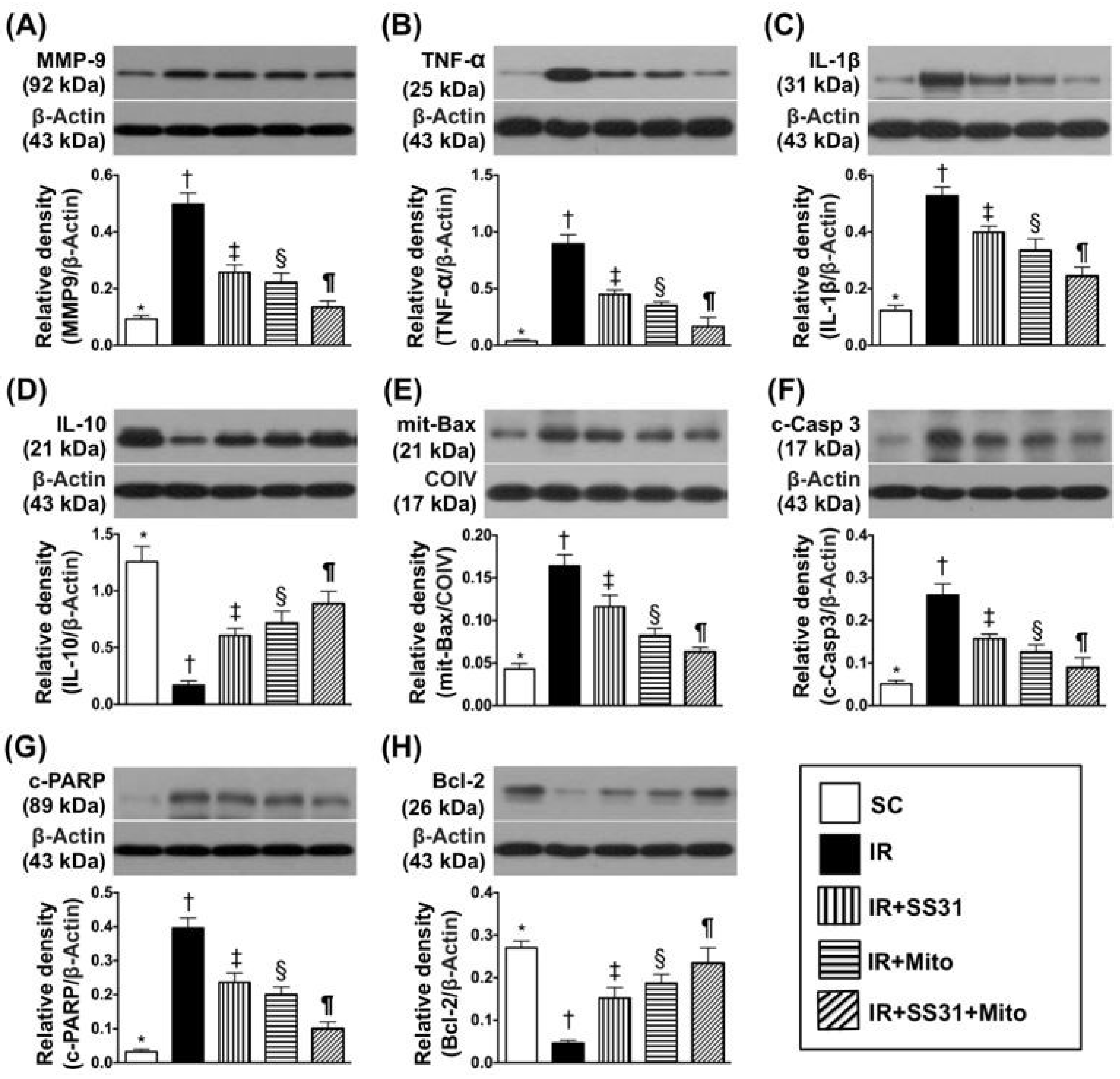
2.4. Inflammatory and Apoptotic Markers in LV Myocardium after the IR Procedure
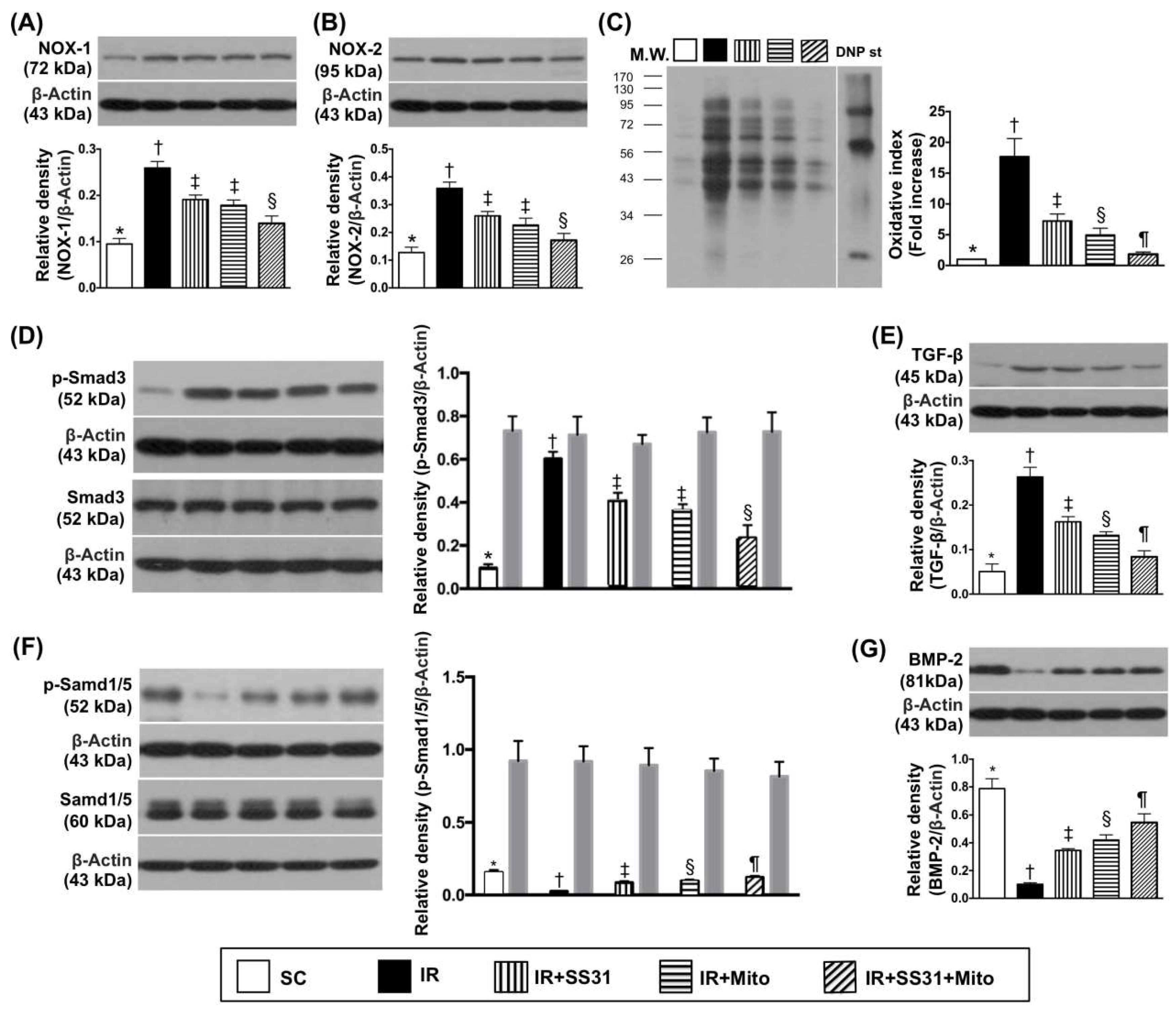
2.5. Oxidative Stress and Fibrosis Markers in LV Myocardium after the IR Procedure
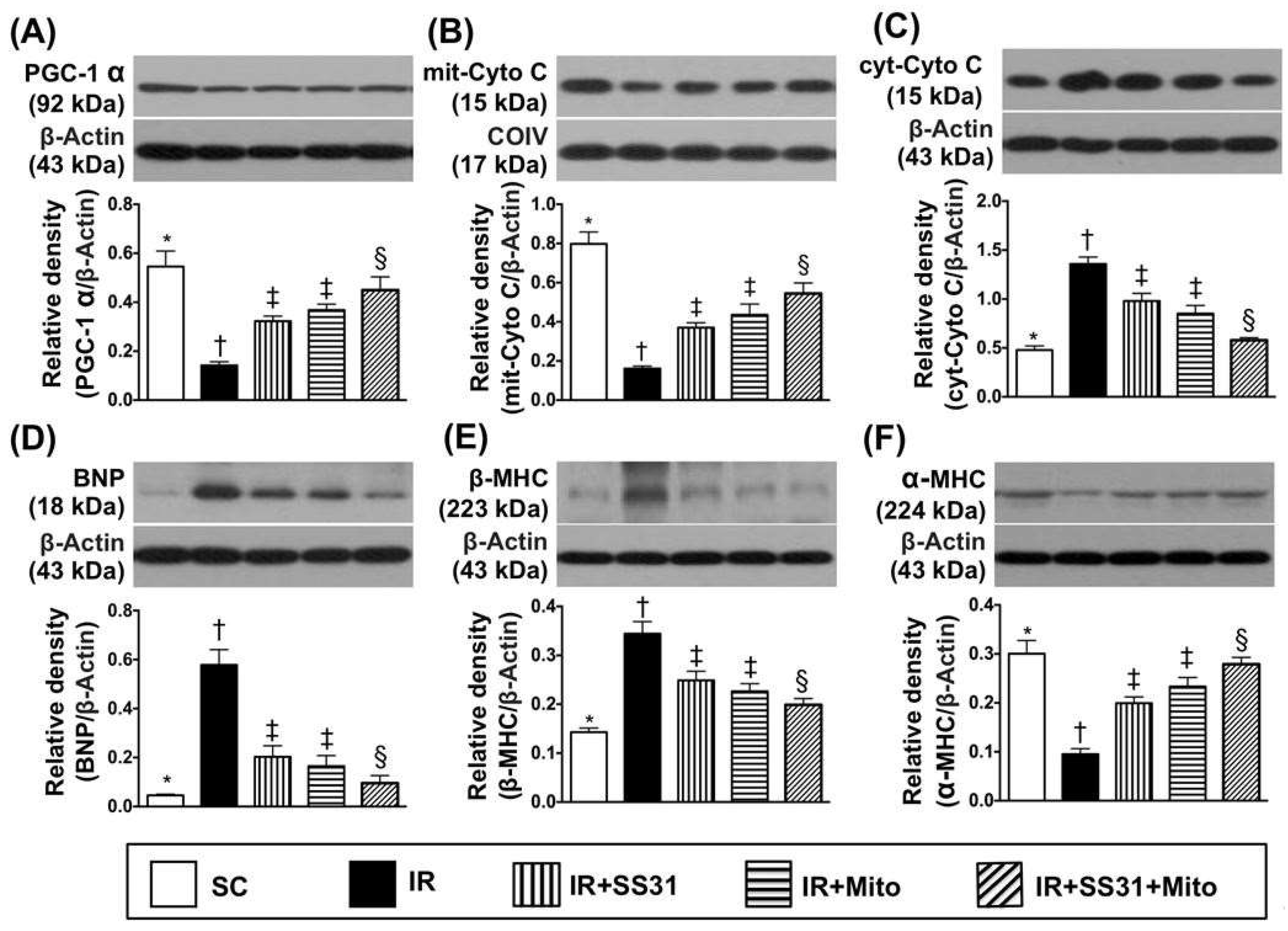
2.6. Energy Integrity and Pressure/Volume Overload Biomarkers in LV Myocardium after IR Procedure
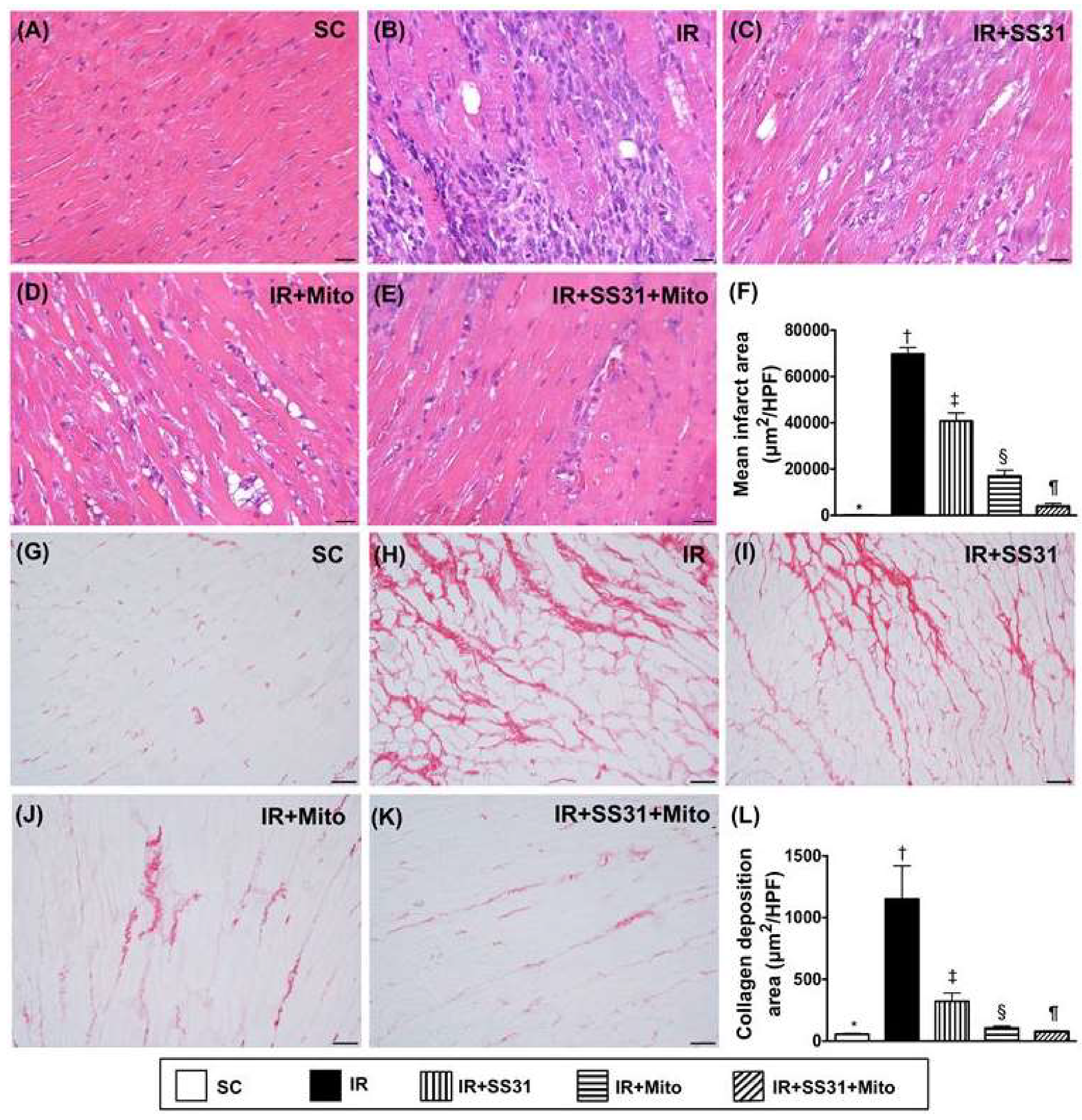
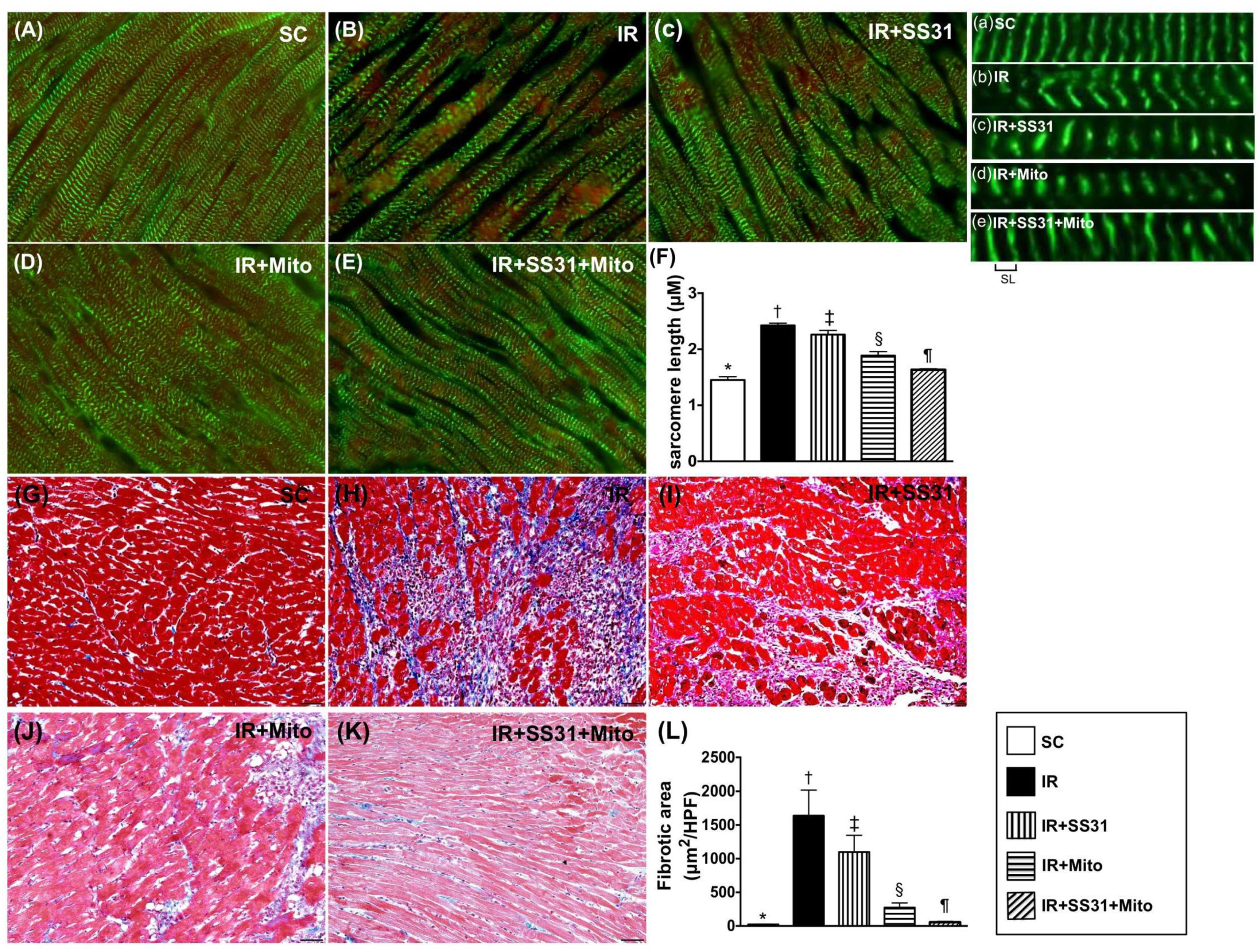
2.7. Infarct and Collagen-Deposition Areas in LV Myocardium after the IR Procedure
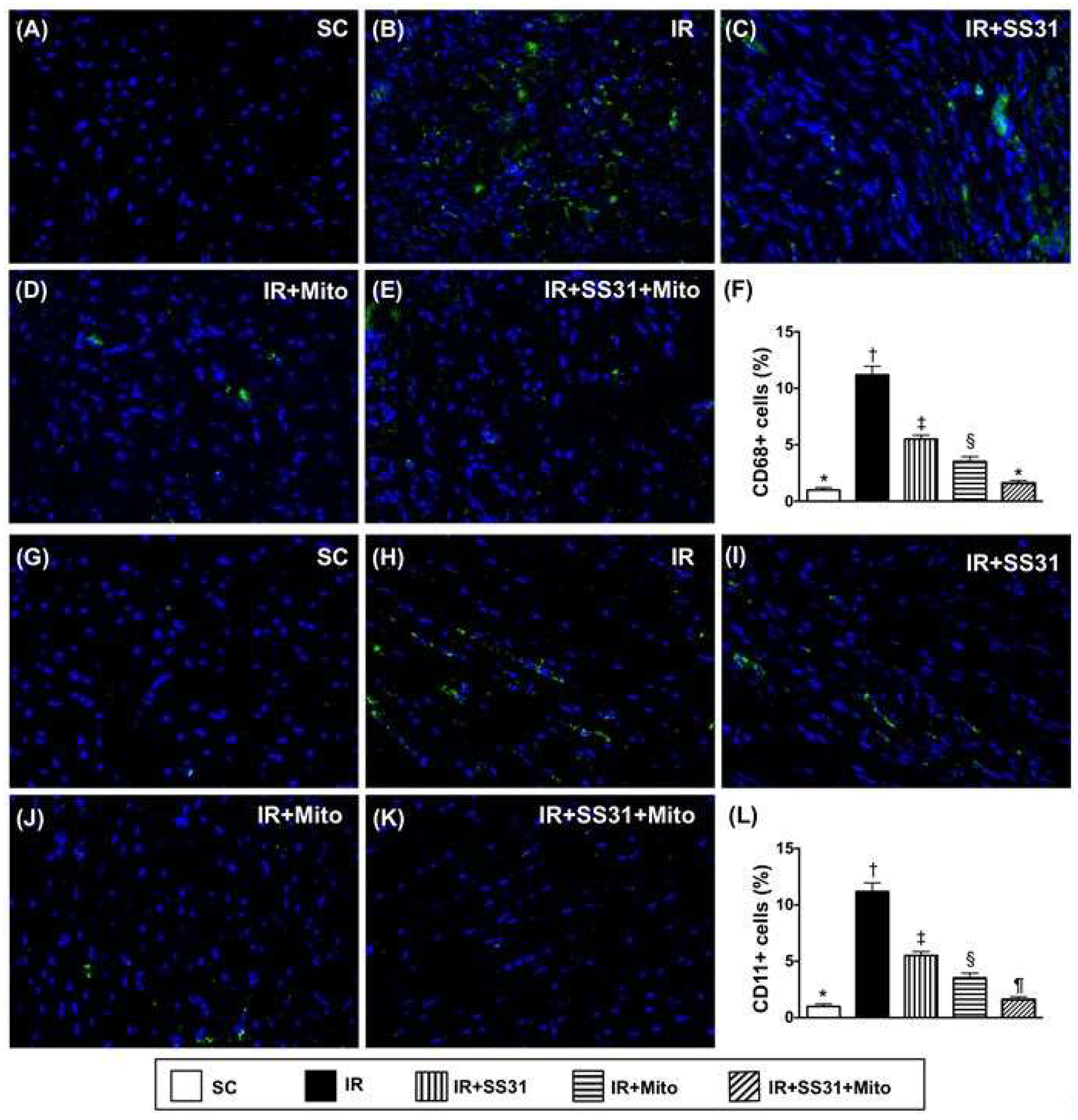
2.8. Cellular Levels of Inflammatory Markers in LV Myocardium after IR Procedure
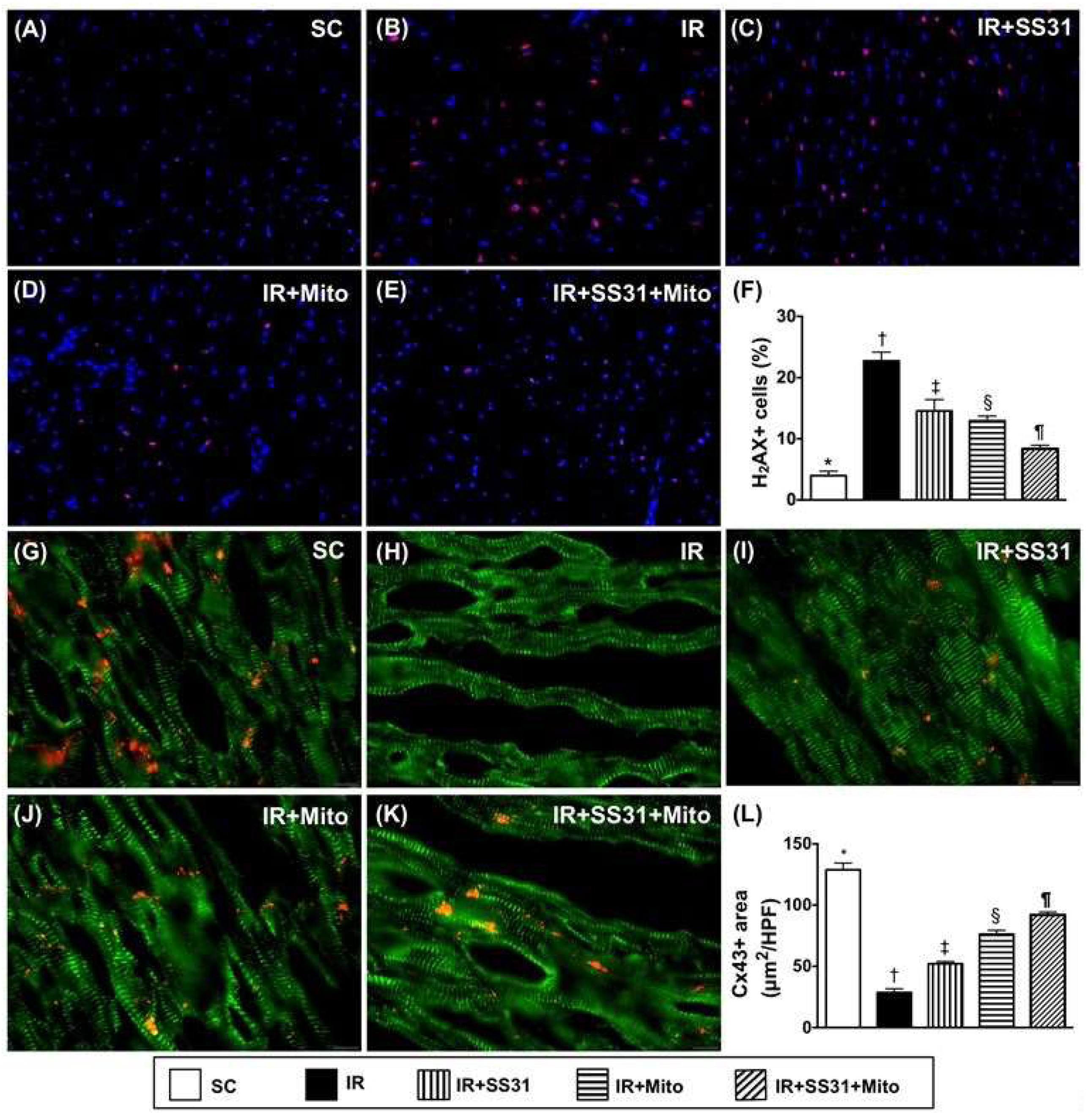
2.9. Cellular Levels of DNA-Damage and Gap Junction Biomarkers in the LV Myocardium after the IR Procedure
2.10. Sarcomere Length of LV Myocardium after the IR Procedure
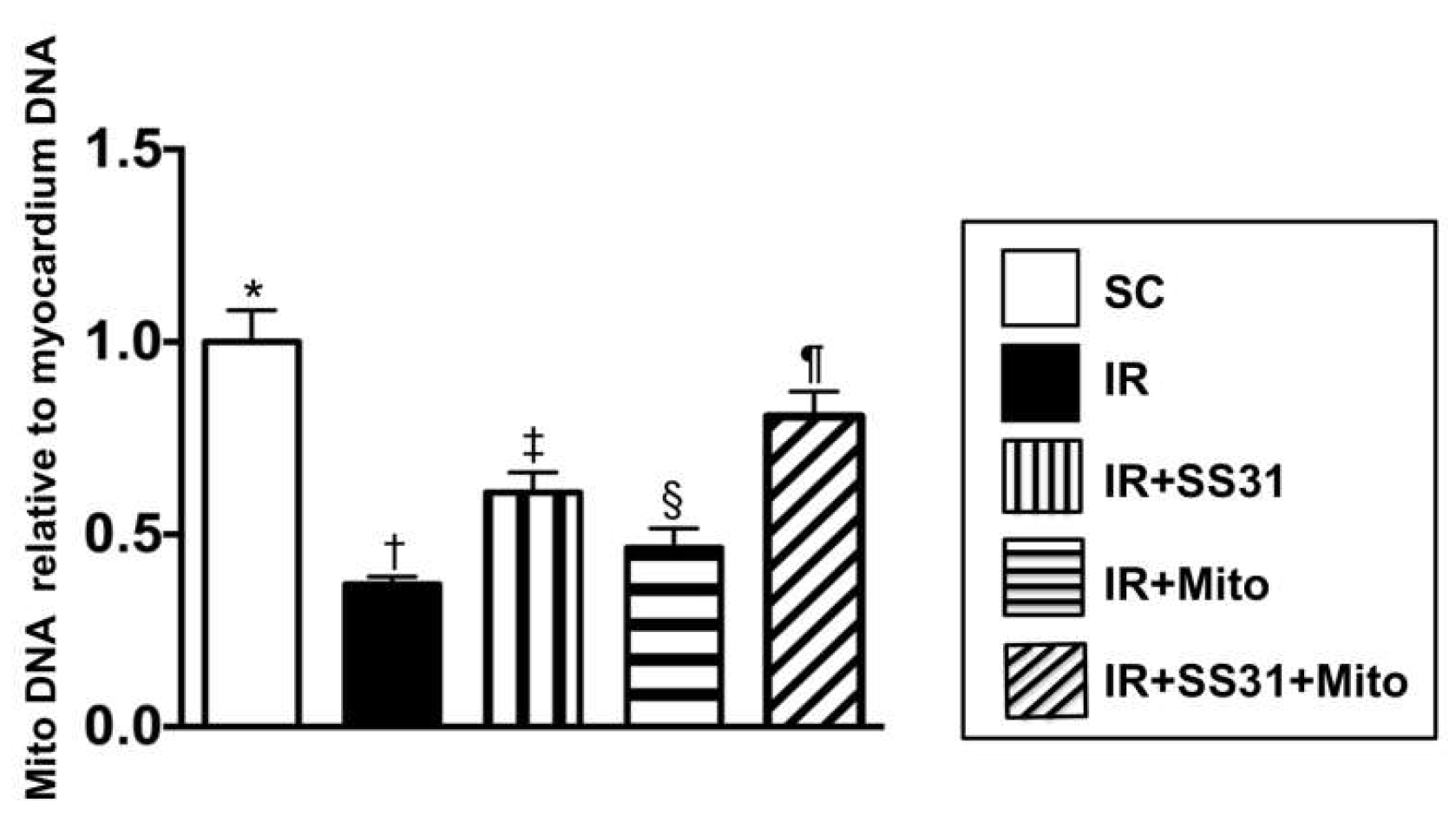
2.11. The Expression of Mitochondrial DNA Copy Number in the IR Area of the Left Ventricular Myocardium
3. Discussion
4. Materials and Methods
4.1. Animal Studies
4.2. Induction of Acute Myocardial Ischemia-Reperfusion Injury
4.3. Functional Assessment with Echocardiography
4.4. Mitochondrial Isolation, Staining, and Transfusion
4.5. Oxygen Consumption Rate (OCR) of the Isolated Mitochondria (Seahorse Method)
4.6. Material and Method for In Vitro Study
4.7. ATP Assay by ELISA
4.8. Assessment of SS31 Role in Protecting Cardiomyocytes from Oxidative Stress
4.9. Western Blot Analysis of Heart Tissues
4.11. Immunohistochemical (IHC) and Immunofluorescent (IF) Staining
4.10. Oxidative Stress Reaction in the LV Myocardium
4.12. Histological Quantification of Myocardial Fibrosis/Infarct Area (IA) and Collagen Deposition Area
4.13. Quantification of Mitochondrial DNA Copy Number by qPCR
4.14. Statistical Analysis
4.15. Availability of Data and Material
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| AMI | acute myocardial infarction |
| BMP | bone morphogenetic protein |
| BNP | brain natriuretic peptide |
| Casp | caspase |
| c | cleaved |
| Cx43 | connexin 43 |
| IL | interleukin |
| IR | ischemia-reperfusion |
| LVEF | left ventricular ejection fraction |
| LVEDd | left ventricular end-diastolic diameter |
| LVESd | left ventricular end-systolic diameter |
| MMP | matrix metalloproteinase |
| Mito | mitochondria |
| MHC | myosin heavy chain |
| NOX | NADPH oxidase |
| NF-κB | nuclear factor-κB |
| OCR | oxygen consumption rate |
| PGC | peroxisome proliferator activated receptor-gamma coactivator |
| PARP | poly(ADP-ribose) polymerase |
| ROS | reactive oxygen species |
| TGF | transforming growth factor |
| TNF-α | tumor necrosis factor-α |
References
- Eltzschig, H.K.; Eckle, T. Ischemia and reperfusion—From mechanism to translation. Nat. Med. 2011, 17, 1391–1401. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.S.; Chao, A.; Yu, H.Y.; Ko, W.J.; Wu, I.H.; Chen, R.J.; Huang, S.C.; Lin, F.Y.; Wang, S.S. Analysis and results of prolonged resuscitation in cardiac arrest patients rescued by extracorporeal membrane oxygenation. J. Am. Coll. Cardiol. 2003, 41, 197–203. [Google Scholar] [CrossRef]
- Ito, H.; Maruyama, A.; Iwakura, K.; Takiuchi, S.; Masuyama, T.; Hori, M.; Higashino, Y.; Fujii, K.; Minamino, T. Clinical implications of the ‘no reflow’ phenomenon. A predictor of complications and left ventricular remodeling in reperfused anterior wall myocardial infarction. Circulation 1996, 93, 223–228. [Google Scholar] [CrossRef] [PubMed]
- Kloner, R.A.; Ellis, S.G.; Lange, R.; Braunwald, E. Studies of experimental coronary artery reperfusion. Effects on infarct size, myocardial function, biochemistry, ultrastructure and microvascular damage. Circulation 1983, 68, 18. [Google Scholar]
- Rokos, I.C.; French, W.J.; Koenig, W.J.; Stratton, S.J.; Nighswonger, B.; Strunk, B.; Jewell, J.; Mahmud, E.; Dunford, J.V.; Hokanson, J.; et al. Integration of pre-hospital electrocardiograms and ST-elevation myocardial infarction receiving center (SRC) networks: Impact on Door-to-Balloon times across 10 independent regions. JACC Cardiovasc. Interv. 2009, 2, 339–346. [Google Scholar] [CrossRef] [PubMed]
- Rokos, I.C.; Larson, D.M.; Henry, T.D.; Koenig, W.J.; Eckstein, M.; French, W.J.; Granger, C.B.; Roe, M.T. Rationale for establishing regional ST-elevation myocardial infarction receiving center (SRC) networks. Am. Heart J. 2006, 152, 661–667. [Google Scholar] [CrossRef] [PubMed]
- De Scheerder, I.; Vandekerckhove, J.; Robbrecht, J.; Algoed, L.; De Buyzere, M.; De Langhe, J.; De Schrijver, G.; Clement, D. Post-cardiac injury syndrome and an increased humoral immune response against the major contractile proteins (actin and myosin). Am. J. Cardiol. 1985, 56, 631–633. [Google Scholar] [CrossRef]
- Frangogiannis, N.G. The immune system and cardiac repair. Pharmacol. Res. 2008, 58, 88–111. [Google Scholar] [CrossRef] [PubMed]
- Frangogiannis, N.G.; Smith, C.W.; Entman, M.L. The inflammatory response in myocardial infarction. Cardiovasc. Res. 2002, 53, 31–47. [Google Scholar] [CrossRef]
- Lambert, J.M.; Lopez, E.F.; Lindsey, M.L. Macrophage roles following myocardial infarction. Int. J. Cardiol. 2008, 130, 147–158. [Google Scholar] [CrossRef] [PubMed]
- Lange, L.G.; Schreiner, G.F. Immune mechanisms of cardiac disease. N. Engl. J. Med. 1994, 330, 1129–1135. [Google Scholar] [PubMed]
- Hoffman, J.W., Jr.; Gilbert, T.B.; Poston, R.S.; Silldorff, E.P. Myocardial reperfusion injury: Etiology, mechanisms, and therapies. J. Extra Corpor. Technol. 2004, 36, 391–411. [Google Scholar] [PubMed]
- Kalogeris, T.; Baines, C.P.; Krenz, M.; Korthuis, R.J. Ischemia/Reperfusion. Compr. Physiol. 2016, 7, 113–170. [Google Scholar] [PubMed]
- Kalogeris, T.; Baines, C.P.; Krenz, M.; Korthuis, R.J. Cell biology of ischemia/reperfusion injury. Int. Rev. Cell Mol. Biol. 2012, 298, 229–317. [Google Scholar] [PubMed]
- Tsutsui, H.; Kinugawa, S.; Matsushima, S. Mitochondrial oxidative stress and dysfunction in myocardial remodelling. Cardiovasc. Res. 2009, 81, 449–456. [Google Scholar] [CrossRef] [PubMed]
- Chuah, S.C.; Moore, P.K.; Zhu, Y.Z. S-allylcysteine mediates cardioprotection in an acute myocardial infarction rat model via a hydrogen sulfide-mediated pathway. Am. J. Physiol. Heart Circ. Physiol. 2007, 293, H2693–H2701. [Google Scholar] [CrossRef] [PubMed]
- Misra, M.K.; Sarwat, M.; Bhakuni, P.; Tuteja, R.; Tuteja, N. Oxidative stress and ischemic myocardial syndromes. Med. Sci. Monit. 2009, 15, RA209–RA219. [Google Scholar] [PubMed]
- Dai, D.F.; Chen, T.; Szeto, H.; Nieves-Cintron, M.; Kutyavin, V.; Santana, L.F.; Rabinovitch, P.S. Mitochondrial targeted antioxidant Peptide ameliorates hypertensive cardiomyopathy. J. Am. Coll. Cardiol. 2011, 58, 73–82. [Google Scholar] [CrossRef] [PubMed]
- Birk, A.V.; Liu, S.; Soong, Y.; Mills, W.; Singh, P.; Warren, J.D.; Seshan, S.V.; Pardee, J.D.; Szeto, H.H. The mitochondrial-targeted compound SS-31 re-energizes ischemic mitochondria by interacting with cardiolipin. J. Am. Soc. Nephrol. 2013, 24, 1250–1261. [Google Scholar] [CrossRef] [PubMed]
- Szeto, H.H. Mitochondria-targeted cytoprotective peptides for ischemia-reperfusion injury. Antioxid Redox Signal 2008, 10, 601–619. [Google Scholar] [CrossRef] [PubMed]
- Lu, H.I.; Huang, T.H.; Sung, P.H.; Chen, Y.L.; Chua, S.; Chai, H.Y.; Chung, S.Y.; Liu, C.F.; Sun, C.K.; Chang, H.W.; et al. Administration of antioxidant peptide SS-31 attenuates transverse aortic constriction-induced pulmonary arterial hypertension in mice. Acta Pharmacol. Sin. 2016, 37, 589–603. [Google Scholar] [CrossRef] [PubMed]
- Zhao, K.; Zhao, G.M.; Wu, D.; Soong, Y.; Birk, A.V.; Schiller, P.W.; Szeto, H.H. Cell-permeable peptide antioxidants targeted to inner mitochondrial membrane inhibit mitochondrial swelling, oxidative cell death, and reperfusion injury. J. Biol. Chem. 2004, 279, 34682–34690. [Google Scholar] [CrossRef] [PubMed]
- Islam, M.N.; Das, S.R.; Emin, M.T.; Wei, M.; Sun, L.; Westphalen, K.; Rowlands, D.J.; Quadri, S.K.; Bhattacharya, S.; Bhattacharya, J. Mitochondrial transfer from bone-marrow-derived stromal cells to pulmonary alveoli protects against acute lung injury. Nat. Med. 2012, 18, 759–765. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.H.; Chen, Y.T.; Yang, C.C.; Chen, K.H.; Sung, P.H.; Chiang, H.J.; Chen, C.H.; Chua, S.; Chung, S.Y.; Chen, Y.L.; et al. Melatonin pretreatment enhances the therapeutic effects of exogenous mitochondria against hepatic ischemia-reperfusion injury in rats through suppression of mitochondrial permeability transition. J. Pineal Res. 2016, 61, 52–68. [Google Scholar] [CrossRef] [PubMed]
- Masuzawa, A.; Black, K.M.; Pacak, C.A.; Ericsson, M.; Barnett, R.J.; Drumm, C.; Seth, P.; Bloch, D.B.; Levitsky, S.; Cowan, D.B.; et al. Transplantation of autologously derived mitochondria protects the heart from ischemia-reperfusion injury. Am. J. Physiol. Heart Circ. Physiol. 2013, 304, H966–H982. [Google Scholar] [CrossRef] [PubMed]
- Lin, H.C.; Liu, S.Y.; Lai, H.S.; Lai, I.R. Isolated mitochondria infusion mitigates ischemia-reperfusion injury of the liver in rats. Shock 2013, 39, 304–310. [Google Scholar] [CrossRef] [PubMed]
- Szeto, H.H.; Liu, S.; Soong, Y.; Wu, D.; Darrah, S.F.; Cheng, F.Y.; Zhao, Z.; Ganger, M.; Tow, C.Y.; Seshan, S.V. Mitochondria-targeted peptide accelerates ATP recovery and reduces ischemic kidney injury. J. Am. Soc. Nephrol. 2011, 22, 1041–1052. [Google Scholar] [CrossRef] [PubMed]
- Kloner, R.A.; Hale, S.L.; Dai, W.; Gorman, R.C.; Shuto, T.; Koomalsingh, K.J.; Gorman, J.H., 3rd; Sloan, R.C.; Frasier, C.R.; Watson, C.A.; et al. Reduction of ischemia/reperfusion injury with bendavia, a mitochondria-targeting cytoprotective Peptide. J. Am. Heart Assoc. 2012, 1, e001644. [Google Scholar] [CrossRef] [PubMed]
- Dai, W.; Cheung, E.; Alleman, R.J.; Perry, J.B.; Allen, M.E.; Brown, D.A.; Kloner, R.A. Cardioprotective Effects of Mitochondria-Targeted Peptide SBT-20 in two Different Models of Rat Ischemia/Reperfusion. Cardiovasc. Drugs Ther. 2016, 30, 559–566. [Google Scholar] [CrossRef] [PubMed]
- Chua, S.; Lee, F.Y.; Chiang, H.J.; Chen, K.H.; Lu, H.I.; Chen, Y.T.; Yang, C.C.; Lin, K.C.; Chen, Y.L.; Kao, G.S.; et al. The cardioprotective effect of melatonin and exendin-4 treatment in a rat model of cardiorenal syndrome. J. Pineal Res. 2016, 61, 438–456. [Google Scholar] [CrossRef] [PubMed]
- Sheu, J.J.; Sung, P.H.; Leu, S.; Chai, H.T.; Zhen, Y.Y.; Chen, Y.C.; Chua, S.; Chen, Y.L.; Tsai, T.H.; Lee, F.Y.; et al. Innate immune response after acute myocardial infarction and pharmacomodulatory action of tacrolimus in reducing infarct size and preserving myocardial integrity. J. Biomed. Sci. 2013, 20, 82. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.L.; Chung, S.Y.; Chai, H.T.; Chen, C.H.; Liu, C.F.; Chen, Y.L.; Huang, T.H.; Zhen, Y.Y.; Sung, P.H.; Sun, C.K.; et al. Early Administration of Carvedilol Protected against Doxorubicin-Induced Cardiomyopathy. J. Pharmacol. Exp. Ther. 2015, 355, 516–527. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.L.; Sun, C.K.; Tsai, T.H.; Chang, L.T.; Leu, S.; Zhen, Y.Y.; Sheu, J.J.; Chua, S.; Yeh, K.H.; Lu, H.I.; et al. Adipose-derived mesenchymal stem cells embedded in platelet-rich fibrin scaffolds promote angiogenesis, preserve heart function, and reduce left ventricular remodeling in rat acute myocardial infarction. Am. J. Transl. Res. 2015, 7, 781–803. [Google Scholar] [PubMed]
- Khalitova, E.B. Viability and retention of virulent properties by shigellae lyophilized in different protective media. Zh. Mikrobiol. Epidemiol. Immunobiol. 1974, 150–151. [Google Scholar]
- Morigi, M.; Perico, L.; Rota, C.; Longaretti, L.; Conti, S.; Rottoli, D.; Novelli, R.; Remuzzi, G.; Benigni, A. Sirtuin 3-dependent mitochondrial dynamic improvements protect against acute kidney injury. J. Clin. Investig. 2015, 125, 715–726. [Google Scholar] [CrossRef] [PubMed]
- Chua, S.; Lee, F.Y.; Tsai, T.H.; Sheu, J.J.; Leu, S.; Sun, C.K.; Chen, Y.L.; Chang, H.W.; Chai, H.T.; Liu, C.F.; et al. Inhibition of dipeptidyl peptidase-IV enzyme activity protects against myocardial ischemia-reperfusion injury in rats. J. Transl. Med. 2014, 12, 357. [Google Scholar] [CrossRef] [PubMed]
- Huang, T.H.; Chung, S.Y.; Chua, S.; Chai, H.T.; Sheu, J.J.; Chen, Y.L.; Chen, C.H.; Chang, H.W.; Tong, M.S.; Sung, P.H.; et al. Effect of early administration of lower dose versus high dose of fresh mitochondria on reducing monocrotaline-induced pulmonary artery hypertension in rat. Am. J. Transl. Res. 2016, 8, 5151–5168. [Google Scholar] [PubMed]
- Sun, C.K.; Lee, F.Y.; Kao, Y.H.; Chiang, H.J.; Sung, P.H.; Tsai, T.H.; Lin, Y.C.; Leu, S.; Wu, Y.C.; Lu, H.I.; et al. Systemic combined melatonin-mitochondria treatment improves acute respiratory distress syndrome in the rat. J. Pineal Res. 2015, 58, 137–150. [Google Scholar] [CrossRef] [PubMed]
- Leu, S.; Sun, C.K.; Sheu, J.J.; Chang, L.T.; Yuen, C.M.; Yen, C.H.; Chiang, C.H.; Ko, S.F.; Pei, S.N.; Chua, S.; et al. Autologous bone marrow cell implantation attenuates left ventricular remodeling and improves heart function in porcine myocardial infarction: An echocardiographic, six-month angiographic, and molecular-cellular study. Int. J. Cardiol. 2011, 150, 156–168. [Google Scholar] [CrossRef] [PubMed]


© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lee, F.-Y.; Shao, P.-L.; Wallace, C.G.; Chua, S.; Sung, P.-H.; Ko, S.-F.; Chai, H.-T.; Chung, S.-Y.; Chen, K.-H.; Lu, H.-I.; et al. Combined Therapy with SS31 and Mitochondria Mitigates Myocardial Ischemia-Reperfusion Injury in Rats. Int. J. Mol. Sci. 2018, 19, 2782. https://doi.org/10.3390/ijms19092782
Lee F-Y, Shao P-L, Wallace CG, Chua S, Sung P-H, Ko S-F, Chai H-T, Chung S-Y, Chen K-H, Lu H-I, et al. Combined Therapy with SS31 and Mitochondria Mitigates Myocardial Ischemia-Reperfusion Injury in Rats. International Journal of Molecular Sciences. 2018; 19(9):2782. https://doi.org/10.3390/ijms19092782
Chicago/Turabian StyleLee, Fan-Yen, Pei-Lin Shao, Christopher Glenn Wallace, Sarah Chua, Pei-Hsun Sung, Sheung-Fat Ko, Han-Tan Chai, Sheng-Ying Chung, Kuan-Hung Chen, Hung-I Lu, and et al. 2018. "Combined Therapy with SS31 and Mitochondria Mitigates Myocardial Ischemia-Reperfusion Injury in Rats" International Journal of Molecular Sciences 19, no. 9: 2782. https://doi.org/10.3390/ijms19092782
APA StyleLee, F.-Y., Shao, P.-L., Wallace, C. G., Chua, S., Sung, P.-H., Ko, S.-F., Chai, H.-T., Chung, S.-Y., Chen, K.-H., Lu, H.-I., Chen, Y.-L., Huang, T.-H., Sheu, J.-J., & Yip, H.-K. (2018). Combined Therapy with SS31 and Mitochondria Mitigates Myocardial Ischemia-Reperfusion Injury in Rats. International Journal of Molecular Sciences, 19(9), 2782. https://doi.org/10.3390/ijms19092782




