Analysis of Genetic Variation and Enhancement of Salt Tolerance in French Pea (Pisum Sativum L.)
Abstract
1. Introduction
2. Results and Discussion
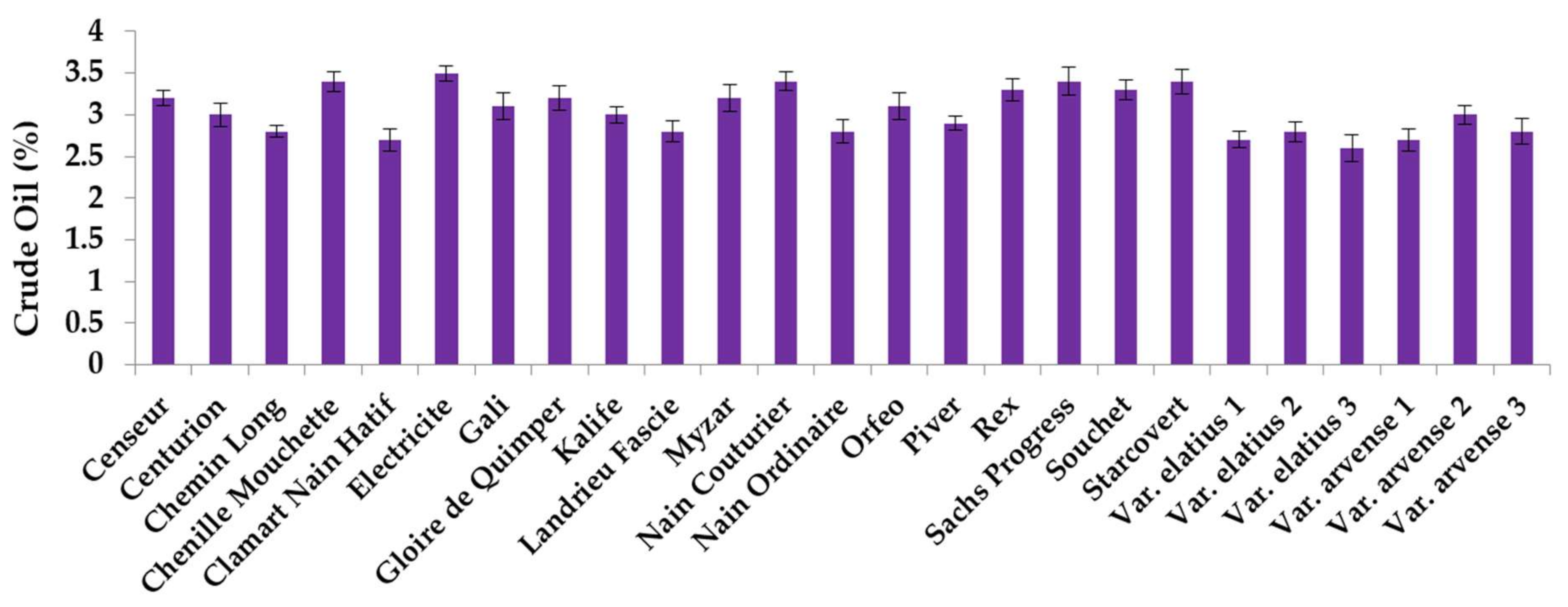
2.1. Crude Oil Content and Fatty Acid Analysis
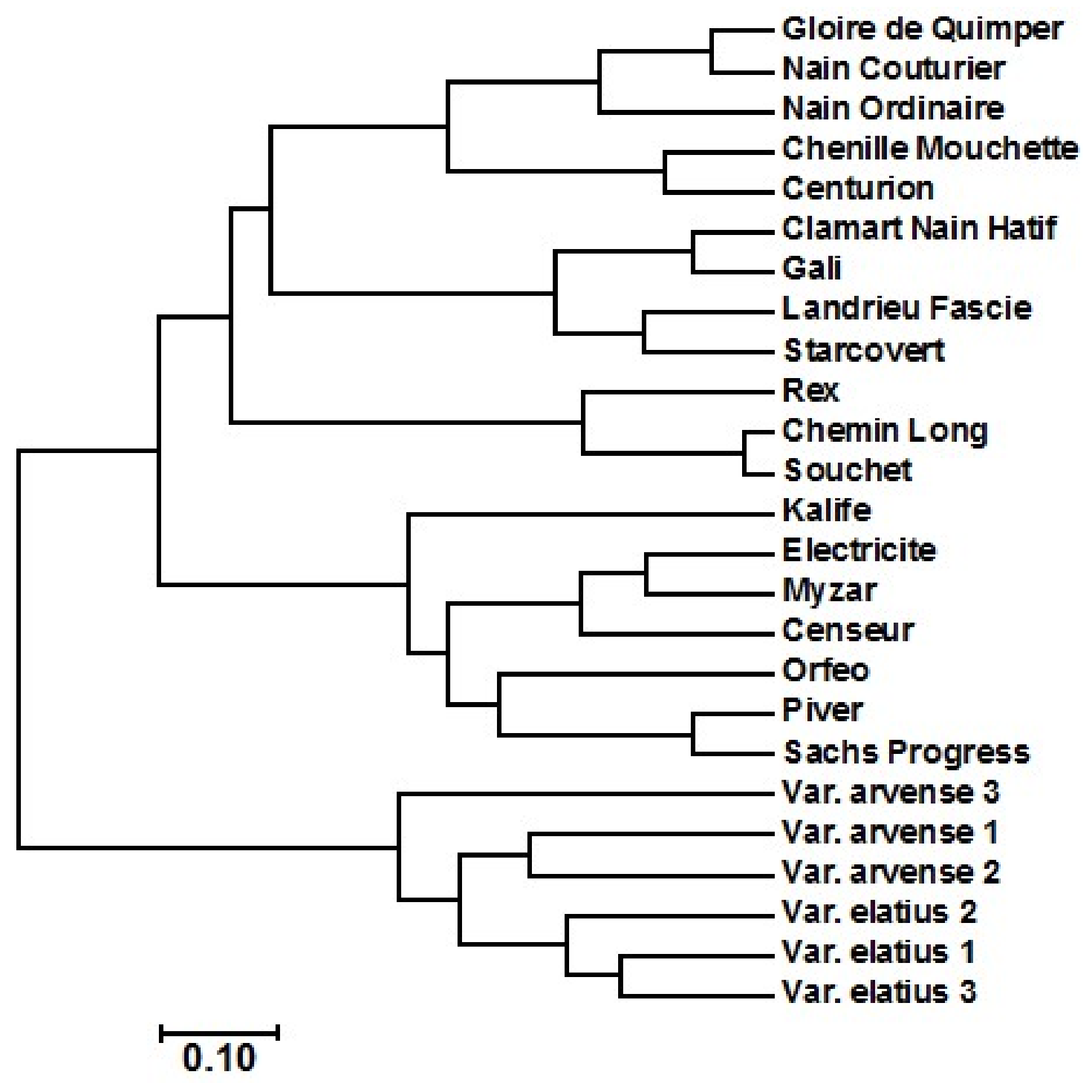
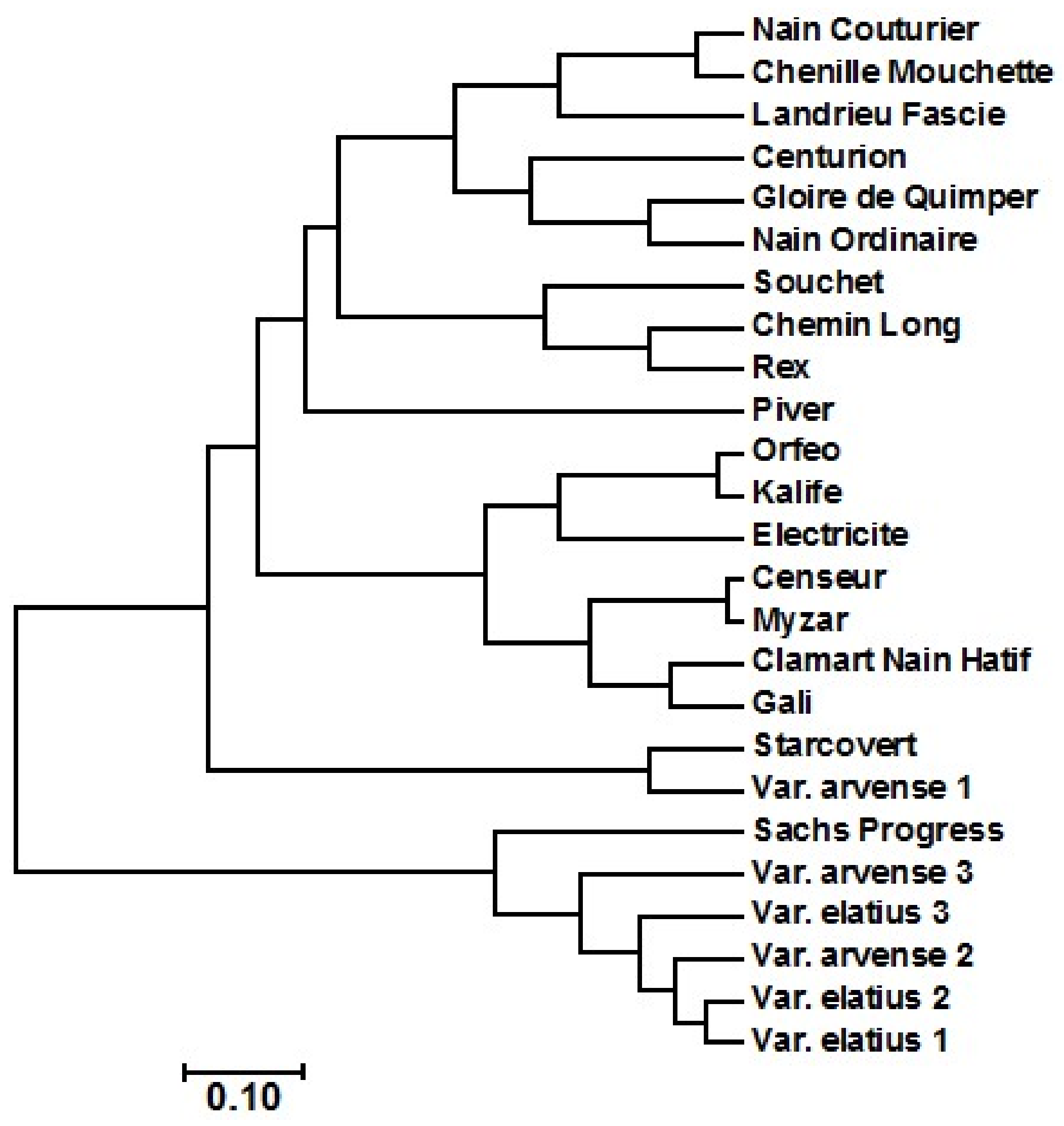
2.2. Amplified Fragment Length Polymorphism (AFLP) Diversity Analysis
2.3. Marker–Trait Association
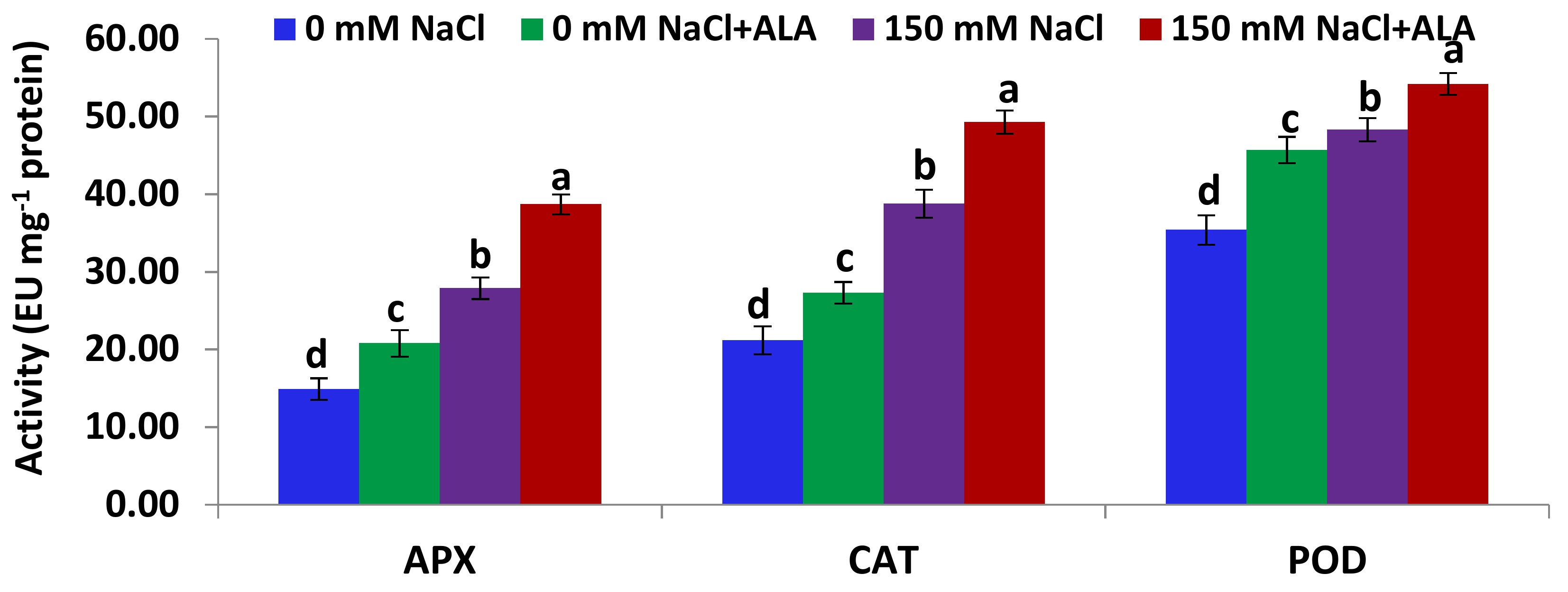
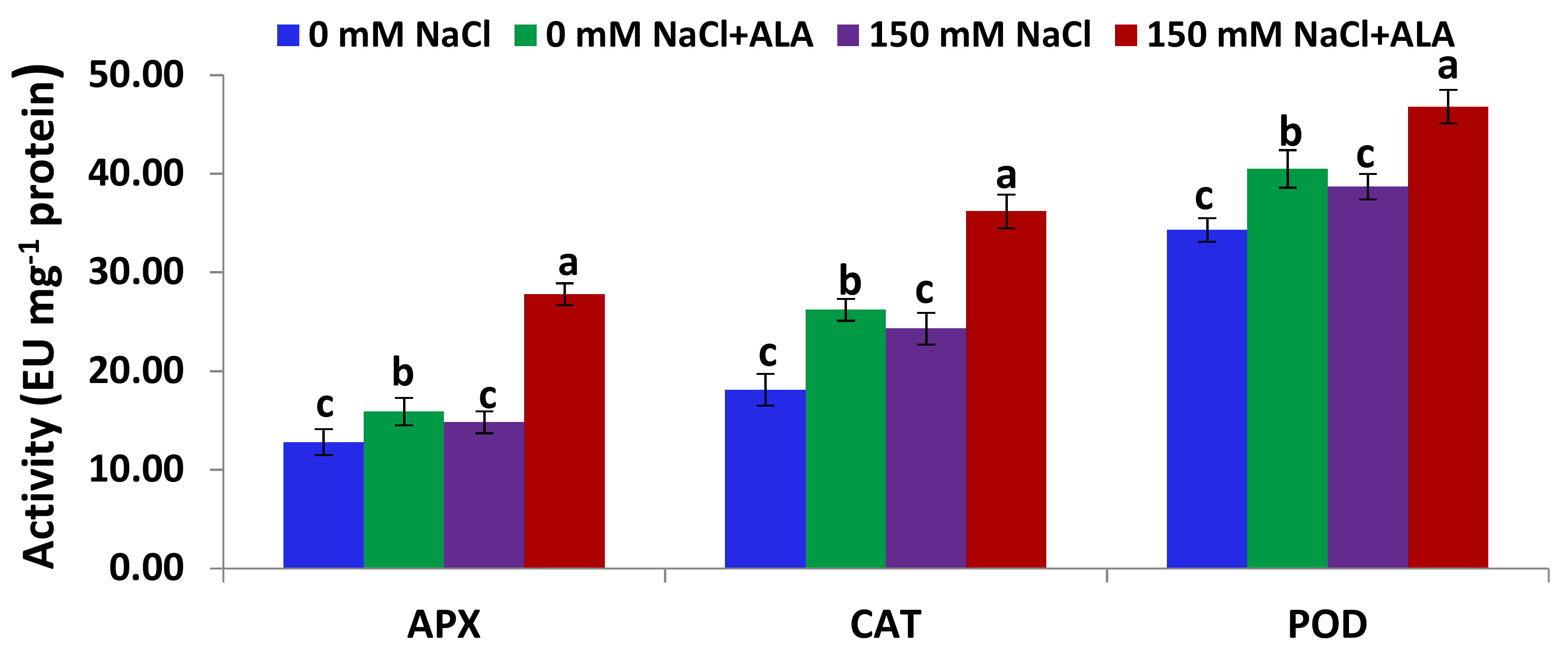
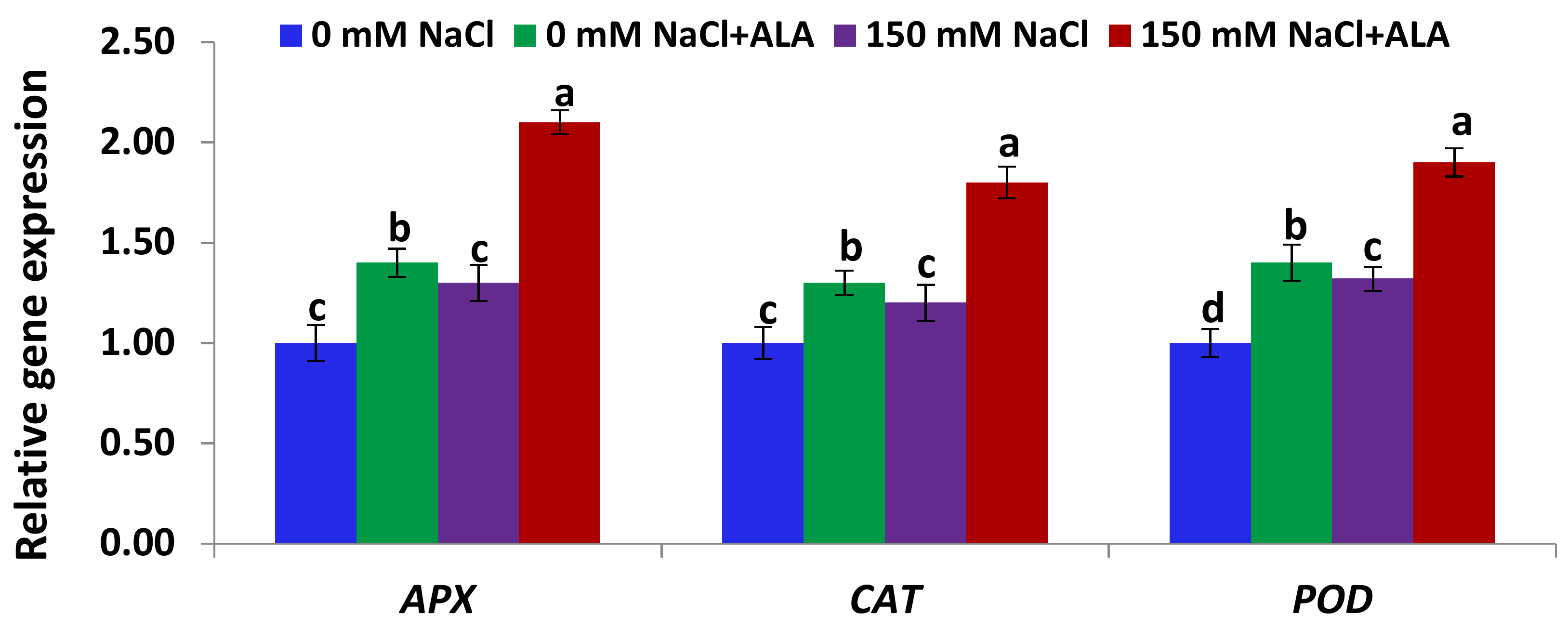
2.4. Salt Tolerance Test and the Effect of 5-Aminolevulinic Foliar Spray
3. Materials and Methods
3.1. Plant Material
3.2. Pea Oil Content Estimation and Fatty Acid Analysis
3.3. Molecular Diversity Analysis
3.3.1. DNA Extraction and AFLP Analysis
3.3.2. AFLP Data Analysis
3.4. Marker–Trait Association
3.5. Salt Tolerance Test and The Effect of 5-Aminolevulinic Foliar Spray in Pea
3.5.1. Growth Conditions and Treatments
3.5.2. Determination of Contents of Photosynthetic Pigments
3.5.3. Antioxidant Enzymes Assay
3.5.4. Gene Expression Analysis
3.5.5. Statistical Analysis
Author Contributions
Acknowledgments
Conflicts of Interest
References
- McPhee, K. Dry pea production and breeding: A mini review. J. Food Agric. Environ. 2003, 1, 64–69. [Google Scholar]
- Ahmad, S.; Kaur, S.; Lamb-Palmer, N.D.; Lefsrud, M.; Sing, J. Genetic diversity and population structure of Pisum sativum accessions for marker-trait association of lipid content. Crop J. 2015, 3, 238–245. [Google Scholar] [CrossRef]
- Yoshida, H.; Tomiyama, Y.; Tanaka, M.; Mizushina, Y. Characteristic profiles of lipid classes, fatty acids and triacylglycerol molecular species of peas (Pisum sativum L.). Eur. J. Lipid Sci. Technol. 2007, 109, 600–607. [Google Scholar] [CrossRef]
- Welch, R.W.; Griffiths, D.W. Variation in the oil content and fatty acid composition of field beans (Vicia faba) and peas (Pisum spp.). J. Sci. Food Agric. 1984, 35, 1282–1289. [Google Scholar] [CrossRef]
- Coxon, D.T.; Wright, D.J. Analysis of pea lipid content by gas chromatographic and microgravimetric methods. Genotype variation in lipid content and fatty acid composition. J. Sci. Food Agric. 1985, 36, 847–856. [Google Scholar] [CrossRef]
- Solis, M.I.V.; Patel, A.; Orsat, V.; Singh, J.; Lefsrud, M. Fatty acid profiling of the seed oils of some varieties of field peas (Pisum sativum) by RPLC/ESI-MS/MS: Towards the development of an oilseed pea. Food Chem. 2013, 139, 986–993. [Google Scholar] [CrossRef] [PubMed]
- Able, J.A.; Langridge, P.; Milligan, A.S. Capturing diversity in the cereals: Many options but little promiscuity. Trends Plant Sci. 2007, 12, 71–79. [Google Scholar] [CrossRef] [PubMed]
- El-Esawi, M.A.; Sammour, R. Karyological and phylogenetic studies in the genus Lactuca L. (asteraceae). Cytologia 2014, 79, 269–275. [Google Scholar] [CrossRef]
- El-Esawi, M.A. Genetic diversity and evolution of Brassica genetic resources: From morphology to novel genomic technologies—A review. Plant Genet. Resour. 2017, 15, 388–399. [Google Scholar] [CrossRef]
- El-Esawi, M.A.; Germaine, K.; Bourke, P.; Malone, R. Genetic diversity and population structure of Brassica oleracea germplasm in Ireland using SSR markers. C. R. Biol. 2016, 339, 133–140. [Google Scholar] [CrossRef] [PubMed]
- El-Esawi, M.A.; Germaine, K.; Bourke, P.; Malone, R. AFLP analysis of genetic diversity and phylogenetic relationships of Brassica oleracea in Ireland. C. R. Biol. 2016, 339, 163–170. [Google Scholar] [CrossRef] [PubMed]
- El-Esawi, M.A.; Mustafa, A.; Badr, S.; Sammour, R. Isozyme analysis of genetic variability and population structure of Lactuca L. germplasm. Biochem. Syst. Ecol. 2017, 70, 73–79. [Google Scholar] [CrossRef]
- El-Esawi, M.A. SSR analysis of genetic diversity and structure of the germplasm of faba bean (Vicia faba L.). C. R. Biol. 2017, 340, 474–480. [Google Scholar] [CrossRef] [PubMed]
- El-Esawi, M.A.; Witczak, J.; Abomohra, A.; Ali, H.M.; Elshikh, M.S.; Ahmad, M. Analysis of the Genetic Diversity and Population Structure of Austrian and Belgian Wheat Germplasm Within a Regional Context Based on DArT Markers. Genes 2018, 9, 47. [Google Scholar] [CrossRef] [PubMed]
- El-Esawi, M.A.; Elkelish, A.; Elansary, H.O.; Ali, H.M.; Elshikh, M.; Witczak, I.; Ahmad, M. Genetic Transformation and Hairy Root Induction Enhance the Antioxidant Potential of Lactuca serriola L. Oxidative Med. Cell. Longev. 2017, 2017, 5604746. [Google Scholar] [CrossRef] [PubMed]
- El-Esawi, M.A.; Elansary, H.O.; El-Shanhorey, N.A.; Abdel-Hamid, A.M.E.; Ali, H.M.; Elshikh, M.S. Salicylic Acid-Regulated Antioxidant Mechanisms and Gene Expression Enhance Rosemary Performance under Saline Conditions. Front. Physiol. 2017, 8, 716. [Google Scholar] [CrossRef] [PubMed]
- Vwioko, E.; Adinkwu, O.; El-Esawi, M.A. Comparative Physiological, Biochemical and Genetic Responses to Prolonged Waterlogging Stress in Okra and Maize Given Exogenous Ethylene Priming. Front. Physiol. 2017, 8, 632. [Google Scholar] [CrossRef] [PubMed]
- Consentino, L.; Lambert, S.; Martino, C.; Jourdan, N.; Bouchet, P.-E.; Witczak, J.; Castello, P.; El-Esawi, M.; Corbineau, F.; d’Harlingue, A.; et al. Blue-light dependent reactive oxygen species formation by Arabidopsis cryptochrome may defi ne a novel evolutionarily conserved signalling mechanism. New Phytol. 2015, 206, 1450–1462. [Google Scholar] [CrossRef] [PubMed]
- Elansary, H.O.; Szopa, A.; Kubica, P.; Ekiert, H.; Ali, H.M.; Elshikh, M.S.; Abdel-Salam, E.M.; El-Esawi, M.; El-Ansary, D.O. Bioactivities of traditional medicinal plants in Alexandria. Evid. Based Complement. Altern. Med. 2018, 2018, 1463579. [Google Scholar] [CrossRef] [PubMed]
- Elansary, H.O.; Yessoufou, K.; Abdel-Hamid, A.M.E.; El-Esawi, M.A.; Ali, H.M.; Elshikh, M.S. Seaweed Extracts Enhance Salam Turfgrass Performance during Prolonged Irrigation Intervals and Saline Shock. Front. Plant. Sci. 2017, 8, 830. [Google Scholar] [CrossRef] [PubMed]
- El-Esawi, M.A. Micropropagation technology and its applications for crop improvement. In Plant Tissue Culture: Propagation, Conservation and Crop Improvement; Anis, M., Ahmad, N., Eds.; Springer: Singapore, 2016; pp. 523–545. [Google Scholar]
- El-Esawi, M.A. Nonzygotic embryogenesis for plant development. In Plant Tissue Culture: Propagation, Conservation and Crop Improvement; Anis, M., Ahmad, N., Eds.; Springer: Singapore, 2016; pp. 583–598. [Google Scholar]
- El-Esawi, M.A. Somatic hybridization and microspore culture in Brassica improvement. In Plant Tissue Culture: Propagation, Conservation and Crop Improvement; Anis, M., Ahmad, N., Eds.; Springer: Singapore, 2016; pp. 599–609. [Google Scholar]
- El-Esawi, M.; Arthaut, L.; Jourdan, N.; d’Harlingue, A.; Martino, C.; Ahmad, M. Blue-light induced biosynthesis of ROS contributes to the signaling mechanism of Arabidopsis cryptochrome. Sci. Rep. 2017, 7, 13875. [Google Scholar] [CrossRef] [PubMed]
- El-Esawi, M.; Glascoe, A.; Engle, D.; Ritz, T.; Link, J.; Ahmad, M. Cellular metabolites modulate in vivo signaling of Arabidopsis cryptochrome-1. Plant Signal. Behav. 2015, 10, e1063758. [Google Scholar] [CrossRef] [PubMed]
- Jourdan, N.; Martino, C.; El-Esawi, M.; Witczak, J.; Bouchet, P.-E.; d’Harlingue, A.; Ahmad, M. Blue light dependent ROS formation by Arabidopsis Cryptochrome-2 may contribute towards its signaling role. Plant Signal. Behav. 2015, 10, e1042647. [Google Scholar] [CrossRef] [PubMed]
- Sallam, A.; Arbaoui, M.; El-Esawi, M.; Abshire, N.; Martsch, R. Identification and Verification of QTL Associated with Frost Tolerance Using Linkage Mapping and GWAS in Winter Faba Bean. Front. Plant Sci. 2016, 7, 1098. [Google Scholar] [CrossRef] [PubMed]
- Becher, S.A.; Steinmetz, K.; Weising, K.; Boury, S.; Peltier, D.; Renou, J.P.; Kahl, G.; Wolff, K. Microsatellites for cultivar identification in Pelargonium. Theor. Appl. Genet. 2000, 101, 643–651. [Google Scholar] [CrossRef]
- Burstin, J.; Deniot, G.; Potier, J.; Weinachter, C.; Aubert, G.; Baranger, A. Microsatellite polymorphism in Pisum sativum. Plant Breed. 2001, 120, 311–317. [Google Scholar] [CrossRef]
- Simioniuc, D.; Uptmoor, R.; Friedt, W.; Ordon, F. Genetic diversity and relationships among pea cultivars revealed by RAPDs and AFLPs. Plant Breed. 2002, 121, 429–435. [Google Scholar] [CrossRef]
- Loridon, K.; McPhee, K.; Morin, J.; Dubreuil, P.; Pilet-Nayel, M.L.; Aubert, G.; Rameau, C.; Baranger, A.; Coyne, C.; Lejeune-Henaut, I.; et al. Microsatellite marker polymorphism and mapping in pea (Pisum sativum L.). Theor. Appl. Genet. 2005, 111, 1022–1031. [Google Scholar] [CrossRef] [PubMed]
- Smýkal, P.; Hýbl, M.; Corander, J.; Jarkovský, J.; Flavell, A.J.; Griga, M. Genetic diversity and population structure of pea (Pisum sativum L.) varieties derived from combined retrotransposon, microsatellite and morphological marker analysis. Theor. Appl. Genet. 2008, 117, 413–424. [Google Scholar] [CrossRef] [PubMed]
- Sarikamis, G.; Yanmaz, R.; Ermis, S.; Bakir, M.; Yuksel, C. Genetic characterization of pea (Pisum sativum) germplasm from Turkey using morphological and SSR markers. Genet. Mol. Res. 2010, 9, 591–600. [Google Scholar] [CrossRef] [PubMed]
- Deulvot, C.; Charrel, H.; Marty, A.; Jacquin, F.; Donnadieu, C.; Lejeune-Hénaut, I.; Burstin, J.; Aubert, G. Highly-multiplexed SNP genotyping for genetic mapping and germplasm diversity studies in pea. BMC Genom. 2010, 11, 468. [Google Scholar] [CrossRef] [PubMed]
- Dyachenko, E.A.; Ryzhova, N.N.; Vishnyakova, M.A.; Kochieva, E.Z. Molecular Genetic Diversity of the Pea (Pisum sativum L.) from the Vavilov Research Institute Collection Detected by the AFLP Analysis. Russ. J. Genet. 2014, 50, 916–924. [Google Scholar] [CrossRef]
- Murcia, M.A.; Rincon, F. Fatty acid composition of pea (Pisum sativum L. var. Citrina) during growth. Grasas Aceites 1991, 42, 444–449. [Google Scholar] [CrossRef]
- Murcia, M.A.; Rincon, F. Size as source of variance in lipid composition of pea. Food Chem. 1992, 44, 29–35. [Google Scholar] [CrossRef]
- Yang, Z.; Chang, Z.; Sun, L.; Yu, J.; Huang, B. Physiological and metabolic effects of 5-aminolevulinic acid for mitigating salinity stress in creeping bentgrass. PLoS ONE 2014, 9, e116283. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.-P.; Miao, M.-M.; Wang, C.-L. Effects of ALA on photosynthesis, antioxidant enzyme activity, and gene expression, and regulation of proline accumulation in tomato seedlings under NaCl stress. J. Plant Growth Regul. 2015, 34, 637–650. [Google Scholar] [CrossRef]
- Liu, L.; Nguyen, N.T.; Ueda, A.; Saneoka, H. Effects of 5-aminolevulinic acid on Swiss chard (Beta vulgaris L. subsp. cicla) seedling growth under saline conditions. Plant Growth Regul. 2014, 74, 219–228. [Google Scholar] [CrossRef]
- Nunkaew, T.; Kantachote, D.; Kanzaki, H.; Nitoda, T.; Ritchie, R.J. Effects of 5-aminolevulinic acid (ALA)-containing supernatants from selected Rhodopseudomonas palustris strains on rice growth under NaCl stress, with mediating effects on chlorophyll, photosynthetic electron transport and antioxidative enzymes. Electron. J. Biotechnol. 2014, 17, 19–26. [Google Scholar] [CrossRef]
- Zhen, A.; Bie, Z.; Huang, Y.; Liu, Z.; Fan, M. Effects of 5-aminolevulinic acid on the H2O2-content and antioxidative enzyme gene expression in NaCl-treated cucumber seedlings. Biol. Plant. 2012, 56, 566–570. [Google Scholar] [CrossRef]
- Zhang, C.-P.; Li, Y.-C.; Yuan, F.-G.; Hu, S.-J.; Liu, H.-Y.; He, P. Role of 5-aminolevulinic acid in the salinity stress response of the seeds and seedlings of the medicinal plant Cassia obtusifolia L. Bot. Stud. 2013, 54, 18. [Google Scholar] [CrossRef] [PubMed]
- Aksakal, O.; Algur, O.F.; Icoglu Aksakal, F.; Aysin, F. Exogenous 5-aminolevulinic acid alleviates the detrimental effects of UV-B stress on lettuce (Lactuca sativa L.) seedlings. Acta Physiol. Plant. 2017, 39, 55. [Google Scholar] [CrossRef]
- Liu, D.; Wu, L.; Naeem, M.S.; Liu, H.; Deng, X.; Xu, L.; Zhang, F.; Zhou, W. 5-Aminolevulinic acid enhances photosynthetic gas exchange, chlorophyll fluorescence and antioxidant system in oilseed rape under drought stress. Acta Physiol. Plant. 2013, 35, 2747–2759. [Google Scholar] [CrossRef]
- Ahmad, R.; Ali, S.; Hannan, F.; Rizwan, M.; Iqbal, M.; Hassan, Z.; Akram, N.A.; Maqbool, S.; Abbas, F. Promotive role of 5-aminolevulinic acid on chromium-induced morphological, photosynthetic, and oxidative changes in cauliflower (Brassica oleracea botrytis L.). Environ. Sci. Pollut. Res. 2017, 24, 8814–8824. [Google Scholar] [CrossRef] [PubMed]
- Liu, D.; Hu, L.Y.; Ali, B.; Yang, A.G.; Wan, G.L.; Xu, L.; Zhou, W.J. Influence of 5-aminolevulinic acid on photosynthetically related parameters and gene expression in Brassica napus L. under drought stress. Soil Sci. Plant Nutr. 2016, 62, 254–262. [Google Scholar] [CrossRef]
- Vos, P.; Mogers, R.M.; Bleeker, M.; Reijans, T.; Vande, L.; Hornes, M.; Frijters, A.; Port, J.; Peleman, J.; Kliper, M.; et al. AFLP: A new technique for DNA fingerprinting. Nucleic Acids Res. 1995, 23, 4407–4417. [Google Scholar] [CrossRef] [PubMed]
- Liu, K.J.; Muse, S.V. PowerMarker: An integrated analysis environment for genetic marker analysis. Bioinformatics 2005, 21, 2128–2129. [Google Scholar] [CrossRef] [PubMed]
- Peakall, R.; Smouse, P.E. GenAlEx 6.5: Genetic analysis in Excel. Population genetic software for teaching and research—An update. Bioinformatics 2012, 28, 2537–2539. [Google Scholar] [CrossRef] [PubMed]
- Nei, M. Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 1978, 89, 583–590. [Google Scholar] [PubMed]
- Zhang, Z.; Ersoz, E.; Lai, C.-Q.; Todhunter, R.J.; Tiwari, H.K.; Gore, M.A.; Bradbury, P.J.; Yu, J.; Arnett, D.K.; Ordovas, J.M. Mixed linear model approach adapted for genome-wide association studies. Nat. Genet. 2010, 42, 355–360. [Google Scholar] [CrossRef] [PubMed]
- Bradbury, P.J.; Zhang, Z.; Kroon, D.E.; Casstevens, T.M.; Ramdoss, Y.; Buckler, E.S. TASSEL: Software for association mapping of complex traits in diverse samples. Bioinformatics 2007, 23, 2633–2635. [Google Scholar] [CrossRef] [PubMed]
- Lichtenthaler, H.K.; Wellburn, R.R. Determination of total carotenoids and chlorophylls a and b of leaf extracts in different solvents. Biochem. Soc. Trans. 1983, 11, 591–592. [Google Scholar] [CrossRef]
- Zhang, J.; Kirkham, M.B. Antioxidant responses to drought in sunflower and sorghum seedlings. New Phytol. 1996, 132, 361–373. [Google Scholar] [CrossRef] [PubMed]
- Sirhindi, G.; Mir, M.A.; Abd-Allah, E.F.; Ahmad, P.; Gucel, S. Jasmonic Acid Modulates the Physio-Biochemical Attributes, Antioxidant Enzyme Activity, and Gene Expression in Glycine max under Nickel Toxicity. Front. Plant Sci. 2016, 7, 591. [Google Scholar] [CrossRef] [PubMed]


| Cultivar/Variety | Palmitic (C16:0) | Stearic (C18:0) | Oleic (C18:1) | Linoleic (C18:2) | Linolenic (C18:3) |
|---|---|---|---|---|---|
| Censeur | 15.8 | 14.7 | 23.2 | 33.4 | 12.4 |
| Centurion | 16.7 | 15.8 | 23.3 | 33.2 | 12.2 |
| Chemin Long | 17.1 | 16.2 | 24.1 | 35.2 | 13.4 |
| Chenille Mouchette | 15.9 | 14.7 | 24.4 | 33.9 | 12.8 |
| Clamart Nain Hatif | 14.8 | 14.3 | 24.2 | 34.1 | 13.2 |
| Electricite | 16.1 | 15.7 | 23.2 | 33.4 | 12.7 |
| Gali | 17.0 | 16.1 | 22.8 | 32.1 | 14.1 |
| Gloire de Quimper | 18.2 | 14.8 | 24.3 | 32.8 | 13.2 |
| Kalife | 15.8 | 15.1 | 22.4 | 31.6 | 11.8 |
| Landrieu Fascie | 16.2 | 13.6 | 25.6 | 30.2 | 13.1 |
| Myzar | 18.4 | 15.2 | 23.7 | 32.8 | 13.3 |
| Nain Couturier | 16.4 | 15.0 | 22.8 | 31.7 | 12.7 |
| Nain Ordinaire | 16.1 | 14.9 | 24.1 | 33.2 | 12.2 |
| Orfeo | 15.6 | 14.7 | 22.8 | 33.6 | 13.4 |
| Piver | 16.8 | 15.8 | 23.3 | 32.4 | 13.6 |
| Rex | 14.9 | 13.9 | 20.6 | 29.8 | 14.0 |
| Sachs Progress | 16.3 | 15.1 | 21.8 | 31.1 | 13.9 |
| Souchet | 15.4 | 14.5 | 20.2 | 28.4 | 14.2 |
| Starcovert | 16.6 | 15.3 | 23.4 | 31.2 | 12.6 |
| Elatius 1 | 14.0 | 13.1 | 19.2 | 27.4 | 11.2 |
| Elatius 2 | 14.2 | 12.8 | 18.7 | 26.2 | 11.0 |
| Elatius 3 | 13.8 | 13.5 | 20.1 | 25.9 | 11.9 |
| Arvense 1 | 14.4 | 14.1 | 20.8 | 26.1 | 12.2 |
| Arvense 2 | 13.6 | 13.2 | 19.4 | 25.8 | 11.7 |
| Arvense 3 | 14.6 | 15.0 | 21.3 | 24.8 | 12.4 |
| AFLP Primer Sets | No. of Fragments | No. of Polymorphic Fragments | Polymorphism (%) | Gene Diversity | PIC |
|---|---|---|---|---|---|
| E-ACA/M-CGA | 86 | 54 | 63.8 | 0.52 | 0.47 |
| E-ACA/M-CTG | 89 | 51 | 57.3 | 0.58 | 0.54 |
| E-AAC/M-CAA | 51 | 35 | 68.6 | 0.43 | 0.38 |
| E-ATG/M-CAA | 56 | 33 | 58.9 | 0.39 | 0.36 |
| E-AAA/M-CAC | 71 | 47 | 66.2 | 0.51 | 0.47 |
| E-AAC/M-CAC | 37 | 23 | 62.2 | 0.38 | 0.34 |
| E-ACA/M-CAC | 51 | 41 | 80.4 | 0.49 | 0.44 |
| E-ATC/M-CAC | 62 | 44 | 70.9 | 0.37 | 0.33 |
| E-ACA/M-CAG | 41 | 21 | 51.2 | 0.28 | 0.23 |
| E-ATG/M-CAG | 31 | 19 | 61.3 | 0.21 | 0.16 |
| E-AAT/M-CAT | 29 | 15 | 51.7 | 0.35 | 0.31 |
| E-ATC/M-CAT | 65 | 40 | 61.5 | 0.41 | 0.37 |
| Mean | 55.75 | 35.25 | 62.83 | 0.41 | 0.37 |
| Cultivar/Variety | Ho | He | F | PIC |
|---|---|---|---|---|
| Censeur | 0.291 | 0.425 | 0.371 | 0.391 |
| Centurion | 0.312 | 0.392 | 0.244 | 0.352 |
| Chemin Long | 0.321 | 0.388 | 0.187 | 0.344 |
| Chenille Mouchette | 0.197 | 0.311 | 0.351 | 0.281 |
| Clamart Nain Hatif | 0.221 | 0.359 | 0.389 | 0.315 |
| Electricite | 0.198 | 0.291 | 0.304 | 0.262 |
| Gali | 0.258 | 0.321 | 0.211 | 0.282 |
| Gloire de Quimper | 0.321 | 0.424 | 0.302 | 0.374 |
| Kalife | 0.294 | 0.356 | 0.182 | 0.311 |
| Landrieu Fascie | 0.281 | 0.399 | 0.332 | 0.355 |
| Myzar | 0.265 | 0.317 | 0.151 | 0.272 |
| Nain Couturier | 0.274 | 0.392 | 0.361 | 0.348 |
| Nain Ordinaire | 0.311 | 0.477 | 0.421 | 0.416 |
| Orfeo | 0.292 | 0.411 | 0.362 | 0.372 |
| Piver | 0.231 | 0.397 | 0.441 | 0.352 |
| Rex | 0.262 | 0.225 | −0.124 | 0.188 |
| Sachs Progress | 0.254 | 0.413 | 0.458 | 0.372 |
| Souchet | 0.199 | 0.314 | 0.361 | 0.272 |
| Starcovert | 0.311 | 0.282 | −0.102 | 0.244 |
| Elatius 1 | 0.281 | 0.391 | 0.331 | 0.344 |
| Elatius 2 | 0.313 | 0.262 | −0.152 | 0.238 |
| Elatius 3 | 0.224 | 0.212 | −0.039 | 0.184 |
| Arvense 1 | 0.314 | 0.427 | 0.334 | 0.387 |
| Arvense 2 | 0.322 | 0.462 | 0.421 | 0.411 |
| Arvense 3 | 0.278 | 0.401 | 0.397 | 0.362 |
| Mean | 0.273 | 0.362 | 0.260 | 0.321 |
| NaCl Treatment | ALA (6 ppm) | Carotenoid (mg g−1 FW) | Chl a (mg g−1 FW) | Chl b (mg g−1 FW) | Total Chl (mg g−1 FW) |
|---|---|---|---|---|---|
| 0 mM | −ALA | 0.28 ± 0.09 c | 2.27 ± 0.14 c | 1.21 ± 0.08 c | 3.48 ± 0.11 c |
| +ALA | 0.38 ± 0.06 a | 2.46 ± 0.18 a | 1.36 ± 0.05 a | 3.82 ± 0.12 a | |
| 150 mM | −ALA | 0.25 ± 0.08 c | 2.21 ± 0.14 c | 1.16 ± 0.07 c | 3.37 ± 0.15 c |
| +ALA | 0.31 ± 0.09 b | 2.29 ± 0.11 b | 1.28 ± 0.05 b | 3.57 ± 0.13 b |
| NaCl Treatment | ALA (6 ppm) | Carotenoid (mg g−1 FW) | Chl a (mg g−1 FW) | Chl b (mg g−1 FW) | Total Chl (mg g−1 FW) |
|---|---|---|---|---|---|
| 0 mM | −ALA | 0.26 ± 0.05 b | 2.18 ± 0.12 b | 1.16 ± 0.07 b | 3.34 ± 0.10 b |
| +ALA | 0.31 ± 0.06 a | 2.42 ± 0.11 a | 1.52 ± 0.09 a | 3.94 ± 0.15 a | |
| 150 mM | −ALA | 0.18 ± 0.07 c | 1.72 ± 0.12 d | 0.94 ± 0.04 d | 2.66 ± 0.12 c |
| + ALA | 0.25 ± 0.05 b | 1.95 ± 0.10 c | 1.13 ± 0.07 c | 2.08 ± 0.11 d |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
El-Esawi, M.A.; Al-Ghamdi, A.A.; Ali, H.M.; Alayafi, A.A.; Witczak, J.; Ahmad, M. Analysis of Genetic Variation and Enhancement of Salt Tolerance in French Pea (Pisum Sativum L.). Int. J. Mol. Sci. 2018, 19, 2433. https://doi.org/10.3390/ijms19082433
El-Esawi MA, Al-Ghamdi AA, Ali HM, Alayafi AA, Witczak J, Ahmad M. Analysis of Genetic Variation and Enhancement of Salt Tolerance in French Pea (Pisum Sativum L.). International Journal of Molecular Sciences. 2018; 19(8):2433. https://doi.org/10.3390/ijms19082433
Chicago/Turabian StyleEl-Esawi, Mohamed A., Abdullah A. Al-Ghamdi, Hayssam M. Ali, Aisha A. Alayafi, Jacques Witczak, and Margaret Ahmad. 2018. "Analysis of Genetic Variation and Enhancement of Salt Tolerance in French Pea (Pisum Sativum L.)" International Journal of Molecular Sciences 19, no. 8: 2433. https://doi.org/10.3390/ijms19082433
APA StyleEl-Esawi, M. A., Al-Ghamdi, A. A., Ali, H. M., Alayafi, A. A., Witczak, J., & Ahmad, M. (2018). Analysis of Genetic Variation and Enhancement of Salt Tolerance in French Pea (Pisum Sativum L.). International Journal of Molecular Sciences, 19(8), 2433. https://doi.org/10.3390/ijms19082433






