Comparative Transcriptome Profiling of Two Tomato Genotypes in Response to Potassium-Deficiency Stress
Abstract
1. Introduction
2. Results
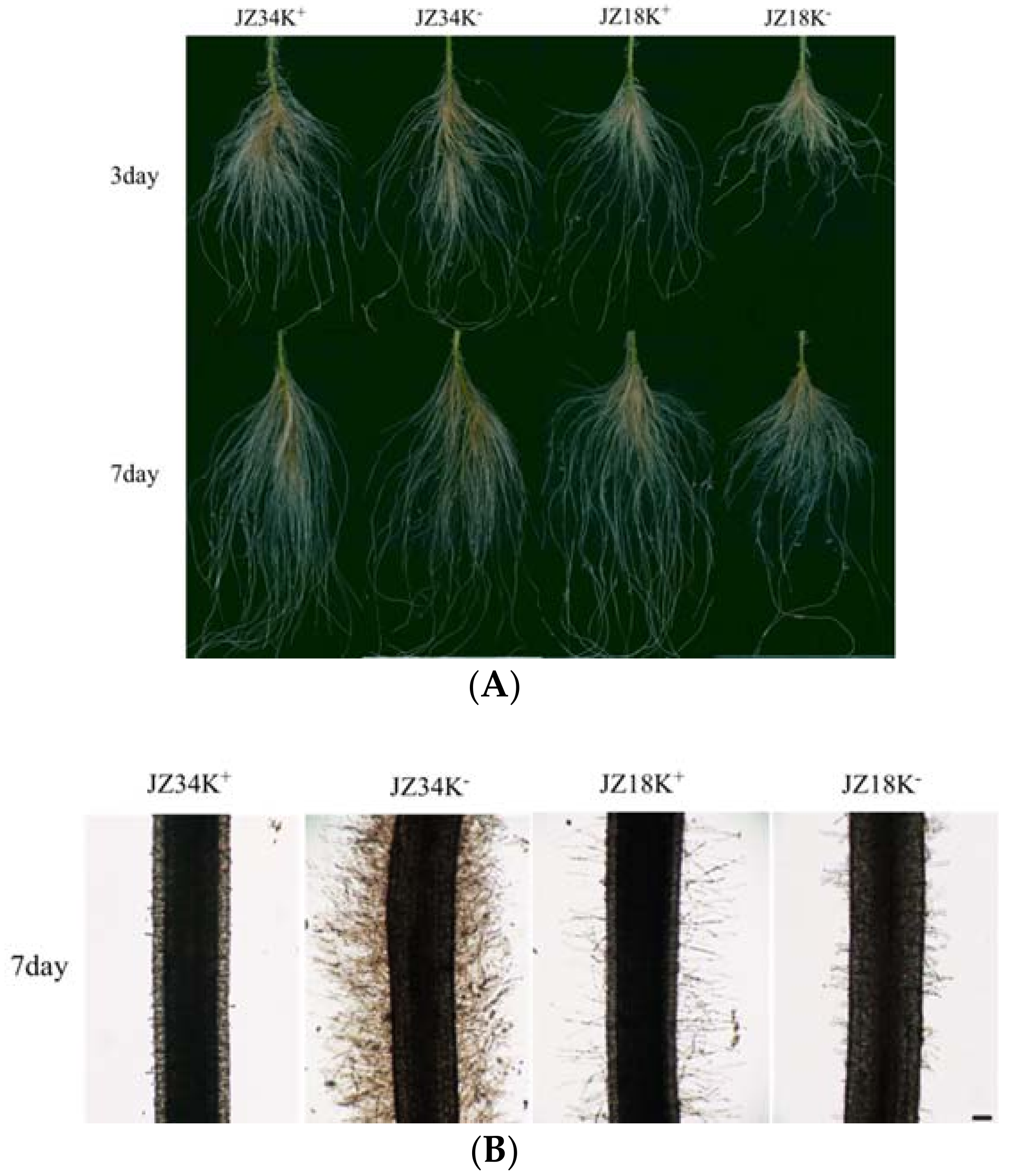
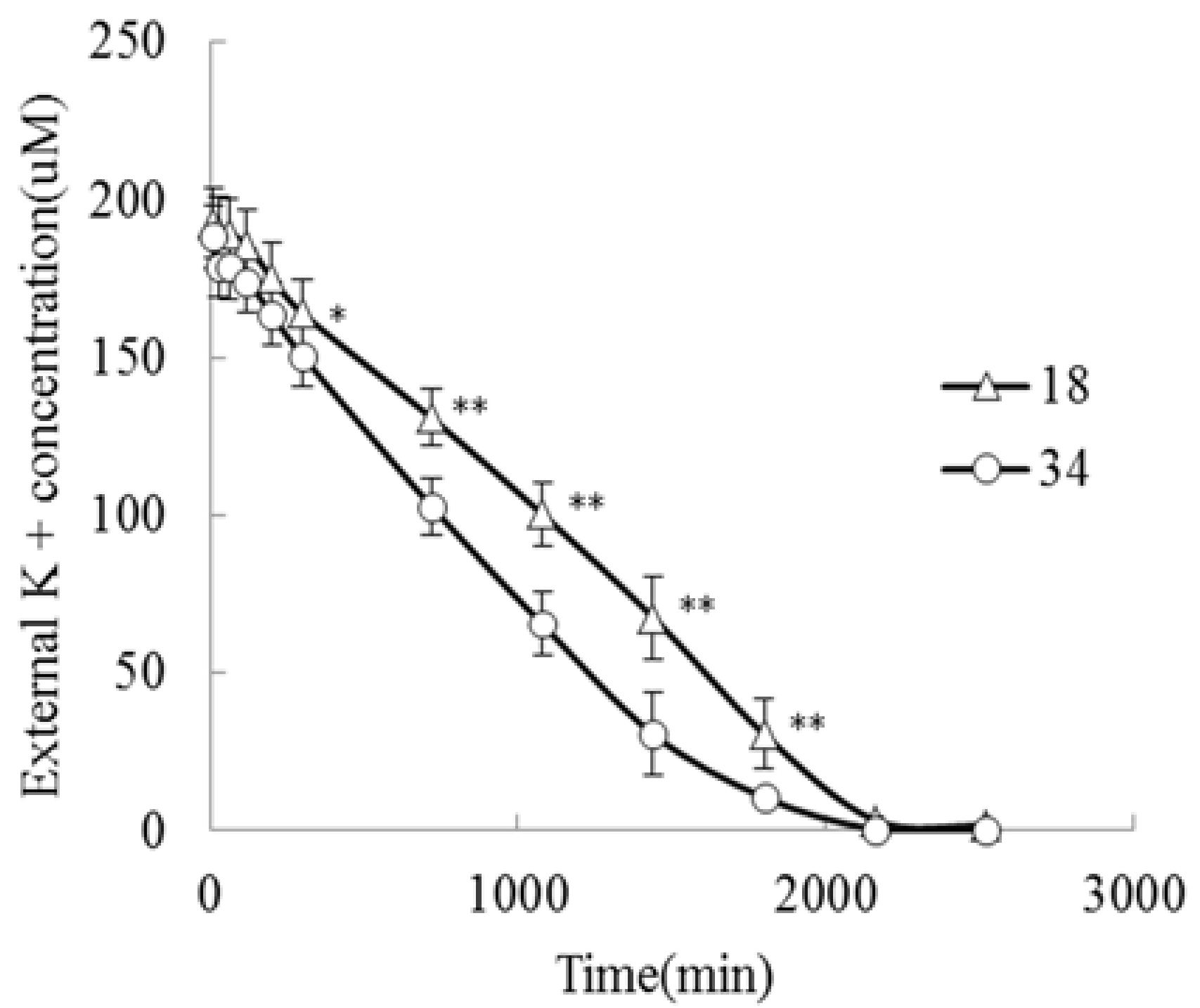
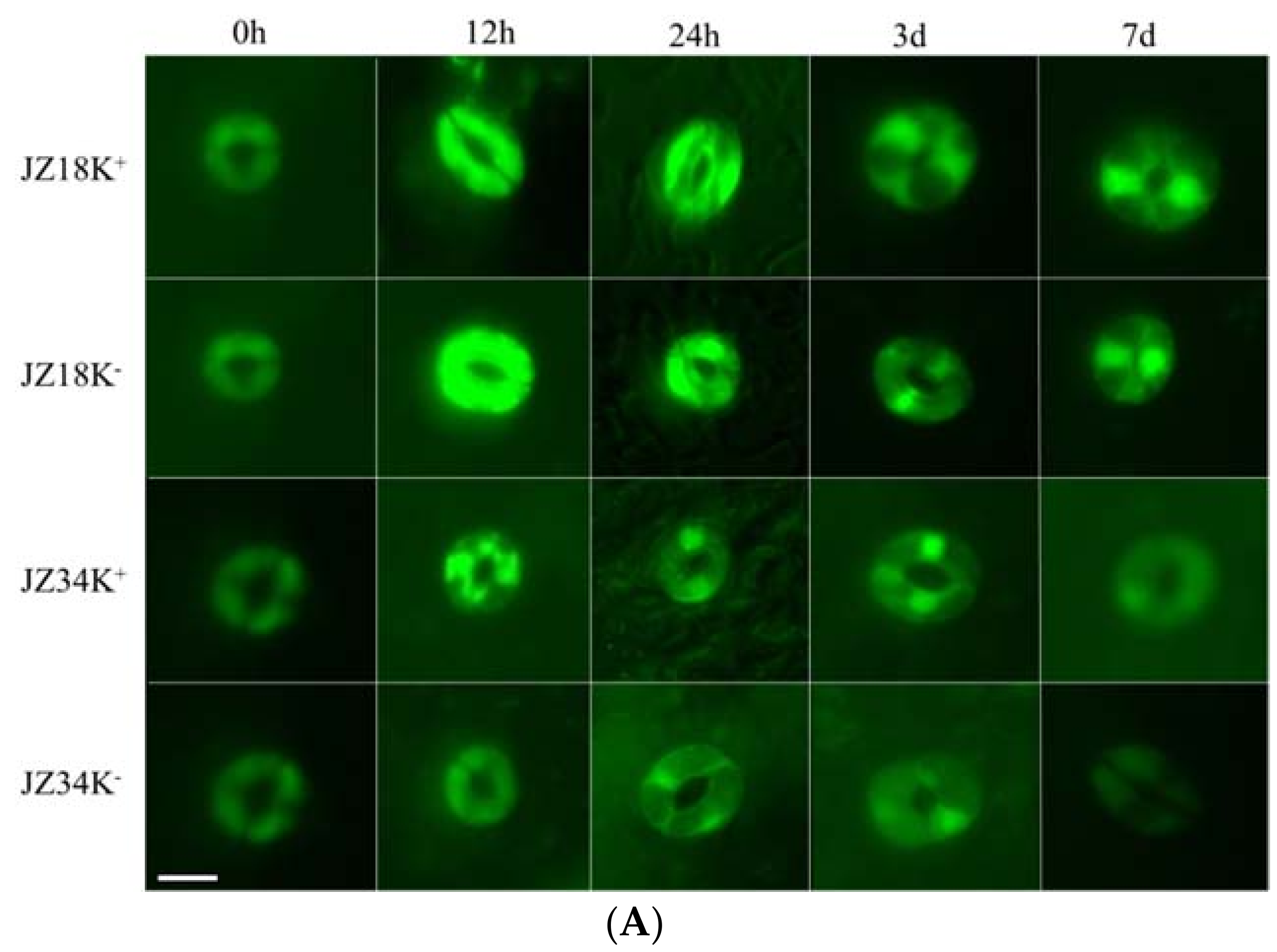
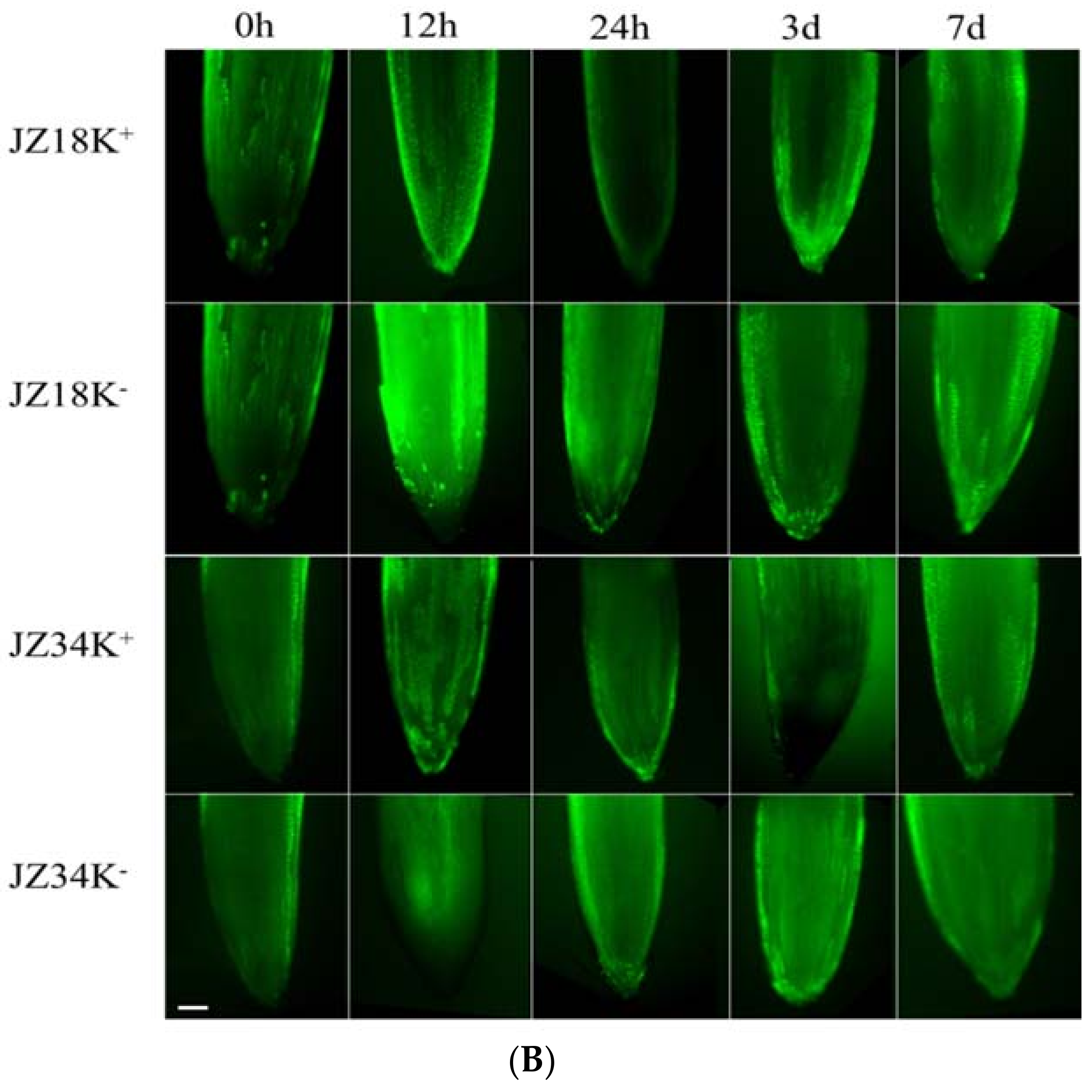
2.1. Analysis of Root Systems in JZ34 and JZ18 under K+-Deficient Conditions
2.2. Overview of the Gene Expression Profile Sequence Data and Mapping Results
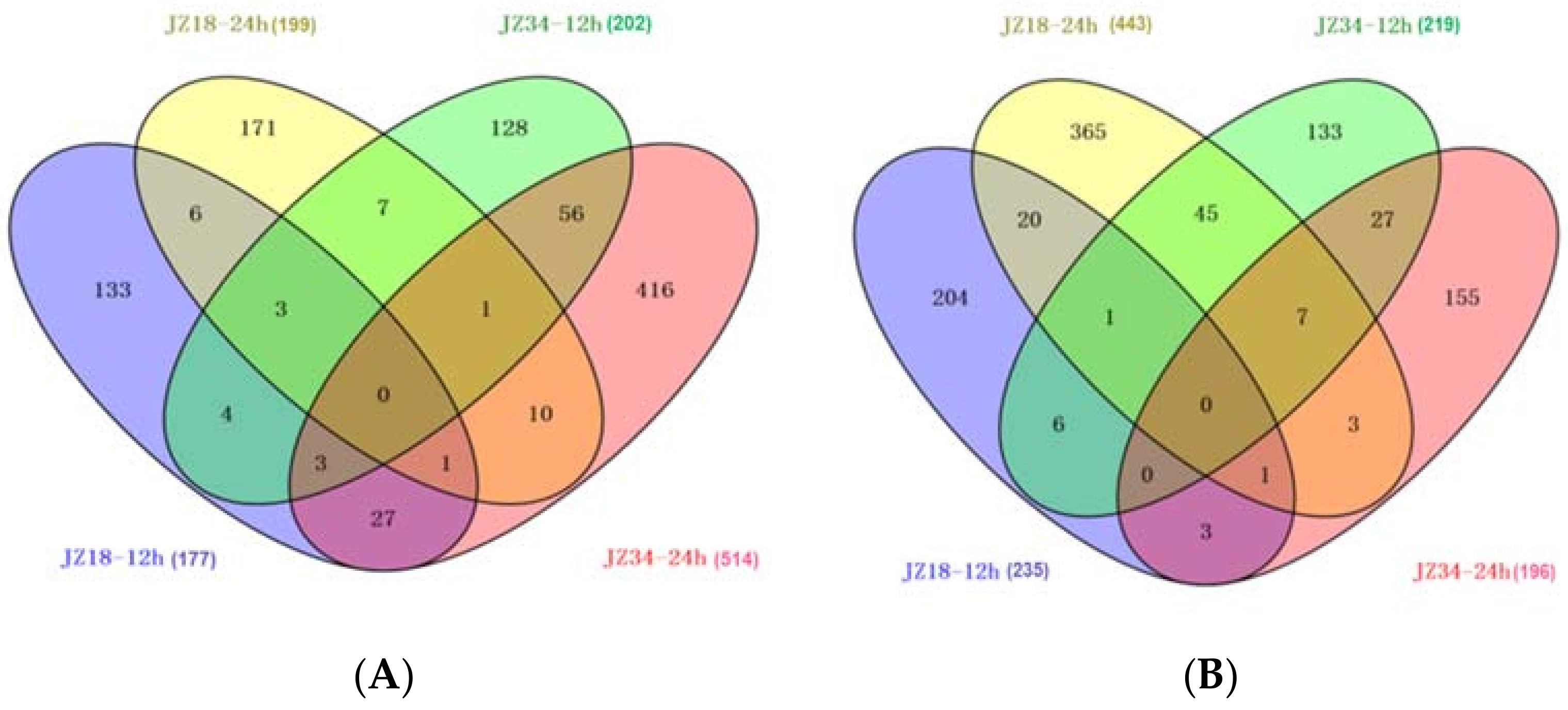
2.3. Global Analysis of DEGs
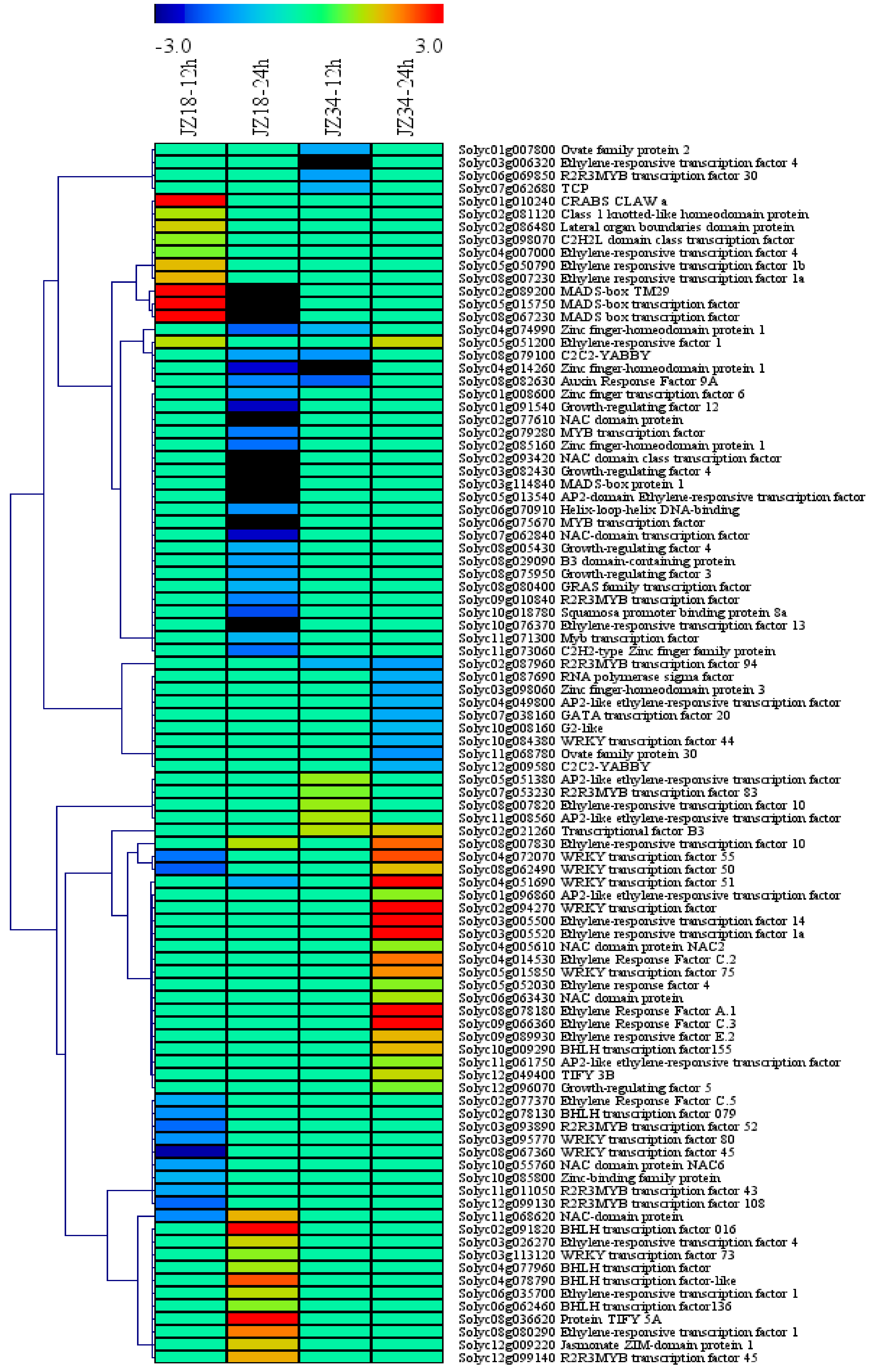
2.4. TFs in the DEGs
2.5. DEGs Encoding Transporters and Kinases
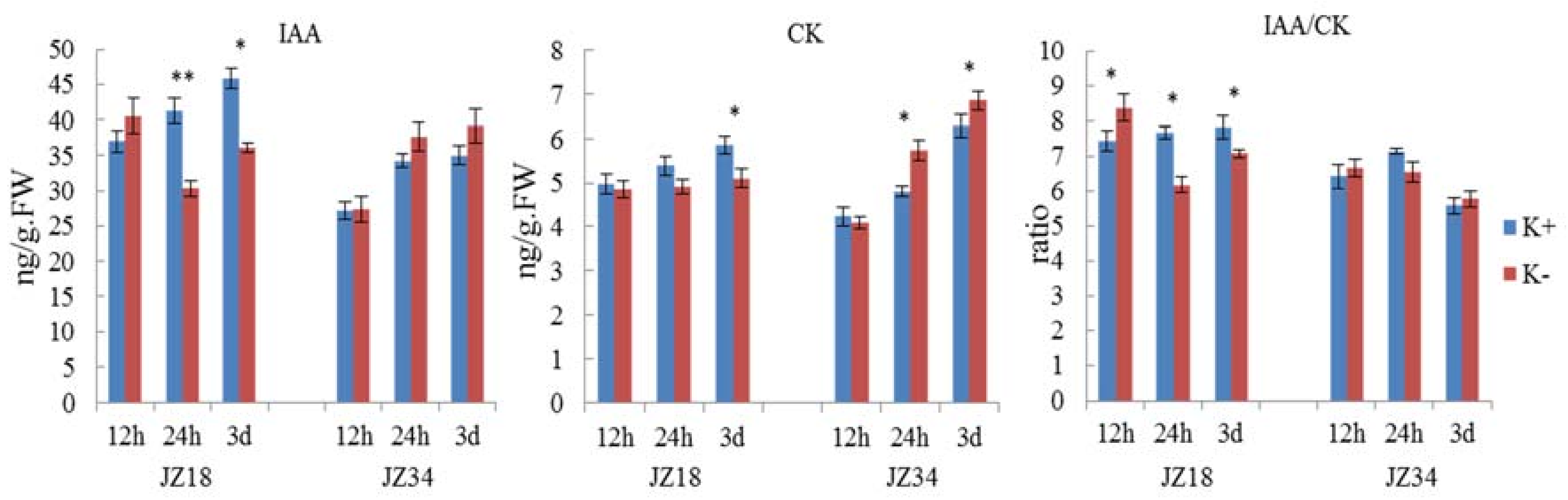
2.6. Differences in the Oxidative Stress, Hormone and Glycometabolism Analyses in JZ34JZ34 and JZ18 under K+ Deficiency
2.7. DEGs Associated with Root Architecture
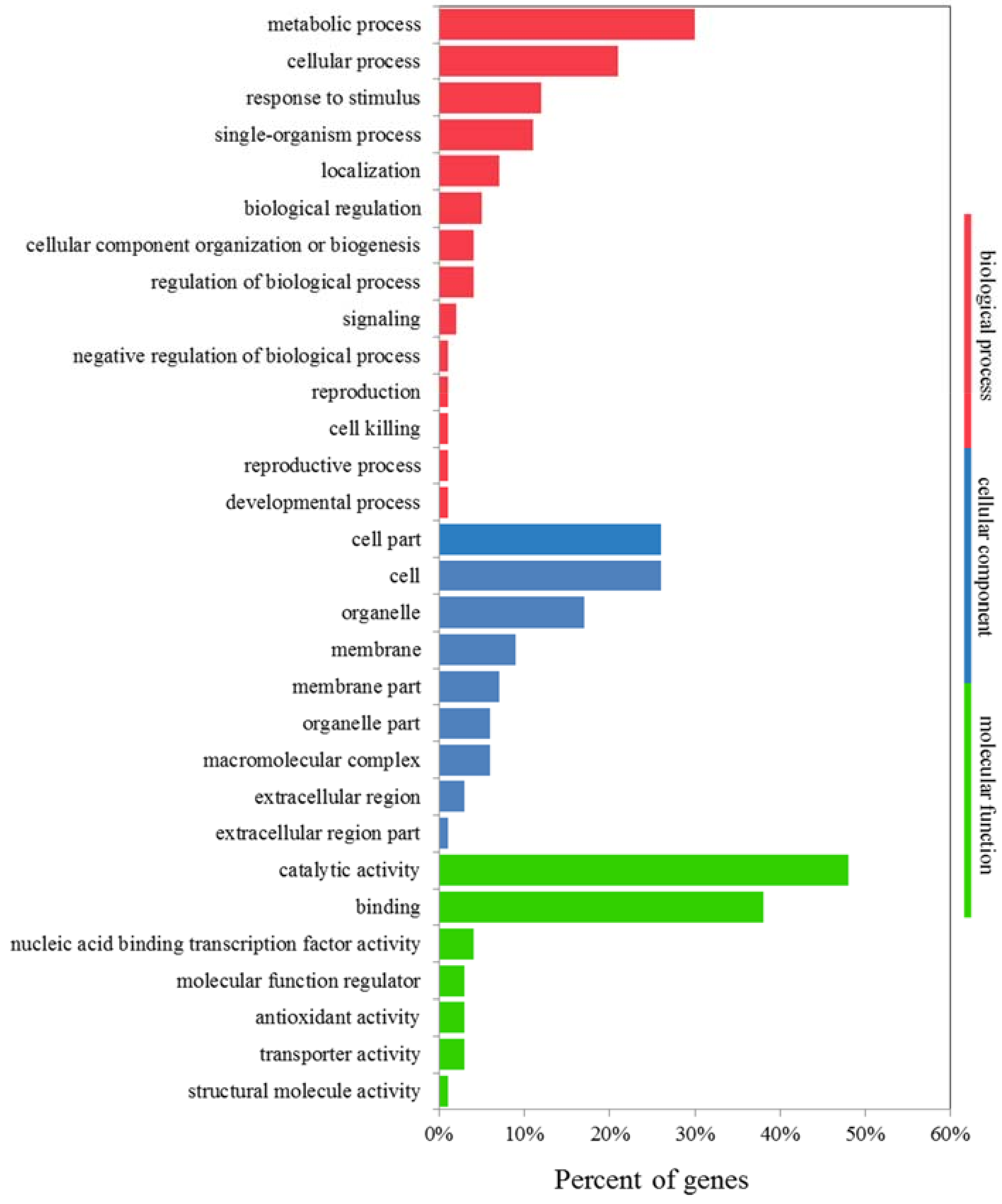
2.8. GO and KEGG Analyses of K+-Deficiency Stress Tolerance-Related DEGs
3. Discussion
3.1. K+ Transporter Expression Results in Differences in Tolerance to K+ Deficiency between the Genotypes
3.2. Root-Related Genes, the ROS Signaling Pathway, and ERFs Cause Changes in Root Growth under K+ D Eficiency
3.3. K+ Deficiency between the Genotypes Leads to Changes in Hormonal Responses and Biosynthesis-Related DEGs
3.4. Glycometabolism and Transport Involved in K+-D Eficiency Stress
4. Materials and Methods
4.1. Plant Materials and K+ Deficiency
4.2. Measurement of K+ Uptake Kinetics
4.3. RNA-seq Sampling and RNA Isolation
4.4. Library Construction, Sequencing, and Data Processing
4.5. Identification of DEGs
4.6. Gene Annotation, GO Enrichment and KEGG Analysis
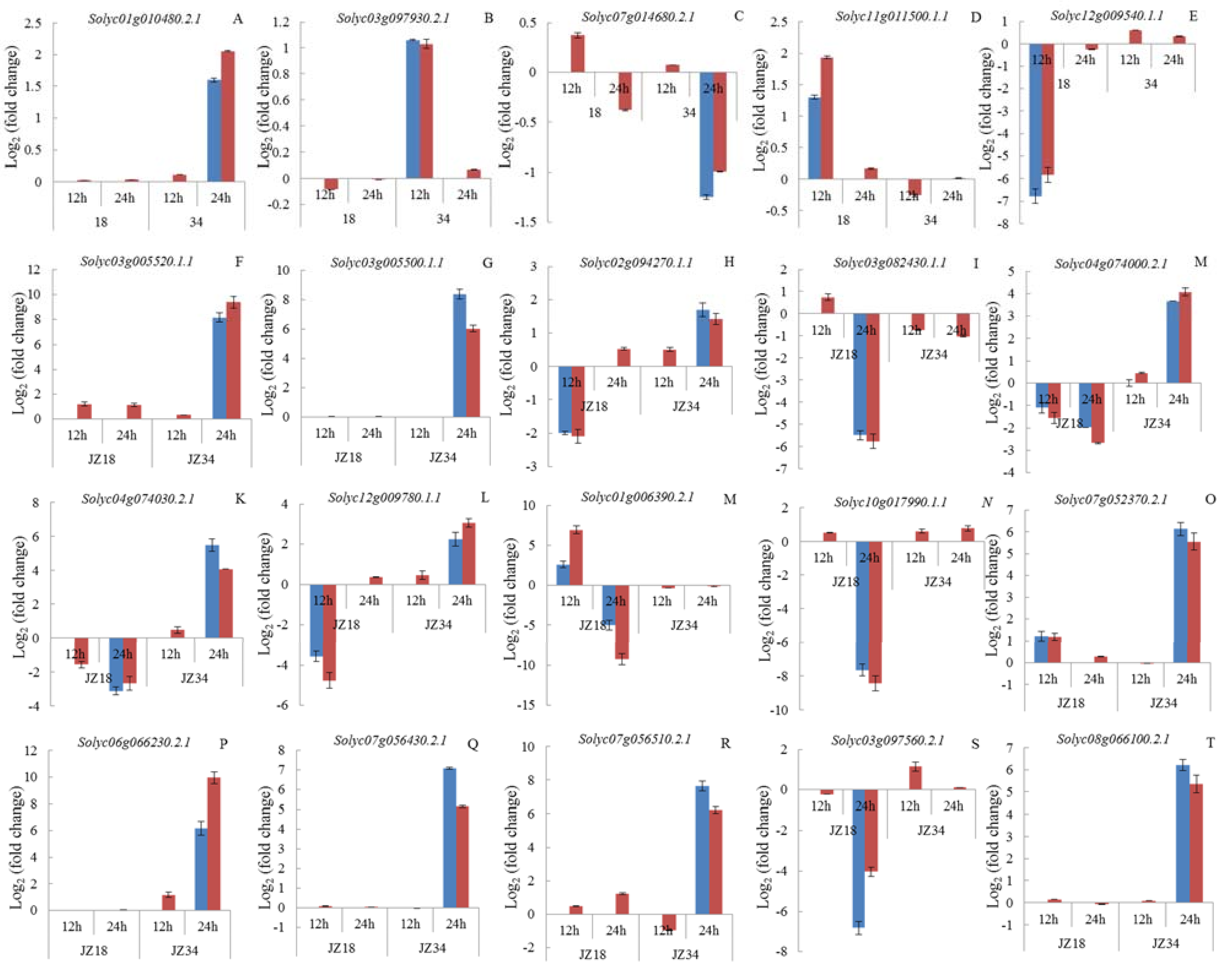
4.7. qRT-PCR Analysis
4.8. Determination of ROS
4.9. Endogenous Hormone Content Determination
4.10. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| ABA | Abscisic acid |
| AP2/ERF | Apetala 2 DNA binding domain transcription factor/ethylene-responsive factor |
| ARF | Auxin response factor |
| BGISEQ-500 | Big sequencing system-500 |
| bHLH | Basic helix-loop-helix |
| BLAST | Basic local alignment search tool |
| cPAS | Combinatorial probe-anchor synthesis |
| CK | Cytokinin |
| CKX | Cytokinin oxidase genes |
| CPA | Cation proton antiporters |
| DEGs | Differentially expressed genes |
| DNB | DNA nanoballs |
| ERF | Ethylene-responsive factor |
| EDTA | Ethylenediamine tetraacetic acid |
| FDR | False discovery rate |
| FPKM | Fragments per kilobase of exon model per million mapped reads |
| GO | Gene ontology |
| GA | Gibberellins |
| GAPDH | Glyceraldehyde-3-phosphate dehydrogenase |
| GPX | Glutathione peroxidase |
| IAA | Auxin |
| Imax | Ion maximum absorption rate |
| Km | Michaelis-Menten constant |
| KEGG | Kyoto encyclopedia of genes and genomes |
| KUE | K use efficiency |
| LR | Lateral root |
| LRR-RLKs | Leucine-rich repeat receptor-like protein kinases |
| GRF | Growth-regulating factor genes |
| OFP | Ovate family proteins |
| RH | Relative humidity |
| PIN | PIN-formed |
| ROS | Reactive oxygen species |
| SWEET | Sugars will eventually be exported transporter |
| SKOR | Stelar K+ outward rectifying channel |
| TFs | Transcription factors. |
References
- Besford, R.; Maw, G. Effect of potassium nutrition on tomato plant growth and fruit development. Plant Soil 1975, 42, 395–412. [Google Scholar] [CrossRef]
- Pujos, A.; Morard, P. Effects of potassium deficiency on tomato growth and mineral nutrition at the early production stage. Plant Soil 1997, 189, 189–196. [Google Scholar] [CrossRef]
- Kanai, S.; Moghaieb, R.E.; El-Shemy, H.A.; Panigrahi, R.; Mohapatra, P.K.; Ito, J.; Nguyen, N.T.; Saneoka, H.; Fujita, K. Potassium deficiency affects water status and photosynthetic rate of the vegetative sink in green house tomato prior to its effects on source activity. Plant Sci. 2011, 180, 368–374. [Google Scholar] [CrossRef] [PubMed]
- Sheldrick, W.F.; Syers, J.K.; Lingard, J. Soil nutrient audits for China to estimate nutrient balances and output/input relationships. Agric. Ecosyst. Environ. 2003, 94, 341–354. [Google Scholar] [CrossRef]
- Schwarz, D.; Öztekin, G.B.; Tüzel, Y.; Brückner, B.; Krumbein, A. Rootstocks can enhance tomato growth and quality characteristics at low potassium supply. Sci. Hortic. 2013, 149, 70–79. [Google Scholar] [CrossRef]
- Jungk, A.; Claassen, N. Ion diffusion in the soil-root system. Adv. Agron. (USA) 1997, 61, 53–110. [Google Scholar]
- Hussain, Z.; Arshad, M. Relationship among root characteristics and differential potassium uptake and use efficiency of selected cotton genotypes under potassium deficiency stress. Pak. J. Bot. 2011, 43, 1831–1935. [Google Scholar]
- Jia, Y.-B.; Yang, X.-E.; Feng, Y.; Jilani, G. Differential response of root morphology to potassium deficient stress among rice genotypes varying in potassium efficiency. J. Zhejiang Univ. Sci. B 2008, 9, 427. [Google Scholar] [CrossRef] [PubMed]
- Pyo, Y.J.; Gierth, M.; Schroeder, J.I.; Cho, M.H. High-affinity K+ transport in Arabidopsis: AtHAK5 and AKT1 are vital for seedling establishment and postgermination growth under low-potassium conditions. Plant Physiol. 2010, 153, 863–875. [Google Scholar] [CrossRef] [PubMed]
- Hyun, T.K.; Rim, Y.; Kim, E.; Kim, J.S. Genome-wide and molecular evolution analyses of the KT/HAK/KUP family in tomato (Solanum lycopersicum L.). Genes Genom. 2014, 36, 365–374. [Google Scholar] [CrossRef]
- Pettersson, S.; Jensen, P. Variation among species and varieties in uptake and utilization of potassium. Plant Soil 1983, 72, 231–237. [Google Scholar] [CrossRef]
- Zeng, J.; He, X.; Wu, D.; Zhu, B.; Cai, S.; Nadira, U.A.; Jabeen, Z.; Zhang, G. Comparative Transcriptome Profiling of Two Tibetan Wild Barley Genotypes in Responses to Low Potassium. PLoS ONE 2014, 9, e100567. [Google Scholar] [CrossRef] [PubMed]
- Gerardeaux, E.; Jordan-Meille, L.; Constantin, J.; Pellerin, S.; Dingkuhn, M. Changes in plant morphology and dry matter partitioning caused by potassium deficiency in Gossypium hirsutum (L.). Environ. Exp. Bot. 2011, 67, 451–459. [Google Scholar] [CrossRef]
- Zhao, X.M.; Jiang, J.; Zhang, Y.C. Different tomato strains growth and development affected by K+ deficiency stress. Jiangsu Agric. Sci. 2011, 39, 219–223. [Google Scholar]
- Armengaud, P.; Breitling, R.; Amtmann, A. The potassium-dependent transcriptome of Arabidopsis reveals a prominent role of jasmonic acid in nutrient signaling. Plant Physiol. 2004, 136, 2556–2576. [Google Scholar] [CrossRef] [PubMed]
- Ma, T.-L.; Wu, W.-H.; Wang, Y. Transcriptome analysis of rice root responses to potassium deficiency. BMC Plant Biol. 2012, 12, 161. [Google Scholar] [CrossRef] [PubMed]
- Lu, L.; Chen, Y.; Lu, L.; Lu, Y.; Li, L. Transcriptome analysis reveals dynamic changes in the gene expression of tobacco seedlings under low potassium stress. J. Genet. 2015, 94, 397–406. [Google Scholar] [CrossRef] [PubMed]
- Shen, C.; Wang, J.; Shi, X.; Kang, Y.; Xie, C.; Peng, L.; Dong, C.; Shen, Q.; Xu, Y. Transcriptome Analysis of Differentially Expressed Genes Induced by Low and High Potassium Levels Provides Insight into Fruit Sugar Metabolism of Pear. Front. Plant Sci. 2017, 8, 938. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Jiang, H.; Wang, H.; Cui, J.; Wang, J.; Hu, J.; Guo, L.; Qian, Q.; Xue, D. Transcriptome Analysis of Rice Seedling Roots in Response to Potassium Deficiency. Sci. Rep. 2017, 7, 5523. [Google Scholar] [CrossRef] [PubMed]
- Claassen, N.; Barber, S.A. A method for characterizing the relation between nutrient concentration and flux into roots of intact plants. Plant Physiol. 1974, 54, 564–568. [Google Scholar] [CrossRef] [PubMed]
- Rengel, Z.; Damon, P.M. Crops and genotypes differ in efficiency of potassium uptake and use. Physiol. Plant. 2008, 133, 624–636. [Google Scholar] [CrossRef] [PubMed]
- Epstein, E.; Rains, D.; Elzam, O. Resolution of dual mechanisms of potassium absorption by barley roots. Proc. Natl. Acad. Sci. USA 1963, 49, 684–692. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Cheng, N.-H.; Motes, C.M.; Blancaflor, E.B.; Moore, M.; Gonzales, N.; Padmanaban, S.; Sze, H.; Ward, J.M.; Hirschi, K.D. AtCHX13 is a plasma membrane K+ transporter. Plant Physiol. 2008, 148, 796–807. [Google Scholar] [CrossRef] [PubMed]
- Gaymard, F.; Pilot, G.; Lacombe, B.; Bouchez, D.; Bruneau, D.; Boucherez, J.; Michaux-Ferriere, N.; Thibaud, J.-B.; Sentenac, H. Identification and disruption of a plant shaker-like outward channel involved in K+ release into the xylem sap. Cell 1998, 94, 647–655. [Google Scholar] [CrossRef]
- Nakamura, R.L.; McKendree, W.L., Jr.; Hirsch, R.E.; Sedbrook, J.C.; Gaber, R.F.; Sussman, M.R. Expression of an Arabidopsis potassium channel gene in guard cells. Plant Physiol. 1995, 109, 371–374. [Google Scholar] [CrossRef] [PubMed]
- Sattelmacher, B.; Horst, W.J.; Becker, H.C. Factors that contribute to genetic variation for nutrient efficiency of crop plants. J. Plant Nutr. Soil Sci. 1994, 157, 215–224. [Google Scholar] [CrossRef]
- Yang, X.; Liu, J.; Wang, W.; Li, H.; Luo, A.; Ye, Z.; Yang, Y. Genotypic differences and some associated plant traits in potassium internal use efficiency of lowland rice (Oryza sativa L.). Nutr. Cycl. Agroecosyst. 2003, 67, 273–282. [Google Scholar] [CrossRef]
- Chen, J.; Gabelman, W. Isolation of tomato strains varying in potassium acquisition using a sand-zeolite culture system. Plant Soil 1995, 176, 65–70. [Google Scholar] [CrossRef]
- Hafsi, C.; Romero-Puertas, M.C.; Luis, A.; Abdelly, C.; Sandalio, L.M. Antioxidative response of Hordeum maritimum L. to potassium deficiency. Acta Physiol. Plant. 2011, 33, 193–202. [Google Scholar] [CrossRef]
- Brouder, S.; Cassman, K. Root development of two cotton cultivars in relation to potassium uptake and plant growth in a vermiculitic soil. Field Crops Res. 1990, 23, 187–203. [Google Scholar] [CrossRef]
- Tanimoto, M.; Roberts, K.; Dolan, L. Ethylene is a positive regulator of root hair development in Arabidopsis thaliana. Plant J. 1995, 8, 943–948. [Google Scholar] [CrossRef] [PubMed]
- Jung, J.-Y.; Shin, R.; Schachtman, D.P. Ethylene mediates response and tolerance to potassium deprivation in Arabidopsis. Plant Cell 2009, 21, 607–621. [Google Scholar] [CrossRef] [PubMed]
- Shin, R.; Schachtman, D.P. Hydrogen peroxide mediates plant root cell response to nutrient deprivation. Proc. Natl. Acad. Sci. USA 2004, 101, 8827–8832. [Google Scholar] [CrossRef] [PubMed]
- Halušková, L.U.; Valentovičová, K.; Huttová, J.; Mistrík, I.; Tamás, L. Effect of abiotic stresses on glutathione peroxidase and glutathione S-transferase activity in barley root tips. Plant Physiol. Biochem. 2009, 47, 1069–1074. [Google Scholar] [CrossRef] [PubMed]
- Manzano, C.; Pallerobaena, M.; Casimiro, I.; De, R.B.; Ormanligeza, B.; Van, I.G.; Beeckman, T.; Draye, X.; Casero, P.; Del Pozo, J.C. The Emerging Role of Reactive Oxygen Species Signaling during Lateral Root Development. Plant Physiol. 2014, 165, 1105–1119. [Google Scholar] [CrossRef] [PubMed]
- Passaia, G.; Queval, G.; Bai, J.; Margispinheiro, M.; Foyer, C.H. The effects of redox controls mediated by glutathione peroxidases on root architecture in Arabidopsis thaliana. J. Exp. Bot. 2014, 65, 1403–1413. [Google Scholar] [CrossRef] [PubMed]
- Vegamás, I.; Pérezdelgado, C.M.; Marino, D.; Fuertesmendizábal, T.; Gonzálezmurua, C.; Márquez, A.J.; Betti, M.; Estavillo, J.M.; Gonzálezmoro, M.B. Elevated CO2 induces root defensive mechanisms in tomato plants when dealing with ammonium toxicity. Plant Cell Physiol. 2017, 58, 2112–2125. [Google Scholar] [CrossRef] [PubMed]
- Ljung, K. Auxin metabolism and homeostasis during plant development. Development 2013, 140, 943–950. [Google Scholar] [CrossRef] [PubMed]
- Ganguly, A.; Sangho, L.; Misuk, C.; Okran, L.; Heejin, Y.; Hyungtaeg, C. Differential auxin-transporting activities of PIN-FORMED proteins in Arabidopsis root hair cells. Plant Physiol. 2010, 153, 1046–1061. [Google Scholar] [CrossRef] [PubMed]
- Evans, N.H. Modulation of guard cell plasma membrane potassium currents by methyl jasmonate. Plant Physiol. 2003, 131, 8–11. [Google Scholar] [CrossRef] [PubMed]
- Blomster, T.; Salojärvi, J.; Sipari, N.; Brosché, M.; Ahlfors, R.; Keinänen, M.; Overmyer, K.; Kangasjärvi, J. Apoplastic Reactive Oxygen Species Transiently Decrease Auxin Signaling and Cause Stress-Induced Morphogenic Response in Arabidopsis. Plant Physiol. 2011, 157, 1866–1883. [Google Scholar] [CrossRef] [PubMed]
- Werner, T.; Nehnevajova, E.; Köllmer, I.; Novák, O.; Strnad, M.; Krämer, U.; Schmülling, T. Root-Specific Reduction of Cytokinin Causes Enhanced Root Growth, Drought Tolerance, and Leaf Mineral Enrichment in Arabidopsis and Tobacco. Plant Cell 2010, 22, 3905–3920. [Google Scholar] [CrossRef] [PubMed]
- Riefler, M.; Novak, O.; Strnad, M.; Schmülling, T. Arabidopsis cytokinin receptor mutants reveal functions in shoot growth, leaf senescence, seed size, germination, root development, and cytokinin metabolism. Plant Cell 2006, 18, 40–54. [Google Scholar] [CrossRef] [PubMed]
- Raskin, I. Role of Salicylic Acid in Plants. Annu. Rev. Plant Physiol. 1992, 43, 439–463. [Google Scholar] [CrossRef]
- Gutiérrez-Coronado, M.A.; Trejo-López, C.; Larqué-Saavedra, A. Effects of salicylic acid on the growth of roots and shoots in soybean. Plant Physiol. Biochem. 1998, 36, 563–565. [Google Scholar] [CrossRef]
- Malamy, J.; Carr, J.P.; Klessig, D.F.; Raskin, I. Salicylic Acid: A Likely Endogenous Signal in the Resistance Response of Tobacco to Viral Infection. Science 1990, 250, 1002–1004. [Google Scholar] [CrossRef] [PubMed]
- Proels, R.K.; Hückelhoven, R. Cell-wall invertases, key enzymes in the modulation of plant metabolism during defence responses. Mol. Plant Pathol. 2014, 15, 858–864. [Google Scholar] [CrossRef] [PubMed]
- Sleiman, M.S.B.; Schüpfer, F.; Lemaitre, B. Transforming Growth Factor β/Activin Signaling Functions as a Sugar-Sensing Feedback Loop to Regulate Digestive Enzyme Expression. Cell Res. 2014, 9, 336–348. [Google Scholar]
- Couée, I.; Sulmon, C.; Gouesbet, G.; El, A.A. Involvement of soluble sugars in reactive oxygen species balance and responses to oxidative stress in plants. J. Exp. Bot. 2006, 57, 449–459. [Google Scholar] [CrossRef] [PubMed]
- Bolourimoghaddam, M.R.; Roy, K.L.; Li, X.A.; Rolland, F.; Ende, W.V.D. Sugar signalling and antioxidant network connections in plant cells. FEBS J. 2010, 277, 2022–2037. [Google Scholar] [CrossRef] [PubMed]
- Matros, A.; Peshev, D.; Peukert, M.; Mock, H.P.; Wim, V.D.E. Sugars as hydroxyl radical scavengers: Proof-of-concept by studying the fate of sucralose in Arabidopsis. Plant J. 2015, 82, 822–839. [Google Scholar] [CrossRef] [PubMed]
- Baker, R.F.; Leach, K.A.; Braun, D.M. SWEET as sugar: New sucrose effluxers in plants. Mol. Plant 2012, 5, 766–768. [Google Scholar] [CrossRef] [PubMed]
- Drew, M.; Saker, L.; Barber, S.; Jenkins, W. Changes in the kinetics of phosphate and potassium absorption in nutrient-deficient barley roots measured by a solution-depletion technique. Planta 1984, 160, 490–499. [Google Scholar] [CrossRef] [PubMed]
- Trapnell, C.; Roberts, A.; Goff, L.; Pertea, G.; Kim, D.; Kelley, D.R.; Pimentel, H.; Salzberg, S.L.; Rinn, J.L.; Pachter, L. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 2012, 7, 562. [Google Scholar] [CrossRef] [PubMed]
- Trapnell, C.; Williams, B.A.; Pertea, G.; Mortazavi, A.; Kwan, G.; Van Baren, M.J.; Salzberg, S.L.; Wold, B.J.; Pachter, L. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 2010, 28, 511–515. [Google Scholar] [CrossRef] [PubMed]
- Audic, S.; Claverie, J.-M. The significance of digital gene expression profiles. Genome Res. 1997, 7, 986–995. [Google Scholar] [CrossRef] [PubMed]
- Zenoni, S.; Ferrarini, A.; Giacomelli, E.; Xumerle, L.; Fasoli, M.; Malerba, G.; Bellin, D.; Pezzotti, M.; Delledonne, M. Characterization of transcriptional complexity during berry development in Vitis vinifera using RNA-Seq. Plant Physiol. 2010, 152, 1787–1795. [Google Scholar] [CrossRef] [PubMed]
- Eisen, M.B.; Spellman, P.T.; Brown, P.O.; Botstein, D. Cluster analysis and display of genome-wide expression patterns. Proc. Natl. Acad. Sci. USA 1998, 95, 14863–14868. [Google Scholar] [CrossRef] [PubMed]
- Mi, H.; Lazareva-Ulitsky, B.; Loo, R.; Kejariwal, A.; Vandergriff, J.; Rabkin, S.; Guo, N.; Muruganujan, A.; Doremieux, O.; Campbell, M.J. The PANTHER database of protein families, subfamilies, functions and pathways. Nucleic Acids Res. 2005, 33 (Suppl. 1), D284–D288. [Google Scholar] [CrossRef] [PubMed]
- Kanehisa, M.; Araki, M.; Goto, S.; Hattori, M.; Hirakawa, M.; Itoh, M.; Katayama, T.; Kawashima, S.; Okuda, S.; Tokimatsu, T. KEGG for linking genomes to life and the environment. Nucleic Acids Res. 2007, 36 (Suppl. 1), D480–D484. [Google Scholar] [CrossRef] [PubMed]
- Patterson, B.D.; Macrae, E.A.; Ferguson, I.B. Estimation of hydrogen peroxide in plant extracts using titanium(IV). Anal. Biochem. 1984, 139, 487–492. [Google Scholar] [CrossRef]
- Elstner, E.F.; Heupel, A. Inhibition of nitrite formation from hydroxylammoniumchloride: A simple assay for superoxide dismutase. Anal. Biochem. 1976, 70, 616–620. [Google Scholar] [CrossRef]
- Sun, C.; Li, Y.; Zhao, W.; Song, X.; Lu, M.; Li, X.; Li, X.; Liu, R.; Yan, L.; Zhang, X. Integration of hormonal and nutritional cues orchestrates progressive corolla opening. Plant Physiol. 2016, 171, 1209–1229. [Google Scholar] [PubMed]

| Genotype | K+ | K− | Relative Index | |
|---|---|---|---|---|
| Stem length (mm) | JZ34 | 36.33a | 34.67a | 0.95 |
| JZ18 | 28.67a | 24.00a | 0.84 | |
| Root length (mm) | JZ34 | 21.67a | 21.50a | 0.99 |
| JZ18 | 25.00a | 21.50b | 0.86 | |
| Fresh weight (mg·plant−1 FW) | JZ34 | 12.08a | 12.44a | 1.03 |
| JZ18 | 10.46a | 10.01a | 0.96 | |
| Dry weight (mg·plant−1 DW) | JZ34 | 0.79a | 0.85a | 1.07 |
| JZ18 | 0.77a | 0.55b | 0.71 | |
| K+ content (mg·g−1 DW) | JZ34 | 367.00a | 325.71a | 0.89 |
| JZ18 | 389.88a | 223. 34b | 0.57 | |
| K accumulation (mg·plant−1) | JZ34 | 289.20a | 276.53a | 0.95 |
| JZ18 | 286.17A | 129.98B | 0.45 |
| Sample | Raw Data Size (nt) | Raw Reads Number | Clean Data Size | Clean Reads Number | Clean Data Rate (%) | Total Mapped Reads (%) | Q20 (%) |
|---|---|---|---|---|---|---|---|
| JZ18-ck12h | 1206807650 | 24136153 | 1204621000 | 24092420 | 99.81 | 80.53 | 96.7 |
| JZ18-lk12h | 1206819150 | 24136383 | 1201320750 | 24026415 | 99.54 | 81.54 | 96.9 |
| JZ18-ck24h | 1206831700 | 24136634 | 1202634100 | 24052682 | 99.65 | 81.69 | 96.7 |
| JZ18-lk24h | 1206825700 | 24136514 | 1202777750 | 24055555 | 99.66 | 80.22 | 96.9 |
| JZ34-ck12h | 1206841700 | 24136834 | 1176860650 | 23537213 | 97.51 | 81.98 | 97.0 |
| JZ34-lk12h | 1206833100 | 24136662 | 1199255700 | 23985114 | 99.37 | 80.78 | 97.5 |
| JZ34-ck24h | 1206848550 | 24136971 | 1203941400 | 24078828 | 99.75 | 83.08 | 97.6 |
| JZ34-lk24h | 1206808900 | 24136178 | 1204460950 | 24089219 | 99.8 | 83.75 | 97.6 |
| Housekeeping Gene | Gene ID | Treatment Time (h) | JZ18 Log2 Fold Change | FDR | Significant | JZ34 Log2 Fold Change | FDR | Significant |
|---|---|---|---|---|---|---|---|---|
| GAPDH | Solyc04g009030.2 | 12 | 0.01 | 1.00 | no | 0.02 | 1.00 | no |
| 24 | 0.03 | 1.00 | no | 0.02 | 1.00 | no | ||
| GAPDH | Solyc12g094640.1 | 12 | −0.10 | 1.00 | no | 0.00 | 0.99 | no |
| 24 | 0.81 | 1.00 | no | −0.76 | 1.00 | no | ||
| Catalase | Solyc02g082760.2 | 12 | 0.13 | 1.00 | no | 0.13 | 1.00 | no |
| 24 | 0.14 | 1.00 | no | 0.05 | 0.07 | no | ||
| Catalase | Solyc12g094620.1 | 12 | 0.49 | 1.00 | no | −0.27 | 1.00 | no |
| 24 | 0.72 | 1.00 | no | 0.21 | 1.00 | no | ||
| Cys protease | Solyc08g082400.1 | 12 | 0.05 | 0.83 | no | −0.14 | 0.15 | no |
| 24 | 0.15 | 0.24 | no | −0.31 | 1.00 | no | ||
| Cys protease | Solyc12g095910.1 | 12 | 0.24 | 1.00 | no | −0.12 | 1.00 | no |
| 24 | 0.04 | 0.70 | no | 0.26 | 1.00 | no | ||
| α-Tubulin | Solyc04g077020.2 | 12 | 0.15 | 1.00 | no | −0.03 | 0.23 | no |
| 24 | −0.06 | 0.11 | no | −0.04 | 0.12 | no | ||
| Ubiquitin | Solyc07g064130.1 | 12 | −0.16 | 1.00 | no | 0.00 | 0.86 | no |
| 24 | −0.05 | 1.00 | no | −0.22 | 1.00 | no | ||
| Actin | Solyc04g011500.2 | 12 | 0.04 | 0.75 | no | 0.13 | 0.19 | no |
| 24 | −0.07 | 0.27 | no | 0.05 | 0.41 | no | ||
| DNAJ chaperone | Solyc11g006460.1 | 12 | 0.19 | 1.00 | no | 0.04 | 0.25 | no |
| 24 | 0.03 | 0.36 | no | −0.02 | 0.71 | no | ||
| Translation initiation factor 5A | Solyc12g010060.1 | 12 | 0.12 | 1.00 | no | 0.11 | 1.00 | no |
| 24 | 0.01 | 0.87 | no | 0.24 | 1.00 | no |
| Group | Gene ID | log2 (Fold Change) | Seq Description | |||
|---|---|---|---|---|---|---|
| JZ18 | JZ34 | |||||
| 12 h | 24 h | 12 h | 24 h | |||
| potassium | Solyc11g011500 | 1.30 | Potassium channel | |||
| Solyc03g097930 | 1.06 | potassium channel SKOR-like | ||||
| Solyc01g010480 | 1.60 | Potassium voltage-gated channel | ||||
| Solyc12g009540 | −6.79 | KUP system potassium uptake protein | ||||
| Solyc07g014680 | −1.25 | Potassium transporter | ||||
| Nitrate | Solyc11g069760 | 1.36 | −2.69 | High affinity nitrate transporter protein | ||
| Solyc08g007430 | 1.23 | Nitrate transporter | ||||
| Solyc06g010250 | −2.20 | −1.85 | Nitrate transporter | |||
| Solyc00g090860 | 1.21 | −1.83 | Nitrate transporter | |||
| Solyc07g032490 | 1.10 | Nitrate transporter | ||||
| Solyc08g078950 | −1.18 | Nitrate transporter | ||||
| Solyc08g007430 | −1.23 | Nitrate transporter | ||||
| Solyc11g069750 | 1.70 | Nitrate transporter | ||||
| Solyc03g113250 | −1.15 | Nitrate transporter | ||||
| Solyc11g069740 | −1.56 | Nitrate transporter | ||||
| Yellow-strike | Solyc03g082620 | 2.10 | Metal-nicotianamine transporter | |||
| Mate | Solyc07g006740 | 1.63 | MATE efflux family protein | |||
| Solyc11g065820 | 1.13 | MATE efflux family protein | ||||
| Solyc03g026230 | 1.57 | MATE efflux family protein | ||||
| Solyc08g080310 | 1.12 | −1.32 | MATE efflux family protein | |||
| LRR | Solyc01g005870 | 1.74 | LRR receptor-like protein kinase | |||
| Solyc06g033920 | −1.01 | LRR receptor-like protein kinase | ||||
| Solyc01g009810 | −2.04 | LRR receptor-like protein kinase | ||||
| Solyc01g091230 | −1.20 | LRR receptor-like protein kinase | ||||
| Solyc01g098690 | 1.56 | LRR receptor-like protein kinase | ||||
| Solyc02g071820 | 1.57 | LRR receptor-like protein kinase | ||||
| Solyc03g005960 | −1.65 | LRR receptor-like protein kinase | ||||
| Solyc03g007050 | −1.41 | LRR receptor-like protein kinase | ||||
| Solyc04g014400 | −1.16 | 1.74 | LRR receptor-like protein kinase | |||
| Solyc04g014900 | −1.06 | LRR receptor-like protein kinase | ||||
| Solyc04g054450 | −1.66 | LRR receptor-like protein kinase | ||||
| Solyc04g074000 | −1.10 | −1.97 | 3.67 | LRR receptor-like protein kinase | ||
| Solyc04g074020 | −1.73 | LRR receptor-like protein kinase | ||||
| Solyc04g074030 | −3.12 | 5.49 | LRR receptor-like protein kinase | |||
| Solyc04g074050 | −2.08 | 2.11 | LRR receptor-like protein kinase | |||
| Solyc04g081080 | −1.10 | LRR receptor-like protein kinase | ||||
| Solyc06g006020 | −2.89 | −1.23 | LRR receptor-like protein kinase | |||
| Solyc09g014480 | 1.14 | LRR receptor-like protein kinase | ||||
| Solyc09g082530 | −1.15 | LRR receptor-like protein kinase | ||||
| Solyc10g007830 | −3.32 | LRR receptor-like protein kinase | ||||
| Solyc12g009730 | −1.43 | LRR receptor-like protein kinase | ||||
| Solyc12g009750 | −1.43 | LRR receptor-like protein kinase | ||||
| Solyc12g009780 | −3.56 | 2.25 | LRR receptor-like protein kinase | |||
| Solyc11g011180 | 2.83 | LRR receptor-like protein kinase | ||||
| CRK | Solyc01g006390 | 2.60 | −5.03 | Cysteine-rich receptor-like protein kinase | ||
| Solyc03g111530 | −4.90 | Cysteine-rich receptor-like protein kinase | ||||
| Solyc11g006430 | −2.52 | −1.29 | Cysteine-rich receptor-like protein kinase | |||
| Solyc07g005110 | 1.34 | Cysteine-rich receptor-like protein kinase | ||||
| Solyc05g018930 | −2.59 | Cysteine-rich receptor-like protein kinase | ||||
| H2O2 Concentration/nmol·min−1·g−1 FW | O2− Concentration/nmol·min−1·g−1 FW | |||||||
|---|---|---|---|---|---|---|---|---|
| JZ18K+ | JZ18K− | JZ34K+ | JZ34K− | JZ18K+ | JZ18K− | JZ34K+ | JZ34K− | |
| Leaf | ||||||||
| 0 h | 31.07 ± 1.73 | 30.66 ± 2.35 | 19.02 ± 1.42 | 19.33 ± 1.43 | 3.91 ± 0.06 | 3.89 ± 0.22 | 3.07 ± 0.05 | 3.21 ± 0.25 |
| 12 h | 30.88 ± 1.09 | 40.10 ± 2.07 ** | 22.00 ± 1.47 | 22.66 ± 1.67 | 3.80 ± 0.14 | 4.72 ± 0.15 * | 2.88 ± 0.14 | 2.87 ± 0.24 |
| 24 h | 31.01 ± 1.33 | 33.40 ± 2.31 | 21.16 ± 2.00 | 23.97 ± 2.05 | 3.49 ± 0.21 | 4.43 ± 0.20 * | 2.66 ± 0.04 | 2.46 ± 0.12 |
| 3 d | 32.00 ± 1.68 | 36.73 ± 0.66 | 18.62 ± 1.01 | 18.12 ± 2.01 | 3.58 ± 0.24 | 4.14 ± 0.24 | 2.78 ± 0.17 | 2.44 ± 0.10 |
| 7 d | 31.58 ± 0.77 | 34.95 ± 2.55 | 19.45 ± 1.65 | 19.11 ± 2.23 | 3.70 ± 0.20 | 4.38 ± 0.12 | 2.87 ± 0.15 | 2.70 ± 0.17 |
| Root | ||||||||
| 0 h | 6.45 ± 0.17 | 6.62 ± 0.13 | 4.42 ± 0.21 | 4.44 ± 0.04 | 0.96 ± 0.03 | 0.88 ± 0.04 | 0.83 ± 0.03 | 0.84 ± 0.02 |
| 12 h | 5.98 ± 0.25 | 8.79 ± 0.25 ** | 4.86 ± 0.23 | 4.25 ± 0.14 | 0.71 ± 0.03 | 1.06 ± 0.04 ** | 0.77 ± 0.05 | 0.70 ± 0.04 |
| 24 h | 6.66 ± 0.18 | 7.75 ± 0.16 * | 3.66 ± 0.16 | 4.07 ± 0.13 | 0.66 ± 0.04 | 0.73 ± 0.01 | 0.75 ± 0.02 | 0.87 ± 0.01 * |
| 3 d | 6.10 ± 0.23 | 6.29 ± 0.09 | 3.78 ± 0.09 | 3.65 ± 0.16 | 0.70 ± 0.02 | 0.75 ± 0.03 | 0.71 ± 0.01 | 0.74 ± 0.03 |
| 7 d | 6.98 ± 0.03 | 7.24 ± 0.11 | 3.62 ± 0.19 | 3.76 ± 0.03 | 0.74 ± 0.04 | 0.77 ± 0.03 | 0.73 ± 0.02 | 0.66 ± 0.01 |
| Transcript ID | JZ18-12 h | JZ18-24 h | JZ34-12 h | JZ34-24 h | Seq Description |
|---|---|---|---|---|---|
| Solyc01g068410 | −2.65 | 2.83 | Auxin Efflux Carrier | ||
| Solyc10g080880 | −1.00 | Auxin Efflux Carrier | |||
| Solyc04g056620 | 2.11 | Auxin Efflux Carrier | |||
| Solyc07g006900 | 1.14 | 1.13 | Auxin Efflux Carrier | ||
| Solyc02g082450 | −1.63 | Auxin Efflux Carrier | |||
| Solyc12g042980 | 1.83 | Gibberellin 20-oxidase | |||
| Solyc03g006880 | 1.14 | Gibberellin 20-oxidase-1 | |||
| Solyc11g072310 | 1.55 | Gibberellin 20-oxidase-3 | |||
| Solyc03g025490 | 1.44 | Gibberellin 20-oxidase-like protein | |||
| Solyc06g066820 | −1.51 | Gibberellin 3-beta-hydroxylase | |||
| Solyc07g056670 | 1.74 | Gibberellin 3-oxidase | |||
| Solyc12g042980 | −3.27 | Gibberellin 3-beta-dioxygenase | |||
| Solyc00g138060 | 1.91 | 1.35 | Gibberellin 2-oxidase | ||
| Solyc01g090630 | −1.04 | Gibberellin 2-oxidase | |||
| Solyc01g079200 | 2.57 | Gibberellin 2-oxidase | |||
| Solyc02g080120 | 1.36 | 1.89 | Gibberellin 2-beta-dioxygenase | ||
| Solyc02g070430 | 1.11 | Gibberellin 2-oxidase 1 | |||
| Solyc06g076550 | −1.54 | Cytokinin-N-glucosyltransferase | |||
| Solyc04g074380 | −2.12 | Cytokinin-N-glucosyltransferase | |||
| Solyc10g085280 | 1.18 | Cytokinin-N-glucosyltransferase | |||
| Solyc12g057080 | −1.39 | Cytokinin-N-glucosyltransferase | |||
| Solyc12g008900 | −4.91 | Cytokinin dehydrogenase 1 | |||
| Solyc10g017990 | −7.65 | Cytokinin dehydrogenase 1 | |||
| Solyc02g079440 | 1.47 | 1.04 | Cytokinin dehydrogenase | ||
| Solyc06g082730 | 1.42 | Cytokinin trans-hydroxylase | |||
| Solyc09g089580 | −1.49 | −1.31 | Ethylene metabolic process | ||
| Solyc02g091990 | 2.15 | Ethylene metabolic process | |||
| Solyc07g049530 | −1.00 | Ethylene metabolic process | |||
| Solyc09g091550 | −2.9 | 2.44 | Salicylic acid carboxyl methyltransferase | ||
| Solyc09g091540 | 6.92 | Salicylic acid carboxyl methyltransferase |
| Transcript ID | JZ18-12 h | JZ18-24 h | JZ34-12 h | JZ34-24 h | Seq Description |
|---|---|---|---|---|---|
| Solyc03g097560 | −6.82 | bidirectional sugar transporter SWEET14 | |||
| Solyc03g097600 | 1.40 | 1.69 | bidirectional sugar transporter SWEET10-like | ||
| Solyc06g072620 | 1.24 | bidirectional sugar transporter SWEET10-like | |||
| Solyc03g097610 | 1.22 | bidirectional sugar transporter SWEET10 | |||
| Solyc06g060590 | 1.41 | bidirectional sugar transporter SWEET1 | |||
| Solyc01g080680 | 1.01 | 1.04 | Glucose transporter 8 | ||
| Solyc06g054270 | 1.05 | Solute carrier family 2 Sugar/inositol transporter | |||
| Solyc09g075820 | 1.81 | Solute carrier family 2 Sugar/inositol transporter | |||
| Solyc11g067340 | 1.02 | Acyltransferase (Fragment) | |||
| Solyc09g092130 | 1.24 | −1.58 | Sucrose phosphate synthase | ||
| Solyc08g066100 | 6.22 | ATP-dependent 6-phosphofructokinase | |||
| Solyc12g009300 | 2.68 | Sucrose synthase | |||
| Solyc08g079080 | −1.33 | 1.17 | vacuolar invertase, Lin9 | ||
| Solyc02g086530 | −3.08 | Alpha-galactosidase |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhao, X.; Liu, Y.; Liu, X.; Jiang, J. Comparative Transcriptome Profiling of Two Tomato Genotypes in Response to Potassium-Deficiency Stress. Int. J. Mol. Sci. 2018, 19, 2402. https://doi.org/10.3390/ijms19082402
Zhao X, Liu Y, Liu X, Jiang J. Comparative Transcriptome Profiling of Two Tomato Genotypes in Response to Potassium-Deficiency Stress. International Journal of Molecular Sciences. 2018; 19(8):2402. https://doi.org/10.3390/ijms19082402
Chicago/Turabian StyleZhao, Xiaoming, Yang Liu, Xin Liu, and Jing Jiang. 2018. "Comparative Transcriptome Profiling of Two Tomato Genotypes in Response to Potassium-Deficiency Stress" International Journal of Molecular Sciences 19, no. 8: 2402. https://doi.org/10.3390/ijms19082402
APA StyleZhao, X., Liu, Y., Liu, X., & Jiang, J. (2018). Comparative Transcriptome Profiling of Two Tomato Genotypes in Response to Potassium-Deficiency Stress. International Journal of Molecular Sciences, 19(8), 2402. https://doi.org/10.3390/ijms19082402





