Muller’s Ratchet and Ribosome Degeneration in the Obligate Intracellular Parasites Microsporidia
Abstract
1. Introduction
2. Materials and Methods
2.1. Analysis of rRNA-Protein Contacts Evolution in the Ribosome Structure
2.2. Preparation of Protein Extracts from Microsporidian Spores
2.3. Mass Spectrometry
2.4. Database Searching to Detect Protein Identity in Microsporidian Bulk Protein Extracts
3. Results
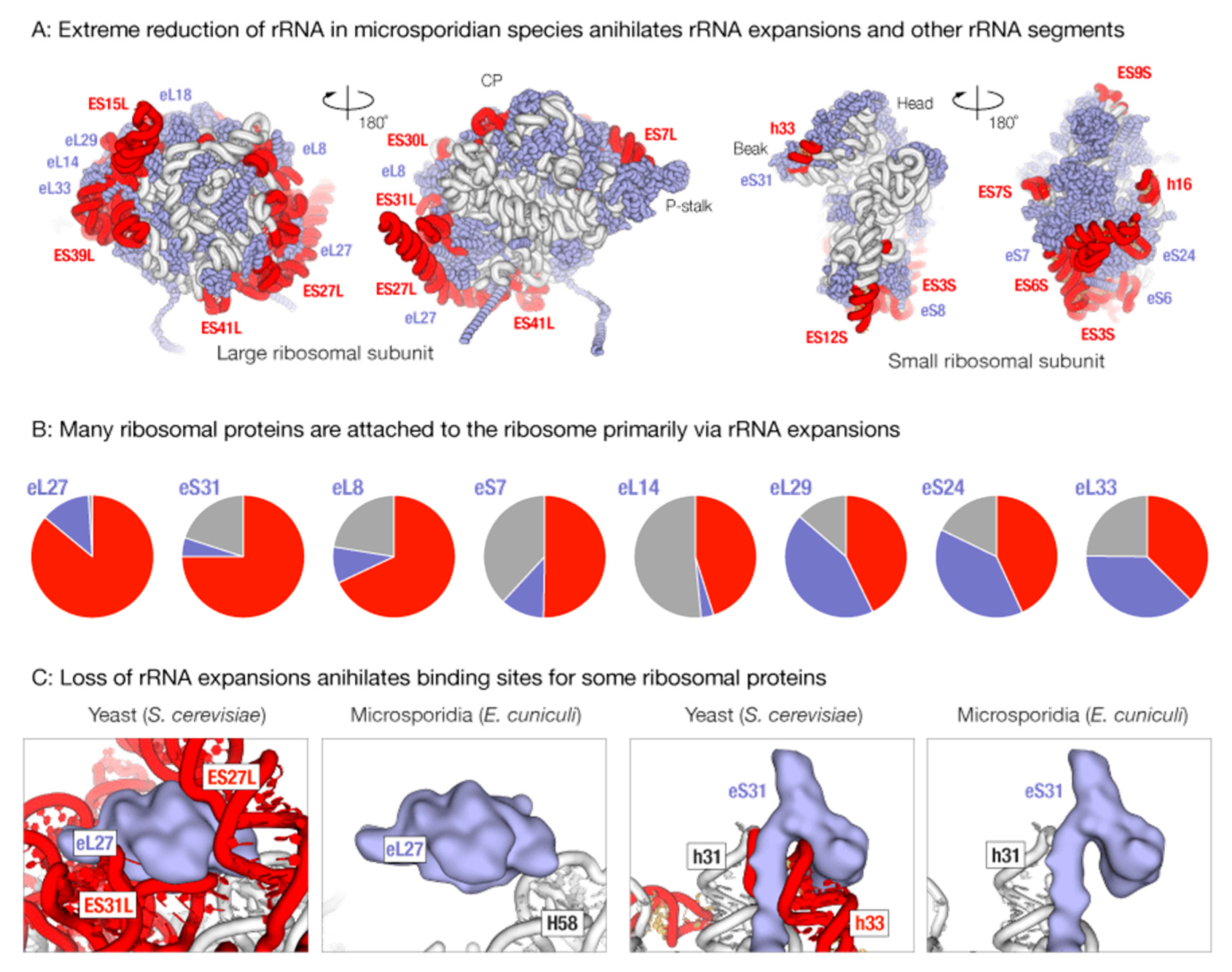
3.1. Massive Loss of rRNA Expansion Segments Degenerates Binding Sites for Ribosomal Proteins in Microsporidian Ribosomes
3.2. Proteins that Appear to Lose Association with the Ribosome also Lose Their Sequence Conservation
3.3. Apparent Rudimentary Ribosomal Proteins are Abundantly Expressed in Microsporidian Cells
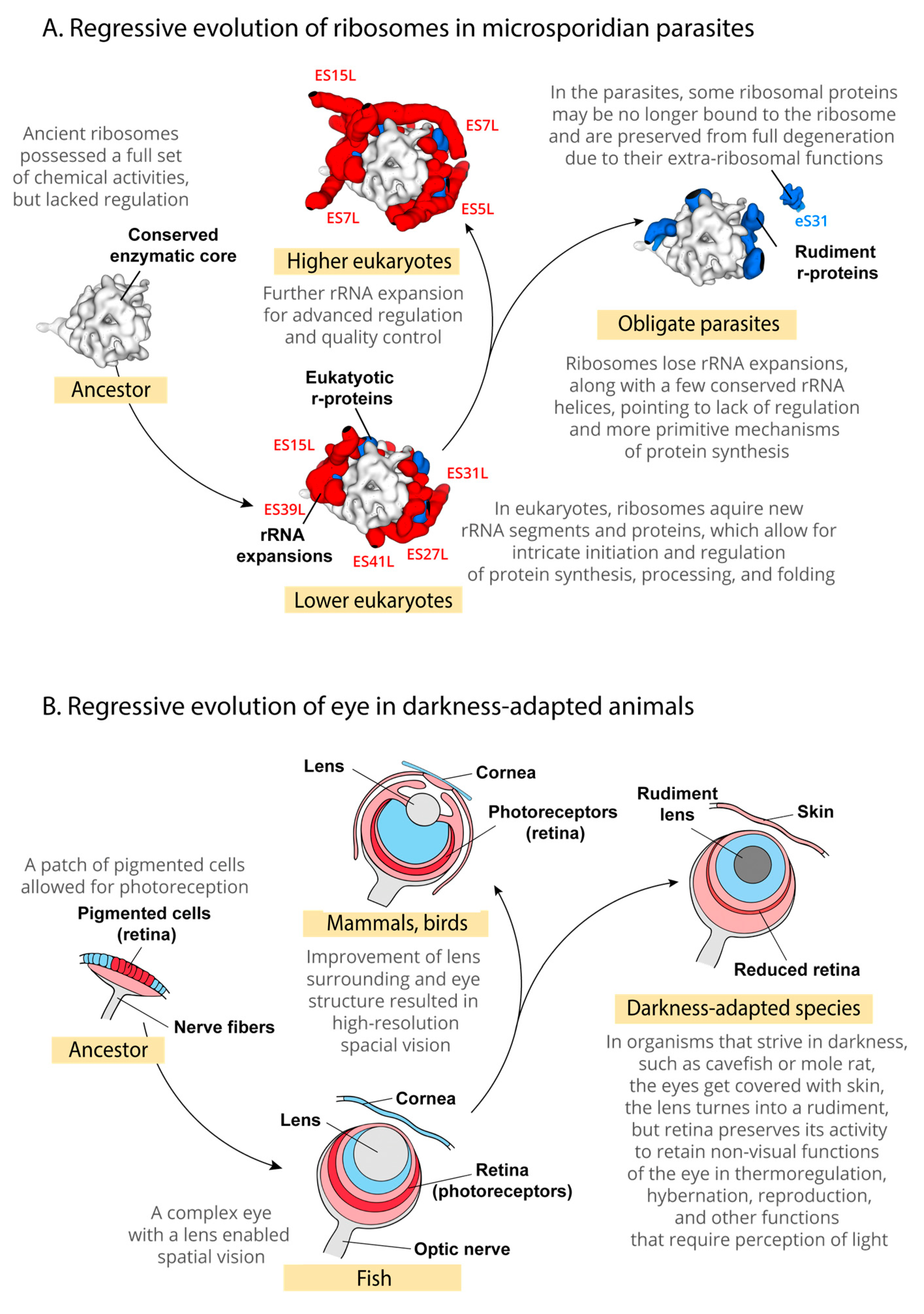
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Moran, N.A. Microbial minimalism: Genome reduction in bacterial pathogens. Cell 2002, 108, 583–586. [Google Scholar] [CrossRef]
- McCutcheon, J.P.; Moran, N.A. Extreme genome reduction in symbiotic bacteria. Nat. Rev. Microbiol. 2011, 10, 13–26. [Google Scholar] [CrossRef] [PubMed]
- Andersson, S.G.; Kurland, C.G. Reductive evolution of resident genomes. Trends Microbiol. 1998, 6, 263–268. [Google Scholar] [CrossRef]
- Muller, H.J. The relation of recombination to mutational advance. Mutat. Res. 1964, 106, 2–9. [Google Scholar] [CrossRef]
- Moran, N.A.; Bennett, G.M. The tiniest tiny genomes. Annu. Rev. Microbiol. 2014, 68, 195–215. [Google Scholar] [CrossRef] [PubMed]
- Moran, N.A. Accelerated evolution and Muller’s rachet in endosymbiotic bacteria. Proc. Natl. Acad. Sci. USA 1996, 93, 2873–2878. [Google Scholar] [CrossRef] [PubMed]
- Melnikov, S.V.; Rivera, K.D.; Ostapenko, D.; Makarenko, A.; Sanscrainte, N.D.; Becnel, J.J.; Solomon, M.J.; Texier, C.; Pappin, D.J.; Soll, D. Error-prone protein synthesis in parasites with the smallest eukaryotic genome. Proc. Natl. Acad. Sci. USA 2018, 115, E6245–E6253. [Google Scholar] [CrossRef] [PubMed]
- Melnikov, S.V.; van den Elzen, A.; Stevens, D.L.; Thoreen, C.C.; Soll, D. Loss of protein synthesis quality control in host-restricted organisms. Proc. Natl. Acad. Sci. USA 2018, 115, E11505–E11512. [Google Scholar] [CrossRef] [PubMed]
- Gill, E.E.; Fast, N.M. Stripped-down DNA repair in a highly reduced parasite. BMC Mol. Biol. 2007, 8, 24. [Google Scholar] [CrossRef] [PubMed]
- Shonhai, A.; Maier, A.G.; Przyborski, J.M.; Blatch, G.L. Intracellular protozoan parasites of humans: The role of molecular chaperones in development and pathogenesis. Protein Pept. Lett. 2011, 18, 143–157. [Google Scholar] [CrossRef] [PubMed]
- Wust, P.; Hildebrandt, B.; Sreenivasa, G.; Rau, B.; Gellermann, J.; Riess, H.; Felix, R.; Schlag, P.M. Hyperthermia in combined treatment of cancer. Lancet Oncol. 2002, 3, 487–497. [Google Scholar] [CrossRef]
- Lambert, J.D.; Moran, N.A. Deleterious mutations destabilize ribosomal RNA in endosymbiotic bacteria. Proc. Natl. Acad. Sci. USA 1998, 95, 4458–4462. [Google Scholar] [CrossRef]
- Escarmis, C.; Perales, C.; Domingo, E. Biological effect of muller’s ratchet: Distant capsid site can affect picornavirus protein processing. J. Virol. 2009, 83, 6748–6756. [Google Scholar] [CrossRef] [PubMed]
- Fares, M.A.; Moya, A.; Barrio, E. Groel and the maintenance of bacterial endosymbiosis. Trends Genet. 2004, 20, 413–416. [Google Scholar] [CrossRef]
- Katinka, M.D.; Duprat, S.; Cornillot, E.; Metenier, G.; Thomarat, F.; Prensier, G.; Barbe, V.; Peyretaillade, E.; Brottier, P.; Wincker, P.; et al. Genome sequence and gene compaction of the eukaryote parasite Encephalitozoon cuniculi. Nature 2001, 414, 450–453. [Google Scholar] [CrossRef]
- Peyretaillade, E.; El Alaoui, H.; Diogon, M.; Polonais, V.; Parisot, N.; Biron, D.G.; Peyret, P.; Delbac, F. Extreme reduction and compaction of microsporidian genomes. Res. Microbiol. 2011, 162, 598–606. [Google Scholar] [CrossRef] [PubMed]
- Didier, E.S.; Weiss, L.M. Microsporidiosis: Not just in AIDs patients. Curr. Opin. Infect Dis. 2011, 24, 490–495. [Google Scholar] [CrossRef]
- NIH: National Institute of Allergy and Infectious Diseases. Available online: https://www.niaid.nih.gov/research/emerging-infectious-diseases-pathogens (accessed on 17 November 2018).
- Melnikov, S.; Ben-Shem, A.; Garreau de Loubresse, N.; Jenner, L.; Yusupova, G.; Yusupov, M. One core, two shells: Bacterial and eukaryotic ribosomes. Nat. Struct. Mol. Biol. 2012, 19, 560–567. [Google Scholar] [CrossRef] [PubMed]
- Peyretaillade, E.; Biderre, C.; Peyret, P.; Duffieux, F.; Metenier, G.; Gouy, M.; Michot, B.; Vivares, C.P. Microsporidian Encephalitozoon cuniculi, a unicellular eukaryote with an unusual chromosomal dispersion of ribosomal genes and a lsu rrna reduced to the universal core. Nucleic Acids Res. 1998, 26, 3513–3520. [Google Scholar] [CrossRef]
- Vossbrinck, C.R.; Maddox, J.V.; Friedman, S.; Debrunner-Vossbrinck, B.A.; Woese, C.R. Ribosomal RNA sequence suggests Microsporidia are extremely ancient eukaryotes. Nature 1987, 326, 411–414. [Google Scholar] [CrossRef]
- Weiss, L.M.; Zhu, X.; Cali, A.; Tanowitz, H.B.; Wittner, M. Utility of microsporidian rRNA in diagnosis and phylogeny: A review. Folia Parasitol. 1994, 41, 81–90. [Google Scholar] [PubMed]
- Dong, S.; Shen, Z.; Xu, L.; Zhu, F. Sequence and phylogenetic analysis of SSU rRNA gene of five Microsporidia. Curr. Microbiol. 2010, 60, 30–37. [Google Scholar] [CrossRef] [PubMed]
- Krissinel, E. Stock-based detection of protein oligomeric states in jsPISA. Nucleic Acids Res. 2015, 43, W314–W319. [Google Scholar] [CrossRef]
- Sievers, F.; Higgins, D.G. Clustal Omega, accurate alignment of very large numbers of sequences. Methods Mol. Biol. 2014, 1079, 105–116. [Google Scholar]
- Brosson, D.; Kuhn, L.; Delbac, F.; Garin, J.; Vivares, C.P.; Texier, C. Proteomic analysis of the eukaryotic parasite Encephalitozoon cuniculi (Microsporidia): A reference map for proteins expressed in late sporogonial stages. Proteomics 2006, 6, 3625–3635. [Google Scholar] [CrossRef]
- Delbac, F.; Duffieux, F.; David, D.; Metenier, G.; Vivares, C.P. Immunocytochemical identification of spore proteins in two Microsporidia, with emphasis on extrusion apparatus. J. Eukaryot. Microbiol. 1998, 45, 224–231. [Google Scholar] [CrossRef]
- Ross, P.L.; Huang, Y.N.; Marchese, J.N.; Williamson, B.; Parker, K.; Hattan, S.; Khainovski, N.; Pillai, S.; Dey, S.; Daniels, S.; et al. Multiplexed protein quantitation in Saccharomyces cerevisiae using amine-reactive isobaric tagging reagents. Mol. Cell. Proteomics 2004, 3, 1154–1169. [Google Scholar] [CrossRef] [PubMed]
- Gilar, M.; Olivova, P.; Daly, A.E.; Gebler, J.C. Two-dimensional separation of peptides using RP-RP-HPLC system with different pH in first and second separation dimensions. J. Sep. Sci. 2005, 28, 1694–1703. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Yang, F.; Gritsenko, M.A.; Wang, Y.; Clauss, T.; Liu, T.; Shen, Y.; Monroe, M.E.; Lopez-Ferrer, D.; Reno, T.; et al. Reversed-phase chromatography with multiple fraction concatenation strategy for proteome profiling of human Mcf10a cells. Proteomics 2011, 11, 2019–2026. [Google Scholar] [CrossRef]
- Perkins, D.N.; Pappin, D.J.; Creasy, D.M.; Cottrell, J.S. Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 1999, 20, 3551–3567. [Google Scholar] [CrossRef]
- Passmore, L.A.; Schmeing, T.M.; Maag, D.; Applefield, D.J.; Acker, M.G.; Algire, M.A.; Lorsch, J.R.; Ramakrishnan, V. The eukaryotic translation initiation factors eIF1 and eIF1a induce an open conformation of the 40S ribosome. Mol. Cell 2007, 26, 41–50. [Google Scholar] [CrossRef] [PubMed]
- Pisareva, V.P.; Pisarev, A.V.; Komar, A.A.; Hellen, C.U.; Pestova, T.V. Translation initiation on mammalian mRNAs with structured 5′utrs requires dexh-box protein Dhx29. Cell 2008, 135, 1237–1250. [Google Scholar] [CrossRef] [PubMed]
- Myasnikov, A.G.; Afonina, Z.A.; Menetret, J.F.; Shirokov, V.A.; Spirin, A.S.; Klaholz, B.P. The molecular structure of the left-handed supra-molecular helix of eukaryotic polyribosomes. Nat. Commun. 2014, 5, 5294. [Google Scholar] [CrossRef] [PubMed]
- Sanyal, S.; Jansen, H.G.; de Grip, W.J.; Nevo, E.; de Jong, W.W. The eye of the blind mole rat, Spalax ehrenbergi. Rudiment with hidden function? Invest. Ophthalmol. Vis. Sci. 1990, 31, 1398–1404. [Google Scholar] [PubMed]
- De Jong, W.W.; Hendriks, W.; Sanyal, S.; Nevo, E. The eye of the blind mole rat (Spalax ehrenbergi): Regressive evolution at the molecular level. Prog. Clin. Biol. Res. 1990, 335, 383–395. [Google Scholar] [PubMed]

© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Melnikov, S.V.; Manakongtreecheep, K.; Rivera, K.D.; Makarenko, A.; Pappin, D.J.; Söll, D. Muller’s Ratchet and Ribosome Degeneration in the Obligate Intracellular Parasites Microsporidia. Int. J. Mol. Sci. 2018, 19, 4125. https://doi.org/10.3390/ijms19124125
Melnikov SV, Manakongtreecheep K, Rivera KD, Makarenko A, Pappin DJ, Söll D. Muller’s Ratchet and Ribosome Degeneration in the Obligate Intracellular Parasites Microsporidia. International Journal of Molecular Sciences. 2018; 19(12):4125. https://doi.org/10.3390/ijms19124125
Chicago/Turabian StyleMelnikov, Sergey V., Kasidet Manakongtreecheep, Keith D. Rivera, Arthur Makarenko, Darryl J. Pappin, and Dieter Söll. 2018. "Muller’s Ratchet and Ribosome Degeneration in the Obligate Intracellular Parasites Microsporidia" International Journal of Molecular Sciences 19, no. 12: 4125. https://doi.org/10.3390/ijms19124125
APA StyleMelnikov, S. V., Manakongtreecheep, K., Rivera, K. D., Makarenko, A., Pappin, D. J., & Söll, D. (2018). Muller’s Ratchet and Ribosome Degeneration in the Obligate Intracellular Parasites Microsporidia. International Journal of Molecular Sciences, 19(12), 4125. https://doi.org/10.3390/ijms19124125




