Morphological Characterization of Root System Architecture in Diverse Tomato Genotypes during Early Growth
Abstract
1. Introduction
2. Results and Discussion
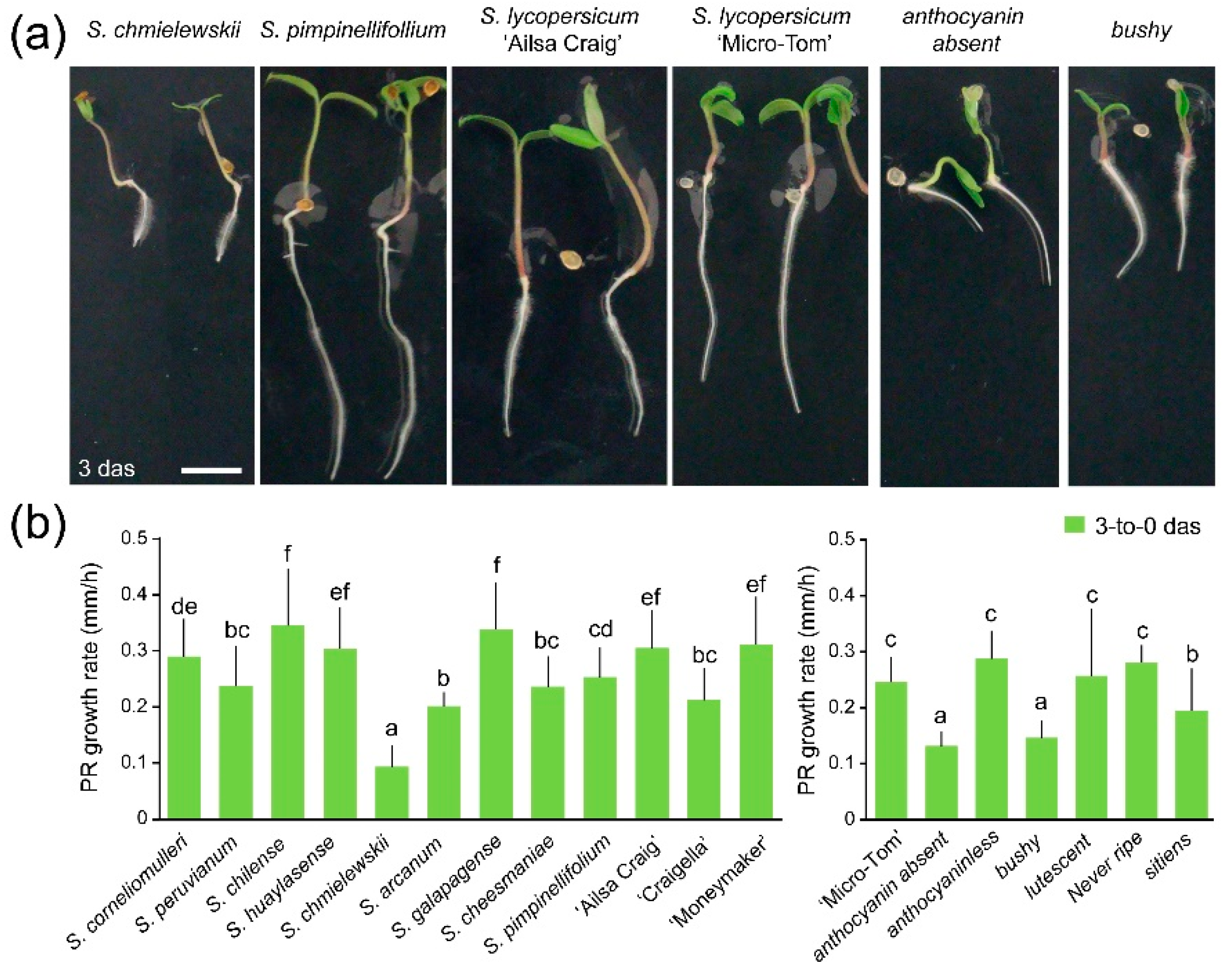
2.1. Germination and Early Root Growth
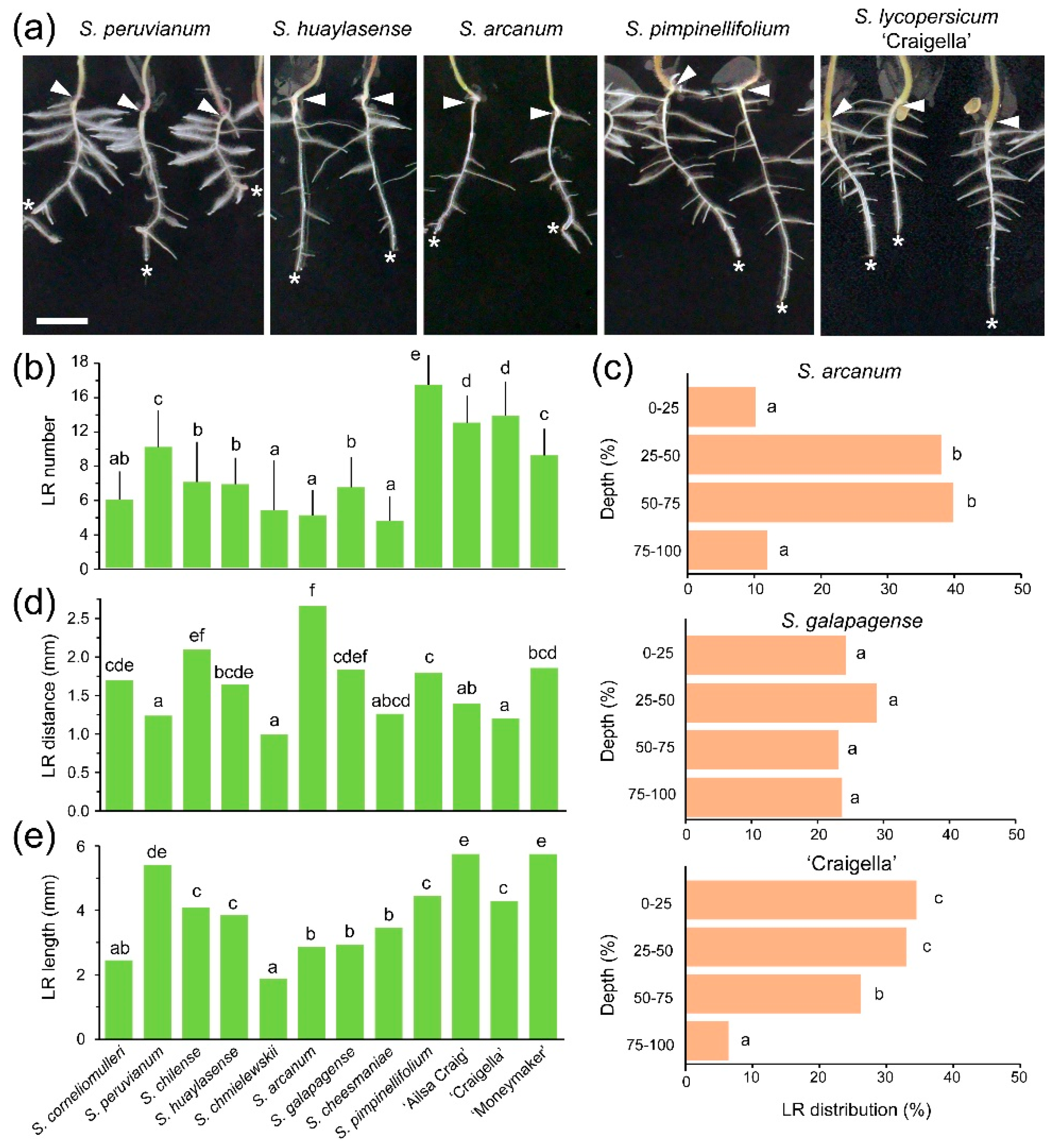
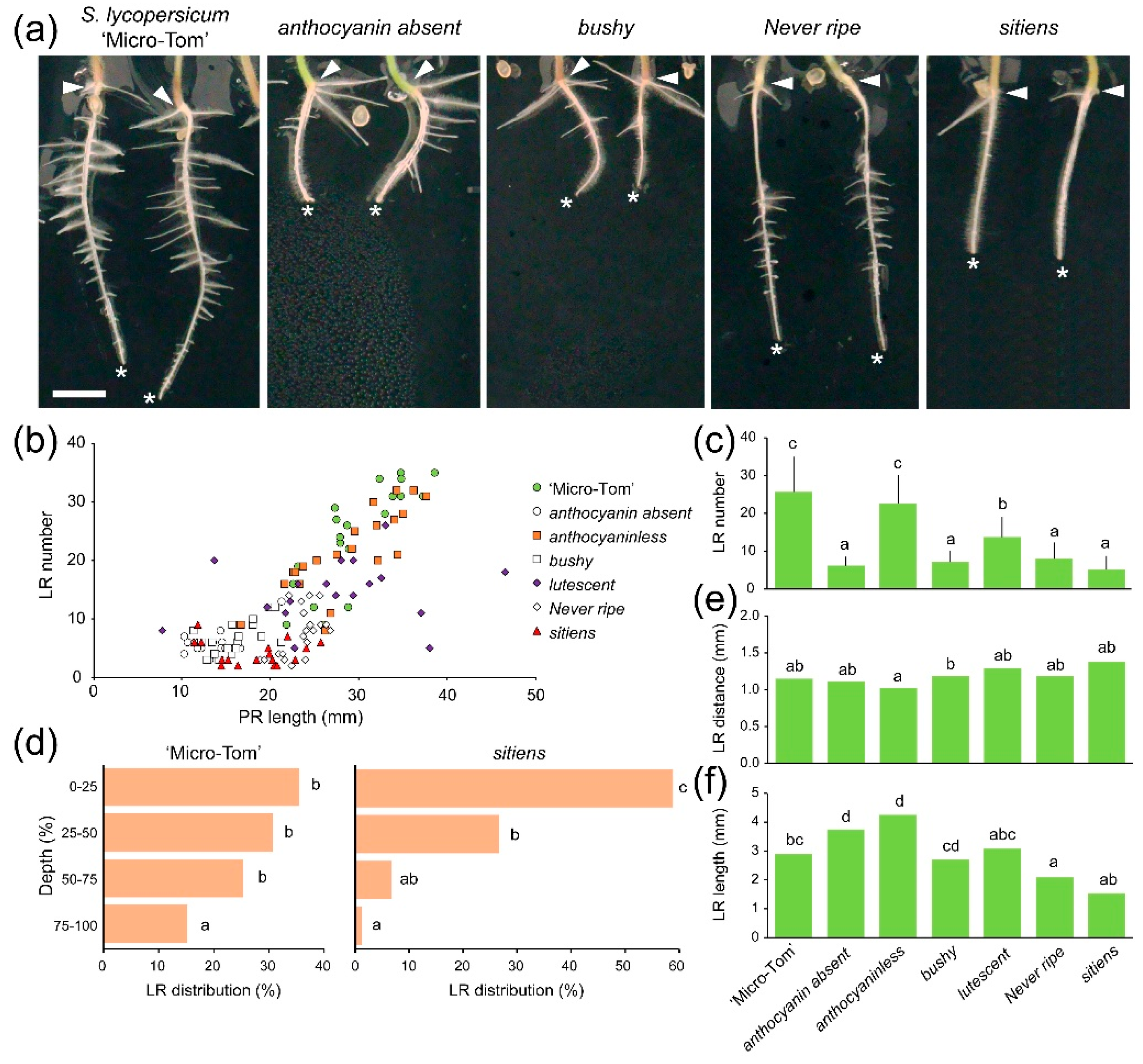
2.2. Lateral Root (LR) Capacity Assay
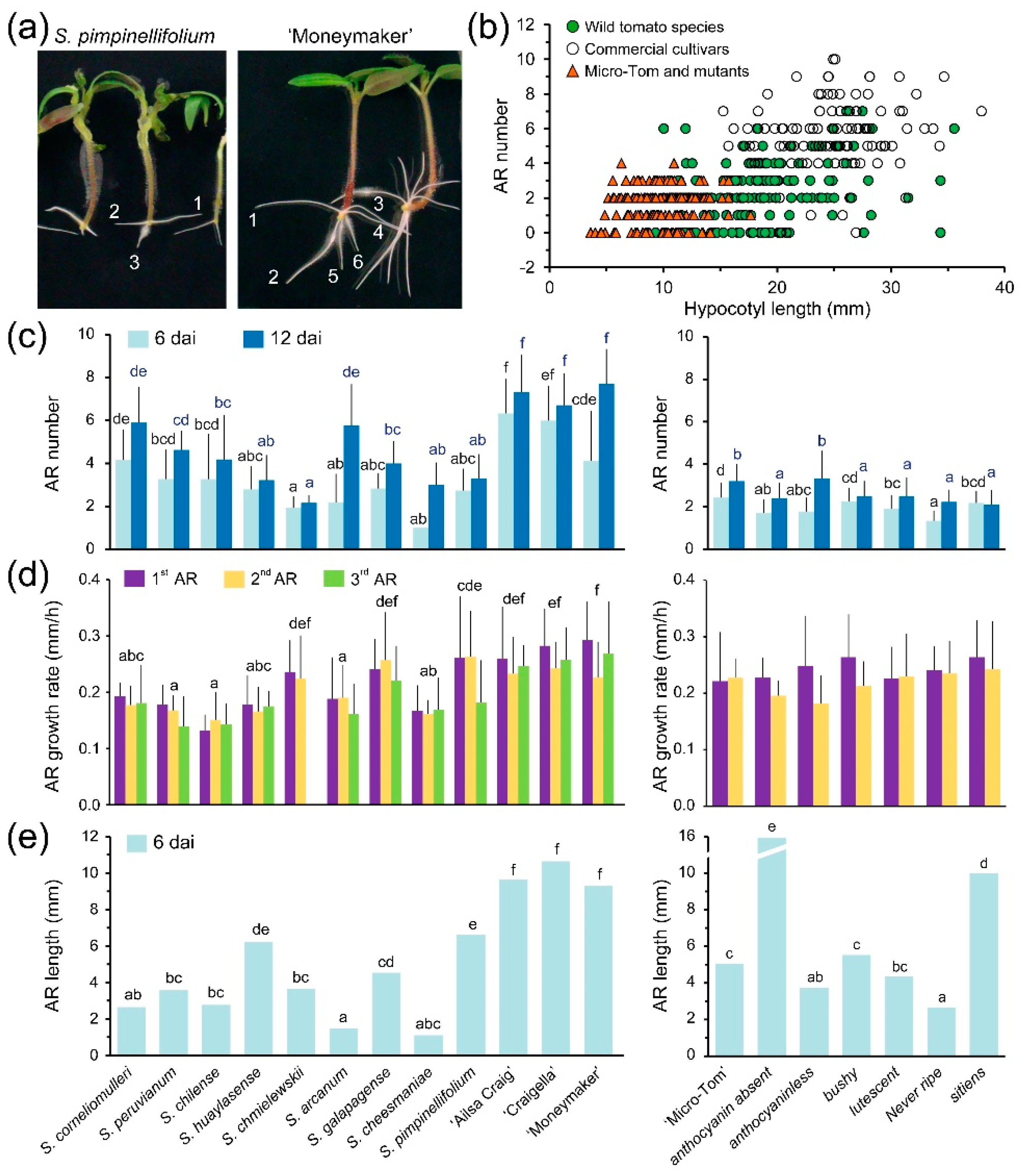
2.3. Adventitious Root (AR) Formation in the Hypocotyl after Wounding
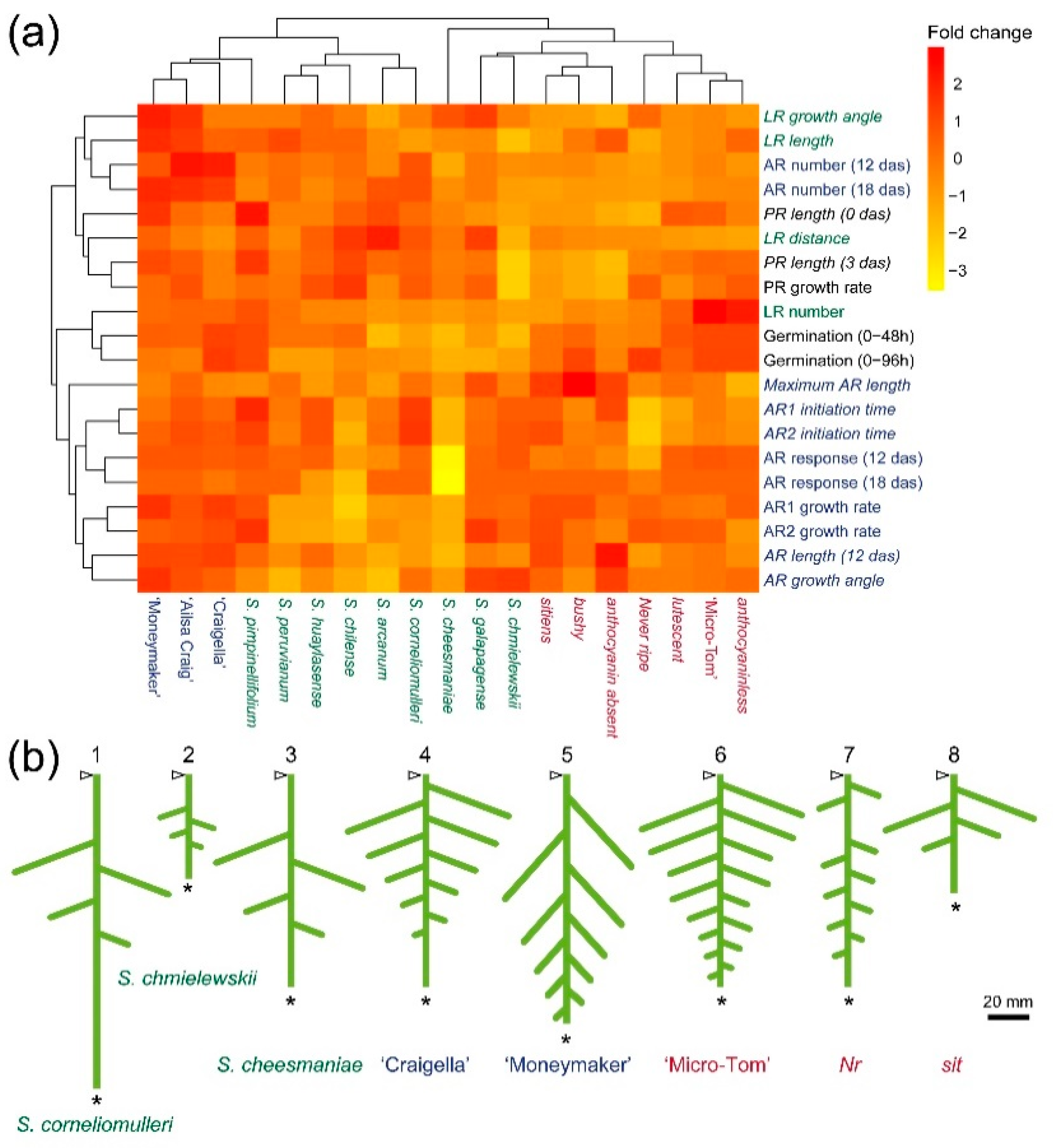
2.4. Capturing Quantitative Variation of Early RSA in Tomato
3. Materials and Methods
3.1. Plant Material and Growth Conditions
3.2. Image Analysis
3.3. Statistical Analyses
3.4. Heat Map Representation
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Petricka, J.J.; Winter, C.M.; Benfey, P.N. Control of Arabidopsis root development. Annu. Rev. Plant Biol. 2012, 63, 563–590. [Google Scholar] [CrossRef] [PubMed]
- Atkinson, J.A.; Rasmussen, A.; Traini, R.; Voss, U.; Sturrock, C.; Mooney, S.J.; Wells, D.M.; Bennett, M.J. Branching out in roots: Uncovering form, function, and regulation. Plant Physiol. 2014, 166, 538–550. [Google Scholar] [CrossRef]
- Coudert, Y.; Périn, C.; Courtois, B.; Khong, N.G.; Gantet, P. Genetic control of root development in rice, the model cereal. Trends Plant Sci. 2010, 15, 219–226. [Google Scholar] [CrossRef] [PubMed]
- Orman-Ligeza, B.; Parizot, B.; Gantet, P.P.; Beeckman, T.; Bennett, M.J.; Draye, X. Post-embryonic root organogenesis in cereals: Branching out from model plants. Trends Plant Sci. 2013, 18, 459–467. [Google Scholar] [CrossRef] [PubMed]
- Hochholdinger, F.; Yu, P.; Marcon, C. Genetic control of root system development in maize. Trends Plant Sci. 2018, 23, 79–88. [Google Scholar] [CrossRef] [PubMed]
- Food and Agriculture Organization of the United Nations (FAO). Available online: http://www.fao.org/land-water/databases-and-software/crop-information/tomato/en/ (accessed on 6 November 2018).
- Patanè, C.; Tringali, S.; Sortino, O. Effects of deficit irrigation on biomass, yield, water productivity and fruit quality of processing tomato under semi-arid Mediterranean climate conditions. Sci. Hortic. 2011, 129, 590–596. [Google Scholar] [CrossRef]
- Wang, X.; Xing, Y. Evaluation of the effects of irrigation and fertilization on tomato fruit yield and quality: A principal component analysis. Sci. Rep. 2017, 7, 350. [Google Scholar] [CrossRef] [PubMed]
- Lynch, J.P.; Chimungu, J.G.; Brown, K.M. Root anatomical phenes associated with water acquisition from drying soil: Targets for crop improvement. J. Exp. Bot. 2014, 65, 6155–6166. [Google Scholar] [CrossRef] [PubMed]
- Koevoets, I.T.; Venema, J.H.; Elzenga, J.T.M.; Testerink, C. Roots withstanding their environment: Exploiting root system architecture responses to abiotic stress to improve crop tolerance. Front. Plant Sci. 2016, 7, 1335. [Google Scholar] [CrossRef] [PubMed]
- Peralta, I.E.; Spooner, D.M.; Knapp, S. Taxonomy of wild tomatoes and their relatives (Solanum Sect. Lycopersicoides, Sect. Juglandifolia, Sect. Lycopersicon; Solanaceae). Syst. Bot. Monogr. 2008, 84, 1–186. [Google Scholar] [CrossRef]
- Aflitos, S.; Schijlen, E.; de Jong, H.; De Rider, D.; Smit, S.; Finkers, R.; Wang, J.; Zhang, G.; Li, N.; Mao, L.; et al. Exploring genetic variation in the tomato (Solanum section Lycopersicon) clade by whole-genome sequencing. Plant J. 2014, 80, 136–148. [Google Scholar] [CrossRef] [PubMed]
- Blanca, J.; Montero-Pau, J.; Sauvage, C.; Bauchet, G.; Illa, E.; Diez, M.J.; Francis, D.; Causse, M.; van der Knaap, E.; Cañizares, J. Genomic variation in tomato, from wild ancestors to contemporary breeding accessions. BMC Genom. 2015, 16, 257. [Google Scholar] [CrossRef]
- Ron, M.; Dorrity, M.W.; de Lucas, M.; Toal, T.; Hernandez, R.I.; Little, S.A.; Maloof, J.N.; Kiebenstein, D.J.; Brady, S.M. Identification of novel loci regulating interspecific variation in root morphology and cellular development in tomato. Plant Physiol. 2013, 162, 755–768. [Google Scholar] [CrossRef] [PubMed]
- Ivanchenko, M.G.; Coffeen, W.C.; Lomax, T.L.; Dubrovsky, J.G. Mutations in the Diageotropica (Dgt) gene uncouple patterned cell division during lateral root initiation from proliferative cell division in the pericycle. Plant J. 2006, 46, 436–447. [Google Scholar] [CrossRef] [PubMed]
- Ivanchenko, M.G.; Zhu, J.; Wang, B.; Medvecká, E.; Du, Y.; Azzarello, E.; Mancuso, S.; Megraw, M.; Filichkin, S.; Dubrovsky, J.G.; et al. The cyclophilin A DIAGEOTROPICA gene affects auxin transport in both root and shoot to control lateral root formation. Development 2015, 142, 712–721. [Google Scholar] [CrossRef]
- Maloney, G.S.; DiNapoli, K.T.; Muday, G.K. The anthocyanin reduced tomato mutant demonstrates the role of flavonols in tomato lateral root and root hair development. Plant Physiol. 2014, 166, 614–631. [Google Scholar] [CrossRef]
- Carvalho, R.F.; Campos, M.L.; Pino, L.E.; Crestana, S.L.; Zsögön, A.; Lima, J.E.; Benedito, V.A.; Peres, L.E. Convergence of developmental mutants into a single tomato model system: “Micro-Tom” as an effective toolkit for plant development research. Plant Methods 2011, 7, 18. [Google Scholar] [CrossRef]
- Martí, E.; Gisbert, C.; Bishop, G.J.; Dixon, M.S.; García-Martínez, J.L. Genetic and physiological characterization of tomato cv. Micro-Tom. J. Exp. Bot. 2006, 57, 2037–2047. [Google Scholar] [CrossRef]
- Zhang, L.; Huang, Z.; Wang, X.; Gao, J.; Guo, Y.; Du, Y.; Hu, H. Fine mapping and molecular marker development of anthocyanin absent, a seedling morphological marker for the selection of male sterile 10 in tomato. Mol. Breed. 2016, 36, 107. [Google Scholar] [CrossRef]
- De Jong, W.S.; Eannetta, N.T.; De Jong, D.M.; Bodis, M. Candidate gene analysis of anthocyanin pigmentation loci in the Solanaceae. Theor. Appl. Genet. 2004, 108, 423–432. [Google Scholar] [CrossRef]
- Wilkinson, J.Q.; Lanahan, M.B.; Yen, H.C.; Giovannoni, J.J.; Klee, H.J. An ethylene-inducible component of signal transduction encoded by Never-ripe. Science 1995, 270, 1807–1809. [Google Scholar] [CrossRef] [PubMed]
- Harrison, E.; Burbidge, A.; Okyere, J.P.; Thompson, A.J.; Taylor, I.B. Identification of the tomato ABA-deficient mutant sitiens as a member of the ABA-aldehyde oxidase gene family using genetic and genomic analysis. Plant Growth Regul. 2011, 64, 301–309. [Google Scholar] [CrossRef]
- Abbo, S.; Pinhasi van-Oss, R.; Gopher, A.; Saranga, Y.; Ofner, I.; Peleg, Z. Plant domestication versus crop evolution: A conceptual framework for cereals and grain legumes. Trends Plant Sci. 2014, 19, 351–360. [Google Scholar] [CrossRef]
- Bai, Y.; Lindhout, P. Domestication and breeding of tomatoes: What have we gained and what can we gain in the future? Ann. Bot. 2007, 100, 1085–1094. [Google Scholar] [CrossRef]
- Postma, J.A.; Dathe, A.; Lynch, J.P. The optimal lateral root branching density for maize depends on nitrogen and phosphorus availability. Plant Physiol. 2014, 166, 590–602. [Google Scholar] [CrossRef]
- Zhan, A.; Lynch, J.P. Reduced frequency of lateral root branching improves N capture from low-N soils in maize. J. Exp. Bot. 2015, 66, 2055–2065. [Google Scholar] [CrossRef]
- Osmont, K.S.; Sibout, R.; Hardtke, C.S. Hidden branches: Developments in root system architecture. Annu. Rev. Plant Biol. 2007, 58, 93–113. [Google Scholar] [CrossRef] [PubMed]
- Trachsel, S.; Kaeppler, S.M.; Brown, K.M.; Lynch, J.P. Shovelomics: High throughput phenotyping of maize (Zea mays L.) root architecture in the field. Plant Soil 2011, 341, 75–87. [Google Scholar] [CrossRef]
- Gruber, B.D.; Giehl, R.F.H.; Friedel, S.; von Wirén, N. Plasticity of the Arabidopsis root system under nutrient deficiencies. Plant Physiol. 2013, 163, 161–179. [Google Scholar] [CrossRef]
- Giehl, R.F.H.; Gruber, B.D.; von Wirén, N. It’s time to make changes: Modulation of root system architecture by nutrient signals. J. Exp. Bot. 2014, 65, 769–778. [Google Scholar] [CrossRef]
- Rellán-Álvarez, R.; Lobet, G.; Dinneny, J.R. Environmental control of root system biology. Annu. Rev. Plant Biol. 2016, 67, 619–642. [Google Scholar] [CrossRef] [PubMed]
- Van Norman, J.M.; Zhang, J.; Cazzonelli, C.I.; Pogson, B.J.; Harrison, P.J.; Bugg, T.D.H.; Chan, K.X.; Thompson, A.J.; Benfey, P.N. Periodic root branching in Arabidopsis requires synthesis of an uncharacterized carotenoid derivative. Proc. Natl. Acad. Sci. USA 2014, 111, E1300–E1309. [Google Scholar] [CrossRef] [PubMed]
- Loudet, O.; Gaudon, V.; Trubuil, A.; Daniel-Vedele, F. Quantitative trait loci controlling root growth and architecture in Arabidopsis thaliana confirmed by heterogeneous inbred family. Theor. Appl. Genet. 2005, 110, 742–753. [Google Scholar] [CrossRef] [PubMed]
- Shi, L.; Shi, T.; Broadley, M.R.; White, P.J.; Long, Y.; Meng, J.; Xu, F.; Hammond, J.P. High-throughput root phenotyping screens identify genetic loci associated with root architectural traits in Brassica napus under contrasting phosphate availabilities. Ann. Bot. 2013, 112, 381–389. [Google Scholar] [CrossRef] [PubMed]
- Song, W.; Wang, B.; Hauck, A.L.; Dong, X.; Li, J.; Lai, J. Genetic dissection of maize seedling root system architecture traits using an ultra-high density bin-map and a recombinant inbred line population. J. Integr. Plant Biol. 2016, 58, 266–279. [Google Scholar] [CrossRef] [PubMed]
- Miyazawa, Y.; Yamazaki, T.; Moriwaki, T.; Takahashi, H. Root tropism: Its mechanism and possible functions in drought avoidance. Adv. Bot. Res. 2011, 57, 349–375. [Google Scholar] [CrossRef]
- Van Norman, J.M.; Xuan, W.; Beeckman, T.; Benfey, P.N.; Poething, S.; Roberts, K.; Scheres, B. To branch or not to branch: The role of pre-patterning in lateral root formation. Development 2013, 140, 4301–4310. [Google Scholar] [CrossRef] [PubMed]
- Kato, Y.; Abe, J.; Kamoshita, A.; Yamagishi, J. Genotypic variation in root growth angle in rice (Oryza sativa L.) and its association with deep root development in upland fields with different water regimes. Plant Soil 2006, 287, 117–129. [Google Scholar] [CrossRef]
- Manschadi, A.M.; Christopher, J.; deVoil, P.; Hammer, G.L. The role of root architectural traits in adaptation of wheat to water-limited environments. Funct. Plant Biol. 2006, 33, 823–837. [Google Scholar] [CrossRef]
- Uga, Y.; Sugimoto, K.; Ogawa, S.; Rane, J.; Ishitani, M.; Hara, N.; Kitomi, Y.; Inukai, Y.; Ono, K.; Kanno, N.; et al. Control of root system architecture by DEEPER ROOTING 1 increases rice yield under drought conditions. Nat. Genet. 2013, 45, 1097–1102. [Google Scholar] [CrossRef] [PubMed]
- Guseman, J.M.; Webb, K.; Srinivasan, C.; Dardick, C. DRO1 influences root system architecture in Arabidopsis and Prunus species. Plant J. 2017, 89, 1093–1105. [Google Scholar] [CrossRef] [PubMed]
- Du, Y.; Scheres, B. Lateral root formation and the multiple roles of auxin. J. Exp. Bot. 2018, 69, 155–167. [Google Scholar] [CrossRef] [PubMed]
- Dubrovsky, J.G.; Gambetta, G.A.; Hernández-Barrera, A.; Shishkova, S.; González, I. Lateral root initiation in Arabidopsis: Developmental window, spatial patterning, density and predictability. Ann. Bot. 2006, 97, 903–915. [Google Scholar] [CrossRef] [PubMed]
- Péret, B.; De Rybel, B.; Casimiro, I.; Benková, E.; Swarup, R.; Laplaze, L.; Beeckman, T.; Bennett, M.J. Arabidopsis lateral root development: An emerging story. Trends Plant Sci. 2009, 14, 399–408. [Google Scholar] [CrossRef] [PubMed]
- Giehl, R.F.H.; von Wirén, N. Root nutrient foraging. Plant Physiol. 2014, 166, 509–517. [Google Scholar] [CrossRef] [PubMed]
- López-Bucio, J.; Hernández-Abreu, E.; Sánchez-Calderón, L.; Nieto-Jacobo, M.F.; Simpson, J.; Herrera-Estrella, L. Phosphate availability alters architecture and causes changes in hormone sensitivity in the Arabidopsis root system. Plant Physiol. 2002, 129, 244–256. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Torres, C.A.; López-Bucio, J.; Cruz-Ramírez, A.; Ibarra-Laclette, E.; Dharmasiri, S.; Estelle, M.; Herrera-Estrella, L. Phosphate availability alters lateral root development in Arabidopsis by modulating auxin sensitivity via a mechanism involving the TIR1 auxin receptor. Plant Cell 2008, 20, 3258–3272. [Google Scholar] [CrossRef]
- Ding, Z.; De Smet, I. Localised ABA signalling mediates root growth plasticity. Trends Plant Sci. 2013, 18, 533–535. [Google Scholar] [CrossRef]
- Zhang, H.; Han, W.; De Smet, I.; Talboys, P.; Loya, R.; Hassan, A.; Rong, H.; Jürgens, G.; Paul Knox, J.; Wang, M.H. ABA promotes quiescence of the quiescent centre and suppresses stem cell differentiation in the Arabidopsis primary root meristem. Plant J. 2010, 64, 764–774. [Google Scholar] [CrossRef]
- Sun, Y.; Li, H.; Huang, J.R. Arabidopsis TT19 functions as a carrier to transport anthocyanin from the cytosol to tonoplasts. Mol. Plant 2012, 5, 387–400. [Google Scholar] [CrossRef]
- Bellini, C.; Pacurar, D.I.; Perrone, I. Adventitious roots and lateral roots: Similarities and differences. Annu. Rev. Plant Biol. 2014, 65, 639–666. [Google Scholar] [CrossRef] [PubMed]
- Steffens, B.; Rasmussen, A. The physiology of adventitious roots. Plant Physiol. 2016, 170, 603–617. [Google Scholar] [CrossRef] [PubMed]
- da Costa, C.T.; de Almeida, M.R.; Ruedell, C.M.; Schwambach, J.; Maraschin, F.S.; Fett-Neto, A.G. When stress and development go hand in hand: Main hormonal controls of adventitious rooting in cuttings. Front. Plant Sci. 2013, 4, 133. [Google Scholar] [CrossRef] [PubMed]
- Druege, U.; Franken, P.; Hajirezaei, M.R. Plant hormone homeostasis, signaling, and function during adventitious root formation in cuttings. Front. Plant Sci. 2016, 7, 381. [Google Scholar] [CrossRef] [PubMed]
- Soyk, S.; Lemmon, Z.H.; Oved, M.; Fisher, J.; Liberatore, K.L.; Park, S.J.; Goren, A.; Jiang, K.; Ramos, A.; van der Knaap, E.; et al. Bypassing negative epistasis on yield in tomato imposed by a domestication gene. Cell 2017, 169, 1142–1155. [Google Scholar] [CrossRef] [PubMed]
- Van de Poel, B.; Smet, D.; Van Der Straeten, D. Ethylene and hormonal cross talk in vegetative growth and development. Plant Physiol. 2015, 169, 61–72. [Google Scholar] [CrossRef] [PubMed]
- Negi, S.; Ivanchenko, M.G.; Muday, G.K. Ethylene regulates lateral root formation and auxin transport in Arabidopsis thaliana. Plant J. 2008, 55, 175–187. [Google Scholar] [CrossRef]
- Negi, S.; Sukumar, P.; Liu, X.; Cohen, J.D.; Muday, G.K. Genetic dissection of the role of ethylene in regulating auxin-dependent lateral and adventitious root formation in tomato. Plant J. 2010, 61, 3–15. [Google Scholar] [CrossRef]
- Kim, H.J.; Lynch, J.P.; Brown, K.M. Ethylene insensitivity impedes a subset of responses to phosphorus deficiency in tomato and petunia. Plant Cell Environ. 2008, 31, 1744–1755. [Google Scholar] [CrossRef]
- Vidoz, M.L.; Loreti, E.; Mensuali, A.; Alpi, A.; Perata, P. Hormonal interplay during adventitious root formation in flooded tomato plants. Plant J. 2010, 63, 551–562. [Google Scholar] [CrossRef]
- Dawood, T.; Yang, X.; Visser, E.J.W.; Te Beek, T.A.; Kensche, P.R.; Cristescu, S.M.; Lee, S.; Floková, K.; Nguyen, D.; Mariani, C.; et al. A co-opted hormonal cascade activates dormant adventitious root primordia upon flooding in Solanum dulcamara. Plant Physiol. 2016, 170, 2351–2364. [Google Scholar] [CrossRef] [PubMed]
- Lynch, J.P.; Wojciechowski, T. Opportunities and challenges in the subsoil: Pathways to deeper rooted crops. J. Exp. Bot. 2015, 66, 2199–2210. [Google Scholar] [CrossRef] [PubMed]
- Henry, A.; Cal, A.J.; Batoto, T.C.; Torres, R.O.; Serraj, R. Root attributes affecting water uptake of rice (Oryza sativa) under drought. J. Exp. Bot. 2012, 63, 4751–4763. [Google Scholar] [CrossRef] [PubMed]
- Burton, A.L.; Brown, K.M.; Lynch, J.P. Phenotypic diversity of root anatomical and architectural traits in Zea species. Crop Sci. 2013, 53, 1042–1055. [Google Scholar] [CrossRef]
- Lynch, J.P. Steep, cheap and deep: An ideotype to optimize water and N acquisition by maize root systems. Ann. Bot. 2013, 112, 347–357. [Google Scholar] [CrossRef] [PubMed]
- Thomas, C.L.; Graham, N.S.; Hayden, R.; Meacham, M.C.; Neugebauer, K.; Nightingale, M.; Dupuy, L.X.; Hammond, J.P.; White, P.J.; Broadley, M.R. High-throughput phenotyping (HTP) identifies seedling root traits linked to variation in seed yield and nutrient capture in field-grown oilseed rape (Brassica napus L.). Ann. Bot. 2016, 118, 655–665. [Google Scholar] [CrossRef] [PubMed]
- Pflugfelder, D.; Metzner, R.; van Dusschoten, D.; Reichel, R.; Jahnke, S.; Koller, R. Non-invasive imaging of plant roots in different soils using magnetic resonance imaging (MRI). Plant Methods 2017, 13, 102. [Google Scholar] [CrossRef]
- Rambla, J.L.; Medina, A.; Fernández-del-Carmen, A.; Barrantes, W.; Grandillo, S.; Cammareri, M.; López-Casado, G.; Rodrigo, G.; Alonso, A.; García-Martínez, S.; et al. Identification, introgression, and validation of fruit volatile QTLs from a red-fruited wild tomato species. J. Exp. Bot. 2016, 68, 429–442. [Google Scholar] [CrossRef]
- Okabe, Y.; Ariizumi, T.; Ezura, H. Updating the Micro-Tom TILLING platform. Breed. Sci. 2013, 63, 42–48. [Google Scholar] [CrossRef]
- Kudo, T.; Kobayashi, M.; Terashima, S.; Katayama, M.; Ozaki, S.; Kanno, M.; Saito, M.; Yokoyama, K.; Ohyanagi, H.; Aoki, K.; et al. TOMATOMICS: A web database for integrated omics information in tomato. Plant Cell Physiol. 2017, 58, e8. [Google Scholar] [CrossRef]
- Shikata, M.; Ezura, H. Micro-Tom tomato as an alternative plant model system: Mutant collection and efficient transformation. Methods Mol. Biol. 2016, 16, 47–55. [Google Scholar] [CrossRef]
- Shikata, M.; Hoshikawa, K.; Ariizumi, T.; Fukuda, N.; Yamazaki, Y.; Ezura, H. TOMATOMA update: Phenotypic and metabolite information in the Micro-Tom mutant resource. Plant Cell Physiol. 2016, 57, e11. [Google Scholar] [CrossRef] [PubMed]
- Garcia, V.; Bres, C.; Just, D.; Fernandez, L.; Tai, F.W.; Mauxion, J.P.; Le Paslier, M.C.; Bèrard, A.; Brunel, D.; Aoki, K.; et al. Rapid identification of causal mutations in tomato EMS populations via mapping-by-sequencing. Nat. Protoc. 2016, 11, 2401–2418. [Google Scholar] [CrossRef] [PubMed]
- Schindelin, J.; Arganda-Carreras, I.; Frise, E.; Kaynig, V.; Logair, M.; Pietzsch, T.; Preibisch, S.; Rueden, C.; Saalfeld, S.; Schmid, B.; et al. Fiji: An open-source platform for biological-image analysis. Nat. Methods 2012, 9, 676–682. [Google Scholar] [CrossRef] [PubMed]
- Armengaud, P.; Zambaux, K.; Hills, A.; Sulpice, R.; Pattison, R.J.; Blatt, M.R.; Amtmann, A. EZ-Rhizo: Integrated software for the fast and accurate measurement of root system architecture. Plant J. 2009, 57, 945–956. [Google Scholar] [CrossRef] [PubMed]
- Aguinis, H.; Gottfredson, R.K.; Joo, H. Best-practice recommendations for defining, identifying, and handling outliers. Organ. Res. Methods 2013, 16, 270–301. [Google Scholar] [CrossRef]
| 1. Wild Tomato Species | ||||||
|---|---|---|---|---|---|---|
| Species Name ‘Cultivar’ | Accession | Collection Site | Latitude | Longitude | Altitude | Other Comments |
| S. corneliomulleri | LA1274 | Lima (Perú) | 11°27′36″ | 76°54′0″ | 1440 m | Fruits from 6 plants |
| S. peruvianum | LA1336 | Arequipa (Perú) | ||||
| S. chilense | LA1932 | Arequipa (Perú) | 15°25′0″ | 74°42′0″ | 1100 m | Fruits from 15 plants, stress tolerant |
| S. huaylasense | LA1983 | Ancash (Perú) | 8°4121″ | 77°58′20″ | 940 m | Fruits from 1 plant, very dry spot |
| S. chmielewskii | LA2663 | Cusco (Perú) | 13°41′44″ | 74°59′39″ | 2500 m | Fruits form 6–7 plants |
| S. arcanum | LA2157 | Cajamarca (Perú) | 6°30′21″ | 78°48′32″ | 1600 m | Fruits from 2 plants |
| S. galapagense | LA1044 | Galapagos Islands (Ecuador) | 0°17′4″ | 90°32′54″ | <100 m | |
| S. cheesmaniae | LA1037 | Galapagos Islands (Ecuador) | 0°25′21″ | 91°7′0″ | 800 m | From bottom of volcano crater |
| S. pimpinelifollium | LA1587 | La Libertat (Perú) | 7°20′0″ | 79°35′0″ | <100 m | Fruits from 20 plants. Grown on river sand |
| 2. Commercial Tomato Cultivars | 3. ‘Micro-Tom’ developmental mutants | |||||
| ‘Cultivar name’ | Accession | Mutant | Phenotype | Gene Product | Gene Function | References |
| ‘Ailsa Craig’ | LA2838A | anthocyanin absent (aa) | Anthocyanin deficient | SlGSTAA | Anthocyanin transport | [18,20] |
| ‘Craigella’ | LA3247 | anthocyaninless (a) | Low anthocyanin levels | F3’5’H | Anthocyanin biosynthesis | [18,21] |
| ‘Moneymaker’ | LA2706 | bushy (bu) | Short internodes | Unknown | [18] | |
| ‘Micro-Tom’ | LA4480 | lutescent (l) | Premature senescence | Unknown | [18] | |
| Never ripe (Nr) | Low ethylene responses | SlETR3 | Ethylene receptor | [18,22] | ||
| sitiens (sit) | ABA deficient | ABA aldehyde oxidase | ABA biosynthesis | [18,23] | ||
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Alaguero-Cordovilla, A.; Gran-Gómez, F.J.; Tormos-Moltó, S.; Pérez-Pérez, J.M. Morphological Characterization of Root System Architecture in Diverse Tomato Genotypes during Early Growth. Int. J. Mol. Sci. 2018, 19, 3888. https://doi.org/10.3390/ijms19123888
Alaguero-Cordovilla A, Gran-Gómez FJ, Tormos-Moltó S, Pérez-Pérez JM. Morphological Characterization of Root System Architecture in Diverse Tomato Genotypes during Early Growth. International Journal of Molecular Sciences. 2018; 19(12):3888. https://doi.org/10.3390/ijms19123888
Chicago/Turabian StyleAlaguero-Cordovilla, Aurora, Francisco Javier Gran-Gómez, Sergio Tormos-Moltó, and José Manuel Pérez-Pérez. 2018. "Morphological Characterization of Root System Architecture in Diverse Tomato Genotypes during Early Growth" International Journal of Molecular Sciences 19, no. 12: 3888. https://doi.org/10.3390/ijms19123888
APA StyleAlaguero-Cordovilla, A., Gran-Gómez, F. J., Tormos-Moltó, S., & Pérez-Pérez, J. M. (2018). Morphological Characterization of Root System Architecture in Diverse Tomato Genotypes during Early Growth. International Journal of Molecular Sciences, 19(12), 3888. https://doi.org/10.3390/ijms19123888







