Overexpression of the DEAD-Box RNA Helicase Gene AtRH17 Confers Tolerance to Salt Stress in Arabidopsis
Abstract
1. Introduction
2. Results
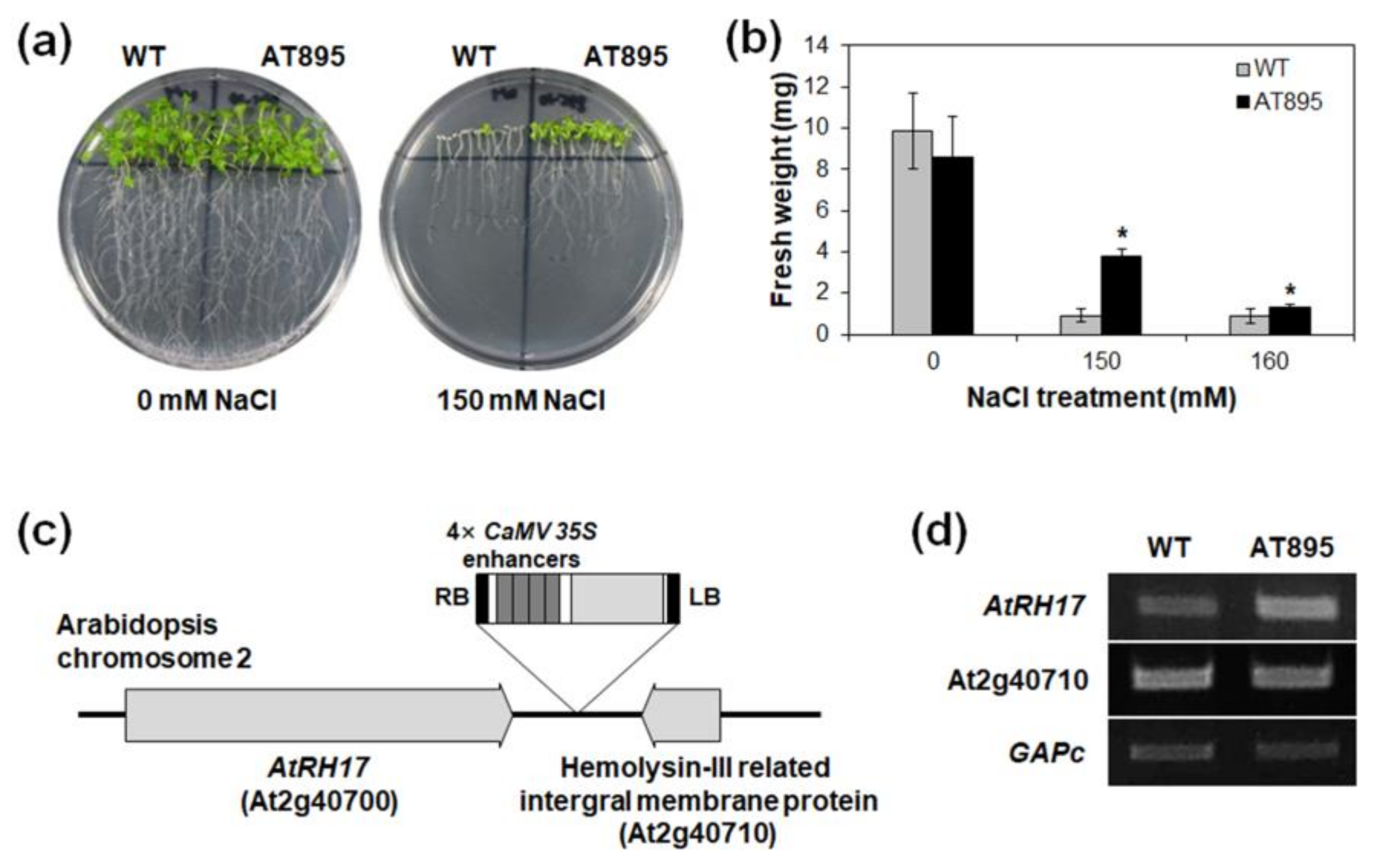
2.1. Isolation of AtRH17 via Activation Tagging
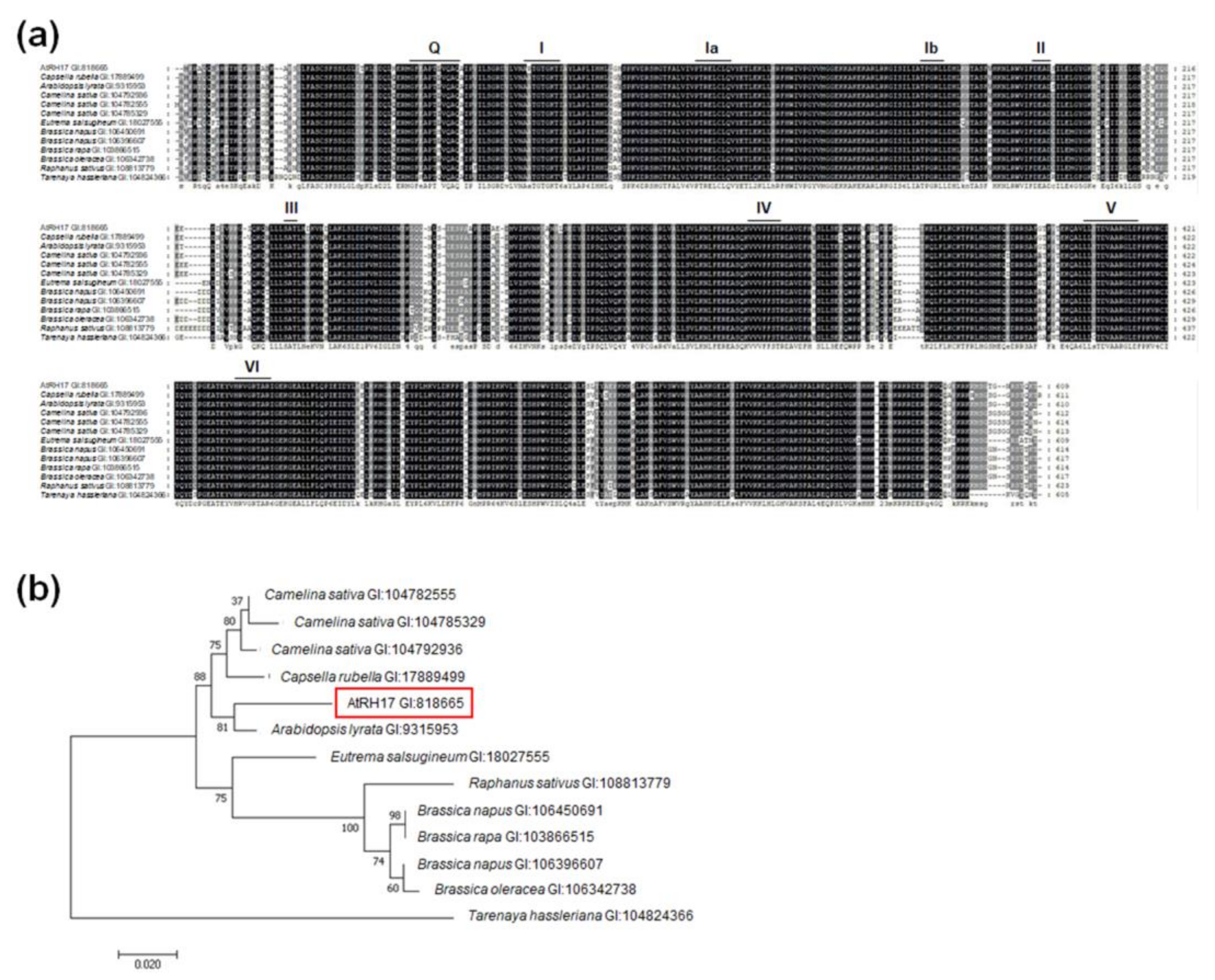
2.2. Phylogenetic Analysis of AtRH17
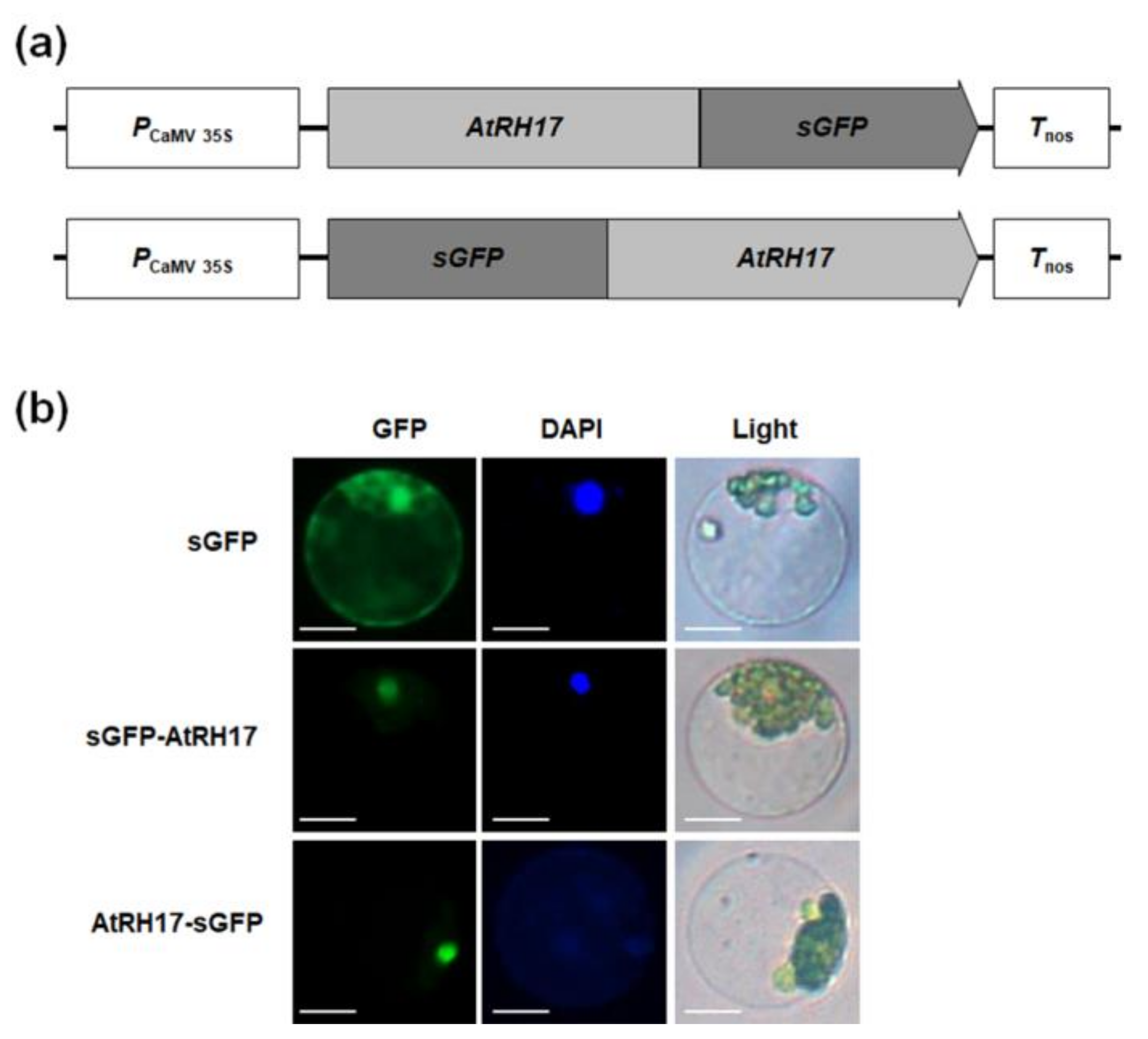
2.3. Subcellular Localization of AtRH17 in the Nucleus
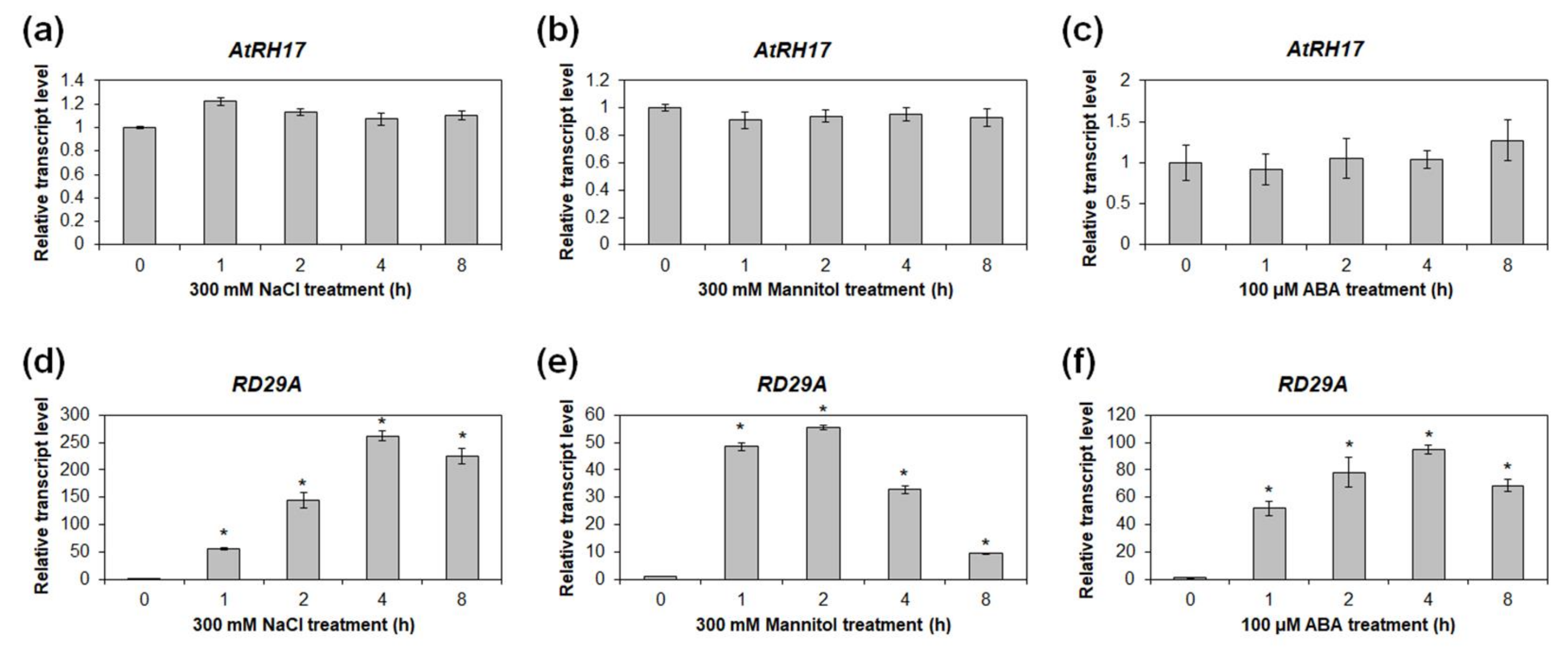
2.4. Expression of AtRH17 Is Unaffected by Osmotic Stress Conditions
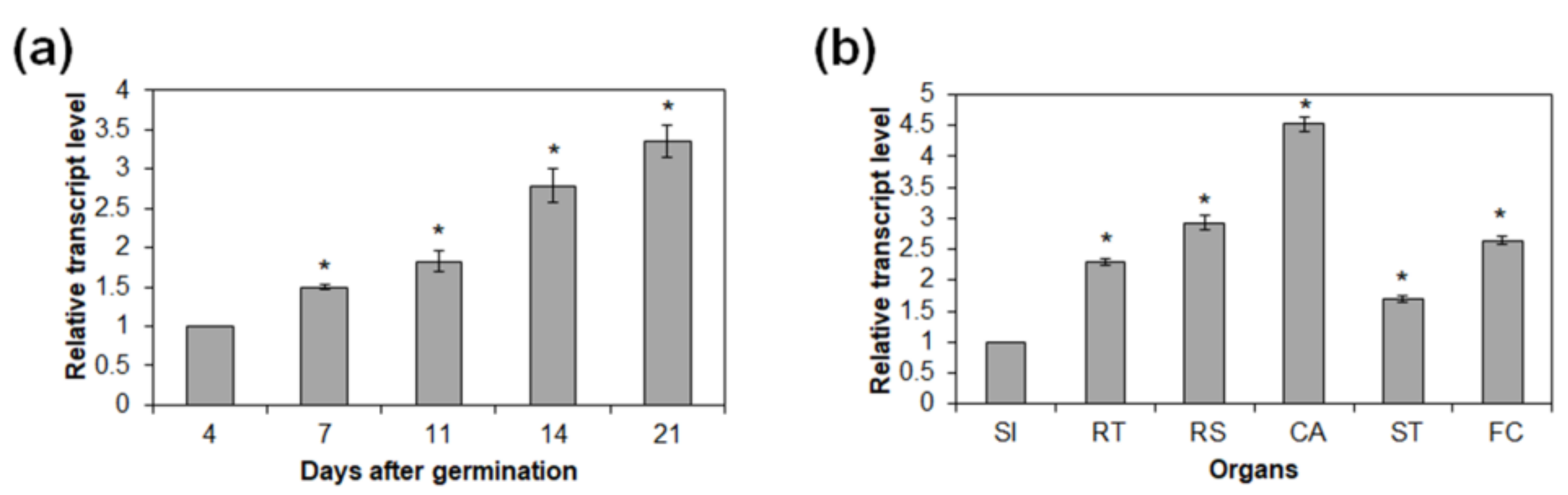
2.5. Expression of AtRH17 during Developmental Stages and in Organs
2.6. AtRH17 OXs Exhibit Tolerance to Salt Stresses, but Not to Mannitol and Freezing Stresses
2.7. Expression of ABA-Dependent and ABA-Independent Salt-Stress-Responsive Genes Does Not Alter, but is Lower in AtRH17 OXs under Salt Stress Conditions
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. Plasmid Construction and Plant Transformation
4.3. Activation Tagging Line Screening and Plant Stress Treatments
4.4. PS II Activity (Fv/Fm) and Chlorophyll Content Measurement
4.5. Histochemical Staining of Superoxide Production
4.6. RNA Extraction and RT-PCR Analysis
4.7. Multiple Sequence Alignment and Phylogenetic Analysis
4.8. Subcellular Localization of the AtRH17-GFP Fusion Protein
4.9. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| ABA | Abscisic acid |
| AT | Activation tagging |
| CaMV | Cauliflower mosaic virus |
| DAG | Days after germination |
| LD | Long-day |
| MS | Murashige and Skoog |
| NBT | Nitro blue tetrazolium |
| OXs | Overexpressing transgenic plants |
| RH | RNA helicase |
| ROS | Reactive oxygen species |
| SD | Short-day |
References
- Nakashima, K.; Takasaki, H.; Mizoi, J.; Shinozaki, K.; Yamaguchi-Shinozaki, K. NAC transcription factors in plant abiotic stress responses. Biochim. Biophys. Acta 2012, 1819, 97–103. [Google Scholar] [CrossRef] [PubMed]
- Borkotoky, S.; Saravanan, V.; Jaiswal, A.; Das, B.; Selvaraj, S.; Murali, A.; Lakshmi, P.T.V. The Arabidopsis stress responsive gene database. Int. J. Plant Genom. 2013, 2013, 949564. [Google Scholar] [CrossRef] [PubMed]
- Aubourg, S.; Kreis, M.; Lecharny, A. The DEAD box RNA helicase family in Arabidopsis thaliana. Nucleic Acids Res. 1999, 27, 628–636. [Google Scholar] [CrossRef] [PubMed]
- Cordin, O.; Banroques, J.; Tanner, N.K.; Linder, P. The DEAD-box protein family of RNA helicases. Gene 2006, 367, 17–37. [Google Scholar] [CrossRef] [PubMed]
- Linder, P.; Jankowsky, E. From unwinding to clamping—The DEAD box RNA helicase family. Nat. Rev. Mol. Cell Biol. 2011, 12, 505–516. [Google Scholar] [CrossRef] [PubMed]
- Owttrim, G.W. RNA helicases: Diverse roles in prokaryotic response to abiotic stress. RNA Biol. 2013, 10, 96–110. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Liu, C.; Li, M.; Li, H.; Hu, J.; Zhu, L.; Zeng, D.; Yang, Y.; Peng, Y.; Ruan, B.; et al. A rice DEAD-box RNA helicase protein, OsRH17, suppresses 16S ribosomal RNA maturation in Escherichia coli. Gene 2015, 555, 318–328. [Google Scholar] [CrossRef] [PubMed]
- Linder, P.; Rocak, S. DEAD-box proteins: The driving forces behind RNA metabolism. Nat. Rev. Mol. Cell Biol. 2004, 5, 232–241. [Google Scholar] [CrossRef]
- Fuller-Pace, F.V. DExD/H box RNA helicases: Multifunctional proteins with important roles in transcriptional regulation. Nucleic Acids Res. 2006, 34, 4206–4215. [Google Scholar] [CrossRef] [PubMed]
- Nawaz, G.; Kang, H. Chloroplast- or mitochondria-targeted DEAD-box RNA helicases play essential roles in organellar RNA metabolism and abiotic stress responses. Front. Plant Sci. 2017, 8, 871. [Google Scholar] [CrossRef] [PubMed]
- Tanner, N.K.; Cordin, O.; Banroques, J.; Doère, M.; Linder, P. The Q motif: A newly identified motif in DEAD box helicases may regulate ATP binding and hydrolysis. Mol. Cell 2003, 11, 127–138. [Google Scholar] [CrossRef]
- Iost, I.; Dreyfus, M. DEAD-box RNA helicases in Escherichia coli. Nucleic Acids Res. 2006, 34, 4189–4197. [Google Scholar] [CrossRef] [PubMed]
- Martin, R.; Straub, A.U.; Doebele, C.; Bohnsack, M.T. DExD/H-box RNA helicases in ribosome biogenesis. RNA Biol. 2013, 10, 4–18. [Google Scholar] [CrossRef] [PubMed]
- Giraud, C.; Hausmann, S.; Lemeille, S.; Prados, J.; Redder, P.; Linder, P. The C-terminal region of the RNA helicase CshA is required for the interaction with the degradosome and turnover of bulk RNA in the opportunistic pathogen Staphylococcus aureus. RNA Biol. 2015, 12, 658–674. [Google Scholar] [CrossRef] [PubMed]
- Banroques, J.; Cordin, O.; Doère, M.; Linder, P.; Tanner, N.K. Analyses of the functional regions of DEAD-box RNA “Helicases” with deletion and chimera constructs tested in vivo and in vitro. J. Mol. Biol. 2011, 413, 451–472. [Google Scholar] [CrossRef] [PubMed]
- Gong, Z.; Dong, C.; Lee, H.; Zhu, J.; Xiong, L.; Gong, D.; Stevenson, B.; Zhu, J.K. A DEAD box RNA helicase is essential for mRNA export and important for development and stress responses in Arabidopsis. Plant Cell 2005, 17, 256–267. [Google Scholar] [CrossRef] [PubMed]
- Gu, L.; Xu, T.; Lee, K.H.; Kang, H. A chloroplast-localized DEAD-box RNA helicase AtRH3 is essential for intron splicing and plays an important role in the growth and stress response in Arabidopsis thaliana. Plant Physiol. Biochem. 2014, 81, 309–318. [Google Scholar] [CrossRef] [PubMed]
- Kanai, M.; Hayashi, M.; Kondo, M.; Nishimura, M. The plastidic DEAD-box RNA helicase 22, HS3, is essential for plastid functions both in seed development and in seedling growth. Plant Cell Physiol. 2013, 54, 1431–1440. [Google Scholar] [CrossRef] [PubMed]
- Nishimura, K.; Ashida, H.; Ogawa, T.; Yokota, A. A DEAD box protein is required for formation of a hidden break in Arabidopsis chloroplast 23S rRNA. Plant J. 2010, 63, 766–777. [Google Scholar] [CrossRef] [PubMed]
- Gong, Z.; Lee, H.; Xiong, L.; Jagendorf, A.; Stevenson, B.; Zhu, J.K. RNA helicase-like protein as an early regulator of transcription factors for plant chilling and freezing tolerance. Proc. Natl. Acad. Sci. USA 2002, 99, 11507–11512. [Google Scholar] [CrossRef] [PubMed]
- Bond, A.T.; Mangus, D.A.; He, F.; Jacobson, A. Absence of Dbp2p alters both nonsense-mediated mRNA decay and rRNA processing. Mol. Cell. Biol. 2001, 21, 7366–7379. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Sie, Y.; Chen, Y.; Huang, T.; Lu, C. Two highly similar DEAD box proteins, OsRH2 and OsRH34, homologous to eukaryotic initiation factor 4AIII, play roles of the exon junction complex in regulating growth and development in rice. BMC Plant Biol. 2016, 16, 84. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.; Kang, H. Emerging roles of RNA-binding proteins in plant growth, development, and stress responses. Mol. Cells 2016, 39, 179–185. [Google Scholar] [CrossRef] [PubMed]
- Stonebloom, S.; Burch-Smith, T.; Kim, I.; Meinke, D.; Mindrinoss, M.; Zambryski, P. Loss of the plant DEAD-box protein ISE1 leads to defective mitochondria and increased cell-to-cell transport via plasmodesmata. Proc. Natl. Acad. Sci. USA 2009, 106, 17229–17234. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Huang, L.; Huang, J.; Wu, S.; Yeh, C.; Lu, C. A DEAD-box protein, AtRH36, is essential for female gametophyte development and is involved in rRNA biogenesis in Arabidopsis. Plant Cell Physiol. 2010, 51, 694–706. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.; Shi, D.Q.; Yuan, L.; Liu, J.; Yang, W.C. SLOW WALKER3, encoding a putative DEAD-box RNA helicase, is essential for female gametogenesis in Arabidopsis. J. Integr. Plant Biol. 2010, 52, 817–828. [Google Scholar] [CrossRef] [PubMed]
- Hsu, Y.; Chen, Y.; Hsiao, Y.; Wang, B.; Lin, S.; Cheng, W.; Jauh, G.; Harada, J.J.; Wang, C. AtRH57, a DEAD-box RNA helicase, is involved in feedback inhibition of glucose-mediated abscisic acid accumulation during seedling development and additively affects pre-ribosomal RNA processing with high glucose. Plant J. 2014, 77, 119–135. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Gao, X.; Wei, Y.; Deng, L.; Ouyang, Y.; Chen, G.; Li, X.; Zhang, O.; Wu, C. Rice APOPTOSIS INHIBITOR5 coupled with two DEAD-box adenosine 5′-triphosphate-dependent RNA helicases regulates tapetum degeneration. Plant Cell 2011, 23, 1416–1434. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Imai, R. Function of plant DExD/H-box RNA helicases associated with ribosomal RNA biogenesis. Front. Plant Sci. 2018, 9, 125. [Google Scholar] [CrossRef] [PubMed]
- Baruah, I.; Debbarma, J.; Boruah, H.P.D.; Keshavaiah, C. The DEAD-box RNA helicases and multiple abiotic stresses in plants: A systematic review of recent advantages and challenges. Plant Omics J. 2017, 10, 252–262. [Google Scholar] [CrossRef]
- Zhu, J.; Dong, C.; Zhu, J.K. Interplay between cold-responsive gene regulation, metabolism and RNA processing during plant cold acclimation. Curr. Opin. Plant Biol. 2007, 10, 290–295. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Shen, Y.; Huang, L.; Wu, S.; Yeh, C.; Lu, C. The DEAD-box RNA helicase AtRH7/PRH75 participates in pre-rRNA processing, plant development and cold tolerance in Arabidopsis. Plant Cell Physiol. 2016, 57, 174–191. [Google Scholar] [CrossRef] [PubMed]
- Guan, Q.; Wu, J.; Zhang, Y.; Jiang, C.; Liu, R.; Chai, C.; Zhu, J. A DEAD box RNA helicase is critical for pre-mRNA splicing, cold-responsive gene regulation, and cold tolerance in Arabidopsis. Plant Cell 2013, 25, 342–356. [Google Scholar] [CrossRef] [PubMed]
- Kant, P.; Kant, S.; Gordon, M.; Shaked, R.; Barak, S. Stress Response Suppressor1 and Stress Response Suppressor2, two DEAD-box RNA helicases that attenuate Arabidopsis responses to multiple abiotic stresses. Plant Physiol. 2007, 145, 814–830. [Google Scholar] [CrossRef] [PubMed]
- Khan, A.; Garbelli, A.; Grossi, S.; Florentin, A.; Batelli, G.; Acuna, T.; Zolla, G.; Kaye, Y.; Paul, L.K.; Zhu, J.K.; et al. The Arabidopsis STRESS RESPONSE SUPPRESSOR DEAD-box RNA helicases are nucleolar- and chromocenter-localized proteins that undergo stress-mediated relocalization and are involved in epigenetic gene silencing. Plant J. 2014, 79, 28–43. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.S.; Kim, K.A.; Oh, T.R.; Park, C.M.; Kang, H. Functional characterization of DEAD-box RNA helicases in Arabidopsis thaliana under abiotic stress conditions. Plant Cell Physiol. 2008, 49, 1563–1571. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Liu, H.; Zhang, H.; Wang, X.; Song, F. OsBIRH1, a DEAD-box RNA helicase with functions in modulating defence responses against pathogen infection and oxidative stress. J. Exp. Bot. 2008, 59, 2133–2146. [Google Scholar] [CrossRef] [PubMed]
- Sanan-Mishra, N.; Pham, X.H.; Sopory, S.K.; Tuteja, N. Pea DNA helicase 45 overexpression in tobacco confers high salinity tolerance without affecting yield. Proc. Natl. Acad. Sci. USA 2005, 102, 509–514. [Google Scholar] [CrossRef] [PubMed]
- Shivakumara, T.N.; Sreevathsa, R.; Dash, P.K.; Sheshshayee, M.S.; Papolu, P.K.; Rao, U.; Tuteja, N.; UdayaKumar, M. Overexpression of pea DNA helicase 45 (PDH45) imparts tolerance to multiple abiotic stresses in chili (Capsicum annuum L.). Sci. Rep. 2017, 7, 2760. [Google Scholar] [CrossRef] [PubMed]
- Jeong, D.H.; An, S.; Kang, H.G.; Moon, S.; Han, J.J.; Park, S.; Lee, H.S.; An, K.; An, G. T-DNA insertional mutagenesis for activation tagging in rice. Plant Physiol. 2002, 130, 1636–1644. [Google Scholar] [CrossRef] [PubMed]
- Marsch-Martinez, N.; Greco, R.; Van Arkel, G.; Herrera-Estrella, L.; Pereira, A. Activation tagging using the En-I maize transposon system in Arabidopsis. Plant Physiol. 2002, 129, 1544–1556. [Google Scholar] [CrossRef] [PubMed]
- Ito, T.; Meyerowitz, E.M. Overexpression of a gene encoding a cytochrome P450, CYP78A9, induces large and seedless fruit in Arabidopsis. Plant Cell 2000, 12, 1541–1550. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.Y.; Seok, H.Y.; Tarte, V.N.; Woo, D.H.; Le, D.H.; Lee, E.H.; Moon, Y.H. The Arabidopsis chloroplast protein S-RBP11 is involved in oxidative and salt stress responses. Plant Cell Rep. 2014, 33, 837–847. [Google Scholar] [CrossRef] [PubMed]
- Ashraf, M.; Harris, P.J.C. Potential biochemical indicators of salinity tolerance in plants. Plant Sci. 2004, 166, 3–16. [Google Scholar] [CrossRef]
- Ashraf, M. Biotechnological approach of improving plant salt tolerance using antioxidants as markers. Biotechnol. Adv. 2009, 27, 84–93. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; McFarlane, H.E.; Persson, S. The impact of abiotic factors on cellulose synthesis. J. Exp. Bot. 2016, 67, 543–552. [Google Scholar] [CrossRef] [PubMed]
- Nakashima, K.; Ito, Y.; Yamaguchi-Shinozaki, K. Transcriptional regulatory networks in response to abiotic stresses in Arabidopsis and grasses. Plant Physiol. 2009, 149, 88–95. [Google Scholar] [CrossRef] [PubMed]
- Barak, S.; Singh Yadav, N.; Khan, A. DEAD-box RNA helicases and epigenetic control of abiotic stress-responsive gene expression. Plant Signal. Behav. 2014, 9, e977729. [Google Scholar] [CrossRef] [PubMed]
- Tarte, V.N.; Seok, H.Y.; Woo, D.H.; Le, D.H.; Tran, H.T.; Baik, J.W.; Kang, I.S.; Lee, S.Y.; Chung, T.; Moon, Y.H. Arabidopsis Qc-SNARE gene AtSFT12 is involved in salt and osmotic stress responses and Na+ accumulation in vacuoles. Plant Cell Rep. 2015, 34, 1127–1138. [Google Scholar] [CrossRef] [PubMed]
- Seok, H.Y.; Woo, D.H.; Nguyen, L.V.; Tran, H.T.; Tarte, V.N.; Mehdi, S.M.M.; Lee, S.Y.; Moon, Y.H. Arabidopsis AtNAP functions as a negative regulator via repression of AREB1 in salt stress response. Planta 2017, 245, 329–341. [Google Scholar] [CrossRef] [PubMed]
- Höfgen, R.; Willmitzer, L. Storage of competent cells for Agrobacterium transformation. Nucleic Acids Res. 1988, 16, 9877. [Google Scholar] [CrossRef] [PubMed]
- Clough, S.J.; Bent, A.F. Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 1998, 16, 735–743. [Google Scholar] [CrossRef] [PubMed]
- Flohé, L.; Otting, F. Superoxide dismutase assays. Methods Enzymol. 1984, 105, 93–104. [Google Scholar] [CrossRef] [PubMed]
- Sheen, J. Signal transduction in maize and Arabidopsis mesophyll protoplasts. Plant Physiol. 2001, 127, 1466–1475. [Google Scholar] [CrossRef] [PubMed]



© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nguyen, L.V.; Seok, H.-Y.; Woo, D.-H.; Lee, S.-Y.; Moon, Y.-H. Overexpression of the DEAD-Box RNA Helicase Gene AtRH17 Confers Tolerance to Salt Stress in Arabidopsis. Int. J. Mol. Sci. 2018, 19, 3777. https://doi.org/10.3390/ijms19123777
Nguyen LV, Seok H-Y, Woo D-H, Lee S-Y, Moon Y-H. Overexpression of the DEAD-Box RNA Helicase Gene AtRH17 Confers Tolerance to Salt Stress in Arabidopsis. International Journal of Molecular Sciences. 2018; 19(12):3777. https://doi.org/10.3390/ijms19123777
Chicago/Turabian StyleNguyen, Linh Vu, Hye-Yeon Seok, Dong-Hyuk Woo, Sun-Young Lee, and Yong-Hwan Moon. 2018. "Overexpression of the DEAD-Box RNA Helicase Gene AtRH17 Confers Tolerance to Salt Stress in Arabidopsis" International Journal of Molecular Sciences 19, no. 12: 3777. https://doi.org/10.3390/ijms19123777
APA StyleNguyen, L. V., Seok, H.-Y., Woo, D.-H., Lee, S.-Y., & Moon, Y.-H. (2018). Overexpression of the DEAD-Box RNA Helicase Gene AtRH17 Confers Tolerance to Salt Stress in Arabidopsis. International Journal of Molecular Sciences, 19(12), 3777. https://doi.org/10.3390/ijms19123777





