Structural Analysis of Variability and Interaction of the N-terminal of the Oncogenic Effector CagA of Helicobacter pylori with Phosphatidylserine
Abstract
1. Introduction
2. Results
2.1. Sequence Selection and Multiple Sequence Alignment (MSA)
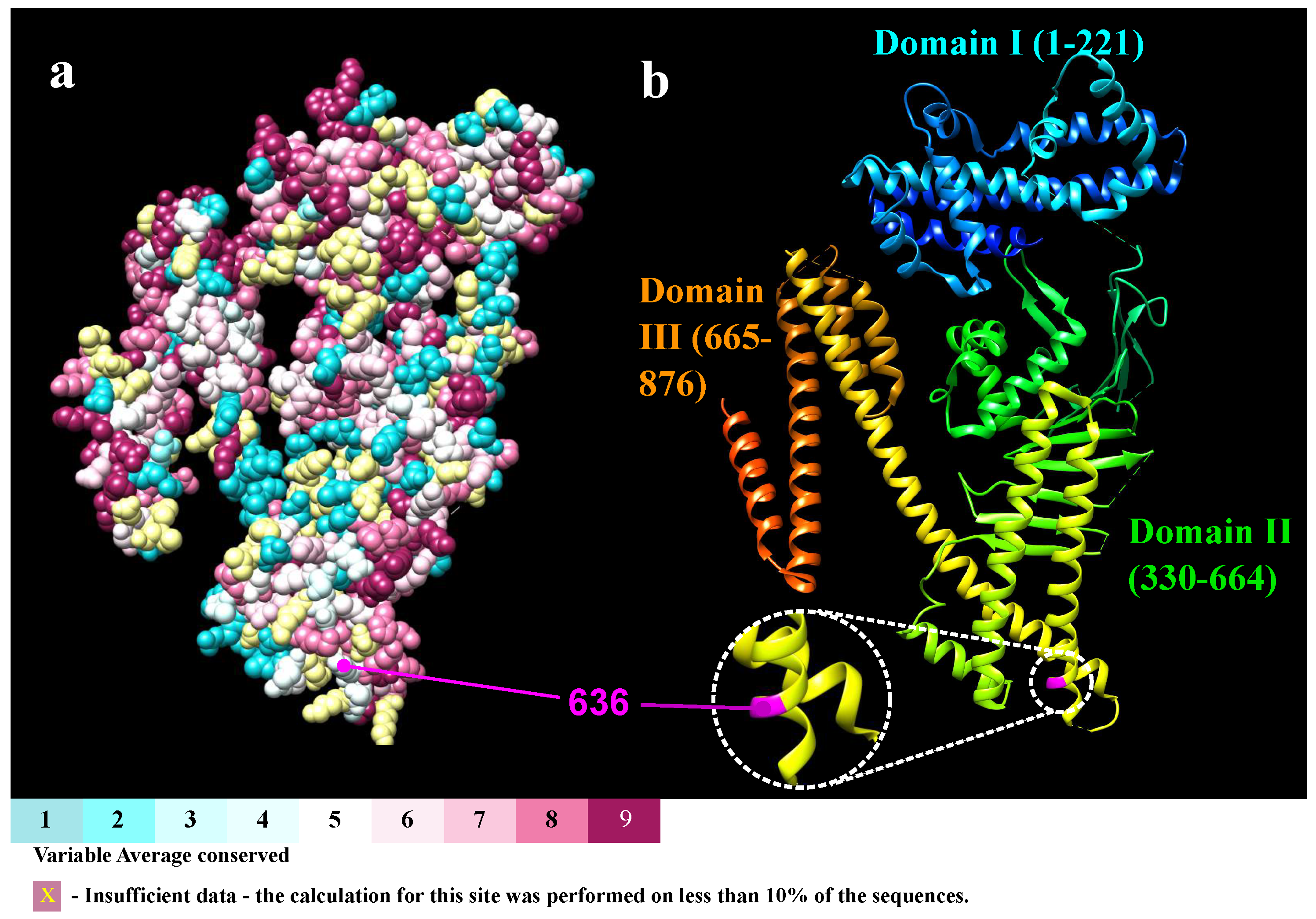
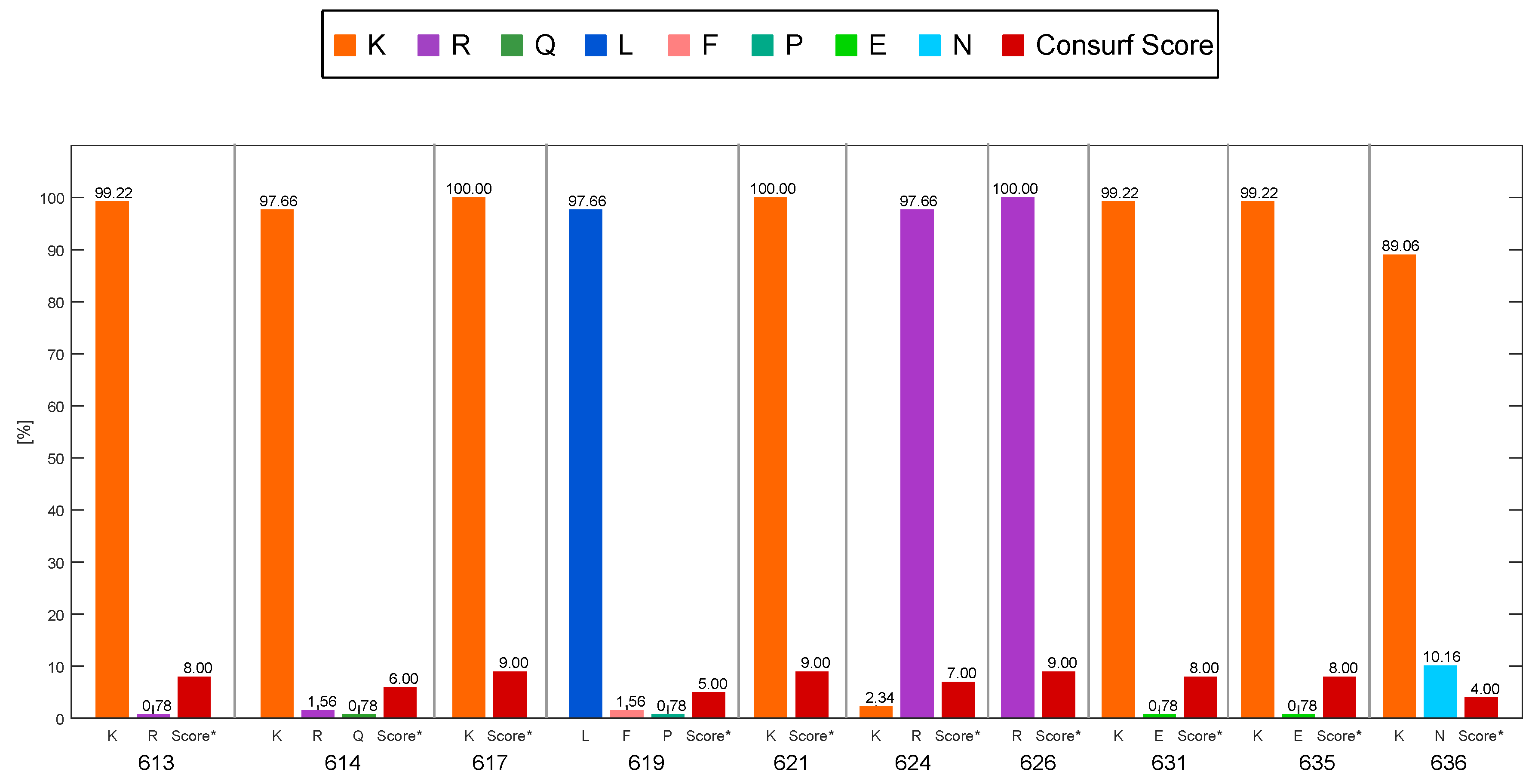
2.2. Consurf Analysis
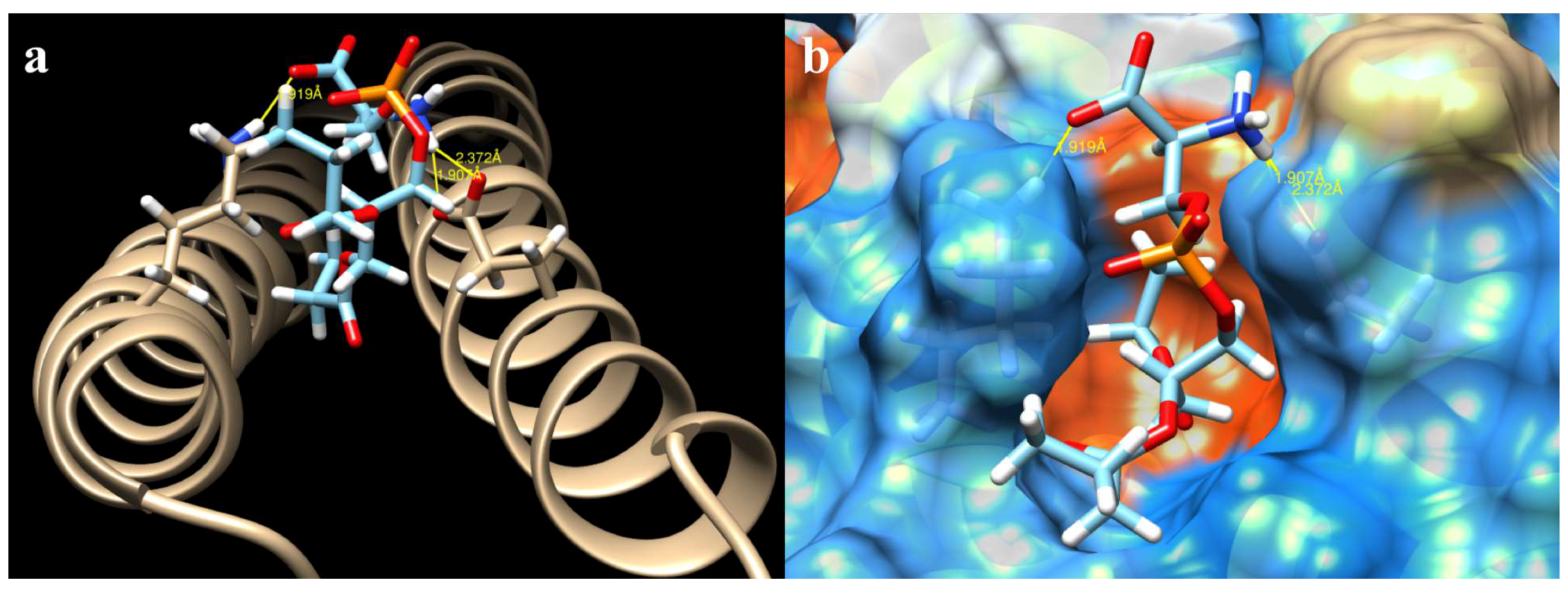
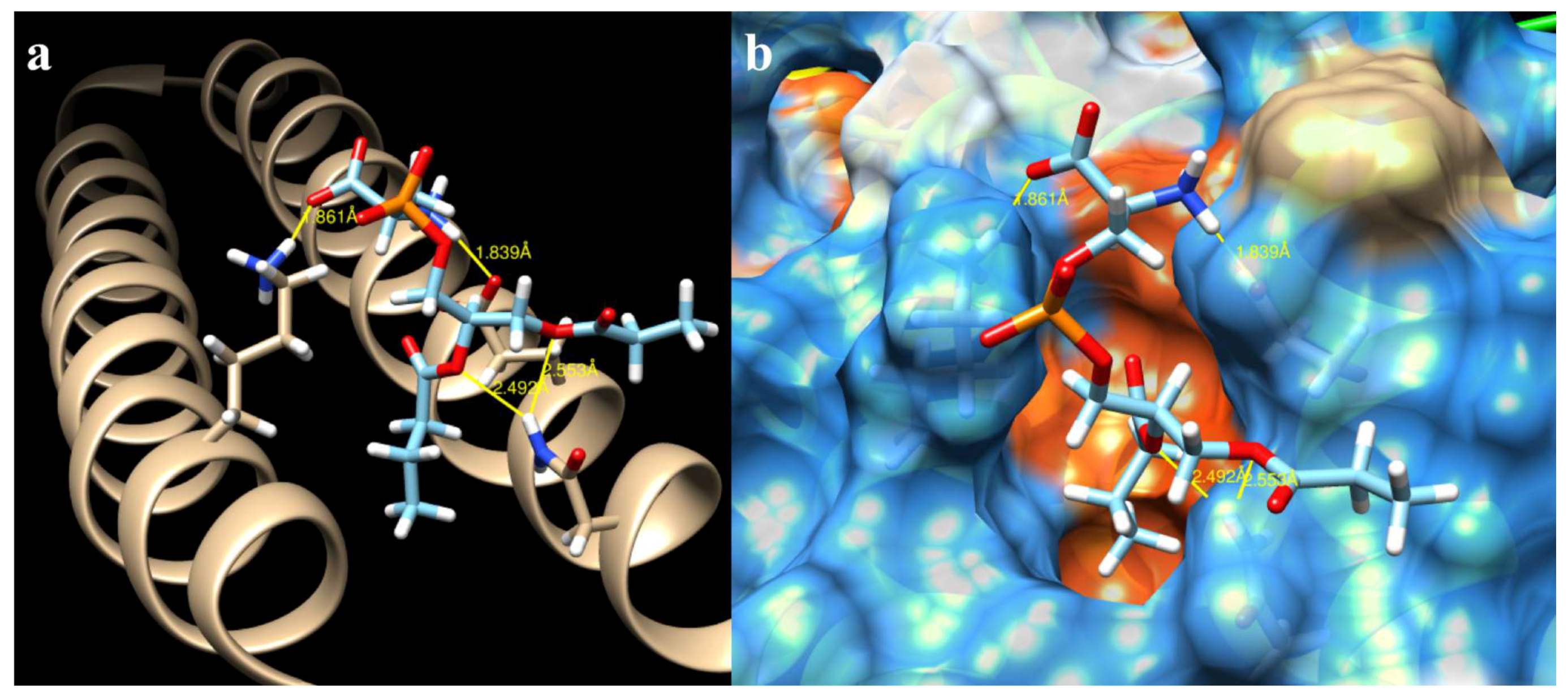
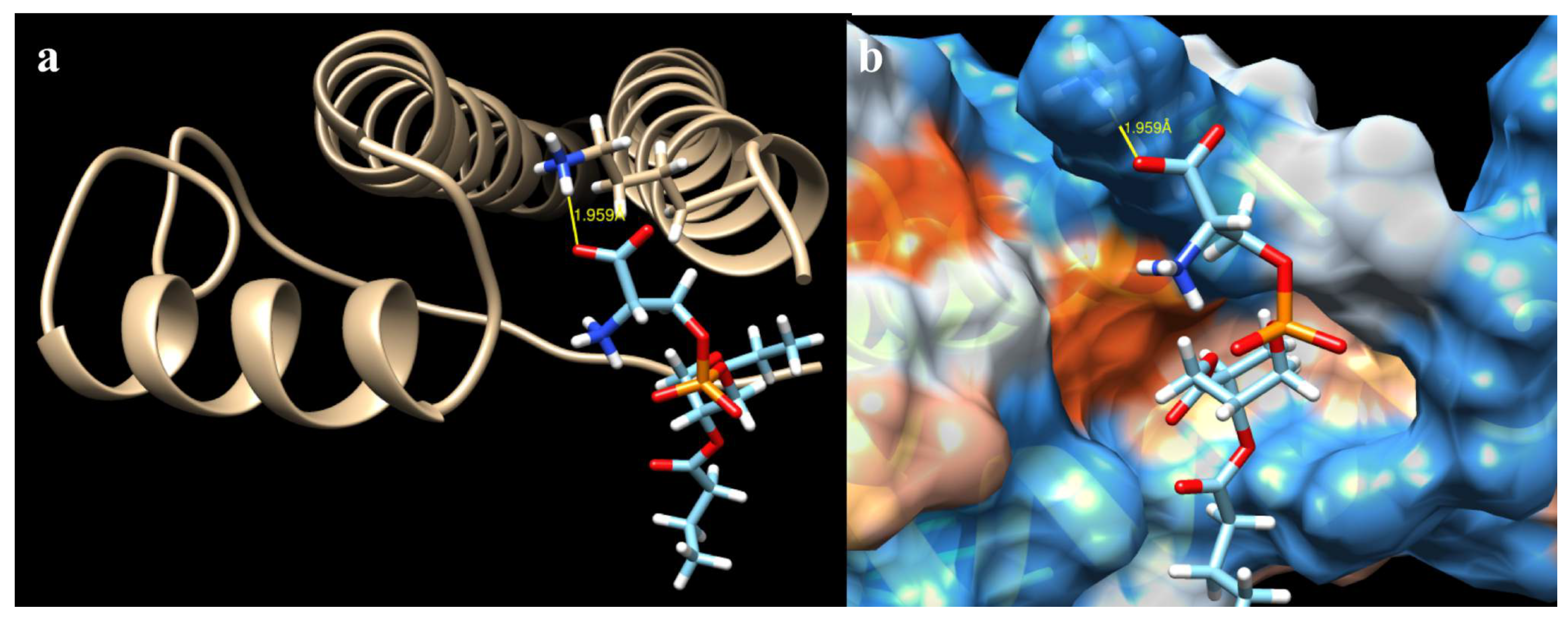
2.3. Docking of All Mutations
3. Discussion
4. Material and Methods
4.1. Database
4.2. Translation of Gene Bank DNA Sequences
4.3. Multiple Sequence Alignment (MSA) of Translated Sequences and Quality Assessment
4.4. Conservation Level and Variability of the Interaction Residues with PS
4.5. Determination of Interaction Forces between CagA N-Terminal and PS
4.6. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| CagA | Cytotoxin-associated gen A |
| Cag PAI | CagA pathogenic island |
| c-Abl | Mammalian Abelson murine leukemia viral oncogene |
| CHARMM | Chemistry at Harvard Macromolecular Mechanics |
| CM | Multimerization sequence |
| c-Met | Tyrosine-protein kinase Met |
| FACTS | Fast Analytical Continuum Treatment of Solvation |
| LMM | Linear Mixed Model |
| MALT | Mucosa-associated lymphoid tissue |
| MARK | Microtubule affinity-regulating kinase |
| MSA | Multiple Sequence Alignment |
| NCBI | The National Center for Biotechnology Information |
| NFB | Nuclear factor B |
| PAR1 | Protease-activated receptor 1 |
| PDB | Protein Databank |
| PS | Phosphatidylserine |
| Src | Proto-oncogene tyrosine-protein kinase |
| TCF | Transcription factor |
| T4SS | Type IV secretion system |
References
- Covacci, A.; Telford, J.L.; Del Giudice, G.; Parsonnet, J.; Rappuoli, R. Helicobacter pylori virulence and genetic geography. Science 1999, 284, 1328–1333. [Google Scholar] [CrossRef] [PubMed]
- Woon, A.P.; Tohidpour, A.; Alonso, H.; Saijo-Hamano, Y.; Kwok, T.; Roujeinikova, A. Conformational analysis of isolated domains of Helicobacter pylori CagA. PLoS ONE 2013, 8, e79367. [Google Scholar] [CrossRef] [PubMed]
- Wroblewski, L.E.; Peek, R.M.; Wilson, K.T. Helicobacter pylori and gastric cancer: Factors that modulate disease risk. Clin. Microbiol. Rev. 2010, 23, 713–739. [Google Scholar] [CrossRef] [PubMed]
- Tsang, Y.H.; Lamb, A.; Romero-Gallo, J.; Huang, B.; Ito, K.; Peek, R.M.J.; Ito, Y.; Chen, L.F. Helicobacter pylori CagA targets gastric tumor suppressor RUNX3 for proteasome-mediated degradation. Oncogene 2010, 29, 5643–5650. [Google Scholar] [CrossRef] [PubMed]
- Pelz, C.; Steininger, S.; Weiss, C.; Coscia, F.; Vogelmann, R. A novel inhibitory domain of Helicobacter pylori protein CagA reduces CagA effects on host cell biology. J. Biol. Chem. 2011, 286, 8999–9008. [Google Scholar] [CrossRef] [PubMed]
- Malfertheiner, P.; Sipponen, P.; Naumann, M.; Moayyedi, P.; Mégraud, F.; Xiao, S.D.; Sugano, K.; Nyrén, O. Helicobacter pylori eradication has the potential to prevent gastric cancer: A state-of-the-art critique. Am. J. Gastroenterol. 2005, 100, 2100–2115. [Google Scholar] [CrossRef] [PubMed]
- Vogiatzi, P.; Cassone, M.; Luzzi, I.; Lucchetti, C.; Otvos, L.; Giordano, A. Helicobacter pylori as a class I carcinogen: Physiopathology and management strategies. J. Cell. Biochem. 2007, 102, 264–273. [Google Scholar] [CrossRef] [PubMed]
- Baek, H.Y.; Lim, J.W.; Kim, H. Interaction between the Helicobacter pylori CagA and alpha-Pix in gastric epithelial AGS cells. Ann. N. Y. Acad. Sci. 2007, 1096, 18–23. [Google Scholar] [CrossRef] [PubMed]
- Churin, Y.; Al-Ghoul, L.; Kepp, O.; Meyer, T.F.; Birchmeier, W.; Naumann, M. Helicobacter pylori CagA protein targets the c-Met receptor and enhances the motogenic response. J. Cell Biol. 2003, 161, 249–255. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, M.; Mimuro, H.; Suzuki, T.; Park, M.; Yamamoto, T.; Sasakawa, C. Interaction of CagA with Crk plays an important role in Helicobacter pylori-induced loss of gastric epithelial cell adhesion. J. Exp. Med. 2005, 202, 1235–1247. [Google Scholar] [CrossRef] [PubMed]
- Tsutsumi, R.; Higashi, H.; Higuchi, M.; Okada, M.; Hatakeyama, M. Attenuation of Helicobacter pylori CagA·SHP-2 signaling by interaction between CagA and C-terminal Src kinase. J. Biol. Chem. 2003, 278, 3664–3670. [Google Scholar] [CrossRef] [PubMed]
- Kurashima, Y.; Murata-Kamiya, N.; Kikuchi, K.; Higashi, H.; Azuma, T.; Kondo, S.; Hatakeyama, M. Deregulation of β-catenin signal by Helicobacter pylori CagA requires the CagA-multimerization sequence. Int. J. Cancer 2008, 122, 823–831. [Google Scholar] [CrossRef] [PubMed]
- Krueger, S.; Hundertmark, T.; Kuester, D.; Kalinski, T.; Peitz, U.; Roessner, A. Helicobacter pylori alters the distribution of ZO-1 and p120ctn in primary human gastric epithelial cells. Pathol. Res. Pract. 2007, 203, 433–444. [Google Scholar] [CrossRef] [PubMed]
- Roujeinikova, A. Phospholipid binding residues of eukaryotic membrane-remodelling F-BAR domain proteins are conserved in Helicobacter pylori CagA. BMC Res. Notes 2014, 7, 525. [Google Scholar] [CrossRef] [PubMed]
- Nešić, D.; Buti, L.; Lu, X.; Stebbins, C.E. Structure of the Helicobacter pylori CagA oncoprotein bound to the human tumor suppressor ASPP2. Proc. Natl. Acad. Sci. USA 2014, 111, 1562–1567. [Google Scholar] [CrossRef] [PubMed]
- Kaplan-Turkoz, B.; Jimenez-Soto, L.F.; Dian, C.; Ertl, C.; Remaut, H.; Louche, A.; Tosi, T.; Haas, R.; Terradot, L. Structural insights into Helicobacter pylori oncoprotein CagA interaction with 1 integrin. Proc. Natl. Acad. Sci. USA 2012, 109, 14640–14645. [Google Scholar] [CrossRef] [PubMed]
- Murata-Kamiya, N.; Kikuchi, K.; Hayashi, T.; Higashi, H.; Hatakeyama, M. Helicobacter pylori exploits host membrane phosphatidylserine for delivery, localization, and pathophysiological action of the CagA oncoprotein. Cell Host Microbe 2010, 7, 399–411. [Google Scholar] [CrossRef] [PubMed]
- Sokolova, O.; Maubach, G.; Naumann, M. MEKK3 and TAK1 synergize to activate IKK complex in Helicobacter pylori infection. Biochim. Biophys. Acta 2014, 1843, 715–724. [Google Scholar] [CrossRef] [PubMed]
- Lamb, A.; Yang, X.D.; Tsang, Y.H.N.; Li, J.D.; Higashi, H.; Hatakeyama, M.; Peek, R.M.; Blanke, S.R.; Chen, L.F. Helicobacter pylori CagA activates NF-kappaB by targeting TAK1 for TRAF6-mediated Lys 63 ubiquitination. EMBO Rep. 2009, 10, 1242–1249. [Google Scholar] [CrossRef] [PubMed]
- Coombs, N.; Sompallae, R.; Olbermann, P.; Gastaldello, S.; Göppel, D.; Masucci, M.G.; Josenhans, C. Helicobacter pylori affects the cellular deubiquitinase USP7 and ubiquitin-regulated components TRAF6 and the tumour suppressor p53. Int. J. Med. Microbiol. 2011, 301, 213–224. [Google Scholar] [CrossRef] [PubMed]
- Backert, S.; Tegtmeyer, N.; Selbach, M. The versatility of helicobacter pylori caga effector protein functions: The master key hypothesis. Helicobacter 2010, 15, 163–176. [Google Scholar] [CrossRef] [PubMed]
- Bagnoli, F.; Buti, L.; Tompkins, L.; Covacci, A.; Amieva, M.R. Helicobacter pylori CagA induces a transition from polarized to invasive phenotypes in MDCK cells. Proc. Natl. Acad. Sci. USA 2005, 102, 16339–16344. [Google Scholar] [CrossRef] [PubMed]
- Amieva, M.R.; Vogelmann, R.; Covacci, A.; Tompkins, L.S.; Nelson, W.J.; Falkow, S. Disruption of the epithelial apical-junctional complex by Helicobacter pylori CagA. Science 2003, 300, 1430–1434. [Google Scholar] [CrossRef] [PubMed]
- Yamaoka, Y. Mechanisms of disease: Helicobacter pylori virulence factors. Nat. Rev. Gastroenterol. Hepatol. 2010, 7, 629–641. [Google Scholar] [CrossRef] [PubMed]
- Hayashi, T.; Senda, M.; Morohashi, H.; Higashi, H.; Horio, M.; Kashiba, Y.; Nagase, L.; Sasaya, D.; Shimizu, T.; Venugopalan, N.; et al. Tertiary Structure-Function Analysis Reveals the Pathogenic Signaling Potentiation Mechanism of Helicobacter pylori Oncogenic Effector CagA. Cell Host Microbe 2012, 12, 20–33. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, R.M.; Machado, J.C.; Leite, M.; Carneiro, F.; Figueiredo, C. The number of Helicobacter pylori CagA EPIYA C tyrosine phosphorylation motifs influences the pattern of gastritis and the development of gastric carcinoma. Histopathology 2012, 60, 992–998. [Google Scholar] [CrossRef] [PubMed]
- Beltrán-Anaya, F.O.; Poblete, T.M.T.M.; Román-Román, A.; Reyes, S.S.; de Sampedro, J.J.; Peralta-Zaragoza, O.; Rodríguez, M.Á.; del Moral-Hernández, O.; Illades-Aguiar, B.; Fernández-Tilapa, G. The EPIYA-ABCC motif pattern in CagA of Helicobacter pylori is associated with peptic ulcer and gastric cancer in Mexican population. BMC Gastroenterol. 2014, 14, 223. [Google Scholar] [CrossRef] [PubMed]
- Batista, S.A.; Rocha, G.A.; Rocha, A.M.C.; Saraiva, I.E.B.; Cabral, M.M.D.A.; Oliveira, R.C.; Queiroz, D.M.M. Higher number of Helicobacter pylori CagA EPIYA C phosphorylation sites increases the risk of gastric cancer, but not duodenal ulcer. BMC Microbiol. 2011, 11, 61. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, R.M.; Machado, J.C.; Figueiredo, C. Clinical relevance of Helicobacter pylori vacA and cagA genotypes in gastric carcinoma. Best Pract. Res. Clin. Gastroenterol. 2014, 28, 1003–1015. [Google Scholar] [CrossRef] [PubMed]
- Sicinschi, L.A.; Correa, P.; Peek, R.M.; Camargo, M.C.; Piazuelo, M.B.; Romero-Gallo, J.; Hobbs, S.S.; Krishna, U.; Delgado, A.; Mera, R.; et al. CagA C-terminal variations in Helicobacter pylori strains from Colombian patients with gastric precancerous lesions. Clin. Microbiol. Infect. 2010, 16, 369–378. [Google Scholar] [CrossRef] [PubMed]
- Breurec, S.; Michel, R.; Seck, A.; Brisse, S.; Côme, D.; Dieye, F.B.; Garin, B.; Huerre, M.; Mbengue, M.; Fall, C.; et al. Clinical relevance of cagA and vacA gene polymorphisms in Helicobacter pylori isolates from Senegalese patients. Clin. Microbiol. Infect. 2012, 18, 153–159. [Google Scholar] [CrossRef] [PubMed]
- Fajardo, C.A.; Quiroga, A.J.; Coronado, A.; Labrador, K.; Acosta, N.; Delgado, P.; Jaramillo, C.; Bravo, M.M. CagA EPIYA polymorphisms in Colombian Helicobacter pylori strains and their influence on disease-associated cellular responses. World J. Gastrointest. Oncol. 2013, 5, 50–59. [Google Scholar] [CrossRef] [PubMed]
- Püls, J.; Fischer, W.; Haas, R. Activation of Helicobacter pylori CagA by tyrosine phosphorylation is essential for dephosphorylation of host cell proteins in gastric epithelial cells. Mol. Microbiol. 2002, 43, 961–969. [Google Scholar] [CrossRef] [PubMed]
- Acosta, N.; Quiroga, A.A.; Delgado, P.; Bravo, M.M.M.M.; Jaramillo, C. Helicobacter pylori CagA protein polymorphisms and their lack of association with pathogenesis. World J. Gastroenterol. 2010, 16, 3936–3943. [Google Scholar] [CrossRef] [PubMed]
- Steininger, S.; Pelz, C.; Vogelmann, R. Purpose of recently detected inhibitory domain of the Helicobacter pylori protein CagA. Gut Microbes 2011, 2, 37–41. [Google Scholar] [CrossRef]
- Lemmon, M.A. Membrane recognition by phospholipid-binding domains. Nat. Rev. Mol. Cell Biol. 2008, 9, 99–111. [Google Scholar] [CrossRef] [PubMed]
- Grosdidier, A.; Zoete, V.; Michielin, O. SwissDock, a protein-small molecule docking web service based on EADock DSS. Nucleic Acids Res. 2011, 39, W270–W277. [Google Scholar] [CrossRef] [PubMed]
- Piana, S.; Klepeis, J.L.; Shaw, D.E. Assessing the accuracy of physical models used in protein-folding simulations: Quantitative evidence from long molecular dynamics simulations. Curr. Opin. Struct. Biol. 2014, 24, 98–105. [Google Scholar] [CrossRef] [PubMed]
- Berman, H.M.; Bhat, T.N.; Bourne, P.E.; Feng, Z.; Gilliland, G.; Weissig, H.; Westbrook, J. The Protein Data Bank and the challenge of structural genomics. Nat. Struct. Biol. 2000, 7, 957. [Google Scholar] [CrossRef] [PubMed]
- Grosdidier, A.; Zoete, V.; Michielin, O. Fast docking using the CHARMM force field with EADock DSS. J. Comput. Chem. 2011, 32, 2149–2159. [Google Scholar] [CrossRef] [PubMed]
- Brooks, B.R.; Brooks, C.L., III; Mackerell, A.D., Jr.; Nilsson, L.; Petrella, R.J.; Roux, B.; Won, Y.; Archontis, G.; Bartels, C.; Boresch, S.; et al. CHARMM: The Biomolecular Simulation Program B. J. Computat. Chem. 2009, 30, 1545–1614. [Google Scholar] [CrossRef] [PubMed]
- Haberthür, U.; Caflisch, A. FACTS: Fast analytical continuum treatment of solvation. J. Comput. Chem. 2008, 29, 701–715. [Google Scholar] [CrossRef]
- Landau, M.; Mayrose, I.; Rosenberg, Y.; Glaser, F.; Martz, E.; Pupko, T.; Ben-Tal, N. ConSurf 2005: The projection of evolutionary conservation scores of residues on protein structures. Nucleic Acids Res. 2005, 33, W299–W302. [Google Scholar] [CrossRef] [PubMed]
- Barnes, M.R.; Gray, I.C. Bioinformatics for Geneticists; John Wiley & Sons, Ltd.: Hoboken, NJ, USA, 2003; Volume 4, Chapter 14; pp. 289–314. [Google Scholar]
- Kay, J.G.; Grinstein, S. Lipid-Mediated Protein Signaling; Springer: Blacksburg, VA, USA, 2013; Chapter 10; pp. 117–193. [Google Scholar]
- Wilchek, M.; Bayer, E.A.; Livnah, O. Essentials of biorecognition: The (strept)avidin-biotin system as a model for protein-protein and protein-ligand interaction. Immunol. Lett. 2006, 103, 27–32. [Google Scholar] [CrossRef] [PubMed]
- Friesner, R.A.; Murphy, R.B.; Repasky, M.P.; Frye, L.L.; Greenwood, J.R.; Halgren, T.A.; Sanschagrin, P.C.; Mainz, D.T. Extra precision glide: Docking and scoring incorporating a model of hydrophobic enclosure for protein-ligand complexes. J. Med. Chem. 2006, 49, 6177–6196. [Google Scholar] [CrossRef] [PubMed]
- Zoete, V.; Michielin, O. EADock: Docking of Small Molecules Into Protein Active Sites with a Multiobjective Evolutionary Optimization. Proteins Struct. Funct. Bioinform. 2007, 1025, 1010–1025. [Google Scholar] [CrossRef]
- Matias, P.M.; Donner, P.; Coelho, R.; Thomaz, M.; Peixoto, C.; Macedo, S.; Otto, N.; Joschko, S.; Scholz, P.; Wegg, A.; et al. Structural evidence for ligand specificity in the binding domain of the human androgen receptor. Implications for pathogenic gene mutations. J. Biol. Chem. 2000, 275, 26164–26171. [Google Scholar] [CrossRef] [PubMed]
- Looger, L.L.; Dwyer, M.A.; Smith, J.J.; Hellinga, H.W. Computational design of receptor and sensor proteins with novel functions. Nature 2003, 423, 185–190. [Google Scholar] [CrossRef] [PubMed]
- Craveur, P.; Joseph, A.P.; Esque, J.; Narwani, T.J.; Noel, F.; Shinada, N.; Goguet, M.; Sylvain, L.; Poulain, P.; Bertrand, O.; et al. Protein flexibility in the light of structural alphabets. Front. Mol. Biosci. 2015, 2, 20. [Google Scholar] [CrossRef] [PubMed]
- Hatakeyama, M. Linking epithelial polarity and carcinogenesis by multitasking Helicobacter pylori virulence factor CagA. Oncogene 2008, 27, 7047–7054. [Google Scholar] [CrossRef] [PubMed]
- Rojas-Rengifo, D.F.; Alvarez-Silva, M.C.; Ulloa-Guerrero, C.P.; Nuñez-Velez, V.L.; del Pilar Delgado, M.; Aguilera, S.M.; Castro, H.; Jaramillo, C.A.; Fernando González Barrios, A. Intramolecular energies of the cytotoxic protein CagA of Helicobacter pylori as a possible descriptor of strains’ pathogenicity level. Comput. Biol. Chem. 2018, 76, 17–22. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef] [PubMed]
- Di Tommaso, P.; Moretti, S.; Xenarios, I.; Orobitg, M.; Montanyola, A.; Chang, J.M.; Taly, J.F.; Notredame, C. T-Coffee: A web server for the multiple sequence alignment of protein and RNA sequences using structural information and homology extension. Nucleic Acids Res. 2011, 39, W13–W17. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Standley, D.M. MAFFT Multiple Sequence Alignment Software Version 7: Improvements in Performance and Usability. Mol. Biol. Evolut. 2013, 30, 772–780. [Google Scholar] [CrossRef] [PubMed]
- Thompson, J.D.; Thierry, J.C.; Poch, O. RASCAL: Rapid scanning and correction of multiple sequence alignments. Bioinformatics 2003, 19, 1155–1161. [Google Scholar] [CrossRef] [PubMed]
- Thompson, J.D.; Plewniak, F.; Ripp, R.; Thierry, J.C.; Poch, O. Towards a reliable objective function for multiple sequence alignments11Edited by J. Karn. J. Mol. Biol. 2001, 314, 937–951. [Google Scholar] [CrossRef] [PubMed]
- Muller, J.; Creevey, C.J.; Thompson, J.D.; Arendt, D.; Bork, P. AQUA: Automated quality improvement for multiple sequence alignments. Bioinformatics 2010, 26, 263–265. [Google Scholar] [CrossRef] [PubMed]
- Biasini, M.; Bienert, S.; Waterhouse, A.; Arnold, K.; Studer, G.; Schmidt, T.; Kiefer, F.; Gallo Cassarino, T.; Bertoni, M.; Bordoli, L.; et al. SWISS-MODEL: Modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Res. 2014, 42, 252–258. [Google Scholar] [CrossRef] [PubMed]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Couch, G.S.; Greenblatt, D.M.; Meng, E.C.; Ferrin, T.E. UCSF Chimera—A visualization system for exploratory research and analysis. J. Comput. Chem. 2004, 25, 1605–1612. [Google Scholar] [CrossRef] [PubMed]
- Team, RDC. R: A Language and Environment for Statistical Computing; Team, RDC: Mangalore, India, 2012. [Google Scholar]


| Mutation | Model Number | Cluster | ΔG (Kcal/mol) | H-Bonds |
|---|---|---|---|---|
| Crystal | 1.81 | 10 | −8.919907 | 1 |
| K636A | 1.69 | 8 | −8.665261 | 0 |
| K636R | 1.74 | 8 | −8.701923 | 3 |
| K636N | 1.60 | 0 | −8.515097 | 4 |
| Sequence Number | Accession Number | Region * | Pathology |
|---|---|---|---|
| 39 | 22,335,784 | Eastern | Severe |
| 104 | 259,123,360 | Western | Severe |
| 106 | 259,123,364 | Eastern | Severe |
| 112 | 307,135,434 | Eastern | Severe |
| 115 | 307,135,440 | Eastern | Severe |
| 116 | 307,135,442 | Eastern | Severe |
| 117 | 307,135,444 | Eastern | Mild |
| 119 | 307,135,448 | Eastern | Severe |
| 121 | 307,135,452 | Eastern | Mild |
| 125 | 307,135,460 | Eastern | Severe |
| 127 | 307,135,464 | Eastern | Severe |
| 135 | 335,335,488 | Western | Severe |
| 150 | 345,421,953 | Eastern | Mild |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ulloa-Guerrero, C.P.; Delgado, M.D.P.; Jaramillo, C.A. Structural Analysis of Variability and Interaction of the N-terminal of the Oncogenic Effector CagA of Helicobacter pylori with Phosphatidylserine. Int. J. Mol. Sci. 2018, 19, 3273. https://doi.org/10.3390/ijms19103273
Ulloa-Guerrero CP, Delgado MDP, Jaramillo CA. Structural Analysis of Variability and Interaction of the N-terminal of the Oncogenic Effector CagA of Helicobacter pylori with Phosphatidylserine. International Journal of Molecular Sciences. 2018; 19(10):3273. https://doi.org/10.3390/ijms19103273
Chicago/Turabian StyleUlloa-Guerrero, Cindy P., Maria Del Pilar Delgado, and Carlos A. Jaramillo. 2018. "Structural Analysis of Variability and Interaction of the N-terminal of the Oncogenic Effector CagA of Helicobacter pylori with Phosphatidylserine" International Journal of Molecular Sciences 19, no. 10: 3273. https://doi.org/10.3390/ijms19103273
APA StyleUlloa-Guerrero, C. P., Delgado, M. D. P., & Jaramillo, C. A. (2018). Structural Analysis of Variability and Interaction of the N-terminal of the Oncogenic Effector CagA of Helicobacter pylori with Phosphatidylserine. International Journal of Molecular Sciences, 19(10), 3273. https://doi.org/10.3390/ijms19103273





