Identification of Proteases and Protease Inhibitors in Allergenic and Non-Allergenic Pollen
Abstract
:1. Introduction
2. Results and Discussion
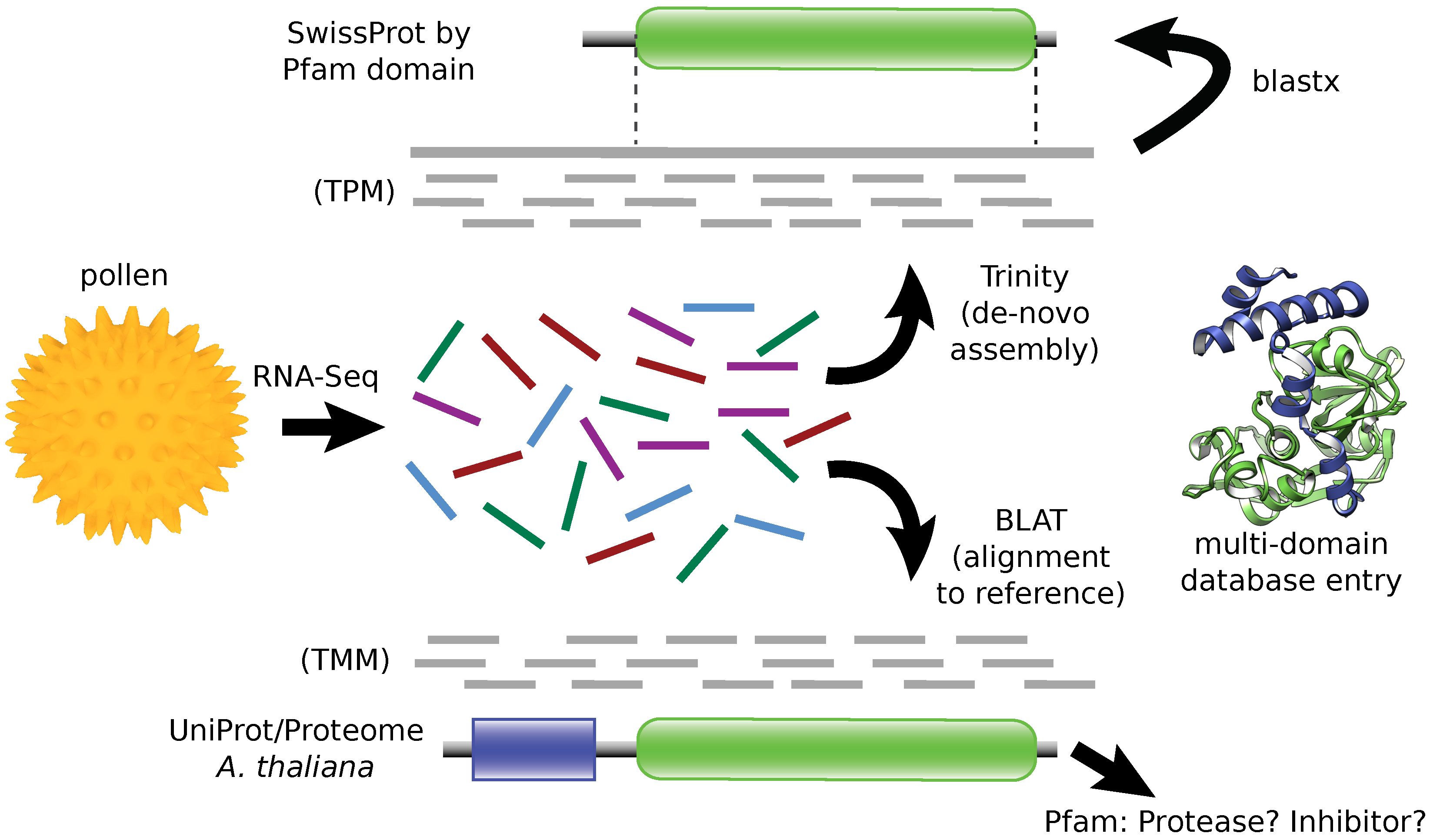
2.1. RNA Sequencing
2.2. Functional Profiling
2.3. Annotation of Proteases and Their Inhibitors
2.3.1. Proteases
2.3.2. Protease Inhibitors
2.4. Towards a Quantitative Comparison
2.5. Allergen Homologous Transcripts
3. Materials and Methods
3.1. RNA Sequencing
3.2. Annotation
3.3. Functional Profiling
3.4. Identification of Proteases and Protease Inhibitors
3.5. Quantitative Inter-Species Comparison
3.6. Identification of Allergen Homologues
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| ACC | Accession code |
| AllFam | Database of allergen families |
| ASDP | Allergen source derived protease |
| COG | Clusters of Orthologous Groups |
| GO | Gene Ontology |
| ID | Identifier |
| SP | SwissProt database entry |
| Pfam | Protein family database entry |
| TPM | Transcripts per million transcripts |
| TMM | Trimmed mean of M values |
References
- Bauchau, V.; Durham, S.R. Prevalence and rate of diagnosis of allergic rhinitis in Europe. Eur. Respir. J. 2004, 24, 758–764. [Google Scholar] [CrossRef] [PubMed]
- Traidl-Hoffmann, C.; Jakob, T.; Behrendt, H. Determinants of allergenicity. J. Allergy Clin. Immunol. 2009, 123, 558–566. [Google Scholar] [CrossRef] [PubMed]
- Davies, J.M. Grass pollen allergens globally: The contribution of subtropical grasses to burden of allergic respiratory diseases. Clin. Exp. Allergy 2014, 44, 790–801. [Google Scholar] [CrossRef] [PubMed]
- Asam, C.; Hofer, H.; Wolf, M.; Aglas, L.; Wallner, M. Tree pollen allergens— An update from a molecular perspective. Allergy 2015, 70, 1201–1211. [Google Scholar] [CrossRef] [PubMed]
- Pablos, I.; Wildner, S.; Asam, C.; Wallner, M.; Gadermaier, G. Pollen Allergens for Molecular Diagnosis. Curr. Allergy Asthma Rep. 2016, 16, 31. [Google Scholar] [CrossRef] [PubMed]
- European Forest Genetic Resources Programme. Available online: http://www.euforgen.org (accessed on 31 May 2017).
- D’Amato, G.; Cecchi, L.; Bonini, S.; Nunes, C.; Annesi-Maesano, I.; Behrendt, H.; Liccardi, G.; Popov, T.; van Cauwenberge, P. Allergenic pollen and pollen allergy in Europe. Allergy 2007, 62, 976–990. [Google Scholar] [CrossRef] [PubMed]
- PalDat – Palynological Database. Available online: https://www.paldat.org (accessed on 31 May 2017).
- Vrtala, S.; Grote, M.; Duchêne, M.; van Ree, R.; Kraft, D.; Scheiner, O.; Valenta, R. Properties of tree and grass pollen allergens: Reinvestigation of the linkage between solubility and allergenicity. Int. Arch. Allergy Immunol. 1993, 102, 160–169. [Google Scholar] [CrossRef] [PubMed]
- Obersteiner, A.; Gilles, S.; Frank, U.; Beck, I.; Häring, F.; Ernst, D.; Rothballer, M.; Hartmann, A.; Traidl-Hoffmann, C.; Schmid, M. Pollen-Associated Microbiome Correlates with Pollution Parameters and the Allergenicity of Pollen. PLoS ONE 2016, 11, e0149545. [Google Scholar] [CrossRef] [PubMed]
- Karle, A.C.; Oostingh, G.J.; Mutschlechner, S.; Ferreira, F.; Lackner, P.; Bohle, B.; Fischer, G.F.; Vogt, A.B.; Duschl, A. Nitration of the pollen allergen bet v 1.0101 enhances the presentation of bet v 1-derived peptides by HLA-DR on human dendritic cells. PLoS ONE 2012, 7, e31483. [Google Scholar] [CrossRef] [PubMed]
- McClain, S.; Bowman, C.; Fernández-Rivas, M.; Ladics, G.S.; van Ree, R. Allergic sensitization: Food- and protein-related factors. Clin. Transl. Allergy 2014, 4, 11. [Google Scholar] [CrossRef] [PubMed]
- Chua, K.Y.; Stewart, G.A.; Thomas, W.R.; Simpson, R.J.; Dilworth, R.J.; Plozza, T.M.; Turner, K.J. Sequence analysis of cDNA coding for a major house dust mite allergen, Der p 1. Homology with cysteine proteases. J. Exp. Med. 1988, 167, 175–182. [Google Scholar] [CrossRef] [PubMed]
- Groeme, R.; Airouche, S.; Kopečný, D.; Jaekel, J.; Savko, M.; Berjont, N.; Bussieres, L.; Le Mignon, M.; Jagic, F.; Zieglmayer, P.; et al. Structural and Functional Characterization of the Major Allergen Amb a 11 from Short Ragweed Pollen. J. Biol. Chem. 2016, 291, 13076–13087. [Google Scholar] [CrossRef] [PubMed]
- Dumez, M.E.; Herman, J.; Campizi, V.; Galleni, M.; Jacquet, A.; Chevigné, A. Orchestration of an uncommon maturation cascade of the house dust mite protease allergen quartet. Front. Immunol. 2014, 5, 138. [Google Scholar] [CrossRef] [PubMed]
- Winningham, K.M.; Fitch, C.D.; Schmidt, M.; Hoffman, D.R. Hymenoptera venom protease allergens. J. Allergy Clin. Immunol. 2004, 114, 928–933. [Google Scholar] [CrossRef] [PubMed]
- Herbert, C.A.; King, C.M.; Ring, P.C.; Holgate, S.T.; Stewart, G.A.; Thompson, P.J.; Robinson, C. Augmentation of permeability in the bronchial epithelium by the house dust mite allergen Der p1. Am. J. Respir. Cell Mol. Biol. 1995, 12, 369–378. [Google Scholar] [CrossRef] [PubMed]
- Wan, H.; Winton, H.L.; Soeller, C.; Gruenert, D.C.; Thompson, P.J.; Cannell, M.B.; Stewart, G.A.; Garrod, D.R.; Robinson, C. Quantitative structural and biochemical analyses of tight junction dynamics following exposure of epithelial cells to house dust mite allergen Der p 1. Clin. Exp. Allergy 2000, 30, 685–698. [Google Scholar] [CrossRef] [PubMed]
- Widmer, F.; Hayes, P.J.; Whittaker, R.G.; Kumar, R.K. Substrate preference profiles of proteases released by allergenic pollens. Clin. Exp. Allergy 2000, 30, 571–576. [Google Scholar] [CrossRef] [PubMed]
- Page, K.; Ledford, J.R.; Zhou, P.; Dienger, K.; Wills-Karp, M. Mucosal sensitization to German cockroach involves protease-activated receptor-2. Respir. Res. 2010, 11, 62. [Google Scholar] [CrossRef] [PubMed]
- Matsuwaki, Y.; Wada, K.; White, T.; Moriyama, H.; Kita, H. Alternaria fungus induces the production of GM-CSF, interleukin-6 and interleukin-8 and calcium signaling in human airway epithelium through protease-activated receptor 2. Int. Arch. Allergy Immunol. 2012, 158 (Suppl. 1), 19–29. [Google Scholar] [CrossRef] [PubMed]
- Florsheim, E.; Yu, S.; Bragatto, I.; Faustino, L.; Gomes, E.; Ramos, R.N.; Barbuto, J.A.M.; Medzhitov, R.; Russo, M. Integrated innate mechanisms involved in airway allergic inflammation to the serine protease subtilisin. J. Immunol. 2015, 194, 4621–4630. [Google Scholar] [CrossRef] [PubMed]
- Matsumura, Y. Role of Allergen Source-Derived Proteases in Sensitization via Airway Epithelial Cells. J. Allergy 2012, 2012, 903659. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, M.; Itoh, M.; Ohta, N.; Nakamura, Y.; Moriyama, A.; Matsumoto, T.; Ohashi, T.; Murakami, S. Blocking of protease allergens with inhibitors reduces allergic responses in allergic rhinitis and other allergic diseases. Acta Oto-laryngol. 2006, 126, 746–751. [Google Scholar] [CrossRef] [PubMed]
- Kato, T.; Takai, T.; Mitsuishi, K.; Okumura, K.; Ogawa, H. Cystatin A inhibits IL-8 production by keratinocytes stimulated with Der p 1 and Der f 1: Biochemical skin barrier against mite cysteine proteases. J. Allergy Clin. Immunol. 2005, 116, 169–176. [Google Scholar] [CrossRef] [PubMed]
- Runswick, S.; Mitchell, T.; Davies, P.; Robinson, C.; Garrod, D.R. Pollen proteolytic enzymes degrade tight junctions. Respirology 2007, 12, 834–842. [Google Scholar] [CrossRef] [PubMed]
- Rutley, N.; Twell, D. A decade of pollen transcriptomics. Plant Reprod. 2015, 28, 73–89. [Google Scholar] [CrossRef] [PubMed]
- Zhao, F.; Durner, J.; Winkler, J.B.; Traidl-Hoffmann, C.; Strom, T.M.; Ernst, D.; Frank, U. Pollen of common ragweed (Ambrosia artemisiifolia L.): Illumina-based de novo sequencing and differential transcript expression upon elevated NO2/O3. Environ. Pollut. 2017, 224, 503–514. [Google Scholar] [CrossRef] [PubMed]
- Lang, V.; Usadel, B.; Obermeyer, G. De novo sequencing and analysis of the lily pollen transcriptome: An open access data source for an orphan plant species. Plant Mol. Biol. 2015, 87, 69–80. [Google Scholar] [CrossRef] [PubMed]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; Zeng, Q.; et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 2011, 29, 644–652. [Google Scholar] [CrossRef] [PubMed]
- Finn, R.D.; Coggill, P.; Eberhardt, R.Y.; Eddy, S.R.; Mistry, J.; Mitchell, A.L.; Potter, S.C.; Punta, M.; Qureshi, M.; Sangrador-Vegas, A.; et al. The Pfam protein families database: Towards a more sustainable future. Nucleic Acids Res. 2016, 44, D279–D285. [Google Scholar] [CrossRef] [PubMed]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [PubMed]
- Bray, N.L.; Pimentel, H.; Melsted, P.; Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 2016, 34, 525–527. [Google Scholar] [CrossRef] [PubMed]
- Tatusov, R.L.; Fedorova, N.D.; Jackson, J.D.; Jacobs, A.R.; Kiryutin, B.; Koonin, E.V.; Krylov, D.M.; Mazumder, R.; Mekhedov, S.L.; Nikolskaya, A.N.; et al. The COG database: An updated version includes eukaryotes. BMC Bioinform. 2003, 4, 41. [Google Scholar] [CrossRef] [PubMed]
- Gene Ontology Consortium. Gene Ontology Consortium: Going forward. Nucleic Acids Res. 2015, 43, D1049–D1056. [Google Scholar]
- Rawlings, N.D.; Barrett, A.J.; Finn, R. Twenty years of the MEROPS database of proteolytic enzymes, their substrates and inhibitors. Nucleic Acids Res. 2016, 44, D343–D350. [Google Scholar] [CrossRef] [PubMed]
- Schaller, A.; Stintzi, A.; Graff, L. Subtilases - versatile tools for protein turnover, plant development, and interactions with the environment. Physiol. Plantarum 2012, 145, 52–66. [Google Scholar] [CrossRef] [PubMed]
- Radauer, C.; Bublin, M.; Wagner, S.; Mari, A.; Breiteneder, H. Allergens are distributed into few protein families and possess a restricted number of biochemical functions. J. Allergy Clin. Immunol. 2008, 121, 847–852. [Google Scholar] [CrossRef] [PubMed]
- Ibrahim, A.R.N.; Kawamoto, S.; Mizuno, K.; Shimada, Y.; Rikimaru, S.; Onishi, N.; Hashimoto, K.; Aki, T.; Hayashi, T.; Ono, K. Molecular cloning and immunochemical characterization of a new Japanese cedar pollen allergen homologous to plant subtilisin-like serine protease. World Allergy Organ. J. 2010, 3, 262–265. [Google Scholar] [CrossRef] [PubMed]
- Verma, S.; Dixit, R.; Pandey, K.C. Cysteine Proteases: Modes of Activation and Future Prospects as Pharmacological Targets. Front. Pharmacol. 2016, 7, 107. [Google Scholar] [CrossRef] [PubMed]
- Rawlings, N.D.; Tolle, D.P.; Barrett, A.J. Evolutionary families of peptidase inhibitors. Biochem. J. 2004, 378, 705–716. [Google Scholar] [CrossRef] [PubMed]
- Koshikawa, N.; Nakamura, T.; Tsuchiya, N.; Isaji, M.; Yasumitsu, H.; Umeda, M.; Miyazaki, K. Purification and identification of a novel and four known serine proteinase inhibitors secreted by human glioblastoma cells. J. Biochem. 1996, 119, 334–339. [Google Scholar] [CrossRef] [PubMed]
- Hedges, S.B.; Marin, J.; Suleski, M.; Paymer, M.; Kumar, S. Tree of life reveals clock-like speciation and diversification. Mol. Biol. Evol. 2015, 32, 835–845. [Google Scholar] [CrossRef] [PubMed]
- Kent, W.J. BLAT–the BLAST-like alignment tool. Genome Res. 2002, 12, 656–664. [Google Scholar] [CrossRef] [PubMed]
- Robinson, M.D.; Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 2010, 11, R25. [Google Scholar] [CrossRef] [PubMed]
- Wallner, M.; Erler, A.; Hauser, M.; Klinglmayr, E.; Gadermaier, G.; Vogel, L.; Mari, A.; Bohle, B.; Briza, P.; Ferreira, F. Immunologic characterization of isoforms of Car b 1 and Que a 1, the major hornbeam and oak pollen allergens. Allergy 2009, 64, 452–460. [Google Scholar] [CrossRef] [PubMed]
- FastQC: A Quality Control Tool for High Throughput Sequence Data. Available online: http://www.bioinformatics.babraham.ac.uk/projects/fastqc (accessed on 31 January 2017).
- Breese, M.R.; Liu, Y. NGSUtils: A software suite for analyzing and manipulating next-generation sequencing datasets. Bioinformatics 2013, 29, 494–496. [Google Scholar] [CrossRef] [PubMed]
- Cock, P.J.A.; Antao, T.; Chang, J.T.; Chapman, B.A.; Cox, C.J.; Dalke, A.; Friedberg, I.; Hamelryck, T.; Kauff, F.; Wilczynski, B.; et al. Biopython: Freely available Python tools for computational molecular biology and bioinformatics. Bioinformatics 2009, 25, 1422–1423. [Google Scholar] [CrossRef] [PubMed]
- Wickham, H. ggplot2: Elegant Graphics for Data Analysis; Springer: New York, NY, USA, 2009. [Google Scholar]
- Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2010, 26, 139–140. [Google Scholar] [CrossRef] [PubMed]
| Attribute | Betula | Pinus |
|---|---|---|
| Total paired reads | 223 M | 221 M |
| GC content (%) | 47 | 46 |
| Read length range | 35–151 | 35–151 |
| Read length median | 150 | 150 |
| Paired reads after filtering: | 218 M | 218 M |
| Assembled Trinity transcripts: | 59,581 | 85,151 |
| Assembled Trinity genes: | 25,263 | 38,064 |
| COG | GO-BP | GO-MF | GO-CC | |||||
|---|---|---|---|---|---|---|---|---|
| Bet | Pin | Bet | Pin | Bet | Pin | Bet | Pin | |
| Lil | 0.80 | 0.82 | 0.52 | 0.52 | 0.43 | 0.43 | 0.87 | 0.87 |
| Bet | 0.98 | 1.00 | 0.99 | 0.98 | ||||
| Pfam ID | Name | ∑ TPM Total | ∑ TPM Betula | ∑ TPM Pinus |
|---|---|---|---|---|
| PF00082 | Subtilase family | 28,858.7 | 21,782.0 | 7076.6 |
| PF00450 | Serine carboxypeptidase | 6259.0 | 4190.4 | 2068.6 |
| PF14543 | Xylanase inhibitor N-terminal | 4466.0 | 1185.8 | 3280.2 |
| PF00112 | Papain family cysteine protease | 4439.9 | 3076.1 | 1363.7 |
| PF00227 | Proteasome subunit | 1451.6 | 319.9 | 1131.7 |
| PF14541 | Xylanase inhibitor C-terminal | 1147.3 | 880.4 | 266.9 |
| PF12146 | Serine aminopeptidase, S33 | 871.5 | 427.1 | 444.4 |
| PF00443 | Ubiquitin carboxyl-terminal hydrolase | 699.6 | 266.3 | 433.3 |
| PF00026 | Eukaryotic aspartyl protease | 554.0 | 240.7 | 313.3 |
| PF00557 | Metallopeptidase family M24 | 547.5 | 183.9 | 363.6 |
| PF01694 | Rhomboid family | 518.0 | 328.8 | 189.2 |
| PF04258 | Signal peptide peptidase | 469.7 | 267.2 | 202.5 |
| PF01434 | Peptidase family M41 | 341.0 | 199.0 | 142.0 |
| PF02338 | OTU-like cysteine protease | 253.4 | 91.8 | 161.6 |
| PF05903 | PPPDE putative peptidase domain | 253.1 | 106.6 | 146.6 |
| PF01433 | Peptidase family M1 | 230.4 | 80.7 | 149.7 |
| PF00574 | Clp protease | 228.3 | 48.9 | 179.5 |
| PF02127 | Aminopeptidase I zinc metalloprotease (M18) | 179.6 | 137.3 | 42.4 |
| PF00717 | Peptidase S24-like | 174.8 | 66.9 | 107.9 |
| PF01088 | Ubiquitin carboxyl-terminal hydrolase, family 1 | 166.1 | 35.3 | 130.8 |
| PF00326 | Prolyl oligopeptidase family | 146.6 | 60.9 | 85.7 |
| PF01965 | DJ-1/PfpI family | 123.3 | 13.7 | 109.6 |
| PF05577 | Serine carboxypeptidase S28 | 106.6 | 21.7 | 84.8 |
| PF10502 | Signal peptidase, peptidase S26 | 103.5 | 66.5 | 37.0 |
| PF02902 | Ulp1 protease family, C-terminal catalytic domain | 99.7 | 20.9 | 78.8 |
| PF02099 | Josephin | 99.5 | 74.6 | 24.9 |
| PF04389 | Peptidase family M28 | 95.9 | 11.2 | 84.8 |
| PF01435 | Peptidase family M48 | 93.2 | 23.0 | 70.2 |
| PF13365 | Trypsin-like peptidase domain | 80.8 | 39.4 | 41.3 |
| PF10275 | Peptidase C65 Otubain | 79.0 | 14.6 | 64.5 |
| PF00883 | Cytosol aminopeptidase family, catalytic domain | 73.5 | 15.6 | 57.9 |
| PF00675 | Insulinase (Peptidase family M16) | 64.7 | 20.1 | 44.6 |
| PF00089 | Trypsin | 60.1 | 13.7 | 46.4 |
| PF01470 | Pyroglutamyl peptidase | 54.5 | 2.0 | 52.5 |
| PF01432 | Peptidase family M3 | 52.8 | 26.0 | 26.8 |
| PF08325 | WLM domain | 52.7 | 39.7 | 13.0 |
| PF03416 | Peptidase family C54 | 45.6 | 5.8 | 39.7 |
| PF09668 | Aspartyl protease | 39.6 | 1.8 | 37.9 |
| PF09768 | Peptidase M76 family | 36.6 | 28.8 | 7.7 |
| PF02163 | Peptidase family M50 | 31.9 | 4.9 | 27.0 |
| PF00246 | Zinc carboxypeptidase | 29.9 | 20.6 | 9.4 |
| PF05362 | Lon protease (S16) C-terminal proteolytic domain | 27.0 | 2.8 | 24.2 |
| PF14538 | Raptor N-terminal CASPase like domain | 26.2 | 16.9 | 9.3 |
| PF07910 | Peptidase family C78 | 18.8 | 3.1 | 15.7 |
| PF03572 | Peptidase family S41 | 12.2 | 5.0 | 7.2 |
| PF03571 | Peptidase family M49 | 11.6 | 5.3 | 6.3 |
| PF02586 | SOS response associated peptidase (SRAP) | 11.6 | 5.9 | 5.6 |
| PF00814 | Glycoprotease family | 10.3 | 3.7 | 6.5 |
| PF01457 | Leishmanolysin | 8.0 | 0.6 | 7.4 |
| PF00648 | Calpain family cysteine protease | 5.5 | 1.8 | 3.7 |
| PF00413 | Matrixin | 5.2 | 5.2 | 0.0 |
| PF01551 | Peptidase family M23 | 4.4 | 0.0 | 4.4 |
| PF06480 | FtsH Extracellular | 1.3 | 0.6 | 0.6 |
| Pfam ID | Name | ∑ TPM Total | ∑ TPM Betula | ∑ TPM Pinus |
|---|---|---|---|---|
| PF00188 | Cysteine-rich secretory protein family | 19,707.4 | 19,611.2 | 96.3 |
| PF05922 | Peptidase inhibitor I9 | 4382.9 | 2866.2 | 1516.7 |
| PF16845 | Aspartic acid proteinase inhibitor | 1328.2 | 1328.2 | 0.0 |
| PF00031 | Cystatin domain | 820.3 | 0.0 | 820.3 |
| PF00280 | Potato inhibitor I family | 154.1 | 121.2 | 32.9 |
| PF00079 | Serpin (serine protease inhibitor) | 84.4 | 31.7 | 52.8 |
| PF00197 | Trypsin and protease inhibitor | 6.5 | 6.5 | 0.0 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Höllbacher, B.; Schmitt, A.O.; Hofer, H.; Ferreira, F.; Lackner, P. Identification of Proteases and Protease Inhibitors in Allergenic and Non-Allergenic Pollen. Int. J. Mol. Sci. 2017, 18, 1199. https://doi.org/10.3390/ijms18061199
Höllbacher B, Schmitt AO, Hofer H, Ferreira F, Lackner P. Identification of Proteases and Protease Inhibitors in Allergenic and Non-Allergenic Pollen. International Journal of Molecular Sciences. 2017; 18(6):1199. https://doi.org/10.3390/ijms18061199
Chicago/Turabian StyleHöllbacher, Barbara, Armin O. Schmitt, Heidi Hofer, Fatima Ferreira, and Peter Lackner. 2017. "Identification of Proteases and Protease Inhibitors in Allergenic and Non-Allergenic Pollen" International Journal of Molecular Sciences 18, no. 6: 1199. https://doi.org/10.3390/ijms18061199
APA StyleHöllbacher, B., Schmitt, A. O., Hofer, H., Ferreira, F., & Lackner, P. (2017). Identification of Proteases and Protease Inhibitors in Allergenic and Non-Allergenic Pollen. International Journal of Molecular Sciences, 18(6), 1199. https://doi.org/10.3390/ijms18061199







