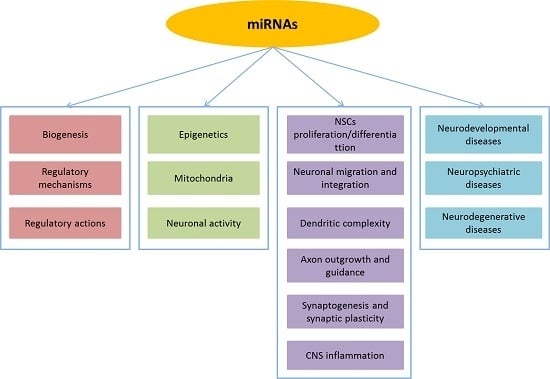
MicroRNAs: Key Regulators in the Central Nervous System and Their Implication in Neurological Diseases
Abstract
:1. Introduction
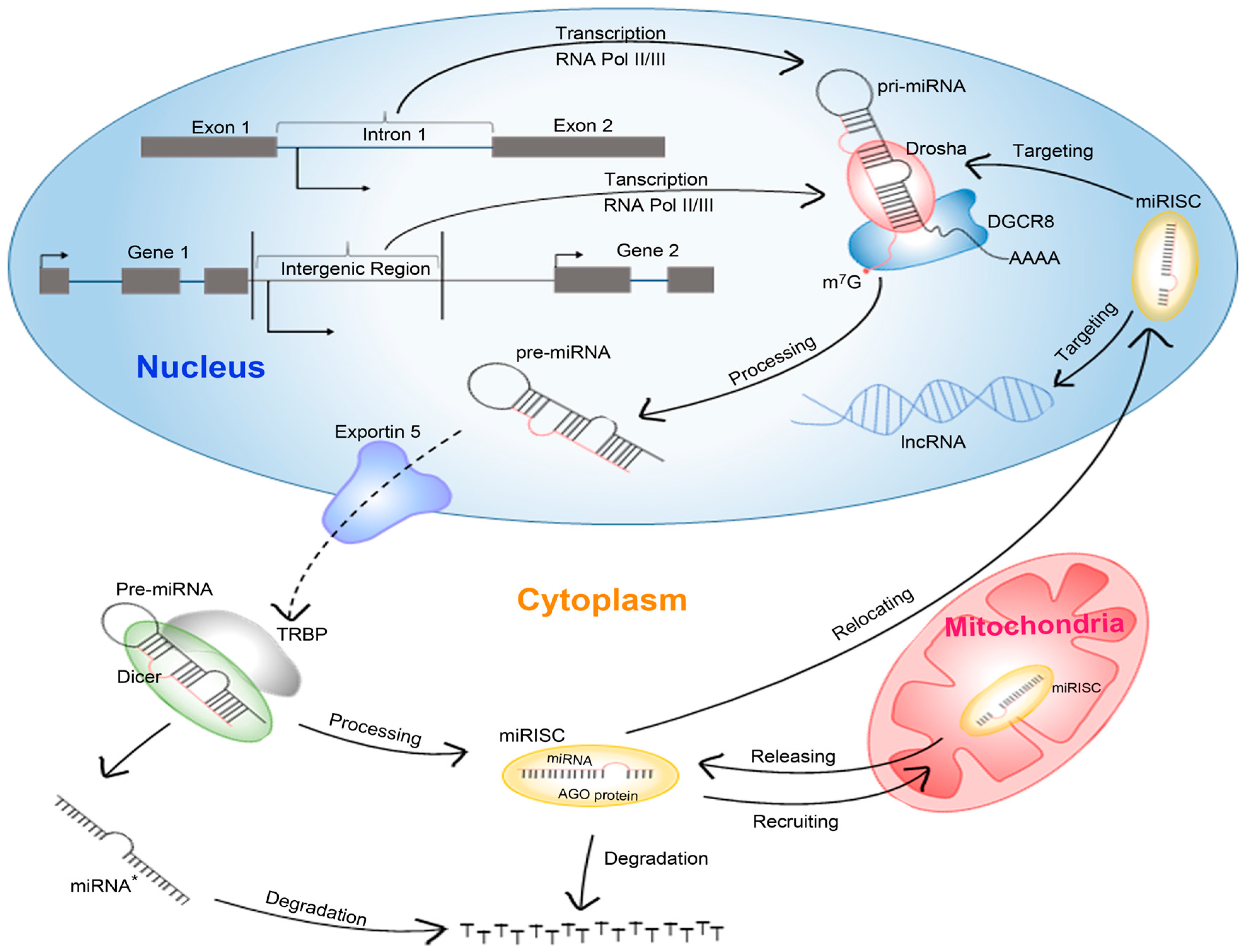
1.1. Biogenesis of miRNAs
1.2. Turnover of miRNAs
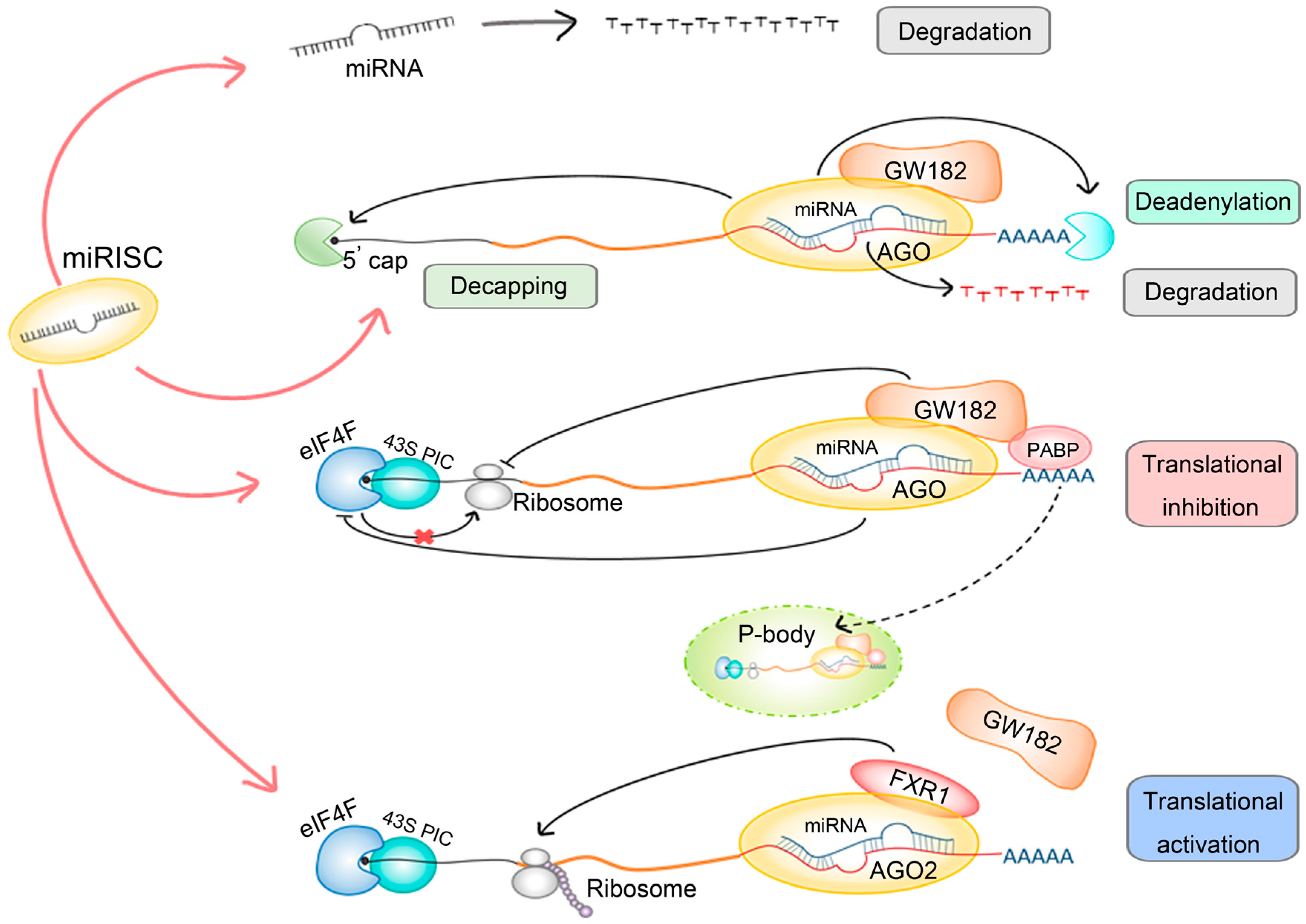
1.3. Mechanisms of miRNA Regulation
1.4. MicroRNA Regulation in Canonical and Non-Canonical Ways
2. MicroRNA Regulation and Function in the Central Nervous System
2.1. MicroRNA Regulation in the CNS
2.1.1. Crosstalk between miRNA and Epigenetics
2.1.2. MicroRNA Regulation by Mitochondria
2.1.3. MicroRNA Regulation by Neuronal Activity
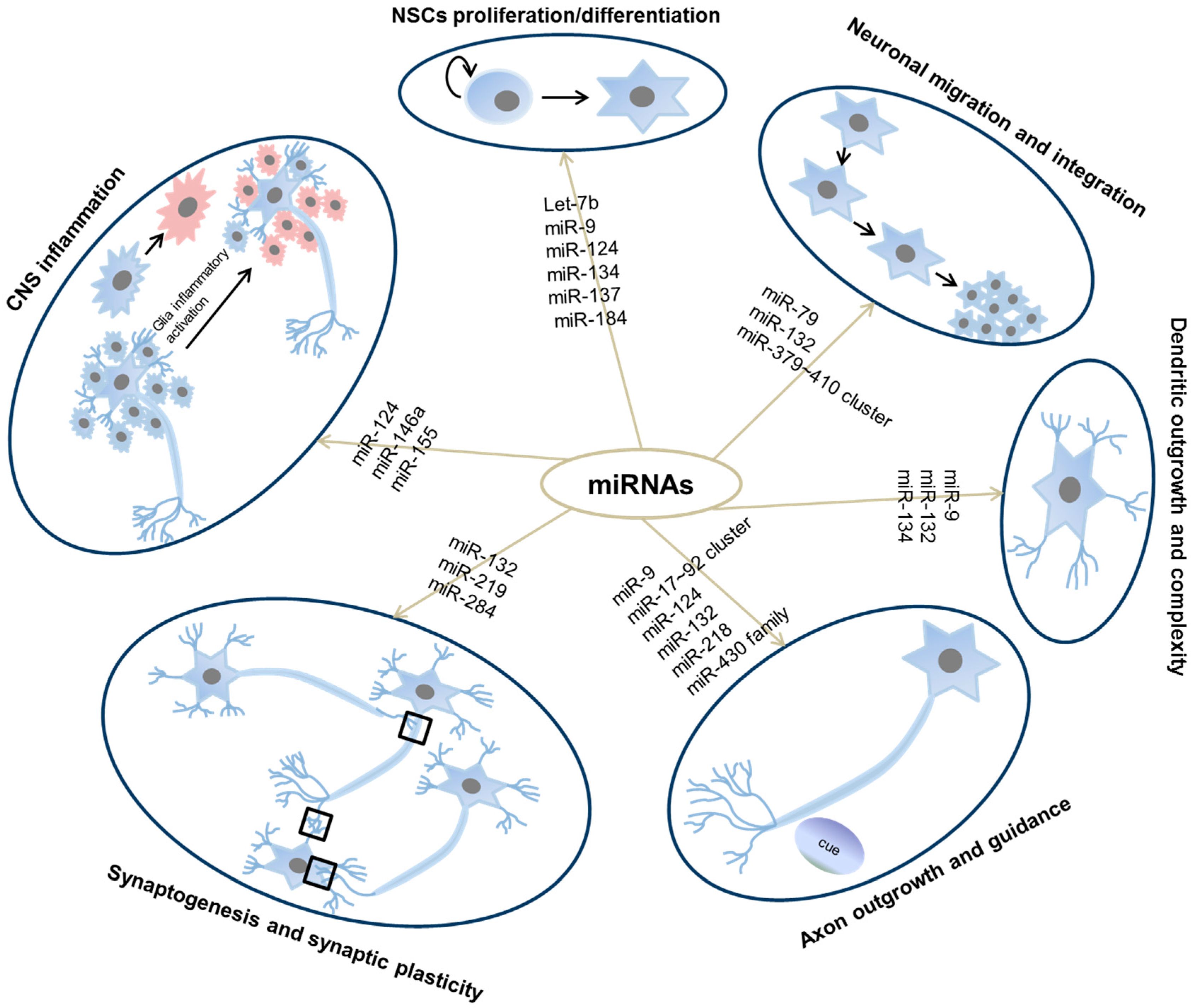
2.2. MicroRNA Function in the CNS
2.2.1. MicroRNA Function in Embryonic Neural Development
2.2.2. MicroRNA Function in NSCs Proliferation and Differentiation
2.2.3. MicroRNA Function in Neuronal Migration and Integration
2.2.4. MicroRNA Function in Dendritic Complexity
2.2.5. MicroRNA Function in Axon Outgrowth and Guidance
2.2.6. MicroRNA Function in Synaptogenesis and Synaptic Plasticity
2.2.7. MicroRNA Function in CNS Inflammation
3. Implications of miRNAs in Neurological Diseases
3.1. Neurodevelopmental Diseases
3.1.1. Fragile X Syndrome
3.1.2. Rett Syndrome
3.1.3. Autism Spectrum Disorders
3.2. Neuropsychiatric Diseases
3.2.1. Major Depression Disorder
3.2.2. Schizophrenia
3.3. Neurodegenerative Diseases
3.3.1. Alzheimer’s Disease
3.3.2. Parkinson’s Disease
4. Therapeutic Applications and Challenges
5. Conclusions and Outlook
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Ramalingam, P.; Palanichamy, J.K.; Singh, A.; Das, P.; Bhagat, M.; Kassab, M.A.; Sinha, S.; Chattopadhyay, P. Biogenesis of intronic miRNAs located in clusters by independent transcription and alternative splicing. RNA 2014, 20, 76–87. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.; Kim, M.; Han, J.; Yeom, K.H.; Lee, S.; Baek, S.H.; Kim, V.N. MicroRNA genes are transcribed by RNA polymerase II. EMBO J. 2004, 23, 4051–4060. [Google Scholar] [CrossRef] [PubMed]
- Borchert, G.M.; Lanier, W.; Davidson, B.L. RNA polymerase III transcribes human microRNAs. Nat. Struct. Mol. Biol. 2006, 13, 1097–1101. [Google Scholar] [CrossRef] [PubMed]
- Han, J.; Lee, Y.; Yeom, K.H.; Kim, Y.K.; Jin, H.; Kim, V.N. The drosha-DGCR8 complex in primary microRNA processing. Genes Dev. 2004, 18, 3016–3027. [Google Scholar] [CrossRef] [PubMed]
- Yi, R.; Qin, Y.; Macara, I.G.; Cullen, B.R. Exportin-5 mediates the nuclear export of pre-microRNAs and short hairpin rnas. Genes Dev. 2003, 17, 3011–3016. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.; Zhang, X.; Graves, P.; Zeng, Y. A comprehensive analysis of precursor microRNA cleavage by human dicer. RNA 2012, 18, 2083–2092. [Google Scholar] [CrossRef] [PubMed]
- Tsutsumi, A.; Kawamata, T.; Izumi, N.; Seitz, H.; Tomari, Y. Recognition of the pre-miRNA structure by drosophila dicer-1. Nat. Struct. Mol. Biol. 2011, 18, 1153–1158. [Google Scholar] [CrossRef] [PubMed]
- Ro, S.; Park, C.; Young, D.; Sanders, K.M.; Yan, W. Tissue-dependent paired expression of miRNAs. Nucleic Acids Res. 2007, 35, 5944–5953. [Google Scholar] [CrossRef] [PubMed]
- Chatterjee, S.; Fasler, M.; Bussing, I.; Grosshans, H. Target-mediated protection of endogenous microRNAs in C. Elegans. Dev. Cell 2011, 20, 388–396. [Google Scholar] [CrossRef] [PubMed]
- Ruegger, S.; Grosshans, H. MicroRNA turnover: When, how, and why. Trends Biochem. Sci. 2012, 37, 436–446. [Google Scholar] [CrossRef] [PubMed]
- Gantier, M.P.; McCoy, C.E.; Rusinova, I.; Saulep, D.; Wang, D.; Xu, D.; Irving, A.T.; Behlke, M.A.; Hertzog, P.J.; Mackay, F.; et al. Analysis of microRNA turnover in mammalian cells following dicer1 ablation. Nucleic Acids Res. 2011, 39, 5692–5703. [Google Scholar] [CrossRef] [PubMed]
- Krol, J.; Busskamp, V.; Markiewicz, I.; Stadler, M.B.; Ribi, S.; Richter, J.; Duebel, J.; Bicker, S.; Fehling, H.J.; Schubeler, D.; et al. Characterizing light-regulated retinal microRNAs reveals rapid turnover as a common property of neuronal microRNAs. Cell 2010, 141, 618–631. [Google Scholar] [CrossRef] [PubMed]
- Kocerha, J.; Faghihi, M.A.; Lopez-Toledano, M.A.; Huang, J.; Ramsey, A.J.; Caron, M.G.; Sales, N.; Willoughby, D.; Elmen, J.; Hansen, H.F.; et al. MicroRNA-219 modulates NMDA receptor-mediated neurobehavioral dysfunction. Proc. Natl. Acad. Sci. USA 2009, 106, 3507–3512. [Google Scholar] [CrossRef] [PubMed]
- Wibrand, K.; Panja, D.; Tiron, A.; Ofte, M.L.; Skaftnesmo, K.O.; Lee, C.S.; Pena, J.T.; Tuschl, T.; Bramham, C.R. Differential regulation of mature and precursor microRNA expression by NMDA and metabotropic glutamate receptor activation during LTP in the adult dentate gyrus In vivo. Eur. J. Neurosci. 2010, 31, 636–645. [Google Scholar] [CrossRef] [PubMed]
- Ivey, K.N.; Srivastava, D. MicroRNAs as developmental regulators. Cold Spring Harb. Perspect. Biol. 2015, 7, a008144. [Google Scholar] [CrossRef] [PubMed]
- Lewis, B.P.; Burge, C.B.; Bartel, D.P. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 2005, 120, 15–20. [Google Scholar] [CrossRef] [PubMed]
- Bushati, N.; Cohen, S.M. MicroRNA functions. Annu. Rev. Cell. Dev. Biol. 2007, 23, 175–205. [Google Scholar] [CrossRef] [PubMed]
- Eulalio, A.; Huntzinger, E.; Nishihara, T.; Rehwinkel, J.; Fauser, M.; Izaurralde, E. Deadenylation is a widespread effect of miRNA regulation. RNA 2009, 15, 21–32. [Google Scholar] [CrossRef] [PubMed]
- Valinezhad Orang, A.; Safaralizadeh, R.; Kazemzadeh-Bavili, M. Mechanisms of miRNA-mediated gene regulation from common downregulation to mRNA-specific upregulation. Int. J. Genom. 2014, 2014, 970607. [Google Scholar] [CrossRef] [PubMed]
- Mathonnet, G.; Fabian, M.R.; Svitkin, Y.V.; Parsyan, A.; Huck, L.; Murata, T.; Biffo, S.; Merrick, W.C.; Darzynkiewicz, E.; Pillai, R.S.; et al. MicroRNA inhibition of translation initiation in vitro by targeting the cap-binding complex eIF4F. Science 2007, 317, 1764–1767. [Google Scholar] [CrossRef] [PubMed]
- Huntzinger, E.; Kuzuoglu-Ozturk, D.; Braun, J.E.; Eulalio, A.; Wohlbold, L.; Izaurralde, E. The interactions of GW182 proteins with pabp and deadenylases are required for both translational repression and degradation of miRNA targets. Nucleic Acids Res. 2013, 41, 978–994. [Google Scholar] [CrossRef] [PubMed]
- Behm-Ansmant, I.; Rehwinkel, J.; Doerks, T.; Stark, A.; Bork, P.; Izaurralde, E. mRNA degradation by miRNAs and GW182 requires both CCR4:Not deadenylase and Dcp1:Dcp2 decapping complexes. Genes Dev. 2006, 20, 1885–1898. [Google Scholar] [CrossRef] [PubMed]
- Kuzuoglu-Ozturk, D.; Huntzinger, E.; Schmidt, S.; Izaurralde, E. The caenorhabditis elegans GW182 protein AIN-1 interacts with PAB-1 and subunits of the PAN2-PAN3 and CCR4-not deadenylase complexes. Nucleic Acids Res. 2012, 40, 5651–5665. [Google Scholar] [CrossRef] [PubMed]
- Fukaya, T.; Iwakawa, H.O.; Tomari, Y. MicroRNAs block assembly of eIF4F translation initiation complex in drosophila. Mol. Cell 2014, 56, 67–78. [Google Scholar] [CrossRef] [PubMed]
- Izaurralde, E. A role for eIF4AII in microRNA-mediated mRNA silencing. Nat. Struct. Mol. Biol. 2013, 20, 543–545. [Google Scholar] [CrossRef] [PubMed]
- Meijer, H.A.; Kong, Y.W.; Lu, W.T.; Wilczynska, A.; Spriggs, R.V.; Robinson, S.W.; Godfrey, J.D.; Willis, A.E.; Bushell, M. Translational repression and EIF4A2 activity are critical for microRNA-mediated gene regulation. Science 2013, 340, 82–85. [Google Scholar] [CrossRef] [PubMed]
- Fukao, A.; Mishima, Y.; Takizawa, N.; Oka, S.; Imataka, H.; Pelletier, J.; Sonenberg, N.; Thoma, C.; Fujiwara, T. MicroRNAs trigger dissociation of EIF4AI and EIF4AII from target mRNAs in humans. Mol. Cell 2014, 56, 79–89. [Google Scholar] [CrossRef] [PubMed]
- Eystathioy, T.; Jakymiw, A.; Chan, E.K.; Seraphin, B.; Cougot, N.; Fritzler, M.J. The GW182 protein colocalizes with mRNA degradation associated proteins hDcp1 and hLSm4 in cytoplasmic GW bodies. RNA 2003, 9, 1171–1173. [Google Scholar] [CrossRef] [PubMed]
- Andrei, M.A.; Ingelfinger, D.; Heintzmann, R.; Achsel, T.; Rivera-Pomar, R.; Luhrmann, R. A role for EIF4E and EIF4E-transporter in targeting mRNPs to mammalian processing bodies. RNA 2005, 11, 717–727. [Google Scholar] [CrossRef] [PubMed]
- Sen, G.L.; Blau, H.M. Argonaute 2/RISC resides in sites of mammalian mRNA decay known as cytoplasmic bodies. Nat. Cell Biol. 2005, 7, 633–636. [Google Scholar] [CrossRef] [PubMed]
- Vasudevan, S.; Tong, Y.; Steitz, J.A. Switching from repression to activation: MicroRNAs can up-regulate translation. Science 2007, 318, 1931–1934. [Google Scholar] [CrossRef] [PubMed]
- Truesdell, S.S.; Mortensen, R.D.; Seo, M.; Schroeder, J.C.; Lee, J.H.; LeTonqueze, O.; Vasudevan, S. MicroRNA-mediated mRNA translation activation in quiescent cells and oocytes involves recruitment of a nuclear microRNP. Sci. Rep. 2012, 2, 842. [Google Scholar] [CrossRef] [PubMed]
- Yoo, A.S.; Staahl, B.T.; Chen, L.; Crabtree, G.R. MicroRNA-mediated switching of chromatin-remodelling complexes in neural development. Nature 2009, 460, 642–646. [Google Scholar] [PubMed]
- Lee, I.; Ajay, S.S.; Yook, J.I.; Kim, H.S.; Hong, S.H.; Kim, N.H.; Dhanasekaran, S.M.; Chinnaiyan, A.M.; Athey, B.D. New class of microRNA targets containing simultaneous 5′-UTR and 3′-UTR interaction sites. Genome Res. 2009, 19, 1175–1183. [Google Scholar] [CrossRef] [PubMed]
- Place, R.F.; Li, L.C.; Pookot, D.; Noonan, E.J.; Dahiya, R. MicroRNA-373 induces expression of genes with complementary promoter sequences. Proc. Natl. Acad. Sci. USA 2008, 105, 1608–1613. [Google Scholar] [CrossRef] [PubMed]
- Qin, W.M.; Shi, Y.; Zhao, B.T.; Yao, C.G.; Jin, L.; Ma, J.X.; Jin, Y.X. miR-24 regulates apoptosis by targeting the open reading frame (ORF) region of FAF1 in cancer cells. PLoS ONE 2010, 5, e9429. [Google Scholar] [CrossRef] [PubMed]
- Loeb, G.B.; Khan, A.A.; Canner, D.; Hiatt, J.B.; Shendure, J.; Darnell, R.B.; Leslie, C.S.; Rudensky, A.Y. Transcriptome-wide miR-155 binding map reveals widespread noncanonical microRNA targeting. Mol. Cell 2012, 48, 760–770. [Google Scholar] [CrossRef] [PubMed]
- Dweep, H.; Gretz, N. MiRwalk2.0: A comprehensive atlas of microRNA-target interactions. Nat. Methods 2015, 12, 697. [Google Scholar] [CrossRef] [PubMed]
- Reczko, M.; Maragkakis, M.; Alexiou, P.; Grosse, I.; Hatzigeorgiou, A.G. Functional microRNA targets in protein coding sequences. Bioinformatics 2012, 28, 771–776. [Google Scholar] [CrossRef] [PubMed]
- Politz, J.C.R.; Hogan, E.M.; Pederson, T. MicroRNAs with a nucleolar location. RNA 2009, 15, 1705–1715. [Google Scholar] [CrossRef] [PubMed]
- Hwang, H.W.; Wentzel, E.A.; Mendell, J.T. A hexanucleotide element directs microRNA nuclear import. Science 2007, 315, 97–100. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.H.; Saetrom, P.; Snove, O.; Rossi, J.J. MicroRNA-directed transcriptional gene silencing in mammalian cells. Proc. Natl. Acad. Sci. USA 2008, 105, 16230–16235. [Google Scholar] [CrossRef] [PubMed]
- Foldes-Papp, Z.; Konig, K.; Studier, H.; Buckle, R.; Breunig, H.G.; Uchugonova, A.; Kostner, G.M. Trafficking of mature miRNA-122 into the nucleus of live liver cells. Curr. Pharm. Biotechnol. 2009, 10, 569–578. [Google Scholar] [CrossRef] [PubMed]
- Jeffries, C.D.; Fried, H.M.; Perkins, D.O. Nuclear and cytoplasmic localization of neural stem cell microRNAs. RNA 2011, 17, 675–686. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Liang, H.W.; Zhang, C.Y.; Zen, K. miRNA regulates noncoding RNA: A noncanonical function model. Trends Biochem. Sci. 2012, 37, 457–459. [Google Scholar] [CrossRef] [PubMed]
- Tang, R.; Li, L.M.; Zhu, D.H.; Hou, D.X.; Cao, T.; Gu, H.W.; Zhang, J.; Chen, J.Y.; Zhang, C.Y.; Zen, K. Mouse miRNA-709 directly regulates miRNA-15a/16-1 biogenesis at the posttranscriptional level in the nucleus: Evidence for a microRNA hierarchy system. Cell Res. 2012, 22, 504–515. [Google Scholar] [CrossRef] [PubMed]
- Hansen, T.B.; Wiklund, E.D.; Bramsen, J.B.; Villadsen, S.B.; Statham, A.L.; Clark, S.J.; Kjems, J. miRNA-dependent gene silencing involving AGO2-mediated cleavage of a circular antisense RNA. EMBO J. 2011, 30, 4414–4422. [Google Scholar] [CrossRef] [PubMed]
- Gotz, M.; Huttner, W.B. The cell biology of neurogenesis. Nat. Rev. Mol. Cell Biol. 2005, 6, 777–788. [Google Scholar] [CrossRef] [PubMed]
- Azevedo, F.A.C.; Carvalho, L.R.B.; Grinberg, L.T.; Farfel, J.M.; Ferretti, R.E.L.; Leite, R.E.P.; Jacob, W.; Lent, R.; Herculano-Houzel, S. Equal numbers of neuronal and nonneuronal cells make the human brain an isometrically scaled-up primate brain. J. Comp. Neurol. 2009, 513, 532–541. [Google Scholar] [CrossRef] [PubMed]
- Narayan, A.; Bommakanti, A.; Patel, A.A. High-throughput RNA profiling via up-front sample parallelization. Nat. Methods 2015, 12, 343–399. [Google Scholar] [CrossRef] [PubMed]
- Zaits, M.N.; Rennert, O.M. Identification of differentially expressed microRNAs across the developing human brain. Mol. Psychiatry 2014, 19, 848–852. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.; Qin, C. General hallmarks of microRNAs in brain evolution and development. RNA Biol. 2015, 12, 701–708. [Google Scholar] [CrossRef] [PubMed]
- Davis, G.M.; Haas, M.A.; Pocock, R. MicroRNAs: Not “fine-tuners” but key regulators of neuronal development and function. Front. Neurol. 2015, 6, 245. [Google Scholar] [CrossRef] [PubMed]
- Fagiolini, M.; Jensen, C.L.; Champagne, F.A. Epigenetic influences on brain development and plasticity. Curr. Opin. Neurobiol. 2009, 19, 207–212. [Google Scholar] [CrossRef] [PubMed]
- Keverne, E.B. Significance of epigenetics for understanding brain development, brain evolution and behavior. Neuroscience 2014, 264, 207–217. [Google Scholar] [CrossRef] [PubMed]
- Han, L.; Witmer, P.D.; Casey, E.; Valle, D.; Sukumar, S. DNA methylation regulates microRNA expression. Cancer Biol. Ther. 2007, 6, 1284–1288. [Google Scholar] [CrossRef] [PubMed]
- Scott, G.K.; Mattie, N.D.; Berger, C.E.; Benz, S.C.; Benz, C.C. Rapid alteration of microRNA levels by histone deacetylase inhibition. Cancer Res. 2006, 66, 1277–1281. [Google Scholar] [CrossRef] [PubMed]
- Saito, Y.; Liang, G.; Egger, G.; Friedman, J.M.; Chuang, J.C.; Coetzee, G.A.; Jones, P.A. Specific activation of microRNA-127 with downregulation of the proto-oncogene BCL6 by chromatin-modifying drugs in human cancer cells. Cancer Cell 2006, 9, 435–443. [Google Scholar] [CrossRef] [PubMed]
- Fabbri, M.; Garzon, R.; Cimmino, A.; Liu, Z.; Zanesi, N.; Callegari, E.; Liu, S.; Alder, H.; Costinean, S.; Fernandez-Cymering, C.; et al. MicroRNA-29 family reverts aberrant methylation in lung cancer by targeting DNA methyltransferases 3a and 3b. Proc. Natl. Acad. Sci. USA 2007, 104, 15805–15810. [Google Scholar] [CrossRef] [PubMed]
- Chen, B.F.; Gu, S.; Suen, Y.K.; Li, L.; Chan, W.Y. MicroRNA-199a-3p, DNMT3A, and aberrant DNA methylation in testicular cancer. Epigenetics 2014, 9, 119–128. [Google Scholar] [CrossRef] [PubMed]
- Duursma, A.M.; Kedde, M.; Schrier, M.; Le Sage, C.; Agami, R. miR-148 targets human DNMT3B protein coding region. RNA 2008, 14, 872–877. [Google Scholar] [CrossRef] [PubMed]
- Szulwach, K.E.; Li, X.K.; Smrt, R.D.; Li, Y.J.; Luo, Y.P.; Lin, L.; Santistevan, N.J.; Li, W.D.; Zhao, X.Y.; Jin, P. Cross talk between microRNA and epigenetic regulation in adult neurogenesis. J. Cell Biol. 2010, 189, U127–U181. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Chen, B.F.; Chan, W.Y. An epigenetic regulator: Methyl-CpG-binding domain protein 1 (MBD1). Int. J. Mol. Sci. 2015, 16, 5125–5140. [Google Scholar] [CrossRef] [PubMed]
- Guy, J.; Cheval, H.; Selfridge, J.; Bird, A. The role of MeCP2 in the brain. Annu Rev. Cell. Dev. Biol. 2011, 27, 631–652. [Google Scholar] [CrossRef] [PubMed]
- Amir, R.E.; Van den Veyver, I.B.; Wan, M.; Tran, C.Q.; Francke, U.; Zoghbi, H.Y. Rett syndrome is caused by mutations in x-linked MeCP2, encoding methyl-CPG-binding protein 2. Nat. Genet. 1999, 23, 185–188. [Google Scholar] [PubMed]
- Liu, Z.; Li, X.; Zhang, J.T.; Cai, Y.J.; Cheng, T.L.; Cheng, C.; Wang, Y.; Zhang, C.C.; Nie, Y.H.; Chen, Z.F.; et al. Autism-like behaviors and germline transmission in transgenic monkeys overexpressing MeCP2. Nature 2016, 530, 98–102. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.M.; Teng, Z.Q.; Santistevan, N.J.; Szulwach, K.E.; Guo, W.X.; Jin, P.; Zhao, X.Y. Epigenetic regulation of miR-184 by MBD1 governs neural stem cell proliferation and differentiation. Cell Stem Cell 2010, 6, 433–444. [Google Scholar] [CrossRef] [PubMed]
- Sullivan, P.G.; Rabchevsky, A.G.; Waldmeier, P.C.; Springer, J.E. Mitochondrial permeability transition in cns trauma: Cause or effect of neuronal cell death? J. Neurosci. Res. 2005, 79, 231–239. [Google Scholar] [CrossRef] [PubMed]
- Bian, Z.; Li, L.M.; Tang, R.; Hou, D.X.; Chen, X.; Zhang, C.Y.; Zen, K. Identification of mouse liver mitochondria-associated miRNAs and their potential biological functions. Cell Res. 2010, 20, 1076–1078. [Google Scholar] [CrossRef] [PubMed]
- Das, S.; Ferlito, M.; Kent, O.A.; Fox-Talbot, K.; Wang, R.; Liu, D.L.; Raghavachari, N.; Yang, Y.Q.; Wheelan, S.J.; Murphy, E.; et al. Nuclear miRNA regulates the mitochondrial genome in the heart. Circ. Res. 2012, 110, 1596–1603. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.R.; Zuo, X.X.; Yang, B.; Li, Z.R.; Xue, Y.C.; Zhou, Y.; Huang, J.; Zhao, X.L.; Zhou, J.; Yan, Y.; et al. MicroRNA directly enhances mitochondrial translation during muscle differentiation. Cell 2014, 158, 607–619. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.X.; Visavadiya, N.P.; Pandya, J.D.; Nelson, P.T.; Sullivan, P.G.; Springer, J.E. Mitochondria-associated microRNAs in rat hippocampus following traumatic brain injury. Exp. Neurol 2015, 265, 84–93. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.X.; Springer, J.E. Role of mitochondria in regulating microRNA activity and its relevance to the central nervous system. Neural Regen Res. 2015, 10, 1026–1028. [Google Scholar] [CrossRef] [PubMed]
- Jung, H.S.; Yoon, B.C.; Holt, C.E. Axonal mRNA localization and local protein synthesis in nervous system assembly, maintenance and repair. Nat. Rev. Neurosci. 2012, 13, 308–324. [Google Scholar] [CrossRef] [PubMed]
- Streck, E.L.; Goncalves, C.L.; Furlanetto, C.B.; Scaini, G.; Dal-Pizzol, F.; Quevedo, J. Mitochondria and the central nervous system: Searching for a pathophysiological basis of psychiatric disorders. Rev. Bras. Psiquiatr. 2014, 36, 156–167. [Google Scholar] [CrossRef] [PubMed]
- Schratt, G.M.; Tuebing, F.; Nigh, E.A.; Kane, C.G.; Sabatini, M.E.; Kiebler, M.; Greenberg, M.E. A brain-specific microRNA regulates dendritic spine development. Nature 2006, 439, 283–289. [Google Scholar] [CrossRef] [PubMed]
- Kye, M.J.; Liu, T.L.; Levy, S.F.; Xu, N.L.; Groves, B.B.; Bonneau, R.; Lao, K.Q.; Kosik, K.S. Somatodendritic microRNAs identified by laser capture and multiplex RT-PCR. RNA 2007, 13, 1224–1234. [Google Scholar] [CrossRef] [PubMed]
- Lugli, G.; Torvik, V.I.; Larson, J.; Smalheiser, N.R. Expression of microRNAs and their precursors in synaptic fractions of adult mouse forebrain. J. Neurochem. 2008, 106, 650–661. [Google Scholar] [CrossRef] [PubMed]
- Siegel, G.; Obernosterer, G.; Fiore, R.; Oehmen, M.; Bicker, S.; Christensen, M.; Khudayberdiev, S.; Leuschner, P.F.; Busch, C.J.L.; Kane, C.; et al. A functional screen implicates microRNA-138-dependent regulation of the depalmitoylation enzyme APT1 in dendritic spine morphogenesis. Nat. Cell Biol. 2009, 11, 705–736. [Google Scholar] [CrossRef] [PubMed]
- Park, C.S.; Tang, S.J. Regulation of microRNA expression by induction of bidirectional synaptic plasticity. J. Mol. Neurosci. 2009, 38, 50–56. [Google Scholar] [CrossRef] [PubMed]
- Kye, M.J.; Neveu, P.; Lee, Y.S.; Zhou, M.; Steen, J.A.; Sahin, M.; Kosik, K.S.; Silva, A.J. NMDA mediated contextual conditioning changes miRNA expression. PLoS ONE 2011, 6. [Google Scholar] [CrossRef] [PubMed]
- Van Spronsen, M.; van Battum, E.Y.; Kuijpers, M.; Vangoor, V.R.; Rietman, M.L.; Pothof, J.; Gumy, L.F.; van IJcken, W.F.J.; Akhmanova, A.; Pasterkamp, R.J.; et al. Developmental and activity-dependent miRNA expression profiling in primary hippocampal neuron cultures. PLoS ONE 2013, 8, e74907. [Google Scholar] [CrossRef] [PubMed]
- Eacker, S.M.; Keuss, M.J.; Berezikov, E.; Dawson, V.L.; Dawson, T.M. Neuronal activity regulates hippocampal miRNA expression. PLoS ONE 2011, 6, e25068. [Google Scholar] [CrossRef] [PubMed]
- Klein, M.E.; Lioy, D.T.; Ma, L.; Impey, S.; Mandel, G.; Goodman, R.H. Homeostatic regulation of MeCP2 expression by a creb-induced microRNA. Nat. Neurosci. 2007, 10, 1513–1514. [Google Scholar] [CrossRef] [PubMed]
- Nudelman, A.S.; DiRocco, D.P.; Lambert, T.J.; Garelick, M.G.; Le, J.; Nathanson, N.M.; Storm, D.R. Neuronal activity rapidly induces transcription of the creb-regulated microRNA-132, in vivo. Hippocampus 2010, 20, 492–498. [Google Scholar] [CrossRef] [PubMed]
- Cheng, H.Y.M.; Papp, J.W.; Varlamova, O.; Dziema, H.; Russell, B.; Curfman, J.P.; Nakazawa, T.; Shimizu, K.; Okamura, H.; Impey, S.; et al. MicroRNA modulation of circadian-clock period and entrainment. Neuron 2007, 54, 813–829. [Google Scholar] [CrossRef] [PubMed]
- Wayman, G.A.; Davare, M.; Ando, H.; Fortin, D.; Varlamova, O.; Cheng, H.Y.M.; Marks, D.; Obrietan, K.; Soclerling, T.R.; Goodman, R.H.; et al. An activity-regulated microRNA controls dendritic plasticity by down-regulating p250GAP. Proc. Natl. Acad. Sci. USA 2008, 105, 9093–9098. [Google Scholar] [CrossRef] [PubMed]
- Sim, S.E.; Bakes, J.; Kaang, B.K. Neuronal activity-dependent regulation of microRNAs. Mol. Cells 2014, 37, 511–517. [Google Scholar] [CrossRef] [PubMed]
- Aksoy-Aksel, A.; Zampa, F.; Schratt, G. MicroRNAs and synaptic plasticity-a mutual relationship. Philos Trans. R. Soc. B 2014, 369. [Google Scholar] [CrossRef] [PubMed]
- Eacker, S.M.; Dawson, T.M.; Dawson, V.L. The interplay of microRNA and neuronal activity in health and disease. Front. Cell Neurosci. 2013, 7, 136. [Google Scholar] [CrossRef] [PubMed]
- Bicker, S.; Khudayberdiev, S.; Weiss, K.; Zocher, K.; Baumeister, S.; Schratt, G. The deah-box helicase DHX36 mediates dendritic localization of the neuronal precursor-microRNA-134. Genes Dev. 2013, 27, 991–996. [Google Scholar] [PubMed]
- Kim, H.H.; Kim, P.; Phay, M.; Yoo, S. Identification of precursor microRNAs within distal axons of sensory neuron. J. Neurochem. 2015, 134, 193–199. [Google Scholar] [CrossRef] [PubMed]
- Vo, N.; Klein, M.E.; Varlamova, O.; Keller, D.M.; Yamamoto, T.; Goodman, R.H.; Impey, S. A camp-response element binding protein-induced microRNA regulates neuronal morphogenesis. Proc. Natl. Acad. Sci. USA 2005, 102, 16426–16431. [Google Scholar] [CrossRef] [PubMed]
- Remenyi, J.; Hunter, C.J.; Cole, C.; Ando, H.; Impey, S.; Monk, C.E.; Martin, K.J.; Barton, G.J.; Hutvagner, G.; Arthur, J.S.C. Regulation of the miR-212/132 locus by MSK1 and CREB in response to neurotrophins. Biochem. J. 2010, 428, 281–291. [Google Scholar] [CrossRef] [PubMed]
- Herbert, K.M.; Pimienta, G.; DeGregorio, S.J.; Alexandrov, A.; Steitz, J.A. Phosphorylation of DGCR8 increases its intracellular stability and induces a progrowth miRNA profile. Cell Rep. 2013, 5, 1070–1081. [Google Scholar] [CrossRef] [PubMed]
- Memczak, S.; Jens, M.; Elefsinioti, A.; Torti, F.; Krueger, J.; Rybak, A.; Maier, L.; Mackowiak, S.D.; Gregersen, L.H.; Munschauer, M.; et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 2013, 495, 333–338. [Google Scholar] [CrossRef] [PubMed]
- Hansen, T.B.; Jensen, T.I.; Clausen, B.H.; Bramsen, J.B.; Finsen, B.; Damgaard, C.K.; Kjems, J. Natural RNA circles function as efficient microRNA sponges. Nature 2013, 495, 384–388. [Google Scholar] [CrossRef] [PubMed]
- Turrigiano, G. Homeostatic synaptic plasticity: Local and global mechanisms for stabilizing neuronal function. Cold Spring Harb. Perspect. Biol. 2012, 4. [Google Scholar] [CrossRef] [PubMed]
- Temple, S. The development of neural stem cells. Nature 2001, 414, 112–117. [Google Scholar] [CrossRef] [PubMed]
- Eriksson, P.S.; Perfilieva, E.; Bjork-Eriksson, T.; Alborn, A.M.; Nordborg, C.; Peterson, D.A.; Gage, F.H. Neurogenesis in the adult human hippocampus. Nat. Med. 1998, 4, 1313–1317. [Google Scholar] [CrossRef] [PubMed]
- Reynolds, B.A.; Weiss, S. Generation of neurons and astrocytes from isolated cells of the adult mammalian central-nervous-system. Science 1992, 255, 1707–1710. [Google Scholar] [CrossRef] [PubMed]
- Bian, S.; Xu, T.L.; Sun, T. Tuning the cell fate of neurons and glia by microRNAs. Curr. Opin. Neurobiol. 2013, 23, 928–934. [Google Scholar] [CrossRef] [PubMed]
- Stappert, L.; Roese-Koerner, B.; Brustle, O. The role of microRNAs in human neural stem cells, neuronal differentiation and subtype specification. Cell. Tissue Res. 2015, 359, 47–64. [Google Scholar] [CrossRef] [PubMed]
- Visvanathan, J.; Lee, S.; Lee, B.; Lee, J.W.; Lee, S.K. The microRNA miR-124 antagonizes the anti-neural REST/SCP1 pathway during embryonic CNS development. Gene Dev. 2007, 21, 744–749. [Google Scholar] [CrossRef] [PubMed]
- Makeyev, E.V.; Zhang, J.W.; Carrasco, M.A.; Maniatis, T. The microRNA miR-124 promotes neuronal differentiation by triggering brain-specific alternative pre-mRNA splicing. Mol. Cell 2007, 27, 435–448. [Google Scholar] [CrossRef] [PubMed]
- Cao, X.W.; Pfaff, S.L.; Gage, F.H. A functional study of miR-124 in the developing neural tube. Gene Dev. 2007, 21, 531–536. [Google Scholar] [CrossRef] [PubMed]
- Cheng, L.C.; Pastrana, E.; Tavazoie, M.; Doetsch, F. miR-124 regulates adult neurogenesis in the subventricular zone stem cell niche. Nat. Neurosci. 2009, 12, 399–408. [Google Scholar] [CrossRef] [PubMed]
- Mondanizadeh, M.; Arefian, E.; Mosayebi, G.; Saidijam, M.; Khansarinejad, B.; Hashemi, S.M. MicroRNA-124 regulates neuronal differentiation of mesenchymal stem cells by targeting SP1 mRNA. J. Cell Biochem. 2015, 116, 943–953. [Google Scholar] [CrossRef] [PubMed]
- Pedersen, M.E.; Snieckute, G.; Kagias, K.; Nehammer, C.; Multhaupt, H.A.B.; Couchman, J.R.; Pocock, R. An epidermal microRNA regulates neuronal migration through control of the cellular glycosylation state. Science 2013, 341, 1404–1408. [Google Scholar] [CrossRef] [PubMed]
- Luikart, B.W.; Bensen, A.L.; Washburn, E.K.; Perederiy, J.V.; Su, K.G.; Li, Y.; Kernie, S.G.; Parada, L.F.; Westbrook, G.L. miR-132 mediates the integration of newborn neurons into the adult dentate gyrus. PLoS ONE 2011, 6, e19077. [Google Scholar] [CrossRef] [PubMed]
- Rago, L.; Beattie, R.; Taylor, V.; Winter, J. MiR379–410 cluster miRNAs regulate neurogenesis and neuronal migration by fine-tuning N-cadherin. EMBO J. 2014, 33, 906–920. [Google Scholar] [CrossRef] [PubMed]
- Giusti, S.A.; Vogl, A.M.; Brockmann, M.M.; Vercelli, C.A.; Rein, M.L.; Trumbach, D.; Wurst, W.; Cazalla, D.; Stein, V.; Deussing, J.M.; et al. MicroRNA-9 controls dendritic development by targeting REST. Elife 2014, 3. [Google Scholar] [CrossRef] [PubMed]
- Magill, S.T.; Cambronne, X.A.; Luikart, B.W.; Lioy, D.T.; Leighton, B.H.; Westbrook, G.L.; Mandel, G.; Goodman, R.H. MicroRNA-132 regulates dendritic growth and arborization of newborn neurons in the adult hippocampus. Proc. Natl. Acad. Sci. USA 2010, 107, 20382–20387. [Google Scholar] [CrossRef] [PubMed]
- Pathania, M.; Torres-Reveron, J.; Yan, L.; Kimura, T.; Lin, T.V.; Gordon, V.; Teng, Z.Q.; Zhao, X.Y.; Fulga, T.A.; Van Vactor, D.; et al. miR-132 enhances dendritic morphogenesis, spine density, synaptic integration, and survival of newborn olfactory bulb neurons. PLoS ONE 2012, 7, e38174. [Google Scholar] [CrossRef] [PubMed]
- Jasinska, M.; Milek, J.; Cymerman, I.A.; Leski, S.; Kaczmarek, L.; Dziembowska, M. miR-132 regulates dendritic spine structure by direct targeting of matrix metalloproteinase 9 mRNA. Mol. Neurobiol. 2015. [Google Scholar] [CrossRef] [PubMed]
- Dhar, M.; Zhu, M.Y.; Impey, S.; Lambert, T.J.; Bland, T.; Karatsoreos, I.N.; Nakazawa, T.; Appleyard, S.M.; Wayman, G.A. Leptin induces hippocampal synaptogenesis via creb-regulated microRNA-132 suppression of p250GAP. Mol. Endocrinol. 2014, 28, 1073–1087. [Google Scholar] [CrossRef] [PubMed]
- Dajas-Bailador, F.; Bonev, B.; Garcez, P.; Stanley, P.; Guillemot, F.; Papalopulu, N. MicroRNA-9 regulates axon extension and branching by targeting map1b in mouse cortical neurons. Nat. Neurosci. 2012, 15, 697–699. [Google Scholar] [CrossRef] [PubMed]
- Shibata, M.; Nakao, H.; Kiyonari, H.; Abe, T.; Aizawa, S. MicroRNA-9 regulates neurogenesis in mouse telencephalon by targeting multiple transcription factors. J. Neurosci. 2011, 31, 3407–3422. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Ueno, Y.; Liu, X.S.; Buller, B.; Wang, X.L.; Chopp, M.; Zhang, Z.G. The microRNA-17–92 cluster enhances axonal outgrowth in embryonic cortical neurons. J. Neurosci. 2013, 33, 6885–6894. [Google Scholar] [CrossRef] [PubMed]
- Baudet, M.L.; Zivraj, K.H.; Abreu-Goodger, C.; Muldal, A.; Armisen, J.; Blenkiron, C.; Goldstein, L.D.; Miska, E.A.; Holt, C.E. miR-124 acts through corest to control onset of SeMA3A sensitivity in navigating retinal growth cones. Nat. Neurosci. 2012, 15, U29–U38. [Google Scholar] [CrossRef] [PubMed]
- Pinter, R.; Hindges, R. Perturbations of microRNA function in mouse dicer mutants produce retinal defects and lead to aberrant axon pathfinding at the optic chiasm. PLoS ONE 2010, 5, e10021. [Google Scholar] [CrossRef] [PubMed]
- Giraldez, A.J.; Cinalli, R.M.; Glasner, M.E.; Enright, A.J.; Thomson, J.M.; Baskerville, S.; Hammond, S.M.; Bartel, D.P.; Schier, A.F. MicroRNAs regulate brain morphogenesis in zebrafish. Science 2005, 308, 833–838. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.Y.; Phang, R.Z.; Hsu, P.H.; Wang, W.H.; Huang, H.T.; Liu, I.Y. In vivo knockdown of hippocampal miR-132 expression impairs memory acquisition of trace fear conditioning. Hippocampus 2013, 23, 625–633. [Google Scholar] [CrossRef] [PubMed]
- Karr, J.; Vagin, V.; Chen, K.Y.; Ganesan, S.; Olenkina, O.; Gvozdev, V.; Featherstone, D.E. Regulation of glutamate receptor subunit availability by microRNAs. J. Cell Biol. 2009, 185, 685–697. [Google Scholar] [CrossRef] [PubMed]
- Ponomarev, E.D.; Veremeyko, T.; Barteneva, N.; Krichevsky, A.M.; Weiner, H.L. MicroRNA-124 promotes microglia quiescence and suppresses eae by deactivating macrophages via the C/EBP-α PU.1 pathway. Nat. Med. 2011, 17, U64–U70. [Google Scholar] [CrossRef] [PubMed]
- Jayadev, S.; Case, A.; Alajajian, B.; Eastman, A.J.; Moller, T.; Garden, G.A. Presenilin 2 influences miR146 level and activity in microglia. J. Neurochem. 2013, 127, 592–599. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.Y.; Cui, J.G.; Dua, P.; Pogue, A.I.; Bhattacharjee, S.; Lukiw, W.J. Differential expression of miRNA-146a-regulated inflammatory genes in human primary neural, astroglial and microglial cells. Neurosci. Lett. 2011, 499, 109–113. [Google Scholar] [CrossRef] [PubMed]
- Alexandrov, P.N.; Dua, P.; Lukiw, W.J. Up-regulation of miRNA-146a in progressive, age-related inflammatory neurodegenerative disorders of the human CNS. Front. Neurol. 2014, 5, 181. [Google Scholar] [CrossRef] [PubMed]
- Su, W.; Hopkins, S.; Nesser, N.K.; Sopher, B.; Silvestroni, A.; Ammanuel, S.; Jayadev, S.; Moller, T.; Weinstein, J.; Garden, G.A. The p53 transcription factor modulates microglia behavior through microRNA-dependent regulation of C-Maf. J. Immunol. 2014, 192, 358–366. [Google Scholar] [CrossRef] [PubMed]
- Kawase-Koga, Y.; Otaegi, G.; Sun, T. Different timings of dicer deletion affect neurogenesis and gliogenesis in the developing mouse central nervous system. Dev. Dynam. 2009, 238, 2800–2812. [Google Scholar] [CrossRef] [PubMed]
- Davis, T.H.; Cuellar, T.L.; Koch, S.M.; Barker, A.J.; Harfe, B.D.; McManus, M.T.; Ullian, E.M. Conditional loss of dicer disrupts cellular and tissue morphogenesis in the cortex and hippocampus. J. Neurosci. 2008, 28, 4322–4330. [Google Scholar] [CrossRef] [PubMed]
- Tao, J.F.; Wu, H.; Lin, Q.; Wei, W.Z.; Lu, X.H.; Cantle, J.P.; Ao, Y.; Olsen, R.W.; Yang, X.W.; Mody, I.; et al. Deletion of astroglial dicer causes non-cell-autonomous neuronal dysfunction and degeneration. J. Neurosci. 2011, 31, 8306–8319. [Google Scholar] [CrossRef] [PubMed]
- Lim, L.P.; Lau, N.C.; Garrett-Engele, P.; Grimson, A.; Schelter, J.M.; Castle, J.; Bartel, D.P.; Linsley, P.S.; Johnson, J.M. Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature 2005, 433, 769–773. [Google Scholar] [CrossRef] [PubMed]
- Conaco, C.; Otto, S.; Han, J.J.; Mandel, G. Reciprocal actions of rest and a microRNA promote neuronal identity. Proc. Natl. Acad. Sci. USA 2006, 103, 2422–2427. [Google Scholar] [CrossRef] [PubMed]
- Dickson, B.J. Molecular mechanisms of axon guidance. Science 2002, 298, 1959–1964. [Google Scholar] [CrossRef] [PubMed]
- Kolodkin, A.L.; Tessier-Lavigne, M. Mechanisms and molecules of neuronal wiring: A primer. Cold Spring Herb. Perspect. Biol. 2011, 3. [Google Scholar] [CrossRef] [PubMed]
- Ashraf, S.I.; McLoon, A.L.; Sclarsic, S.M.; Kunes, S. Synaptic protein synthesis associated with memory is regulated by the risc pathway in drosophila. Cell 2006, 124, 191–205. [Google Scholar] [CrossRef] [PubMed]
- Neves, G.; Cooke, S.F.; Bliss, T.V.P. Synaptic plasticity, memory and the hippocampus: A neural network approach to causality. Nat. Rev. Neurosci. 2008, 9, 65–75. [Google Scholar] [CrossRef] [PubMed]
- Schratt, G. microRNAs at the synapse. Nat. Rev. Neurosci. 2011, 12, 182. [Google Scholar] [CrossRef]
- Ju, W.; Morishita, W.; Tsui, J.; Gaietta, G.; Deerinck, T.J.; Adams, S.R.; Garner, C.C.; Tsien, R.Y.; Ellisman, M.H.; Malenka, R.C. Activity-dependent regulation of dendritic synthesis and trafficking of AMPA receptors. Nat. Neurosci. 2004, 7, 244–253. [Google Scholar] [CrossRef] [PubMed]
- Su, W.; Aloi, M.S.; Garden, G.A. MicroRNAs mediating CNS inflammation: Small regulators with powerful potential. Brain Behav. Immun. 2016, 52, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Jovicic, A.; Roshan, R.; Moisoi, N.; Pradervand, S.; Moser, R.; Pillai, B.; Luthi-Carter, R. Comprehensive expression analyses of neural cell-type-specific miRNAs identify new determinants of the specification and maintenance of neuronal phenotypes. J. Neurosci. 2013, 33, 5127–5137. [Google Scholar] [CrossRef] [PubMed]
- Butovsky, O.; Jedrychowski, M.P.; Moore, C.S.; Cialic, R.; Lanser, A.J.; Gabriely, G.; Koeglsperger, T.; Dake, B.; Wu, P.M.; Doykan, C.E.; et al. Identification of a unique TGF-β-dependent molecular and functional signature in microglia. Nat. Neurosci. 2014, 17, 131–143. [Google Scholar] [CrossRef] [PubMed]
- Freilich, R.W.; Woodbury, M.E.; Ikezu, T. Integrated expression profiles of mRNA and miRNA in polarized primary murine microglia. PLoS ONE 2013, 8, e79416. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.L.; Li, Y.; Wang, F.; Gao, F.B. The steady-state level of the nervous-system-specific microRNA-124a is regulated by dFMR1 in drosophila. J. Neurosci. 2008, 28, 11883–11889. [Google Scholar] [CrossRef] [PubMed]
- Edbauer, D.; Neilson, J.R.; Foster, K.A.; Wang, C.F.; Seeburg, D.P.; Batterton, M.N.; Tada, T.; Dolan, B.M.; Sharp, P.A.; Sheng, M. Regulation of synaptic structure and function by fmrp-associated microRNAs miR-125b and miR-132. Neuron 2010, 65, 373–384. [Google Scholar] [CrossRef] [PubMed]
- Hansen, K.F.; Sakamoto, K.; Wayman, G.A.; Impey, S.; Obrietan, K. Transgenic miR132 alters neuronal spine density and impairs novel object recognition memory. PLoS ONE 2010, 5, e15497. [Google Scholar] [CrossRef] [PubMed]
- Nomura, T.; Kimura, M.; Horii, T.; Morita, S.; Soejima, H.; Kudo, S.; Hatada, I. MeCP2-dependent repression of an imprinted miR-184 released by depolarization. Hum. Mol. Genet. 2008, 17, 1192–1199. [Google Scholar] [CrossRef] [PubMed]
- Fiore, R.; Khudayberdiev, S.; Christensen, M.; Siegel, G.; Flavell, S.W.; Kim, T.K.; Greenberg, M.E.; Schratt, G. MEF2-mediated transcription of the miR379-410 cluster regulates activity-dependent dendritogenesis by fine-tuning PUMILIO2 protein levels. EMBO J. 2009, 28, 697–710. [Google Scholar] [CrossRef] [PubMed]
- Xu, B.; Hsu, P.K.; Karayiorgou, M.; Gogos, J.A. MicroRNA dysregulation in neuropsychiatric disorders and cognitive dysfunction. Neurobiol. Dis. 2012, 46, 291–301. [Google Scholar] [CrossRef] [PubMed]
- Miller, D.T.; Shen, Y.; Weiss, L.A.; Korn, J.; Anselm, I.; Bridgemohan, C.; Cox, G.F.; Dickinson, H.; Gentile, J.; Harris, D.J.; et al. Microdeletion/duplication at 15q13.2q13.3 among individuals with features of autism and other neuropsychiatric disorders. J. Med. Genet. 2009, 46, 242–248. [Google Scholar] [CrossRef] [PubMed]
- Packer, A.N.; Xing, Y.; Harper, S.Q.; Jones, L.; Davidson, B.L. The bifunctional microRNA miR-9/miR-9* regulates rest and corest and is downregulated in huntington’s disease. J. Neurosci. 2008, 28, 14341–14346. [Google Scholar] [CrossRef] [PubMed]
- Bai, M.; Zhu, X.Z.; Zhang, Y.; Zhang, S.; Zhang, L.; Xue, L.; Yi, J.Y.; Yao, S.Q.; Zhang, X.W. Abnormal hippocampal BDNF and miR-16 expression is associated with depression-like behaviors induced by stress during early life. PLoS ONE 2012, 7, e46921. [Google Scholar] [CrossRef] [PubMed]
- Lopez, J.P.; Lim, R.; Cruceanu, C.; Crapper, L.; Fasano, C.; Labonte, B.; Maussion, G.; Yang, J.P.; Yerko, V.; Vigneault, E.; et al. miR-1202 is a primate-specific and brain-enriched microRNA involved in major depression and antidepressant treatment. Nat. Med. 2014, 20, 764–768. [Google Scholar] [CrossRef] [PubMed]
- Stark, K.L.; Xu, B.; Bagchi, A.; Lai, W.S.; Liu, H.; Hsu, R.; Wan, X.; Pavlidis, P.; Mills, A.A.; Karayiorgou, M.; et al. Altered brain microRNA biogenesis contributes to phenotypic deficits in a 22q11-deletion mouse model. Nat. Genet. 2008, 40, 751–760. [Google Scholar] [CrossRef] [PubMed]
- Stone, J.L.; O’Donovan, M.C.; Gurling, H.; Kirov, G.K.; Blackwood, D.H.R.; Corvin, A.; Craddock, N.J.; Gill, M.; Hultman, C.M.; Lichtenstein, P.; et al. Rare chromosomal deletions and duplications increase risk of schizophrenia. Nature 2008, 455, 237–241. [Google Scholar] [CrossRef] [PubMed]
- Ingason, A.; Rujescu, D.; Cichon, S.; Sigurdsson, E.; Sigmundsson, T.; Pietilainen, O.P.H.; Buizer-Voskamp, J.E.; Strengman, E.; Francks, C.; Muglia, P.; et al. Copy number variations of chromosome 16p13.1 region associated with schizophrenia. Mol. Psychiatry 2011, 16, 17–25. [Google Scholar] [CrossRef] [PubMed]
- Kwon, E.; Wang, W.; Tsai, L.H. Validation of schizophrenia-associated genes CSMD1, C10ORF26, CACNA1C and TCF4 as miR-137 targets. Mol. Psychiatry 2013, 18, 11–12. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.K.; Wang, X.; Li, L.; Du, Y.H.; Ye, H.T.; Li, C.Y. MicroRNA-98 induces an Alzheimer’s disease-like disturbance by targeting insulin-like growth factor 1. Neurosci. Bull. 2013, 29, 745–751. [Google Scholar] [CrossRef] [PubMed]
- Smith, P.; Al Hashimi, A.; Girard, J.; Delay, C.; Hebert, S.S. In vivo regulation of amyloid precursor protein neuronal splicing by microRNAs. J. Neurochem. 2011, 116, 240–247. [Google Scholar] [CrossRef] [PubMed]
- Absalon, S.; Kochanek, D.M.; Raghavan, V.; Krichevsky, A.M. MiR-26b, upregulated in Alzheimer’s disease, activates cell cycle entry, tau-phosphorylation, and apoptosis in postmitotic neurons. J. Neurosci. 2013, 33, 14645–14659. [Google Scholar] [CrossRef] [PubMed]
- Dickson, J.R.; Kruse, C.; Montagna, D.R.; Finsen, B.; Wolfe, M.S. Alternative polyadenylation and miR-34 family members regulate tau expression. J. Neurochem. 2013, 127, 739–749. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.H.; Pogue, A.I.; Lukiw, W.J. MicroRNA (miRNA) signaling in the human CNS in sporadic Alzheimer’s disease (AD)-novel and unique pathological features. Int. J. Mol. Sci. 2015, 16, 30105–30116. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Inoue, K.; Ishii, J.; Vanti, W.B.; Voronov, S.V.; Murchison, E.; Hannon, G.; Abeliovich, A. A microRNA feedback circuit in midbrain dopamine neurons. Science 2007, 317, 1220–1224. [Google Scholar] [CrossRef] [PubMed]
- Gehrke, S.; Imai, Y.; Sokol, N.; Lu, B. Pathogenic LRRK2 negatively regulates microRNA-mediated translational repression. Nature 2010, 466, 637–641. [Google Scholar] [CrossRef] [PubMed]
- Cho, H.J.; Liu, G.; Jin, S.M.; Parisiadou, L.; Xie, C.; Yu, J.; Sun, L.; Ma, B.; Ding, J.; Vancraenenbroeck, R.; et al. MicroRNA-205 regulates the expression of parkinson’s disease-related leucine-rich repeat kinase 2 protein. Hum. Mol. Genet. 2013, 22, 608–620. [Google Scholar] [CrossRef] [PubMed]
- Doxakis, E. Post-transcriptional regulation of α-synuclein expression by miR-7 and miR-153. J. Biol. Chem. 2010, 285, 12726–12734. [Google Scholar] [CrossRef] [PubMed]
- Gupta, S.; Verma, S.; Mantri, S.; Berman, N.E.; Sandhir, R. Targeting microRNAs in prevention and treatment of neurodegenerative disorders. Drug Dev. Res. 2015, 76, 397–418. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; van der Walt, J.M.; Mayhew, G.; Li, Y.J.; Zuchner, S.; Scott, W.K.; Martin, E.R.; Vance, J.M. Variation in the miRNA-433 binding site of FGF20 confers risk for parkinson disease by overexpression of α-synuclein. Am. J. Hum. Genet. 2008, 82, 283–289. [Google Scholar] [CrossRef] [PubMed]
- Penagarikano, O.; Mulle, J.G.; Warren, S.T. The pathophysiology of fragile X syndrome. Annu. Rev. Genom. Hum. G 2007, 8, 109–129. [Google Scholar] [CrossRef] [PubMed]
- Dictenberg, J.B.; Swanger, S.A.; Antar, L.N.; Singer, R.H.; Bassell, G.J. A direct role for FMRP in activity-dependent dendritic mRNA transport links filopodial-spine morphogenesis to fragile X syndrome. Dev. Cell 2008, 14, 926–939. [Google Scholar] [CrossRef] [PubMed]
- Huber, K.M.; Gallagher, S.M.; Warren, S.T.; Bear, M.F. Altered synaptic plasticity in a mouse model of fragile X mental retardation. Proc. Natl. Acad. Sci. USA 2002, 99, 7746–7750. [Google Scholar] [CrossRef] [PubMed]
- Urdinguio, R.G.; Fernandez, A.F.; Lopez-Nieva, P.; Rossi, S.; Huertas, D.; Kulis, M.; Liu, C.G.; Croce, C.; Calin, G.A.; Esteller, M. Disrupted microRNA expression caused by MeCP2 loss in a mouse model of rett syndrome. Epigenetics 2010, 5, 656–663. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Tao, J.F.; Chen, P.J.; Shahab, A.; Ge, W.H.; Hart, R.P.; Ruan, X.A.; Ruan, Y.J.; Sun, Y.E. Genome-wide analysis reveals methyl-CPG-binding protein 2-dependent regulation of microRNAs in a mouse model of rett syndrome. Proc. Natl. Acad. Sci. USA 2010, 107, 18161–18166. [Google Scholar] [CrossRef] [PubMed]
- Percy, A.K. Rett syndrome exploring the autism link. Arch. Neurol. Chic. 2011, 68, 985–989. [Google Scholar] [CrossRef] [PubMed]
- Talebizadeh, Z.; Butler, M.G.; Theodoro, M.F. Feasibility and relevance of examining lymphoblastoid cell lines to study role of microRNAs in autism. Autism Res. 2008, 1, 240–250. [Google Scholar] [CrossRef] [PubMed]
- Abu-Elneel, K.; Liu, T.; Gazzaniga, F.S.; Nishimura, Y.; Wall, D.P.; Geschwind, D.H.; Lao, K.Q.; Kosik, K.S. Heterogeneous dysregulation of microRNAs across the autism spectrum. Neurogenetics 2008, 9, 153–161. [Google Scholar] [CrossRef] [PubMed]
- Sarachana, T.; Zhou, R.L.; Chen, G.; Manji, H.K.; Hu, V.W. Investigation of post-transcriptional gene regulatory networks associated with autism spectrum disorders by microRNA expression profiling of lymphoblastoid cell lines. Genome Med. 2010, 2. [Google Scholar] [CrossRef] [PubMed]
- Seno, M.M.G.; Hu, P.Z.; Gwadry, F.G.; Pinto, D.; Marshall, C.R.; Casallo, G.; Scherer, S.W. Gene and miRNA expression profiles in autism spectrum disorders. Brain Res. 2011, 1380, 85–97. [Google Scholar] [CrossRef] [PubMed]
- Marshall, C.R.; Noor, A.; Vincent, J.B.; Lionel, A.C.; Feuk, L.; Skaug, J.; Shago, M.; Moessner, R.; Pinto, D.; Ren, Y.; et al. Structural variation of chromosomes in autism spectrum disorder. Am. J. Hum. Genet. 2008, 82, 477–488. [Google Scholar] [CrossRef] [PubMed]
- Glessner, J.T.; Wang, K.; Cai, G.Q.; Korvatska, O.; Kim, C.E.; Wood, S.; Zhang, H.T.; Estes, A.; Brune, C.W.; Bradfield, J.P.; et al. Autism genome-wide copy number variation reveals ubiquitin and neuronal genes. Nature 2009, 459, 569–573. [Google Scholar] [CrossRef] [PubMed]
- Leistedt, S.J.; Linkowski, P. Brain, networks, depression, and more. Eur. Neuropsychopharmacol. 2013, 23, 55–62. [Google Scholar] [CrossRef] [PubMed]
- Ota, K.T.; Duman, R.S. Environmental and pharmacological modulations of cellular plasticity: Role in the pathophysiology and treatment of depression. Neurobiol. Dis. 2013, 57, 28–37. [Google Scholar] [CrossRef] [PubMed]
- Smalheiser, N.R.; Lugli, G.; Rizavi, H.S.; Torvik, V.I.; Turecki, G.; Dwivedi, Y. MicroRNA expression is down-regulated and reorganized in prefrontal cortex of depressed suicide subjects. PLoS ONE 2012, 7, e33201. [Google Scholar]
- He, Y.; Zhou, Y.; Xi, Q.; Cui, H.Y.; Luo, T.X.; Song, H.L.; Nie, X.; Wang, L.L.; Ying, B.W. Genetic variations in microRNA processing genes are associated with susceptibility in depression. DNA Cell Biol. 2012, 31, 1499–1506. [Google Scholar] [CrossRef] [PubMed]
- Meerson, A.; Cacheaux, L.; Goosens, K.A.; Sapolsky, R.M.; Soreq, H.; Kaufer, D. Changes in brain microRNAs contribute to cholinergic stress reactions. J. Mol. Neurosci. 2010, 40, 47–55. [Google Scholar] [CrossRef] [PubMed]
- Uchida, S.; Hara, K.; Kobayashi, A.; Funato, H.; Hobara, T.; Otsuki, K.; Yamagata, H.; McEwen, B.S.; Watanabe, Y. Early life stress enhances behavioral vulnerability to stress through the activation of REST4-mediated gene transcription in the medial prefrontal cortex of rodents. J. Neurosci. 2010, 30, 15007–15018. [Google Scholar] [CrossRef] [PubMed]
- Perkins, D.O.; Jeffries, C.D.; Jarskog, L.F.; Thomson, J.M.; Woods, K.; Newman, M.A.; Parker, J.S.; Jin, J.P.; Hammond, S.M. MicroRNA expression in the prefrontal cortex of individuals with schizophrenia and schizoaffective disorder. Genome Biol. 2007, 8, R27. [Google Scholar] [CrossRef] [PubMed]
- Beveridge, N.J.; Gardiner, E.; Carroll, A.P.; Tooney, P.A.; Cairns, M.J. Schizophrenia is associated with an increase in cortical microRNA biogenesis. Mol. Psychiatry 2010, 15, 1176–1189. [Google Scholar] [CrossRef] [PubMed]
- Kim, A.H.; Reimers, M.; Mahera, B.; Williamson, V.; McMichael, O.I.; McClay, J.L.; van den Oord, E.J.C.G.; Riley, B.P.; Kendler, K.S.; Vladimirov, V.I. MicroRNA expression profiling in the prefrontal cortex of individuals affected with schizophrenia and bipolar disorders. Schizophr. Res. 2010, 124, 183–191. [Google Scholar] [CrossRef] [PubMed]
- Moreau, M.P.; Bruse, S.E.; David-Rus, R.; Buyske, S.; Brzustowicz, L.M. Altered microRNA expression profiles in postmortem brain samples from individuals with schizophrenia and bipolar disorder. Biol. Psychiatry 2011, 69, 188–193. [Google Scholar] [CrossRef] [PubMed]
- Santarelli, D.M.; Beveridge, N.J.; Tooney, P.A.; Cairns, M.J. Upregulation of dicer and microRNA expression in the dorsolateral prefrontal cortex brodmann area 46 in schizophrenia. Biol. Psychiatry 2011, 69, 180–187. [Google Scholar] [CrossRef] [PubMed]
- Drew, L.J.; Crabtree, G.W.; Markx, S.; Stark, K.L.; Chaverneff, F.; Xu, B.; Mukai, J.; Fenelon, K.; Hsu, P.K.; Gogos, J.A.; et al. The 22q11.2 microdeletion: Fifteen years of insights into the genetic and neural complexity of psychiatric disorders. Int. J. Dev. Neurosci. 2011, 29, 259–281. [Google Scholar] [CrossRef] [PubMed]
- Stefansson, H.; Rujescu, D.; Cichon, S.; Pietilainen, O.P.H.; Ingason, A.; Steinberg, S.; Fossdal, R.; Sigurdsson, E.; Sigmundsson, T.; Buizer-Voskamp, J.E.; et al. Large recurrent microdeletions associated with schizophrenia. Nature 2008, 455, U232–U261. [Google Scholar] [CrossRef] [PubMed]
- Xu, B.; Roos, J.L.; Levy, S.; Van Rensburg, E.J.; Gogos, J.A.; Karayiorgou, M. Strong association of de novo copy number mutations with sporadic schizophrenia. Nat. Genet. 2008, 40, 880–885. [Google Scholar] [CrossRef] [PubMed]
- O’Brien, R.J.; Wong, P.C. Amyloid precursor protein processing and Alzheimer’s disease. Annu. Rev. Neurosci. 2011, 34, 185–204. [Google Scholar] [CrossRef] [PubMed]
- Cogswell, J.P.; Ward, J.; Taylor, I.A.; Waters, M.; Shi, Y.L.; Cannon, B.; Kelnar, K.; Kemppainen, J.; Brown, D.; Chen, C.; et al. Identification of miRNA changes in Alzheimer’s disease brain and CSF yields putative biomarkers and insights into disease pathways. J. Alzheimer’s Dis. 2008, 14, 27–41. [Google Scholar]
- Tan, L.; Yu, J.T.; Tan, M.S.; Liu, Q.Y.; Wang, H.F.; Zhang, W.; Jiang, T.; Tan, L. Genome-wide serum microRNA expression profiling identifies serum biomarkers for Alzheimer’s disease. J. Alzheimer’s Dis. 2014, 40, 1017–1027. [Google Scholar]
- Denk, J.; Boelmans, K.; Siegismund, C.; Lassner, D.; Arlt, S.; Jahn, H. MicroRNA profiling of CSF reveals potential biomarkers to detect Alzheimer’s disease. PLoS ONE 2015, 10, e0126423. [Google Scholar] [CrossRef] [PubMed]
- Itoh, N.; Ohta, H. Roles of FGF20 in dopaminergic neurons and parkinson’s disease. Front. Mol. Neurosci. 2013, 6, 15. [Google Scholar] [CrossRef] [PubMed]
- Hayes, J.; Peruzzi, P.P.; Lawler, S. MicroRNAs in cancer: Biomarkers, functions and therapy. Trends Mol. Med. 2014, 20, 460–469. [Google Scholar] [CrossRef] [PubMed]
- Haqqani, A.S.; Delaney, C.E.; Tremblay, T.L.; Sodja, C.; Sandhu, J.K.; Stanimirovic, D.B. Method for isolation and molecular characterization of extracellular microvesicles released from brain endothelial cells. Fluids Barriers CNS 2013, 10, 4. [Google Scholar] [CrossRef] [PubMed]
- Alvarez-Erviti, L.; Seow, Y.; Yin, H.; Betts, C.; Lakhal, S.; Wood, M.J. Delivery of siRNA to the mouse brain by systemic injection of targeted exosomes. Nat. Biotechnol. 2011, 29, 341–345. [Google Scholar] [CrossRef] [PubMed]
- Cardo, L.F.; Coto, E.; de Mena, L.; Ribacoba, R.; Moris, G.; Menendez, M.; Alvarez, V. Profile of microRNAs in the plasma of parkinson’s disease patients and healthy controls. J. Neurol. 2013, 260, 1420–1422. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Yu, J.T.; Tan, L.; Tian, Y.; Ma, J.; Tan, C.C.; Wang, H.F.; Liu, Y.; Tan, M.S.; Jiang, T.; et al. Genome-wide circulating microRNA expression profiling indicates biomarkers for epilepsy. Sci. Rep. 2015, 5, 9522. [Google Scholar] [CrossRef] [PubMed]
- Grasso, M.; Piscopo, P.; Confaloni, A.; Denti, M.A. Circulating miRNAs as biomarkers for neurodegenerative disorders. Molecules 2014, 19, 6891–6910. [Google Scholar] [CrossRef] [PubMed]
- Miller, J.D.; Ganat, Y.M.; Kishinevsky, S.; Bowman, R.L.; Liu, B.; Tu, E.Y.; Mandal, P.K.; Vera, E.; Shim, J.W.; Kriks, S.; et al. Human iPSC-based modeling of late-onset disease via progerin-induced aging. Cell Stem Cell 2013, 13, 691–705. [Google Scholar] [CrossRef] [PubMed]
- Sancho-Martinez, I.; Nivet, E.; Xia, Y.; Hishida, T.; Aguirre, A.; Ocampo, A.; Ma, L.; Morey, R.; Krause, M.N.; Zembrzycki, A.; et al. Establishment of human iPSC-based models for the study and targeting of glioma initiating cells. Nat. Commun. 2016, 7, 10743. [Google Scholar] [CrossRef] [PubMed]
- Ebert, A.D.; Liang, P.; Wu, J.C. Induced pluripotent stem cells as a disease modeling and drug screening platform. J. Cardiovasc. Pharm. 2012, 60, 408–416. [Google Scholar] [CrossRef] [PubMed]
| miRNA | Species | Targets/Pathway | Function | References |
|---|---|---|---|---|
| NSC Proliferation/Differentiation | ||||
| let-7b | Mouse | TLX, Cyclin D1 | NSC proliferation and neuronal differentiation | [102] |
| miR-9 | Mouse | TLX | NSC expansion and differentiation | [102,103] |
| miR-134 | Mouse | Dcx, Chrdl-1 | cortical NPC proliferation, neuron migration | [102] |
| miR-137 | Mouse | LSD1 | NSC expansion and differentiation | [102] |
| miR-124 | Chick | SCP1 | neuronal differentiation | [104] |
| Mouse | PTBP1 | neuronal differentiation | [105] | |
| Chick | LAMC1, ITGB1 | neuronal differentiation | [106] | |
| Mouse | SOX9 | neuronal differentiation | [107] | |
| Mouse | DLX2, JAG1 | – | [107] | |
| Human | SP1 | neuronal differentiation | [108] | |
| miR-184 | Mouse | Numbl | adult NSC proliferation and differentiation | [67] |
| Neuronal Migration and Integration | ||||
| miR-79 | C. elegans | SVQ-5, SVQ-7 | neuronal migration | [109] |
| miR-132 | Mouse | – | newborn neuron integration, dnedritic spine density | [110] |
| miR-379~410 cluster | Mouse | N-cadherin | NSC differentiation and neuronal migration | [111] |
| Dendritic Outgrowth and Complexity | ||||
| miR-9 | Mouse | REST | dendritic development, total dendritic length and complexity | [112] |
| miR-132 | Mouse | – | dendrite length, arborization, and spine density | [113] |
| Mouse | – | dendritic complexity and spine density | [114] | |
| Mouse | MMP-9 | dendritic spine structure | [115] | |
| Mouse | p250GAP | activity-dependent dendritic growth | [116] | |
| miR-134 | Rat | Limk1 | the size of dendritic spines | [76] |
| Axon Outgrowth and Guidance | ||||
| miR-9 | Mouse | Map1b | axonal extension and branching | [117] |
| Mouse | – | axon guidance | [118] | |
| miR-17~92 cluster | Rat | PTEN | axon outgrowth | [119] |
| miR-124 | Xenopus laevis | coREST | axon targeting | [120] |
| miR-132 | Rat | p250GAP | neurite outgrowth | [93] |
| miR-218 | – | Robo1/2 | axon guidance | [121] |
| miR-430 family | Zebrafish | – | axon fasciculation | [122] |
| Synaptogenesis and Synaptic Plasticity | ||||
| miR-132 | Mouse | – | the storage of temporally associated information | [123] |
| Mouse | – | light-induced clock resetting | [86] | |
| miR-219 | Mouse | – | circadian period length | [86] |
| miR-284 | Drosophila | GluRIIA and GluRIIB | postsynaptic strength | [124] |
| CNS Inflammation | ||||
| miR-124 | Mouse | CEBPα-PU.1 | microglia quiescence | [125] |
| miR-146a | Mouse & Human | NF-κB | pro-inflammatory cytokine expression | [126,127,128] |
| miR-155 | Mouse | cMAF | microglia pro-inflammatory function | [129] |
| Neurological Diseases | miRNA | Evidence | References |
|---|---|---|---|
| Fragile X Syndrome (FXS) | miR-124a | Involved in FMRP gene regulatory pathway | [145] |
| miR-125b, miR-132 | [146] | ||
| Rett Syndrome (RTT) | miR-132 | Involved in MeCP2 gene regulatory pathway | [84,93,147] |
| miR-184 | [148] | ||
| miR-379-410 | [149] | ||
| Autism spectrum disorder (ASD) | miR-185 | Located in ASD associated microduplication 22q11.2 | [150] |
| miR-211 | Located in ASD associated variant 15q13.2-q13.3 | [151] | |
| Major depression disorder (MDD) | miR-9 | Involved in REST gene regulatory pathway | [152] |
| miR-124 | [134] | ||
| miR-16 | Involved in BDNF regulatory pathway | [153] | |
| miR-1202 | biomarker of antidepressant response of MDD | [154] | |
| Schizophrenia (SCZ) | miR-185 | Located in SCZ associated microdeletion 22q11.2 | [155] |
| miR-211 | Encompassing SCZ associated genetic variants | [156] | |
| miR-484 | [157] | ||
| miR-137 | [158] | ||
| Alzheimer’s disease (AD) | miR-98 | Involved in APP processing by targeting IGF1 | [159] |
| miR-124 | Regulating production of APP isoforms by targeting PTBP1 | [160] | |
| miR-26b | Regulating Tau protein | [161] | |
| miR-34a | [162] | ||
| miR-146a | Regulated by Presenilin | [126] | |
| miR-7, miR-9, miR-34a, miR-125b, miR-146a | Involved in NF-κB regulated signaling pathway | [163] | |
| Parkinson disease (PD) | miR-133b | Regulating dopaminergic neurons by targeting Pitx3 | [164] |
| let-7, miR-184* | Involved in LRRK2 regulatory pathway | [165] | |
| miR-205 | [166] | ||
| miR-7, miR-153 | Predicted to target SNCA | [167] | |
| miR-433 | Regulating SNCA indirectly | [168,169] |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cao, D.-D.; Li, L.; Chan, W.-Y. MicroRNAs: Key Regulators in the Central Nervous System and Their Implication in Neurological Diseases. Int. J. Mol. Sci. 2016, 17, 842. https://doi.org/10.3390/ijms17060842
Cao D-D, Li L, Chan W-Y. MicroRNAs: Key Regulators in the Central Nervous System and Their Implication in Neurological Diseases. International Journal of Molecular Sciences. 2016; 17(6):842. https://doi.org/10.3390/ijms17060842
Chicago/Turabian StyleCao, Dan-Dan, Lu Li, and Wai-Yee Chan. 2016. "MicroRNAs: Key Regulators in the Central Nervous System and Their Implication in Neurological Diseases" International Journal of Molecular Sciences 17, no. 6: 842. https://doi.org/10.3390/ijms17060842
APA StyleCao, D.-D., Li, L., & Chan, W.-Y. (2016). MicroRNAs: Key Regulators in the Central Nervous System and Their Implication in Neurological Diseases. International Journal of Molecular Sciences, 17(6), 842. https://doi.org/10.3390/ijms17060842