AfAP2-1, An Age-Dependent Gene of Aechmea fasciata, Responds to Exogenous Ethylene Treatment
Abstract
:1. Introduction
2. Results
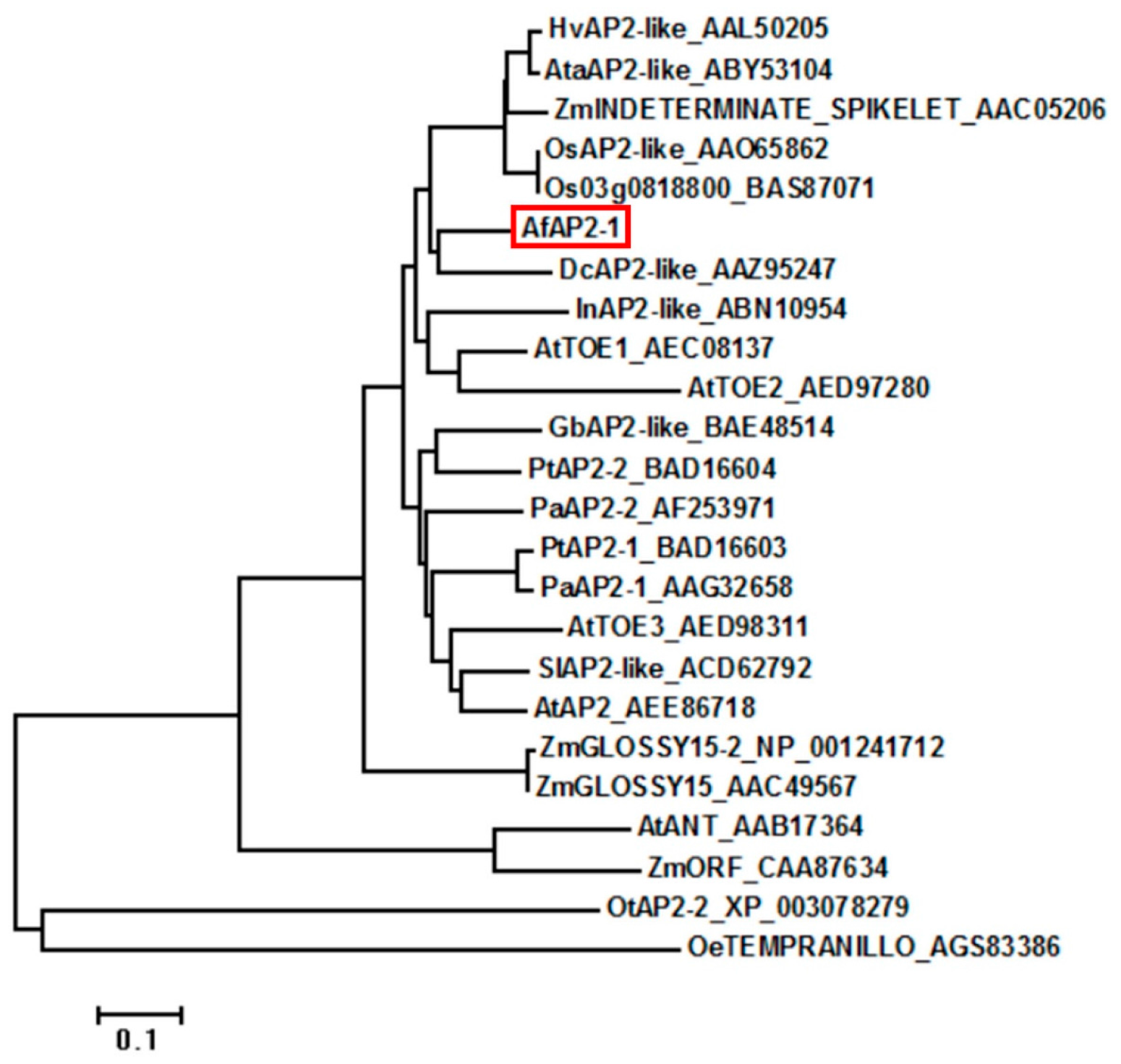
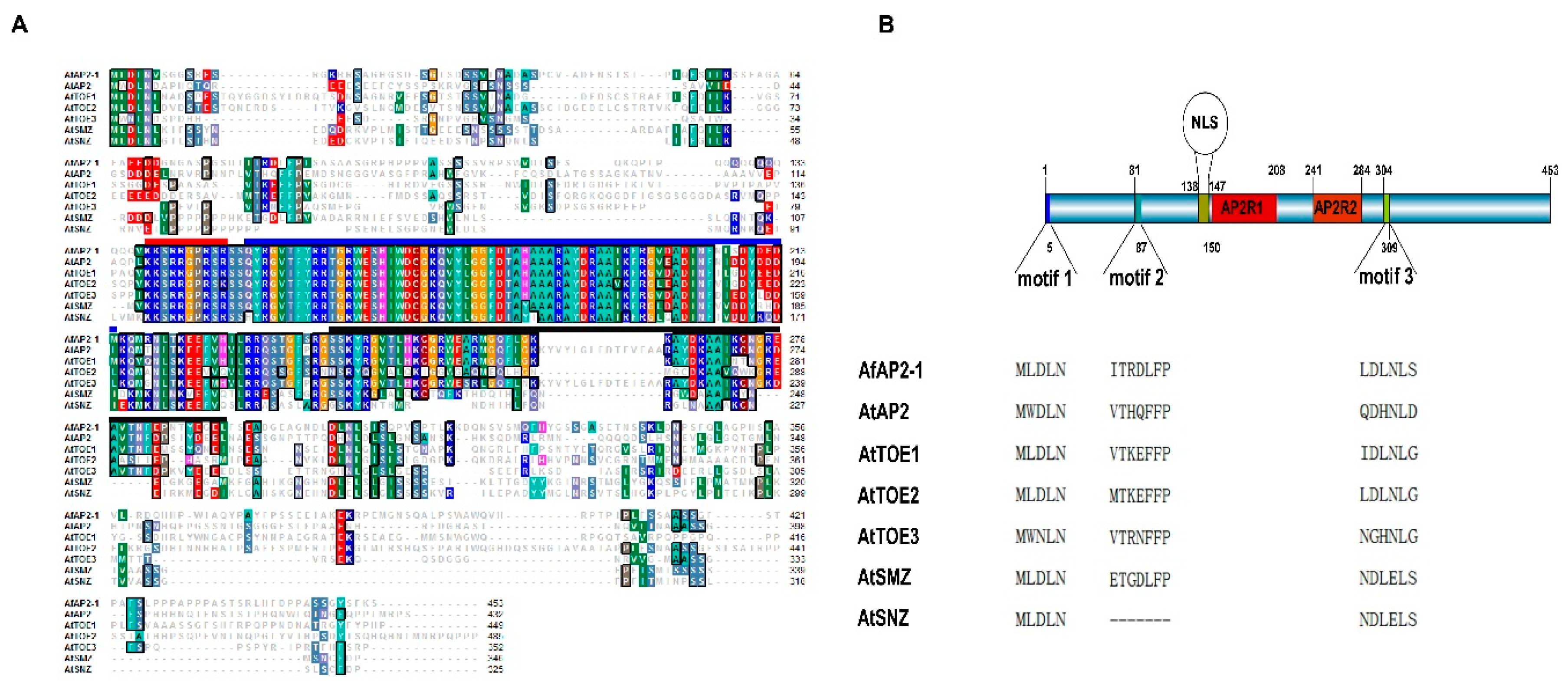
2.1. Isolation and Sequence Analysis of AfAP2-1
2.2. Transcript Profiling of AfAP2-1 in A. fasciata
2.3. Relative Expression of AfAP2-1 under the Control of the Circadian Rhythm
2.4. Response to Exogenous Ethylene Treatment
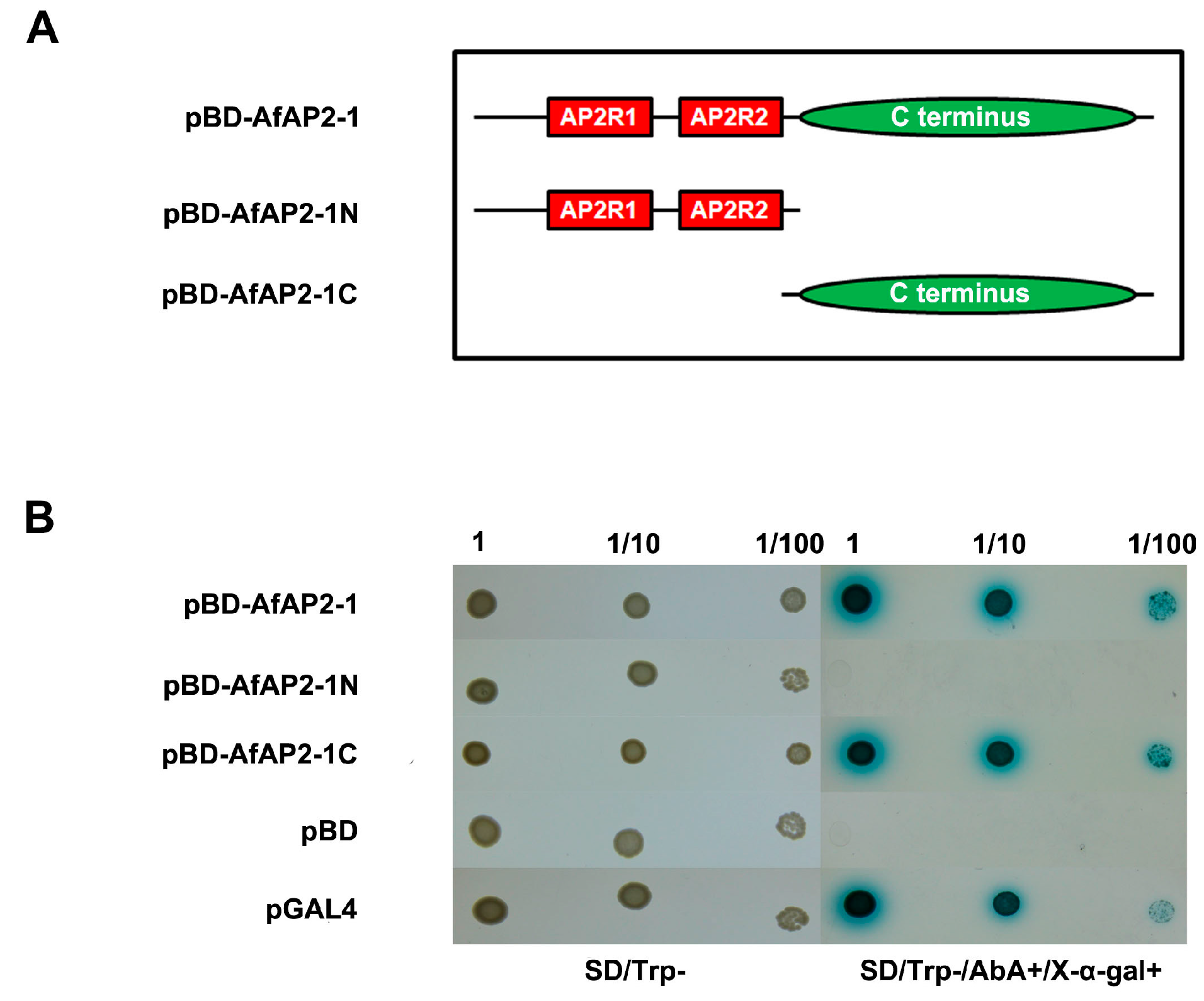
2.5. AfAP2-1 Exhibits Transactivation Activity in Yeast
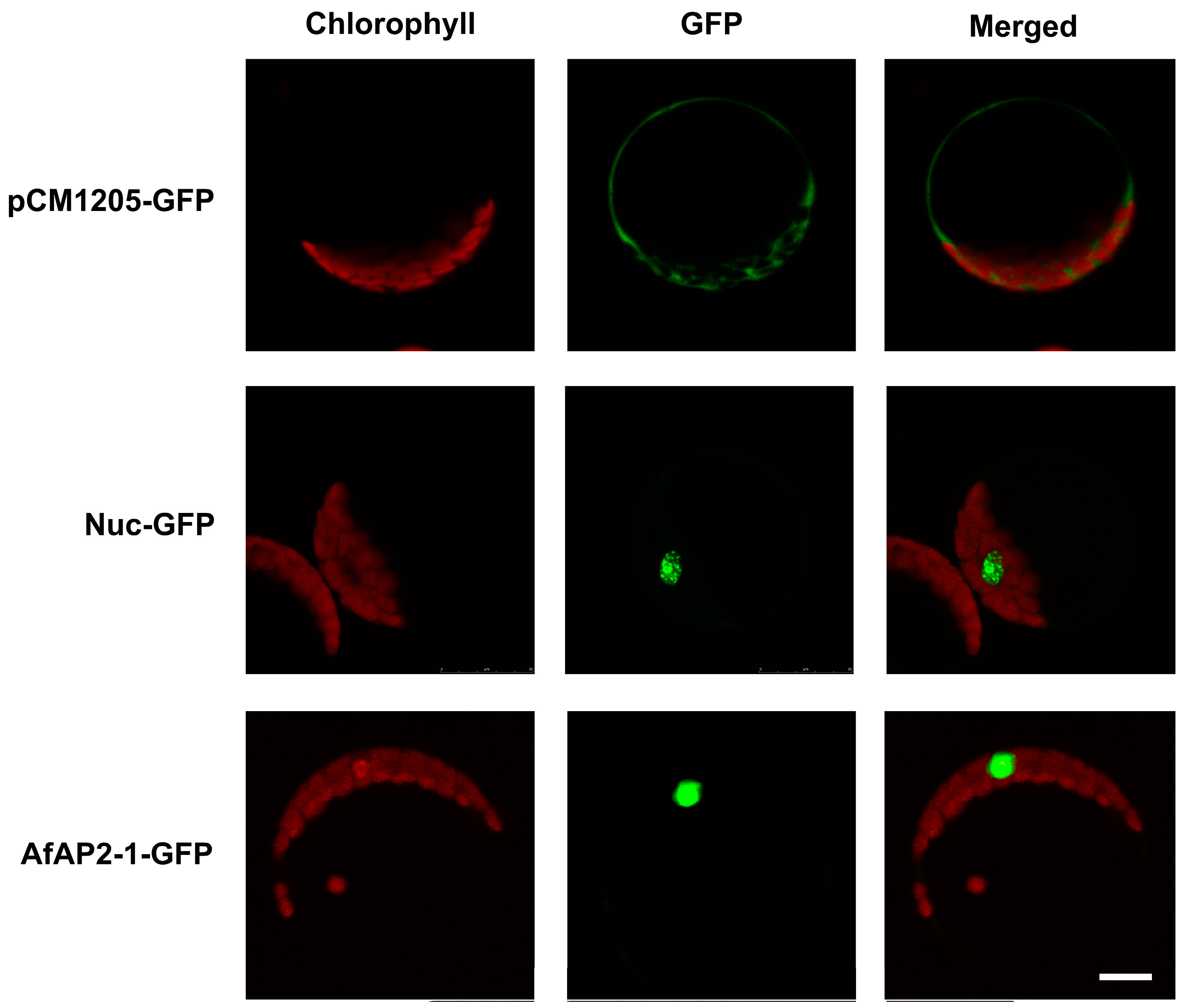
2.6. Subcellular Localization of AfAP2-1 Protein
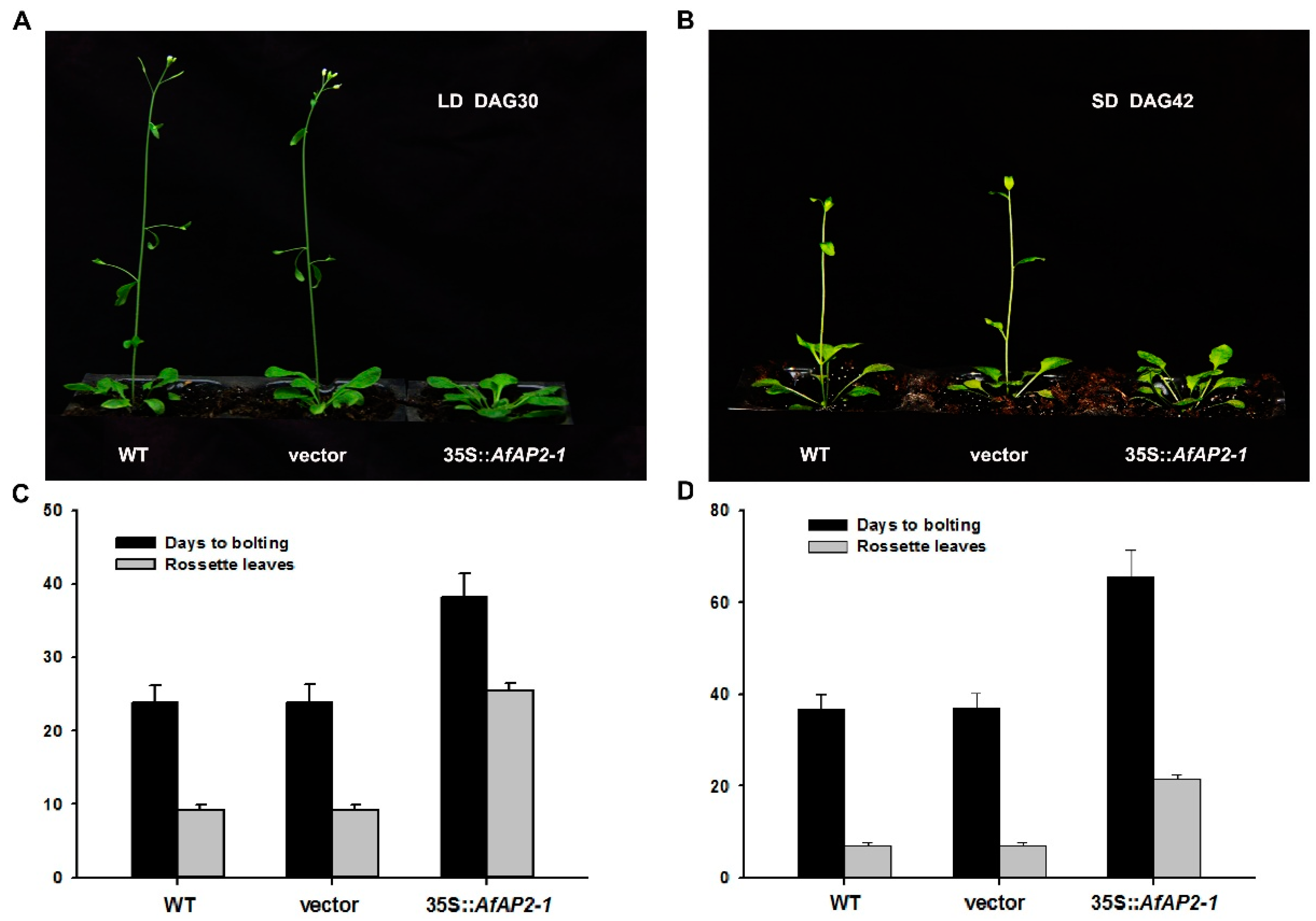
2.7. Constitutive Overexpression of AfAP2-1 in Arabidopsis Causes Significantly Delayed Flowering Phenotype
3. Discussion
3.1. Overexpression of AfAP2-1 in Arabidopsis Strongly Delayed Flowering under both LD and SD Conditions
3.2. The Abundance of AfAP2-1 Transcripts Is Negatively Regulated by Exogenous Ethylene Treatment
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. Isolation and Sequencing of the AfAP2-1 Gene
4.3. Bioinformatic Analysis
4.4. qRT-PCR Analysis
4.5. Plant Expression Vector Construction and Arabidopsis Transformation
4.6. Transactivation Analysis of AfAP2-1 in Yeast Cells
4.7. Subcellular Localization of AfAP2-1 Protein
4.8. Exogenous Ethylene Treatment of A. fasciata
4.9. Data Analysis
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
References
- Luther, H.E. An Alphabetical List of Bromeliad Binomials; The Bromeliad Society International: Sarasota, FL, USA, 2000; p. 4. [Google Scholar]
- Versieux, L.M.; Barbará, T.; das Graças Lapa Wanderley, M.D.; Calvente, A.; Fay, M.F.; Lexer, C. Molecular phylogenetics of the Brazilian giant bromeliads (Alcantarea, Bromeliaceae): Implications for morphological evolution and biogeography. Mol. Phylogenet. Evol. 2012, 64, 177–189. [Google Scholar] [CrossRef] [PubMed]
- Givnish, T.J.; Barfuss, M.H.J.; Ee, B.V.; Riina, R.; Schulte, K. Phylogeny, adaptive radiation, and historical biogeography in Bromeliaceae: Insights from an Eight-Locus Plastid Phylogeny. Am. J. Bot. 2011, 98, 872–895. [Google Scholar] [CrossRef] [PubMed]
- Clement, C.R.; de Cristo-Araújo, M.; Coppens D’Eeckenbrugge, G.; Pereira, A.A.; Picanço-Rodrigues, D. Origin and domestication of native Amazonian crops. Diversity 2010, 2, 72–106. [Google Scholar] [CrossRef]
- Ming, R.; VanBuren, R.; Wai, C.M.; Tang, H.B.; Schatz, M.C.; Bowers, J.E.; Lyons, E.; Wang, M.L.; Chen, J.; Biggers, E.; et al. The pineapple genome and the evolution of CAM photosynthesis. Nat. Genet. 2015, 47, 1435–1442. [Google Scholar] [CrossRef] [PubMed]
- Neuteboom, L.W.; Kunimitsu, W.Y.; Webb, D.; Christopher, D.A. Characterization and tissue-regulated expression of genes involved in pineapple (Ananas. comosus L.) root development. Plant Sci. 2002, 163, 1021–1035. [Google Scholar] [CrossRef]
- Zanella, C.M.; Janke, A.; Palma-Silva, C.; Kaltchuk-Santos, E.; Pinheiro, F.G.; Paggi, G.M.; Soares, L.E.S.; Goetze, M.; Büttow, M.V.; Bered, F. Genetics, evolution and conservation of Bromeliaceae. Genet. Mol. Biol. 2012, 35, 1020–1026. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.W.; Czech, B.; Weigel, D. miR156-Regulated SPL transcription factors define an endogenous flowering pathway in Arabidopsis thaliana. Cell 2009, 138, 738–749. [Google Scholar] [CrossRef] [PubMed]
- Srikanth, A.; Schmid, M. Regulation of flowering time: All roads lead to Rome. Cell. Mol. Life Sci. 2011, 68, 2013–2037. [Google Scholar] [CrossRef] [PubMed]
- Jofuku, K.D.; den Boer, B.G.; van Montagu, M.; Okamuro, J.K. Control of Arabidopsis flower and seed development by the homeotic gene APETALA2. Plant Cell 1994, 6, 1211–1225. [Google Scholar] [CrossRef] [PubMed]
- Jung, J.H.; Seo, Y.H.; Seo, P.J.; Reyes, J.L.; Yun, J.; Chua, N.H.; Park, C.M. The GIGANTEA-regulated microRNA172 mediates photoperiodic flowering independent of CONSTANS in Arabidopsis. Plant Cell 2007, 19, 2736–2748. [Google Scholar] [CrossRef] [PubMed]
- Mathieu, J.; Yant, L.J.; Mürdter, F.; Küttner, F.; Schmid, M. Repression of flowering by the miR172 target SMZ. PLoS Biol. 2009, 7, e1000148. [Google Scholar] [CrossRef] [PubMed]
- Zhang, B.; Wang, L.; Zeng, L.; Zhang, C.; Ma, H. Arabidopsis TOE proteins convey a photoperiodic signal to antagonize CONSTANS and regulate flowering time. Genes Dev. 2015, 29, 975–987. [Google Scholar] [CrossRef] [PubMed]
- Aukerman, M.J.; Sakai, H. Regulation of flowering time and floral organ identity by a microRNA and its APETALA2-like target genes. Plant Cell 2003, 15, 2730–2741. [Google Scholar] [CrossRef] [PubMed]
- Chen, X. A microRNA as a translational repressor of APETALA2 in Arabidopsis flower development. Science 2004, 303, 2022–2025. [Google Scholar] [CrossRef] [PubMed]
- Achard, P.; Herr, A.; Baulcombe, D.C.; Harberd, N.P. Modulation of floral development by a gibberellin-regulated microRNA. Development 2004, 131, 3357–3365. [Google Scholar] [CrossRef] [PubMed]
- Schwab, R.; Palatnik, J.F.; Riester, M.; Schommer, C.; Schmid, M.; Weigel, D. Specific effects of microRNAs on the plant transcriptome. Dev. Cell 2005, 8, 517–527. [Google Scholar] [CrossRef] [PubMed]
- Yant, L.; Mathieu, J.; Dinh, T.T.; Ott, F.; Lanz, C.; Wollmann, H.; Chen, X.; Schmid, M. Orchestration of the floral transition and floral development in Arabidopsis by the bifunctional transcription factor APETALA2. Plant Cell 2010, 22, 2156–2170. [Google Scholar] [CrossRef] [PubMed]
- Ogawara, T.; Higashi, K.; Kamada, H.; Ezura, H. Ethylene advances the transition from vegetative growth to flowering in Arabidopsis thaliana. J. Plant Physiol. 2003, 160, 1335–1340. [Google Scholar] [CrossRef] [PubMed]
- Achard, P.; Baghour, M.; Chapple, A.; Hedden, P.; van der Straeten, D.; Genschik, P.; Moritz, T.; Harberd, N.P. The plant stress hormone ethylene controls floral transition via DELLA-dependent regulation of floral meristem-identity genes. Proc. Natl. Acad. Sci. USA 2007, 104, 6484–6489. [Google Scholar] [CrossRef] [PubMed]
- Switzenberg, J.A.; Little, H.A.; Hammar, S.A.; Grumet, R. Floral primordia-targeted ACS (1-aminocyclopropane-1-carboxylate synthase) expression in transgenic Cucumis melo implicates fine tuning of ethylene production mediating unisexual flower development. Planta 2014, 240, 797–808. [Google Scholar] [CrossRef] [PubMed]
- Achard, P.; Cheng, H.; de Grauwe, L.; Decat, J.; Schoutteten, H.; Moritz, T.; van der Straeten, D.; Peng, J.; Harberd, N.P. Integration of plant responses to environmentally activated phytohormonal signals. Science 2006, 311, 91–94. [Google Scholar] [CrossRef] [PubMed]
- Kuan, C.S.; Yu, C.W.; Lin, M.L.; Hsu, H.T. Foliar application of aviglycine reduces natural flowering in pineapple. Hortscience 2005, 40, 123–126. [Google Scholar]
- Wang, R.H.; Hsu, Y.M. Delaying natural flowering in pineapple through foliar application of aviglycine, an inhibitor of ethylene biosynthesis. Hortscience 2007, 42, 1188–1191. [Google Scholar]
- Trusov, Y.; Botella, J.R. Silencing of the ACC synthase gene ACACS2 causes delayed flowering in pineapple (Ananas. comosus (L.) Merr.). J. Exp. Bot. 2006, 57, 3953–3960. [Google Scholar] [CrossRef] [PubMed]
- Wang, K. Agrobacterium Protocols. In Methods in Molecular Biology, 3rd ed.; Walker, J.M., Ed.; Humana Press: New York, NY, USA, 2014; Volume 2, pp. 293–294. [Google Scholar]
- Wessler, S.R. Homing into the origin of the AP2 DNA binding domain. Trends Plant Sci. 2005, 10, 54–56. [Google Scholar] [CrossRef] [PubMed]
- Licausi, F.; Ohme-Takagi, M.; Perata, P. APETALA2/Ethylene responsive factor (AP2/ERF) transcription factors: Mediators of stress responses and developmental programs. New Phytol. 2013, 199, 639–649. [Google Scholar] [CrossRef] [PubMed]
- Shigyo, M.; Ito, M. Analysis of gymnosperm two-AP2-domain-containing genes. Dev. Genes Evol. 2004, 214, 105–114. [Google Scholar] [CrossRef] [PubMed]
- Vahala, T.; Oxelman, B.; von Arnold, S. Two APETALA2-like genes of Picea. abies are differentially expressed during development. J. Exp. Bot. 2001, 52, 1111–1115. [Google Scholar] [CrossRef] [PubMed]
- Tang, M.; Li, G.; Chen, M. The phylogeny and expression pattern of APETALA2-like genes in rice. J. Genet. Genom. 2007, 34, 930–938. [Google Scholar] [CrossRef]
- Okamuro, J.K.; Caster, B.; Villarroel, R.; van Montagu, M.; Jofuku, K.D. The AP2 domain of APETALA2 defines a large new family of DNA binding proteins in Arabidopsis. Proc. Natl. Acad. Sci. USA 1997, 94, 7076–7081. [Google Scholar] [CrossRef] [PubMed]
- Yan, X.; Zhang, L.; Chen, B.; Xiong, Z.; Chen, C.; Wang, L.; Yu, J.; Lu, C.; Wei, W. Functional identification and characterization of the Brassica napus transcription factor gene BnAP2, the ortholog of Arabidopsis thaliana APETALA2. PLoS ONE 2012, 7, e33890. [Google Scholar]
- Krishnaswamy, S.; Verma, S.; Rahman, M.H.; Kav, N.N.V. Functional characterization of four APETALA2-family genes (RAP2.6, RAP2.6L, DREB19 and DREB26) in Arabidopsis. Plant Mol. Biol. 2011, 75, 107–127. [Google Scholar] [CrossRef] [PubMed]
- Krogan, N.T.; Hogan, K.; Long, J.A. APETALA2 negatively regulates multiple floral organ identity genes in Arabidopsis by recruiting the co-repressor TOPLESS and the histone deacetylase HDA19. Development 2012, 139, 4180–4190. [Google Scholar] [CrossRef] [PubMed]
- Vogel, M.O.; Gomez-Perez, D.; Probst, N. Combinatorial signal integration by APETALA2/Ethylene response factor (ERF)-transcription factors and the involvement of AP2-2 in starvation response. Int. J. Mol. Sci. 2012, 13, 5933–5951. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Wang, J.; Zhang, X.; Lei, M.; Fu, Y.; Zhang, J.; Wang, Z.; Xu, L. Transcriptome sequencing determined flowering pathway genes in Aechmea. fasciata treated with ethylene. J. Plant. Growth Regul. 2015, 1–14. [Google Scholar] [CrossRef]
- Zhou, C.M.; Zhang, T.Q.; Wang, X.; Yu, S.; Lian, H.; Tang, H.; Feng, Z.Y.; Zozomova-Lihová, J.; Wang, J.W. Molecular basis of age-dependent vernalization in Cardamine flexuosa. Science 2013, 340, 1097–1100. [Google Scholar] [CrossRef] [PubMed]
- Cong, H.; Li, Z.; Xu, L. Characterizing developmental and inducible differentiation between juvenile and adult plants of Aechmea fasciata treated with ethylene by transcriptomic analysis. Plant Growth Regul. 2013, 69, 247–257. [Google Scholar] [CrossRef]
- The ProtParam tool. Avaliable online: http://web.expasy.org/protparam/ (accessed on 1 January 2016).
- ORF Finder. Avaliable online: http://www.ncbi.nlm.nih.gov/projects/gorf/ (accessed on 1 January 2016).
- BLAST. Avaliable online: http://blast.ncbi.nlm.nih.gov/Blast.cgi (accessed on 1 January 2016).
- Cluster Omega. Avaliable online: http://www.ebi.ac.uk/Tools/msa/clustalo/ (accessed on 1 January 2016).
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular evolutionary genetics analysis Version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCt method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Clough, S.J.; Bent, A.F. Floral dip: A simplified method for agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 1998, 16, 735–743. [Google Scholar] [CrossRef] [PubMed]


© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lei, M.; Li, Z.-Y.; Wang, J.-B.; Fu, Y.-L.; Ao, M.-F.; Xu, L. AfAP2-1, An Age-Dependent Gene of Aechmea fasciata, Responds to Exogenous Ethylene Treatment. Int. J. Mol. Sci. 2016, 17, 303. https://doi.org/10.3390/ijms17030303
Lei M, Li Z-Y, Wang J-B, Fu Y-L, Ao M-F, Xu L. AfAP2-1, An Age-Dependent Gene of Aechmea fasciata, Responds to Exogenous Ethylene Treatment. International Journal of Molecular Sciences. 2016; 17(3):303. https://doi.org/10.3390/ijms17030303
Chicago/Turabian StyleLei, Ming, Zhi-Ying Li, Jia-Bin Wang, Yun-Liu Fu, Meng-Fei Ao, and Li Xu. 2016. "AfAP2-1, An Age-Dependent Gene of Aechmea fasciata, Responds to Exogenous Ethylene Treatment" International Journal of Molecular Sciences 17, no. 3: 303. https://doi.org/10.3390/ijms17030303
APA StyleLei, M., Li, Z.-Y., Wang, J.-B., Fu, Y.-L., Ao, M.-F., & Xu, L. (2016). AfAP2-1, An Age-Dependent Gene of Aechmea fasciata, Responds to Exogenous Ethylene Treatment. International Journal of Molecular Sciences, 17(3), 303. https://doi.org/10.3390/ijms17030303




