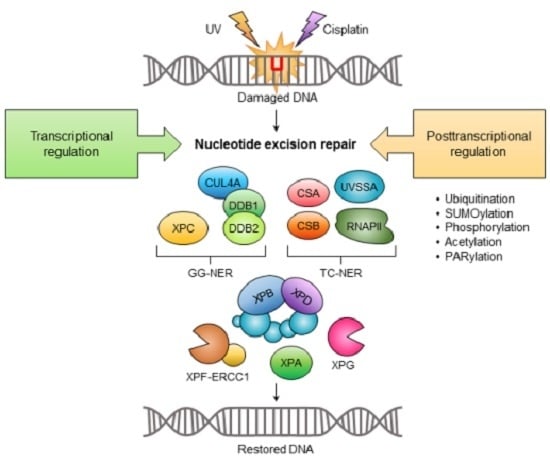
Transcriptional and Posttranslational Regulation of Nucleotide Excision Repair: The Guardian of the Genome against Ultraviolet Radiation
Abstract
:1. Introduction
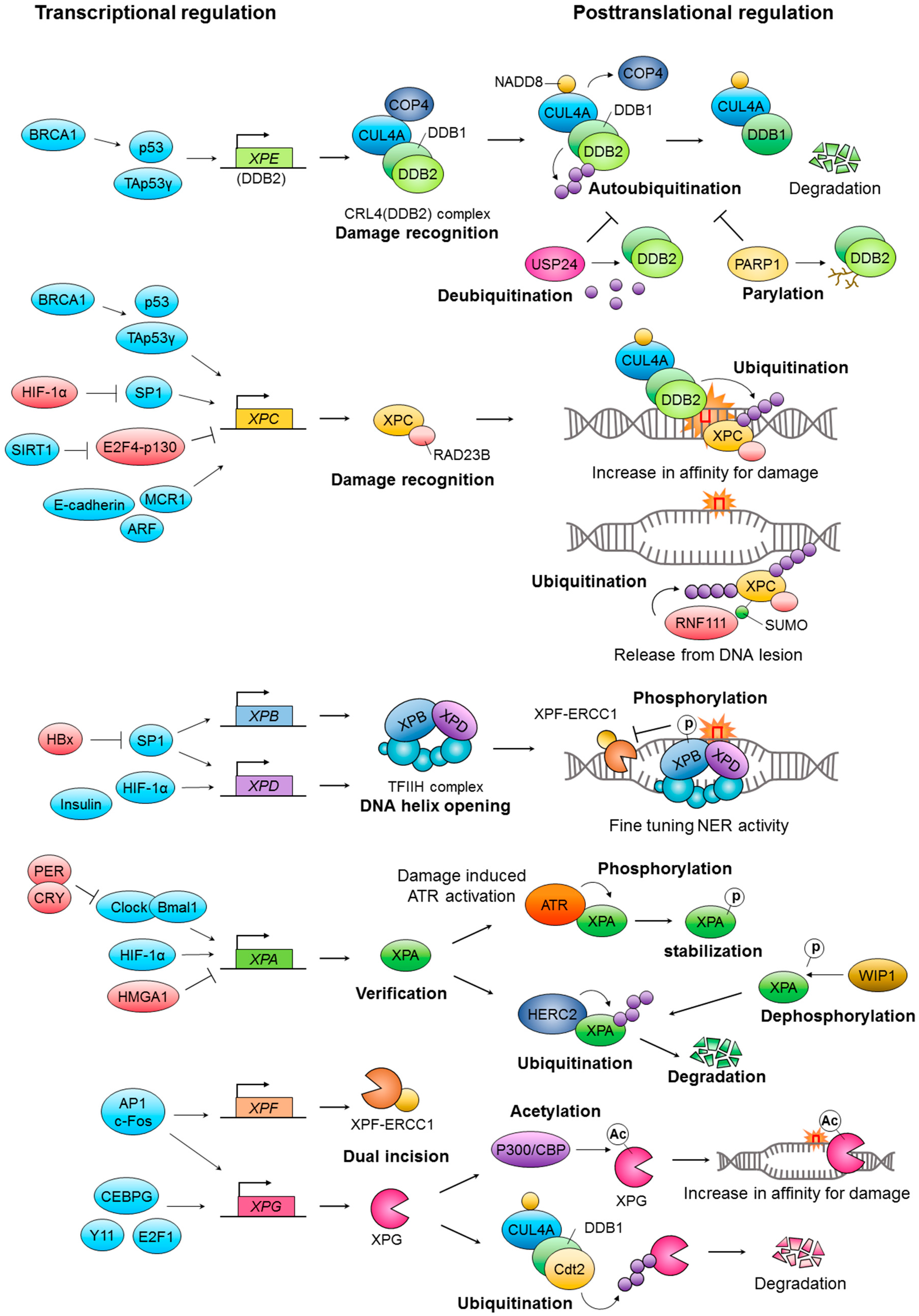
2. Transcriptional Regulation of Nucleotide Excision Repair (NER)
Transcriptional Regulation of Core NER Factors
3. Posttranslational Regulation of NER Factors
3.1. Ubiquitination and SUMOylation
3.2. Phosphorylation, Acetylation, and PARylation
4. Concluding Remark
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| 6-4PP | Pyrimidine-pyrimidone (6-4) photoproduct |
| ADP | Adenosine diphosphate |
| ALC1 | Amplified in liver cancer protein 1 |
| AP-1 | Activator protein 1 (also known as c-Jun) |
| ARF | Alternative reading frame |
| ATR | Ataxia-telangiectasia mutated and RAD3-related protein kinase |
| BRCA1 | Breast cancer 1 |
| c-Abl | Abelson murine leukemia viral oncogene homolog 1 |
| c-Fos | Proto-oncogene c-Fos |
| CBP | CREB-binding protein |
| CEBPG | CCAAT/enhancer-binding protein gamma |
| COP9 | Constitutive photomorphogenesis 9 |
| CPD | Cyclobutane pyrimidine dimer |
| CRL4 | Cullin 4A (CUL4A)–regulator of cullins 1 (ROC1) E3 ubiquitin ligase |
| Cry1 | Cryptochrome 1 |
| CSB | Cockayne syndrome protein B |
| CUL4A | Cullin 4A |
| DDB1 | DNA damage binding protein 1 |
| ERCC1 | Excision repair cross-complementation group 1 |
| GG-NER | Global genome repair |
| Gli1 | Glioma-associated oncogene homolog 1 |
| HBx | Hepatitis B virus x protein |
| HECT | Homologous to the E6-AP carboxyl terminus |
| HERC2 | HECT domain and RCC1-like domain-containing protein 2 |
| hHR23B | Human UV excision repair protein RAD23 homolog B |
| HIF-1α | Hypoxia-inducible factor-1α |
| HMGA1 | High-mobility group A 1 |
| HRE | Hypoxia response element |
| ISGylation | Protein conjugation by interferon-stimulated gene 15 |
| MAPK | Mitogen-activated protein kinase |
| MC1R | Melanocortin 1 receptor |
| NAD | Nicotinamide adenine dinucleotide |
| NDR1 | Nuclear-Dbf2-related family of Ser/Thr kinases 1 |
| NEDD8 | Neural precursor cell expressed developmentally down-regulated protein 8 |
| NER | Nucleotide excision repair |
| PARP | Poly(ADP ribose) polymerases |
| PARylation | Poly(ADP-ribosyl)ation |
| RASSF1A | Ras association domain-containing protein 1 |
| RLD | RCC1 like domain |
| RNAPII | RNA polymerase II |
| RNF111 | RING finger protein 111 |
| ROC1 | Regulator of cullins 1 |
| SIRT1 | Sirtuin 1 |
| Sp1 | Specificity protein 1 |
| SUMO | Small ubiquitin-related modifier |
| TAp63γ | Transactivation isoforms of p63γ |
| TC-NER | Transcription-coupled repair |
| TGF-β | Transforming growth factor-β |
| UBD | Ubiquitin-binding domain |
| USP | Ubiquitin-specific processing protease |
| UV | Ultraviolet |
| UVRAG | UV radiation resistance-associated gene protein |
| UVSSA | UV-sensitive syndrome A |
| WIP1 | Wild-type p53-induced phosphatase 1 |
| XPA | Xeroderma pigmentosum complementation group A |
| XPB | Xeroderma pigmentosum complementation group B (also known as ERCC3) |
| XPC | Xeroderma pigmentosum complementation group C |
| XPD | Xeroderma pigmentosum complementation group D (also known as ERCC2) |
| XPE | Xeroderma pigmentosum complementation group E (also known as DDB2) |
| XPF | Xeroderma pigmentosum complementation group F (also known as ERCC4) |
| XPG | Xeroderma pigmentosum complementation group G (also known as ERCC5) |
| YY1 | Yin Yang 1 |
References
- D’Orazio, J.; Jarrett, S.; Amaro-Ortiz, A.; Scott, T. UV radiation and the skin. Int. J. Mol. Sci. 2013, 14, 12222–12248. [Google Scholar] [CrossRef] [PubMed]
- Seebode, C.; Lehmann, J.; Emmert, S. Photocarcinogenesis and skin cancer prevention strategies. Anticancer Res. 2016, 36, 1371–1378. [Google Scholar] [PubMed]
- Hofbauer, G. Phototherapy and carcinogenesis. Hautarzt 2013, 64, 349–353. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, D.L.; Karentz, D. The Induction and Repair of DNA Photodamage in the Environment. In Environmental UV Photobiology; Young, A.R., Moan, J., Björn, L.O., Nultsch, W., Eds.; Springer: Boston, MA, USA, 1993; pp. 345–377. [Google Scholar]
- Latonen, L.; Laiho, M. Cellular UV damage responses—Functions of tumor suppressor p53. Biochim. Biophys. Acta-Rev. Cancer 2005, 1755, 71–89. [Google Scholar] [CrossRef] [PubMed]
- Daya-Grosjean, L.; Dumaz, N.; Sarasin, A. The specificity of p53 mutation spectra in sunlight induced human cancers. J. Photochem. Photobiol. B 1995, 28, 115–124. [Google Scholar] [CrossRef]
- Brash, D.E. UV signature mutations. Photochem. Photobiol. 2015, 91, 15–26. [Google Scholar] [CrossRef] [PubMed]
- Vrouwe, M.G.; Pines, A.; Overmeer, R.M.; Hanada, K.; Mullenders, L.H.F. UV-induced photolesions elicit ATR-kinase-dependent signaling in non-cycling cells through nucleotide excision repair-dependent and -independent pathways. J. Cell Sci. 2011, 124, 435–446. [Google Scholar] [CrossRef] [PubMed]
- Scharer, O.D. Nucleotide excision repair in eukaryotes. Cold Spring Harb. Perspect. Biol. 2013, 5, a012609. [Google Scholar] [CrossRef] [PubMed]
- Marteijn, J.A.; Lans, H.; Vermeulen, W.; Hoeijmakers, J.H.J. Understanding nucleotide excision repair and its roles in cancer and ageing. Nat. Rev. Mol. Cell Biol. 2014, 15, 465–481. [Google Scholar] [CrossRef] [PubMed]
- Hanawalt, P.C.; Spivak, G. Transcription-coupled DNA repair: Two decades of progress and surprises. Nat. Rev. Mol. Cell Biol. 2008, 9, 958–970. [Google Scholar] [CrossRef] [PubMed]
- Cleaver, J.E.; Lam, E.T.; Revet, I. Disorders of nucleotide excision repair: The genetic and molecular basis of heterogeneity. Nat. Rev. Genet. 2009, 10, 756–768. [Google Scholar] [CrossRef] [PubMed]
- De Boer, J.; Hoeijmakers, J.H.J. Nucleotide excision repair and human syndromes. Carcinogenesis 2000, 21, 453–460. [Google Scholar] [CrossRef] [PubMed]
- Batty, D.P.; Wood, R.D. Damage recognition in nucleotide excision repair of DNA. Gene 2000, 241, 193–204. [Google Scholar] [CrossRef]
- Kang, T.H.; Reardon, J.T.; Kemp, M.; Sancar, A. Circadian oscillation of nucleotide excision repair in mammalian brain. Proc. Natl. Acad. Sci. USA 2009, 106, 2864–2867. [Google Scholar] [CrossRef] [PubMed]
- Kang, T.H.; Lindsey-Boltz, L.A.; Reardon, J.T.; Sancar, A. Circadian control of XPA and excision repair of cisplatin-DNA damage by cryptochrome and HERC2 ubiquitin ligase. Proc. Natl. Acad. Sci. USA 2010, 107, 4890–4895. [Google Scholar] [CrossRef] [PubMed]
- Adair, J.E.; Maloney, S.C.; Dement, G.A.; Wertzler, K.J.; Smerdon, M.J.; Reeves, R. High-mobility group A1 proteins inhibit expression of nucleotide excision repair factor xeroderma pigmentosum group A. Cancer Res. 2007, 67, 6044–6052. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.B.; Bernauer, A.M.; Yingling, C.M.; Belinsky, S.A. HIF1α regulated expression of XPA contributes to cisplatin resistance in lung cancer. Carcinogenesis 2012, 33, 1187–1192. [Google Scholar] [CrossRef] [PubMed]
- Ma, L.; Weeda, G.; Jochemsen, A.G.; Bootsma, D.; Hoeijmakers, J.H.J.; van der Eb, A.J. Molecular and functional analysis of the XPBC/ERCC-3 promoter: Transcription activity is dependent on the integrity of an Sp1-binding site. Nucleic Acids Res. 1992, 20, 217–224. [Google Scholar] [CrossRef] [PubMed]
- Jaitovich-Groisman, I.; Benlimame, N.; Slagle, B.L.; Perez, M.H.; Alpert, L.; Song, D.J.; Fotouhi-Ardakani, N.; Galipeau, J.; Alaoui-Jamali, M.A. Transcriptional regulation of the TFIIH transcription repair components XPB and XPD by the hepatitis B virus x protein in liver cells and transgenic liver tissue. J. Biol. Chem. 2001, 276, 14124–14132. [Google Scholar] [PubMed]
- Adimoolam, S.; Ford, J.M. p53 and DNA damage-inducible expression of the xeroderma pigmentosum group C gene. Proc. Natl. Acad. Sci. USA 2002, 99, 12985–12990. [Google Scholar] [CrossRef] [PubMed]
- Barckhausen, C.; Roos, W.P.; Naumann, S.C.; Kaina, B. Malignant melanoma cells acquire resistance to DNA interstrand cross-linking chemotherapeutics by p53-triggered upregulation of DDB2/XPC-mediated DNA repair. Oncogene 2014, 33, 1964–1974. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Lin, M.H.; Zhang, C.; Wang, D.D.; Feng, Z.H.; Hu, W.W. TAp63γ enhances nucleotide excision repair through transcriptional regulation of DNA repair genes. DNA Repair 2012, 11, 167–176. [Google Scholar] [CrossRef] [PubMed]
- Hartman, A.R.; Ford, J.M. BRCA1 induces DNA damage recognition factors and enhances nucleotide excision repair. Nat. Genet. 2002, 32, 180–184. [Google Scholar] [CrossRef] [PubMed]
- Piao, M.J.; Hewage, S.; Han, X.; Kang, K.A.; Kang, H.K.; Lee, N.H.; Hyun, J.W. Protective effect of diphlorethohydroxycarmalol against ultraviolet B radiation-Induced DNA damage by inducing the nucleotide excision repair system in HaCaT human keratinocytes. Mar. Drugs 2015, 13, 5629–5641. [Google Scholar] [CrossRef] [PubMed]
- Rezvani, H.R.; Mahfouf, W.; Ali, N.; Chemin, C.; Ged, C.; Kim, A.L.; de Verneuil, H.; Taieb, A.; Bickers, D.R.; Mazurier, F. Hypoxia-inducible factor-1α regulates the expression of nucleotide excision repair proteins in keratinocytes. Nucleic Acids Res. 2010, 38, 797–809. [Google Scholar] [CrossRef] [PubMed]
- Ming, M.; Shea, C.R.; Guo, X.M.; Li, X.L.; Soltani, K.; Han, W.N.; He, Y.Y. Regulation of global genome nucleotide excision repair by SIRT1 through xeroderma pigmentosum C. Proc. Natl. Acad. Sci. USA 2010, 107, 22623–22628. [Google Scholar] [CrossRef] [PubMed]
- Dominguez-Brauer, C.; Chen, Y.J.; Brauer, P.M.; Pimkina, J.; Raychaudhuri, P. ARF stimulates XPC to trigger nucleotide excision repair by regulating the repressor complex of E2F4. EMBO Rep. 2009, 10, 1036–1042. [Google Scholar] [CrossRef] [PubMed]
- Qiang, L.; Shah, P.; Barcellos-Hoff, M.H.; He, Y.Y. TGF-β signaling links E-cadherin loss to suppression of nucleotide excision repair. Oncogene 2016, 35, 3293–3302. [Google Scholar] [CrossRef] [PubMed]
- Swope, V.; Alexander, C.; Starner, R.; Schwemberger, S.; Babcock, G.; Abdel-Malek, Z.A. Significance of the melanocortin 1 receptor in the DNA damage response of human melanocytes to ultraviolet radiation. Pigment Cell Melanoma Res. 2014, 27, 601–610. [Google Scholar] [CrossRef] [PubMed]
- Merkel, P.; Khoury, N.; Bertolotto, C.; Perfetti, R. Insulin and glucose regulate the expression of the DNA repair enzyme XPD. Mol. Cell. Endocrinol. 2003, 201, 75–85. [Google Scholar] [CrossRef]
- Tan, T.; Chu, G. p53 binds and activates the xeroderma pigmentosum DDB2 gene in humans but not mice. Mol. Cell. Biol. 2002, 22, 3247–3254. [Google Scholar] [CrossRef] [PubMed]
- Christmann, M.; Tomicic, M.T.; Origer, J.; Aasland, D.; Kaina, B. c-Fos is required for excision repair of UV-light induced DNA lesions by triggering the re-synthesis of XPF. Nucleic Acids Res. 2006, 34, 6530–6539. [Google Scholar] [CrossRef] [PubMed]
- Tomicic, M.T.; Reischmann, P.; Rasenberger, B.; Meise, R.; Kaina, B.; Christmann, M. Delayed c-Fos activation in human cells triggers XPF induction and an adaptive response to UVC-induced DNA damage and cytotoxicity. Cell. Mol. Life Sci. 2011, 68, 1785–1798. [Google Scholar] [CrossRef] [PubMed]
- Crawford, E.L.; Blomquist, T.; Mullins, D.N.; Yoon, Y.; Hernandez, D.R.; Al-Bagdhadi, M.; Ruiz, J.; Hammersley, J.; Willey, J.C. CEBPG regulates ERCC5/XPG expression in human bronchial epithelial cells and this regulation is modified by E2F1/YY1 interactions. Carcinogenesis 2007, 28, 2552–2559. [Google Scholar] [CrossRef] [PubMed]
- Kang, T.H.; Reardon, J.T.; Sancar, A. Regulation of nucleotide excision repair activity by transcriptional and post-transcriptional control of the XPA protein. Nucleic Acids Res. 2011, 39, 3176–3187. [Google Scholar] [CrossRef] [PubMed]
- Camenisch, U.; Dip, R.; Schumacher, S.B.; Schuler, B.; Naegeli, H. Recognition of helical kinks by xeroderma pigmentosum group A protein triggers DNA excision repair. Nat. Struct. Mol. Biol. 2006, 13, 278–284. [Google Scholar] [CrossRef] [PubMed]
- Orelli, B.; McClendon, T.B.; Tsodikov, O.V.; Ellenberger, T.; Niedernhofer, L.J.; Scharer, O.D. The XPA-binding domain of ERCC1 is required for nucleotide excision repair but not other DNA repair pathways. J. Biol. Chem. 2010, 285, 3705–3712. [Google Scholar] [CrossRef] [PubMed]
- Kang, T.H.; Sancar, A. Circadian regulation of DNA excision repair implications for chrono-chemotherapy. Cell Cycle 2009, 8, 1665–1667. [Google Scholar] [CrossRef] [PubMed]
- Reeves, R. High mobility group (HMG) proteins: Modulators of chromatin structure and DNA repair in mammalian cells. DNA Repair 2015, 36, 122–136. [Google Scholar] [CrossRef] [PubMed]
- Compe, E.; Egly, J.M. TFIIH: When transcription met DNA repair. Nat. Rev. Mol. Cell Biol. 2012, 13, 343–354. [Google Scholar] [CrossRef] [PubMed]
- Kaczynski, J.; Cook, T.; Urrutia, R. Sp1- and Kruppel-like transcription factors. Genome Biol. 2003, 4, 206. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.I.; Lee, S.; Lee, Y.; Bong, Y.S.; Hyun, S.W.; Yoo, Y.D.; Kim, S.J.; Kim, Y.W.; Poo, H.R. The human hepatitis B virus transactivator X gene product regulates Sp1 mediated transcription of an insulin-like growth factor II promoter 4. Oncogene 1998, 16, 2367–2380. [Google Scholar] [CrossRef] [PubMed]
- Sugasawa, K.; Okamoto, T.; Shimizu, Y.; Masutani, C.; Iwai, S.; Hanaoka, F. A multistep damage recognition mechanism for global genomic nucleotide excision repair. Genes Dev. 2001, 15, 507–521. [Google Scholar] [CrossRef] [PubMed]
- Min, J.H.; Pavletich, N.P. Recognition of DNA damage by the Rad4 nucleotide excision repair protein. Nature 2007, 449, 570–575. [Google Scholar] [CrossRef] [PubMed]
- Wakasugi, M.; Kawashima, A.; Morioka, H.; Linn, S.; Sancar, A.; Mori, T.; Nikaido, O.; Matsunaga, T. DDB accumulates at DNA damage sites immediately after UV irradiation and directly stimulates nucleotide excision repair. J. Biol. Chem. 2002, 277, 1637–1640. [Google Scholar] [CrossRef] [PubMed]
- Dotsch, V.; Bernassola, F.; Coutandin, D.; Candi, E.; Melino, G. p63 and p73, the ancestors of p53. Cold Spring Harb. Perspect. Biol. 2010, 2, a004887. [Google Scholar] [CrossRef] [PubMed]
- Fagbemi, A.F.; Orelli, B.; Scharer, O.D. Regulation of endonuclease activity in human nucleotide excision repair. DNA Repair 2011, 10, 722–729. [Google Scholar] [CrossRef] [PubMed]
- Christmann, M.; Kaina, B. Transcriptional regulation of human DNA repair genes following genotoxic stress: Trigger mechanisms, inducible responses and genotoxic adaptation. Nucleic Acids Res. 2013, 41, 8403–8420. [Google Scholar] [CrossRef] [PubMed]
- Youn, C.K.; Kim, M.H.; Cho, H.J.; Kim, H.B.; Chang, I.Y.; Chung, M.H.; You, H.J. Oncogenic H-Ras up-regulates expression of ERCC1 to protect cells from platinum-based anticancer agents. Cancer Res. 2004, 64, 4849–4857. [Google Scholar] [CrossRef] [PubMed]
- Kudo, K.; Gavin, E.; Das, S.; Amable, L.; Shevde, L.A.; Reed, E. Inhibition of Gli1 results in altered c-Jun activation, inhibition of cisplatin-induced upregulation of ERCC1, XPD and XRCC1, and inhibition of platinum-DNA adduct repair. Oncogene 2012, 31, 4718–4724. [Google Scholar] [CrossRef] [PubMed]
- Bergink, S.; Jentsch, S. Principles of ubiquitin and SUMO modifications in DNA repair. Nature 2009, 458, 461–467. [Google Scholar] [CrossRef] [PubMed]
- Sugasawa, K.; Okuda, Y.; Saijo, M.; Nishi, R.; Matsuda, N.; Chu, G.; Mori, T.; Iwai, S.; Tanaka, K.; Tanaka, K.; et al. UV-induced ubiquitylation of XPC protein mediated by UV-DDB-ubiquitin ligase complex. Cell 2005, 121, 387–400. [Google Scholar] [CrossRef] [PubMed]
- Fischer, E.S.; Scrima, A.; Bohm, K.; Matsumoto, S.; Lingaraju, G.M.; Faty, M.; Yasuda, T.; Cavadini, S.; Wakasugi, M.; Hanaoka, F.; et al. The molecular basis of CRL4DDB2/CSA ubiquitin ligase architecture, targeting, and activation. Cell 2011, 147, 1024–1039. [Google Scholar] [CrossRef] [PubMed]
- Hannah, J.; Zhou, P.B. Regulation of DNA damage response pathways by the cullin-RING ubiquitin ligases. DNA Repair 2009, 8, 536–543. [Google Scholar] [CrossRef] [PubMed]
- Scrima, A.; Konickova, R.; Czyzewski, B.K.; Kawasaki, Y.; Jeffrey, P.D.; Groisman, R.; Nakatani, Y.; Iwai, S.; Pavletich, N.P.; Thoma, N.H. Structural basis of UV DNA-damage recognition by the DDB1-DDB2 complex. Cell 2008, 135, 1213–1223. [Google Scholar] [CrossRef] [PubMed]
- Guerrero-Santoro, J.; Kapetanaki, M.G.; Hsieh, C.L.; Gorbachinsky, I.; Levine, A.S.; Rapic-Otrin, V. The cullin 4B-based UV-damaged DNA-binding protein ligase binds to UV-damaged chromatin and ubiquitinates histone H2A. Cancer Res. 2008, 68, 5014–5022. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Zhai, L.; Xu, J.; Joo, H.Y.; Jackson, S.; Erdjument-Bromage, H.; Tempst, P.; Xiong, Y.; Zhang, Y. Histone H3 and H4 ubiquitylation by the CUL4-DDB-ROC1 ubiquitin ligase facilitates cellular response to DNA damage. Mol. Cell 2006, 22, 383–394. [Google Scholar] [CrossRef] [PubMed]
- Takedachi, A.; Saijo, M.; Tanaka, K. DDB2 complex-mediated ubiquitylation around DNA damage is oppositely regulated by XPC and Ku and contributes to the recruitment of XPA. Mol. Cell. Biol. 2010, 30, 2708–2723. [Google Scholar] [CrossRef] [PubMed]
- Nag, A.; Bondar, T.; Shiv, S.; Raychaudhuri, P. The xeroderma pigmentosum group E gene product DDB2 is a specific target of cullin 4A in mammalian cells. Mol. Cell. Biol. 2001, 21, 6738–6747. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Lubin, A.; Chen, H.; Sun, Z.; Gong, F. The deubiquitinating protein USP24 interacts with DDB2 and regulates DDB2 stability. Cell Cycle 2012, 11, 4378–4384. [Google Scholar] [CrossRef] [PubMed]
- Puumalainen, M.R.; Ruthemann, P.; Min, J.H.; Naegeli, H. Xeroderma pigmentosum group C sensor: Unprecedented recognition strategy and tight spatiotemporal regulation. Cell. Mol. Life Sci. 2016, 73, 547–566. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Q.; Barakat, B.M.; Qin, S.; Ray, A.; El-Mahdy, M.A.; Wani, G.; Arafa, E.S.; Mir, S.N.; Wang, Q.E.; Wani, A.A. The p38 mitogen-activated protein kinase augments nucleotide excision repair by mediating DDB2 degradation and chromatin relaxation. J. Biol. Chem. 2008, 283, 32553–32561. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.F.; He, S.S.; Wang, Q.X.; Li, F.; Kwak, M.J.; Chen, S.; O’Connell, D.; Zhang, T.; Pirooz, S.D.; Jeon, Y.; et al. Autophagic UVRAG promotes UV-induced photolesion repair by activation of the CRL4(DDB2) E3 ligase. Mol. Cell 2016, 62, 507–519. [Google Scholar] [CrossRef] [PubMed]
- Van Cuijk, L.; van Belle, G.J.; Turkyilmaz, Y.; Poulsen, S.L.; Janssens, R.C.; Theil, A.F.; Sabatella, M.; Lans, H.; Mailand, N.; Houtsmuller, A.B.; et al. SUMO and ubiquitin-dependent XPC exchange drives nucleotide excision repair. Nat. Commun. 2015, 6, 7499. [Google Scholar] [CrossRef] [PubMed]
- Poulsen, S.L.; Hansen, R.K.; Wagner, S.A.; van Cuijk, L.; van Belle, G.J.; Streicher, W.; Wikstrom, M.; Choudhary, C.; Houtsmuller, A.B.; Marteijn, J.A.; et al. RNF111/Arkadia is a SUMO-targeted ubiquitin ligase that facilitates the DNA damage response. J. Cell Biol. 2013, 201, 797–807. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.E.; Zhu, Q.Z.; Wani, G.; El-Mahdy, M.A.; Li, J.Y.; Wani, A.A. DNA repair factor XPC is modified by SUMO-1 and ubiquitin following UV irradiation. Nucleic Acids Res. 2005, 33, 4023–4034. [Google Scholar] [CrossRef] [PubMed]
- Akita, M.; Tak, Y.S.; Shimura, T.; Matsumoto, S.; Okuda-Shimizu, Y.; Shimizu, Y.; Nishi, R.; Saitoh, H.; Iwai, S.; Mori, T.; et al. SUMOylation of xeroderma pigmentosum group C protein regulates DNA damage recognition during nucleotide excision repair. Sci. Rep. 2015, 5, 10984. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.E.; Praetorius-Ibba, M.; Zhu, Q.; El-Mahdy, M.A.; Wani, G.; Zhao, Q.; Qin, S.; Patnaik, S.; Wani, A.A. Ubiquitylation-independent degradation of Xeroderma pigmentosum group C protein is required for efficient nucleotide excision repair. Nucleic Acids Res. 2007, 35, 5338–5350. [Google Scholar] [CrossRef] [PubMed]
- Groisman, R.; Kuraoka, I.; Chevallier, O.; Gaye, N.; Magnaldo, T.; Tanaka, K.; Kisselev, A.F.; Harel-Bellan, A.; Nakatani, Y. CSA-dependent degradation of CSB by the ubiquitin-proteasome pathway establishes a link between complementation factors of the Cockayne syndrome. Genes Dev. 2006, 20, 1429–1434. [Google Scholar] [CrossRef] [PubMed]
- Anindya, R.; Mari, P.O.; Kristensen, U.; Kool, H.; Giglia-Mari, G.; Mullenders, L.H.; Fousteri, M.; Vermeulen, W.; Egly, J.M.; Svejstrup, J.Q. A ubiquitin-binding domain in cockayne syndrome B required for transcription-coupled nucleotide excision repair. Mol. Cell 2010, 38, 637–648. [Google Scholar] [CrossRef] [PubMed]
- Schwertman, P.; Lagarou, A.; Dekkers, D.H.; Raams, A.; van der Hoek, A.C.; Laffeber, C.; Hoeijmakers, J.H.; Demmers, J.A.; Fousteri, M.; Vermeulen, W.; et al. UV-sensitive syndrome protein UVSSA recruits USP7 to regulate transcription-coupled repair. Nat. Genet. 2012, 44, 598–602. [Google Scholar] [CrossRef] [PubMed]
- Sin, Y.; Tanaka, K.; Saijo, M. The C-terminal region and SUMOylation of cockayne syndrome group B protein play critical roles in transcription-coupled nucleotide excision repair. J. Biol. Chem. 2016, 291, 1387–1397. [Google Scholar] [CrossRef] [PubMed]
- Perez-Oliva, A.B.; Lachaud, C.; Szyniarowski, P.; Munoz, I.; Macartney, T.; Hickson, I.; Rouse, J.; Alessi, D.R. USP45 deubiquitylase controls ERCC1-XPF endonuclease-mediated DNA damage responses. EMBO J. 2015, 34, 326–343. [Google Scholar] [CrossRef] [PubMed]
- Han, C.H.; Wani, G.; Zhao, R.; Qian, J.; Sharma, N.; He, J.S.; Zhu, Q.Z.; Wang, Q.E.; Wani, A.A. Cdt2-mediated XPG degradation promotes gap-filling DNA synthesis in nucleotide excision repair. Cell Cycle 2015, 14, 1103–1115. [Google Scholar] [CrossRef] [PubMed]
- Shell, S.M.; Li, Z.K.; Shkriabai, N.; Kvaratskhelia, M.; Brosey, C.; Serrano, M.A.; Chazin, W.J.; Musich, P.R.; Zou, Y. Checkpoint kinase ATR promotes nucleotide excision repair of UV-induced DNA damage via physical interaction with xeroderma pigmentosum group A. J. Biol. Chem. 2009, 284, 24213–24222. [Google Scholar] [CrossRef] [PubMed]
- Lee, T.H.; Park, J.M.; Leem, S.H.; Kang, T.H. Coordinated regulation of XPA stability by ATR and HERC2 during nucleotide excision repair. Oncogene 2014, 33, 19–25. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, T.A.; Slattery, S.D.; Moon, S.H.; Darlington, Y.F.; Lu, X.B.; Donehower, L.A. The oncogenic phosphatase WIP1 negatively regulates nucleotide excision repair. DNA Repair 2010, 9, 813–823. [Google Scholar] [CrossRef] [PubMed]
- Fan, W.; Luo, J.Y. SIRT1 regulates UV-induced DNA repair through deacetylating XPA. Mol. Cell 2010, 39, 247–258. [Google Scholar] [CrossRef] [PubMed]
- Coin, F.; Auriol, J.; Tapias, A.; Clivio, P.; Vermeulen, W.; Egly, J.M. Phosphorylation of XPB helicase regulates TFIIH nucleotide excision repair activity. EMBO J. 2004, 23, 4835–4846. [Google Scholar] [CrossRef] [PubMed]
- Takeuchi, T.; Inoue, S.; Yokosawa, H. Identification and Herc5-mediated ISGylation of novel target proteins. Biochem. Biophys. Res. Commun. 2006, 348, 473–477. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.S.; Hahn, Y. Gains of ubiquitylation sites in highly conserved proteins in the human lineage. BMC Bioinform. 2012, 13, 306. [Google Scholar] [CrossRef] [PubMed]
- Pines, A.; Vrouwe, M.G.; Marteijn, J.A.; Typas, D.; Luijsterburg, M.S.; Cansoy, M.; Hensbergen, P.; Deelder, A.; de Groot, A.; Matsumoto, S.; et al. PARP1 promotes nucleotide excision repair through DDB2 stabilization and recruitment of ALC1. J. Cell Biol. 2012, 199, 235–249. [Google Scholar] [CrossRef] [PubMed]
- Robu, M.; Shah, R.G.; Petitclerc, N.; Brind′Amour, J.; Kandan-Kulangara, F.; Shah, G.M. Role of poly(ADP-ribose) polymerase-1 in the removal of UV-induced DNA lesions by nucleotide excision repair. Proc. Natl. Acad. Sci. USA 2013, 110, 1658–1663. [Google Scholar] [CrossRef] [PubMed]
- Tillhon, M.; Cazzalini, O.; Nardo, T.; Necchi, D.; Sommatis, S.; Stivala, L.A.; Scovassi, A.I.; Prosperi, E. p300/CBP acetyl transferases interact with and acetylate the nucleotide excision repair factor XPG. DNA Repair 2012, 11, 844–852. [Google Scholar] [CrossRef] [PubMed]
- Imam, S.Z.; Indig, F.E.; Cheng, W.H.; Saxena, S.P.; Stevnsner, T.; Kufe, D.; Bohr, V.A. Cockayne syndrome protein B interacts with and is phosphorylated by c-Abl tyrosine kinase. Nucleic Acids Res. 2007, 35, 4941–4951. [Google Scholar] [CrossRef] [PubMed]
- Thorslund, T.; von Kobbe, C.; Harrigan, J.A.; Indig, F.E.; Christiansen, M.; Stevnsner, T.; Bohr, V.A. Cooperation of the Cockayne syndrome group B protein and poly(ADP-ribose) polymerase 1 in the response to oxidative stress. Mol. Cell. Biol. 2005, 25, 7625–7636. [Google Scholar] [CrossRef] [PubMed]
- Cimprich, K.A.; Cortez, D. ATR: An essential regulator of genome integrity. Nat. Rev. Mol. Cell Biol. 2008, 9, 616–627. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.K.; Musich, P.R.; Cartwright, B.M.; Wang, H.; Zou, Y. UV-induced nuclear import of XPA is mediated by importin-α4 in an ATR-dependent manner. PLoS ONE 2013, 8, e68297. [Google Scholar] [CrossRef] [PubMed]
- Park, J.M.; Choi, J.Y.; Yi, J.M.; Chung, J.W.; Leem, S.H.; Koh, S.S.; Kang, T.H. NDR1 modulates the UV-induced DNA-damage checkpoint and nucleotide excision repair. Biochem. Biophys. Res. Commun. 2015, 461, 543–548. [Google Scholar] [CrossRef] [PubMed]
- Guo, R.F.; Chen, J.; Zhu, F.; Biswas, A.K.; Berton, T.R.; Mitchell, D.L.; Johnson, D.G. E2F1 localizes to sites of UV-induced DNA damage to enhance nucleotide excision repair. J. Biol. Chem. 2010, 285, 19308–19315. [Google Scholar] [CrossRef] [PubMed]
- Biswas, A.K.; Mitchell, D.L.; Johnson, D.G. E2F1 responds to ultraviolet radiation by directly stimulating DNA repair and suppressing carcinogenesis. Cancer Res. 2014, 74, 3369–3377. [Google Scholar] [CrossRef] [PubMed]
- Guo, R.F.; Chen, J.; Mitchell, D.L.; Johnson, D.G. GCN5 and E2F1 stimulate nucleotide excision repair by promoting H3K9 acetylation at sites of damage. Nucleic Acids Res. 2011, 39, 1390–1397. [Google Scholar] [CrossRef] [PubMed]
- Lake, R.J.; Fan, H.Y. Structure, function and regulation of CSB: A multi-talented gymnast. Mech. Ageing Dev. 2013, 134, 202–211. [Google Scholar] [CrossRef] [PubMed]
- Matsuoka, S.; Ballif, B.A.; Smogorzewska, A.; McDonald, E.R.; Hurov, K.E.; Luo, J.; Bakalarski, C.E.; Zhao, Z.M.; Solimini, N.; Lerenthal, Y.; et al. ATM and ATR substrate analysis reveals extensive protein networks responsive to DNA damage. Science 2007, 316, 1160–1166. [Google Scholar] [CrossRef] [PubMed]
- Choi, J.Y.; Park, J.M.; Yi, J.M.; Leem, S.H.; Kang, T.H. Enhanced nucleotide excision repair capacity in lung cancer cells by preconditioning with DNA-damaging agents. Oncotarget 2015, 6, 22575–22586. [Google Scholar] [CrossRef] [PubMed]
- Donninger, H.; Clark, J.; Rinaldo, F.; Nelson, N.; Barnoud, T.; Schmidt, M.L.; Hobbing, K.R.; Vos, M.D.; Sils, B.; Clark, G.J. The RASSF1A tumor suppressor regulates XPA-mediated DNA repair. Mol. Cell. Biol. 2015, 35, 277–287. [Google Scholar] [CrossRef] [PubMed]
- Feltes, B.C.; Bonatto, D. Overview of xeroderma pigmentosum proteins architecture, mutations and post-translational modifications. Mutat. Res. Rev. Mutat. Res. 2015, 763, 306–320. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.E.; Han, C.H.; Zhao, R.; Wani, G.; Zhu, Q.Z.; Gong, L.; Battu, A.; Racoma, I.; Sharma, N.; Wani, A.A. p38 MAPK- and Akt-mediated p300 phosphorylation regulates its degradation to facilitate nucleotide excision repair. Nucleic Acids Res. 2013, 41, 1722–1733. [Google Scholar] [CrossRef] [PubMed]
- Huang, W.C.; Chen, C.C. Akt-phosphorylation of p300 at Ser-1834 is essential for its histone acetyltransferase and transcriptional activity. Mol. Cell. Biol. 2005, 25, 6592–6602. [Google Scholar] [CrossRef] [PubMed]
- Ame, J.C.; Spenlehauer, C.; de Murcia, G. The PARP superfamily. Bioessays 2004, 26, 882–893. [Google Scholar] [CrossRef] [PubMed]
- Rouleau, M.; Patel, A.; Hendzel, M.J.; Kaufmann, S.H.; Poirier, G.G. PARP inhibition: PARP1 and beyond. Nat. Rev. Cancer 2010, 10, 293–301. [Google Scholar] [CrossRef] [PubMed]
- Choi, J.Y.; Joh, H.M.; Park, J.M.; Kim, M.J.; Chung, T.H.; Kang, T.H. Non-thermal plasma-induced apoptosis is modulated by ATR-and PARP1-mediated DNA damage responses and circadian clock. Oncotarget 2016, 7, 32980–32989. [Google Scholar] [CrossRef] [PubMed]
- Pines, A.; Mullenders, L.H.; van Attikum, H.; Luijsterburg, M.S. Touching base with PARPs: Moonlighting in the repair of UV lesions and double-strand breaks. Trends Biochem. Sci. 2013, 38, 321–330. [Google Scholar] [CrossRef] [PubMed]
- Fahrer, J.; Kranaster, R.; Altmeyer, M.; Marx, A.; Burkle, A. Quantitative analysis of the binding affinity of poly(ADP-ribose) to specific binding proteins as a function of chain length. Nucleic Acids Res. 2007, 35, e143. [Google Scholar] [CrossRef] [PubMed]
- Fischer, J.M.F.; Popp, O.; Gebhard, D.; Veith, S.; Fischbach, A.; Beneke, S.; Leitenstorfer, A.; Bergemann, J.; Scheffner, M.; Ferrando-May, E.; et al. Poly(ADP-ribose)-mediated interplay of XPA and PARP1 leads to reciprocal regulation of protein function. FEBS J. 2014, 281, 3625–3641. [Google Scholar] [CrossRef] [PubMed]
- King, B.S.; Cooper, K.L.; Liu, K.J.; Hudson, L.G. Poly(ADP-ribose) contributes to an association between poly(ADP-ribose) polymerase-1 and xeroderma pigmentosum complementation group A in nucleotide excision repair. J. Biol. Chem. 2012, 287, 10. [Google Scholar] [CrossRef] [PubMed]
- Perera, D.; Poulos, R.C.; Shah, A.; Beck, D.; Pimanda, J.E.; Wong, J.W.H. Differential DNA repair underlies mutation hotspots at active promoters in cancer genomes. Nature 2016, 532, 259–263. [Google Scholar] [CrossRef] [PubMed]
- Sabarinathan, R.; Mularoni, L.; Deu-Pons, J.; Gonzalez-Perez, A.; Lopez-Bigas, N. Nucleotide excision repair is impaired by binding of transcription factors to DNA. Nature 2016, 532, 264–267. [Google Scholar] [CrossRef] [PubMed]
| XP 1 | Function in NER | Transcriptional Regulation | ||
|---|---|---|---|---|
| Factor | Regulation | Reference | ||
| XPA | Damage verification | Cry1 | Down | [15,16] |
| HMGA1 | Down | [17] | ||
| HIF-1α | Up | [18] | ||
| XPB | ATP-dependent helicase | Sp1 | Up | [19] |
| HBx | Down | [20] | ||
| XPC | Damage recognition | p53 | Up | [21,22] |
| TAp63γ | Up | [23] | ||
| BRCA1 | UP | [24] | ||
| Sp1 | Up | [25,26] | ||
| HIF-1α | Down | [26] | ||
| E2F4-p130 | Down | [27,28] | ||
| SIRT1 | UP | [27] | ||
| ARF | UP | [28] | ||
| E-cadherin | Up | [29] | ||
| MC1R | Up | [30] | ||
| XPD | ATP-dependent helicase | HBx | Down | [20] |
| HIF-1α | Up | [26] | ||
| Insulin | Up | [31] | ||
| XPE | Damage recognition | p53 | Up | [22,32] |
| TAp63γ | Up | [23] | ||
| BRCA1 | Up | [24] | ||
| XPF | 5′ incision endonuclease | c-Fos, AP-1 | Up | [33,34] |
| XPG | 3′ incision endonuclease | c-Fos, AP-1 | Up | [33] |
| CEBPG, E2F1, YY1 | UP | [35] | ||
| XP | Modification | Factor | Residue | Effect | Ref. |
|---|---|---|---|---|---|
| XPA | Phosphorylation | ATR | Ser196 | Stabilization | [76,77] |
| Dephosphorylation | WIP1 | Ser196 | Inactivation | [78] | |
| Ubiquitination | HERC2 | - | Degradation | [16,36] | |
| Deacetylation | SIRT1 | Lys63, Lys67 | Stabilization, Interaction with RPA32 | [79] | |
| XPB | Phosphorylation | - | Ser751 | Inhibition of 5′ incision by XPF/ERCC1 | [80] |
| XPC | Ubiquitination | CRL4(DDB2) | - | Affinity for DNA lesions | [53,59] |
| Ubiquitination | RNF111 | - | Release from DNA lesions | [66] | |
| SUMOylation | SUMO2 | - | RNF111 induced ubiquitination | [65,66] | |
| SUMOylation | SUMO1 | Lys81, Lys89, Lys183 | Damage recognition | [67,68] | |
| SUMOylation | - | Lys655 | Degradation | [69] | |
| Dephosphorylation | WIP1 | Ser892 | Inactivation | [78] | |
| XPD | ISGylation | HERC5 | - | Not investigated | [81] |
| Ubiquitination | - | Lys 701 | Possible ubiquitination site | [82] | |
| XPE (DDB2) | Ubiquitination | CRL4(DDB2) | Lys5, Lys11, Lys35, Lys40, Lys151 | Autoubiquitination, Degradation | [54,60] |
| Deubiquitination | USP24 | - | Stabilization | [61] | |
| PARylation | PARP1 | - | Stabilization | [83,84] | |
| XPG | Ubiquitination | CRL4(Cdt2) | Degradation | [75] | |
| Acetylation | p300/CBP | Affinity for DNA lesions | [85] | ||
| CSB | Ubiquitination | - | - | Degradation | [70] |
| Deubiquitination | USP7 | - | Stabilization | [72] | |
| SUMOylation | SUMO2/3 | Lys205 | Efficient TC-NER | [73] | |
| Phosphorylation | c-Abl | Tyr932 | Nuclear localization | [86] | |
| PARylation | PARP1 | - | Repair for oxidative DNA damage | [87] | |
| ERCC1 | Deubiquitination | USP45 | - | Access to damage site | [74] |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Park, J.-M.; Kang, T.-H. Transcriptional and Posttranslational Regulation of Nucleotide Excision Repair: The Guardian of the Genome against Ultraviolet Radiation. Int. J. Mol. Sci. 2016, 17, 1840. https://doi.org/10.3390/ijms17111840
Park J-M, Kang T-H. Transcriptional and Posttranslational Regulation of Nucleotide Excision Repair: The Guardian of the Genome against Ultraviolet Radiation. International Journal of Molecular Sciences. 2016; 17(11):1840. https://doi.org/10.3390/ijms17111840
Chicago/Turabian StylePark, Jeong-Min, and Tae-Hong Kang. 2016. "Transcriptional and Posttranslational Regulation of Nucleotide Excision Repair: The Guardian of the Genome against Ultraviolet Radiation" International Journal of Molecular Sciences 17, no. 11: 1840. https://doi.org/10.3390/ijms17111840
APA StylePark, J.-M., & Kang, T.-H. (2016). Transcriptional and Posttranslational Regulation of Nucleotide Excision Repair: The Guardian of the Genome against Ultraviolet Radiation. International Journal of Molecular Sciences, 17(11), 1840. https://doi.org/10.3390/ijms17111840