Milk—A Nutrient System of Mammalian Evolution Promoting mTORC1-Dependent Translation
Abstract
:1. Introduction
2. mTORC1: Key Regulator of Cell Growth and Proliferation
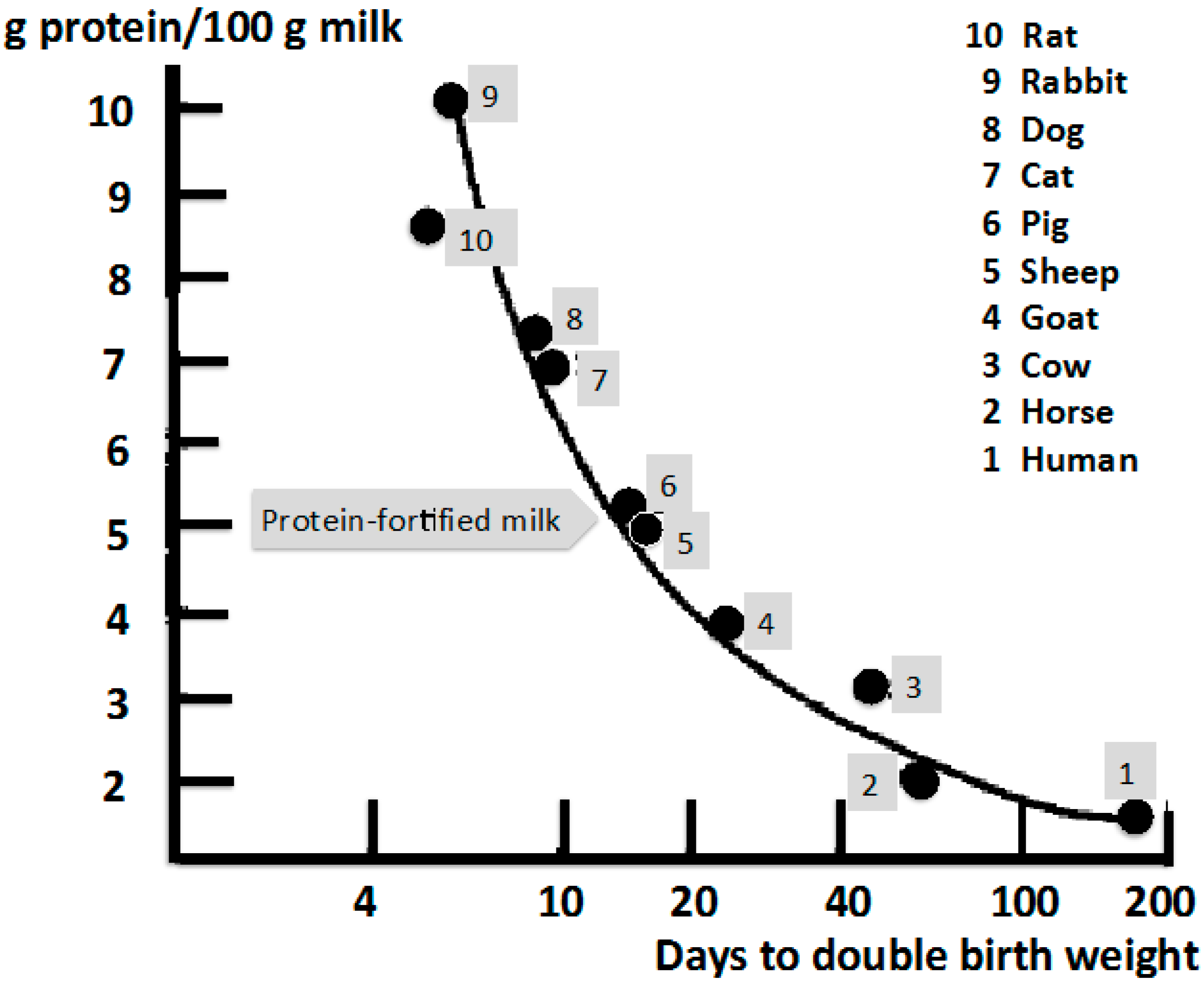
3. Amino Acids: Milk’s Hardware Activating mTORC1
| Species | Total Amino Acids * g/100 mL Whole Milk | BCAA mg/g Total Amino Acids * | Leucine mg/g Total Amino Acids * |
|---|---|---|---|
| Rat | 8.69 ± 0.77 | 176 ± 4 | 92 ± 2 |
| Cat | 7.57 ± 1.27 | 208 ± 3 | 118 ± 1 |
| Sheep | 5.41 ± 0.24 | 196 ± 5 | 90 ± 4 |
| Cow | 3.36 ± 0.48 | 199 ± 3 | 99 ± 1 |
| Horse | 1.58 ± 0.35 | 178 ± 3 | 93 ± 3 |
| Man | 0.85 ± 0.09 | 209 ± 5 | 104 ± 1 |
4. Milk Induces mTORC1 Activating Growth Hormones
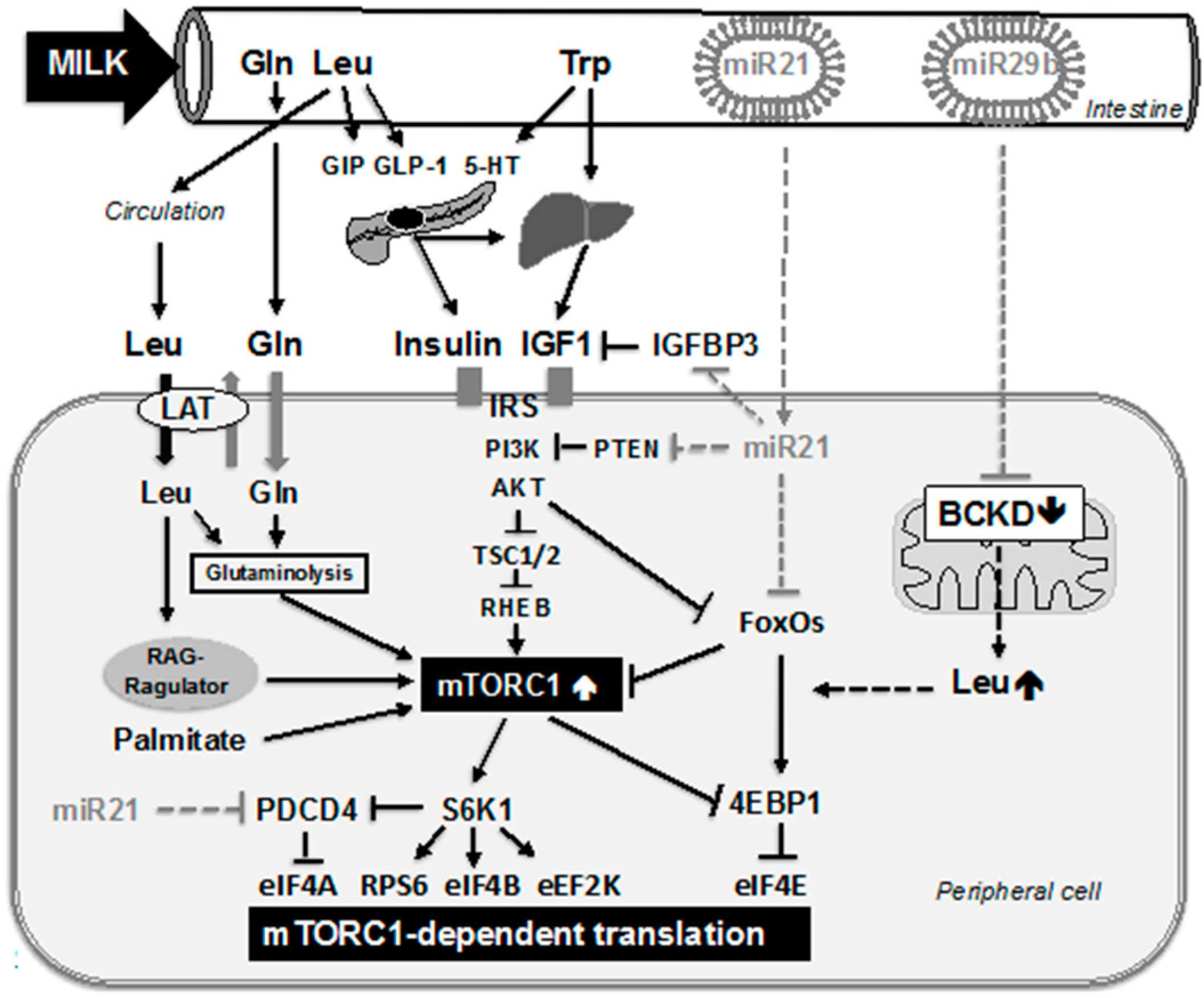
| Milk Amino Acid | Function | References |
|---|---|---|
| Leucine (Leu) | Leu stimulates intestinal production of GIP by K-cells augmenting insulin production. Insulin stimulates mTORC1 of peripheral cells of the body. | [72,75,76] |
| Leu stimulates intestinal production of GLP-1 by L-cells promoting the production of insulin activating mTORC1. GLP-1 stimulates islet cell DNA replication via activation of mTORC1 involving IGF-1 signaling. | [73,75,77] | |
| Leu stimulates insulin production of pancreatic β-cells. Insulin stimulates mTORC1 of peripheral cells. | [57,58,62,63] | |
| Leu allosterically activates GDH, the rate-limiting enzyme of glutaminolysis, which activates mTORC1. GDH contributes to Leu sensing in the regulation of autophagy. | [55,56] | |
| Glutamine (Gln) | Gln promotes cellular uptake of Leu that is the primary amino acid for mTORC1 activation. mTORC1 activation stimulates the uptake of Gln by positive regulation of glutaminase. | [52,53,91] |
| Gln is the precursor of the glutaminolysis pathway that activates mTORC1 and mTORC1-dependent insulin synthesis. | [44,54,57] | |
| Gln controls the activity of the β-cell IGF-2/IGF1R autocrine loop by increasing biosynthesis and secretion of IGF-2. | [90] | |
| Tryptophan (Trp) | Trp induces hepatic IGF1 gene and IGF-1 protein expression. Insulin co-stimulates hepatic IGF-1 secretion. | [81,82] |
| Trp via conversion to 5-HT enhances glucose-stimulated insulin secretion, thereby promotes mTORC1 activation. | [78,79,80] | |
| Arginine (Arg) | The amino acid transporter SLC38A9 is a key component of a lysosomal membrane complex that signals Arg sufficiency to mTORC1. | [60] |
5. Milk-Derived Palmitic Acid Activates mTORC1/S6K1 Signaling
6. Exosomal MicroRNAs: Milk’s Software Activating mTORC1
6.1. Milk MicroRNA-29b: Activator of mTORC1 by Attenuating BCAA Catabolism?
6.2. Milk MicroRNA-21 Enhances mTORC1-Dependent Translation
| Milk MicroRNA | Function | References |
|---|---|---|
| MicroRNA-29b | Targets the mRNA of dihydrolipoyl branched-chain acyltransferase, the core of the BCKD complex. This may attenuate BCAA catabolism increasing BCAA/leucine availability for mTORC1 activation. | [11,67,123] |
| MicroRNA-21 | Targets the mRNA of IGFBP3. Resulting increase in free IGF-1 may enhance IGF-1-mediated mTORC1 activation. | [143] |
| MicroRNA-21 | Targets the mRNA of PTEN, thus enhances the activation of AKT, which via phosphorylation of PRAS40 and TSC2 activates mTORC1 and inhibits nuclear FoxO activity. This leads to mTORC1 activation as FoxOs are negative regulators of mTORC1. | [130,131,132,133,144,145] |
| MicroRNA-21 | Targets the mRNA of Sprouty1 and -2 thereby increasing RAS-RAF-ERK signaling. ERK-mediated phosphorylation of TSC2 activates mTORC1. | [135,136,137] |
| MicroRNA-21 | Targets the mRNA of FoxO1, which downregulates the FoxO-promoted expression of the translational repressor 4E-BP-1. | [140,141,142,146,147] |
| MicroRNA-21 | Targets the mRNA of PDCD4, which is a suppressor of translation initiation inhibiting the RNA helicase eIF4A. | [149,150,151,152] |
| MicroRNA-21 | Is upregulated by TGFβ, a component of bovine milk exosomes. | [111,153] |
7. mTORC1: Key Regulator of Translation
8. Milk’s Hardware and Software Synergize in mTORC1-Driven Translation
9. Milk-Mediated Over-Activation of mTORC1 and Age-Related Diseases
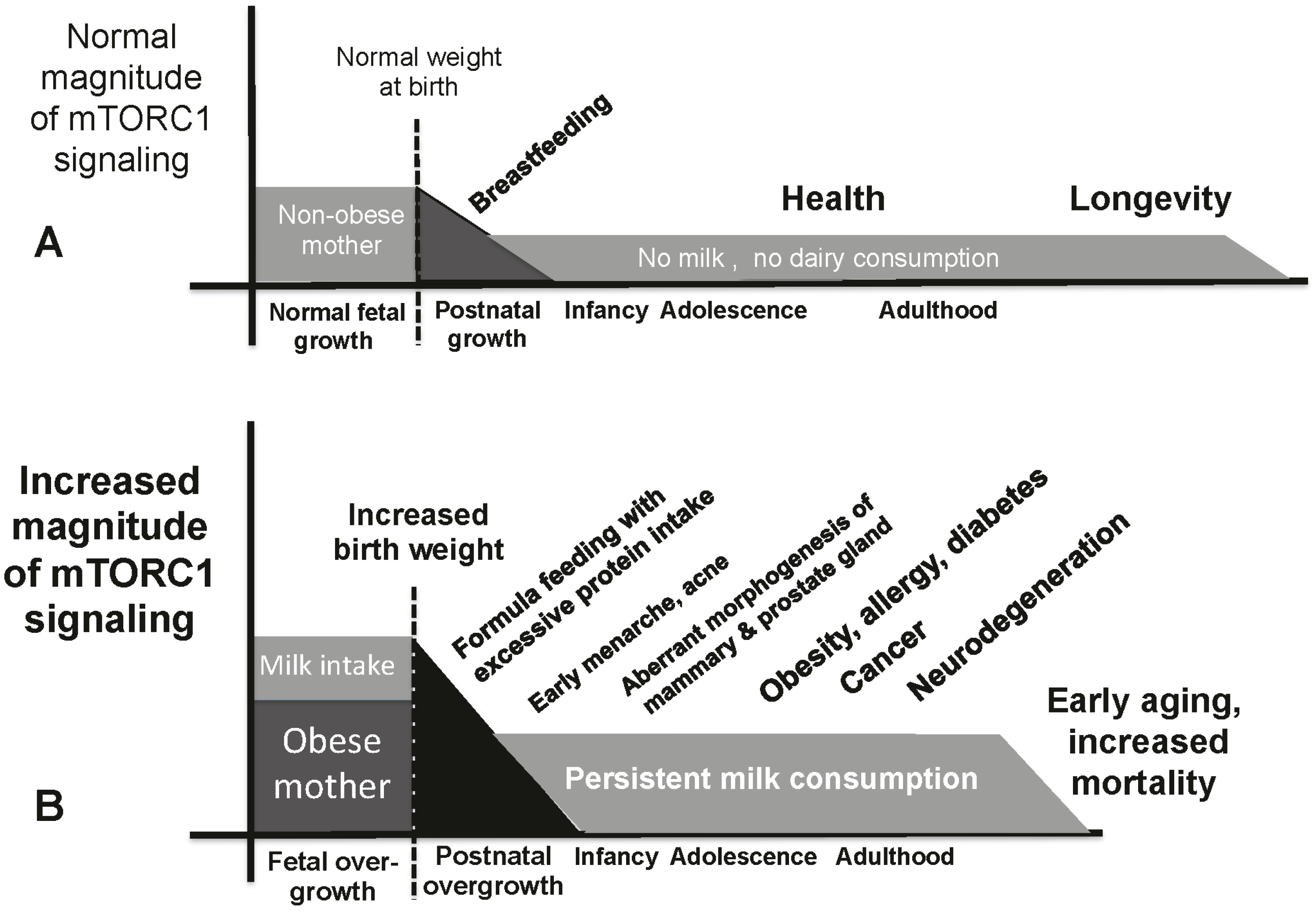
10. Conclusions
Acknowledgments
Conflicts of Interest
References
- Landecker, H. Food as exposure: Nutritional epigenetics and the new metabolism. Biosocieties 2011, 6, 167–194. [Google Scholar] [CrossRef] [PubMed]
- Dunne, J.; Evershed, R.P.; Salque, M.; Cramp, L.; Bruni, S.; Ryan, K.; Biagetti, S.; di Lernia, S. First dairying in green Saharan Africa in the fifth millennium BC. Nature 2012, 486, 390–394. [Google Scholar] [CrossRef] [PubMed]
- Curry, A. The milk revolution. Nature 2013, 500, 20–22. [Google Scholar] [CrossRef] [PubMed]
- Food and Agriculture Organization of the United Nations. Food Outlook-Global Market Analysis; Food and Agriculture Organization of the United Nations: Roma, Italy, 2012; pp. 51–54. [Google Scholar]
- Agriculture and Horticulture Development Board. Dairy Co. World dairy cow numbers. Available online: http://www.dairyco.org.uk/market-information/farming-data/cow-numbers/world-cow-numbers/#.VTfift7-BW_ (assessed on 15 April 2015).
- Laplante, M.; Sabatini, D.M. mTOR signaling in growth control and disease. Cell 2012, 149, 274–293. [Google Scholar] [CrossRef] [PubMed]
- Dazert, E.; Hall, M.N. mTOR signaling in disease. Curr. Opin. Cell Biol. 2011, 23, 744–755. [Google Scholar] [CrossRef] [PubMed]
- Efeyan, A.; Comb, W.C.; Sabatini, D.M. Nutrient-sensing mechanisms and pathways. Nature 2015, 517, 302–310. [Google Scholar] [CrossRef] [PubMed]
- Melnik, B.C.; John, S.M.; Schmitz, G. Milk is not just food but most likely a genetic transfection system activating mTORC1 signaling for postnatal growth. Nutr. J. 2013, 12, 103. [Google Scholar] [CrossRef] [PubMed]
- Wiley, A.S. Cow milk consumption, insulin-like growth factor-I, and human biology: A life history approach. Am. J. Hum. Biol. 2012, 24, 130–138. [Google Scholar] [CrossRef] [PubMed]
- Baier, S.R.; Nguyen, C.; Xie, F.; Wood, J.R.; Zempleni, J. MicroRNAs are absorbed in biologically meaningful amounts from nutritionally relevant doses of cow milk and affect gene expression in peripheral blood mononuclear cells, HEK-293 kidney cell cultures, and mouse livers. J. Nutr. 2014, 144, 1495–1500. [Google Scholar] [CrossRef] [PubMed]
- Mendelsohn, A.R.; Larrick, J.W. Dissecting mammalian target of rapamycin to promote longevity. Rejuvenation Res. 2012, 15, 334–337. [Google Scholar] [CrossRef] [PubMed]
- Xu, S.; Cai, Y.; Wie, Y. mTOR signaling from cellular senescence to organismal aging. Aging Dis. 2014, 5, 263–273. [Google Scholar] [PubMed]
- Johnson, S.C.; Rabinovitch, P.S.; Kaeberlein, M. mTOR is a key modulator of ageing and age-related disease. Nature 2013, 493, 338–345. [Google Scholar] [CrossRef] [PubMed]
- Melnik, B.C.; Zouboulis, C.C. Potential role of FoxO1 and mTORC1 in the pathogenesis of Western diet-induced acne. Exp. Dermatol. 2013, 22, 311–315. [Google Scholar] [CrossRef] [PubMed]
- Zoncu, R.; Efeyan, A.; Sabatini, D.M. mTOR: From growth signal integration to cancer, diabetes and ageing. Nat. Rev. Mol. Cell Biol. 2011, 12, 21–35. [Google Scholar] [CrossRef] [PubMed]
- Cornu, M.; Albert, V.; Hall, M.N. mTOR in aging, metabolism, and cancer. Curr. Opin. Genet. Dev. 2013, 23, 53–62. [Google Scholar] [CrossRef] [PubMed]
- Oddo, S. The role of mTOR signaling in Alzheimer disease. Front. Biosci. 2012, 4, 941–952. [Google Scholar] [CrossRef]
- Tang, Z.; Bereczki, E.; Zhang, H.; Wang, S.; Li, C.; Ji, X.; Branca, R.M.; Lehtiö, J.; Guan, Z.; Filipcik, P.; et al. Mammalian target of rapamycin (mTor) mediates tau protein dyshomeostasis: Implication for Alzheimer disease. J. Biol. Chem. 2013, 288, 15556–15570. [Google Scholar] [CrossRef] [PubMed]
- Michaëlsson, K.; Wolk, A.; Langenskiöld, S.; Basu, S.; Warensjö Lemming, E.; Melhus, H.; Byberg, L. Milk intake and risk of mortality and fractures in women and men: Cohort studies. BMJ 2014, 349, g6015. [Google Scholar] [CrossRef] [PubMed]
- Foster, K.G.; Fingar, D.C. Mammalian target of rapamycin (mTOR): Conducting the cellular signaling symphony. J. Biol. Chem. 2010, 285, 14071–14077. [Google Scholar] [CrossRef] [PubMed]
- Inoki, K.; Ouyang, H.; Li, Y.; Guan, K.L. Signaling by target of rapamycin proteins in cell growth control. Microbiol. Mol. Biol. Rev. 2005, 69, 79–100. [Google Scholar] [CrossRef] [PubMed]
- Avruch, J.; Long, X.; Ortiz-Vega, S.; Rapley, J.; Papageorgiou, A.; Dai, N. Amino acid regulation of TOR complex 1. Am. J. Physiol. Endocrinol. Metab. 2009, 296, E592–E602. [Google Scholar] [CrossRef] [PubMed]
- Sengupta, S.; Peterson, T.; Sabatini, D.M. Regulation of the mTOR complex 1 pathway by nutrients, growth factors, and stress. Mol. Cell 2010, 40, 310–322. [Google Scholar] [CrossRef] [PubMed]
- Laplante, M.; Sabatini, D.M. mTOR signaling. Cold Spring Harb. Perspect. Biol. 2012, 4, a011593. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Guan, K.L. Amino acid signaling in TOR activation. Annu. Rev. Biochem. 2011, 80, 1001–1032. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Buel, G.R.; Blenis, J. Nutrient regulation of the mTOR complex 1 signaling pathway. Mol. Cells 2013, 35, 463–473. [Google Scholar] [CrossRef] [PubMed]
- Jewell, J.L.; Guan, K.L. Nutrient signaling to mTOR and cell growth. Trends Biochem. Sci. 2013, 38, 233–242. [Google Scholar] [CrossRef] [PubMed]
- Efeyan, A.; Sabatini, D.M. Nutrients and growth factors in mTORC1 activation. Biochem. Soc. Trans. 2013, 41, 902–905. [Google Scholar] [CrossRef] [PubMed]
- Laplante, M.; Sabatini, D.M. Regulation of mTORC1 and its impact on gene expression at a glance. J. Cell Sci. 2013, 126, 1713–1719. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Proud, C.G. Nutrient control of mTORC1, a cell-cycle regulator. Trends Cell Biol. 2009, 19, 260–267. [Google Scholar] [CrossRef] [PubMed]
- Chakrabarti, P.; English, T.; Shi, J.; Smas, C.M.; Kandror, K.V. Mammalian target of rapamycin complex 1 suppresses lipolysis, stimulates lipogenesis, and promotes fat storage. Diabetes 2010, 59, 775–781. [Google Scholar] [CrossRef] [PubMed]
- Ricoult, S.J.; Manning, B.D. The multifaceted role of mTORC1 in the control of lipid metabolism. EMBO Rep. 2013, 14, 242–251. [Google Scholar] [CrossRef] [PubMed]
- Yamin, H.B.; Barnea, M.; Genzer, Y.; Chapnik, N.; Froy, O. Long-term commercial cow’s milk consumption and its effects on metabolic parameters associated with obesity in young mice. Mol. Nutr. Food Res. 2014, 58, 1061–1068. [Google Scholar] [CrossRef] [PubMed]
- Howell, J.J.; Ricoult, S.J.; Ben-Sahra, I.; Manning, B.D. A growing role for mTOR in promoting anabolic metabolism. Biochem. Soc. Trans. 2013, 41, 906–912. [Google Scholar] [CrossRef] [PubMed]
- Yoon, M.S.; Zhang, C.; Sun, Y.; Schoenherr, C.J.; Chen, J. Mechanistic target of rapamycin controls homeostasis of adipogenesis. J. Lipid Res. 2013, 54, 2166–2173. [Google Scholar] [CrossRef] [PubMed]
- Goodman, C.A. The role of mTORC1 in regulating protein synthesis and skeletal muscle mass in response to various mechanical stimuli. Rev. Physiol. Biochem. Pharmacol. 2014, 166, 43–95. [Google Scholar] [PubMed]
- Hatfield, I.; Harvey, I.; Yates, E.R.; Redd, J.R.; Reiter, L.T.; Bridges, D. The role of TORC1 in muscle development in Drosophila. Sci. Rep. 2015, 5. [Google Scholar] [CrossRef] [PubMed]
- Yogev, O.; Williams, V.C.; Hinits, Y.; Hughes, S.M. eIF4EBP3L acts as a gatekeeper of TORC1 in activity-dependent muscle growth by specifically regulating Mef2ca translational initiation. PLoS Biol. 2013, 11, e1001679. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Ji, J.; Yan, X.H. Cross-talk between AMPK and mTOR in regulating energy balance. Crit. Rev. Food Sci. Nutr. 2012, 52, 373–381. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.; Qian, S.B. Translational regulation in nutrigenomics. Adv. Nutr. 2011, 2, 511–519. [Google Scholar] [CrossRef] [PubMed]
- Oshiro, N.; Rapley, J.; Avruch, J. Amino acids activate mammalian target of rapamycin (mTOR) complex 1 without changing Rag GTPase guanyl nucleotide charging. J. Biol. Chem. 2014, 289, 2658–2674. [Google Scholar] [CrossRef] [PubMed]
- Dodd, K.M.; Tee, A.R. Leucine and mTORC1: A complex relationship. Am. J. Physiol. Endocrinol. Metab. 2012, 302, E1329–E1342. [Google Scholar] [CrossRef] [PubMed]
- Jewell, J.L.; Kim, Y.C.; Russell, R.C.; Yu, F.X.; Park, H.W.; Plouffe, S.W.; Tagliabracci, V.S.; Guan, K.L. Metabolism. Differential regulation of mTORC1 by leucine and glutamine. Science 2015, 347, 194–198. [Google Scholar] [CrossRef] [PubMed]
- Millward, D.J.; Layman, D.K.; Tomé, D.; Schaafsma, G. Protein quality assessment: Impact of expanding understanding of protein and amino acid needs for optimal health. Am. J. Clin. Nutr. 2008, 87, 1576S–1581S. [Google Scholar] [PubMed]
- Boirie, Y.; Dangin, M.; Gachon, P.; Vasson, M.P.; Maubois, J.L.; Beaufrère, B. Slow and fast dietary proteins differently modulate postprandial protein accretion. Proc. Natl. Acad. Sci. USA 1997, 94, 14930–14935. [Google Scholar] [CrossRef] [PubMed]
- He, T.; Giuseppin, M.L. Slow and fast dietary proteins differentially modulate postprandial metabolism. Int. J. Food Sci. Nutr. 2014, 65, 386–390. [Google Scholar] [CrossRef] [PubMed]
- Boutrou, R.; Gaudichon, C.; Dupont, D.; Jardin, J.; Airinei, G.; Marsset-Baglieri, A.; Benamouzig, R.; Tomé, D.; Leonil, J. Sequential releases of milk protein-derived bioactive peptides in the jejunum in healthy humans. Am. J. Clin. Nutr. 2013, 97, 1414–1423. [Google Scholar] [CrossRef] [PubMed]
- Mahé, S.; Roos, N.; Benamouzig, R.; Davin, L.; Luengo, C.; Gagnon, L.; Gaussergès, N.; Rautureau, J.; Tomé, D. Gastrojejunal kinetics and the digestion of [15N]β-lactoglobulin and casein in humans: The influence of the nature and quantity of the protein. Am. J. Clin. Nutr. 1996, 63, 546–552. [Google Scholar] [PubMed]
- Bounous, G.; Kongshavn, P.A.; Taveroff, A.; Gold, P. Evolutionary traits in human milk proteins. Med. Hypotheses 1988, 27, 133–140. [Google Scholar] [CrossRef]
- Davis, T.A.; Nguyen, H.V.; Garcia-Bravo, R.; Fiorotto, M.L.; Jackson, E.M.; Lewis, D.S.; Lee, D.R.; Reeds, P.J. Amino acid composition of human milk is not unique. J. Nutr. 1994, 124, 1126–1132. [Google Scholar] [PubMed]
- Nicklin, P.; Bergman, P.; Zhang, B.; Triantafellow, E.; Wang, H.; Nyfeler, B.; Yang, H.; Hild, M.; Kung, C.; Wilson, C.; et al. Bidirectional ransport of amino acids regulates mTOR and autophagy. Cell 2009, 136, 521–534. [Google Scholar] [CrossRef] [PubMed]
- Cohen, A.; Hall, M.N. An amino acid shuffle activates mTORC1. Cell 2009, 136, 399–400. [Google Scholar] [CrossRef] [PubMed]
- Duran, R.V.; Oppliger, W.; Robitaille, A.M.; Heiserich, L.; Skendaj, R.; Gottlieb, E.; Hall, M.N. Glutaminolysis activates Rag-mTORC1 signaling. Mol. Cell 2012, 47, 349–358. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Li, C.; Allen, A.; Stanley, C.A.; Smith, T.J. The structure and allosteric regulation of mammalian glutamate dehydrogenase. Arch. Biochem. Biophys. 2012, 519, 69–80. [Google Scholar] [CrossRef] [PubMed]
- Lorin, S.; Tol, M.J.; Bauvy, C.; Strijland, A.; Poüs, C.; Verhoeven, A.J.; Codogno, P.; Meijer, A.J. Glutamate dehydrogenase contributes to leucine sensing in the regulation of autophagy. Autophagy 2013, 9, 850–860. [Google Scholar] [CrossRef] [PubMed]
- McDaniel, M.L.; Marshall, C.A.; Pappan, K.L.; Kwon, G. Metabolic and autocrine regulation of the mammalian target of rapamycin by pancreatic β-cells. Diabetes 2002, 51, 2877–2885. [Google Scholar] [CrossRef] [PubMed]
- Xu, G.; Kwon, G.; Cruz, W.S.; Marshall, C.A.; McDaniel, M.L. Metabolic regulation by leucine of translation initiation through the mTOR-signaling pathway by pancreatic β-cells. Diabetes 2001, 50, 353–360. [Google Scholar] [CrossRef] [PubMed]
- Lenders, C.M.; Liu, S.; Wilmore, D.W.; Sampson, L.; Dougherty, L.W.; Spiegelman, D.; Willett, W.C. Evaluation of a novel food composition database that includes glutamine and other amino acids derived from gene sequencing data. Eur. J. Clin. Nutr. 2009, 63, 1433–1439. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Tsun, Z.Y.; Wolfson, R.; Shen, K.; Wyant, G.A.; Plovanich, M.E.; Yuan, E.D.; Jones, T.D.; Chantranupong, L.; Comb, W.; et al. The amino acid transporter SLC38A9 is a key component of a lysosomal membrane complex that signals arginine sufficiency to mTORC1. Science 2015, 347, 188–194. [Google Scholar] [CrossRef] [PubMed]
- Leibowitz, G.; Cerasi, E.; Ketzinel-Gilad, M. The role of mTOR in the adaptation and failure of β-cells in type 2 diabetes. Diabetes Obes. Metab. 2008, 10, 157–169. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Chi, Y.; Burkhardt, B.R.; Guan, Y.; Wolf, B.A. Leucine metabolism in regulation of insulin secretion from pancreatic β cells. Nutr. Rev. 2010, 68, 270–279. [Google Scholar] [CrossRef] [PubMed]
- Le Bacquer, O.; Queniat, G.; Gmyr, V.; Kerr-Conte, J.; Lefebvre, B.; Pattou, F. mTORC1 and mTORC2 regulate insulin secretion through Akt in INS-1 cells. J. Endocrinol. 2013, 216, 21–29. [Google Scholar] [CrossRef] [PubMed]
- Axelsson, I.E.; Ivarsson, S.A.; Räihä, N.C. Protein intake in early infancy: Effects on plasma amino acid concentrations, insulin metabolism, and growth. Pediatr. Res. 1989, 26, 614–617. [Google Scholar] [CrossRef] [PubMed]
- O’Sullivan, A.; He, X.; McNiven, E.M.; Haggarty, N.W.; Lönnerdal, B.; Slupsky, C.M. Early diet impacts infant Rhesus gut microbiome, immunity, and metabolism. J. Proteome Res. 2013, 12, 2833–2845. [Google Scholar] [CrossRef] [PubMed]
- Socha, P.; Grote, V.; Gruszfeld, D.; Janas, R.; Demmelmair, H.; Closa-Monasterolo, R.; Subías, J.E.; Scaglioni, S.; Verduci, E.; Dain, E.; et al. Milk protein intake, the metabolic-endocrine response, and growth in infancy: Data from a randomized clinical trial. Am. J. Clin. Nutr. 2011, 94, 1776S–1784S. [Google Scholar] [CrossRef] [PubMed]
- Melnik, B.C. The pathogenic role of persistent milk signaling in mTORC1- and milk-microRNA-driven type 2 diabetes mellitus. Curr. Diabetes Rev. 2015, 11, 46–62. [Google Scholar] [CrossRef] [PubMed]
- Holt, S.; Brand Miller, J.; Petocz, P. An insulin index of foods. The insulin demand generated by 1000-kK portions of common foods. Am. J. Clin. Nutr. 1997, 66, 1264–1276. [Google Scholar] [PubMed]
- Hoyt, G.; Hickey, M.S.; Cordain, L. Dissociation of the glycaemic and insulinaemic responses to whole and skimmed milk. Br. J. Nutr. 2005, 93, 175–177. [Google Scholar] [CrossRef] [PubMed]
- Hoppe, C.; Mølgaard, C.; Vaag, A.; Barkholt, V.; Michaelsen, K.F. High intakes of milk, but not meat, increases s-insulin and insulin resistance in 8-year-old boys. Eur. J. Clin. Nutr. 2005, 59, 393–398. [Google Scholar] [CrossRef] [PubMed]
- Hoppe, C.; Mølgaard, C.; Dalum, C.; Vaag, A.; Michaelsen, K.F. Differential effects of casein versus whey on fasting plasma levels of insulin, IGF-1 and IGF-1/IGFBP-3: Results from a randomized 7-day supplementation study in prepubertal boys. Eur. J. Clin. Nutr. 2009, 63, 1076–1083. [Google Scholar] [CrossRef] [PubMed]
- Thomas, F.B.; Sinar, D.; Mazzaferri, E.L.; Cataland, S.; Mekhjian, H.S.; Caldwell, J.H.; Fromkess, J.J. Selective release of gastric inhibitory polypeptide by intraduodenal amino acid perfusion in man. Gastroenterology 1978, 74, 1261–1265. [Google Scholar] [PubMed]
- Chen, Q.; Reimer, R.A. Dairy protein and leucine alter GLP-1 release and mRNA of genes involved in intestinal lipid metabolism in vitro. Nutrition 2009, 25, 340–349. [Google Scholar] [CrossRef] [PubMed]
- Nilsson, M.; Stenberg, M.; Frid, A.H.; Holst, J.J.; Björck, I.M. Glycemia and insulinemia in healthy subjects after lactose-equivalent meals of milk and other food proteins: The role of plasma amino acids and incretins. Am. J. Clin. Nutr. 2004, 80, 1246–2353. [Google Scholar] [PubMed]
- Nilsson, M.; Holst, J.J.; Björck, I.M. Metabolic effects of amino acid mixtures and whey protein in healthy subjects: Studies using glucose-equivalent drinks. Am. J. Clin. Nutr. 2007, 85, 996–1004. [Google Scholar] [PubMed]
- Salehi, A.; Gunnerud, U.; Muhammed, S.J.; Ostman, E.; Holst, J.J.; Björck, I.; Rorsman, P. The insulinogenic effects of whey protein is partially mediated by a direct effect of amino acids and GIP on β-cells. Nutr. Metab. 2012, 9. [Google Scholar] [CrossRef] [PubMed]
- Xie, J.; El Sayed, N.M.; Qi, C.; Zhao, X.; Moore, C.E.; Herbert, T.P. Exendin-4 stimulates islet cell replication via the IGF1 receptor activation of mTORC1/S6K1. J. Mol. Endocrinol. 2014, 53, 105–115. [Google Scholar] [CrossRef] [PubMed]
- Ohara-Imaizumi, M.; Kim, H.; Yoshida, M.; Fujiwara, T.; Aoyagi, K.; Toyofuku, Y.; Nakamichi, Y.; Nishiwaki, C.; Okamura, T.; Uchida, T.; et al. Serotonin regulates glucose-stimulated insulin secretion from pancreatic β cells during pregnancy. Proc. Natl. Acad. Sci. USA 2013, 110, 19420–19425. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.; Oh, C.M.; Ohara-Imaizumi, M.; Park, S.; Namkung, J.; Yadav, V.K.; Tamarina, N.A.; Roe, M.W.; Philipson, L.H.; Karsenty, G.; et al. Functional role of serotonin in insulin secretion in a diet-induced insulin-resistant state. Endocrinology 2015, 156, 444–452. [Google Scholar] [CrossRef] [PubMed]
- Gallegos-Perez, J.L.; Limon, A.; Reyes-Ruiz, J.M.; Alshanqeeti, A.S.; Aljohi, M.A.; Miledi, R. Analysis of free ACh and 5-HT in milk from four different species and their bioactivity on 5-HT(3) and nACh receptors. Food Funct. 2014, 5, 1489–1494. [Google Scholar] [CrossRef] [PubMed]
- Wheelhouse, N.M.; Stubbs, A.K.; Lomax, M.A.; MacRae, J.C.; Hazlerigg, D.G. Growth hormone and amino acid supply interact synergistically to control insulin-like growth factor-I production and gene expression in cultured ovine hepatocytes. J. Endocrinol. 1999, 163, 353–361. [Google Scholar] [CrossRef] [PubMed]
- Harp, J.B.; Goldstein, S.; Phillips, L.S. Nutrition and somatomedin. XXIII. Molecular regulation of IGF-I by amino acid availability in cultured hepatocytes. Diabetes 1991, 40, 95–101. [Google Scholar] [CrossRef] [PubMed]
- Heine, W.; Radke, M.; Wutzke, K.D.; Peters, E.; Kundt, G. α-lactalbumin enriched low-protein infant formulas: A comparison to breast milk feeding. Acta Paediatr. 1996, 85, 1024–1028. [Google Scholar] [CrossRef] [PubMed]
- Markus, C.R.; Olivier, B.; Panhuysen, G.E.; van der Gugten, J.; Alles, M.S.; Tuiten, A.; Westenberg, H.G.; Fekkes, D.; Koppeschaar, H.F.; de Haan, E.E. The bovine protein α-lactalbumin increases the plasma ratio of tryptophan to the other large neutral amino acids, and in vulnerable subjects raises brain serotonin activity, reduces cortisol concentration, and improves mood under stress. Am. J. Clin. Nutr. 2000, 71, 1536–1544. [Google Scholar] [PubMed]
- Qin, L.Q.; He, K.; Xu, J.Y. Milk consumption and circulating insulin-like growth factor-I level: A systematic literature review. Int. J. Food Sci. Nutr. 2009, 60, 330–340. [Google Scholar] [CrossRef] [PubMed]
- Norat, T.; Dossus, L.; Rinaldi, S.; Overvad, K.; Grønbaek, H.; Tjønneland, A.; Olsen, A.; Clavel-Chapelon, F.; Boutron-Ruault, M.C.; Boeing, H.; et al. Diet, serum insulin-like growth factor-I and IGF-binding protein-3 in European women. Eur. J. Clin. Nutr. 2007, 61, 91–98. [Google Scholar] [CrossRef] [PubMed]
- Rich-Edwards, J.W.; Ganmaa, D.; Pollak, M.N.; Nakamoto, E.K.; Kleinman, K.; Tserendolgor, U.; Willett, W.C.; Frazier, A.L. Milk consumption and the prepubertal somatotropic axis. Nutr. J. 2007, 6. [Google Scholar] [CrossRef] [PubMed]
- Larnkjær, A.; Arnberg, K.; Michaelsen, K.F.; Jensen, S.M.; Mølgaard, C. Effect of milk proteins on linear growth and IGF variables in overweight adolescents. Growth Horm. IGF Res. 2014, 24, 54–59. [Google Scholar] [CrossRef] [PubMed]
- Melnik, B.C. Leucine signaling in the pathogenesis of type 2 diabetes and obesity. World J. Diabetes 2012, 3, 38–53. [Google Scholar] [CrossRef] [PubMed]
- Modi, H.; Cornu, M.; Thorens, B. Glutamine stimulates biosynthesis and secretion of insulin-like growth factor 2 (IGF2), an autocrine regulator of β cell mass and function. J. Biol. Chem. 2014, 289, 31972–31982. [Google Scholar] [CrossRef] [PubMed]
- Csibi, A.; Lee, G.; Yoon, S.O.; Tong, H.; Ilter, D.; Elia, I.; Fendt, S.M.; Roberts, T.M.; Blenis, J. The mTORC1/S6K1 pathway regulates glutamine metabolism through the eIF4B-dependent control of c-Myc translation. J. Curr. Biol. 2014, 24, 2274–2280. [Google Scholar] [CrossRef] [PubMed]
- Yasuda, M.; Tanaka, Y.; Kume, S.; Morita, Y.; Chin-Kanasaki, M.; Araki, H.; Isshiki, K.; Araki, S.; Koya, D.; Haneda, M.; et al. Fatty acids are novel nutrient factors to regulate mTORC1 lysosomal localization and apoptosis in podocytes. Biochim. Biophys. Acta 2014, 1842, 1097–1108. [Google Scholar] [CrossRef] [PubMed]
- Mordier, S.; Iynedjian, P.B. Activation of mammalian target of rapamycin complex 1 and insulin resistance induced by palmitate in hepatocytes. Biochem. Biophys. Res. Commun. 2007, 362, 206–211. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Yu, W.; Nawaz, A.; Guan, F.; Sun, S.; Wang, C. Palmitate induced insulin resistance by PKCtheta-dependent activation of mTOR/S6K pathway in C2C12 myotubes. Exp. Clin. Endocrinol. Diabetes 2010, 118, 657–661. [Google Scholar] [CrossRef] [PubMed]
- Hatanaka, M.; Maier, B.; Sims, E.K.; Templin, A.T.; Kulkarni, R.N.; Evans-Molina, C.; Mirmira, R.G. Palmitate induces mRNA translation and increases ER protein load in islet β-cells via activation of the mammalian target of rapamycin pathway. Diabetes 2014, 63, 3404–3415. [Google Scholar] [CrossRef] [PubMed]
- Pardo, V.; González-Rodríguez, Á.; Muntané, J.; Kozma, S.C.; Valverde, Á.M. Role of hepatocyte S6K1 in palmitic acid-induced endoplasmic reticulum stress, lipotoxicity, insulin resistance and in oleic acid-induced protection. Food Chem. Toxicol. 2015, 80, 298–309. [Google Scholar] [CrossRef] [PubMed]
- Um, S.H.; D’Alessio, D.; Thomas, G. Nutrient overload, insulin resistance, and ribosomal protein S6 kinase 1, S6K1. Cell Metab. 2006, 3, 393–402. [Google Scholar] [CrossRef] [PubMed]
- Jensen, R.G.; Ferris, A.M.; Lammi-Keefe, C.J. The composition of milk fat. J. Dairy Sci. 1991, 74, 3228–3243. [Google Scholar] [CrossRef]
- Bitman, J.; Wood, D.L. Changes in milk fat phospholipids during lactation. J. Dairy Sci. 1990, 73, 1208–1216. [Google Scholar] [CrossRef]
- Valadi, H.; Ekstöm, K.; Bossios, A.; Sjöstrand, M.; Lee, J.J.; Lötvall, J.O. Exosome-mediated transfer of mRNA and microRNAs is novel mechanism of genetic exchange between cells. Nat. Cell Biol. 2007, 9, 654–659. [Google Scholar] [CrossRef] [PubMed]
- Liang, H.; Huang, L.; Cao, J.; Zen, K.; Chen, X.; Zhang, C.Y. Regulation of mammalian gene expression by exogeneous microRNAs. Wiley Interdiscip. Rev. RNA 2012, 3, 733–742. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Liang, H.; Zhang, J.; Zen, K.; Thang, C.Y. Secreted microRNAs: A new form of intercellular communication. Trends Cell Biol. 2012, 22, 125–132. [Google Scholar] [CrossRef] [PubMed]
- Ludwig, A.K.; Giebel, B. Exosomes: Small vesicles participating in intercellular communication. Int. J. Biochem. Cell Biol. 2012, 44, 11–15. [Google Scholar] [CrossRef] [PubMed]
- Ambros, V. The functions of animal microRNAs. Nature 2004, 431, 350–355. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Liang, H.; Zhang, J.; Zen, K.; Zhang, C.Y. Horizontal transfer of microRNAs: Molecular mechanism and clinical applications. Protein Cell 2012, 3, 28–37. [Google Scholar] [CrossRef] [PubMed]
- Weber, J.A.; Baxter, D.H.; Zhang, S.; Huang, D.Y.; Huang, K.H.; Lee, M.J.; Galas, D.J.; Wang, K. The microRNA spectrum in 12 body fluids. Clin. Chem. 2010, 56, 1733–1741. [Google Scholar] [CrossRef] [PubMed]
- Reinhardt, T.A.; Lippolis, J.D.; Nonnecke, B.J.; Sacco, R.E. Bovine milk exosome proteome. J. Proteomics 2012, 75, 1486–1492. [Google Scholar] [CrossRef] [PubMed]
- Hata, T.; Murakami, K.; Nakatani, H.; Yamamoto, Y.; Matsuda, T.; Aoki, N. Isolation of bovine milk-derived microvesicles carrying mRNAs and microRNAs. Biochem. Biophys. Res. Commun. 2010, 396, 528–533. [Google Scholar] [CrossRef] [PubMed]
- Izumi, H.; Kosaka, N.; Shimizu, T.; Sekine, K.; Ochiya, T.; Takase, M. Purification of RNA from milk whey. Methods Mol. Biol. 2013, 1024, 191–201. [Google Scholar] [PubMed]
- Izumi, H.; Tsuda, M.; Sato, Y.; Kosaka, N.; Ochiya, T.; Iwamoto, H.; Namba, K.; Takeda, Y. Bovine milk exosomes contain microRNA and mRNA and are taken up by human macrophages. J. Dairy Sci. 2015, 98, 2920–2933. [Google Scholar] [CrossRef] [PubMed]
- Pieters, B.C.; Arntz, O.J.; Bennink, M.B.; Broeren, M.G.; van Caam, A.P.; Koenders, M.I.; van Lent, P.L.; van den Berg, W.B.; de Vries, M.; van der Kraan, P.M.; et al. Commercial cow milk contains physically stable extracellular vesicles expressing immunoregulatory TGF-β. PLoS ONE 2015, 10, e0121123. [Google Scholar] [CrossRef] [PubMed]
- Sun, Q.; Chen, X.; Yu, J.; Zen, K.; Zhang, C.Y.; Li, L. Immune modulatory function of abundant immune-related microRNAs in microvesicles from bovine colostrum. Protein Cell 2013, 4, 197–210. [Google Scholar] [CrossRef] [PubMed]
- Kosaka, N.; Izumi, H.; Sekine, K.; Ochiya, T. microRNA as a new immune-regulatory agent in breast milk. Silence 2010, 1. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Q.; Li, M.; Wang, X.; Li, Q.; Wang, T.; Zhu, Q.; Zhou, X.; Wang, X.; Gao, X.; Li, X. Immune-related microRNAs are abundant in breast milk exosomes. Int. J. Biol. Sci. 2012, 8, 118–123. [Google Scholar] [CrossRef] [PubMed]
- Melnik, B.C.; John, S.M.; Schmitz, G. Milk: An exosomal microRNA transmitter promoting thymic regulatory T cell maturation preventing the development of atopy? J. Transl. Med. 2014, 12. [Google Scholar] [CrossRef] [PubMed]
- Fernández-Messina, L.; Gutiérrez-Vázquez, C.; Rivas-García, E.; Sánchez-Madrid, F.; de la Fuente, H. Immunomodulatory role of microRNAs transferred by extracellular vesicles. Biol. Cell 2015, 107, 61–77. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Gao, C.; Li, H.; Huang, L.; Sun, Q.; Dong, Y.; Guan, D.; Hu, X.; Zhao, S.; Li, L.; et al. Identification and characterization of microRNAs in raw milk during different periods of lactation, commercial fluid, and powdered milk products. Cell Res. 2010, 20, 1128–1137. [Google Scholar] [CrossRef] [PubMed]
- Izumi, H.; Kosaka, N.; Shimizu, T.; Sekine, K.; Ochiya, T.; Takase, M. Bovine milk contains microRNA and messenger RNA that are stable under degradative conditions. J. Dairy Sci. 2012, 95, 4831–4841. [Google Scholar] [CrossRef] [PubMed]
- Howard, K.M.; Jati Kusuma, R.; Baier, S.R.; Friemel, T.; Markham, L.; Vanamala, J.; Zempleni, J. Loss of miRNAs during processing and storage of cow’s (Bos taurus) milk. J. Agric. Food Chem. 2015, 63, 588–592. [Google Scholar] [CrossRef] [PubMed]
- Brosnan, J.T.; Brosnan, M.E. Branched-chain amino acids: Enzyme and substrate regulation. J. Nutr. 2006, 136, 207S–211S. [Google Scholar] [PubMed]
- Islam, M.M.; Nautiyal, M.; Wynn, R.M.; Mobley, J.A.; Chuang, D.T.; Hutson, S.M. Branched-chain amino acid metabolon: Interaction of glutamate dehydrogenase with the mitochondrial branched-chain aminotransferase (BCATm). J. Biol. Chem. 2010, 285, 265–276. [Google Scholar] [CrossRef] [PubMed]
- AEvarsson, A.; Chuang, J.L.; Wynn, R.M.; Turley, S.; Chuang, D.T.; Hol, W.G. Crystal structure of human branched-chain α-ketoacid dehydrogenase and the molecular basis of multienzyme complex deficiency in maple syrup urine disease. Structure 2000, 8, 277–291. [Google Scholar] [CrossRef]
- Mersey, B.D.; Jin, P.; Danner, D.J. Human microRNA (miR29b) expression controls the amount of branched chain α ketoacid dehydrogenase complex in a cell. Hum. Mol. Genet. 2005, 14, 3371–3377. [Google Scholar] [CrossRef] [PubMed]
- Bian, Y.; Lei, Y.; Wang, C.; Wang, J.; Wang, L.; Liu, L.; Liu, L.; Gao, X.; Li, Q. Epigenetic regulation of miR-29s affects the lactation activity of dairy cow mammary epithelial cells. J. Cell. Physiol. 2015, 230, 2152–2163. [Google Scholar] [CrossRef] [PubMed]
- Davie, J.R.; Wynn, R.M.; Meng, M.; Huang, Y.S.; Aalund, G.; Chuang, D.T.; Lau, K.S. Expression and characterization of branched-chain α-ketoacid dehydrogenase kinase from the rat. Is it a histidine-protein kinase? J. Biol.Chem. 1995, 270, 19861–19867. [Google Scholar] [CrossRef] [PubMed]
- Chou, P.Y.; Fasman, G.D. Structural and functional role of leucine residues in proteins. J. Mol. Biol. 1973, 74, 263–281. [Google Scholar] [CrossRef]
- Glover, J.N.; Harrison, S.C. Crystal structure of the heterodimeric bZIP transcription factor c-Fos-c-Jun bound to DNA. Nature 1995, 373, 257–261. [Google Scholar] [CrossRef] [PubMed]
- Hawgood, S.; Derrick, M.; Poulain, F. Structure and properties of surfactant protein B. Biochim. Biophys. Acta 1998, 1408, 150–160. [Google Scholar] [CrossRef]
- Chung, J.; Bauer, D.E.; Ghamari, A.; Nizzi, C.P.; Deck, K.M.; Kingsley, P.D.; Yien, Y.Y.; Huston, N.C.; Chen, C.; Schultz, I.J.; et al. The mTORC1/4E-BP pathway coordinates hemoglobin production with L-leucine availability. Sci. Signal. 2015, 8, ra34. [Google Scholar] [CrossRef] [PubMed]
- Meng, F.; Henson, R.; Wehbe-Janek, H.; Ghoshal, K.; Jacob, S.T.; Patel, T. MicroRNA-21 regulates expression of the PTEN tumor suppressor gene in human hepatocellular cancer. Gastroenterology 2007, 133, 647–658. [Google Scholar] [CrossRef] [PubMed]
- Han, M.; Liu, M.; Wang, Y.; Chen, X.; Xu, J.; Sun, Y.; Zhao, L.; Qu, H.; Fan, Y.; Wu, C. Antagonism of miR-21 reverses epithelial-mesenchymal transition and cancer stem cell phenotype through AKT/ERK1/2 incactivation by targeting PTEN. PLoS ONE 2012, 7, e39520. [Google Scholar] [CrossRef] [PubMed]
- Dey, N.; Ghosh-Choudhury, N.; Kasinath, B.S.; Choudhury, G.G. TGFβ-stimulated microRNA-21 utilizes PTEN to orchestrate AKT/mTORC1 signaling for mesangial cell hypertrophy and matrix expansion. PLoS ONE 2012, 7, e42316. [Google Scholar] [CrossRef] [PubMed]
- Xiong, B.; Cheng, Y.; Ma, L.; Zhang, C. MiR-21 regulates biological behavior through the PTEN/PI-3K/Akt signaling pathway in human colorectal cancer cells. Int. J. Oncol. 2013, 42, 219–228. [Google Scholar] [PubMed]
- Huang, J.; Manning, B.D. A complex interplay between Akt, TSC2 and the two mTOR complexes. Biochem. Soc. Trans. 2009, 37, 217–222. [Google Scholar] [CrossRef] [PubMed]
- Sayed, D.; Rane, S.; Lypowy, J.; He, M.; Chen, I.Y.; Vashistha, H.; Yan, L.; Malhotra, A.; Vatner, D.; Abdellatif, M. MicroRNA-21 targets Sprouty2 and promotes cellular outgrouths. Mol. Biol. Cell 2008, 19, 3272–3282. [Google Scholar] [CrossRef] [PubMed]
- Dariminpourain, M.; Wang, S.; Ittmann, M.; Kwabi-Addo, B. Transcriptional and post-transcriptional regulation of Sprouty1, a receptor tyrosine kinase inhibitor in prostate cancer. Prostate Cancer Prostatic Dis. 2011, 14, 279–285. [Google Scholar] [CrossRef] [PubMed]
- Frey, M.R.; Carraro, G.; Batra, R.K.; Polk, D.B.; Warburton, D. Sprouty keeps bowel kinases regular in colon cancer, while miR-21 targets Sprouty. Cancer Biol. Ther. 2011, 11, 122–124. [Google Scholar] [CrossRef] [PubMed]
- Van der Heide, L.P.; Hoekman, M.F.; Smidt, M.P. The ins and outs of FoxO shuttling: Mechanisms of FoxO translocation and transcriptional regulation. Biochem. J. 2004, 380, 297–309. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.; Tindall, D.J. Dynamic FoxO transcription factors. J. Cell Sci. 2007, 120, 2479–2487. [Google Scholar] [CrossRef] [PubMed]
- Lei, B.X.; Liu, Z.H.; Li, Z.J.; Li, C.; Deng, Y.F. miR-21 induces cell proliferation and suppresses the chemosensitivity in glioblastoma cells via downregulation of FOXO1. Int. J. Clin. Exp. Med. 2014, 7, 2060–2066. [Google Scholar] [PubMed]
- Song, W.; Wang, L.; Wang, L.; Li, Q. Interplay of miR-21 and FoxO1 modulates growth of pancreatic ductal adenocarcinoma. Tumour Biol. 2015. [Google Scholar] [CrossRef] [PubMed]
- Song, W.; Li, Q.; Wang, L.; Wang, L. Modulation of FoxO1 expression by miR-21 to promote growth of pancreatic ductal adenocarcinoma. Cell. Physiol. Biochem. 2015, 35, 184–190. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.H.; Yue, J.; Pfeffer, S.R.; Fan, M.; Paulus, E.; Hosni-Ahmed, A.; Sims, M.; Qayyum, S.; Davidoff, A.M.; Handorf, C.R.; et al. MicroRNA-21 promotes glioblastoma tumorigenesis by down-regulating insulin-like growth factor-binding protein-3 (IGFBP3). J. Biol. Chem. 2014, 289, 25079–25087. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.C.; Jeon, S.M.; Bhaskar, P.T.; Nogueira, V.; Sundararajan, D.; Tonic, I.; Park, Y.; Hay, N. FoxOs inhibit mTORC1 and activate Akt by inducing the expression of Sestrin3 and Rictor. Dev. Cell 2010, 18, 592–604. [Google Scholar] [CrossRef] [PubMed]
- Hay, N. Interplay between FOXO, TOR, and Akt. Biochim. Biophys. Acta 2011, 1813, 1965–1970. [Google Scholar] [CrossRef] [PubMed]
- Jünger, M.A.; Rintelen, F.; Stocker, H.; Wasserman, J.D.; Végh, M.; Radimerski, T.; Greenberg, M.E.; Hafen, E. Drosophila forkhead transcription factor FOXO mediates the reduction in cell number associated with reduced insulin signaling. J. Biol. 2003, 2, 20. [Google Scholar] [CrossRef] [PubMed]
- Puig, O.; Marr, M.T.; Ruhf, M.L.; Tjian, R. Control of cell number by Drosophila FOXO: Downstream and feedback regulation of the insulin receptor pathway. Genes Dev. 2003, 17, 2006–2020. [Google Scholar] [CrossRef] [PubMed]
- Thoreen, C.C.; Chantranupong, L.; Keys, H.R.; Wang, T.; Gray, N.S.; Sabatini, D.M. A unifying model for mTORC1-mediated regulation of mRNA translation. Nature 2012, 485, 109–113. [Google Scholar] [CrossRef] [PubMed]
- Asangani, I.A.; Rasheed, S.A.; Nikolova, D.A.; Leupold, J.H.; Colburn, N.H.; Post, S.; Allgayer, H. MicroRNA-21 (miR-21) post-transcriptionally downregulates tumor suppressor Pdcd4 and stimulates invasion, intravasation and metastasis in colorectal cancer. Oncogene 2008, 27, 2128–2136. [Google Scholar] [CrossRef] [PubMed]
- Lu, Z.; Liu, M.; Stribinskis, V.; Klinge, C.M.; Ramos, K.S.; Colburn, N.H.; Li, Y. MicroRNA-21 promotes cell transformation by targeting the programmed cell death 4 gene. Oncogene 2008, 27, 4373–4379. [Google Scholar] [CrossRef] [PubMed]
- Carayol, N.; Katsoulidis, E.; Sassano, A.; Altman, J.K.; Druker, B.J.; Platanias, L.C. Suppression of programmed cell death 4 (PDCD4) protein expression by BCR-ABL-regulated engagement of the mTOR/p70 S6 kinase pathway. J. Biol. Chem. 2008, 28, 8601–8610. [Google Scholar] [CrossRef] [PubMed]
- Dennis, M.D.; Jefferson, L.S.; Kimball, S.R. Role of p70S6K1-mediated phosphorylation of eIF4B and PDCD4 proteins in the regulation of protein synthesis. J. Biol. Chem. 2012, 287, 42890–42899. [Google Scholar] [CrossRef] [PubMed]
- Davis, B.N.; Hilyard, A.C.; Lagna, G.; Hata, A. SMAD proteins control DROSHA-mediated microRNA maturation. Nature 2008, 454, 56–61. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.M.; Blenis, J. Molecular mechanisms of mTOR-mediated translational control. Nat. Rev. Mol. Cell Biol. 2009, 10, 307–318. [Google Scholar] [CrossRef] [PubMed]
- Szymańska, P.; Martin, K.R.; MacKeigan, J.P.; Hlavacek, W.S.; Lipniacki, T. Computational analysis of an autophagy/translation switch based on mutual inhibition of MTORC1 and ULK1. PLoS ONE 2015, 10, e0116550. [Google Scholar] [CrossRef] [PubMed]
- Haghighat, A.; Mader, S.; Pause, A.; Sonenberg, N. Repression of cap-dependent translation by 4E-binding protein 1: Competition with p220 for binding to eukaryotic initiation factor-4E. EMBO J. 1995, 14, 5701–5709. [Google Scholar] [PubMed]
- Gross, J.D.; Moerke, N.J.; von der Haar, T.; Lugovskoy, A.A.; Sachs, A.B.; McCarthy, J.E.; Wagner, G. Ribosome loading onto the mRNA cap is driven by conformational coupling between eIF4G and eIF4E. Cell 2003, 115, 739–750. [Google Scholar] [CrossRef]
- Moerke, N.J.; Aktas, H.; Chen, H.; Cantel, S.; Reibarkh, M.Y.; Fahmy, A.; Gross, J.D.; Degterev, A.; Yuan, J.; Chorev, M.; et al. Small-molecule inhibition of the interaction between the translation initiation factors eIF4E and eIF4G. Cell 2007, 128, 257–267. [Google Scholar] [CrossRef] [PubMed]
- Schütz, P.; Bumann, M.; Oberholzer, A.E.; Bieniossek, C.; Trachsel, H.; Altmann, M.; Baumann, U. Crystal structure of the yeast eIF4A-eIF4G complex: An RNA-helicase controlled by protein-protein interactions. Proc. Natl. Acad. Sci. USA 2008, 105, 9564–9569. [Google Scholar] [CrossRef] [PubMed]
- Yanagiya, A.; Svitkin, Y.V.; Shibata, S.; Mikami, S.; Imataka, H.; Sonenberg, N. Requirement of RNA binding of mammalian eukaryotic translation initiation factor 4GI (eIF4GI) for efficient interaction of eIF4E with the mRNA cap. Mol. Cell. Biol. 2009, 29, 1661–1669. [Google Scholar] [CrossRef] [PubMed]
- Marintchev, A.; Edmonds, K.A.; Marintcheva, B.; Hendrickson, E.; Oberer, M.; Suzuki, C.; Herdy, B.; Sonenberg, N.; Wagner, G. Topology and regulation of the human eIF4A/4G/4H helicase complex in translation initiation. Cell 2009, 136, 447–460. [Google Scholar] [CrossRef] [PubMed]
- Raught, B.; Peiretti, F.; Gingras, A.C.; Livingstone, M.; Shahbazian, D.; Mayeur, G.L.; Polakiewicz, R.D.; Sonenberg, N.; Hershey, J.W. Phosphorylation of eucaryotic translation initiation factor 4B Ser422 is modulated by S6 kinases. EMBO J. 2004, 23, 1761–1769. [Google Scholar] [CrossRef] [PubMed]
- Holz, M.K.; Ballif, B.A.; Gygi, S.P.; Blenis, J. mTOR and S6K1 mediate assembly of the translation preinitiation complex through dynamic protein interchange and ordered phosphorylation events. Cell 2005, 123, 569–580. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, C.; Garces, R.G.; Edmonds, K.A.; Hiller, S.; Hyberts, S.G.; Marintchev, A.; Wagner, G. PDCD4 inhibits translation initiation by binding to eIF4A using both its MA3 domains. Proc. Natl. Acad. Sci. USA 2008, 105, 3274–3279. [Google Scholar] [CrossRef] [PubMed]
- Chang, J.H.; Cho, Y.H.; Sohn, S.Y.; Choi, J.M.; Kim, A.; Kim, Y.C.; Jang, S.K.; Cho, Y. Crystal structure of the eIF4A-PDCD4 complex. Proc. Natl. Acad. Sci. USA 2009, 106, 3148–3153. [Google Scholar] [CrossRef] [PubMed]
- Rahman, H.; Qasim, M.; Oellerich, M.; Asif, A.R. Crosstalk between Edc4 and mammalian target of rapamycin complex 1 (mTORC1) signaling in mRNA decapping. Int. J. Mol. Sci. 2014, 15, 23179–23195. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.H.; Yang, W.H.; Gulick, T.; Bloch, K.D.; Bloch, D.B. Ge-1 is a central component of the mammalian cytoplasmic mRNA processing body. RNA 2005, 11, 1795–1802. [Google Scholar] [CrossRef] [PubMed]
- Brodersen, P.; Sakvarelidze-Achard, L.; Bruun-Rasmussen, M.; Dunoyer, P.; Yamamoto, Y.Y.; Sieburth, L.; Voinnet, O. Widespread translational inhibition by plant miRNAs and siRNAs. Science 2008, 320, 1185–1190. [Google Scholar] [CrossRef] [PubMed]
- Bossé, G.D.; Rüegger, S.; Ow, M.C.; Vasquez-Rifo, A.; Rondeau, E.L.; Ambros, V.R.; Grosshans, H.; Simard, M.J. The decapping scavenger enzyme DCS-1 controls microRNA levels in Caenorhabditis elegans. Mol. Cell 2013, 50, 28128–28127. [Google Scholar] [CrossRef] [PubMed]
- Faller, W.J.; Jackson, T.J.; Knight, J.R.; Ridgway, R.A.; Jamieson, T.; Karim, S.A.; Jones, C.; Radulescu, S.; Huels, D.J.; Myant, K.B.; et al. mTORC1-mediated translational elongation limits intestinal tumour initiation and growth. Nature 2015, 517, 497–500. [Google Scholar] [CrossRef] [PubMed]
- Csibi, A.; Fendt, S.M.; Li, C.; Poulogiannis, G.; Choo, A.Y.; Chapski, D.J.; Jeong, S.M.; Dempsey, J.M.; Parkhitko, A.; Morrison, T.; et al. The mTORC1 pathway stimulates glutamine metabolism and cell proliferation by repressing SIRT4. Cell 2013, 153, 840–854. [Google Scholar] [CrossRef] [PubMed]
- Workman, J.J.; Chen, H.; Laribee, R.N. Environmental signaling through the mechanistic target of rapamycin complex 1: mTORC1 goes nuclear. Cell Cycle 2014, 13, 714–725. [Google Scholar] [CrossRef] [PubMed]
- Smith, E.M.; Proud, C.G. Cdc2-cyclin B regulates eEF2 kinase activity in a cell cycle- and amino acid-dependent manner. EMBO J. 2008, 27, 1005–1016. [Google Scholar] [CrossRef] [PubMed]
- Kimball, S.R. Regulation of translation initiation by amino acids in eukaryotic cells. Prog. Mol. Subcell. Biol. 2001, 26, 155–184. [Google Scholar] [PubMed]
- Proud, C.G. Regulation of mammalian translation factors by nutrients. Eur. J. Biochem. 2002, 269, 5338–5349. [Google Scholar] [CrossRef] [PubMed]
- Schriever, S.C.; Deutsch, M.J.; Adamski, J.; Roscher, A.A.; Ensenauer, R. Cellular signaling of amino acids towards mTORC1 activation in impaired human leucine catabolism. J. Nutr. Biochem. 2013, 24, 824–831. [Google Scholar] [CrossRef] [PubMed]
- Proud, C.G. Control of the translational machinery by amino acids. Am. J. Clin. Nutr. 2014, 99, 231S–236S. [Google Scholar] [CrossRef] [PubMed]
- Blahna, M.T.; Hata, A. Smad-mediated regulation of microRNA biosynthesis. FEBS Lett. 2012, 586, 1906–1912. [Google Scholar] [CrossRef] [PubMed]
- Zavadil, J.; Narasimhan, M.; Blumenberg, M.; Schneider, R.J. Transforming growth factor-β and microRNA:mRNA regulatory networks in epithelial plasticity. Cells Tissues Organs 2007, 185, 157–161. [Google Scholar] [CrossRef] [PubMed]
- Patursky-Polischuk, I.; Kasir, J.; Miloslavski, R.; Hayouka, Z.; Hausner-Hanochi, M.; Stolovich-Rain, M.; Tsukerman, P.; Biton, M.; Mudhasani, R.; Jones, S.N.; et al. Reassessment of the role of TSC, mTORC1 and microRNAs in amino acids mediated translational control of TOP mRNAs. PLoS ONE 2014, 9, e109410. [Google Scholar] [CrossRef] [PubMed]
- Das, F.; Ghosh-Choudhury, N.; Mahimainathan, L.; Venkatesan, B.; Feliers, D.; Riley, D.J.; Kasinath, B.S.; Choudhury, G.G. Raptor-rictor axis in TGFβ-induced protein synthesis. Cell Signal. 2008, 20, 409–423. [Google Scholar] [CrossRef] [PubMed]
- Das, F.; Bera, A.; Ghosh-Choudhury, N.; Abboud, H.E.; Kasinath, B.S.; Choudhury, G.G. TGFβ-induced deptor suppression recruits mTORC1 and not mTORC2 to enhance collagen I (α2) gene expression. PLoS ONE 2014, 9, e109608. [Google Scholar] [CrossRef] [PubMed]
- Das, F.; Ghosh-Choudhury, N.; Bera, A.; Dey, N.; Abboud, H.E.; Kasinath, B.S.; Choudhury, G.G. Transforming growth factor β integrates Smad 3 to mechanistic target of rapamycin complexes to arrest deptor abundance for glomerular mesangial cell hypertrophy. J. Biol. Chem. 2013, 288, 7756–7768. [Google Scholar] [CrossRef] [PubMed]
- Olsen, S.F.; Halldorsson, T.I.; Willett, W.C.; Knudsen, V.K.; Gillman, M.W.; Mikkelsen, T.B.; Olsen, J.; NUTRIX Consortium. Milk consumption during pregnancy is associated with increased infant size at birth: Prospective cohort study. Am. J. Clin. Nutr. 2007, 86, 1104–1110. [Google Scholar] [PubMed]
- Heppe, D.H.; van Dam, R.M.; Willemsen, S.P.; den Breeijen, H.; Raat, H.; Hofman, A.; Steegers, E.A.; Jaddoe, V.W. Maternal milk consumption, fetal growth, and the risks of neonatal complications: The Generation R Study. Am. J. Clin. Nutr. 2011, 94, 501–509. [Google Scholar] [CrossRef] [PubMed]
- Melnik, B.C.; John, S.M.; Schmitz, G. Milk consumption during pregnancy increases birth weight, a risk factor for the development of diseases of civilization. J. Transl. Med. 2015, 13. [Google Scholar] [CrossRef] [PubMed]
- Roos, S.; Jansson, N.; Palmberg, I.; Säljö, K.; Powell, T.L.; Jansson, T. Mammalian target of rapamycin in the human placenta regulates leucine transport and is down-regulated in restricted growth. J. Physiol. 2007, 582, 449–459. [Google Scholar] [CrossRef] [PubMed]
- Jansson, T.; Aye, I.L.; Goberdhan, D.C. The emerging role of mTORC1 signaling in placental nutrient-sensing. Placenta 2012, 33, e23–e29. [Google Scholar] [CrossRef] [PubMed]
- Roos, S.; Lagerlöf, O.; Wennergren, M.; Powell, T.L.; Jansson, T. Regulation of amino acid transporters by glucose and growth factors in cultured primary human trophoblast cells is mediated by mTOR signaling. Am. J. Physiol. Cell Physiol. 2009, 297, C723–C731. [Google Scholar] [CrossRef] [PubMed]
- Jansson, N.; Rosario, F.J.; Gaccioli, F.; Lager, S.; Jones, H.N.; Roos, S.; Jansson, T.; Powell, T.L. Activation of placental mTOR signaling and amino acid transporters in obese women giving birth to large babies. J. Clin. Endocrinol. Metab. 2013, 98, 105–113. [Google Scholar] [CrossRef] [PubMed]
- Jiang, H.; Wu, W.; Zhang, M.; Li, J.; Peng, Y.; Miao, T.T.; Zhu, H.; Xu, G. Aberrant upregulation of miR-21 in placental tissues of macrosomia. J. Perinatol. 2014, 34, 658–663. [Google Scholar] [CrossRef] [PubMed]
- Gaccioli, F.; White, V.; Capobianco, E.; Powell, T.L.; Jawerbaum, A.; Jansson, T. Maternal overweight induced by a diet with high content of saturated fat activates placental mTOR and eIF2α signaling and increases fetal growth in rats. Biol. Reprod. 2013, 89. [Google Scholar] [CrossRef]
- Jung, C.H.; Kim, H.; Ahn, J.; Jeon, T.I.; Lee, D.H.; Ha, T.Y. Fisetin regulates obesity by targeting mTORC1 signaling. J. Nutr. Biochem. 2013, 24, 1547–1554. [Google Scholar] [CrossRef] [PubMed]
- Wiley, A.S. Dairy and milk consumption and child growth: Is BMI involved? An analysis of NHANES 1999–2004. Am. J. Hum. Biol. 2010, 22, 517–525. [Google Scholar] [CrossRef] [PubMed]
- Berkey, C.S.; Rocket, H.R.; Willet, W.C.; Colditz, G.A. Milk, dairy fat, dietary calcium, and weight gain. Arch. Pediatr. Adolesc. Med. 2005, 159, 543–550. [Google Scholar] [CrossRef] [PubMed]
- Matthews, V.L.; Wien, M.; Sabaté, J. The risk of child and adolescent overweight is related to types of food consumed. Nutr. J. 2011, 10, 71–74. [Google Scholar] [CrossRef] [PubMed]
- Arnberg, K.; Mølgaard, C.; Michaelsen, K.F.; Jensen, S.M.; Trolle, E.; Larnkjær, A. Skim milk, whey, and casein increase body weight and whey and casein increase plasma C-peptide concentration in overweight adolescents. J. Nutr. 2012, 142, 2083–2090. [Google Scholar] [CrossRef] [PubMed]
- Barr, S.I.; McCarron, D.A.; Heaney, R.P.; Dawson-Hughes, B.; Berga, S.L.; Stern, J.S.; Oparil, S. Effects of increased consumption of fluid milk on energy and nutrient intake, body weight, and cardiovascular risk factors in healthy older adults. Am. J. Diet. Assoc. 2000, 100, 810–817. [Google Scholar] [CrossRef]
- Benatar, J.R.; Sidhu, K.; Stewart, R.A. Effects of high and low fat dairy food on cardio-metabolic risk factors: A meta-analysis of randomized studies. PLoS ONE 2013, 8, e76480. [Google Scholar] [CrossRef] [PubMed]
- Paul, I.M.; Camera, L.; Zeiger, R.S.; Guilbert, T.W.; Bacharier, L.B.; Taussig, L.M.; Morgan, W.J.; Covar, R.A.; Krawiec, M.; Bloomberg, G.R.; et al. Relationship between infant weight gain and later asthma. Pediatr. Allergy Immunol. 2010, 21, 82–89. [Google Scholar] [CrossRef] [PubMed]
- Hersoug, L.G.; Linneberg, A. The link between the epidemics of obesity and allergic diseases: Does obesity induce decreased immune tolerance? Allergy 2007, 62, 1205–1213. [Google Scholar] [CrossRef] [PubMed]
- Gorgievska-Sukaroovska, B.; Lipozencić, J.; Susac, A. Obesity and allergic diseases. Acta Dermatovenerol. Croat. 2008, 16, 231–235. [Google Scholar]
- Ford, E.S. The epidemiology of obesity and asthma. J. Allergy Clin. Immunol. 2005, 115, 897–909. [Google Scholar] [CrossRef] [PubMed]
- Koletzko, B.; von Kries, R.; Closa, R.; Escribano, J.; Scaglioni, S.; Giovannini, M.; Beyer, J.; Demmelmair, H.; Gruszfeld, D.; Dobrzanska, A.; et al. Lower protein in infant formula is associated with lower weight up tp age 2 year: A randomized clinical trial. Am. J. Clin. Nutr. 2009, 89, 1836–1845. [Google Scholar] [CrossRef] [PubMed]
- Escribano, J.; Luque, V.; Ferre, N.; Mendez-Riera, G.; Koletzko, B.; Grote, V.; Demmelmair, H.; Bluck, L.; Wright, A.; Closa-Monasterolo, R.; et al. Effect of protein intake and weight gain velocity on body fat mass at 6 months of age: The EU Childhood Obesity Programme. Int. J. Obes. 2012, 36, 548–553. [Google Scholar] [CrossRef] [PubMed]
- Weber, M.; Grote, V.; Closa-Monasterolo, R.; Escribano, J.; Langhendries, J.P.; Dain, E.; Giovannini, M.; Verduci, E.; Gruszfeld, D.; Socha, P.; et al. Lower protein content in infant formula reduces BMI and obesity risk at school age: Follow-up of a randomized trial. Am. J. Clin. Nutr. 2014, 99, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Melnik, B.C. Excessive leucine-mTORC1-signalling of cow milk-based infant formula: The missing link to understand early childhood obesity. J. Obes. 2012, 2012. [Google Scholar] [CrossRef] [PubMed]
- Brüske, I.; Flexeder, C.; Heinrich, J. Body mass index and the incidence of asthma in children. Curr. Opin. Allergy Clin. Immunol. 2014, 14, 155–160. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Kong, H.; Zeng, X.; Guo, L.; Sun, X.; He, S. Subsets of regulatory T cells and their roles in allergy. J. Transl. Med. 2014, 12. [Google Scholar] [CrossRef] [PubMed]
- Powell, J.D.; Delgoffe, G.M. The mammalian target of rapamycin: Linking T cell differentiation, function, and metabolism. Immunity 2010, 33, 301–311. [Google Scholar] [CrossRef] [PubMed]
- Delgoffe, G.M.; Kole, T.P.; Zheng, Y.; Zarek, P.E.; Matthews, K.L.; Xiao, B.; Worley, P.F.; Kozma, S.C.; Powell, J.D. The mTOR kinase differentially regulates effector and regulatory T cell lineage commitment. Immunity 2009, 30, 832–844. [Google Scholar] [CrossRef] [PubMed]
- Cobbold, S.P.; Adams, E.; Farquhar, C.A.; Nolan, K.F.; Howie, D.; Lui, K.O.; Fairchild, P.J.; Mellor, A.L.; Ron, D.; Waldmann, H. Infectious tolerance via the consumption of essential amino acids and mTOR signaling. Proc. Natl. Acad. Sci. USA 2009, 106, 12055–12060. [Google Scholar] [CrossRef] [PubMed]
- Haxhinasto, S.; Mathis, D.; Benoist, C. The AKT-mTOR axis regulates de novo differentiation of CD4+FoxP3+ cells. J. Exp. Med. 2008, 205, 565–574. [Google Scholar] [CrossRef] [PubMed]
- Sauer, S.; Bruno, L.; Hertweck, A.; Finlay, D.; Leleu, M.; Spivakov, M.; Knight, Z.A.; Cobb, B.S.; Cantrell, D.; O’Connor, E.; et al. T cell receptor signaling controls FoxP3 expression via PI3K, Akt, and mTOR. Proc. Natl. Acad. Sci. USA 2008, 105, 7797–7802. [Google Scholar] [CrossRef] [PubMed]
- Josefowicz, S.Z.; Lu, L.F.; Rudensky, A.Y. Regulatory T cells: Mechanisms of differentiation and function. Annu. Rev. Immunol. 2012, 30, 531–564. [Google Scholar] [CrossRef] [PubMed]
- Xu, G.; Xia, J.; Hua, X.; Zhou, H.; Yu, C.; Liu, Z.; Cai, K.; Shi, J.; Li, H. Activated mammalian target of rapamycin is associated with T regulatory cell insufficiency in nasal polyps. Respir. Res. 2009, 10. [Google Scholar] [CrossRef] [PubMed]
- Melnik, B.C. The potential mechanistic link between allergy and obesity development and infant formula feeding. Allergy Asthma Clin. Immunol. 2014, 10. [Google Scholar] [CrossRef] [PubMed]
- Sawant, D.V.; Yao, W.; Wright, Z.; Sawyers, C.; Tepper, R.S.; Gupta, S.K.; Kaplan, M.H.; Dent, A.L. Serum microRNA-21 as a biomarker for allergic inflammatory disease in children. Microrna 2015, in press. [Google Scholar]
- Melnik, B.C. Evidence for acne-promoting effects of milk and other insulinotropic dairy products. Nestle Nutr. Workshop Ser. Pediatr. Program 2011, 67, 131–145. [Google Scholar] [PubMed]
- Melnik, B.C. Linking diet to acne metabolomics, inflammation, and comedogenesis: An update. Clin. Cosmet. Investig. Dermatol. 2015, 8, 371–388. [Google Scholar] [CrossRef] [PubMed]
- Melnik, B.C.; John, S.M.; Plewig, G. Acne: Risk indicator for increased body mass index and insulin resistance. Acta Derm. Venereol. 2013, 93, 644–649. [Google Scholar] [CrossRef] [PubMed]
- Melnik, B.C. The role of mTORC1 in acne pathogenesis and treatment. Exp. Rev. Dermatol. 2013, 8, 617–622. [Google Scholar] [CrossRef]
- Di Landro, A.; Cazzaniga, S.; Parazzini, F.; Ingordo, V.; Cusano, F.; Atzori, L.; Cutrì, F.T.; Musumeci, M.L.; Zinetti, C.; Pezzarossa, E.; et al. Family history, body mass index, selected dietary factors, menstrual history, and risk of moderate to severe acne in adolescents and young adults. J. Am. Acad. Dermatol. 2012, 67, 1129–1135. [Google Scholar] [CrossRef] [PubMed]
- Grossi, E.; Cazzaniga, S.; Crotti, S.; Naldi, L.; di Landro, A.; Ingordo, V.; Cusano, F.; Atzori, L.; Tripodi Cutrì, F.; Musumeci, M.L.; et al. The constellation of dietary factors in adolescent acne: A semantic connectivity map approach. J. Eur. Acad. Dermatol. Venereol. 2014. [Google Scholar] [CrossRef] [PubMed]
- Burris, J.; Rietkerk, W.; Woolf, K. Relationships of self-reported dietary factors and perceived acne severity in a cohort of New York young adults. J. Acad. Nutr. Diet. 2014, 114, 384–392. [Google Scholar] [CrossRef] [PubMed]
- Cordain, L.; Lindeberg, S.; Hurtado, M.; Hill, K.; Eaton, S.B.; Brand-Miller, J. Acne vulgaris: A disease of Western civilization. Arch. Dermatol. 2002, 138, 1584–1590. [Google Scholar] [CrossRef] [PubMed]
- Wiley, A.S. Milk intake and total dairy consumption: Associations with early menarche in NHANES 1999–2004. PLoS ONE 2011, 6, e14685. [Google Scholar] [CrossRef] [PubMed]
- Gomes, M.B.; Negrato, C.A.; Calliari, L.E.; Brazilian Type 1 Diabetes Study Group (BrazDiab1SG). Early age at menarche: a risk factor for overweight or obesity in patients with type 1 diabetes living in urban areas? Diabetes Res. Clin. Pract. 2015, 107, 23–30. [Google Scholar] [CrossRef] [PubMed]
- Janghorbani, M.; Mansourian, M.; Hosseini, E. Systematic review and meta-analysis of age at menarche and risk of type 2 diabetes. Acta Diabetol. 2014, 51, 519–528. [Google Scholar] [CrossRef] [PubMed]
- Lim, J.S.; Lee, H.S.; Kim, E.Y.; Yi, K.H.; Hwang, J.S. Early menarche increases the risk of type 2 diabetes in young and middle-aged Korean women. Diabet. Med. 2015, 32, 521–525. [Google Scholar] [CrossRef] [PubMed]
- Sluijs, I.; Forouhi, N.G.; Beulens, J.W.; van der Schouw, Y.T.; Agnoli, C.; Arriola, L.; Balkau, B.; Barricarte, A.; Boeing, H.; Bueno-de-Mesquita, H.B.; et al. The amount and type of dairy product intake and incident type 2 diabetes: Results from the EPIC-InterAct Study. Am. J. Clin. Nutr. 2012, 96, 382–390. [Google Scholar] [CrossRef] [PubMed]
- Bartolome, A.; Guillén, C. Role of the mammalian target of rapamycin (mTOR) complexes in pancreatic β-cell mass regulation. Vitam. Horm. 2014, 95, 425–469. [Google Scholar] [PubMed]
- Back, S.H.; Kang, S.W.; Han, J.; Chung, H.A.T. Endoplasmic reticulum stress in the β-cell cell pathogenesis of type 2 diabetes. Exp. Diabetes Res. 2012, 2012. [Google Scholar] [CrossRef] [PubMed]
- Back, S.H.; Kaufman, R.J. Endoplasmic reticulum stress and type 2 diabetes. Annu. Rev. Biochem. 2012, 81, 767–793. [Google Scholar] [CrossRef] [PubMed]
- Reiling, J.H.; Sabatini, D.M. Increased mTORC1 singaling UPRegulates stress. Mol. Cell 2008, 29, 533–535. [Google Scholar] [CrossRef] [PubMed]
- Kato, H.; Nakajima, S.; Saito, Y.; Takahashi, S.; Katoh, R.; Kitamura, M. mTORC1 serves ER stress-triggered apoptosis via selective activation of the IRE-JNK pathway. Cell Death Differ. 2012, 19, 310–320. [Google Scholar] [CrossRef] [PubMed]
- Ozcan, U.; Ozcan, L.; Yilmaz, E.; Duvel, K.; Sahin, M.; Manning, B.D.; Hotamisligil, G.S. Loss of the tuberous sclerosis complex tumor suppressors triggers the unfolded protein response to regulate insulin singaling and apoptosis. Mol. Cell 2008, 29, 541–551. [Google Scholar] [CrossRef] [PubMed]
- Appenzeller-Herzog, C.; Hall, M.N. Bidirectional crosstalk between endoplasmic reticulum stress and mTOR signaling. Trends Cell Biol. 2012, 22, 274–282. [Google Scholar] [CrossRef] [PubMed]
- Guan, B.J.; Krokowski, D.; Majumder, M.; Schmotzer, C.L.; Kimball, S.R.; Merrick, W.C.; Koromilas, A.E.; Hatzolou, M. Translational control during endoplasmic reticulum stress beyond phosphorylation of the translation initiation factor eIF2α. J. Biol. Chem. 2014, 289, 12593–12611. [Google Scholar] [CrossRef] [PubMed]
- Han, J.; Back, S.H.; Hur, J.; Lin, Y.H.; Gildersleeve, R.; Shan, J.; Yuan, C.L.; Krokowski, D.; Wang, S.; Hatzoglou, M.; et al. ER-stress-induced transcriptional regulation increases protein synthesis leading to cell death. Nat. Cell Biol. 2013, 15, 481–490. [Google Scholar] [CrossRef] [PubMed]
- Elghazi, L.; Balcazar, N.; Blandino-Rosano, M.; Cras-Méneur, C.; Fatrai, S.; Gould, A.P.; Chi, M.M.; Moley, K.H.; Bernal-Mizrachi, E. Decreased IRS signaling impairs β-cell cycle progression and survival in transgenic mice overexpressing S6K in β-cells. Diabetes 2010, 59, 2390–2399. [Google Scholar] [CrossRef] [PubMed]
- Yamaguchi, S.; Ishihara, H.; Yamada, T.; Tamura, A.; Usui, M.; Tominaga, R.; Munakata, Y.; Satake, C.; Katagiri, H.; Tashiro, F.; et al. ATF4-mediated induction of 4E-BP1 contributes to pancreatic β cell survival under endoplasmic reticulum stress. Cell Metab. 2008, 7, 269–276. [Google Scholar] [CrossRef] [PubMed]
- Krokowski, D.; Han, J.; Saikia, M.; Majumder, M.; Yuan, C.L.; Guan, B.J.; Bevilacqua, E.; Bussolati, O.; Bröer, S.; Arvan, P.; et al. A self-defeating anabolic program leads to β-cell apoptosis in endoplasmic reticulum stress-induced diabetes via regulation of amino acid flux. J. Biol. Chem. 2013, 288, 17202–17213. [Google Scholar] [CrossRef] [PubMed]
- Shigeyama, Y.; Kobayashi, T.; Kido, Y.; Hashimoto, N.; Asahara, S.; Matsuda, T.; Takeda, A.; Inoue, T.; Shibutani, Y.; Koyanagi, M.; et al. Biphasic response of pancreatic β-cell mass to ablation of tuberous sclerosis complex 2 in mice. Mol. Cell. Biol. 2008, 28, 2971–2979. [Google Scholar] [CrossRef] [PubMed]
- Hsieh, A.C.; Liu, Y.; Edlind, M.P.; Ingolia, N.T.; Janes, M.R.; Sher, A.; Shi, E.Y.; Stumpf, C.R.; Christensen, C.; Bonham, M.J.; et al. The translational landscape of mTOR signalling steers cancer initiation and metastasis. Nature 2012, 485, 55–61. [Google Scholar] [CrossRef] [PubMed]
- Melnik, B.C.; John, S.M.; Carrera-Bastos, P.; Cordain, L. The impact of cow’s milk-mediated mTORC1-signaling in the initiation and progression of prostate cancer. Nutr. Metab. 2012, 9. [Google Scholar] [CrossRef] [PubMed]
- Edlind, M.P.; Hsieh, A.C. PI3K-AKT-mTOR signaling in prostate cancer progression and androgen deprivation therapy resistance. Asian J. Androl. 2014, 16, 378–386. [Google Scholar] [PubMed]
- Sutherland, S.I.; Pe Benito, R.; Henshall, S.M.; Horvath, L.G.; Kench, J.G. Expression of phosphorylated-mTOR during the development of prostate cancer. Prostate 2014, 74, 1231–1239. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Cheng, H.; Pan, T.; Liu, Y.; Su, Y.; Ren, C.; Huang, D.; Zha, X.; Liang, C. mTOR regulate EMT through RhoA and Rac1 pathway in prostate cancer. Mol. Carcinog. 2014. [Google Scholar] [CrossRef] [PubMed]
- Marqués, N.; Sesé, M.; Cánovas, V.; Valente, F.; Bermudo, R.; de Torres, I.; Fernández, Y.; Abasolo, I.; Fernández, P.L.; Contreras, H.; et al. Regulation of protein translation and c-Jun expression by prostate tumor overexpressed 1. Oncogene 2014, 33, 1124–1134. [Google Scholar] [CrossRef] [PubMed]
- Song, Y.; Chavarro, J.E.; Cao, Y.; Qiu, W.; Mucci, L.; Sesso, H.D.; Stampfer, M.J.; Giovannucci, E.; Pollak, M.; Liu, S.; et al. Whole milk intake is associated with prostate cancer-specific mortality among U.S. male physicians. J. Nutr. 2013, 143, 189–196. [Google Scholar] [CrossRef] [PubMed]
- Allen, N.E.; Key, T.J.; Appleby, P.N.; Travis, R.C.; Roddam, A.W.; Tjønneland, A.; Johnsen, N.F.; Overvad, K.; Linseisen, J.; Rohrmann, S.; et al. Animal foods, protein, calcium and prostate cancer risk: The European Prospective Investigation into Cancer and Nutrition. Br. J. Cancer 2008, 98, 1574–1581. [Google Scholar] [CrossRef] [PubMed]
- Torfadottir, J.E.; Steingrimsdottir, L.; Mucci, L.; Aspelund, T.; Kasperzyk, J.L.; Olafsson, O.; Fall, K.; Tryggvadottir, L.; Harris, T.B.; Launer, L.; et al. Milk intake in early life and risk of advanced prostate cancer. Am. J. Epidemiol. 2012, 175, 144–153. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, S.; Lau, H.; Simons, B.W.; Powell, J.D.; Meyers, D.J.; de Marzo, A.M.; Berman, D.M.; Lotan, T.L. PI3K/mTOR signaling regulates prostatic branching morphogenesis. Dev. Biol. 2011, 360, 329–342. [Google Scholar] [CrossRef] [PubMed]
- Ribas, J.; Lupold, S.E. The transcriptional regulation of miR-21, its multiple transcripts, and their implication in prostate cancer. Cell Cycle 2010, 9, 923–929. [Google Scholar] [CrossRef] [PubMed]
- Mishra, S.; Lin, C.L.; Huang, T.H.; Bouamar, H.; Sun, L.Z. MicroRNA-21 inhibits p57Kip2 expression in prostate cancer. Mol. Cancer 2014, 13, 212. [Google Scholar] [CrossRef] [PubMed]
- Leite, K.R.; Reis, S.T.; Viana, N.; Morais, D.R.; Moura, C.M.; Silva, I.A.; Pontes, J., Jr.; Katz, B.; Srougi, M. Controlling RECK miR21 promotes tumor cell invasion and is related to biochemical recurrence in prostate cancer. J. Cancer 2015, 6, 292–301. [Google Scholar] [CrossRef] [PubMed]
- Mishra, S.; Deng, J.J.; Gowda, P.S.; Rao, M.K.; Lin, C.L.; Chen, C.L.; Huang, T.; Sun, L.Z. Androgen receptor and microRNA-21 axis downregulates transforming growth factor β receptor II (TGFBR2) expression in prostate cancer. Oncogene 2014, 33, 4097–4106. [Google Scholar] [CrossRef] [PubMed]
- Tate, P.L.; Bibb, R.; Larcom, L.L. Milk stimulates growth of prostate cancer cells in culture. Nutr. Cancer 2011, 63, 1361–1366. [Google Scholar] [CrossRef] [PubMed]
- Bernichtein, S.; Pigat, N.; Capiod, T.; Boutillon, F.; Verkarre, V.; Camparo, P.; Viltard, M.; Méjean, A.; Oudard, S.; Souberbielle, J.C.; et al. High milk consumption does not affect prostate tumor progression in two mouse models of benign and neoplastic lesions. PLoS ONE 2015, 10, e0125423. [Google Scholar] [CrossRef] [PubMed]
- Gaard, M.; Tretli, S.; Løken, E.B. Dietary fat and the risk of breast cancer; A prospective study of 25,892 Norwegian women. Int. J. Cancer 1995, 63, 13–17. [Google Scholar] [CrossRef] [PubMed]
- Qin, L.Q.; Xu, J.Y.; Tezuka, H.; Li, J.; Arita, J.; Hoshi, K.; Sato, A. Consumption of commercial whole and non-fat milk increases the incidence of 7,12-dimethylbenz(α)anthracene-induced mammary tumors in rats. Cancer Detect. Prev. 2007, 31, 339–343. [Google Scholar] [CrossRef] [PubMed]
- Qin, L.Q.; Xu, J.Y.; Tezuka, H.; Wang, P.Y.; Hoshi, K. Milk inhibits the regression of 7,12-dimethylbenz(α)anthracene-induced mammary tumors in ovariectomized rats. Nutr. Cancer 2008, 60, 505–510. [Google Scholar] [CrossRef] [PubMed]
- Macias, H.; Hinck, L. Mammary gland development. Wiley Interdiscip. Rev. Dev. Biol. 2012, 1, 533–557. [Google Scholar]
- Martin, L.J.; Boyd, N.F. Mammographic density. Potential mechanisms of breast cancer risk associated with mammographic density: Hypotheses based on epidemiological evidence. Breast Cancer Res. 2008, 10. [Google Scholar] [CrossRef] [PubMed]
- Frydenberg, H.; Flote, V.G.; Iversen, A.; Finstad, S.E.; Furberg, A.S.; Torjesen, P.A.; Wilsgaard, T.; Schlichting, E.; Ellison, P.T.; Ursin, G.; et al. Insulin-like growth factor-1, growth hormone, and daily cycling estrogen are associated with mammographic density in premenopausal women. Cancer Causes Control. 2014, 25, 891–903. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Wu, Z.; Chen, H.; Zhu, Q.; Gao, G.; Hu, L.; Negi, H.; Kamle, S.; Li, D. Induction of anterior gradient 2 (AGR2) plays a key role in insulin-like growth factor-1 (IGF-1)-induced breast cancer cell proliferation and migration. Med. Oncol. 2015, 32, 577. [Google Scholar] [CrossRef] [PubMed]
- Renehan, A.G.; Zwahlen, M.; Minder, C.; O’Dwyer, S.T.; Shalet, S.M.; Egger, M. Insulin-like growth factor (IGF)-I, IGF binding protein-3, and cancer risk: Systematic review and meta-regression analysis. Lancet 2004, 363, 1346–1353. [Google Scholar] [CrossRef]
- Cao, Y.; Nimptsch, K.; Shui, I.M.; Platz, E.A.; Wu, K.; Pollak, M.N.; Kenfield, S.A.; Stampfer, M.J.; Giovannucci, E.L. Prediagnostic plasma IGFBP-1, IGF-1 and risk of prostate cancer. Int. J. Cancer 2015, 136, 2418–2426. [Google Scholar] [CrossRef] [PubMed]
- Renehan, A.G.; Harvie, M.; Howell, A. Insulin-like growth factor (IGF)-I, IGF binding protein-3, and breast cancer risk: Eight years on. Endocr. Relat. Cancer 2006, 13, 273–278. [Google Scholar] [CrossRef] [PubMed]
- Christopoulos, P.F.; Msaouel, P.; Koutsilieris, M. The role of the insulin-like growth factor-1 system in breast cancer. Mol. Cancer 2015, 14. [Google Scholar] [CrossRef] [PubMed]
- Worster, D.T.; Schmelzle, T.; Solimini, N.L.; Lightcap, E.S.; Millard, B.; Mills, G.B.; Brugge, J.S.; Albeck, J.G. Akt and ERK control the proliferative response of mammary epithelial cells to the growth factors IGF-1 and EGF through the cell cycle inhibitor p57Kip2. Sci. Signal. 2012, 5, ra19. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Yang, X.; Yang, J.; Zhen, J.; Zhang, D. Serum microRNA-21 as a potential diagnostic biomarker for breast cancer: A systematic review and meta-analysis. Clin. Exp. Med. 2014. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Wang, L.; Sun, S.; Wu, J.; Wang, Q. Quantitative measurement of serum microRNA-21 expression in relation to breast cancer metastasis in Chinese females. Ann. Lab. Med. 2015, 35, 226–232. [Google Scholar] [CrossRef] [PubMed]
- Pan, F.; Mao, H.; Deng, L.; Li, G.; Geng, P. Prognostic and clinicopathological significance of microRNA-21 overexpression in breast cancer: A meta-analysis. Int. J. Clin. Exp. Pathol. 2014, 7, 5622–5633. [Google Scholar] [PubMed]
- Chen, J.; Wang, X. MicroRNA-21 in breast cancer: Diagnostic and prognostic potential. Clin. Transl. Oncol. 2014, 16, 225–233. [Google Scholar] [CrossRef] [PubMed]
- Parvani, J.G.; Taylor, M.A.; Schiemann, W.P. Noncanonical TGF-β signaling during mammary tumorigenesis. J. Mammary Gland Biol. Neoplasia 2011, 16, 127–146. [Google Scholar] [CrossRef] [PubMed]
- Wazir, U.; Newbold, R.F.; Jiang, W.G.; Sharma, A.K.; Mokbel, K. Prognostic and therapeutic implications of mTORC1 and Rictor expression in human breast cancer. Oncol. Rep. 2013, 29, 1969–1974. [Google Scholar] [PubMed]
- Calip, G.S.; Yu, O.; Hoskins, K.F.; Boudreau, D.M. Associations between diabetes medication use and risk of second breast cancer events and mortality. Cancer Causes Control 2015, 26, 1065–1077. [Google Scholar] [CrossRef] [PubMed]
- Gao, S.; Jiang, J.; Li, P.; Song, H.; Wang, W.; Li, C.; Kong, D. Attenuating tumour angiogenesis: A preventive role of metformin against breast cancer. Biomed. Res. Int. 2015, 2015. [Google Scholar] [CrossRef] [PubMed]
- Hatoum, D.; McGowan, E.M. Recent advances in the use of metformin: Can treating diabetes prevent breast cancer? Biomed. Res. Int. 2015, 2015. [Google Scholar] [CrossRef] [PubMed]
- Melnik, B.C.; Schmitz, G. Metformin: An inhibitor of mTORC1 signaling. J. Endocrinol. Diabetes Obes. 2014, 2, 1029. [Google Scholar]
- Campdelacreu, J. Parkinson disease and Alzheimer disease: Environmental risk factors. Neurologia 2014, 29, 541–549. [Google Scholar] [CrossRef] [PubMed]
- Jiang, W.; Ju, C.; Jiang, H.; Zhang, D. Dairy foods intake and risk of Parkinson’s disease: A dose-response meta-analysis of prospective cohort studies. Eur. J. Epidemiol. 2014, 29, 613–619. [Google Scholar] [CrossRef] [PubMed]
- Kyrozis, A.; Ghika, A.; Stathopoulos, P.; Vassilopoulos, D.; Trichopoulos, D.; Trichopoulou, A. Dietary and lifestyle variables in relation to incidence of Parkinson’s disease in Greece. Eur. J. Epidemiol. 2013, 28, 67–77. [Google Scholar] [CrossRef] [PubMed]
- Perluigi, M.; di domenico, F.; Butterfield, D.A. mTOR signaling in aging and neurodegeneration: At the crossroad between metabolism dysfunction and impairment of autophagy. Neurobiol. Dis. 2015, in press. [Google Scholar] [CrossRef] [PubMed]
- Tenreiro, S.; Eckermann, K.; Outeiro, T.F. Protein phosphorylation in neurodegeneration: Friend or foe? Front. Mol. Neurosci. 2014, 7. [Google Scholar] [CrossRef] [PubMed]
- Caccamo, A.; Magrì, A.; Medina, D.X.; Wisely, E.V.; López-Aranda, M.F.; Silva, A.J.; Oddo, S. mTOR regulates tau phosphorylation and degradation: Implications for Alzheimer’s disease and other tauopathies. Aging Cell 2013, 12, 370–380. [Google Scholar] [CrossRef] [PubMed]
- Spilman, P.; Podlutskaya, N.; Hart, M.J.; Debnath, J.; Gorostiza, O.; Bredesen, D.; Richardson, A.; Strong, R.; Galvan, V. Inhibition of mTOR by rapamycin abolishes cognitive deficits and reduces amyloid-β levels in a mouse model of Alzheimer’s disease. PLoS ONE 2010, 5, e9979. [Google Scholar] [CrossRef] [PubMed]
- Richardson, A.; Galvan, V.; Lin, A.L.; Oddo, S. How longevity research can lead to therapies for Alzheimer’s disease: The rapamycin story. Exp. Gerontol. 2015, 68, 51–58. [Google Scholar] [CrossRef] [PubMed]
- Kickstein, E.; Krauss, S.; Thornhill, P.; Rutschow, D.; Zeller, R.; Sharkey, J.; Williamson, R.; Fuchs, M.; Köhler, A.; Glossmann, H.; et al. Biguanide metformin acts on tau phosphorylation via mTOR/protein phosphatase 2A (PP2A) signaling. Proc. Natl. Acad. Sci. USA 2010, 107, 21830–21835. [Google Scholar] [CrossRef] [PubMed]
- Melnik, B.C. Formula feeding promotes adipogenic, diabetogenic, hypertonic and allergic mTORC1-programming. In Handbook of Dietary and Nutritional Aspects of Bottle Feeding, 1st ed.; Preedy, V.R., Watson, R.R., Zibadi, S., Eds.; Wagenigen Academic Publishers: Wageningen, The Netherlands, 2014; pp. 545–568. [Google Scholar]
- Gould, S.J.; Booth, A.M.; Hildreth, J.E. The Trojan exosome hypothesis. Proc. Natl. Acad. Sci. USA 2003, 100, 10592–10597. [Google Scholar] [CrossRef] [PubMed]
- Modepalli, V.; Kumar, A.; Hinds, L.A.; Sharp, J.A.; Nicholas, K.R.; Lefevre, C. Differential temporal expression of milk miRNA during the lactation cycle of the marsupial tammar wallaby (Macropus eugenii). BMC Genomics 2014, 15, 1012. [Google Scholar] [CrossRef] [PubMed]
- Marriott, M.; Schoenthal, J. An experimental study of the use of unsweetened evaporated milk for the preparation of infant feeding formulas. Arch. Pediatr. 1929, 46, 135–148. [Google Scholar]
- Bryder, L. From breast to bottle: A history of modern infant feeding. Endeavour 2009, 54–59. [Google Scholar] [CrossRef] [PubMed]
- McCay, C.M.; Crowell, M.F. Prolonging the life span. Sci. Mon. 1934, 39, 405–414. [Google Scholar]
- Segall, J.J. Hypothesis: Is lactose a dietary risk factor for ischaemic heart disease? Int. J. Epidemiol. 2008, 37, 1204–1208. [Google Scholar] [CrossRef] [PubMed]
- Ji, J.; Sundquist, J.; Sundquist, K. Lactose intolerance and risk of lung, breast and ovarian cancers: Aetiological clues from a population-based study in Sweden. Br. J. Cancer 2015, 112, 149–152. [Google Scholar] [CrossRef] [PubMed]
- Bannister, C.A.; Holden, S.E.; Jenkins-Jones, S.; Morgan, C.L.; Halcox, J.P.; Schernthaner, G.; Mukherjee, J.; Currie, C.J. Can people with type 2 diabetes life longer than those without? A comparison of mortality in people initiatd with metformin or sulphonylurea monotherapy and matched, non-diabetic controls. Diabetes Obes. Metab. 2014, 16, 1165–1173. [Google Scholar] [CrossRef] [PubMed]
- Blagosklonny, M.V. Revisiting the antagonistic peiotropy theory of aging. TOR-driven program and quasi-program. Cell Cycle 2010, 9, 3151–3156. [Google Scholar] [CrossRef] [PubMed]
- Schooling, C.M. Milk and mortality. BMJ 2014, 349, g6205. [Google Scholar] [CrossRef] [PubMed]
- Dellago, H.; Preschitz-Kammerhofer, B.; Terlecki-Zaniewicz, L.; Schreiner, C.; Fortschegger, K.; Chang, M.W.; Hackl, M.; Monteforte, R.; Kühnel, H.; Schosserer, M.; et al. High levels of oncomiR-21 contribute to the senescence-induced growth arrest in normal human cells and its knock-down increases the replicative lifespan. Aging Cell 2013, 12, 446–458. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.Y.; Sun, G.; Luo, H.; Wang, X.F.; Lan, F.M.; Yue, X.; Fu, L.S.; Pu, P.Y.; Kang, C.S.; Liu, N.; et al. MiR-21 modulates hTERT through a STAT3-dependent manner on glioblastoma cell growth. CNS Neurosci. Ther. 2012, 18, 722–728. [Google Scholar] [CrossRef] [PubMed]
- Olivieri, F.; Spazzafumo, L.; Santini, G.; Lazzarini, R.; Albertini, M.C.; Rippo, M.R.; Galeazzi, R.; Abbatecola, A.M.; Marcheselli, F.; Monti, D.; et al. Age-related differences in the expression of circulating microRNAs: miR-21 as a new circulating marker of inflammaging. Mech. Ageing Dev. 2012, 133, 675–685. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y.; Zhang, C. MicroRNA-21 in cardiovascular disease. J. Cardiovasc. Transl. Res. 2010, 3, 251–255. [Google Scholar] [CrossRef] [PubMed]
- Hulsmans, M.; Holvoet, P. MicroRNA-containing microvesicles regulating inflammation in association with atherosclerotic diesease. Cardiovasc. Res. 2013, 100, 7–18. [Google Scholar] [CrossRef] [PubMed]
- Kang, M.; Yan, L.M.; Zhang, W.Y.; Li, Y.M.; Tang, A.Z.; Ou, H.S. Role of microRNA-21 in regulating 3T3-L1 adipocyte differentiation and adiponectin expression. Mol. Biol. Rep. 2013, 40, 5027–5034. [Google Scholar] [CrossRef] [PubMed]
- Keller, P.; Gburcik, V.; Petrovic, N.; Gallagher, I.J.; Nedergaard, J.; Cannon, B.; Timmons, J.A. Gene-chip studies of adipogenesis-regulated microRNAs in mouse primary adipocytes and human obesity. BMC Endocr. Disord. 2011, 11, 7. [Google Scholar] [CrossRef] [PubMed]
- Seeger, T.; Fischer, A.; Muhly-Reinholz, M.; Zeiher, A.M.; Dimmeler, S. Long-term inhibition of miR-21 leads to reduction of obesity in db/db mice. Obesity (Silver Spring) 2014, 22, 2352–2360. [Google Scholar] [CrossRef] [PubMed]
- Medawar, P.B. An Unsolved Problem in Biology; HK Lewis: London, UK, 1952. [Google Scholar]
- Williams, G.C. Pleiotropy, natural selection and the evolution of senescence. Evolution 1957, 11, 398–411. [Google Scholar] [CrossRef]
- Blagosklonny, M.V. Aging is not programmed: Genetic pseudo-program is a shadow of developmental growth. Cell Cycle 2013, 12, 3736–3742. [Google Scholar] [CrossRef] [PubMed]
- Kapahi, P.; Chen, D.; Rogers, A.N.; Katewa, S.D.; Li, P.W.; Thomas, E.L.; Kockel, L. With TOR less is more: A key role for the conserved nutrient sensing TOR pathway in aging. Cell Metab. 2010, 11, 453–465. [Google Scholar] [CrossRef] [PubMed]
- Alsaweed, M.; Hepworth, A.R.; Lefèvre, C.; Hartmann, P.E.; Geddes, D.T.; Hassiotou, F. Human milk microRNA and total RNA differ depending on milk fractionation. J. Cell. Biochem. 2015, in press. [Google Scholar] [CrossRef] [PubMed]
- Melnik, B.C. Milk-the promoter of chronic Western diseases. Med. Hypotheses 2009, 72, 631–639. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Melnik, B.C. Milk—A Nutrient System of Mammalian Evolution Promoting mTORC1-Dependent Translation. Int. J. Mol. Sci. 2015, 16, 17048-17087. https://doi.org/10.3390/ijms160817048
Melnik BC. Milk—A Nutrient System of Mammalian Evolution Promoting mTORC1-Dependent Translation. International Journal of Molecular Sciences. 2015; 16(8):17048-17087. https://doi.org/10.3390/ijms160817048
Chicago/Turabian StyleMelnik, Bodo C. 2015. "Milk—A Nutrient System of Mammalian Evolution Promoting mTORC1-Dependent Translation" International Journal of Molecular Sciences 16, no. 8: 17048-17087. https://doi.org/10.3390/ijms160817048
APA StyleMelnik, B. C. (2015). Milk—A Nutrient System of Mammalian Evolution Promoting mTORC1-Dependent Translation. International Journal of Molecular Sciences, 16(8), 17048-17087. https://doi.org/10.3390/ijms160817048





