Meta-Analysis of miR-146a Polymorphisms Association with Coronary Artery Diseases and Ischemic Stroke
Abstract
:1. Introduction
2. Results
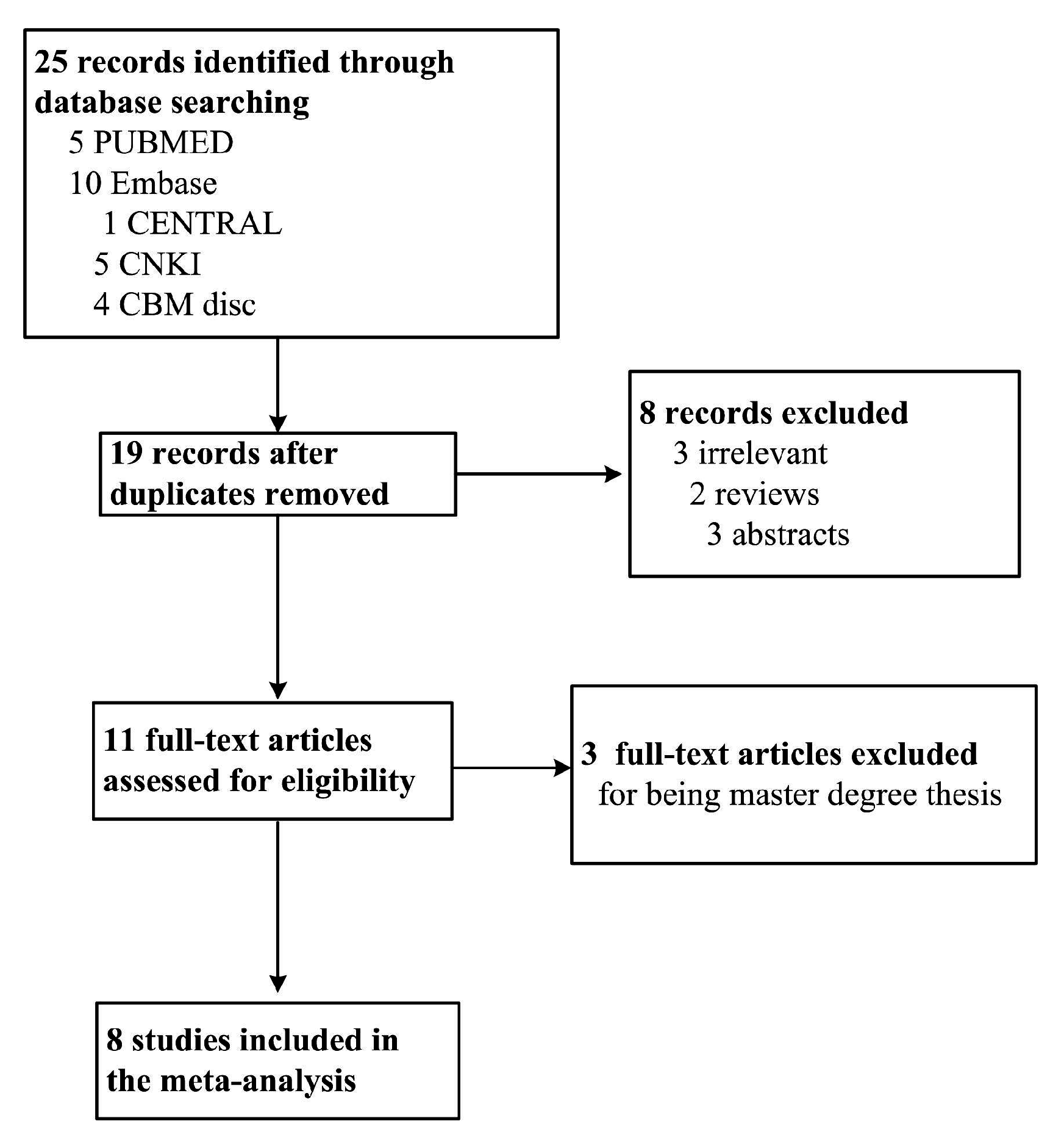
2.1. Characteristics of Eligible Studies
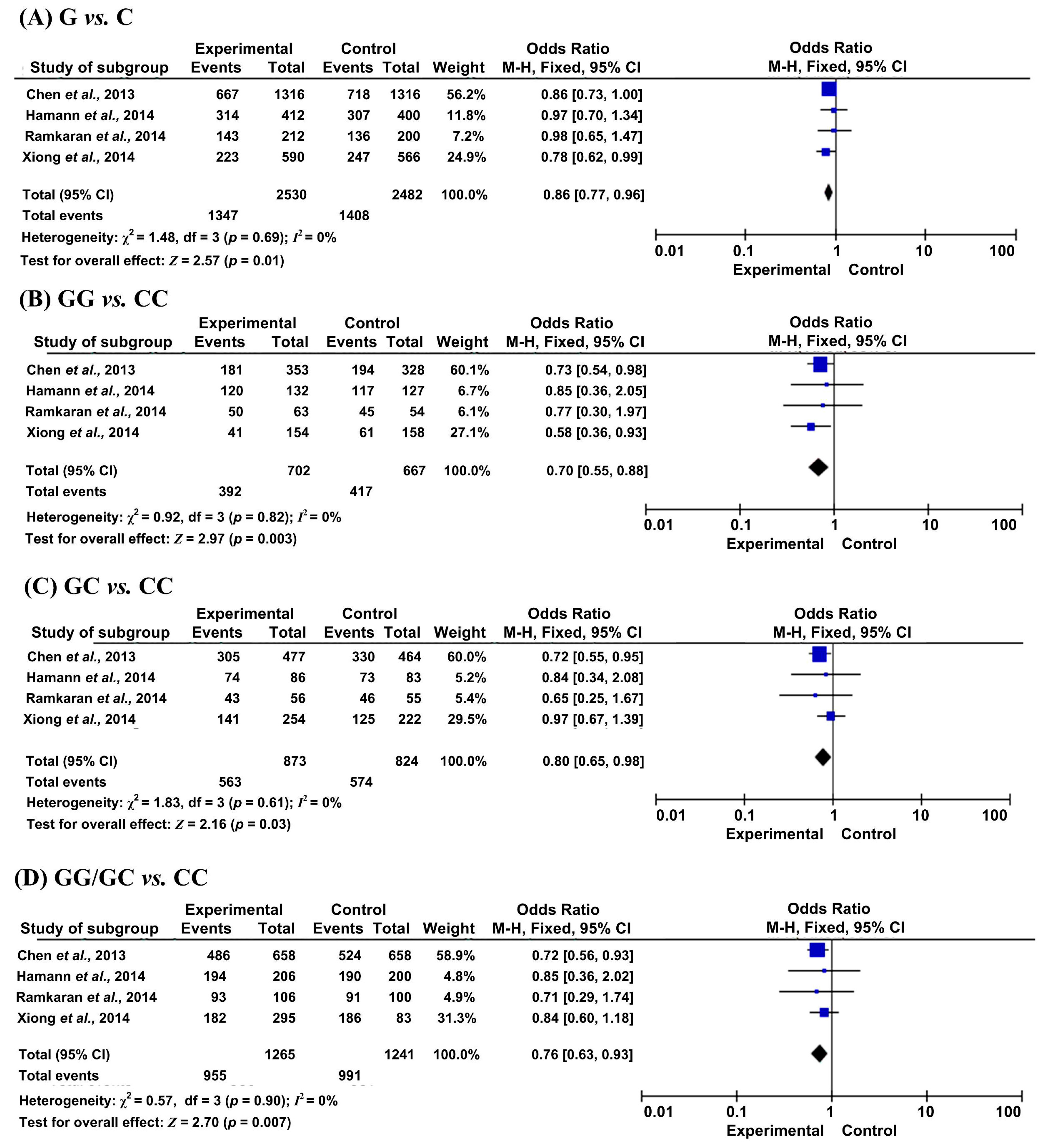
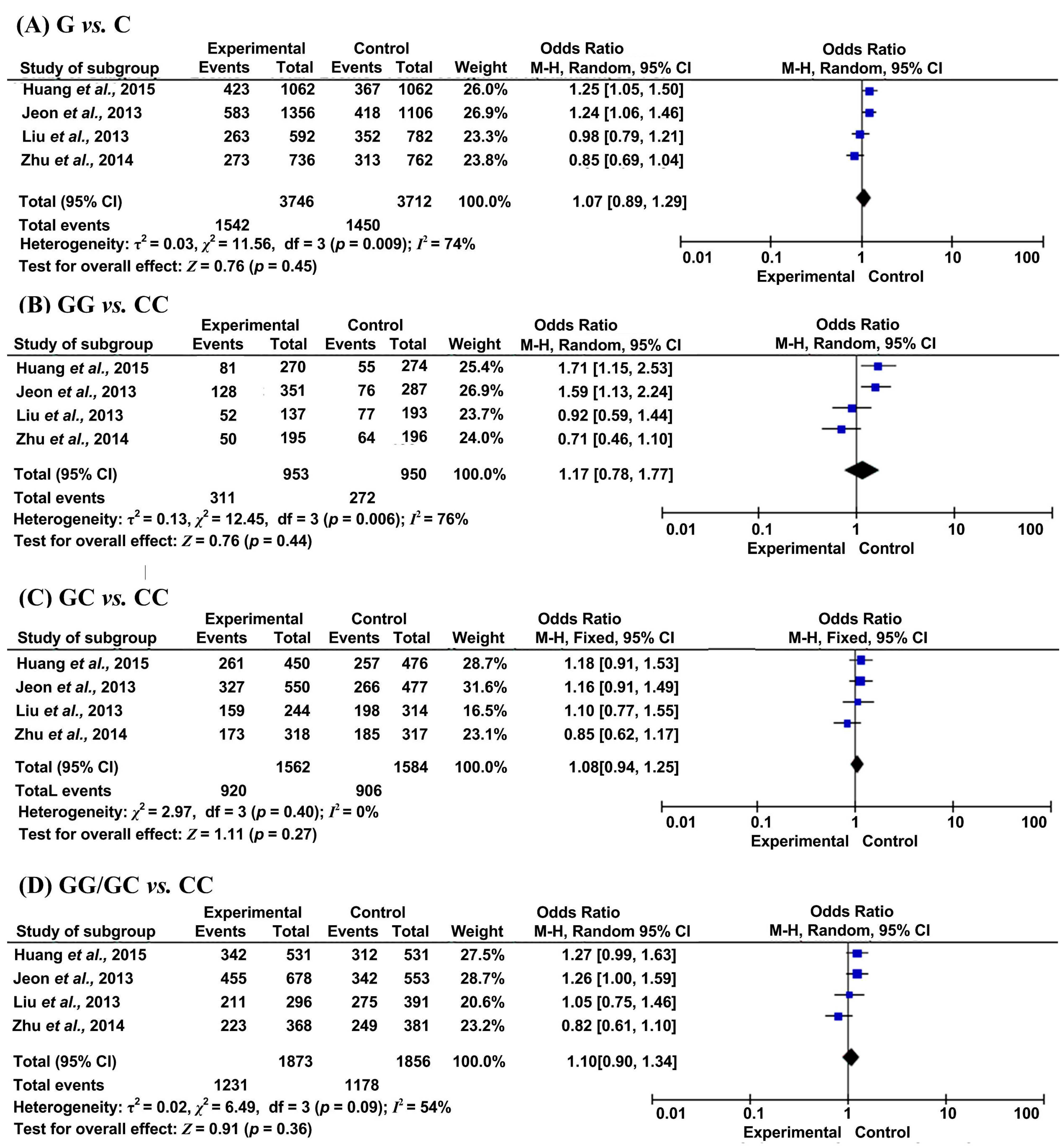
2.2. Results of Meta-Analysis
| Author | Year | Country | Ethnicity | Disease | Genotyping Methods | Sex Ratio (Male:Female) (Case/Control) | Age (Case/Control) | Quality Score | Sample Size (Case/Control) | GG (Case/Control) | GC (Case/Control) | CC (Case/Control) | HWE of Control |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Chen et al. | 2013 | China | Asian | CAD | Taqman | 1:1.79/1:1.184 | 64.57 ± 10.33/ 64.08 ± 11.21 | 14 | 658/658 | 181/194 | 305/330 | 172/134 | 0.769 |
| Hamann et al. | 2014 | German | Caucasian | CAD | HRM | Unavailable | Unavailable | 12 | 206/200 | 120/117 | 74/73 | 12/10 | 0.748 |
| Ramkaran et al. | 2014 | South Africa | Indian | CAD | PCR-RFLP | 1:0/1:0 | 37.6 ± 0.40/ 37.5 ± 0.44 | 12 | 106/100 | 50/45 | 43/46 | 13/9 | 0.569 |
| Xiong et al. | 2014 | Chian | Asian | CAD | PCR-RFLP | 1:0.661/1:0.601 | 65.13 ± 11.86/ 62.59 ± 12.89 | 11 | 295/283 | 41/61 | 141/125 | 113/97 | 0.086 |
| Huang et al. | 2015 | China | Asian | IS | Taqman | 1:0.616/1:0.616 | 61.0 (54, 68)/ 63.0 (54, 70) | 14 | 531/531 | 81/55 | 261/257 | 189/219 | 0.106 |
| Jeon et al. | 2013 | Korea | Korean | IS | Taqman | 1:0.441/1:0.496 | 64.16 ± 11.90/ 63.14 ± 10.19 | 15 | 678/553 | 128/76 | 327/266 | 223/211 | 0.589 |
| Liu et al. | 2013 | China | Asian | IS | PCR-RFLP | 1:0.608/1:0.581 | 67.52 ± 10.29/ 66.34 ± 11.07 | 12 | 296/391 | 52/77 | 159/198 | 85/116 | 0.65 |
| Zhu et al. | 2014 | China | Asian | IS | PCR-LDR | 1:0.688/1:0.685 | 61.62 ± 0.986/ 62.05 ± 0.982 | 12 | 368/381 | 50/64 | 173/185 | 145/132 | 0.952 |
| Genetic Model | Overall or Subgroup | CAD | IS | ||||||
|---|---|---|---|---|---|---|---|---|---|
| n | OR (95% CI) | p-Value | I2 (%) | n | OR (95% CI) | p-Value | I2 (%) | ||
| G vs. C | Overall | 5 | 0.86 (0.77, 0.96) | 0.01 | 0% | 4 | 1.07 (0.89, 1.29) | 0.45 | 74% |
| Large size | 2 | 0.86 (0.73, 1.00) | 0.05 | NA | 2 | 1.25 (1.11, 1.41) | 0.0003 | 0% | |
| Small size | 3 | 0.87 (0.73, 1.03) | 0.10 | 0% | 2 | 0.91 (0.78, 1.05) | 0.20 | 0% | |
| Chinese | 2 | 0.83 (0.73, 0.95) | 0.006 | 0% | 3 | 1.02 (0.80, 1.29) | 0.89 | 76% | |
| Other (Caucasian, Indian, Korean) | 2 | 0.97 (0.75, 0.83) | 0.83 | 0% | 1 | 1.24 (1.06, 1.46) | 0.009 | NA | |
| GG vs. CC | Overall | 4 | 0.70 (0.55, 0.88) | 0.003 | 0% | 4 | 1.17 (0.78, 1.77) | 0.44 | 76% |
| Large size | 1 | 0.73 (0.54, 0.98) | 0.04 | NA | 2 | 1.64 (1.27, 2.12) | 0.0002 | 0% | |
| Small size | 3 | 0.65 (0.44, 0.96) | 0.03 | 0% | 2 | 0.81 (0.59, 1.10) | 0.18 | 0% | |
| Chinese | 2 | 0.68 (0.53, 0.88) | 0.003 | 0% | 3 | 1.05 (0.62, 1.77) | 0.87 | 78% | |
| Other (Caucasian, Indian, Korean) | 2 | 0.81 (0.43, 1.54) | 0.53 | 0% | 1 | 1.59 (1.13, 2.24) | 0.007 | NA | |
| GC vs. CC | Overall | 4 | 0.80 (0.65, 0.98) | 0.03 | 0% | 4 | 1.08 (0.94, 1.25) | 0.27 | 0% |
| Large size | 1 | 0.72 (0.55, 1.39) | 0.02 | NA | 2 | 1.17 (0.98, 1.40) | 0.09 | 0% | |
| Small size | 3 | 0.91 (0.66, 1.25) | 0.56 | 0% | 2 | 0.95 (0.75, 1.22) | 0.71 | 10% | |
| Chinese | 2 | 0.80 (0.64, 1.00) | 0.05 | 39% | 3 | 1.04 (0.86, 1.27) | 0.67 | 20% | |
| Other (Caucasian, Indian, Korean) | 2 | 0.74 (0.39, 1.43) | 0.37 | 0% | 1 | 1.16 (0.91, 1.49) | 0.23 | NA | |
| GG/GC vs. CC | Overall | 4 | 0.76 (0.63, 0.93) | 0.007 | 0% | 4 | 1.10 (0.90, 1.34) | 0.36 | 54% |
| Large size | 1 | 0.72 (0.56, 0.93) | 0.01 | NA | 2 | 1.26 (1.07, 1.50) | 0.007 | 0% | |
| Small size | 3 | 0.83 (0.61, 1.11) | 0.21 | 0% | 2 | 0.91 (0.73, 1.14) | 0.41 | 17% | |
| Chinese | 2 | 0.76 (0.62, 0.94) | 0.01 | 0% | 3 | 1.04 (0.79, 1.35) | 0.79 | 60% | |
| Other (Caucasian, Indian, Korean) | 2 | 0.78 (0.42, 1.45) | 0.43 | 0% | 1 | 1.26 (1.00, 1.59) | 0.05 | NA | |
| Selected Variables | Number of Studies | OR (95% CI) | p-Value | I2 (%) | |
|---|---|---|---|---|---|
| TOAST | LAA | 3 | 0.80 (0.51, 1.27) | 0.34 | 71% |
| SVD | 3 | 1.29 (0.90, 1.84) | 0.17 | 58% | |
| Gender | Male | 2 | 1.06 (0.82, 1.38) | 0.65 | 0% |
| Female | 2 | 1.16 (0.45, 2.97) | 0.76 | 86% | |
| Smoker | Yes | 2 | 1.36 (0.91, 2.03) | 0.14 | 0% |
| No | 2 | 1.09 (0.86, 1.38 ) | 0.86 | 69% | |
| Hypertension | Yes | 2 | 0.87 (0.42, 1.82) | 0.72 | 78% |
| No | 2 | 1.30 (0.97, 1.75) | 0.08 | 0% | |
| Diabetes mellitus | Yes | 2 | 1.76 (0.96, 3.21) | 0.07 | 0% |
| No | 2 | 1.25 (0.95, 1.63) | 0.11 | 0% | |
| Hyperlipidemia | Yes | 2 | 1.16 (0.78, 1.71) | 0.47 | 17% |
| No | 2 | 1.23 (0.80, 1.88) | 0.34 | 42% | |
2.3. Sources of Heterogeneity
2.4. Sensitivity Analysis
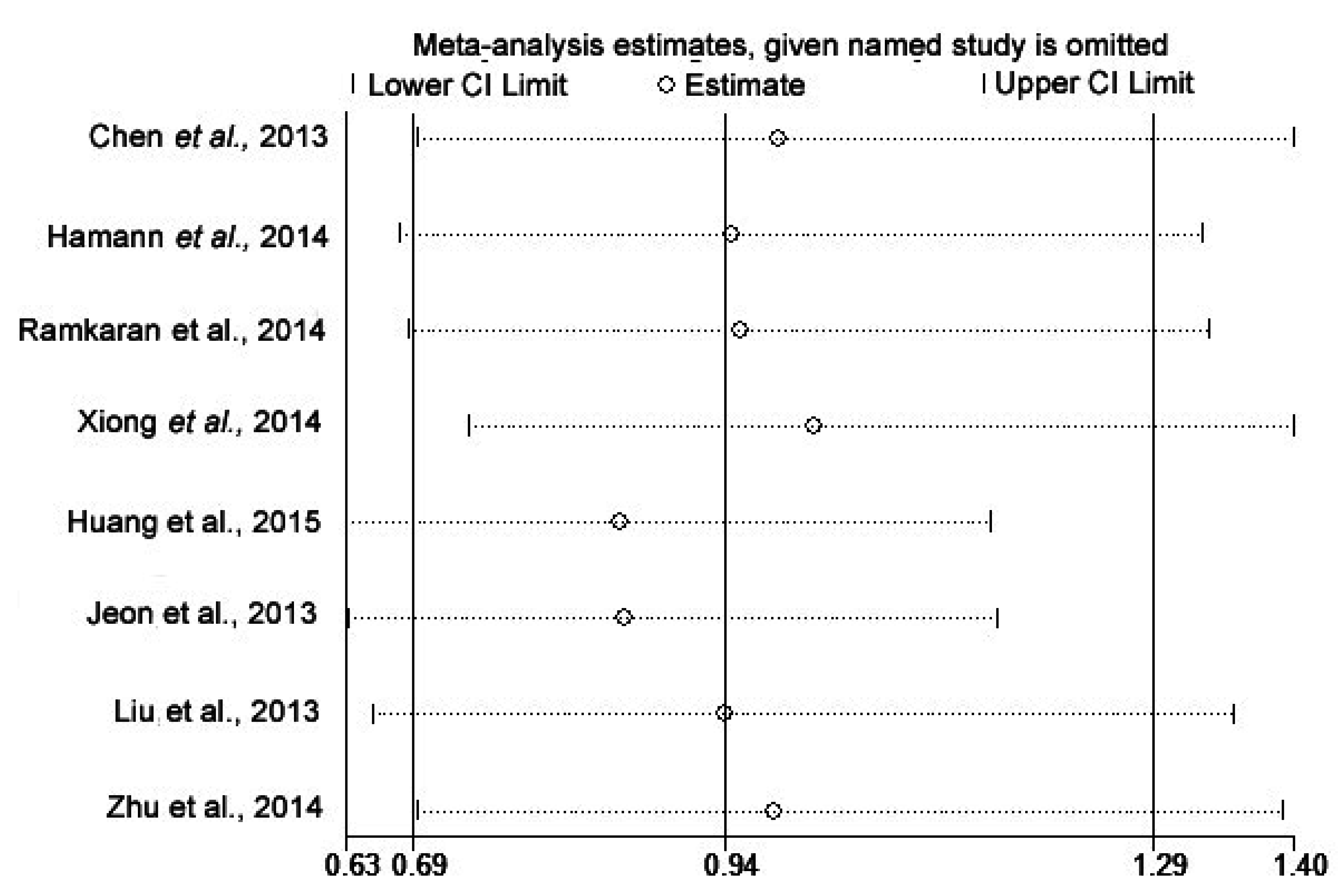
2.5. Publication Bias
| Group | p-Value | |||
|---|---|---|---|---|
| G vs. C | GG vs. CC | GC vs. CC | GG/GC vs. CC | |
| CAD | 0.240 | 0.137 | 0.965 | 0.967 |
| IS | 0.110 | 0.191 | 0.420 | 0.276 |
3. Discussion
4. Methods
4.1. Publication Search Strategy and Inclusion Criteria
4.2. Data Extraction
4.3. Quality Assessment
4.4. Statistical Methods
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Liu, L.; Wang, D.; Wong, K.S.; Wang, Y. Stroke and stroke care in China: Huge burden, significant workload, and a national priority. Stroke 2011, 42, 3651–3654. [Google Scholar] [CrossRef] [PubMed]
- Goldstein, L.B.; Adams, R.; Becker, K.; Furberg, C.D.; Gorelick, P.B.; Hademenos, G.; Hill, M.; Howard, G.; Howard, V.J.; Jacobs, B.; et al. Primary prevention of ischemic stroke: A statement for healthcare professionals from the Stroke Council of the American Heart Association. Stroke 2001, 32, 280–299. [Google Scholar] [CrossRef] [PubMed]
- Marshall, H.W.; Morrison, L.C.; Wu, L.L. Apolipoprotein polymorphisms pail to define risk of coronary-artery disease-results of a prospective, angiographically controlled-study. Circulation 1994, 89, 567–577. [Google Scholar] [CrossRef] [PubMed]
- Fedele, F.; Mancone, M.; Chilian, W.M.; Severino, P.; Canali, E.; Logan, S.; de Marchis, M.L.; Volterrani, M.; Palmirotta, R.; Guadagni, F. Role of genetic polymorphisms of ion channels in the pathophysiology of coronary microvascular dysfunction and ischemic heart disease. Basic Res. Cardiol. 2013, 108, 387. [Google Scholar] [CrossRef] [PubMed]
- Lewis, B.P.; Burge, C.B.; Bartel, D.P. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 2005, 120, 15–20. [Google Scholar] [CrossRef] [PubMed]
- Roldan, V.; Arroyo, A.B.; Salloum-Asfar, S.; Manzano-Fernandez, S.; Garcia-Barbera, N.; Marin, F.; Vicente, V.; Gonzalez-Conejero, R.; Martinez, C. Prognostic role of miR146a polymorphisms for cardiovascular events in atrial fibrillation. Thromb. Haemost. 2014, 112, 781–788. [Google Scholar] [CrossRef] [PubMed]
- Jazdzewski, K.; Murray, E.L.; Franssila, K.; Jarzab, B.; Schoenberg, D.R.; de la Chapelle, A. Common SNP in pre-miR-146a decreases mature miR expression and predisposes to papillary thyroid carcinoma. Proc. Natl. Acad. Sci. USA 2008, 105, 7269–7274. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Wu, Y.T. Association of genetic polymorphisms in microRNAs precursor with the risk and prognosis of coronary heart diseases. J. Xi’an Jiaotong Univ. 2013, 34, 495–499. [Google Scholar]
- Ramkaran, P.; Khan, S.; Phulukdaree, A.; Moodley, D.; Chuturgoon, A.A. miR-146a polymorphism influences levels of miR-146a, IRAK-1, and TRAF-6 in young patients with coronary artery disease. Cell Biochem. Biophys. 2014, 68, 259–266. [Google Scholar] [CrossRef] [PubMed]
- Xiong, X.D.; Cho, M.; Cai, X.P.; Cheng, J.; Jing, X.; Cen, J.M.; Liu, X.; Yang, X.L.; Suh, Y. A common variant in pre-miR-146 is associated with coronary artery disease risk and its mature miRNA expression. Mutat. Res. 2014, 761, 15–20. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.; Zhou, S.; Zhang, Y.; Lv, Z.; Li, S.; Xie, C.; Ke, Y.; Deng, P.; Geng, Y.; Zhang, Q.; et al. Association of the genetic polymorphisms in pre-microRNAs with risk of ischemic stroke in a Chinese population. PLoS ONE 2015, 10, e0117007. [Google Scholar] [CrossRef] [PubMed]
- Jeon, Y.J.J.; Kim, O.J.; Kim, S.Y.; Oh, S.H.; Oh, D.; Kim, O.J.; Shin, B.S.; Kim, N.K. Association of the miR-146a, miR-149, miR-196a2, and miR-499 polymorphisms with ischemic stroke and silent brain infarction risk Genetic polymorphisms in pre-microRNAs and risk of ischemic stroke in a Chinese population. Arterioscler. Thromb. Vasc. Biol. 2013, 33, 420–430. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Ma, Y.; Zhang, B.; Wang, S.X.; Wang, X.M.; Yu, J.M. Genetic polymorphisms in pre-microRNAs and risk of ischemic stroke in a Chinese population. J. Mol. Neurosci. 2014, 52, 473–480. [Google Scholar] [CrossRef] [PubMed]
- Hamann, L.; Glaeser, C.; Schulz, S.; Gross, M.; Franke, A.; Nothlings, U.; Schumann, R.R. A micro RNA-146a polymorphism is associated with coronary restenosis. Int. J. Immunogenet. 2014, 41, 393–396. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Yang, J.; Kong, D.; Lin, J.; Xu, C.; Ren, H.; Ouyang, P.; Ding, Y.; Wang, K. Association of miR-146a rs2910164 polymorphism with cardio-cerebrovascular diseases: A systematic review and meta-analysis. Gene 2011, 565, 171–179. [Google Scholar] [CrossRef] [PubMed]
- Zhu, R.; Liu, X.; He, Z.; Li, Q. miR-146a and miR-196a2 polymorphisms in patients with ischemic stroke in the northern Chinese Han population. Neurochem. Res. 2014, 39, 1709–1716. [Google Scholar] [CrossRef] [PubMed]
- Guo, M.; Mao, X.; Ji, Q.; Lang, M.; Li, S.; Peng, Y.; Zhou, W.; Xiong, B.; Zeng, Q. miR-146a in PBMCs modulates Th1 function in patients with acute coronary syndrome. Immunol. Cell Biol. 2010, 88, 555–564. [Google Scholar] [CrossRef] [PubMed]
- Cheng, H.S.; Sivachandran, N.; Lau, A.; Boudreau, E.; Zhao, J.L.; Baltimore, D.; Delgado-Olguin, P.; Cybulsky, M.I.; Fish, J.E. MicroRNA-146 represses endothelial activation by inhibiting pro-inflammatory pathways. EMBO Mol. Med. 2013, 5, 949–966. [Google Scholar] [CrossRef] [PubMed]
- El, G.M.; Church, A.; Liu, T.; McCall, C.E. MicroRNA-146a regulates both transcription silencing and translation disruption of TNF-α during TLR4-induced gene reprogramming. J. Leukoc. Biol. 2011, 90, 509–519. [Google Scholar]
- Cui, G.; Wang, H.; Li, R.; Zhang, L.; Li, Z.; Wang, Y.; Hui, R.; Ding, H.; Wang, D.W. Polymorphism of tumor necrosis factor α (TNF-α) gene promoter, circulating TNF-α level, and cardiovascular risk factor for ischemic stroke. J. Neuroinflamm. 2012, 9, 235. [Google Scholar] [CrossRef] [PubMed]
- Jiang, D.K.; Wang, W.Z.; Ren, W.H.; Yao, L.; Peng, B.; Yu, L. TP53 Arg72Pro polymorphism and skin cancer risk: A meta-analysis. J. Investig. Dermatol. 2011, 131, 220–228. [Google Scholar] [CrossRef] [PubMed]
- Higgins, J.P.; Thompson, S.G.; Deeks, J.J.; Altman, D.G. Measuring inconsistency in meta-analyses. BMJ 2003, 327, 557–560. [Google Scholar] [CrossRef] [PubMed]
- DerSimonian, R.; Laird, N. Meta-analysis in clinical trials. Control Clin. Trials 1986, 7, 177–188. [Google Scholar] [CrossRef]
- Mantel, N.; Haenszel, W. Statistical aspects of the analysis of data from retrospective studies of disease. J. Natl. Cancer Inst. 1959, 22, 719–748. [Google Scholar] [PubMed]
- Peters, J.L.; Sutton, A.J.; Jones, D.R.; Abrams, K.R.; Rushton, L. Comparison of two methods to detect publication bias in meta-analysis. JAMA 2006, 295, 676–680. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bao, M.-H.; Xiao, Y.; Zhang, Q.-S.; Luo, H.-Q.; Luo, J.; Zhao, J.; Li, G.-Y.; Zeng, J.; Li, J.-M. Meta-Analysis of miR-146a Polymorphisms Association with Coronary Artery Diseases and Ischemic Stroke. Int. J. Mol. Sci. 2015, 16, 14305-14317. https://doi.org/10.3390/ijms160714305
Bao M-H, Xiao Y, Zhang Q-S, Luo H-Q, Luo J, Zhao J, Li G-Y, Zeng J, Li J-M. Meta-Analysis of miR-146a Polymorphisms Association with Coronary Artery Diseases and Ischemic Stroke. International Journal of Molecular Sciences. 2015; 16(7):14305-14317. https://doi.org/10.3390/ijms160714305
Chicago/Turabian StyleBao, Mei-Hua, Yan Xiao, Qing-Song Zhang, Huai-Qing Luo, Ji Luo, Juan Zhao, Guang-Yi Li, Jie Zeng, and Jian-Ming Li. 2015. "Meta-Analysis of miR-146a Polymorphisms Association with Coronary Artery Diseases and Ischemic Stroke" International Journal of Molecular Sciences 16, no. 7: 14305-14317. https://doi.org/10.3390/ijms160714305
APA StyleBao, M.-H., Xiao, Y., Zhang, Q.-S., Luo, H.-Q., Luo, J., Zhao, J., Li, G.-Y., Zeng, J., & Li, J.-M. (2015). Meta-Analysis of miR-146a Polymorphisms Association with Coronary Artery Diseases and Ischemic Stroke. International Journal of Molecular Sciences, 16(7), 14305-14317. https://doi.org/10.3390/ijms160714305





