Acute Toxicity-Supported Chronic Toxicity Prediction: A k-Nearest Neighbor Coupled Read-Across Strategy
Abstract
:1. Introduction
2. Results and Discussion
2.1. k-NN Classification
| Entry | Fingerprint | NER | k | Sensitivity | Specificity | |||
|---|---|---|---|---|---|---|---|---|
| Class 1 | Class 2 | Class 1 | Class 2 | |||||
| 1 | CDK | Fitting | 0.69 | 9 | 0.77 | 0.61 | 0.61 | 0.77 |
| CV | 0.76 | 9 | 0.84 | 0.68 | 0.68 | 0.84 | ||
| External | 0.74 | 9 | 0.79 | 0.70 | 0.70 | 0.79 | ||
| 2 | Estate | Fitting | 0.75 | 3 | 0.73 | 0.76 | 0.76 | 0.73 |
| CV | 0.74 | 3 | 0.77 | 0.71 | 0.71 | 0.77 | ||
| External | 0.81 | 3 | 0.71 | 0.90 | 0.90 | 0.71 | ||
| 3 | Extended CDK | Fitting | 0.70 | 9 | 0.80 | 0.61 | 0.61 | 0.80 |
| CV | 0.74 | 9 | 0.80 | 0.68 | 0.68 | 0.80 | ||
| External | 0.79 | 9 | 0.79 | 0.80 | 0.80 | 0.79 | ||
| 4 | Graph | Fitting | 0.68 | 3 | 0.75 | 0.61 | 0.61 | 0.75 |
| CV | 0.70 | 3 | 0.79 | 0.61 | 0.61 | 0.79 | ||
| External | 0.76 | 3 | 0.71 | 0.80 | 0.80 | 0.71 | ||
| 5 | Klekoth-Roth | Fitting | 0.68 | 10 | 0.70 | 0.66 | 0.66 | 0.70 |
| CV | 0.77 | 10 | 0.77 | 0.76 | 0.76 | 0.77 | ||
| External | 0.72 | 10 | 0.64 | 0.80 | 0.80 | 0.64 | ||
| 6 | MACCS | Fitting | 0.76 | 7 | 0.79 | 0.74 | 0.74 | 0.79 |
| CV | 0.73 | 7 | 0.77 | 0.68 | 0.68 | 0.77 | ||
| External | 0.72 | 7 | 0.64 | 0.80 | 0.80 | 0.64 | ||
| 7 | Pubchem | Fitting | 0.77 | 1 | 0.80 | 0.74 | 0.74 | 0.80 |
| CV | 0.73 | 1 | 0.77 | 0.68 | 0.68 | 0.77 | ||
| External | 0.77 | 1 | 0.64 | 0.90 | 0.90 | 0.64 | ||
| 8 | Substructure | Fitting | 0.68 | 5 | 0.73 | 0.63 | 0.63 | 0.73 |
| CV | 0.66 | 5 | 0.80 | 0.53 | 0.53 | 0.80 | ||
| External | 0.71 | 5 | 0.71 | 0.70 | 0.70 | 0.71 | ||
2.2. Read-Across for LOEL Prediction
| Entry | LOEL | Fold_diff | ||||
|---|---|---|---|---|---|---|
| Query | Analog 1 | Analog 2 | Analog 3 | Predicted | ||
| 1 | 30 | 50 | 625 | 1.2 | 225.40 | 7.51 |
| 2 | 50 | 30 | 625 | 1.2 | 218.73 | 4.37 |
| 3 | 200 | 100 | 750 | 150 | 333.33 | 1.67 |
| 4 | 10 | 10 | 250 | 30 | 96.67 | 9.67 |
| 5 | 30 | 20 | 30 | 30 | 26.67 | 1.12 |
| 6 | 70 | 300 | 20 | 20 | 113.33 | 1.62 |
| 7 | 5 | 150 | 200 | 6 | 118.67 | 23.73 |
| 8 | 100 | 200 | 750 | 200 | 383.33 | 3.83 |
| 9 | 150 | 200 | 100 | 30 | 110.00 | 1.36 |
| 10 | 1000 | 1000 | 11 | 100 | 370.33 | 2.70 |
| 11 | 30 | 1000 | 100 | 150 | 416.67 | 13.89 |
| 12 | 0.75 | 5 | 6 | 50 | 20.33 | 27.11 |
| 13 | 30 | 20 | 200 | 10 | 76.67 | 2.56 |
| 14 | 3130 | 1000 | 100 | 600 | 566.67 | 1.84 |
| 15 | 100 | 300 | 750 | 40 | 363.33 | 3.63 |
| 16 | 10 | 10 | 250 | 30 | 96.67 | 9.67 |
| 17 | 30 | 60 | 60 | 50 | 56.67 | 1.89 |
| 18 | 1000 | 30 | 1000 | 1000 | 676.67 | 1.47 |
| 19 | 60 | 30 | 60 | 50 | 46.67 | 1.29 |
| 20 | 600 | 3130 | 160 | 62.5 | 1117.50 | 1.86 |
| 21 | 20 | 30 | 200 | 10 | 80.00 | 4.00 |
| 22 | 750 | 1000 | 100 | 200 | 433.33 | 1.73 |
| 23 | 25 | 30 | 250 | 10 | 96.67 | 3.87 |
| 24 | 200 | 300 | 100 | 750 | 383.33 | 1.92 |
| 25 | 1000 | 3130 | 100 | 100 | 1110.00 | 1.11 |
| 26 | 250 | 200 | 3130 | 70 | 1133.33 | 4.53 |
| 27 | 200 | 30 | 20 | 200 | 83.33 | 2.40 |
| 28 | 300 | 200 | 200 | 100 | 166.67 | 1.80 |
| 29 | 160 | 600 | 1000 | 1000 | 866.67 | 5.42 |
| 30 | 350 | 1000 | 625 | 625 | 750.00 | 2.14 |
| 31 | 60 | 200 | 750 | 40 | 330.00 | 5.50 |
| 32 | 100 | 240 | 250 | 10 | 166.67 | 1.67 |
| 33 | 100 | 200 | 100 | 1000 | 433.33 | 4.33 |
| 34 | 30 | 30 | 30 | 100 | 53.33 | 1.78 |
| 35 | 3130 | 2500 | 10 | 350 | 953.33 | 1.09 |
| 36 | 1000 | 1000 | 11 | 30 | 347.00 | 2.88 |
| 37 | 40 | 30 | 60 | 60 | 50.00 | 1.25 |
| 38 | 100 | 40 | 20 | 30 | 30.00 | 1.11 |
| 39 | 1000 | 750 | 100 | 100 | 316.67 | 1.05 |
| 40 | 300 | 70 | 20 | 250 | 113.33 | 2.65 |
| 41 | 200 | 150 | 100 | 30 | 93.33 | 2.14 |
| 42 | 30 | 100 | 300 | 200 | 200.00 | 6.67 |
| 43 | 30 | 30 | 20 | 30 | 26.67 | 1.12 |
| 44 | 5 | 10 | 3130 | 2500 | 1880.00 | 376.00 |
| 45 | 40 | 100 | 300 | 1000 | 466.67 | 11.67 |
| 46 | 2 | 20 | 30 | 30 | 26.67 | 13.33 |
| 47 | 1.2 | 30 | 625 | 500 | 385.00 | 320.83 |
| 48 | 240 | 100 | 250 | 10 | 120.00 | 2 |
| 49 | 6 | 11 | 100 | 150 | 87.00 | 14.50 |
| 50 | 250 | 100 | 240 | 40 | 126.67 | 1.97 |
| 51 | 11 | 1000 | 6 | 1000 | 668.67 | 60.79 |
| 52 | 2 | 30 | 1000 | 30 | 353.33 | 176.67 |
| 53 | 62.5 | 6 | 600 | 3130 | 1245.33 | 19.93 |
| 54 | 100 | 1000 | 1000 | 300 | 766.67 | 7.67 |
| 55 | 40 | 200 | 100 | 150 | 150.00 | 3.75 |
| 56 | 10 | 200 | 100 | 200 | 166.67 | 16.67 |
| 57 | 20 | 70 | 300 | 1000 | 456.67 | 22.83 |
| 58 | 200 | 300 | 100 | 100 | 166.67 | 1.20 |
| 59 | 300 | 100 | 1000 | 40 | 380.00 | 1.27 |
| 60 | 100 | 1000 | 30 | 1000 | 676.67 | 6.77 |
| 61 | 30 | 30 | 1000 | 0.78 | 343.59 | 11.45 |
| 62 | 20 | 70 | 20 | 300 | 130.00 | 6.50 |
| 63 | 20 | 30 | 30 | 100 | 53.33 | 2.67 |
| 64 | 30 | 100 | 150 | 30 | 93.33 | 3.11 |
| 65 | 500 | 250 | 1.2 | 10 | 87.07 | 1.91 |
| 66 | 200 | 781 | 40 | 60 | 293.67 | 1.47 |
| 67 | 1000 | 1000 | 100 | 240 | 446.67 | 2.23 |
| 68 | 625 | 1.2 | 30 | 60 | 30.40 | 6.85 |
| 69 | 10 | 2500 | 3130 | 1000 | 2210.00 | 221.00 |
| 70 | 2500 | 10 | 3130 | 1000 | 1380.00 | 1.81 |
| 71 | 100 | 100 | 150 | 30 | 93.33 | 1.07 |
| 72 | 60 | 30 | 60 | 50 | 46.67 | 1.29 |
| 73 | 50 | 30 | 60 | 60 | 50.00 | 1.00 |
| 74 | 1000 | 350 | 1000 | 30 | 460.00 | 2.17 |
| 75 | 625 | 625 | 625 | 350 | 533.33 | 1.17 |
| 76 | 0.78 | 350 | 625 | 625 | 533.33 | 683.76 |
| 77 | 40 | 100 | 240 | 250 | 196.67 | 4.92 |
| 78 | 5 | 5 | 10 | 3130 | 1048.33 | 209.67 |
| 79 | 30 | 1.2 | 625 | 500 | 375.40 | 12.51 |
| 80 | 250 | 20 | 100 | 500 | 206.67 | 1.21 |
| 81 | 2 | 30 | 60 | 60 | 50.00 | 25.00 |
| 82 | 250 | 10 | 100 | 240 | 116.67 | 2.14 |
| 83 | 30 | 10 | 10 | 250 | 90.00 | 3.00 |
| 84 | 20 | 100 | 100 | 750 | 316.67 | 15.83 |
| 85 | 6 | 62.5 | 3130 | 70 | 1087.50 | 181.25 |
| 86 | 60 | 781 | 200 | 350 | 443.67 | 7.39 |
| 87 | 6 | 625 | 625 | 15 | 421.67 | 70.28 |
| 88 | 30 | 30 | 60 | 60 | 50.00 | 1.67 |
| 89 | 100 | 625 | 625 | 15 | 421.67 | 4.22 |
| 90 | 100 | 150 | 30 | 200 | 126.67 | 1.27 |
| 91 | 781 | 60 | 200 | 30 | 96.67 | 2.69 |
| 92 | 625 | 625 | 625 | 350 | 533.33 | 1.17 |
| 93 | 15 | 100 | 625 | 6 | 243.67 | 16.24 |
| 94 | 625 | 625 | 625 | 350 | 533.33 | 1.17 |
| Sr. | Query | Analog-1 | LOEL Analog-2 | Analog-3 | Predicted | Fold_diff |
|---|---|---|---|---|---|---|
| 1 | 30 | 30 | 30 | 20 | 26.67 | 1.13 |
| 2 | 30 | 30 | 1000 | 6 | 345.33 | 11.51 |
| 3 | 1.5 | 200 | 5 | 0.75 | 68.58 | 45.72 |
| 4 | 1250 | 750 | 1000 | 100 | 616.67 | 2.03 |
| 5 | 50 | 781 | 60 | 250 | 363.67 | 7.27 |
| 6 | 0.1 | 30 | 10 | 10 | 16.67 | 166.67 |
| 7 | 1000 | 1000 | 1000 | 11 | 670.33 | 1.49 |
| 8 | 20 | 150 | 200 | 6 | 118.67 | 5.93 |
| 9 | 20 | 5 | 10 | 5 | 6.67 | 3.00 |
| 10 | 100 | 250 | 100 | 500 | 283.33 | 2.83 |
| 11 | 110 | 1000 | 1000 | 11 | 670.33 | 6.09 |
| 12 | 1000 | 1000 | 100 | 750 | 616.67 | 1.62 |
| 13 | 33 | 30 | 10 | 10 | 16.67 | 1.98 |
| 14 | 30 | 3130 | 2500 | 200 | 1943.33 | 64.78 |
| 15 | 10 | 781 | 60 | 30 | 290.33 | 29.03 |
| 16 | 300 | 100 | 300 | 40 | 146.67 | 2.05 |
| 17 | 2 | 30 | 30 | 10 | 23.33 | 11.67 |
| 18 | 200 | 350 | 1000 | 1000 | 783.33 | 3.92 |
| 19 | 125 | 10 | 250 | 40 | 100.00 | 1.25 |
| 20 | 50 | 1000 | 100 | 30 | 376.67 | 7.53 |
| 21 | 100 | 350 | 625 | 6 | 327.00 | 3.27 |
| 22 | 10 | 6 | 15 | 10 | 10.33 | 1.03 |
| 23 | 150 | 1000 | 750 | 200 | 650.00 | 4.33 |
| 24 | 4 | 6 | 625 | 625 | 418.67 | 104.67 |
| Fold_diff# | Number of Queries | Total | |
|---|---|---|---|
| Qualified Category | Non-Qualified Category | ||
| <10 | 54 | 17 | 71 |
| 10–100 | 12 | 4 | 16 |
| >100 | 4 | 3 | 7 |
| Total | 70 | 24 | 94 |
| Fold_diff# | Number of Queries | Total | |
|---|---|---|---|
| Qualified Category | Non-Qualified Category | ||
| <10 | 14 | 3 | 17 |
| 10–100 | 4 | 1 | 5 |
| >100 | 1 | 1 | 2 |
| Total | 19 | 5 | 24 |
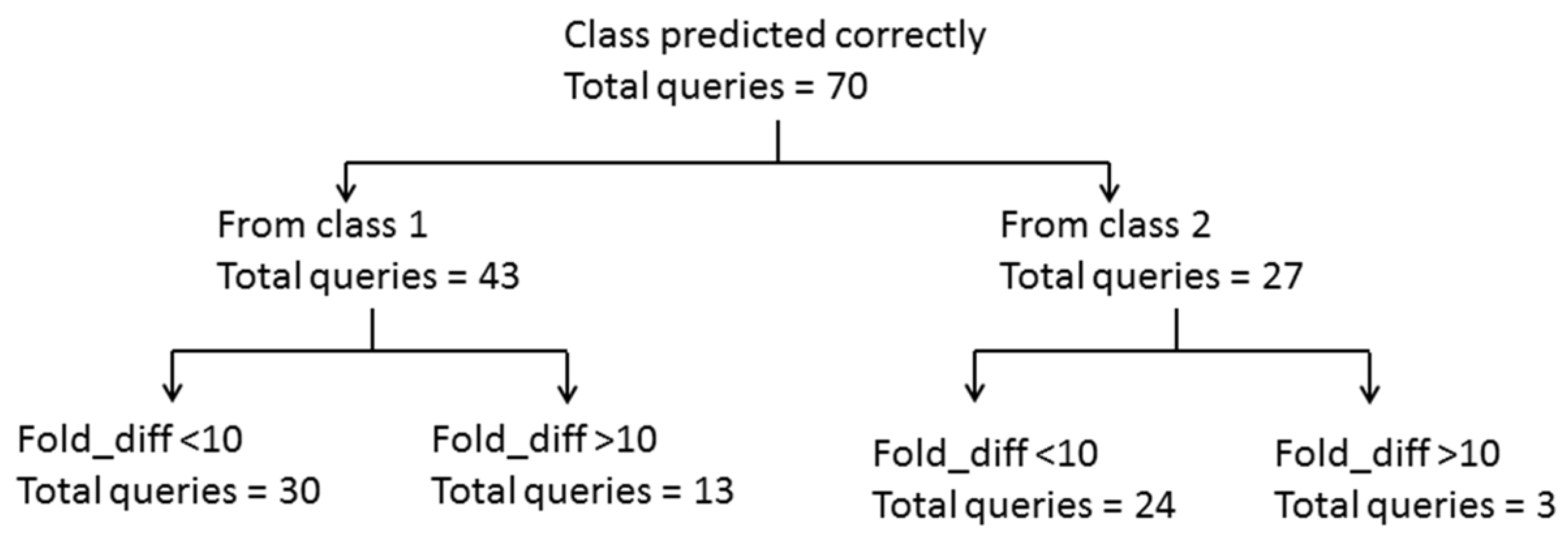
2.3. Mechanistic Interpretation
| Entry | Data | Query | Analog 1 | Analog 2 | Analog 3 | LOEL Predicted | Fold_diff |
|---|---|---|---|---|---|---|---|
| 3 | Structure | 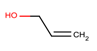 | 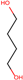 | 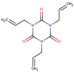 |  | ||
| LD50 | 64 | 1525 | 1000 | 26 | |||
| LOEL | 1.5 | 200 | 5 | 0.75 | 68.58 | 45.72 | |
| 22 | Structure | 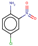 | 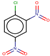 | 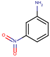 | 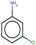 | ||
| LD50 | 400 | 640 | 535 | 256 | |||
| LOEL | 10 | 6 | 15 | 10 | 10.33 | 1.03 | |
| 24 | Structure |  |  |  |  | ||
| LD50 | 953 | 640 | 891 | 1072 | |||
| LOEL | 4 | 6 | 625 | 625 | 418.67 | 104.67 |
2.4. Comparison with Previously Published Models for Repeated Dose Toxicity Prediction
| Method | Training Set Chemicals | Test Set Chemicals | Training Set Prediction | Test Set Prediction | Comment | Reference |
|---|---|---|---|---|---|---|
| Multivariate analysis | 234 | none | 95% within factor of 5 | none | No external prediction | [32] |
| MLR | 234 | none | R2 = 0.52 | none | [29] | |
| MLR | 86 | 16 | R2 = 0.79 | R2 = 0.71 | [33] | |
| PLS | 445 | none | R2 = 0.54 | none | No external prediction | [30] |
| Read-across | 500 | none | none | none | 33 chemical categories formed | [31] |
| k-NN | 254 | 179 | q2 = 0.63 | R2 = 0.54 | [24] |
2.5. Toxicological Significance
3. Experimental Section
3.1. Software and Modules
3.2. Setting of the Dataset
| Description | LD50 (mg/kg/day) | Number of Entries | Training Set Entries | Test Set Entries | |
|---|---|---|---|---|---|
| Class 1 | Highly toxic, toxic and harmful | ≤2000 | 70 | 56 | 14 |
| Class 2 | Non-harmful | >2000 | 48 | 38 | 10 |
3.3. Fingerprint Calculations
3.4. Development of the Classification Model
3.5. External Validation
3.6. Model Selection and Read-Across
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Abbreviations
| CAS | Chemical Abstracts Service |
| CDK | Chemistry Developmental Kit |
| CV | Cross Validation |
| GHS | Globally Harmonized Scheme |
| IUPAC | International Union of Pure and Applied Chemistry |
| k-NN | k Nearest Neighbor |
| LOEL | Lowest Observed Effect Level |
| MACCS | Molecular ACCess System |
| NEDO | New Energy and industrial technology Development Organization |
| NER | Non-Error Rate |
| OECD | Organization for Economic Cooperation and Development |
| QSAR | Quantitative Structure-Activity Relationship |
| RDT | Repeated Dose Toxicity |
| SMILES | Simplified Molecular-Input Line-Entry System |
Conflicts of Interest
References
- Anighoro, A.; Bajorath, J.R.; Rastelli, G. Polypharmacology: Challenges and opportunities in drug discovery. J. Med. Chem. 2014, 57, 7874–7887. [Google Scholar] [CrossRef] [PubMed]
- Stegmeier, F.; Warmuth, M.; Sellers, W.; Dorsch, M. Targeted cancer therapies in the twenty-first century: Lessons from imatinib. Clin. Pharmacol. Ther. 2010, 87, 543–552. [Google Scholar] [CrossRef] [PubMed]
- Abassi, Y.A.; Xi, B.; Zhang, W.; Ye, P.; Kirstein, S.L.; Gaylord, M.R.; Feinstein, S.C.; Wang, X.; Xu, X. Kinetic cell-based morphological screening: Prediction of mechanism of compound action and off-target effects. Chem. Biol. 2009, 16, 712–723. [Google Scholar] [CrossRef] [PubMed]
- Keiser, M.J.; Setola, V.; Irwin, J.J.; Laggner, C.; Abbas, A.I.; Hufeisen, S.J.; Jensen, N.H.; Kuijer, M.B.; Matos, R.C.; Tran, T.B. Predicting new molecular targets for known drugs. Nature 2009, 462, 175–181. [Google Scholar] [CrossRef] [PubMed]
- Enoch, S. Chemical Category Formation and Read-Across for the Prediction of Toxicity. Springer: Dordrecht, The Netherlands, 2010; pp. 209–219. [Google Scholar]
- Patlewicz, G.; Simon, T.W.; Rowlands, J.C.; Budinsky, R.A.; Becker, R.A. Proposing a scientific confidence framework to help support the application of adverse outcome pathways for regulatory purposes. Regul. Toxicol. Pharmacol. 2015, 71, 463–477. [Google Scholar] [CrossRef] [PubMed]
- European Chemical Agency; Helsinki, Finland. Grouping of substances and read-across approach. Available online: http://echa.europa.eu/documents/10162/13628/read_across_introductory_note_en.pdf (accessed on 17 March 2015).
- Patlewicz, G.; Ball, N.; Booth, E.D.; Hulzebos, E.; Zvinavashe, E.; Hennes, C. Use of category approaches, read-across and (Q)SAR: General considerations. Regul. Toxicol. Pharmacol. 2013, 67, 1–12. [Google Scholar] [CrossRef] [PubMed]
- De Raat, K.; Netzava, T. Assessment of read-across in ECHA, European Chemical Agency. Available online: http://echa.europa.eu/documents/10162/5649897/ws_raa_20121003_assessment_of_read-across_in_echa_de_raat_en.pdf (accessed on 18 February 2015).
- Russell, W.M.S.; Burch, R.L.; Hume, C.W. The Principles of Humane Experimental Technique; Methuen: London, UK, 1959; pp. 1–238. [Google Scholar]
- Spielmann, H.; Sauer, U.G.; Mekenyan, O. A critical evaluation of the 2011 ECHA reports on compliance with the REACH and CLP regulations and on the use of alternatives to testing on animals for compliance with the REACH regulation. Altern. Lab. Anim. 2011, 39, 481–493. [Google Scholar] [PubMed]
- Lapenna, S.; Fuart-Gatnik, M.; Worth, A. Review of QSAR models and software tools for predicting acute and chronic systemic toxicity. Joint Research Centre: Ispra, Italy. Available online: http://s-ihcpiprwb002p.jrc.it/our_labs/eurl-ecvam/laboratories-research/predictive_toxicology/doc/EUR_24639_EN.pdf (accessed on 17 March 2015).
- Kowalski, B.; Bender, C. k-Nearest Neighbor Classification Rule (pattern recognition) applied to nuclear magnetic resonance spectral interpretation. Anal. Chem. 1972, 44, 1405–1411. [Google Scholar] [CrossRef]
- Hellman, M.E. The nearest neighbor classification rule with a reject option. IEEE Trans. Syst. Sci. Cybern. 1970, 6, 179–185. [Google Scholar] [CrossRef]
- Patrick, E.A.; Fischer Iii, F.P. A generalized k-nearest neighbor rule. Inf. Control 1970, 16, 128–152. [Google Scholar] [CrossRef]
- Jeng-Shyang, P.; Yu-Long, Q.; Sheng-He, S. A fast k-nearest neighbors classification algorithm. IEICE Trans. Fundam. Electron. 2004, 87, 961–963. [Google Scholar]
- Chavan, S.; Nicholls, I.A.; Karlsson, B.C.; Rosengren, A.M.; Ballabio, D.; Consonni, V.; Todeschini, R. Towards global QSAR model building for acute toxicity: Munro database case study. Int. J. Mol. Sci. 2014, 15, 18162–18174. [Google Scholar] [CrossRef] [PubMed]
- Todeschini, R. k-nearest neighbour method: The influence of data transformations and metrics. Chemom. Intell. Lab. Syst. 1989, 6, 213–220. [Google Scholar] [CrossRef]
- Cover, T.; Hart, P. Nearest neighbor pattern classification. IEEE Trans. Inf. Theory 1967, 13, 21–27. [Google Scholar] [CrossRef]
- Zhang, B.; Srihari, S.N. Properties of binary vector dissimilarity measures. Available online: http://www.cedar.buffalo.edu/papers/articles/CVPRIP03_propbina.pdf (accessed on 18 February 2015).
- Cheetham, A.H.; Hazel, J.E. Binary (presence-absence) similarity coefficients. J. Paleontol. 1969, 43, 1130–1136. [Google Scholar]
- Cramer, G.; Ford, R.; Hall, R. Estimation of toxic hazard—A decision tree approach. Food Chem. Toxicol. 1976, 16, 255–276. [Google Scholar] [CrossRef]
- Nelson, S.D. Metabolic activation and drug toxicity. J. Med. Chem. 1982, 25, 753–765. [Google Scholar] [CrossRef] [PubMed]
- Gadaleta, D.; Pizzo, F.; Lombardo, A.; Carotti, A.; Escher, S.E.; Nicolotti, O.; Benfenati, E. A k-NN algorithm for predicting oral sub-chronic toxicity in the rat. Altex 2014, 31, 423–432. [Google Scholar] [CrossRef] [PubMed]
- Hall, L.H.; Kier, L.B. Electrotopological state indices for atom types: A novel combination of electronic, topological, and valence state information. J. Chem. Inf. Comput. Sci. 1995, 35, 1039–1045. [Google Scholar] [CrossRef]
- Enoch, S.J.; Cronin, M.T.D.; Schultz, T.W.; Madden, J.C. Quantitative and mechanistic read across for predicting the skin sensitization potential of alkenes acting via Michael addition. Chem. Res. Toxicol. 2008, 21, 513–520. [Google Scholar] [CrossRef] [PubMed]
- Yoshii, E. Cytotoxic effects of acrylates and methacrylates: Relationships of monomer structures and cytotoxicity. J. Biomed. Mater. Res. A 1997, 37, 517–524. [Google Scholar] [CrossRef]
- Freidig, A.P.; Verhaar, H.J.; Hermens, J.L. Comparing the potency of chemicals with multiple modes of action in aquatic toxicology: Acute toxicity due to narcosis versus reactive toxicity of acrylic compounds. Environ. Sci. Technol. 1999, 33, 3038–3043. [Google Scholar] [CrossRef]
- De Julian-Ortiz, J.; Garcia-Domenech, R.; Galvez, J.; Pogliani, L. Predictability and prediction of lowest observed adverse effect levels in a structurally heterogeneous set of chemicals. SAR QSAR Environ. Res. 2005, 16, 263–272. [Google Scholar] [CrossRef] [PubMed]
- Mazzatorta, P.; Estevez, M.D.; Coulet, M.; Schilter, B. Modeling oral rat chronic toxicity. J. Chem. Info. Model. 2008, 48, 1949–1954. [Google Scholar] [CrossRef]
- Sakuratani, Y.; Zhang, H.; Nishikawa, S.; Yamazaki, K.; Yamada, T.; Yamada, J.; Gerova, K.; Chankov, G.; Mekenyan, O.; Hayashi, M. Hazard Evaluation Support System (HESS) for predicting repeated dose toxicity using toxicological categories. SAR QSAR Environ. Res. 2013, 24, 351–363. [Google Scholar] [CrossRef]
- Mumtaz, M.; Knauf, L.; Reisman, D.; Peirano, W.; Derosa, C.; Gombar, V.; Enslein, K.; Carter, J.; Blake, B.; Huque, K. Assessment of effect levels of chemicals from quantitative structure-activity relationship (QSAR) models. I. Chronic lowest-observed-adverse-effect level (LOAEL). Toxicol. Lett. 1995, 79, 131–143. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Domenech, R.; De Julian-Ortiz, J.; Besalu, E. True prediction of lowest observed adverse effect levels. Molec. Divers. 2006, 10, 159–168. [Google Scholar] [CrossRef]
- Yap, C.W. PaDEL-descriptor: An open source software to calculate molecular descriptors and fingerprints. J. Comp. Chem. 2011, 32, 1466–1474. [Google Scholar] [CrossRef]
- Kenaga, E.E. Predictability of chronic toxicity from acute toxicity of chemicals in fish and aquatic invertebrates. Environ. Toxicol. Chem. 1982, 1, 347–358. [Google Scholar] [CrossRef]
- Rand, G.M.; Petrocelli, S.R. Fundamentals of aquatic toxicology: Methods and applications; FMC Corp., Princeton, NJ: Washington DC, USA, 1985; pp. 1–666. [Google Scholar]
- Kumar, A.; Correll, R.; Grocke, S.; Bajet, C. Toxicity of selected pesticides to freshwater shrimp, Paratya australiensis (Decapoda: Atyidae): Use of time series acute toxicity data to predict chronic lethality. Ecotoxicol. Environ. Saf. 2010, 73, 360–369. [Google Scholar] [CrossRef] [PubMed]
- Ballabio, D.; Consonni, V. Classification tools in chemistry. Part 1: Linear models. PLS-DA. Anal. Methods 2013, 3790–3798. [Google Scholar] [CrossRef]
- National Institute of Technology and Evaluation. Hazard Evaluation Support System Integrated Platform. 2012. http://www.nite.go.jp/en/chem/qsar/hess-e.html (accessed on 13 March 2015).
- The OECD QSAR toolbox. http://www.qsartoolbox.org/ (accessed on 17 March 2017).
- The OECD QSAR toolbox version 2.2 release note. http://www.oecd.org/chemicalsafety/risk-assessment/48413109.pdf (accessed on 17 March 2017).
- Pence, H.E.; Williams, A. ChemSpider: An online chemical information resource. J. Chem. Educ. 2010, 87, 1123–1124. [Google Scholar] [CrossRef]
- Wang, Y.; Xiao, J.; Suzek, T.O.; Zhang, J.; Wang, J.; Bryant, S.H. PubChem: A public information system for analyzing bioactivities of small molecules. Nucl. Acids Res. 2009, 37, 623–633. [Google Scholar] [CrossRef]
- Lenga, R.E.; Votoupal, K.L. The Sigma-Aldrich library of regulatory and safety data. https://www.sigmaaldrich.com/ (accessed on 18 February 2015).
- Wexler, P. TOXNET: An evolving web resource for toxicology and environmental health information. Toxicology 2001, 157, 3–10. [Google Scholar] [CrossRef] [PubMed]
- United Nations. Globally harmonized system of classification and labelling of chemicals (GHS). Geneva, Switzerland, 2011. http://www.unece.org/fileadmin/DAM/trans/danger/publi/ghs/ghs_rev04/English/ST-SG-AC10-30-Rev4e.pdf (accessed on 17 March 2015).
- Steinbeck, C.; Han, Y.; Kuhn, S.; Horlacher, O.; Luttmann, E.; Willighagen, E. The Chemistry Development Kit (CDK): An open-source Java library for chemo-and bioinformatics. J. Chem. Inform. Comput. Sci. 2003, 43, 493–500. [Google Scholar] [CrossRef]
- Dancet, G. The use of alternatives to testing on animals for the REACH regulation. European Chemical Agency; pp. 1–64. Available online: http://echa.europa.eu/documents/10162/13639/alternatives_test_animals_2014_en.pdf (accessed on 18 February 2015).
- Stephens, M.L.; Fowle, J.; Boverhof, D.; Richmond, J.; Recio, L.; Zhu, H.; Alepee, N.; Curren, R.; Basketter, D.; Natsch, A. Reducing, Refining and Replacing the Use of Animals in Toxicity Testing; Royal Society of Chemistry: Cambridge, UK, 2013; pp. 1–347. [Google Scholar]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chavan, S.; Friedman, R.; Nicholls, I.A. Acute Toxicity-Supported Chronic Toxicity Prediction: A k-Nearest Neighbor Coupled Read-Across Strategy. Int. J. Mol. Sci. 2015, 16, 11659-11677. https://doi.org/10.3390/ijms160511659
Chavan S, Friedman R, Nicholls IA. Acute Toxicity-Supported Chronic Toxicity Prediction: A k-Nearest Neighbor Coupled Read-Across Strategy. International Journal of Molecular Sciences. 2015; 16(5):11659-11677. https://doi.org/10.3390/ijms160511659
Chicago/Turabian StyleChavan, Swapnil, Ran Friedman, and Ian A. Nicholls. 2015. "Acute Toxicity-Supported Chronic Toxicity Prediction: A k-Nearest Neighbor Coupled Read-Across Strategy" International Journal of Molecular Sciences 16, no. 5: 11659-11677. https://doi.org/10.3390/ijms160511659
APA StyleChavan, S., Friedman, R., & Nicholls, I. A. (2015). Acute Toxicity-Supported Chronic Toxicity Prediction: A k-Nearest Neighbor Coupled Read-Across Strategy. International Journal of Molecular Sciences, 16(5), 11659-11677. https://doi.org/10.3390/ijms160511659







