Mass Spectrometry-Based N-Glycomics of Colorectal Cancer
Abstract
:1. Introduction
2. Colorectal Cancer (CRC)
3. Overview of N-Glycosylation and Its Biological Roles
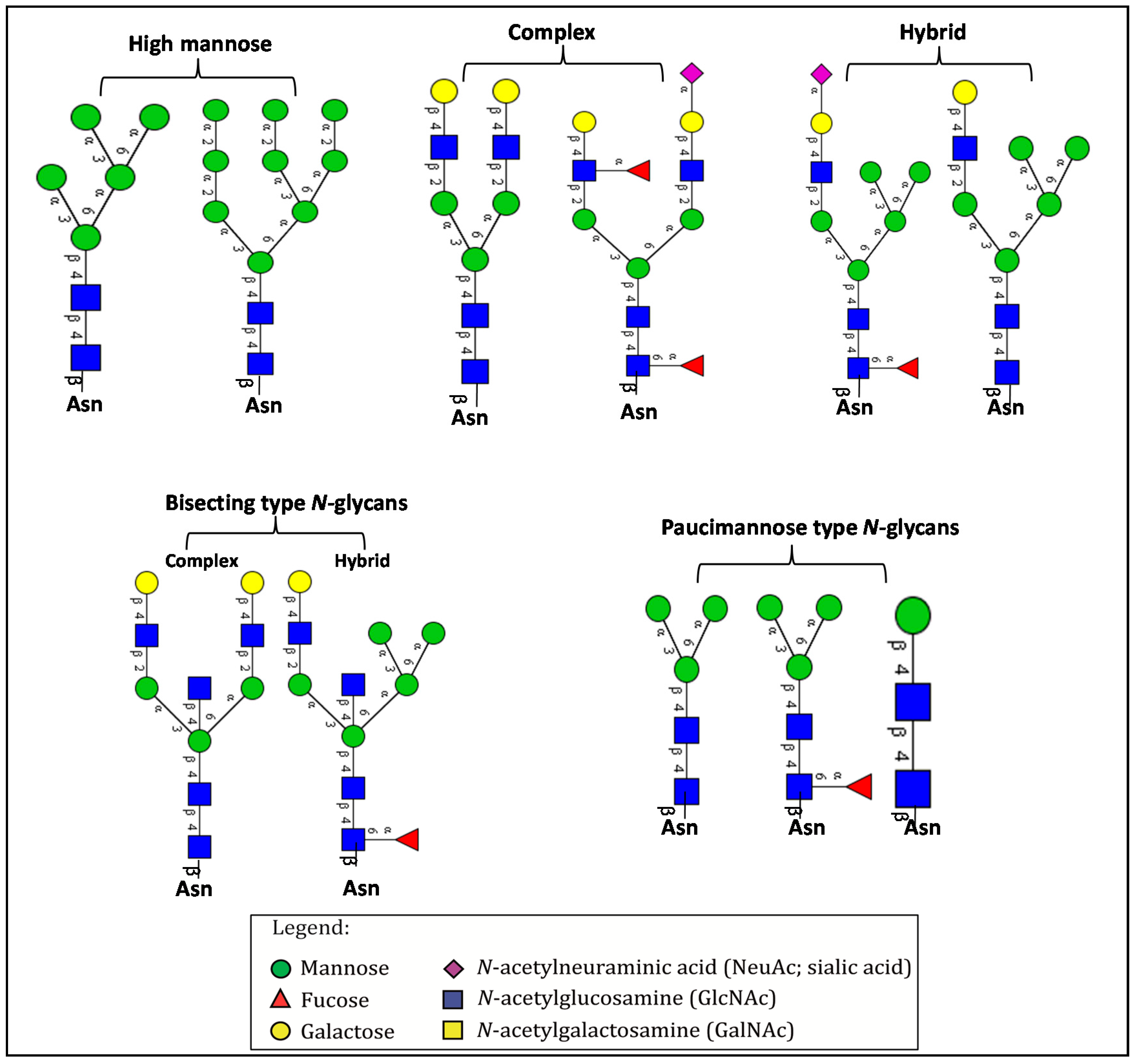
4. Aberrant N-Glycosylation in CRC and Other Cancers
Altered N-Glycosylation in Colorectal Cancer
| Aim of the Study | Finding | Altered N-glycan Structures | Reference | |
|---|---|---|---|---|
| To elucidate differential expression of β1,6-branching in two variants of HCT116 CRC lines (HCT116a (more aggressive subline) and HCT116b). | Increased expression of β1,6-linked GlcNAc branching in HCT116a. | ↑ | 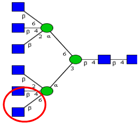 | Li et al. [50] |
| To map the differences in N-glycans attached to lysosomal membrane glycoproteins isolated from CRC sublines exhibiting different metastatic potentials. | Increased poly-N-acetyl lactosamine (LacNAc) units and sialyl Lex, decreased fucosylation on LacNAc units of highly metastatic CRC cells relative to cells with less metastatic potential. | ↑ | 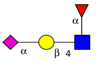 | Saitoh et al. [90] |
| ↑ | 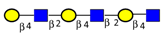 | |||
| ↓ | 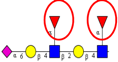 | |||
| To compare sialyltransferase activities in CRC tumor and adjacent normal mucosa. | Increased α2,6-sialyltransferase activity in CRC tumor relative to normal mucosa. | ↑ | 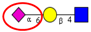 | Dall’Olio et al. [55] |
| To compare the activity of sialyltranferases with different linkage specificities (α2,6- and α2,3-sialyltransferases) in different tissues including human CRC, normal mucosa, liver and liver metastases, and CRC patient serum samples. | Increased activity of α2,6-specific sialyltranferase in tumor tissue and serum of patients with metastatic tumors. α2,3-sialyltransferase activity was unchanged. | ↑ | 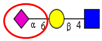 | Gessner et al. [56] |
| To investigate the expression of α2,6- and α2,3-sialylation in CRC tumor tissues from different stages. | Increased α2,3-linked sialylation in stage I and II tumors, with a decrease in advanced CRC. Significant increase in α2,6-sialylation and in metastatic tumors. | ↑ | 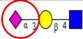 | Vierbuchen et al. [91] |
| ↑ | 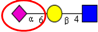 | |||
| To investigate the relationship between N-acetylglucosaminyl-transferase V (GnT-V) and metastasis in CRC tissues. | Expression of GnT-V significantly correlated with distant metastasis. | ↑ | 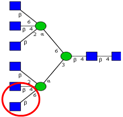 | Murata et al. [51] |
| To compare the expression and activity of α1,6-fucosyltransferase in CRC tumor and healthy tissues. | Increased expression and activity of α1,6-fucosyltransferase expression and activity in CRC tumor compared to healthy tissues. | ↑ | 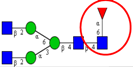 | Muinelo-Romay et al. [92] |
| To compare the expression of sialo- and fucosyl-glycoconjugates in a panel of normal mucosa and adenocarcinoma samples, by lectin immunohistochemical analysis. | Increased expression of α2,6-linked sialic acid residues (as evident by strong staining of CRC tumor tissues with Sambucusnigra Lectin) in CRC tissue. | ↑ | 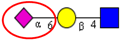 | Fernández-Rodríguez et al. [93] |
| To detect glycosylation changes during colon epithelium differentiation and proliferation. | Significant decrease in high mannose type N-glycans and increase in atypical GlcNAc-ended N-glycans in differentiating HT-29 cells. | ↓ | 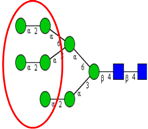 | Vercoutter-Edouart et al. [94] |
| ↑ | 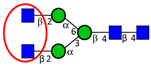 | |||
| To screen N-glycosylation changes in plasma samples from CRC patients. | Significantly higher levels of sialylation and fucosylation in patients with CRC or adenomas, compared to normal controls. | ↓ | 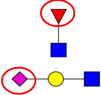 | Qiu et al. [95] |
| To profile serum N-glycans in samples from healthy individuals and patients with CRC and adenomas, to identify potential N-glycan markers for prediction and detection of CRC. | Decreased total core α1,6 fucose residues and fucosyltransferase in CRC compared to adenomas and normal controls; Increased bi-galacto biantennary glycan and α1,3-fucosylated triantennary and decreased single and bi-galacto α1,6 fucosylated biantennary in CRC groups. | ↓ | 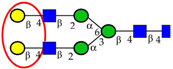 | Zhao et al. [96] |
| ↑ | 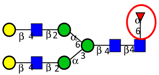 | |||
| ↑ | 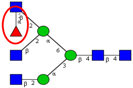 | |||
| To investigate alterations in sialylation and fucosylation in CRC patient tissues, by lectin immunohistochemical staining. | Predominant expression of α2,3 sialylated type 2 chain structures in CRC tissues associated with malignant transformation, in particular lymphatic spread. | ↑ | 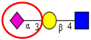 | Fukasawa et al. [97] |
| Comparison of N-glycan profiles from a panel of CRC tumor tissues and corresponding control colon tissues, by hydrophilic interactionliquidchromatography and MALDI-TOF-MS. | Significant increase in sulfated, paucimannosidic and sialylated glycans, in particular glycanswithsialylLewis type epitopes, and decrease in bisecting GlcNAc type N-glycans in CRC tumor tissues relative to normal tissues. | ↑ |  | Balog et al. [35] |
| ↑ | 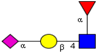 | |||
| ↓ | 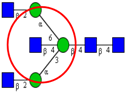 | |||
| Comparative N-glycan analysis of three pathologically and phenotypically different CRC cell lines (LIM1215, LIM1899, LIM2405). | Dominance of high mannose type and α2,6-sialylated glycans in all three cell lines; exclusive expression of bisecting GlcNAc and α2,3-sialylated N-glycans in metastatic (LIM1215) and aggressive (LIM2405) CRC cell lines, respectively. | ↑ | 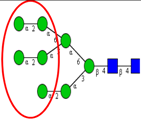 | Sethi et al. [33] |
| ↑ | 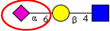 | |||
| Comparative glycomic analysis of CRC cell lines (SW1116, SW480, SW620, SW837, LS174) and CRC tissue samples. | Elevated high mannose type N-glycans in both CRC cell lines and tumor samples. | ↑ | 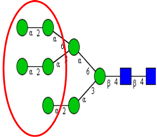 | Chik et al. [98] |
| N-glycomic profiling of rectal adenomas and carcinomas by MALDI-TOF-MS, followed by IHC expression studies of sialyl Lewis a, and paucimannose glycans in a panel of CRC patients. | Mono-antennary, sialylated, paucimannose and small high mannose N-glycan structures were more common in carcinomas than in adenomas; correlation between poor prognosis and elevated expression of sialyl Lea and paucimannosidic N-glycans in CRC and advanced CRC, respectively. | ↑ | 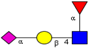 | Kaprio et al. [99] |
| ↑ | 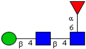 | |||
| ↑ | 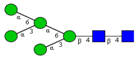 | |||
| Comparison of N-glycan profiles of membrane proteins from paired tumorigenic and adjacent non-tumorigenic CRC tissues. | Overrepresentation of high mannose, hybrid and paucimannosidic type N-glycans and under-representation of complex N-glycans in CRC tissues; higher sialylation, in particular α2,6-sialylation, in CRC tissues, coupled with down-regulation of α2,3-sialylation; high α2,3-sialylation and low bisecting β1,4-GlcNAcylation and Lewis-type fucosylation in mid-late stage CRC tissues, relative to early stage CRC;high bisecting β1,4-GlcNAcylation and low α2,3-sialylation in EGFR-positive tissues. | ↑ | 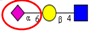 | Sethi et al. [57] |
| ↑ | 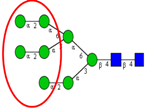 | |||
| ↑ | 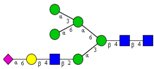 | |||
| ↑ | 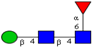 | |||
| ↓ | 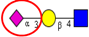 | |||
5. LC-MS/MS-Based CRC N-Glycomics
5.1. Sample Handling for N-Glycan Analysis
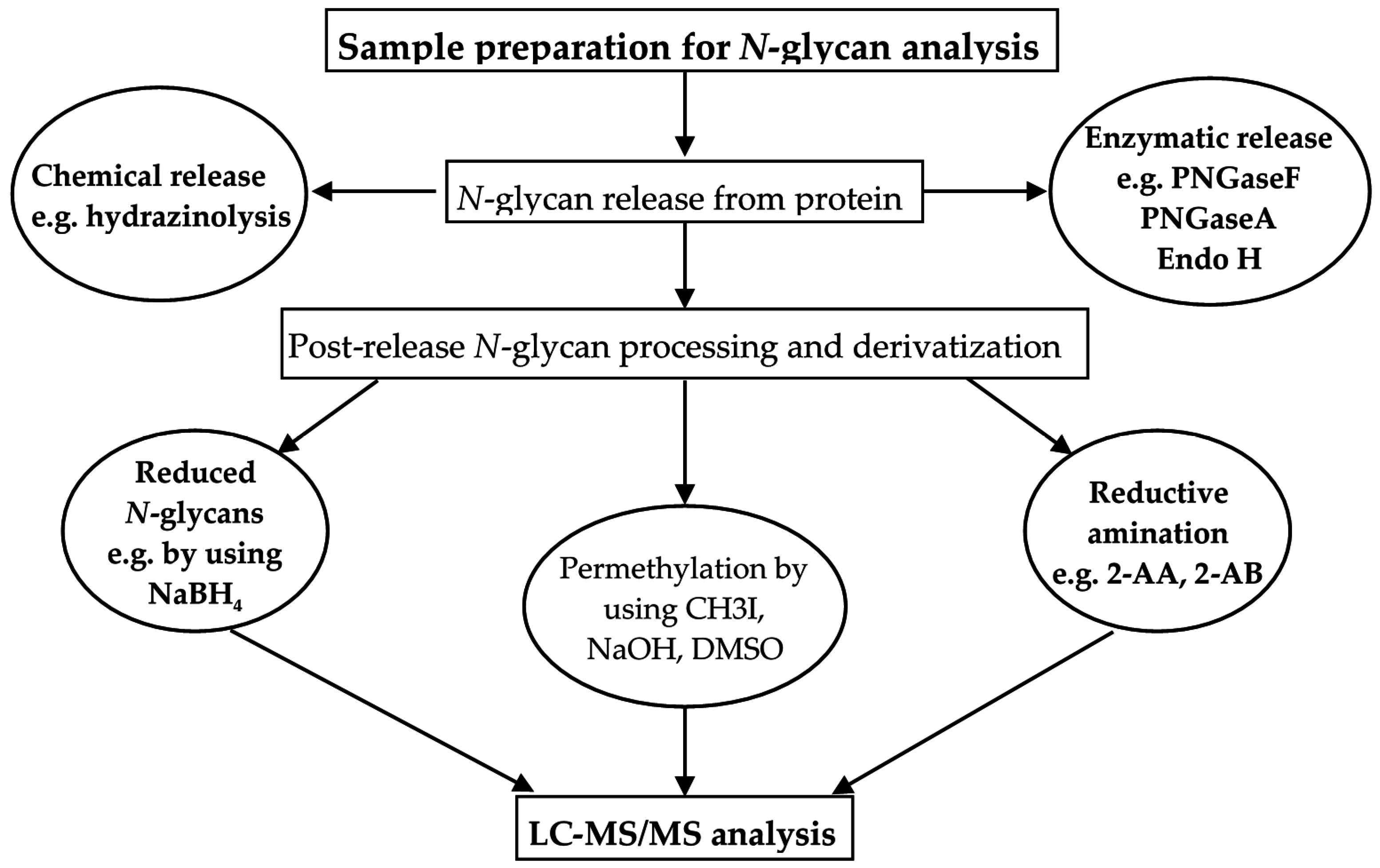
5.2. LC-Based Separation of N-Glycans
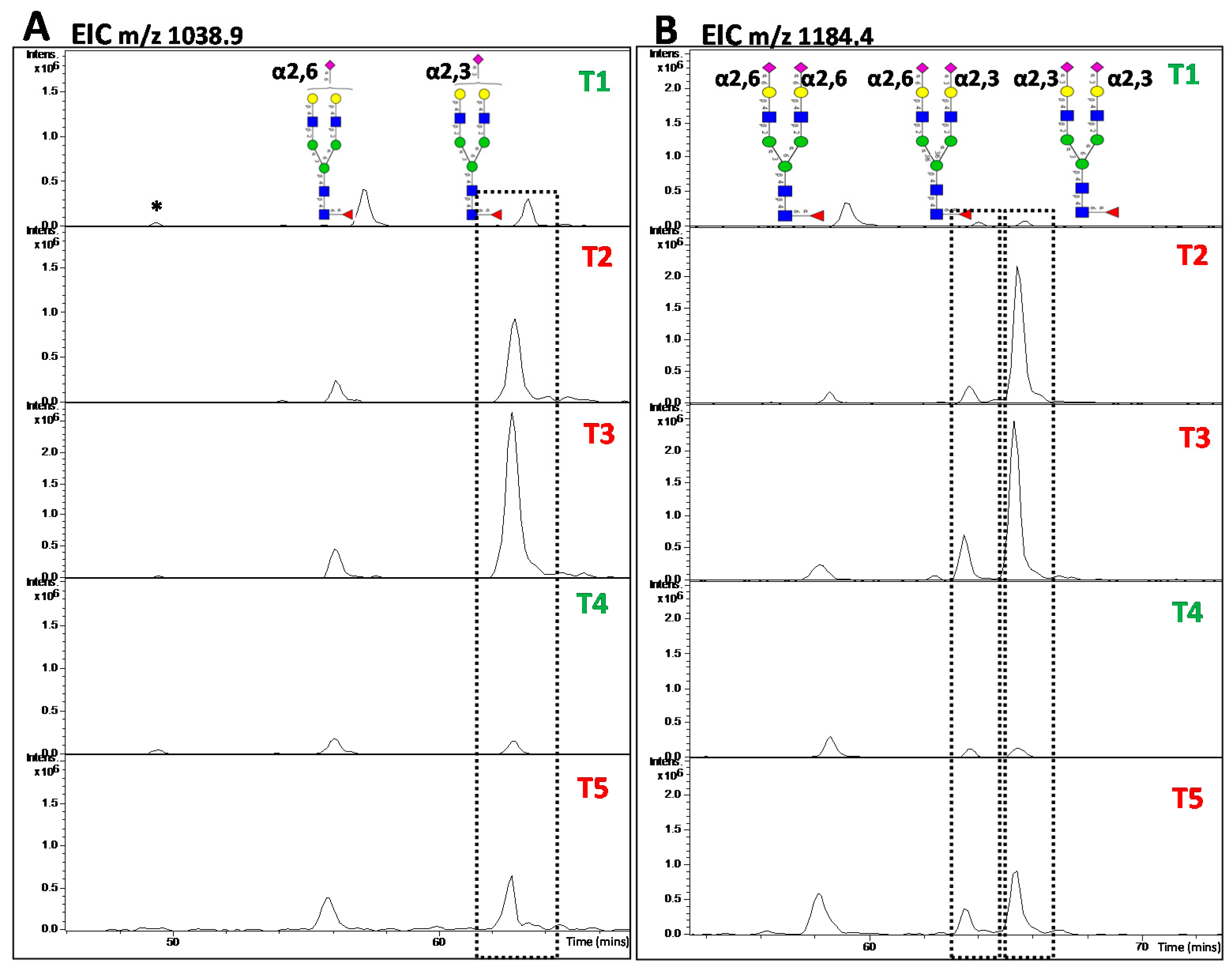
5.3. MS and MS/MS of N-Glycans
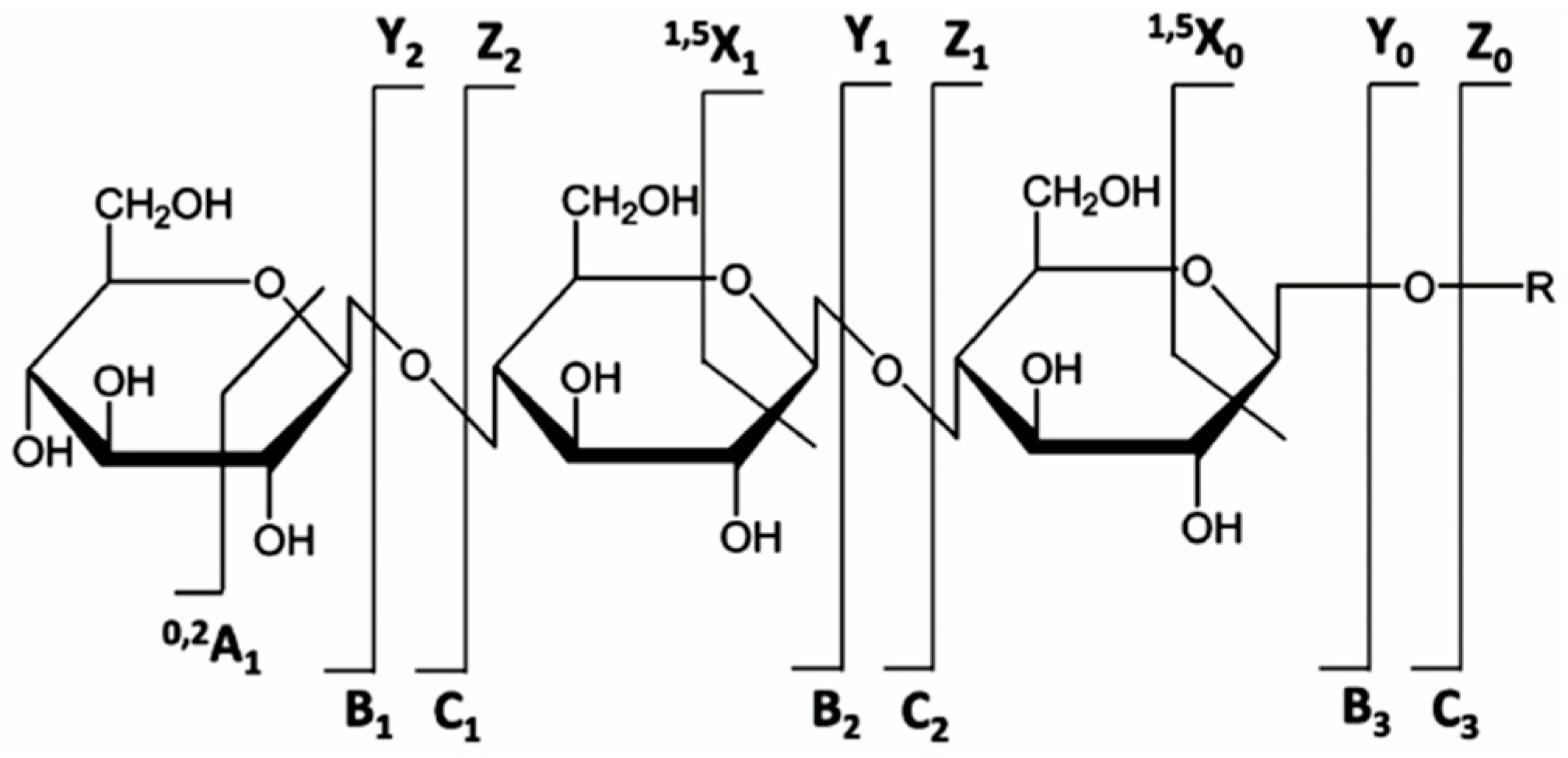
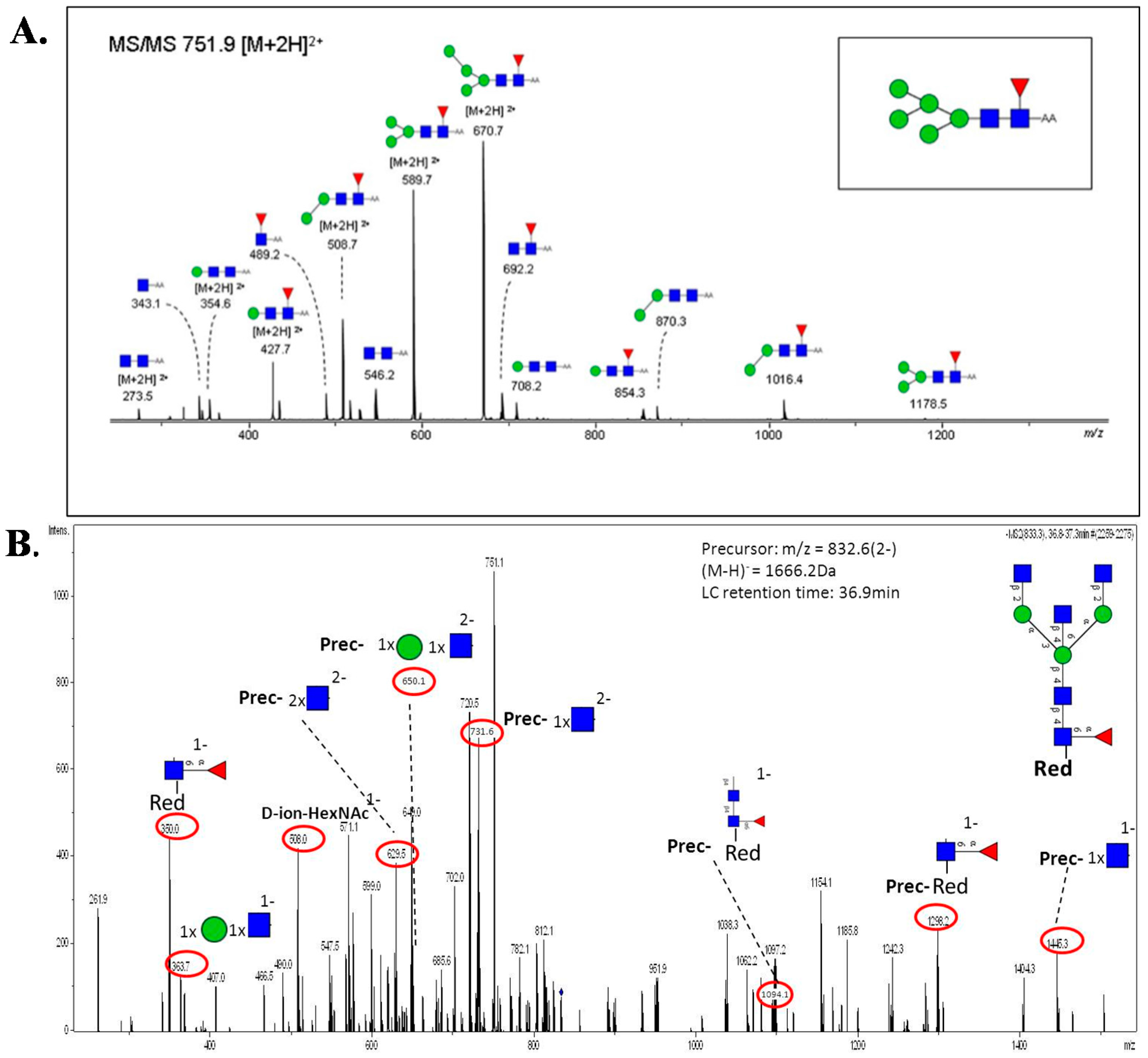
6. Quantitative Glycomics
7. Bioinformatics Tools and Glycome-Centric Databases and Resources
8. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
References
- Bretthauer, M.; Hoff, G. Prevention and early diagnosis of colorectal cancer. Tidsskr. Nor. Laegeforen. 2007, 127, 2688–2691. [Google Scholar] [PubMed]
- Davies, R.J.; Miller, R.; Coleman, N. Colorectal cancer screening: Prospects for molecular stool analysis. Nat. Rev. Cancer 2005, 5, 199–209. [Google Scholar] [CrossRef] [PubMed]
- Terdiman, J.P. Colonoscopy is superior to flexible sigmoidoscopy for colorectal cancer screening: Now beyond a reasonable doubt? Gastroenterology 2005, 129, 1793–1794. [Google Scholar] [CrossRef] [PubMed]
- Winawer, S.; Fletcher, R.; Rex, D.; Bond, J.; Burt, R.; Ferrucci, J.; Ganiats, T.; Levin, T.; Woolf, S.; Johnson, D.; et al. Gastrointestinal Consortium Panel. Colorectal cancer screening and surveillance: Clinical guidelines and rationale-Update based on new evidence. Gastroenterology 2003, 124, 544–560. [Google Scholar] [CrossRef] [PubMed]
- Wulfkuhle, J.D.; Liotta, L.A.; Petricoin, E.F. Proteomic applications for the early detection of cancer. Nat. Rev. Cancer 2003, 3, 267–275. [Google Scholar] [CrossRef] [PubMed]
- Apweiler, R.; Hermjakob, H.; Sharon, N. On the frequency of protein glycosylation, as deduced from analysis of the SWISS-PROT database. Biochim. Biophys. Acta 1999, 1473, 4–8. [Google Scholar] [CrossRef]
- Ohtsubo, K.; Marth, J.D. Glycosylation in cellular mechanisms of health and disease. Cell 2006, 126, 855–867. [Google Scholar] [CrossRef] [PubMed]
- Varki, A.; Cummings, R.D.; Esko, J.D.; Freeze, H.H.; Stanley, P.; Bertozzi, C.R.; Hart, G.W.; Etzler, M.E. Essentials of Glycobiology, 2nd ed.; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 2009. [Google Scholar]
- Hakomori, S. Glycosylation defining cancer malignancy: New wine in an old bottle. Proc. Natl. Acad. Sci. USA 2002, 99, 10231–10233. [Google Scholar] [CrossRef] [PubMed]
- Drake, P.M.; Cho, W.; Li, B.; Prakobphol, A.; Johansen, E.; Anderson, N.L.; Regnier, F.E.; Gibson, B.W.; Fisher, S.J. Sweetening the pot: Adding glycosylation to the biomarker discovery equation. Clin. Chem. 2010, 56, 223–236. [Google Scholar] [CrossRef] [PubMed]
- Hakomori, S. Aberrant glycosylation in tumors and tumor-associated carbohydrate antigens. Adv. Cancer Res. 1989, 52, 257–331. [Google Scholar]
- Fuster, M.M.; Esko, J.D. The sweet and sour of cancer: Glycans as novel therapeutic targets. Nat. Rev. Cancer 2005, 5, 526–542. [Google Scholar] [CrossRef] [PubMed]
- Christiansen, M.N.; Chik, J.; Lee, L.; Anugraham, M.; Abrahams, J.L.; Packer, N.H. Cell surface protein glycosylation in cancer. Proteomics 2014, 14, 525–546. [Google Scholar] [CrossRef] [PubMed]
- Shelton, B.K. Introduction to colorectal cancer. In Seminars in Oncology Nursing; Elsevier: Amsterdam, The Netherlands, 2002; pp. 2–12. [Google Scholar]
- Vogelstein, B.; Kinzler, K.W. The multistep nature of cancer. Trends Genet. 1993, 9, 138–141. [Google Scholar] [CrossRef]
- Behrens, J. The role of the Wnt signalling pathway in colorectal tumorigenesis. Biochem. Soc. Trans. 2005, 33, 672–675. [Google Scholar] [CrossRef] [PubMed]
- Schwitalle, Y.; Linnebacher, M.; Ripberger, E.; Gebert, J.; von Knebel Doeberitz, M. Immunogenic peptides generated by frameshift mutations in DNA mismatch repair-deficient cancer cells. Cancer Immun. 2004, 4, 14. [Google Scholar] [PubMed]
- Davies, H.; Bignell, G.R.; Cox, C.; Stephens, P.; Edkins, S.; Clegg, S.; Teague, J.; Woffendin, H.; Garnett, M.J.; Bottomley, W.; et al. Mutations of the BRAF gene in human cancer. Nature 2002, 417, 949–954. [Google Scholar] [CrossRef] [PubMed]
- Munemitsu, S.; Albert, I.; Souza, B.; Rubinfeld, B.; Polakis, P. Regulation of intracellular beta-catenin levels by the adenomatous polyposis coli (APC) tumor-suppressor protein. Proc. Natl. Acad. Sci. USA 1995, 92, 3046–3050. [Google Scholar] [CrossRef] [PubMed]
- Jemal, A.; Bray, F.; Center, M.M.; Ferlay, J.; Ward, E.; Forman, D. Global cancer statistics. CA Cancer J. Clin. 2011, 61, 69–90. [Google Scholar] [CrossRef] [PubMed]
- Majumdar, S.R.; Fletcher, R.H.; Evans, A.T. How does colorectal cancer present? symptoms, duration, and clues to location. Am. J. Gastroenterol. 1999, 94, 3039–3045. [Google Scholar] [PubMed]
- Astin, M.; Griffin, T.; Neal, R.D.; Rose, P.; Hamilton, W. The diagnostic value of symptoms for colorectal cancer in primary care: A systematic review. Br. J. Gen. Pract. 2011, 61, e231–e243. [Google Scholar] [CrossRef] [PubMed]
- Hamilton, W.; Sharp, D. Diagnosis of colorectal cancer in primary care: The evidence base for guidelines. Fam. Pract. 2004, 21, 99–106. [Google Scholar] [CrossRef] [PubMed]
- Douillard, J.Y.; Cunningham, D.; Roth, A.D.; Navarro, M.; James, R.D.; Karasek, P.; Jandik, P.; Iveson, T.; Carmichael, J.; Alakl, M.; et al. Irinotecan combined with fluorouracil compared with fluorouracil alone as first-line treatment for metastatic colorectal cancer: A multicentre randomised trial. Lancet 2000, 355, 1041–1047. [Google Scholar] [CrossRef]
- Saltz, L.B.; Cox, J.V.; Blanke, C.; Rosen, L.S.; Fehrenbacher, L.; Moore, M.J.; Maroun, J.A.; Ackland, S.P.; Locker, P.K.; Pirotta, N.; et al. Irinotecan plus fluorouracil and leucovorin for metastatic colorectal cancer. Irinotecan Study Group. N. Engl. J. Med. 2000, 343, 905–914. [Google Scholar] [CrossRef] [PubMed]
- Savary, J.B.; Vasseur, F.; Flactif, M.; Willatt, L.; Lefebvre, J.; Ferguson-Smith, M.A.; Deminatti, M.M. Cytogenetic and molecular investigations of an abnormal Y chromosome: Evidence for a pseudo-dicentric (Yq) isochromosome. Ann. Genet. 1992, 35, 134–139. [Google Scholar] [PubMed]
- Van Cutsem, E.; Geboes, K. The multidisciplinary management of gastrointestinal cancer. The integration of cytotoxics and biologicals in the treatment of metastatic colorectal cancer. Best Pract. Res. Clin. Gastroenterol. 2007, 21, 1089–1108. [Google Scholar] [CrossRef] [PubMed]
- Punt, C. New options and old dilemmas in the treatment of patients with advanced colorectal cancer. Ann. Oncol. 2004, 15, 1453–1459. [Google Scholar] [CrossRef] [PubMed]
- Varki, A. Biological roles of oligosaccharides: All of the theories are correct. Glycobiology 1993, 3, 97–130. [Google Scholar] [CrossRef] [PubMed]
- Dube, D.H.; Bertozzi, C.R. Glycans in cancer and inflammation—Potential for therapeutics and diagnostics. Nat. Rev. Drug Discov. 2005, 4, 477–488. [Google Scholar] [CrossRef] [PubMed]
- Stanley, P.; Schachter, H.; Taniguchi, N. N-Glycans. In Essentials of Glycobiology, 2nd ed.; Varki, A., Cummings, R.D., Esko, J.D., Freeze, H.H., Stanley, P., Bertozzi, C.R., Hart, G.W., Etzler, M.E., Eds.; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 2009. [Google Scholar]
- Schachter, H. Paucimannose N-glycans in Caenorhabditis elegans and Drosophila melanogaster. Carbohydr. Res. 2009, 344, 1391–1396. [Google Scholar] [CrossRef] [PubMed]
- Sethi, M.K.; Thaysen-Andersen, M.; Smith, J.T.; Baker, M.S.; Packer, N.H.; Hancock, W.S.; Fanayan, S. Comparative N-glycan profiling of colorectal cancer cell lines reveals unique bisecting GlcNAc and alpha-2,3-linked sialic acid determinants are associated with membrane proteins of the more metastatic/aggressive cell lines. J. Proteome Res. 2014, 13, 277–288. [Google Scholar] [CrossRef] [PubMed]
- Everest-Dass, A.V.; Jin, D.; Thaysen-Andersen, M.; Nevalainen, H.; Kolarich, D.; Packer, N.H. Comparative structural analysis of the glycosylation of salivary and buccal cell proteins: Innate protection against infection by Candida albicans. Glycobiology 2012, 22, 1465–1479. [Google Scholar] [CrossRef] [PubMed]
- Balog, C.I.; Stavenhagen, K.; Fung, W.L.; Koeleman, C.A.; McDonnell, L.A.; Verhoeven, A.; Mesker, W.E.; Tollenaar, R.A.; Deelder, A.M.; Wuhrer, M. N-glycosylation of colorectal cancer tissues: A liquid chromatography and mass spectrometry-based investigation. Mol. Cell. Proteom. 2012, 11, 571–585. [Google Scholar] [CrossRef] [PubMed]
- Venkatakrishnan, V.; Thaysen-Andersen, M.; Chen, S.C.; Nevalainen, H.; Packer, N.H. Cystic fibrosis and bacterial colonization define the sputum N-glycosylation phenotype. Glycobiology 2015, 25, 88–100. [Google Scholar] [CrossRef] [PubMed]
- Thaysen-Andersen, M.; Venkatakrishnan, V.; Loke, I.; Laurini, C.; Diestel, S.; Parker, B.L.; Packer, N.H. Human neutrophils secrete bioactive paucimannosidic proteins from azurophilic granules into pathogen-infected sputum. J. Biol. Chem. 2015, 290, 8789–8802. [Google Scholar] [CrossRef] [PubMed]
- Varki, A.; Lowe, J.B. Biological Roles of Glycans. In Essentials of Glycobiology, 2nd ed.; Varki, A., Cummings, R.D., Esko, J.D., Freeze, H.H., Stanley, P., Bertozzi, C.R., Hart, G.W., Etzler, M.E., Eds.; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 2009. [Google Scholar]
- Rogers, G.N.; Daniels, R.S.; Skehel, J.J.; Wiley, D.C.; Wang, X.F.; Higa, H.H.; Paulson, J.C. Host-mediated selection of influenza virus receptor variants. Sialic acid-alpha 2,6Gal-specific clones of A/duck/Ukraine/1/63 revert to sialic acid-alpha 2,3Gal-specific wild type in ovo. J. Biol. Chem. 1985, 260, 7362–7367. [Google Scholar] [PubMed]
- Kansas, G.S. Selectins and their ligands: Current concepts and controversies. Blood 1996, 88, 3259–3287. [Google Scholar] [PubMed]
- Helenius, A. How N-linked oligosaccharides affect glycoprotein folding in the endoplasmic reticulum. Mol. Biol. Cell 1994, 5, 253–265. [Google Scholar] [CrossRef] [PubMed]
- Gabel, C.A.; Foster, S.A. Lysosomal enzyme trafficking in mannose 6-phosphate receptor-positive mouse L-cells: Demonstration of a steady state accumulation of phosphorylated acid hydrolases. J. Cell Biol. 1986, 102, 943–950. [Google Scholar] [CrossRef] [PubMed]
- Herzog, V.; Neumuller, W.; Holzmann, B. Thyroglobulin, the major and obligatory exportable protein of thyroid follicle cells, carries the lysosomal recognition marker mannose-6-phosphate. EMBO J. 1987, 6, 555–560. [Google Scholar] [PubMed]
- Zhao, Y.Y.; Takahashi, M.; Gu, J.G.; Miyoshi, E.; Matsumoto, A.; Kitazume, S.; Taniguchi, N. Functional roles of N-glycans in cell signaling and cell adhesion in cancer. Cancer Sci. 2008, 99, 1304–1310. [Google Scholar] [CrossRef] [PubMed]
- Fiedler, K.; Simons, K. The role of N-glycans in the secretory pathway. Cell 1995, 81, 309–312. [Google Scholar] [CrossRef]
- Holst, S.; Wuhrer, M.; Rombouts, Y. Glycosylation characteristics of colorectal cancer. Adv. Cancer Res. 2015, 126, 203–256. [Google Scholar] [PubMed]
- Meany, D.L.; Chan, D.W. Aberrant glycosylation associated with enzymes as cancer biomarkers. Clin. Proteom. 2011, 8, 7. [Google Scholar] [CrossRef] [PubMed]
- Pinho, S.S.; Oliveira, P.; Cabral, J.; Carvalho, S.; Huntsman, D.; Gartner, F.; Seruca, R.; Reis, C.A.; Oliveira, C. Loss and recovery of Mgat3 and GnT-III Mediated E-cadherin N-glycosylation is a mechanism involved in epithelial-mesenchymal-epithelial transitions. PLoS ONE 2012, 7, e33191. [Google Scholar] [CrossRef] [PubMed]
- Taniguchi, N.; Korekane, H. Branched N-glycans and their implications for cell adhesion, signaling and clinical applications for cancer biomarkers and in therapeutics. BMB Rep. 2011, 44, 772–781. [Google Scholar] [CrossRef] [PubMed]
- Li, W.P.; Zuber, C.; Heitz, P.U.; Roth, J. Cytochemical staining for beta 1,6 branching of asparagine-linked oligosaccharides in variants of metastatic human colon carcinoma cells. Am. J. Pathol. 1994, 145, 470–480. [Google Scholar] [PubMed]
- Murata, K.; Miyoshi, E.; Kameyama, M.; Ishikawa, O.; Kabuto, T.; Sasaki, Y.; Hiratsuka, M.; Ohigashi, H.; Ishiguro, S.; Ito, S.; et al. Expression of N-acetylglucosaminyltransferase V in colorectal cancer correlates with metastasis and poor prognosis. Clin. Cancer Res. 2000, 6, 1772–1777. [Google Scholar] [PubMed]
- Dennis, J.W.; Laferte, S.; Waghorne, C.; Breitman, M.L.; Kerbel, R.S. Beta 1–6 branching of Asn-linked oligosaccharides is directly associated with metastasis. Science 1987, 236, 582–585. [Google Scholar] [CrossRef] [PubMed]
- Dall’Olio, F.; Chiricolo, M. Sialyltransferases in cancer. Glycoconjug. J. 2001, 18, 841–850. [Google Scholar] [CrossRef]
- Harduin-Lepers, A.; Krzewinski-Recchi, M.A.; Colomb, F.; Foulquier, F.; Groux-Degroote, S.; Delannoy, P. Sialyltransferases functions in cancers. Front. Biosci. 2012, 4, 499–515. [Google Scholar] [CrossRef]
- Dall’Olio, F.; Malagolini, N.; di Stefano, G.; Minni, F.; Marrano, D.; Serafini-Cessi, F. Increased CMP-NeuAc:Gal beta 1,4GlcNAc-R alpha 2,6 sialyltransferase activity in human colorectal cancer tissues. Int. J. Cancer 1989, 44, 434–439. [Google Scholar] [PubMed]
- Gessner, P.; Riedl, S.; Quentmaier, A.; Kemmner, W. Enhanced activity of CMP-neuAc:Gal beta 1,4GlcNAc:alpha 2,6-sialyltransferase in metastasizing human colorectal tumor tissue and serum of tumor patients. Cancer Lett. 1993, 75, 143–149. [Google Scholar] [CrossRef]
- Sethi, M.K.; Kim, H.; Park, C.K.; Baker, M.S.; Paik, Y.K.; Packer, N.H.; Hancock, W.S.; Fanayan, S.; Thaysen-Andersen, M. In-depth N-glycome profiling of paired colorectal cancer and non-tumorigenic tissues reveals cancer-, stage- and EGFR-specific protein N-glycosylation. Glycobiology 2015, 25, 1064–1078. [Google Scholar] [CrossRef] [PubMed]
- Harvey, B.E.; Toth, C.A.; Wagner, H.E.; Steele, G.D., Jr.; Thomas, P. Sialyltransferase activity and hepatic tumor growth in a nude mouse model of colorectal cancer metastases. Cancer Res. 1992, 52, 1775–1779. [Google Scholar] [PubMed]
- Park, J.J.; Lee, M. Increasing the alpha 2, 6 sialylation of glycoproteins may contribute to metastatic spread and therapeutic resistance in colorectal cancer. Gut Liver 2013, 7, 629–641. [Google Scholar] [CrossRef] [PubMed]
- Cazet, A.; Julien, S.; Bobowski, M.; Krzewinski-Recchi, M.A.; Harduin-Lepers, A.; Groux-Degroote, S.; Delannoy, P. Consequences of the expression of sialylated antigens in breast cancer. Carbohydr.Res. 2010, 345, 1377–1383. [Google Scholar] [CrossRef] [PubMed]
- Itzkowitz, S.H.; Bloom, E.J.; Kokal, W.A.; Modin, G.; Hakomori, S.; Kim, Y.S. Sialosyl-Tn. A novel mucin antigen associated with prognosis in colorectal cancer patients. Cancer 1990, 66, 1960–1966. [Google Scholar] [CrossRef]
- Marcos, N.T.; Bennett, E.P.; Gomes, J.; Magalhaes, A.; Gomes, C.; David, L.; Dar, I.; Jeanneau, C.; DeFrees, S.; Krustrup, D.; et al. ST6GalNAc-I controls expression of sialyl-Tn antigen in gastrointestinal tissues. Front. Biosci. 2011, 3, 1443–1455. [Google Scholar] [CrossRef]
- Varki, A. Selectin ligands. Proc. Natl. Acad. Sci. USA 1994, 91, 7390–7397. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.J.; Borsig, L.; Varki, N.M.; Varki, A. P-selectin deficiency attenuates tumor growth and metastasis. Proc. Natl. Acad. Sci. USA 1998, 95, 9325–9330. [Google Scholar] [CrossRef] [PubMed]
- Borsig, L.; Wong, R.; Hynes, R.O.; Varki, N.M.; Varki, A. Synergistic effects of L- and P-selectin in facilitating tumor metastasis can involve non-mucin ligands and implicate leukocytes as enhancers of metastasis. Proc. Natl. Acad. Sci. USA 2002, 99, 2193–2198. [Google Scholar] [CrossRef] [PubMed]
- Perez-Garay, M.; Arteta, B.; Llop, E.; Cobler, L.; Pages, L.; Ortiz, R.; Ferri, M.J.; de Bolo, C.; Figueras, J.; de Llorens, R.; et al. alpha2,3-Sialyltransferase ST3Gal IV promotes migration and metastasis in pancreatic adenocarcinoma cells and tends to be highly expressed in pancreatic adenocarcinoma tissues. Int. J. Biochem. Cell Biol. 2013, 45, 1748–1757. [Google Scholar] [CrossRef] [PubMed]
- Nakamori, S.; Kameyama, M.; Imaoka, S.; Furukawa, H.; Ishikawa, O.; Sasaki, Y.; Izumi, Y.; Irimura, T. Involvement of carbohydrate antigen sialyl Lewis(x) in colorectal cancer metastasis. Dis. Colon Rectum 1997, 40, 420–431. [Google Scholar] [CrossRef] [PubMed]
- Grabowski, P.; Mann, B.; Mansmann, U.; Lovin, N.; Foss, H.D.; Berger, G.; Scherubl, H.; Riecken, E.O.; Buhr, H.J.; Hanski, C. Expression of SIALYL-Le(x) antigen defined by MAb AM-3 is an independent prognostic marker in colorectal carcinoma patients. Int. J. Cancer 2000, 88, 281–286. [Google Scholar] [CrossRef]
- Kumamoto, K.; Goto, Y.; Sekikawa, K.; Takenoshita, S.; Ishida, N.; Kawakita, M.; Kannagi, R. Increased expression of UDP-galactose transporter messenger RNA in human colon cancer tissues and its implication in synthesis of Thomsen-Friedenreich antigen and sialyl Lewis A/X determinants. Cancer Res. 2001, 61, 4620–4627. [Google Scholar] [PubMed]
- Yusa, A.; Miyazaki, K.; Kimura, N.; Izawa, M.; Kannagi, R. Epigenetic silencing of the sulfate transporter gene DTDST induces sialyl Lewisx expression and accelerates proliferation of colon cancer cells. Cancer Res. 2010, 70, 4064–4073. [Google Scholar] [CrossRef] [PubMed]
- Sasai, K.; Ikeda, Y.; Eguchi, H.; Tsuda, T.; Honke, K.; Taniguchi, N. The action of N-acetylglucosaminyltransferase-V is prevented by the bisecting GlcNAc residue at the catalytic step. FEBS Lett. 2002, 522, 151–155. [Google Scholar] [CrossRef]
- Zhao, Y.; Nakagawa, T.; Itoh, S.; Inamori, K.; Isaji, T.; Kariya, Y.; Kondo, A.; Miyoshi, E.; Miyazaki, K.; Kawasaki, N.; et al. N-acetylglucosaminyltransferase III antagonizes the effect of N-acetylglucosaminyltransferase V on alpha3beta1 integrin-mediated cell migration. J. Biol. Chem. 2006, 281, 32122–32130. [Google Scholar] [CrossRef] [PubMed]
- Dalziel, M.; Whitehouse, C.; McFarlane, I.; Brockhausen, I.; Gschmeissner, S.; Schwientek, T.; Clausen, H.; Burchell, J.M.; Taylor-Papadimitriou, J. The relative activities of the C2GnT1 and ST3Gal-I glycosyltransferases determine O-glycan structure and expression of a tumor-associated epitope on MUC1. J. Biol. Chem. 2001, 276, 11007–11015. [Google Scholar] [CrossRef] [PubMed]
- Schneider, F.; Kemmner, W.; Haensch, W.; Franke, G.; Gretschel, S.; Karsten, U.; Schlag, P.M. Overexpression of sialyltransferase CMP-sialic acid:Galbeta1,3GalNAc-R alpha6-Sialyltransferase is related to poor patient survival in human colorectal carcinomas. Cancer Res. 2001, 61, 4605–4611. [Google Scholar] [PubMed]
- Miyagi, T.; Wada, T.; Yamaguchi, K.; Shiozaki, K.; Sato, I.; Kakugawa, Y.; Yamanami, H.; Fujiya, T. Human sialidase as a cancer marker. Proteomics 2008, 8, 3303–3311. [Google Scholar] [CrossRef] [PubMed]
- Kato, T.; Wang, Y.; Yamaguchi, K.; Milner, C.M.; Shineha, R.; Satomi, S.; Miyagi, T. Overexpression of lysosomal-type sialidase leads to suppression of metastasis associated with reversion of malignant phenotype in murine B16 melanoma cells. Int. J. Cancer 2001, 92, 797–804. [Google Scholar] [CrossRef] [PubMed]
- Sawada, M.; Moriya, S.; Saito, S.; Shineha, R.; Satomi, S.; Yamori, T.; Tsuruo, T.; Kannagi, R.; Miyagi, T. Reduced sialidase expression in highly metastatic variants of mouse colon adenocarcinoma 26 and retardation of their metastatic ability by sialidase overexpression. Int. J. Cancer 2002, 97, 180–185. [Google Scholar] [CrossRef] [PubMed]
- Reis, C.A.; Osorio, H.; Silva, L.; Gomes, C.; David, L. Alterations in glycosylation as biomarkers for cancer detection. J. Clin. Pathol. 2010, 63, 322–329. [Google Scholar] [CrossRef] [PubMed]
- Adamczyk, B.; Tharmalingam, T.; Rudd, P.M. Glycans as cancer biomarkers. Biochim. Biophys. Acta 2012, 1820, 1347–1353. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; LaRoche, T.; Hamelinck, D.; Bergsma, D.; Brenner, D.; Simeone, D.; Brand, R.E.; Haab, B.B. Multiplexed analysis of glycan variation on native proteins captured by antibody microarrays. Nat. Methods 2007, 4, 437–444. [Google Scholar] [CrossRef] [PubMed]
- Tanahashi, T.; Namba, K.; Murao, T. Studies on relationship between histology, tumor markers (prostatic acid phosphatase.prostate specific antigen.gamma-seminoprotein.leu-7) and clinical course in prostate cancer. Jpn. J. Urol. 1990, 81, 680–685. [Google Scholar] [CrossRef]
- Peracaula, R.; Barrabés, S.; Sarrats, A.; Rudd, P.M.; de Llorens, R. Altered glycosylation in tumours focused to cancer diagnosis. Dis. Markers 2008, 25, 207–218. [Google Scholar] [CrossRef] [PubMed]
- Luque-Garcia, J.L.; Martinez-Torrecuadrada, J.L.; Epifano, C.; Canamero, M.; Babel, I.; Casal, J.I. Differential protein expression on the cell surface of colorectal cancer cells associated to tumor metastasis. Proteomics 2010, 10, 940–952. [Google Scholar] [PubMed]
- Fanayan, S.; Smith, J.T.; Lee, L.Y.; Yan, F.; Snyder, M.; Hancock, W.S.; Nice, E. Proteogenomic analysis of human colon carcinoma cell lines LIM1215, LIM1899, and LIM2405. J. Proteome Res. 2013, 12, 1732–1742. [Google Scholar] [CrossRef] [PubMed]
- Gan, Y.; Chen, D.; Li, X. Proteomic analysis reveals novel proteins associated with progression and differentiation of colorectal carcinoma. J. Cancer Res. Ther. 2014, 10, 89–96. [Google Scholar] [PubMed]
- Sethi, M.K.; Thaysen-Andersen, M.; Kim, H.; Park, C.K.; Baker, M.S.; Packer, N.H.; Paik, Y.K.; Hancock, W.S.; Fanayan, S. Quantitative proteomic analysis of paired colorectal cancer and non-tumorigenic tissues reveals signature proteins and perturbed pathways involved in CRC progression and metastasis. J. Proteom. 2015, 126, 54–67. [Google Scholar] [CrossRef] [PubMed]
- Thaysen-Andersen, M.; Packer, N.H. Advances in LC-MS/MS-based glycoproteomics: Getting closer to system-wide site-specific mapping of the N- and O-glycoproteome. Biochim. Biophys. Acta 2014, 1844, 1437–1452. [Google Scholar] [CrossRef] [PubMed]
- Jensen, P.H.; Karlsson, N.G.; Kolarich, D.; Packer, N.H. Structural analysis of N- and O-glycans released from glycoproteins. Nat. Protoc. 2012, 7, 1299–1310. [Google Scholar] [CrossRef] [PubMed]
- Frost, D.C.; Li, L. Recent advances in mass spectrometry-based glycoproteomics. Adv. Protein Chem. Struct. Biol. 2014, 95, 71–123. [Google Scholar] [PubMed]
- Saitoh, O.; Wang, W.C.; Lotan, R.; Fukud, M. Differential glycosylation and cell surface expression of lysosomal membrane glycoproteins in sublines of a human colon cancer exhibiting distinct metastatic potentials. J. Biol. Chem. 1992, 267, 5700–5711. [Google Scholar] [PubMed]
- Vierbuchen, M.J.; Fruechtnicht, W.; Brackrock, S.; Krause, K.T.; Zienkiewicz, T.J. Quantitative lectin-histochemical and immunohistochemical studies on the occurrence of alpha(2,3)- and alpha(2,6)-linked sialic acid residues in colorectal carcinomas. Relation to clinicopathologic features. Cancer 1995, 76, 727–735. [Google Scholar] [CrossRef]
- Muinelo-Romay, L.; Vazquez-Martin, C.; Villar-Portela, S.; Cuevas, E.; Gil-Martin, E.; Fernandez-Briera, A. Expression and enzyme activity of alpha(1,6)fucosyltransferase in human colorectal cancer. Int. J. Cancer 2008, 123, 641–646. [Google Scholar] [CrossRef] [PubMed]
- Fernandez-Rodriguez, J.; Feijoo-Carnero, C.; Merino-Trigo, A.; Paez de la Cadena, M.; Rodriguez-Berrocal, F.J.; de Carlos, A.; Butron, M.; Martinez-Zorzano, V.S. Immunohistochemical analysis of sialic acid and fucose composition in human colorectal adenocarcinoma. Tumour Biol. 2000, 21, 153–164. [Google Scholar] [PubMed]
- Vercoutter-Edouart, A.S.; Slomianny, M.C.; Dekeyzer-Beseme, O.; Haeuw, J.F.; Michalsk, J.C. Glycoproteomics and glycomics investigation of membrane N-glycosylproteins from human colon carcinoma cells. Proteomics 2008, 8, 3236–3256. [Google Scholar] [CrossRef] [PubMed]
- Qiu, Y.; Patwa, T.H.; Xu, L.; Shedden, K.; Misek, D.E.; Tuck, M.; Jin, G.; Ruffin, M.T.; Turgeon, D.K.; Synal, S.; et al. Plasma glycoprotein profiling for colorectal cancer biomarker identification by lectin glycoarray and lectin blot. J. Proteome Res. 2008, 7, 1693–1703. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.P.; Ruan, C.P.; Wang, H.; Hu, Z.Q.; Fang, M.; Gu, X.; Ji, J.; Zhao, J.Y.; Gao, C.F. Identification and assessment of new biomarkers for colorectal cancer with serum N-glycan profiling. Cancer 2012, 118, 639–650. [Google Scholar] [CrossRef] [PubMed]
- Fukasawa, T.; Asao, T.; Yamauchi, H.; Ide, M.; Tabe, Y.; Fujii, T.; Yamaguchi, S.; Tsutsumi, S.; Yazawa, S.; Kuwano, H. Associated expression of alpha2,3sialylated type 2 chain structures with lymph node metastasis in distal colorectal cancer. Surg. Today 2013, 43, 155–162. [Google Scholar] [CrossRef] [PubMed]
- Chik, J.H.; Zhou, J.; Moh, E.S.; Christopherson, R.; Clarke, S.J.; Molloy, M.P.; Packer, N.H. Comprehensive glycomics comparison between colon cancer cell cultures and tumours: Implications for biomarker studies. J. Proteom. 2014, 108, 146–162. [Google Scholar] [CrossRef] [PubMed]
- Kaprio, T.; Satomaa, T.; Heiskanen, A.; Hokke, C.H.; Deelder, A.M.; Mustone, H.; Hagstrom, J.; Carpen, O.; Saarinen, J.; Haglund, C. N-glycomic profiling as a tool to separate rectal adenomas from carcinomas. Mol. Cell. Proteom. 2014, 14, 277–288. [Google Scholar] [CrossRef] [PubMed]
- Maley, F.; Trimble, R.B.; Tarentino, A.L.; Plummer, T.H., Jr. Characterization of glycoproteins and their associated oligosaccharides through the use of endoglycosidases. Anal. Biochem. 1989, 180, 195–204. [Google Scholar] [CrossRef]
- Tretter, V.; Altmann, F.; Marz, L. Peptide-N4-(N-acetyl-beta-glucosaminyl)asparagine amidase F cannot release glycans with fucose attached alpha 1–3 to the asparagine-linked N-acetylglucosamine residue. Eur. J. Biochem. 1991, 199, 647–652. [Google Scholar] [CrossRef] [PubMed]
- O’Neill, R.A. Enzymatic release of oligosaccharides from glycoproteins for chromatographic and electrophoretic analysis. J. Chromatogr. A 1996, 720, 201–215. [Google Scholar] [CrossRef]
- Mechref, Y.; Novotny, M.V. Structural investigations of glycoconjugates at high sensitivity. Chem. Rev. 2002, 102, 321–369. [Google Scholar] [CrossRef] [PubMed]
- Geyer, H.; Geyer, R. Strategies for analysis of glycoprotein glycosylation. Biochim. Biophys. Acta 2006, 1764, 1853–1869. [Google Scholar] [CrossRef] [PubMed]
- Ruhaak, L.R.; Zauner, G.; Huhn, C.; Bruggink, C.; Deelder, A.M.; Wuhrer, M. Glycan labeling strategies and their use in identification and quantification. Anal. Bioanal. Chem. 2010, 397, 3457–3481. [Google Scholar] [CrossRef] [PubMed]
- Anugraham, M.; Jacob, F.; Nixdorf, S.; Everest-Dass, A.V.; Heinzelmann-Schwarz, V.; Packer, N.H. Specific glycosylation of membrane proteins in epithelial ovarian cancer cell lines: Glycan structures reflect gene expression and DNA methylation status. Mol. Cell. Proteom. 2014, 13, 2213–2232. [Google Scholar] [CrossRef] [PubMed]
- Lee, L.Y.; Thaysen-Andersen, M.; Baker, M.S.; Packer, N.H.; Hancock, W.S.; Fanayan, S. Comprehensive N-glycome profiling of cultured human epithelial breast cells identifies unique secretome N-glycosylation signatures enabling tumorigenic subtype classification. J. Proteome Res. 2014, 13, 4783–4795. [Google Scholar] [CrossRef] [PubMed]
- Dell, A.; Reason, A.J.; Khoo, K.H.; Panico, M.; McDowell, R.A.; Morris, H.R. Mass spectrometry of carbohydrate-containing biopolymers. Methods Enzymol. 1994, 230, 108–132. [Google Scholar] [PubMed]
- Alvarez-Manilla, G.; Warren, N.L.; Abney, T.; Atwood, J., III; Azadi, P.; York, W.S.; Pierce, M.; Orlando, R. Tools for glycomics: Relative quantitation of glycans by isotopic permethylation using 13CH3I. Glycobiology 2007, 17, 677–687. [Google Scholar] [CrossRef] [PubMed]
- Bigge, J.C.; Patel, T.P.; Bruce, J.A.; Goulding, P.N.; Charles, S.M.; Parekh, R.B. Nonselective and efficient fluorescent labeling of glycans using 2-amino benzamide and anthranilic acid. Anal. Biochem. 1995, 230, 229–238. [Google Scholar] [CrossRef] [PubMed]
- Abd Hamid, U.M.; Royle, L.; Saldova, R.; Radcliffe, C.M.; Harvey, D.J.; Storr, S.J.; Pardo, M.; Antrobus, R.; Chapman, C.J.; Zitzmann, N.; et al. A strategy to reveal potential glycan markers from serum glycoproteins associated with breast cancer progression. Glycobiology 2008, 18, 1105–1118. [Google Scholar] [CrossRef] [PubMed]
- Arnold, J.N.; Saldova, R.; Galligan, M.C.; Murphy, T.B.; Mimura-Kimura, Y.; Telford, J.E.; Godwin, A.K.; Rudd, P.M. Novel glycan biomarkers for the detection of lung cancer. J. Proteome Res. 2011, 10, 1755–1764. [Google Scholar] [CrossRef] [PubMed]
- Wuhrer, M.; Deelder, A.M.; Hokke, C.H. Protein glycosylation analysis by liquid chromatography-mass spectrometry. J. Chromatogr. B Anal. Technol. Biomed. Life Sci. 2005, 825, 124–133. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Flynn, G.C. Analysis of N-glycans from recombinant immunoglobulin G by on-line reversed-phase high-performance liquid chromatography/mass spectrometry. Anal. Biochem. 2007, 370, 147–161. [Google Scholar] [CrossRef] [PubMed]
- Wuhrer, M.; Koeleman, C.A.; Deelder, A.M. Two-dimensional HPLC separation with reverse-phase-nano-LC-MS/MS for the characterization of glycan pools after labeling with 2-aminobenzamide. Methods Mol. Biol. 2009, 534, 79–91. [Google Scholar] [PubMed]
- Delaney, J.; Vouros, P. Liquid chromatography ion trap mass spectrometric analysis of oligosaccharides using permethylated derivatives. Rapid Commun. Mass Spectrom. 2001, 15, 325–334. [Google Scholar] [CrossRef] [PubMed]
- Hemstrom, P.; Irgum, K. Hydrophilic interaction chromatography. J. Sep. Sci. 2006, 29, 1784–1821. [Google Scholar] [CrossRef] [PubMed]
- Han, X.; Aslanian, A.; Yates, J.R., III. Mass spectrometry for proteomics. Curr. Opin. Chem. Biol. 2008, 12, 483–490. [Google Scholar] [CrossRef] [PubMed]
- Hao, Z.; Xiao, B.; Weng, N. Impact of column temperature and mobile phase components on selectivity of hydrophilic interaction chromatography (HILIC). J. Sep. Sci. 2008, 31, 1449–1464. [Google Scholar] [CrossRef] [PubMed]
- Rohrer, J.S.; Basumallick, L.; Hurum, D. High-performance anion-exchange chromatography with pulsed amperometric detection for carbohydrate analysis of glycoproteins. Biochemistry 2013, 78, 697–709. [Google Scholar] [CrossRef] [PubMed]
- Gilbert, M.T.; Knox, J.H. Preparation of porous carbon. Available online: https://www.google.com/patents/US4263268 (acessed on 30 October 2015).
- Knox, J.H.; Kaur, B.; Millward, G. Structure and performance of porous graphitic carbon in liquid chromatography. J. Chromatogr. A 1986, 352, 3–25. [Google Scholar] [CrossRef]
- Stavenhagen, K.; Kolarich, D.; Wuhrer, M. Clinical Glycomics Employing Graphitized Carbon Liquid Chromatography—Mass Spectrometry. Chromatographia 2014, 78, 307–320. [Google Scholar] [CrossRef] [PubMed]
- Pabst, M.; Grass, J.; Toegel, S.; Liebminger, E.; Strasser, R.; Altmann, F. Isomeric analysis of oligomannosidic N-glycans and their dolichol-linked precursors. Glycobiology 2012, 22, 389–399. [Google Scholar] [CrossRef] [PubMed]
- Dole, M.; Mack, L.; Hines, R.; Mobley, R.C.; Ferguson, L.D.; Alice, M.B. Molecular beams of macroions. J. Chem. Phys. 1968, 49, 2240–2249. [Google Scholar] [CrossRef]
- Fenn, J.B.; Mann, M.; Meng, C.K.; Wong, S.F.; Whitehouse, C.M. Electrospray ionization for mass spectrometry of large biomolecules. Science 1989, 246, 64–71. [Google Scholar] [CrossRef] [PubMed]
- Karas, M.; Bachmann, D.; Bahr, U.; Hillenkamp, F. Matrix-assisted ultraviolet laser desorption of non-volatile compounds. Int. J. Mass Spectrom. Ion Process. 1987, 78, 53–68. [Google Scholar] [CrossRef]
- Tanaka, K.; Waki, H.; Ido, Y.; Akita, S.; Yoshida, Y.; Yoshida, T.; Matsuo, T. Protein and polymer analyses up to m/z 100 000 by laser ionization time-of-flight mass spectrometry. Rapid Commun. Mass Spectrom. 1988, 2, 151–153. [Google Scholar] [CrossRef]
- Emmett, M.R.; Caprioli, R.M. Micro-electrospray mass spectrometry: Ultra-high-sensitivity analysis of peptides and proteins. J. Am. Soc. Mass Spectrom. 1994, 5, 605–613. [Google Scholar] [CrossRef]
- Yates, J.R. Mass spectrometry and the age of the proteome. J. Mass Spectrom. 1998, 33, 1–19. [Google Scholar] [CrossRef]
- Beavis, R.C.; Chait, B.T. High-accuracy molecular mass determination of proteins using matrix-assisted laser desorption mass spectrometry. Anal. Chem. 1990, 62, 1836–1840. [Google Scholar] [CrossRef] [PubMed]
- Silvertand, L.H.; Torano, J.S.; de Jong, G.J.; van Bennekom, W.P. Improved repeatability and matrix-assisted desorption/ionization—Time of flight mass spectrometry compatibility in capillary isoelectric focusing. Electrophoresis 2008, 29, 1985–1996. [Google Scholar] [CrossRef] [PubMed]
- Zheng, J.; Li, N.; Ridyard, M.; Dai, H.; Robbins, S.M.; Li, L. Simple and robust two-layer matrix/sample preparation method for MALDI MS/MS analysis of peptides. J. Proteome Res. 2005, 4, 1709–1716. [Google Scholar] [CrossRef] [PubMed]
- Zaia, J. Mass spectrometry and the emerging field of glycomics. Chem. Biol. 2008, 15, 881–892. [Google Scholar] [CrossRef] [PubMed]
- Han, L.; Costello, C.E. Mass spectrometry of glycans. Biochemistry 2013, 78, 710–720. [Google Scholar] [CrossRef] [PubMed]
- Domon, B.; Costello, C.E. A systematic nomenclature for carbohydrate fragmentations in FAB-MS/MS spectra of glycoconjugates. Glycoconjug. J. 1988, 5, 397–409. [Google Scholar] [CrossRef]
- Leymarie, N.; Zaia, J. Effective use of mass spectrometry for glycan and glycopeptide structural analysis. Anal. Chem. 2012, 84, 3040–3048. [Google Scholar] [CrossRef] [PubMed]
- Costello, C.E.; Contado-Miller, J.M.; Cipollo, J.F. A glycomics platform for the analysis of permethylated oligosaccharide alditols. J. Am. Soc. Mass Spectrom. 2007, 18, 1799–1812. [Google Scholar] [CrossRef] [PubMed]
- Mechref, Y.; Kang, P.; Novotny, M.V. Solid-phase permethylation for glycomic analysis. Methods Mol. Biol. 2009, 534, 53–64. [Google Scholar] [PubMed]
- Harvey, D.J. Fragmentation of negative ions from carbohydrates: part 3. fragmentation of hybrid and complex N-linked glycans. J. Am. Soc. Mass Spectrom. 2005, 16, 631–646. [Google Scholar] [CrossRef] [PubMed]
- Orlando, R. Quantitative glycomics. In Functional Glycomics; Springer: Berlin, Germany, 2010; pp. 31–49. [Google Scholar]
- Mechref, Y.; Hu, Y.; Desantos-Garcia, J.L.; Hussein, A.; Tang, H. Quantitative glycomics strategies. Mol. Cell. Proteom. 2013, 12, 874–884. [Google Scholar] [CrossRef] [PubMed]
- Moh, E.S.; Thaysen-Andersen, M.; Packer, N.H. Relative vs absolute quantitation in disease glycomics. Proteom. Clin. Appl. 2015, 9, 368–382. [Google Scholar] [CrossRef] [PubMed]
- Goldberg, D.; Sutton-Smith, M.; Paulson, J.; Dell, A. Automatic annotation of matrix-assisted laser desorption/ionization N-glycan spectra. Proteomics 2005, 5, 865–875. [Google Scholar] [CrossRef] [PubMed]
- GlycoWorkbench. Available online: http://www.eurocarbdb.org/applications/ms-tools (accessed on 15 August 2015).
- Ceroni, A.; Maass, K.; Geyer, H.; Geyer, R.; Dell, A.; Haslam, S.M. GlycoWorkbench: A tool for the computer-assisted annotation of mass spectra of glycans. J. Proteome Res. 2008, 7, 1650–1659. [Google Scholar] [CrossRef] [PubMed]
- GlycoMod. Available online: http://www.expasy.ch/tools/glycomod (accessed on 20 May 2015).
- Cooper, C.A.; Gasteiger, E.; Packer, N.H. GlycoMod—A software tool for determining glycosylation compositions from mass spectrometric data. Proteomics 2001, 1, 340–349. [Google Scholar] [CrossRef]
- UnicarbKB. Available online: http://www.unicarbkb.org/ (accessed on 4 April 2015).
- UnicarbDB. Available online: http://www.unicarb-db.org (accessed on 4 April2015).
- Campbell, M.P.; Peterson, R.; Mariethoz, J.; Gasteiger, E.; Akune, Y.; Aoki-Kinoshita, K.F.; Lisacek, F.; Packer, N.H. UniCarbKB: Building a knowledge platform for glycoproteomics. Nucleic Acids Res. 2014, 42, D215–D221. [Google Scholar] [CrossRef] [PubMed]
- Hayes, C.A.; Karlsson, N.G.; Struwe, W.B.; Lisacek, F.; Rudd, P.M.; Packer, N.H.; Campbell, M.P. UniCarb-DB:A database resource for glycomic discovery. Bioinformatics 2011, 27, 1343–1344. [Google Scholar] [CrossRef] [PubMed]
- Aoki-Kinoshita, K.; Agravat, S.; Aoki, N.P.; Arpinar, S.; Cummings, R.D.; Fujita, A.; Fujita, N.; Hart, G.M.; Haslam, S.M.; Kawasaki, T.; et al. GlyTouCan 1.0—The international glycan structure repository. Nucleic Acids Res. 2015. [Google Scholar] [CrossRef] [PubMed]
- Kam, R.K.; Poon, T.C. The Potentials of Glycomics in Biomarker Discovery. Clin. Proteom. 2008, 4, 67–79. [Google Scholar] [CrossRef]
- Zahradnikova, M.; Vojtesek, B.; Hernychova, L. Sugars Interfere or Glycomics in the Field of Cancer Biomarkers. Klin. Onkol. 2015, 28, S20–S25. [Google Scholar] [CrossRef]
- Cools-Lartigue, J.; Spicer, J.; Najmeh, S.; Ferri, L. Neutrophil extracellular traps in cancer progression. Cell. Mol. Life Sci. 2014, 71, 4179–4194. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sethi, M.K.; Fanayan, S. Mass Spectrometry-Based N-Glycomics of Colorectal Cancer. Int. J. Mol. Sci. 2015, 16, 29278-29304. https://doi.org/10.3390/ijms161226165
Sethi MK, Fanayan S. Mass Spectrometry-Based N-Glycomics of Colorectal Cancer. International Journal of Molecular Sciences. 2015; 16(12):29278-29304. https://doi.org/10.3390/ijms161226165
Chicago/Turabian StyleSethi, Manveen K., and Susan Fanayan. 2015. "Mass Spectrometry-Based N-Glycomics of Colorectal Cancer" International Journal of Molecular Sciences 16, no. 12: 29278-29304. https://doi.org/10.3390/ijms161226165
APA StyleSethi, M. K., & Fanayan, S. (2015). Mass Spectrometry-Based N-Glycomics of Colorectal Cancer. International Journal of Molecular Sciences, 16(12), 29278-29304. https://doi.org/10.3390/ijms161226165






