Endophytic Fungi from Lycium chinense Mill and Characterization of Two New Korean Records of Colletotrichum
Abstract
:1. Introduction
2. Results
| Isolate No. | GenBank Closest Hit (Accession Number) | Similarity (%) | Sequence Based Identification | Host Tissue | Accssion Number |
|---|---|---|---|---|---|
| CNU122031 | C. fructicola C1263.3 (JX010164) | 100 | C. fructicola | Fruit | KJ651254 |
| CNU122032 | C. brevisporum LC0600 (KC790943) | 99 | C. brevisporum | Leaf | KJ651255 |
| Glomerella magna L2.5 (DQ003103) | 99 | ||||
| CNU122033 | Acremonium sp. r116 (HQ649797) | 100 | Acremonium sp. | Fruit | KJ651256 |
| Acremonium strictum F21 (EU497953) | 99 | ||||
| CNU122034 | Cochliobolus lunatus Cs-1C (JN107740) | 99 | Cochliobolus lunatus | Leaf | KJ651257 |
| Cochliobolus sp. P2E4 (JN207244) | 99 | ||||
| CNU122035 | Fusarium cf. equiseti AM-48 (JN038489) | 99 | Fusarium equiseti | Leaf | KJ651258 |
| Fusarium equiseti ATT040 (HQ607811) | 99 | ||||
| CNU122036 | C. truncatum tc-1 (KC460308) | 99 | Colletotrichum sp. | Leaf | KJ651259 |
| Colletotrichum sp. ITCC 2041 (JN390888) | 99 | ||||
| CNU122037 | Hypocrea citrina GJS 91-61 (DQ000630) | 99 | Hypocrea citrina | Fruit | KJ651260 |
| CNU122038 | Nemania sp. NDJL-2009a (GU166482) | 99 | Nemania sp. | Leaf | KJ651261 |
| Nemania serpens BF330 (EF155504) | 95 | ||||
| CNU122039 | Nemania sp. AX48 (KC507255) | 100 | Nemania sp. | Leaf | KJ651262 |
| Nemania diffusa Z26 (JN198514) | 100 | ||||
| CNU122040 | Nemania sp. AX48 (KC507255) | 99 | Nemania sp. | Leaf | KJ651263 |
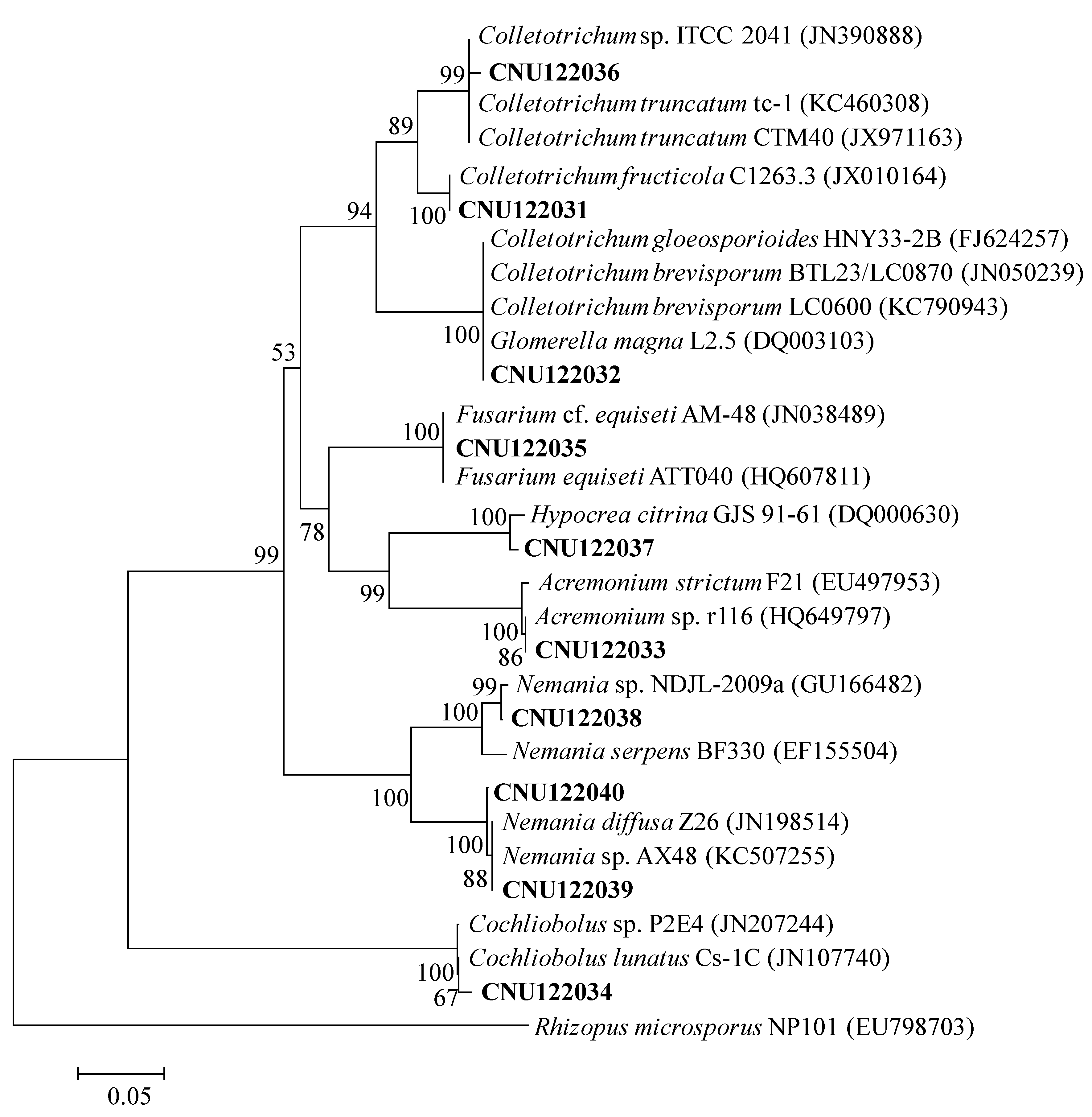
2.1. Taxonomy of Two Colletotrichum Species
2.1.1. Molecular Phylogeny
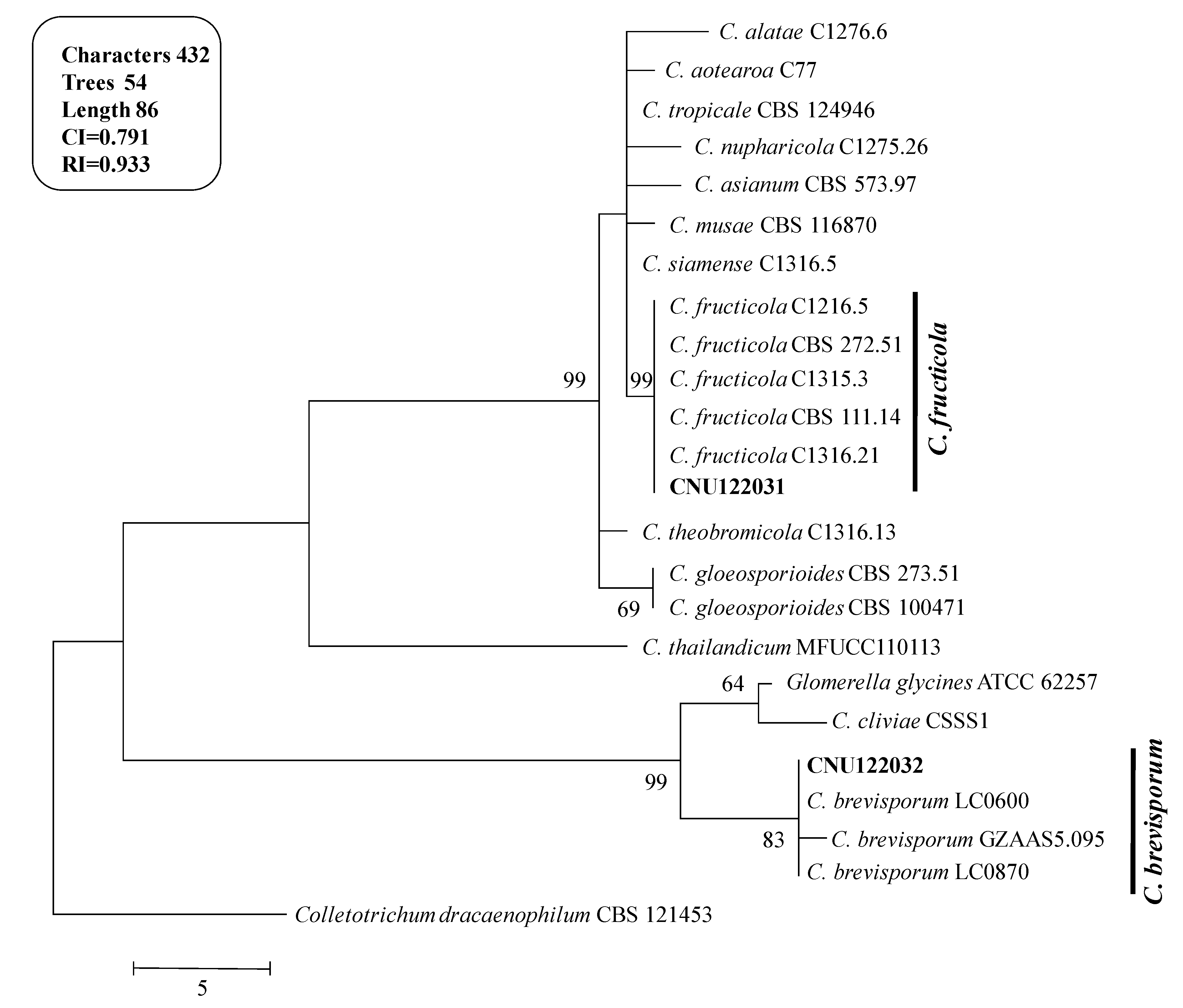
2.1.2. Morphological Characterization
2.1.3. CNU122031-Colletotrichum fructicola
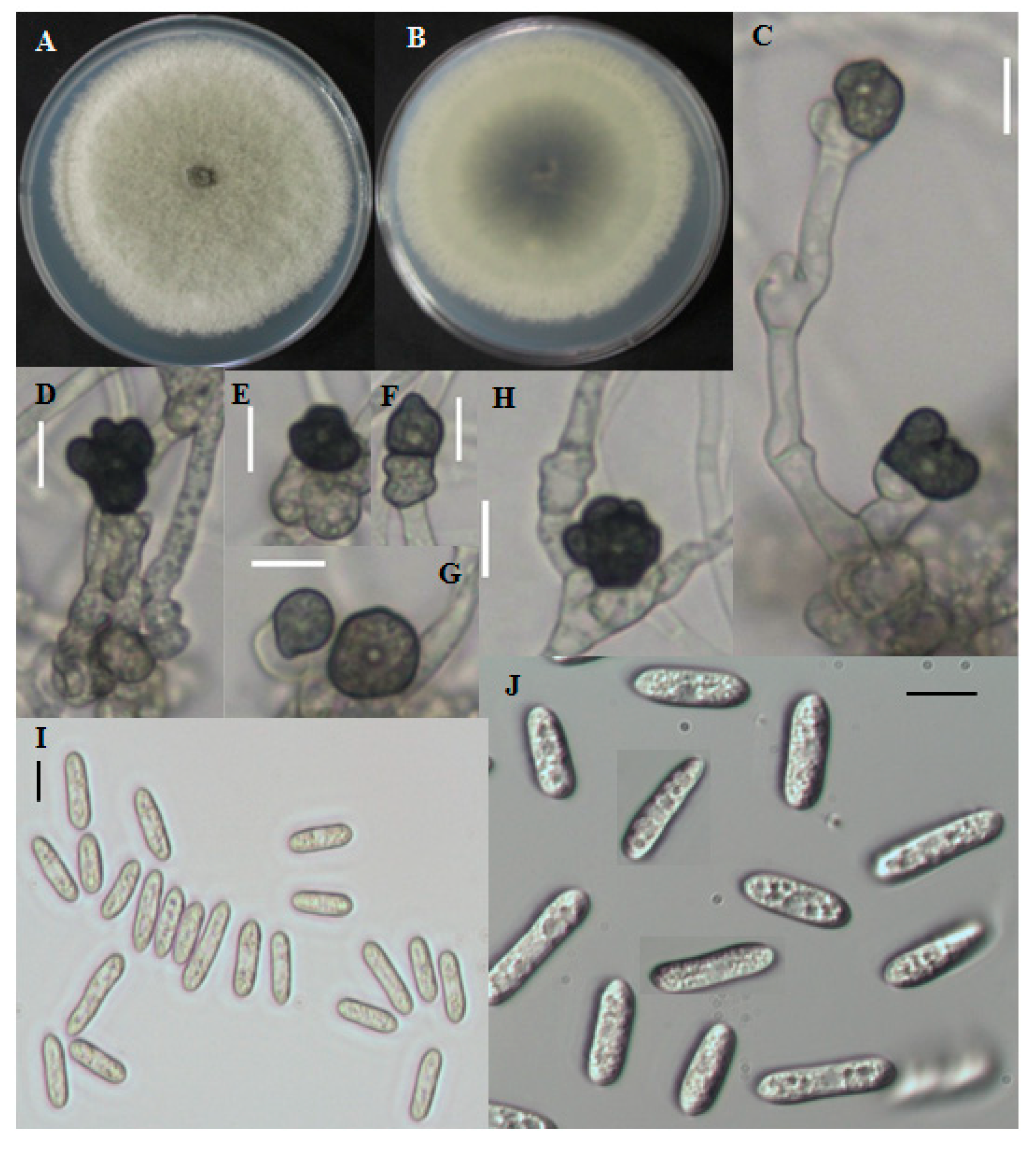
2.1.4. Distinguishing Characters
| Taxa | Colony | Conidia Shape and Size (μm) | Appresoria Size (μm) | Reference |
|---|---|---|---|---|
| C. brevisporum | White, mycelium in small tufts, reverse dark in middle | Cylindrical with round ends. 12.2–24.2 × 2.6–6 | 10–16.8 × 5–9.4 | This study |
| C. brevisporum | Aerial mycelia in small tufts, white, sparse with conidial masses, reverse dark green | Cylindrical with round ends, smooth-walled, hyaline. 12–17 × 5–6 | 10.5–14.5 × 8−11 | [13] |
| C. cliviae | White to gray, white at margin, reverse dark brown to greenish black | Cylindrical, straight or slightly curved, obtuse at the ends. 19.5–24.5 × 4.5–7 | 10.5–14.5 × 6−11 | [19] |
| C. fructicola | White, becoming gray to dark gray at the centre with age, dark circular margin at the center in reverse | Cylindrical with obtuse to rounded ends. 8.7–29.5 × 2.8–5.9 | 4.1–5.4 × 3−4.9 | This study |
| C. fructicola | White, becoming gray to dark gray at the centre with age, dark circular around the growing margin at the center in reverse | Cylindrical with obtuse to slightly rounded ends, sometimes oblong, hyaline. 9.7–14 × 3–4.3 | 4.7–8.3 × 3.5−5 | [18] |
| C. siamense | White, becoming pale brownish to pinkish, pale yellowish to pinkish colonies in reverse | Fusiform, sometimes with obtuse to slightly rounded ends, sometimes oblong, hyaline. 7–18.3 × 3–4.3 | 4.7–10.7 × 3.3–6.7 | [18] |
2.1.5. CNU122032-Colletotrichum bresvisporum
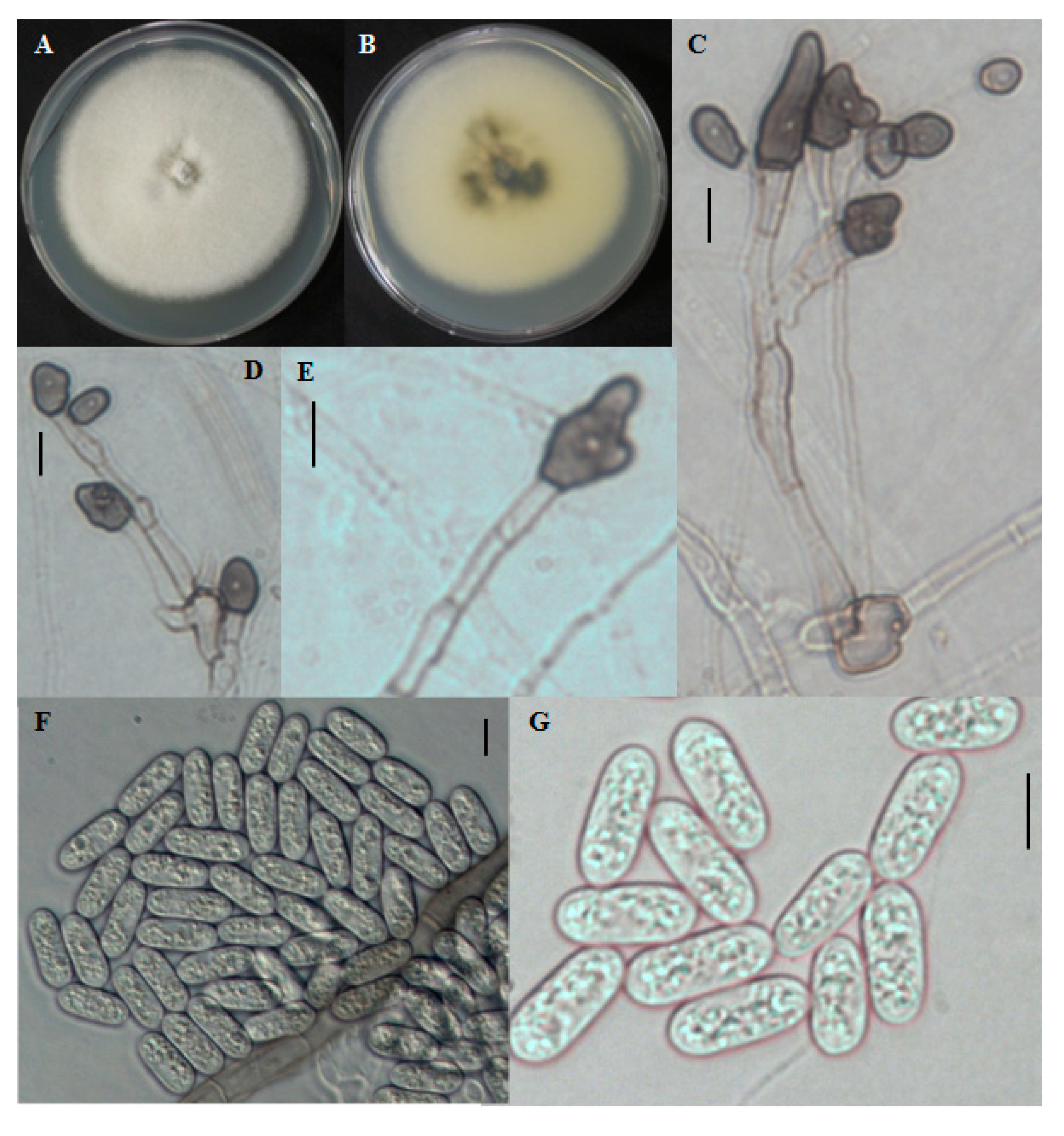
2.1.6. Distinguishing Characters
3. Discussion
4. Experimental Section
4.1. Sampling

4.2. Isolation of Endophytic Fungi
4.3. Genomic DNA Extraction, PCR and Sequencing
| Gene | Product Name | Primer | Direction | Sequence (5'–3') | Reference |
|---|---|---|---|---|---|
| ACT | Actin | ACT-512F | Forward | ATGTGCAAGGCCGGTTTCGC | [29] |
| ACT-783R | Reverse | TACGAGTCCTTCTGGCCCAT | [29] | ||
| GAPDH | Glyceraldehyde-3-phosphate dehydrogenase | GDF | Forward | GCCGTCAACGACCCCTTCATTGA | [30] |
| GDR | Reverse | GGGTGGAGTCGTACTTGAGCATGT | [30] | ||
| ITS | Internal transcribed spacer | ITS-1F | Forward | CTTGGTCATTTAGAGGAAGTAA | [28] |
| ITS-4 | Reverse | TCCTCCGCTTATTGATATGC | [28] |
4.4. Phylogenetic Analysis
| Species | Isolate | Host | Origin | Accession No. | ||
|---|---|---|---|---|---|---|
| ITS | gpd | ACT | ||||
| C. fructicola | CNU122031 | Lycium chinense | Korea | KJ651254 | KJ651266 | KJ651264 |
| C. brevisporum | CNU122032 | Lycium chinense | Korea | KJ651255 | KJ651267 | KJ651265 |
| C. alatae | C1276.6 | Discorea alata | India | JX010191 | JX010011 | JX009470 |
| C. aotearoa | C77 | Vitex lucens | NZ b | JX010221 | JX010023 | JX009500 |
| C. asianum | CBS 573.97 | Mangifera indica | Brazil | KC566732 | KC566586 | KC566878 |
| C. brevisporum | GZAAS5 | Citrus sp. | China | JQ247623 | JQ247599 | JQ247647 |
| C. brevisporum | LC0600 | Neoregalia sp. | Thailand | JN050238 | JN050227 | JN050216 |
| C. brevisporum | LC0870 | Pandanas pygmaeus | Thailand | JN050239 | JN050228 | JN050217 |
| C. cliviae | CSSS1 | Clivia miniata | China | GQ485607 | JX546611 | GU085861 |
| C. dracaenophilum | CBS 121453 | Dracaena sanderiana | Bulgaria | EU003533 | NA a | NA a |
| C. fructicola | C1316.21 | Theobroma cacao | Panama | JX010173 | JX009992 | JX009581 |
| C. fructicola | CBS 111.14 | NA a | Brazil | KC566785 | KC566639 | KC566931 |
| C. fructicola | CBS 272.51 | NA a | Brazil | KC566783 | KC566637 | KC566929 |
| C. fructicola | C1315.3 | Coffea arabica | Thailand | JX010165 | JX010033 | JX009501 |
| C. fructicola | C1216.5 | Persea americana | Australia | JX010166 | JX009946 | JX009529 |
| C. gloeosporioides | CBS 273.51 | Citrus limon | Italy | JX010148 | JX010054 | JX009558 |
| C. gloeosporioides | CBS 100471 | NA | Brazil | KC566719 | KC566573 | KC566865 |
| C. musae | CBS 116870 | Musa sp. | USA | JX010146 | JX010050 | JQ005840 |
| C. nupharicola | C1275.26 | Nuphar lutea | USA | JX010188 | JX010031 | JX009582 |
| C. siamense | C1316.5 | Hymenocallis americana | China | JX010278 | JX010019 | JX009441 |
| C. thailandicum | MFUCC1101 | Hibiscus rosa-sinensis | Thailand | JN050242 | JN050231 | JN050220 |
| C. theobromicola | C1316.13 | T. cacao | Panama | JX010294 | JX010006 | JX009444 |
| C. tropicale | CBS 124946 | T. cacao | Panama | KC566806 | KC566660 | KC566952 |
| Glomerella glycines | ATCC 62257 | Glycine max | USA | KC110794 | KC110812 | KC110830 |
4.5. Morphological Observation
5. Conclusions
Acknowledgments
Author Contributions
Conflict of Interest
References
- Fisher, P.J.; Petrini, O. Location of fungal endophytes in tissues of Suaeda fruiticosa: apreliminary study. Trans. Br. Mycol. Soc. 1987, 89, 246–249. [Google Scholar] [CrossRef]
- Spurr, H.W.; Welty, R.E. Characterization of endophytic fungi in healthy leaves of Nicotiana spp. Phytopathology 1975, 65, 417–422. [Google Scholar] [CrossRef]
- Fisher, P.J.; Petrini, O.; Lappin, S.H.M. The distribution of some fungal and bacterial endophytes in maize (Zea mays L.). New Phytol. 1992, 122, 299–305. [Google Scholar] [CrossRef]
- Larran, S.; Perello, A.; Simon, M.R.; Moreno, V. Isolation and analysis of endophytic microorganisms in wheat (Triticum aestivum L.) leaves. World J. Microbiol. Biotechnol. 2002, 18, 683–686. [Google Scholar]
- Paul, N.C.; Kim, W.K.; Woo, S.K.; Park, M.S.; Yu, S.H. Fungal endophytes in roots of Aralia species and their antifungal activity. Plant Pathol. J. 2007, 23, 287–294. [Google Scholar] [CrossRef]
- Chen, X.M.; Dong, H.L.; Hu, K.X.; Sun, Z.R.; Chen, J.; Guo, S.X. Diversity and antimicrobial and plant-growth-promoting activities of endophytic fungi in Dendrobium loddigesii Rolfe. J. Plant Growth Regul. 2010, 29, 328–337. [Google Scholar] [CrossRef]
- Margherita, D.; Salvatore, F.; Matteo, C. Endophytic fungi occurring in fennel, lettuce, chicory and celery-commercial crops in Southern Italy. Mycol. Res. 2008, 112, 100–107. [Google Scholar]
- Baltruschat, H.; Fodor, J.; Harrach, B.D.; Niemczyk, E.; Barna, B.; Gullner, G.; Janeczko, A.; Kogel, K.H.; Schafer, P.; Schwarczinger, I.; et al. Salt tolerance of barley induced by the root endophyte Piriformospora indica is associated with a strong increase in antioxidants. New Phytol. 2008, 180, 500–510. [Google Scholar]
- Tanaka, A.; Christensen, M.J.; Takemoto, D.; Park, P.; Scott, B. Reactive oxygen species play a role in regulating a fungus—Perennial ryegrass mutualistic interaction. Plant Cell 2006, 18, 1052–1066. [Google Scholar] [CrossRef]
- Tudzynski, B.; Sharon, A. Biosynthesis, biological role and application of fungal phyto-hormones. In Osiewacz of Pestalotiopsis isolated as endophytes from medicinal plants. Fungal Div. 2002, 24, 37–54. [Google Scholar]
- Potterat, O. Goji (Lycium barbarum and L. chinense): Phytochemistry, pharmacology and safety in the perspective of traditional uses and recent popularity. Planta Med. 2010, 76, 7–19. [Google Scholar] [CrossRef]
- Ballarin, S.M.; Lopez-Mataz, M.A.; Saenz, D.A.; Perez-Cinto, N.; Carnes, J. Anaphylaxis associated with the ingestion of Goji Berries (Lycium barbarum). J. Investig. Allergol. Clin. Immunol. 2011, 21, 567–570. [Google Scholar]
- Noireung, P.; Phoulivong, S.; Fang, L.; Cai, L.; Eric, H.C.M.; Ekachai, C.; Jones, E.B.G.; Ali, H.B.; Hyde, D.K. Novel species of Colletotrichum revealed by morphology and molecular analysis. Cryptogam. Mycol. 2012, 33, 347–362. [Google Scholar] [CrossRef]
- Cai, L.; Udayanga, D.; Manamgoda, D.S.; Maharachchikumbura, S.S.N.; Mckenzie, E.H.C.; Guo, L.D.; Liu, X.X.; Bahkali, A.H.; Hyde, K.D. The need to carry out re-inventory of plant pathogens. Trop. Plant Pathol. 2011, 36, 205–213. [Google Scholar]
- Hyde, K.D.; Cai, L.; Mckenzie, E.H.C.; Yang, Y.L.; Zhang, J.Z.; Prihastuti, H. Colletotrichum: A catalogue of confusion. Fungal Div. 2009, 39, 1–17. [Google Scholar]
- Phoulivong, S.; Cai, L.; Chen, H.; Mckenzie, E.H.C.; Abd-Elsalam, K.; Chukeatirote, E.; Hyde, K.D. Colletotrichum gloeosporioides is not a common pathogen on tropical fruits. Fungal Div. 2010, 44, 33–43. [Google Scholar] [CrossRef]
- Cai, L.; Hyde, K.D.; Taylor, P.W.J.; Weir, B.; Waller, J.; Abang, M.M.; Zhang, J.Z.; Yang, Y.L.; Phoulivong, S.; Liu, Z.Y.; et al. A polyphasic approach for studying Colletotrichum. Fung. Div. 2009, 39, 183–204. [Google Scholar]
- Prihastuti, H.; Cai, L.; Chen, H.; Mckenzie, E.H.C.; Hude, K.D. Characterization of Colletotrhichum species with coffee barriers in northern Thailand. Fungal Div. 2009, 39, 89–109. [Google Scholar]
- Yang, Y.L.; Liu, Z.Y.; Cai, L.; Hyde, K.D.; Yu, Z.N.; Mckenzie, E.H.C. Colletotrichum anthracnose of Amaryllidaceae. Fungal Div. 2009, 39, 123–149. [Google Scholar]
- Rubini, M.R.; Silva-Ribeiro, R.T.; Pomella, A.W.V.; Maki, C.S.; Araujo, W.L.; Santos, D.R.; Azevedo, J.L. Diversity of endophytic fungal community of cacao (Theobroma cacao L.) and biological control of Crinipellis perniciosa, causal agent of witches broom disease. Int. J. Biol. Sci. 2005, 1, 24–33. [Google Scholar]
- Rosa, L.H.; Vieira, M.L.A.; Santiago, I.F.; Rosa, C.A. Endophytic fungi community associated with the dicotyledonous plant Colobanthus quitensis (Kunth) Bartl. (Caryophyllaceae) in Antarctica. FEMS Microbiol. Ecol. 2010, 73, 178–189. [Google Scholar]
- Flor, N.R.; Roberto, A.S.; Zolia, N.G.; Luis, B.F. Diversity of endophytic fungi of Taxus globosa (Mexican yew). Fungal Div. 2010, 47, 65–74. [Google Scholar]
- Huang, W.Y.; Cai, Y.Z.; Surveswaran, S.; Hyde, K.D.; Corke, H.; Sun, M. Molecular phylogenetic identification of endophytic fungi isolated from three Artemisia species. Fungal Div. 2009, 36, 69–88. [Google Scholar]
- Kumar, S.; Kaushik, N.; Ebel, R.E.; Ebel, R.; Proksch, P. Isolation, characterization, and bioactivity of endophytic fungi of Tylophora indica. World J. Microbiol. Biotechnol. 2011, 27, 571–577. [Google Scholar] [CrossRef]
- Sanchez, M.S.; Bills, G.F.; Zabalgogeazcoa, I. Diversity and structure of the fungal endophytic assemblages from two sympatric coastal grasses. Fungal Div. 2008, 33, 87–100. [Google Scholar]
- Sette, L.D.; Passarini, M.R.Z.; Delarmelina, C.; Salati, F.; Duarte, M.C.T. Molecular characterization and antimicrobial activity of endophytic fungi from coffee plants. World J. Microbiol. Biotechnol. 2006, 22, 1185–1195. [Google Scholar] [CrossRef]
- Park, M.S.; Seo, G.S.; Bae, K.S.; Yu, S.H. Characterization of Trichoderma spp. Associated with green mold oyster mushroom by PCR-RFLP and sequence analysis of ITS regions of rDNA. Plant Pathol. J. 2005, 21, 229–236. [Google Scholar] [CrossRef]
- White, T.J.; Bruns, T.D.; Lee, S.B.; Taylor, J.W. PCR Protocols: A Guide to the Methods and Applications; Innes, M.A., Gelfand, D.H., Sninsky, J.J., White, T.J., Eds.; Academic Press: New York, NY, USA, 1990; p. 315. [Google Scholar]
- Carbone, I.; Kohn, L.M. A method for designing primer sets for speciation studies in filamentous ascomycetes. Mycologia 1999, 91, 553–556. [Google Scholar] [CrossRef]
- Templeton, M.D.; Rikkerink, E.H.; Solon, S.L.; Crowhurst, R.N. Cloning and molecular characterization of the glyceraldehyde-3-phosphate dehydrogenase-encoding gene and cDNA from the plant pathogenic fungus Glomerella cingulata. Gene 1992, 122, 225–230. [Google Scholar] [CrossRef]
- Thompson, J.D.; Gibson, T.J.; Plewniak, F.; Jeanmougin, F.; Higgins, D.G. ClustalX: Windows interface: Flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 1997, 25, 487–4878. [Google Scholar]
- Chun, J. Computer-Assisted Classification and Identification of Actinomycetes. Ph.D. Thesis, University of Newcastle, New Castle Upon Tyne, UK, July 1995. [Google Scholar]
- Kimura, M. A simple method for estimating evolutionary rate of base substitution through comparative studies of nucleotide sequence. J. Mol. Evol. 1980, 16, 111–120. [Google Scholar] [CrossRef]
- Felsenstein, J. Confidence limits of phylogenies: An approach using the bootstrap. Evolution 1985, 39, 783–791. [Google Scholar] [CrossRef]
- Johnston, P.R.; Jones, D. Relationship among Colletotrichum isolates from fruit-rots assessed using rDNA sequences. Mycologia 1997, 89, 420–430. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Paul, N.C.; Lee, H.B.; Lee, J.H.; Shin, K.S.; Ryu, T.H.; Kwon, H.R.; Kim, Y.K.; Youn, Y.N.; Yu, S.H. Endophytic Fungi from Lycium chinense Mill and Characterization of Two New Korean Records of Colletotrichum. Int. J. Mol. Sci. 2014, 15, 15272-15286. https://doi.org/10.3390/ijms150915272
Paul NC, Lee HB, Lee JH, Shin KS, Ryu TH, Kwon HR, Kim YK, Youn YN, Yu SH. Endophytic Fungi from Lycium chinense Mill and Characterization of Two New Korean Records of Colletotrichum. International Journal of Molecular Sciences. 2014; 15(9):15272-15286. https://doi.org/10.3390/ijms150915272
Chicago/Turabian StylePaul, Narayan Chandra, Hyang Burm Lee, Ji Hye Lee, Kyu Seop Shin, Tae Hee Ryu, Hye Ri Kwon, Yeong Kuk Kim, Young Nam Youn, and Seung Hun Yu. 2014. "Endophytic Fungi from Lycium chinense Mill and Characterization of Two New Korean Records of Colletotrichum" International Journal of Molecular Sciences 15, no. 9: 15272-15286. https://doi.org/10.3390/ijms150915272
APA StylePaul, N. C., Lee, H. B., Lee, J. H., Shin, K. S., Ryu, T. H., Kwon, H. R., Kim, Y. K., Youn, Y. N., & Yu, S. H. (2014). Endophytic Fungi from Lycium chinense Mill and Characterization of Two New Korean Records of Colletotrichum. International Journal of Molecular Sciences, 15(9), 15272-15286. https://doi.org/10.3390/ijms150915272




