Abstract
The air that we breathe contains nearly 21% oxygen, most of which is utilized by mitochondria during respiration. While we cannot live without it, it was perceived as a bane to aerobic organisms due to the generation of reactive oxygen and nitrogen metabolites by mitochondria and other cellular compartments. However, this dogma was challenged when these species were demonstrated to modulate cellular responses through altering signaling pathways. In fact, since this discovery of a dichotomous role of reactive species in immune function and signal transduction, research in this field grew at an exponential pace and the pursuit for mechanisms involved began. Due to a significant number of review articles present on the reactive species mediated cell death, we have focused on emerging novel pathways such as autophagy, signaling and maintenance of the mitochondrial network. Despite its role in several processes, increased reactive species generation has been associated with the origin and pathogenesis of a plethora of diseases. While it is tempting to speculate that anti-oxidant therapy would protect against these disorders, growing evidence suggests that this may not be true. This further supports our belief that these reactive species play a fundamental role in maintenance of cellular and tissue homeostasis.
1. Introduction
Mitochondria were first identified over a century ago and were initially termed as “bioblasts” by Richard Altmann who described them as “elementary organisms” living inside cells [1]. In fact this theory of endosymbiosis that mitochondria are the direct descendants of a bacterial endosymbiont is still one of the most widely accepted theories of mitochondrial evolution [2]. The term mitochondria, was later coined by Carl Benda and literally means “mitos-thread” and “chondrion-granule” [1]. The role of mitochondria in the cell was initially presumed to be only to generate energy in the form of adenosine triphosphate (ATP) and is still referred to as the “powerhouse of the cell”. However, research in the past few decades has provided compelling evidence to suggest that mitochondria are actively involved in a multitude of cellular activities including, signaling, proliferation and death. In fact, while most eukaryotic cells contain mitochondria, the size, number and location of mitochondria in a cell vary significantly based on the cellular needs. For instance, in neuronal cells, mitochondria accumulate predominantly at high energy demanding sites such as presynaptic terminals, nodes of Ranvier and active growth cones and branches [3]. Given the role of mitochondria in a variety of cellular processes, it is not surprising that damage to the mitochondria has been implicated in the pathogenesis of end-organ injury in a variety of diseases [4–23].
How does a single organelle control the fate of the cell? This question has captivated scientists and studies over the past few decades have revealed fascinating mechanisms. An extensively studied and established mechanism relies on the generation of free radical species that determine the outcome of the cellular processes involved. This review will mainly focus on the role of these reactive oxygen/nitrogen species in the modulation of cellular activities. We will summarize some of the major pathways and molecular mechanisms that are regulated by reactive oxygen species (ROS) and reactive nitrogen species (RNS) and given the wealth of knowledge that exists for these species, this review will highlight recent developments in ROS/RNS mediated signaling pathways and their role in the regulation of cellular processes, such as autophagy, mitochondria fusion and fission.
2. Mitochondria Structure and Function
Mitochondria are unique organelles as its structure provides compartmentalization of metabolism (Figure 1A). They are very complex organelles that contain two phospholipid bilayers, by virtue of which they can be categorized into 4 different segments: the outer membrane, inter-membrane space, inner membrane and matrix [24–26].
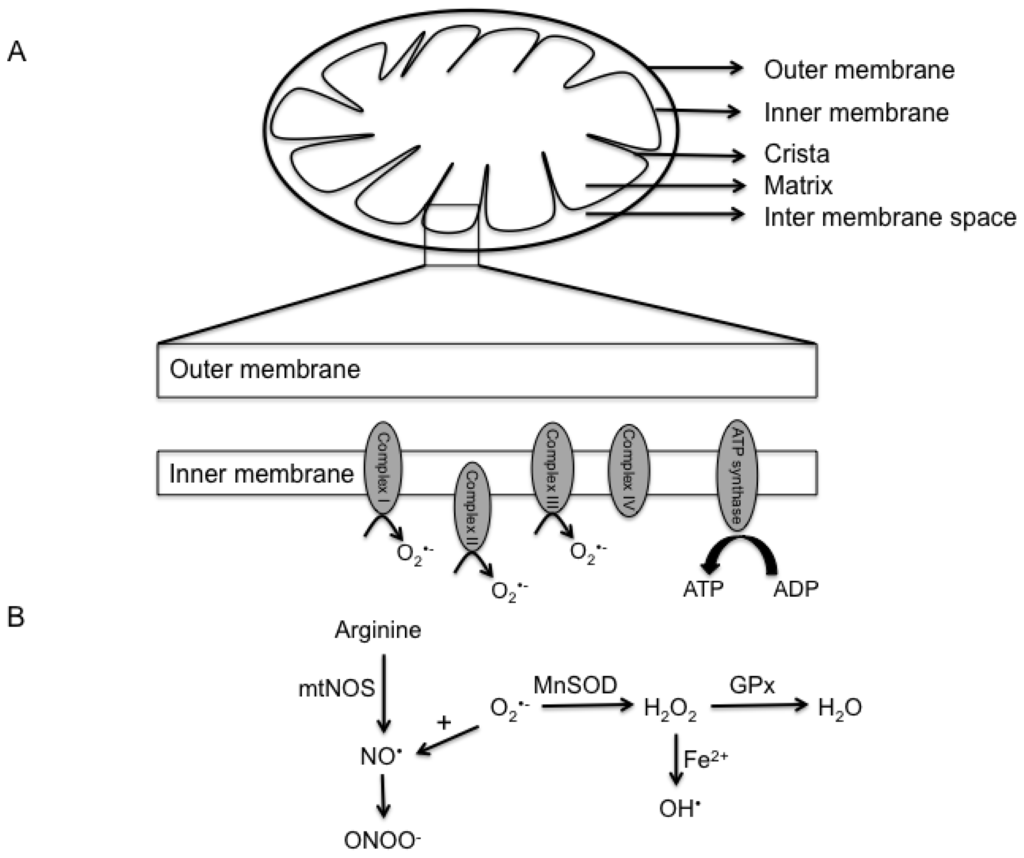
Figure 1.
Mitochondria structure and generation of mitochondrial reactive species. (A) A schematic of a typical mitochondrion is represented. By virtue of its lipid bilayers, the mitochondrion can be subdivided into the outer membrane, inter-membrane space, inner membrane and matrix. The lower panel demonstrates the generation of superoxide anion through the different complexes of the electron transport chain; (B) Amplification of the free radical cycle. Superoxide generated during the electron transport chain can react with nitric oxide to form peroxy nitrite species. Alternatively, superoxide is converted by manganese superoxide dismutase to hydrogen peroxide, which is subsequently converted to water by glutathione peroxidase. In the presence of iron, hydrogen peroxide is rapidly converted to the highly reactive hydroxyl ion.
The outer membrane of the organelle is identical to the plasma membrane in its content (equal ratio of protein to phospholipid content by weight). It contains porins that allow molecules that are less than 5 KDa to freely diffuse through. However, larger proteins require the presence of a mitochondria targeted sequence that will enable binding to specific transporters (translocase of the outer membrane—TOM and inner membrane—TIM) on the membrane for entry into the organelle [26–29]. The outer membrane therefore mainly serves as a permeability barrier to the cytosolic components. Until recently, it was presumed that the inter membrane space had no specific function and was identical to the cytosol in its contents. However, emerging studies have suggested an important role for this space in maintaining mitochondrial homeostasis, including protein sorting and lipid homeostasis (reviewed in [30]). The inner membrane of the mitochondria is perhaps the single most extensively studied cell membrane component due to its relative importance in oxidative phosphorylation. This membrane comprises of the highest number of proteins per phospholipid moiety in a cell. These proteins are integral to the electron transport chain, ATP synthesis and transport [31,32]. The inner membrane is also distinct from other membranes by the presence of cristae (invaginations of the membrane), which allow for compartmentalization and increases the surface area. The inner membrane is also less permeable to ions and molecules and helps in compartmentalization through separation of the mitochondrial matrix from the cytosolic environment, thereby acting as an electrical insulator and chemical barrier [32]. This helps in maintenance of the electron gradient across the membrane, which enables generation of ATP. The mitochondrial matrix of mammalian cells contains the mitochondrial DNA (16.5 kilobase genome) that encodes for nearly 13 proteins, some of which are involved in oxidative phosphorylation. The remaining proteins required for the normal function of the mitochondria are encoded by the nuclear genome and imported into the mitochondria [33]. The matrix also contains a majority of the enzymes required for the citric acid cycle, which oxidizes acetyl coenzyme A and in the process generates energy in the form of nicotinamide adenine dinucleotide (NADH) and flavin adenine dinucleotide (FADH2). These molecules then serve as substrates for oxidative phosphorylation by the proteins in the inner membrane to generate cellular energy in the form of ATP.
3. Reactive Oxygen and Nitrogen Species
Halliwell and colleagues described free radicals as “any species capable of independent existence that contains one or more unpaired electrons” [34]. The term ROS simply refers to a variety of reactive molecules that are derived from oxygen and can be free radicals (superoxide (O2•−) or hydroxyl radical (OH•)) or non-radicals (hydrogen peroxide (H2O2)). Similarly, they can be further classified as ions (O2•−) or non-ions (H2O2). On the other hand, RNS refers to reactive species derived from nitrogen and can be broadly classified as ions (peroxynitrite (ONOO−)) or non-ions (Nitric Oxide (NO•)). These reactive species are formed at low levels during the execution of physiological functions of the cell. In fact, the electron transport chain is responsible for most of the superoxide that is generated through partial reduction of oxygen (Figure 1A) and is reviewed in detail in the following sections. Furthermore, they are formed at different rates in a cell and differ in their activity. In terms of activity, hydroxyl radical is the most reactive species known and is by in large responsible for the cytotoxic effects of ROS. In contrast, reactive species such as nitric oxide and hydrogen peroxide are less reactive and have shown to play an important role in several cellular activities. This is discussed in detail in the following sections.
Other enzymatic systems responsible for the generation of these reactive species include but are not limited to respiratory burst enzymes (such as NADPH oxidases –Nox1-5), amino acid oxidases, cytochrome P450 enzymes, cycloxygenases, lipoxygenases, xanthine oxidase [35–41]. Addressing each of these systems is beyond the scope of this review and hence we will focus specifically on mitochondria associated reactive species.
As discussed above, mitochondria are one of the most active organelles in the cell, consuming nearly 90% of the total oxygen content in the cell to enable oxidative phosphorylation and ATP synthesis [42]. Given that low levels of superoxide are constantly generated during normal respiration by healthy mitochondria, several pathways evolved to detoxify this anion. For example, manganese superoxide dismutase (MnSOD) is a mitochondrial matrix enzyme that rapidly converts superoxide to hydrogen peroxide, another reactive species. This molecule can then be converted to water by catalase or glutathione peroxidase in the mitochondria or following diffusion into the cytosol. In addition to these enzymes, cells are equipped with a variety of antioxidant molecules, such as glutathione, ascorbic acid and α-tocopherol, which are capable or reducing ROS. Gluthathione contains a sulphahydryl group that is oxidized by ROS. Therefore, glutathione protects the vital mitochondrial components from being targeted by ROS by serving as a substrate. However, damaged and dysregulated mitochondria generate excessive amounts of superoxide which can damage several mitochondrial components, including proteins, lipids and DNA. These reactions then lead to a vicious cycle of further generation of reactive species and ultimately cell death. While hydrogen peroxide is a reactive molecule, in the presence of transition metals, such as iron, it can be converted to hydroxyl ion via the Fenton reaction. This iron is thought to be released through destabilization of ferritin and other iron containing proteins by superoxide anion [43–46]. Alternatively, superoxide may react with nitric oxide to generate peroxynitrite species. The details of these reactions are presented in Figure 1B.
Over the past few decades, research on these reactive species has soared in both physiology and pathology. With the advent of novel technology, researchers have now been able to shed light on the source of reactive species generation in the different sub-compartments of the mitochondria which has been extensively reviewed by Lenoz [47]. To summarize, it was demonstrated that complex I, also referred to as NADH CoQ reductase, catalyzes the transfer of electrons from NADH to coenzyme Q, which is accompanied by translocation of protons from the matrix to the intermembrane space. There is now evidence to suggest that complex I is involved in ROS production, specifically superoxide [48–50]. Similarly, succinate dehydrogenase, complex II enzyme is responsible for the reduction of CoQ and has also shown to be involved in generating low levels of superoxide anion [48,51]. Complex III (ubiquinol cytochrome c reductase), on the other hand has shown to be responsible for the superoxide generation in the intermembrane space. Superoxide generation by this complex is significantly enhanced when the electron transfer is reduced, either due to inhibition in respiration (actinamycin A) or an increase in membrane potential [52,53]. Interestingly, the contribution of each of these enzymes to ROS production is different in different tissues and during disease conditions. For instance, while complex III has been implicated as the major source of superoxide in the heart, complex I seem to be of prime importance in the brain [54–57]. Additionally, enzymes such as Glycerol-3-phosphate dehydrogenase, Monoamine oxidase, Dihydrolipoamide dehydrogenase and Electron-transferring-flavoprotein dehydrogenase have also been implicated in ROS production [58–63].
As evident from the reactions described above, most of the superoxide generated is either in the matrix or on the inner membrane of the mitochondria that faces the matrix. While most of the reactive species generated within the mitochondria is superoxide anion, MnSOD rapidly converts it to hydrogen peroxide. Although hydrogen peroxide is more stable than superoxide, it can freely diffuse out of the mitochondria into the cytosol, thereby reducing the harmful effects of these reactive species to the mitochondria. It has also been suggested that in the presence of excessive superoxide, MnSOD is oxidized and this further compromises the antioxidant capacity of the mitochondria and enhances oxidative stress [64]. Alternatively, superoxide may be carried to the cytoplasm by voltage-dependent anion channels [65].
Nitric oxide (NO•) is another reactive species that is generated by the mitochondria and research on this molecule has gained momentum over the past few decades. Nitric oxide is generated during the breakdown of arginine to citrulline by a family of NADPH-dependent enzymes called nitric oxide synthases (NOS). The importance of this enzyme in physiology is underscored by the fact there are several isoforms of NOS, including an endothelial constitutive isoform (NOS3), an inducible isoform that is expressed in several cell types (NOS2) in response to pro-inflammatory stimuli and produces large amounts of NO• and a neuronal isoform (NOS1). Recently, a mitochondrial isoform of NOS referred to as mtNOS has been described [66]. This isoform was found to be responsive to changes in calcium concentration in the matrix and to play an important role in modulating mitochondrial respiration. Following this initial study, several groups have identified the presence of mtNOS in the mitochondria of cells from different tissues including liver, brain and kidney [67–72]. However, and in spite of these studies there is still debate on the existence of this isoform and indeed some studies suggest that the presence of NO• in the mitochondria may reflect pathways independent of NOS activity that may include nitrite reductase activity and the electron transport chain [65,72–75]. Once generated, NO• can inhibit respiration by binding to heme groups in the proteins of the electron transport chain, including cytochrome c oxidase [76–81]. Irrespective of the source of NO•, its presence in the mitochondria can alter the activity of a number of processes including respiration, mitochondrial biogenesis and oxidative stress through increased production of reactive oxygen and nitrogen species and thereby impact cell physiology [82–94].
As described above, mitochondria are a major source of reactive oxygen and nitrogen species. Interestingly, these species in turn regulate the mitochondrial activity through several mechanisms, including mitochondrial biogenesis, mtDNA damage, lipid peroxidation and mitochondrial membrane permeability transition (reviewed in [83,86,89–98]). The physiological and pathological role of these reactive species in cellular activities will be discussed below.
4. Autophagy
Autophagy (meaning self-eating) is an evolutionarily conserved catabolic process that involves an intracellular degradation system in which cytoplasmic components, such as organelles, protein aggregates and other macromolecules are directed to the lysosome through a physiologically regulated process to maintain cell homeostasis. While three different types of autophagy (micro-, macro- and chaperone-mediated autophagy) have been described, the most studied process in mammalian cells is macro-autophagy. As the name suggests, it involves degradation of large moieties such as organelles and protein aggregates through a tightly regulated process. This process begins with the isolation and sequestration of the cytoplasmic components by a double-layered lipid membrane that forms an autophagosome. This vesicle then fuses with a lysosome to form an autolysosomes where the sequestered cellular components are degraded by the lysosomal enzymes (Figure 2A) [99–102]. Although this process may appear to be self-destructive, it is an extremely efficient recycling strategy that is vital to maintain cell homeostasis and generates amino acids, fatty acids and energy (ATP) that are used for macromolecular synthesis. There have been at least 35 Atg (Autophagy) related genes identified in yeast whereas their mammalian orthologs have not been completely characterized [103,104].
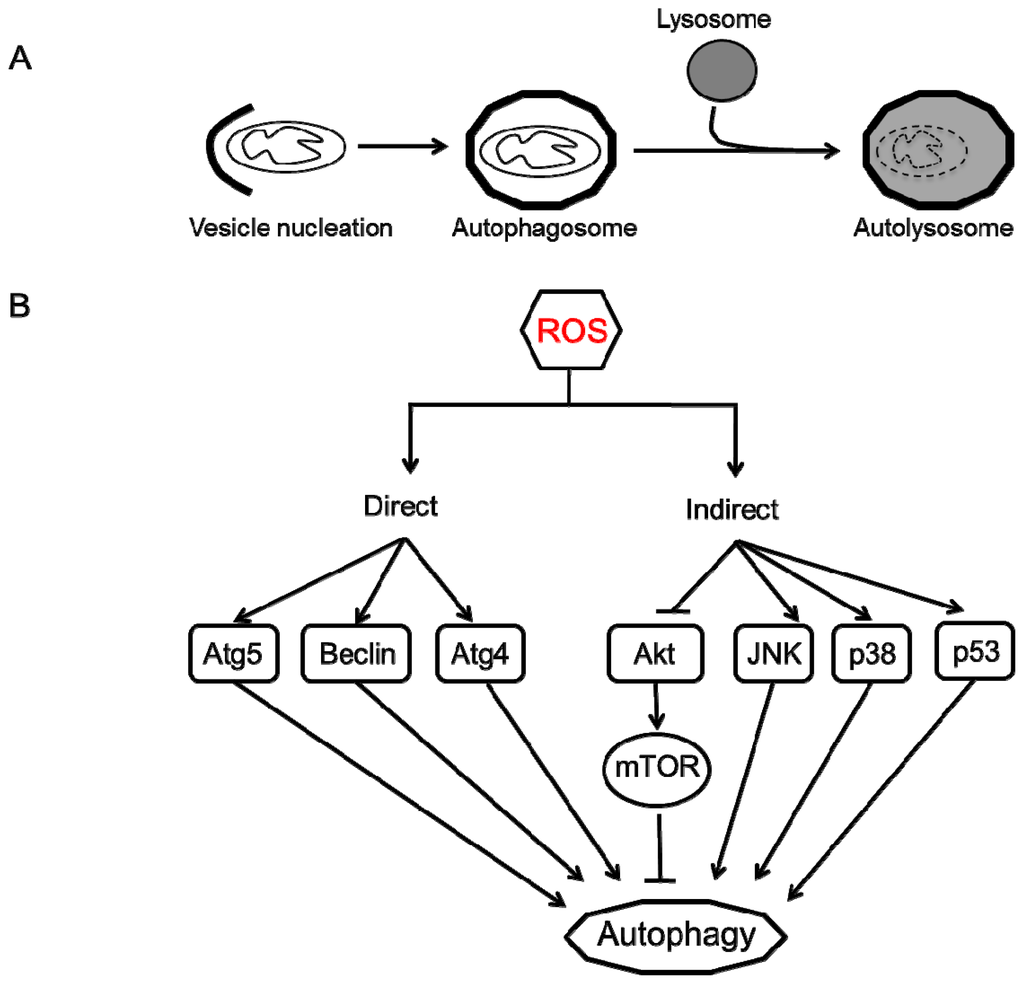
Figure 2.
Autophagy process and regulation by ROS. (A) Autophagy begins with vesicle nucleation where the damaged organelles (mitochondria) are sequestered to form an autophagosome. This vesicle fuses with the lysosome to form an autolysosome where the contents are degraded by lysosomal hydrolases and nutrients are recycled to the cytoplasm; (B) ROS can regulate autophagy in two ways: direct and indirect. Direct regulation involves modification of key proteins involved in the autophagy process including Atg4, Atg5 and Beclin. Indirect regulation by ROS involves alteration of signaling pathways such as JNK, p38 that can induce autophagy. On the other hand, ROS may inhibit Akt signaling and downstream mTOR and thereby induce autophagy.
Autophagy has been associated with a number of diseases, including but not limited to cancer (breast, ovarian, prostate and colon), neurodegenerative diseases (Alzheimer’s, Parkinson’s and Huntington’s disease), myodegenerative diseases (muscular dystrophy, X-linked myopathy), Crohn’s disease, diabetes and several other inherited diseases such as Danon disease, Pompe disease. However, the role of autophagy in protection or disease progression has remained a conundrum [101,105–116]. Nevertheless, research in the past decade has unraveled the autophagy process and provided compelling evidence to suggest a protective role for regulated and controlled autophagy [106,107]. Nutrient-starvation ROS generation is one of the key modulators of autophagy in normal and cancerous cells. In a comprehensive study, Chen and colleagues provide irrefutable evidence to suggest that mitochondrial ROS are important mediators of autophagy and autophagic cell death in transformed cells and cancer cells [117]. They demonstrate that overexpression of SOD or use of ROS scavengers are capable of lowering autophagy and cell death in the presence of electron transport chain inhibitors, rotenone and thenoyltrifluoroacetone [117]. Mitochondrial ROS can regulate autophagy in two major pathways: by direct modification of the autophagy proteins or by altering the proteins that are indirectly involved in the autophagy process.
ROS, specifically, hydrogen peroxide that is generated during starvation, modulate the activity of Atg4, an essential cysteine protease in the autophagic pathway, through a series of redox reactions. Atg4 cleaves the c-terminus of Atg8, enabling the addition of phosphatidylethanolamine (PE) to Atg8 and subsequent conjugation of this protein on the autophagosomal membrane, leading to autophagosome maturation. However, Atg8-PE also serves as a substrate for Atg4, which allows for efficient recycling of Atg8. This protease is therefore tightly regulated and research now points to hydrogen peroxide as a mediator of this redox signaling. Hydrogen peroxide oxidizes Atg4 following the initial cleavage, thereby allowing the autophagosome completion. Once the lysosome fuses with this vesicle, Atg4 is re-activated and recycles Atg8 for another cycle of autophagy. This is the first description of ROS as a signaling molecule that triggers autophagy as a cell survival mechanism [107]. While the mechanism is not clearly understood, several studies have demonstrated an upregulation of beclin 1, a protein involved in autophagy initiation, in the presence of ROS [118–121]. However, it is still unknown whether this cysteine rich protein is also modulated by redox activity.
With regard to ROS-mediated indirect regulation of autophagy, hydrogen peroxide and superoxide anion can modulate the activity of a number of signaling pathways that induce autophagy. Using glioma cells as a model to unravel autophagic mechanism in cancerous cells, several groups have identified key regulatory effectors of ROS that play a major role in the autophagy pathway. One such effector is the mammalian target of rapamycin (mTOR) that is actively involved in a variety of cellular processes including transcription, proliferation, motility and survival. Hydrogen peroxide can disrupt the mitochondrial membrane potential, leading to an inhibition in Akt/mTOR signaling pathway that is capable of inducing autophagy [122,123]. Similarly, several studies have demonstrated the role of ROS in regulation of MAPK (mitogen activated protein kinase) pathways. Increased ROS (hydrogen peroxide and nitric oxide) levels in cardiomyocytes or skeletal muscle, induces autophagy that is dependent on p38 signaling [115,124]. Additionally, using a number of different tumor cell lines, Wong and colleagues demonstrated that ROS and downstream activation of ERK and JNK pathways were responsible for autophagy induction [125]. In another study, ROS mediated glycogen synthase kinase-3 activity was responsible for cadmium induced autophagy in mesangial cells [126]. Of note, even non-mitochondria associated NADPH oxidase–generated ROS can induce autophagy, implying that irrespective of the source, ROS can act as signaling molecules. This phenomenon has been observed in several immune cells (macrophages and neutrophils) that induce ROS-mediated autophagy to enable destruction of phagocytosed microbes [127–130].
While the majority of the research discussed above demonstrates a role for ROS in inducing autophagy, there is strong evidence to suggest that autophagy may in turn regulate mitochondrial network by eliminating damaged mitochondria. Kissova and colleagues were the first to demonstrate in yeast that an outer mitochondrial protein, Uth1p was responsible for the early selective degradation of mitochondria by autophagy during stress induced by nutrient deprivation or rapamycin [131]. This phenomenon gave rise to “mitophagy”, a term coined by Leimaster to describe a sub-type of macroautophagy where damaged mitochondria were sequestered by autophagosomes and removed to maintain mitochondrial homeostasis [132]. Since this discovery, the molecular mechanisms regulating mitophagy in yeast have been identified. However, in the mammalian system, research has been mainly focused on parkin/pink1 (PTEN-induced putative kinase protein 1) mechanism. PINK1 is a mitochondrial kinase that may serve as a guardian to the mitochondria through its ability to identify depolarized mitochondria. Under normal conditions, PINK1 undergoes voltage-dependent lysis and is removed from the mitochondria. However, in damaged mitochondria that have low membrane potential, PINK1 accumulates on the mitochondrial surface and recruits parkin [133,134]. Parkin ubiquitinates mitochondrial proteins, including voltage-dependent anion channel 1 (VDAC1), and recruits autophagic machinery to the damaged mitochondria for removal [133,135–138]. Recently, Egan and colleagues demonstrated that mammalian ortholog of Atg1, ULK-1 is required for mitophagy. In the absence of this protein, there was a decrease in starvation-induced mitophagy and an associated increase in the number of aberrant mitochondria in mouse embryonic fibroblasts and hepatocytes, suggesting that ULK-1 is a key regulator of mitochondrial homeostasis [139].
In summary, these studies suggest that autophagy is a complex process that is regulated by ROS in multiple ways (Figure 2B). Autophagy is a double-edged sword, with both cytoprotective and cytotoxic capabilities. Whether it inhibits cell death by removing damaged organelles (mitochondria) or induces cell death depends on many factors, including ROS. Mild ROS induces autophagy (or mitophagy) as a survival mechanism to eliminate damaged mitochondria that are responsible for uncontrolled ROS generation. On the contrary, high levels of ROS or sustained exposure to ROS can alter signaling pathways that converge to induce autophagic or apoptotic cell death [116,140].
5. Mitochondria Network
Mitochondria are complex organelles that exist as a tubular network in cells. Two opposing pathways that must exist in equilibrium sustain this dynamic filamentous network: mitochondria fusion and fission. While the molecular mechanism and signaling involved in these two processes is only now being understood, it is well established that dysregulation of these pathways can lead to cellular dysfunction. Mitochondria fusion, mediated by mitofusin-1 and mitofusin-2 are responsible for the lengthening and tethering of adjacent mitochondria to form a network. On the contrary, mitochondria fission involves division of mitochondria and is mediated by fission-1 and dynamin-related protein 1 (Drp-1). Under healthy conditions, the mitochondrial tubular network is established through increased fusion. However, during oxidative stress, mitochondria fission prevails and the filamentous network is broken down to fragmented mitochondrial puncta [141,142].
Hydrogen peroxide, a major contributor to oxidative damage in cells, was shown to induce mitochondria fission in a variety of cells, including fibroblasts. This fragmentation has been shown to be dose- and time-dependent and reversible, suggesting that mitochondria dynamics may be involved in signaling during cellular stress [143–145].
In a recent study, Makino and colleagues demonstrated that superoxide anion was able to induce mitochondrial fragmentation in coronary endothelial cells isolated from a diabetic mouse. Supplementation with TEMPOL, a superoxide scavenger was able to inhibit mitochondrial fragmentation, demonstrating a causal role for superoxide in this process [146]. This was also corroborated by another study that treated HUVEC cells with hydrogen peroxide and demonstrate mitochondria fragmentation, which was suppressed in the presence of an anti-oxidant, N-acetylcysteine (Figure 3) [145]. In another study using neuronal cells, nitric oxide had a profound fission effect that was rescued by antioxidants (Figure 4) [147,148]. Furthermore, mitochondrial fission in neurons occurs prior to the onset of neuronal loss in an animal model of stroke [147]. This was further demonstrated in the kidney following ischemia reperfusion and cisplatin nephrotoxicity [149,150]. Conversely, Yu et al. demonstrated that mitochondria fission plays an important role in ROS overproduction and demonstrate that inhibition of fission reduces ROS generation by the mitochondria [151]. While all these studies implied a role for ROS/RNS in mitochondria fission, a recent study by Giedt and colleagues demonstrated that during simulated ischemia reperfusion in endothelial cells, DRP-1 activation (phosphorylation and oligomerization) was enhanced. This was inhibited by N-acetylcysteine or l-NG-Nitroarginine Methyl Ester (l-NAME, a NOS inhibitor), suggesting that oxidative/nitrosative stress drives Drp1 activation and translocation to mitochondria, which could be the underlying mechanism that induces mitochondrial fission in these studies [152,153]. These studies suggest that different species of ROS can initiate or act as a messenger in the signaling process leading to mitochondrial fragmentation and when persistent, it can lead to chronic mitochondria fission and apoptosis.
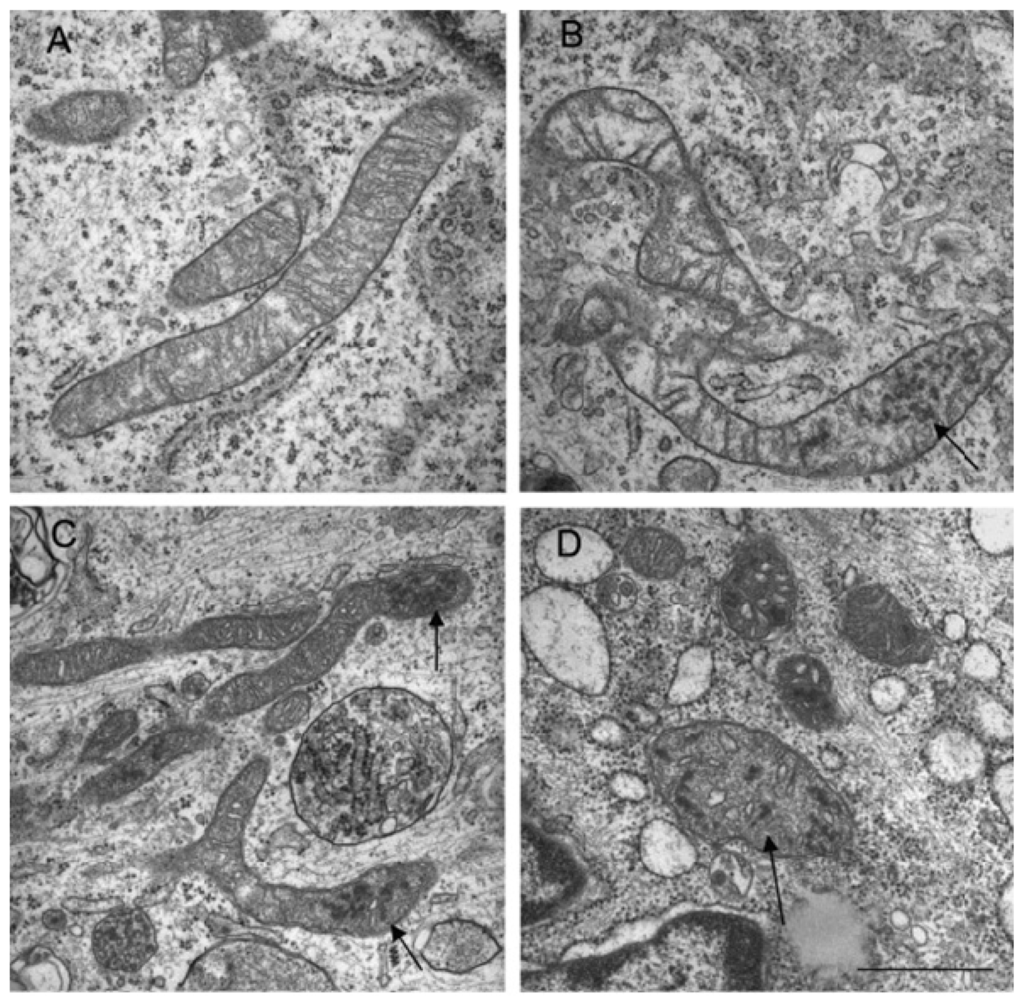
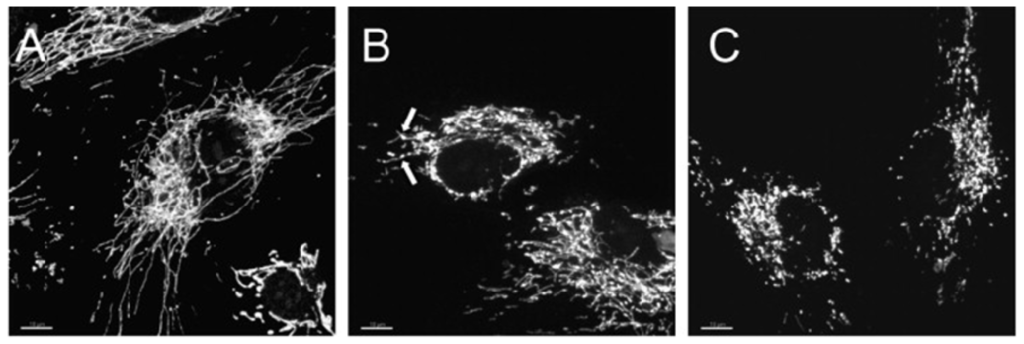
Figure 3.
Changes in mitochondrial structure following hydrogen peroxide treatment. (Top panels) Transmission electron micrographs of mitochondria in untreated (A) and hydrogen peroxide treated cells (B,C,D). Electron dense granules (arrows) and fragmented mitochondria were observed. Bar = 1 μm. (Bottom panels) Mitochondria were stained with Mitotracker Red and existed as tubular (A), intermediate (B) (tubular with swollen regions) and fragmented (C). Bar = 10 μm.
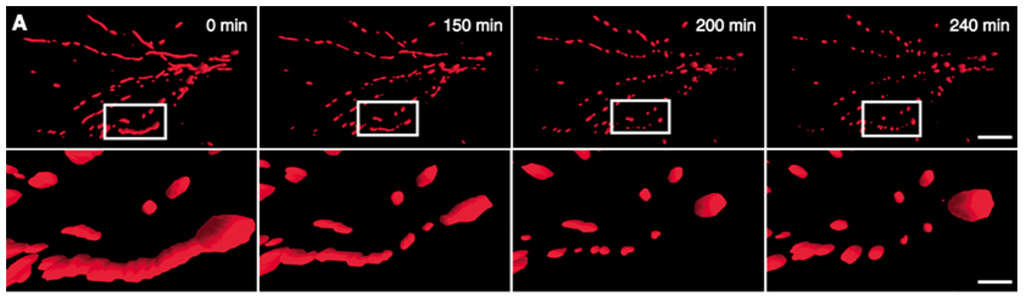
Figure 4.
NO• triggers mitochondrial fission. (A) 3D time-lapse microscopy of mitochondria undergoing fission in a dendritic arbor of a neuron. Neurons were transfected with Mito-DsRed2, pretreated with the pan-caspase inhibitor zVAD-fmk methyl ester (100 M), and exposed to SNOC (200 M). Images were 3D iso-surface rendered. Frames depict representative time points of the movie demonstrating mitochondrial fragmentation within 3 h of NO• exposure (upper panels; scale bar, 15 μm) and closeup views (lower panels; scale bar = 3 μm).
While a majority of these studies point to the role of ROS/RNS to activating fission, there is also evidence suggesting that inhibition of fission may be detrimental to the mitochondria network and cellular homeostasis. Several studies have demonstrated that inhibition of fission led to a decrease in cytochrome c release and delayed apoptosis [154–158]. Similarly, proteins involved in mitochondria fusion have shown to play an anti-apoptotic role [157,159]. However, recent evidence in a number of models suggests that inhibition of fission can lead to mitochondrial dysfunction, increase in ROS levels accompanied by loss of mtDNA and concomitant reduction in energy production and autophagy [160,161].
It is now evident from a variety of studies that ROS affects mitochondrial dynamics and leads to fragmentation of mitochondria tubules. It is also apparent that the mitochondria network plays an important role in mitochondrial function, including respiration, ATP production, apoptosis and functional complementation of mitochondrial DNA mutations and/or damaged proteins. However, it has been suggested that transient or low levels of ROS may in fact benefit the network by selectively removing the dysfunctional mitochondria through autophagy. This idea was proposed using a simulation model that suggested that the vulnerability of the mitochondria network to the harmful effects of ROS is dependent on the dynamics of the network. As one might expect, the healthy mitochondria in close association with the ROS producing dysfunctional mitochondria would be the first to be affected, starting a vicious cycle of uncontrolled and amplified ROS generation. As a result, if these damaged mitochondria were not isolated by fission, they would contaminate the entire network leading to a decrease in energy production and cell proliferation ultimately leading to cell death [162–164].
Nevertheless, it is now clear that defects in mitochondrial fusion or fission alter the susceptibility of cells to undergo apoptosis as documented in a variety of disorders, including heart failure, ischemia reperfusion injury, diabetes, Parkinson’s disease, muscle atrophy, Alzheimer’s disease and aging [141,149,165–172]. Most importantly, these studies provide mechanistic evidence for a role for ROS/RNS and redox signaling in contributing to perturbations in the mitochondrial dynamics and suggest the potential of mitochondria-targeted therapeutics in diseases that involve mitochondrial fragmentation due to uncontrolled ROS/RNS generation.
6. Signaling
Until recently, ROS generation was considered a bane to all aerobic organisms. In fact, this theory was sustained by research that continued to demonstrate generation of high levels of superoxide and nitric oxide by phagocytic cells and its role in host defense [173,174]. In stark contrast, non-immunity based research provided evidence to suggest that these molecules (nitric oxide and hydrogen peroxide) were generated in low levels in other cells, suggesting a distinctive role from its phagocytic activity. In fact, more than three decades ago, pioneers in nitric oxide biology such as Moncada, Ignarro and Furchgott demonstrated a vasodilatory property for this molecule, a discovery that led to their Nobel Prize in 1998 [175–177]. Following this observation, several investigators have established the importance of nitric oxide and other reactive oxygen intermediates in vascular homeostasis [178–187]. Indeed, there is compelling evidence today that demonstrates that these reactive species function as a second messenger in signal transduction and affect cellular function in different tissues [188–205]. Here in we will only focus on the pathways that best illustrate the role of these species in modulating signal transduction.
Since the discovery of a dichotomous role of reactive species in immune function and signal transduction, research in this field grew at an exponential pace and the pursuit for the mechanisms involved began. In a simplistic manner, these regulations can be hypothesized to occur through simple oxidative/nitrosative reactions. For instance, cysteine residues on proteins may be oxidized and hence significantly alter the activity of these proteins. These proteins (e.g., transcription factors, kinases, phosphatases) may then affect downstream signaling cascades that affect cellular responses to stimuli. Alternatively, these residues may be modified through nitrosylation by reactive nitrogen species. More importantly, this redox mechanism may occur at multiple levels in the signal transduction cascade and thereby ultimately alter the fate of a cell.
Earlier research was primarily focused on the ability of hydrogen peroxide to mediate cellular signaling due to its relative stability and ease of measurement. Hydrogen peroxide can also be converted to the highly reactive hydroxyl ion in the presence of a transition metal, such as iron, to amplify oxidative stress and activate cell death pathways. However, there is some consensus that the level of ROS itself may dictate the fate of the cell by modulating different redox-sensitive transcription factors and hence lead to diverse biological responses. For instance, low ROS levels induce Nuclear factor erythroid 2-related factor 2 (Nrf2), a potent transcription factor responsible for the induction of several antioxidant enzymes, including but not limited to NADPH quinone oxidoreductase (NQO1), glutathione S-transferase, heme oxygenase-1 (HO-1), ferritin and γ-glutamylcysteine synthetase [206,207]. When there is a moderate increase in ROS levels, NF-κB and AP-1 are activated and when there are extremely high levels or persistent ROS accumulation in the cell, membrane permeability transition pore is opened, cytochrome c is released from the mitochondria and apoptosis is triggered. These signaling pathways are regulated in multiple ways, some of which are independent of ROS.
7. Modulation of Nrf2 by Reactive Oxygen and Nitrogen Species
Under normal conditions, low levels of ROS generated by the mitochondria are neutralized or scavenged by anti-oxidants and scavengers present in the cell, many of which under the regulation of Nrf2. During mild increases in ROS, Nrf2 translocates to the nucleus, binds to the anti-oxidant response element (ARE) present on stress responsive genes and activates the promoters [207]. It has been suggested that more than 200 genes involved in the cellular antioxidant and anti-inflammatory defense were regulated by Nrf2 (reviewed in [208–214]) and suggesting that the activation of Nrf2 is tightly regulated. Indeed, the regulation of Nrf2 is complex and involves multiple factors including phosphorylation, protein interaction and stability [215–217]. Nrf2 stability is the most widely studied and involves a zinc zinc metalloprotein, Kelch-like ECH associated protein1 (Keap1). Under unstimulated conditions, Nrf2 is sequestered in the cytosol by Keap1 and is targeted for ubiquitin-dependent proteosomal degradation [218–221]. A variety of stimuli can induce Nrf2 by disrupting the Keap1-Nrf2 interaction and inhibiting Nrf2 ubiquitination (reviewed in [215,222–228]. With regard to hydrogen peroxide and nitric oxide, both have shown to oxidize several cysteine residues on Keap1, thereby forming intra- and intermolecular disulphides and inactivating Keap1 [229–234]. Interestingly, the majority of Nrf2 activators increase reactive oxygen and nitrogen species. It is therefore tempting to speculate that Nrf2 activation necessitates the presence of reactive species. While Nrf2 activation and its downstream signaling are important mediators of anti-oxidant signaling during exposure to low levels of reactive species, it is widely believed that NF-κB and AP-1 signaling pathways are switched on to protect against increased cellular stress.
8. NFκB Activation by Reactive Species
Another interesting signaling pathway that is modulated by reactive species is NFκB. Since its discovery by Baltimore almost three decades ago [235], it has been implicated in the regulation of various cellular responses to stress such as apoptosis and inflammation [236–251]. Generation of various NFκB transgenic knockout mice further established its role in these processes [252]. Although it was initially presumed that these factors were expressed only in B lymphocytes, growing evidence indicates its presence in most mammalian cells. Furthermore, aberrant NFκB signaling has been associated with the pathogenesis of a number of diseases, including cancer, atherosclerosis and schizophrenia [253–256]. NFκB signaling is one of the most complex pathways and consists of five different transcription factors known as p65, p50, p52, c-Rel ad RelB. These factors contain a homology domain in their N-terminus (Rel homology domain, RHD) that enables interaction with DNA and dimerization with other RHD containing factors. NFκB is predominantly regulated by a family of inhibitory proteins, IκB that sequester this protein in the cytoplasm using this domain [236,257,258]. The IκB family consists of seven IκB proteins that can modulate NFκB signaling. The contributory role of the NFκB family to transcriptional regulation of genes is complex and continues to grow. For instance, while p50 and p52 homodimers have inhibitory effect on transcription on numerous genes, heterodimers of these factors stimulate transcription [259–261]. NFκB is induced by a variety of stimuli, including cytokines and oxidative stress and plays an important role in a variety of processes such as cell proliferation, inflammation and apoptosis. It achieves this versatility through its regulation of a multitude of genes, including those involved in reactive species generation such as NOS and cyclooxgenase 2 [262–268]. Herein we will highlight the role of reactive species in regulating NFκB signaling.
NFκB was one of the first transcription factors to be described as an oxidative stress responsive transcription factor. This was first demonstrated by Schreck and colleagues in T cells using hydrogen peroxide [269] and further confirmed by incubating cells with scavengers of ROS and demonstrating that ROS-mediated NFκB activation was inhibited [269,270]. Later studies demonstrated that hydrogen peroxide and nitric oxide could activate NFκB signaling in a variety of cells, including cancer cell lines (MCF-7, HeLa, LNCaP), fibroblasts, chondrocytes, lymphocytes, macrophages, epithelial and endothelial cells [269,271–281]. Similarly, NFκB activation was impaired in the presence of anti-oxidants, such as NAC, MnSOD and glutathione peroxidase [274,282–285]. On the other hand, high levels of nitric oxide have shown to inhibit NFκB activation in endothelial cells, hepatocytes, cancerous cells, macrophages and T cells [286–292].
While activation of NFκB signaling promotes survival during stress mediated by a variety of insults, it has also shown to induce death (reviewed in [293]). It seems plausible that while ROS activates NFκB, NFκB signaling may in turn inhibit ROS production to promote survival. The list of anti-oxidant genes induced by NFκB is extensive and comprises of MnSOD, Ferritin, catalase, glutathione s-transferase, heme oxygenase-1, glutathione peroxidase and many others. On the contrary, NFκB plays an important role in inflammation through upregulation of ROS producing enzymes such as NADPH oxidase, NOS, xanthine oxidase, cyclooxygenases. Interestingly, another mechanism by which reactive species may affect signaling is through its oxidation potential. While on one hand, oxidative stress increases NFκB activity, higher levels of ROS may lead to oxidation of NFκB and reduce its activity. For instance, transcriptional activation of NOS requires NFκB [288,294], and once NO• is formed, it can either activate or inhibit NFκB signaling [272,289,294–299].
It now seems more certain that reducing conditions are required in the nucleus for NF-κB DNA binding [300,301], whereas oxidizing conditions in the cytoplasm promote NF-κB activation [275,302,303]. It has been suggested that reactive species stimulate NFκB in the cytosol whereas inhibit its activity in the nucleus [304]. The DNA binding ability of this transcription factor has shown to be modulated by redox status in the cell [300,305,306]. There is evidence to suggest that redox factor protein, Ref-1 reduces cysteine 62 in NFκB in the nucleus and this reaction is required for NFκB to bind to DNA [305]. Conversely, oxidation of this residue inhibits binding to DNA [306]. In addition, glutathionylation of NFκB in the presence of reactive oxygen species led to a decrease in its DNA binding ability and downstream transcriptional activity [307].
Another mechanism by which NFκB can be modulated is through phosphorylation of NFκB or its inhibitor, IκB. Hydrogen peroxide has shown to alter the activity of IκK, a kinase that phosphorylates IκB and hence allows NFκB translocation to the nucleus [308]. Alternatively, ROS can increase the activity of IκK indirectly through modulation of Akt signaling [309]. Furthermore, it was recently demonstrated that ROS mediated phosphorylation of RelA on serine 276 is essential for TNF-α induced NFκB activation and signaling [310]. To summarize, the regulation of NFκB by reactive species and the inverse is extremely complex and more research is warranted for a complete understanding of the mechanisms involved.
9. Other Signaling Pathways
Due to the vast wealth of knowledge available on ROS and RNS signaling, we only focused on two of the most important pathways. However, these species have been implicated in MAPK [311–315], HIF-1 α [316,317], p53 [318–323], AP-1 [313,324,325], SP-1 [313,326,327], apoptosis (caspase regulation) [328–332], cytokine [333–337] and fibroblast-derived growth factor [338–340] and platelet derived growth factor [97,340–344]. Of note, this list is not all encompassing and is constantly growing as research in this field progresses. Furthermore, ROS and RNS mediate mitochondrial cell death signaling pathways and have been extensively reviewed and hence not discussed in this review [345–353].
10. Antioxidants
As discussed in the previous sections, while hydrogen peroxide and NO• are important mediators of cellular processes, higher levels of these species or the presence of highly reactive species such as hydroxyl radical and peroxynitrite can potentially damage cellular components, leading to death. Nature has evolved to combat this stress by developing antioxidants that enable removal of these oxidative species. These antioxidants can be broadly classified as enzymatic (superoxide dismutase, catalase) or non-enzymatic antioxidants (bilirubin, glutathione). While they are directly involved in eliminating reactive species, cells have evolved other mechanisms that can indirectly reduce or inhibit generation of reactive species, such as heme oxygenase-1, ferritin, ceruloplasmin and glutathione transferase.
A beneficial role for antioxidants has been extensively investigated in both cell culture and animal models of injury and disease (reviewed in [354–376]. However, translation of these studies to the clinical setting has yielded confounding results (reviewed in [360,377–385]). In summary, it appears that supplementation of exogenous antioxidants in several clinical trials had no effect or led to an increase in mortality. Several explanations have been suggested to explain these findings. One school of thought believes that cells employ homeostatic mechanisms to restrict the total allowable antioxidant activity. Therefore, supplying exogenous antioxidants may decrease the rate of synthesis or uptake of antioxidants, so that the total antioxidant potential remains unaltered. Yet another explanation could be simply that the amount of antioxidant is insufficient and is not targeted to the site of excessive reactive species generation. In addition, given the importance of these molecules in signaling and other cellular activities, it may be plausible that their complete removal may lead to altered cellular mechanisms and hence worse outcomes. Furthermore, the relative specificity and efficiency of exogenous antioxidants to reduce each of these reactive species may be different. More importantly, the oxidants responsible for injury must be evaluated. For instance, in a model of cisplatin-mediated renal epithelial injury, overexpression of MnSOD was protective whereas, catalase overexpression was ineffective [386]. This supports the notion that injury may be mediated through different oxidative and nitrosative species and future therapies must be targeted based on the detrimental species generated. Therefore, despite the advances made in deciphering the molecular mechanisms that are regulated by oxidants and antioxidants, translation of these pathways to benefit mankind is still in its infancy and more studies are warranted.
11. Concluding Remarks
The field of free radicals has evolved over the past few decades and has significantly contributed to understanding normal physiology and pathophysiology. Research has provided irrefutable evidence that reactive oxygen and nitrogen species are important mediators of cellular response to stress and they function through several mechanisms including, modulation of autophagy, mitochondrial network, signaling and apoptosis. However, high levels of certain reactive species can contribute to cell injury and progression of diseases. These studies granted opportunities for implementing the use of antioxidants in clinical trials, only some of which provided promising results. Therefore, there is an urgent need to comprehensively assess the amounts of different species generated during injury and their relative role in the pathogenesis of disease. Targeting the detrimental reactive species through antioxidant therapy would perhaps yield better outcomes in the clinical trials. In this review, we provided a brief discussion of some of the major pathways that are regulated by ROS and RNS. This review has focused on the complexities and dual roles of these species in cellular activities that affect health and disease. This review will not only provide an understanding of the intricate underlying mechanisms of the reactive species, but will also enable opportunities for the design and development of effective novel therapeutic strategies.
Acknowledgments
This work was supported by an AHA grant 11POST7600074 (S.B.) a Veterans Affairs Program Project Award 1IP1BX001595 (E.A.J) a Merit Review Award of the Veterans Affairs Administration 1I01BX001073 (E.A.J) and an NIH Grant R01ES014948 (E.A.J).
Conflict of Interest
The authors declare no conflict of interest.
References
- Ernster, L.; Schatz, G. Mitochondria: A historical review. J. Cell Biol 1981, 91, 227s–255s. [Google Scholar]
- Gray, M.W.; Burger, G.; Lang, B.F. Mitochondrial evolution. Science 1999, 283, 1476–1481. [Google Scholar]
- Hollenbeck, P.J.; Saxton, W.M. The axonal transport of mitochondria. J. Cell Sci 2005, 118, 5411–5419. [Google Scholar]
- Lenaz, G. The mitochondrial production of reactive oxygen species: Mechanisms and implications in human pathology. IUBMB Life 2001, 52, 159–164. [Google Scholar]
- Swerdlow, R.H. Brain aging, Alzheimer’s disease, and mitochondria. Biochim. Biophys. Acta 2011, 1812, 1630–1639. [Google Scholar]
- Griffiths, E.J. Mitochondria and heart disease. Adv. Exp. Med. Biol 2012, 942, 249–267. [Google Scholar]
- Birch-Machin, M.A. Mitochondria and skin disease. Clin. Exp. Dermatol 2000, 25, 141–146. [Google Scholar]
- Frohman, M.A. Mitochondria as integrators of signal transduction and energy production in cardiac physiology and disease. J. Mol. Med. (Berl. ) 2010, 88, 967–970. [Google Scholar]
- Grattagliano, I.; Russmann, S.; Diogo, C.; Bonfrate, L.; Oliveira, P.J.; Wang, D.Q.; Portincasa, P. Mitochondria in chronic liver disease. Curr. Drug Targets 2011, 12, 879–893. [Google Scholar]
- Duchen, M.R. Mitochondria in health and disease: Perspectives on a new mitochondrial biology. Mol. Aspects Med 2004, 25, 365–451. [Google Scholar]
- Smith, R.A.; Adlam, V.J.; Blaikie, F.H.; Manas, A.R.; Porteous, C.M.; James, A.M.; Ross, M.F.; Logan, A.; Cocheme, H.M.; Trnka, J.; et al. Mitochondria-targeted antioxidants in the treatment of disease. Ann. N. Y. Acad. Sci 2008, 1147, 105–111. [Google Scholar]
- Gobe, G.; Crane, D. Mitochondria, reactive oxygen species and cadmium toxicity in the kidney. Toxicol. Lett 2010, 198, 49–55. [Google Scholar]
- Cloonan, S.M.; Choi, A.M. Mitochondria: Commanders of innate immunity and disease? Curr. Opin. Immunol 2012, 24, 32–40. [Google Scholar]
- Nunnari, J.; Suomalainen, A. Mitochondria: In sickness and in health. Cell 2012, 148, 1145–1159. [Google Scholar]
- Schapira, A.H. Mitochondrial diseases. Lancet 2012, 379, 1825–1834. [Google Scholar]
- Armstrong, J.S. Mitochondrial medicine: Pharmacological targeting of mitochondria in disease. Br. J. Pharmacol 2007, 151, 1154–1165. [Google Scholar]
- Minocherhomji, S.; Tollefsbol, T.O.; Singh, K.K. Mitochondrial regulation of epigenetics and its role in human diseases. Epigenetics 2012, 7, 326–334. [Google Scholar]
- Rocha, M.; Apostolova, N.; Hernandez-Mijares, A.; Herance, R.; Victor, V.M. Oxidative stress and endothelial dysfunction in cardiovascular disease: Mitochondria-targeted therapeutics. Curr. Med. Chem 2010, 17, 3827–3841. [Google Scholar]
- Diogo, C.V.; Grattagliano, I.; Oliveira, P.J.; Bonfrate, L.; Portincasa, P. Re-wiring the circuit: Mitochondria as a pharmacological target in liver disease. Curr. Med. Chem 2011, 18, 5448–5465. [Google Scholar]
- Johannsen, D.L.; Ravussin, E. The role of mitochondria in health and disease. Curr. Opin. Pharmacol 2009, 9, 780–786. [Google Scholar]
- Duchen, M.R. Roles of mitochondria in health and disease. Diabetes 2004, 53, S96–S102. [Google Scholar]
- Duchen, M.R.; Szabadkai, G. Roles of mitochondria in human disease. Essays Biochem 2010, 47, 115–137. [Google Scholar]
- Hedskog, L.; Zhang, S.; Ankarcrona, M. Strategic role for mitochondria in Alzheimer’s disease and cancer. Antioxid. Redox Signal 2012, 16, 1476–1491. [Google Scholar]
- Watson, K.; Haslam, J.M.; Linnane, A.W. Biogenesis of mitochondria. 13. The isolation of mitochondrial structures from anaerobically grown Saccharomyces cerevisiae. J. Cell Biol 1970, 46, 88–96. [Google Scholar]
- Green, D.E.; Oda, T. On the unit of mitochondrial structure and function. J. Biochem 1961, 49, 742–757. [Google Scholar]
- Neupert, W. Protein import into mitochondria. Annu. Rev. Biochem 1997, 66, 863–917. [Google Scholar]
- Pfanner, N.; Craig, E.A.; Honlinger, A. Mitochondrial preprotein translocase. Annu. Rev. Cell Dev. Biol 1997, 13, 25–51. [Google Scholar]
- Dekker, P.J.; Ryan, M.T.; Brix, J.; Muller, H.; Honlinger, A.; Pfanner, N. Preprotein translocase of the outer mitochondrial membrane: Molecular dissection and assembly of the general import pore complex. Mol. Cell. Biol 1998, 18, 6515–6524. [Google Scholar]
- Yamamoto, H.; Itoh, N.; Kawano, S.; Yatsukawa, Y.; Momose, T.; Makio, T.; Matsunaga, M.; Yokota, M.; Esaki, M.; Shodai, T.; et al. Dual role of the receptor Tom20 in specificity and efficiency of protein import into mitochondria. Proc. Natl. Acad. Sci. USA 2011, 108, 91–96. [Google Scholar]
- Herrmann, J.M.; Riemer, J. The intermembrane space of mitochondria. Antioxid. Redox Signal 2010, 13, 1341–1358. [Google Scholar]
- Davies, K.M.; Strauss, M.; Daum, B.; Kief, J.H.; Osiewacz, H.D.; Rycovska, A.; Zickermann, V.; Kuhlbrandt, W. Macromolecular organization of ATP synthase and complex I in whole mitochondria. Proc. Natl. Acad. Sci. USA 2011, 108, 14121–14126. [Google Scholar]
- O’Rourke, B. Mitochondrial ion channels. Annu. Rev. Physiol 2007, 69, 19–49. [Google Scholar]
- Asin-Cayuela, J.; Gustafsson, C.M. Mitochondrial transcription and its regulation in mammalian cells. Trends Biochem. Sci 2007, 32, 111–117. [Google Scholar]
- Halliwell, B.; Gutteridge, J.M. Oxygen toxicity, oxygen radicals, transition metals and disease. Biochem. J 1984, 219, 1–14. [Google Scholar]
- Cline, M.J.; Lehrer, R.I. d-amino acid oxidase in leukocytes: A possible d-amino-acid-linked antimicrobial system. Proc. Natl. Acad. Sci. USA 1969, 62, 756–763. [Google Scholar]
- Ji, L.L. Antioxidant signaling in skeletal muscle: A brief review. Exp. Gerontol 2007, 42, 582–593. [Google Scholar]
- Kim, C.; Kim, J.Y.; Kim, J.H. Cytosolic phospholipase A2, lipoxygenase metabolites, and reactive oxygen species. BMB Rep 2008, 41, 555–559. [Google Scholar]
- Nishino, T. The conversion of xanthine dehydrogenase to xanthine oxidase and the role of the enzyme in reperfusion injury. J. Biochem 1994, 116, 1–6. [Google Scholar]
- Nishino, T.; Okamoto, K.; Eger, B.T.; Pai, E.F. Mammalian xanthine oxidoreductase—Mechanism of transition from xanthine dehydrogenase to xanthine oxidase. FEBS J 2008, 275, 3278–3289. [Google Scholar]
- Sumimoto, H.; Miyano, K.; Takeya, R. Molecular composition and regulation of the Nox family NAD(P)H oxidases. Biochem. Biophys. Res. Commun 2005, 338, 677–686. [Google Scholar]
- Zangar, R.C.; Davydov, D.R.; Verma, S. Mechanisms that regulate production of reactive oxygen species by cytochrome P450. Toxicol. Appl. Pharmacol 2004, 199, 316–331. [Google Scholar]
- Brown, G.C. Control of respiration and ATP synthesis in mammalian mitochondria and cells. Biochem. J 1992, 284, 1–13. [Google Scholar]
- Paul, T. Effect of a prolonged superoxide flux on transferrin and ferritin. Arch. Biochem. Biophys 2000, 382, 253–261. [Google Scholar]
- Biemond, P.; Swaak, A.J.; van Eijk, H.G.; Koster, J.F. Superoxide dependent iron release from ferritin in inflammatory diseases. Free Radic. Biol. Med 1988, 4, 185–198. [Google Scholar]
- Liochev, S.L. The role of iron-sulfur clusters in in vivo hydroxyl radical production. Free Radic. Res 1996, 25, 369–384. [Google Scholar]
- Flint, D.H.; Tuminello, J.F.; Emptage, M.H. The inactivation of Fe–S cluster containing hydro-lyases by superoxide. J. Biol. Chem 1993, 268, 22369–22376. [Google Scholar]
- Lenaz, G. Mitochondria and reactive oxygen species. Which role in physiology and pathology? Adv. Exp. Med. Biol 2012, 942, 93–136. [Google Scholar]
- McLennan, H.R.; Esposti, M.D. The contribution of mitochondrial respiratory complexes to the production of reactive oxygen species. J. Bioenerg. Biomembr 2000, 32, 153–162. [Google Scholar]
- Fato, R.; Bergamini, C.; Leoni, S.; Strocchi, P.; Lenaz, G. Generation of reactive oxygen species by mitochondrial complex I: Implications in neurodegeneration. Neurochem. Res 2008, 33, 2487–2501. [Google Scholar]
- Murphy, M.P. How mitochondria produce reactive oxygen species. Biochem. J 2009, 417, 1–13. [Google Scholar]
- Yankovskaya, V.; Horsefield, R.; Tornroth, S.; Luna-Chavez, C.; Miyoshi, H.; Leger, C.; Byrne, B.; Cecchini, G.; Iwata, S. Architecture of succinate dehydrogenase and reactive oxygen species generation. Science 2003, 299, 700–704. [Google Scholar]
- Casteilla, L.; Rigoulet, M.; Penicaud, L. Mitochondrial ROS metabolism: Modulation by uncoupling proteins. IUBMB Life 2001, 52, 181–188. [Google Scholar]
- Starkov, A.A.; Fiskum, G. Myxothiazol induces H2O2 production from mitochondrial respiratory chain. Biochem. Biophys. Res. Commun 2001, 281, 645–650. [Google Scholar]
- Barja, G. Mitochondrial oxygen radical generation and leak: Sites of production in states 4 and 3, organ specificity, and relation to aging and longevity. J. Bioenerg. Biomembr 1999, 31, 347–366. [Google Scholar]
- Kushnareva, Y.; Murphy, A.N.; Andreyev, A. Complex I-mediated reactive oxygen species generation: Modulation by cytochrome c and NAD(P)+ oxidation-reduction state. Biochem. J 2002, 368, 545–553. [Google Scholar]
- Barja, G.; Herrero, A. Localization at complex I and mechanism of the higher free radical production of brain nonsynaptic mitochondria in the short-lived rat than in the longevous pigeon. J. Bioenerg. Biomembr 1998, 30, 235–243. [Google Scholar]
- Turrens, J.F.; Boveris, A. Generation of superoxide anion by the NADH dehydrogenase of bovine heart mitochondria. Biochem. J 1980, 191, 421–427. [Google Scholar]
- Miwa, S.; St-Pierre, J.; Partridge, L.; Brand, M.D. Superoxide and hydrogen peroxide production by Drosophila mitochondria. Free Radic. Biol. Med 2003, 35, 938–948. [Google Scholar]
- Drahota, Z.; Chowdhury, S.K.; Floryk, D.; Mracek, T.; Wilhelm, J.; Rauchova, H.; Lenaz, G.; Houstek, J. Glycerophosphate-dependent hydrogen peroxide production by brown adipose tissue mitochondria and its activation by ferricyanide. J. Bioenerg. Biomembr 2002, 34, 105–113. [Google Scholar]
- Seifert, E.L.; Estey, C.; Xuan, J.Y.; Harper, M.E. Electron transport chain-dependent and -independent mechanisms of mitochondrial H2O2 emission during long-chain fatty acid oxidation. J. Biol. Chem 2010, 285, 5748–5758. [Google Scholar]
- Maurel, A.; Hernandez, C.; Kunduzova, O.; Bompart, G.; Cambon, C.; Parini, A.; Frances, B. Age-dependent increase in hydrogen peroxide production by cardiac monoamine oxidase A in rats. Am. J. Physiol. Heart Circ. Physiol 2003, 284, H1460–H1467. [Google Scholar]
- Starkov, A.A.; Fiskum, G.; Chinopoulos, C.; Lorenzo, B.J.; Browne, S.E.; Patel, M.S.; Beal, M.F. Mitochondrial alpha-ketoglutarate dehydrogenase complex generates reactive oxygen species. J. Neurosci 2004, 24, 7779–7788. [Google Scholar]
- Tretter, L.; Adam-Vizi, V. Generation of reactive oxygen species in the reaction catalyzed by alpha-ketoglutarate dehydrogenase. J. Neurosci 2004, 24, 7771–7778. [Google Scholar]
- MacMillan-Crow, L.A.; Crow, J.P.; Thompson, J.A. Peroxynitrite-mediated inactivation of manganese superoxide dismutase involves nitration and oxidation of critical tyrosine residues. Biochemistry 1998, 37, 1613–1622. [Google Scholar]
- Lacza, Z.; Snipes, J.A.; Zhang, J.; Horvath, E.M.; Figueroa, J.P.; Szabo, C.; Busija, D.W. Mitochondrial nitric oxide synthase is not eNOS, nNOS or iNOS. Free Radic. Biol. Med 2003, 35, 1217–1228. [Google Scholar]
- Ghafourifar, P.; Richter, C. Nitric oxide synthase activity in mitochondria. FEBS Lett 1997, 418, 291–296. [Google Scholar]
- Giulivi, C.; Poderoso, J.J.; Boveris, A. Production of nitric oxide by mitochondria. J. Biol. Chem 1998, 273, 11038–11043. [Google Scholar]
- Alvarez, S.; Valdez, L.B.; Zaobornyj, T.; Boveris, A. Oxygen dependence of mitochondrial nitric oxide synthase activity. Biochem. Biophys. Res. Commun 2003, 305, 771–775. [Google Scholar]
- Lacza, Z.; Puskar, M.; Figueroa, J.P.; Zhang, J.; Rajapakse, N.; Busija, D.W. Mitochondrial nitric oxide synthase is constitutively active and is functionally upregulated in hypoxia. Free Radic. Biol. Med 2001, 31, 1609–1615. [Google Scholar]
- Bates, T.E.; Loesch, A.; Burnstock, G.; Clark, J.B. Immunocytochemical evidence for a mitochondrially located nitric oxide synthase in brain and liver. Biochem. Biophys. Res. Commun 1995, 213, 896–900. [Google Scholar]
- Haynes, V.; Elfering, S.; Traaseth, N.; Giulivi, C. Mitochondrial nitric-oxide synthase: Enzyme expression, characterization, and regulation. J. Bioenerg. Biomembr 2004, 36, 341–346. [Google Scholar]
- Giulivi, C. Mitochondria as generators and targets of nitric oxide. Novartis Found. Symp. 2007, 287, 92–100, discussion 100–104. [Google Scholar]
- Lacza, Z.; Kozlov, A.V.; Pankotai, E.; Csordas, A.; Wolf, G.; Redl, H.; Kollai, M.; Szabo, C.; Busija, D.W.; Horn, T.F. Mitochondria produce reactive nitrogen species via an arginine-independent pathway. Free Radic. Res 2006, 40, 369–378. [Google Scholar]
- Kozlov, A.V.; Staniek, K.; Nohl, H. Nitrite reductase activity is a novel function of mammalian mitochondria. FEBS Lett 1999, 454, 127–130. [Google Scholar]
- Lacza, Z.; Pankotai, E.; Csordas, A.; Gero, D.; Kiss, L.; Horvath, E.M.; Kollai, M.; Busija, D.W.; Szabo, C. Mitochondrial NO and reactive nitrogen species production: Does mtNOS exist? Nitric. Oxide 2006, 14, 162–168. [Google Scholar]
- Mason, M.G.; Nicholls, P.; Wilson, M.T.; Cooper, C.E. Nitric oxide inhibition of respiration involves both competitive (heme) and noncompetitive (copper) binding to cytochrome c oxidase. Proc. Natl. Acad. Sci. USA 2006, 103, 708–713. [Google Scholar]
- Cooper, C.E.; Giulivi, C. Nitric oxide regulation of mitochondrial oxygen consumption II: Molecular mechanism and tissue physiology. Am. J. Physiol. Cell Physiol 2007, 292, C1993–C2003. [Google Scholar]
- Gladwin, M.T.; Shiva, S. The ligand binding battle at cytochrome c oxidase: How NO regulates oxygen gradients in tissue. Circ. Res 2009, 104, 1136–1138. [Google Scholar]
- Brunori, M.; Forte, E.; Arese, M.; Mastronicola, D.; Giuffre, A.; Sarti, P. Nitric oxide and the respiratory enzyme. Biochim. Biophys. Acta 2006, 1757, 1144–1154. [Google Scholar]
- Sarti, P.; Forte, E.; Giuffre, A.; Mastronicola, D.; Magnifico, M.C.; Arese, M. The chemical interplay between nitric oxide and mitochondrial cytochrome c oxidase: Reactions, effectors and pathophysiology. Int. J. Cell Biol 2012, 2012, 571067. [Google Scholar]
- Poderoso, J.J.; Carreras, M.C.; Lisdero, C.; Riobo, N.; Schopfer, F.; Boveris, A. Nitric oxide inhibits electron transfer and increases superoxide radical production in rat heart mitochondria and submitochondrial particles. Arch. Biochem. Biophys 1996, 328, 85–92. [Google Scholar]
- Brown, G.C.; Borutaite, V. Inhibition of mitochondrial respiratory complex I by nitric oxide, peroxynitrite and S-nitrosothiols. Biochim. Biophys. Acta 2004, 1658, 44–49. [Google Scholar]
- Nisoli, E.; Falcone, S.; Tonello, C.; Cozzi, V.; Palomba, L.; Fiorani, M.; Pisconti, A.; Brunelli, S.; Cardile, A.; Francolini, M.; et al. Mitochondrial biogenesis by NO yields functionally active mitochondria in mammals. Proc. Natl. Acad. Sci. USA 2004, 101, 16507–16512. [Google Scholar]
- Shen, W.; Hintze, T.H.; Wolin, M.S. Nitric oxide. An important signaling mechanism between vascular endothelium and parenchymal cells in the regulation of oxygen consumption. Circulation 1995, 92, 3505–3512. [Google Scholar]
- Brown, G.C. Nitric oxide and mitochondrial respiration. Biochim. Biophys. Acta 1999, 1411, 351–369. [Google Scholar]
- Nisoli, E.; Clementi, E.; Paolucci, C.; Cozzi, V.; Tonello, C.; Sciorati, C.; Bracale, R.; Valerio, A.; Francolini, M.; Moncada, S.; et al. Mitochondrial biogenesis in mammals: The role of endogenous nitric oxide. Science 2003, 299, 896–899. [Google Scholar]
- Xie, Y.W.; Kaminski, P.M.; Wolin, M.S. Inhibition of rat cardiac muscle contraction and mitochondrial respiration by endogenous peroxynitrite formation during posthypoxic reoxygenation. Circ. Res 1998, 82, 891–897. [Google Scholar]
- Radi, R.; Rodriguez, M.; Castro, L.; Telleri, R. Inhibition of mitochondrial electron transport by peroxynitrite. Arch. Biochem. Biophys 1994, 308, 89–95. [Google Scholar]
- Piantadosi, C.A.; Suliman, H.B. Redox regulation of mitochondrial biogenesis. Free Radic. Biol. Med 2012, 53, 2043–2053. [Google Scholar]
- Bourens, M.; Fontanesi, F.; Soto, I.C.; Liu, J.; Barrientos, A. Redox and reactive oxygen species regulation of mitochondrial cytochrome c oxidase biogenesis. Antioxid. Redox Signal. 2012. [Google Scholar] [CrossRef]
- Nisoli, E.; Tonello, C.; Cardile, A.; Cozzi, V.; Bracale, R.; Tedesco, L.; Falcone, S.; Valerio, A.; Cantoni, O.; Clementi, E.; et al. Calorie restriction promotes mitochondrial biogenesis by inducing the expression of eNOS. Science 2005, 310, 314–317. [Google Scholar]
- Nisoli, E.; Clementi, E.; Moncada, S.; Carruba, M.O. Mitochondrial biogenesis as a cellular signaling framework. Biochem. Pharmacol 2004, 67, 1–15. [Google Scholar]
- Lee, H.C.; Wei, Y.H. Mitochondrial biogenesis and mitochondrial DNA maintenance of mammalian cells under oxidative stress. Int. J. Biochem. Cell Biol 2005, 37, 822–834. [Google Scholar]
- Kong, X.; Wang, R.; Xue, Y.; Liu, X.; Zhang, H.; Chen, Y.; Fang, F.; Chang, Y. Sirtuin 3, a new target of PGC-1alpha, plays an important role in the suppression of ROS and mitochondrial biogenesis. PLoS One 2010, 5, e11707. [Google Scholar]
- Piantadosi, C.A.; Carraway, M.S.; Haden, D.W.; Suliman, H.B. Protecting the permeability pore and mitochondrial biogenesis. Novartis Found. Symp. 2007, 280, 266–276, discussion 276–280. [Google Scholar]
- Handy, D.E.; Loscalzo, J. Redox regulation of mitochondrial function. Antioxid. Redox Signal 2012, 16, 1323–1367. [Google Scholar]
- Le Bras, M.; Clement, M.V.; Pervaiz, S.; Brenner, C. Reactive oxygen species and the mitochondrial signaling pathway of cell death. Histol. Histopathol 2005, 20, 205–219. [Google Scholar]
- Ma, Z.A. The role of peroxidation of mitochondrial membrane phospholipids in pancreatic beta-cell failure. Curr. Diabetes Rev 2012, 8, 69–75. [Google Scholar]
- Mizushima, N.; Levine, B.; Cuervo, A.M.; Klionsky, D.J. Autophagy fights disease through cellular self-digestion. Nature 2008, 451, 1069–1075. [Google Scholar]
- Mizushima, N. Autophagy: Process and function. Genes Dev 2007, 21, 2861–2873. [Google Scholar]
- Kiffin, R.; Bandyopadhyay, U.; Cuervo, A.M. Oxidative stress and autophagy. Antioxid. Redox Signal 2006, 8, 152–162. [Google Scholar]
- Mizushima, N.; Klionsky, D.J. Protein turnover via autophagy: Implications for metabolism. Annu. Rev. Nutr 2007, 27, 19–40. [Google Scholar]
- Nakatogawa, H.; Suzuki, K.; Kamada, Y.; Ohsumi, Y. Dynamics and diversity in autophagy mechanisms: Lessons from yeast. Nat. Rev. Mol. Cell Biol 2009, 10, 458–467. [Google Scholar]
- Klionsky, D.J.; Cregg, J.M.; Dunn, W.A., Jr; Emr, S.D.; Sakai, Y.; Sandoval, I.V.; Sibirny, A.; Subramani, S.; Thumm, M.; Veenhuis, M.; et al. Cell 2003, 5, 539–545.
- Levine, B.; Mizushima, N.; Virgin, H.W. Autophagy in immunity and inflammation. Nature 2011, 469, 323–335. [Google Scholar]
- Azad, M.B.; Chen, Y.; Gibson, S.B. Regulation of autophagy by reactive oxygen species (ROS): Implications for cancer progression and treatment. Antioxid. Redox Signal 2009, 11, 777–790. [Google Scholar]
- Scherz-Shouval, R.; Shvets, E.; Fass, E.; Shorer, H.; Gil, L.; Elazar, Z. Reactive oxygen species are essential for autophagy and specifically regulate the activity of Atg4. EMBO J 2007, 26, 1749–1760. [Google Scholar]
- Mathew, R.; White, E. Autophagy, stress, and cancer metabolism: What doesn’t kill you makes you stronger. Cold Spring Harb. Symp. Quant. Biol 2011, 76, 389–396. [Google Scholar]
- Li, Z.Y.; Yang, Y.; Ming, M.; Liu, B. Mitochondrial ROS generation for regulation of autophagic pathways in cancer. Biochem. Biophys. Res. Commun 2011, 414, 5–8. [Google Scholar]
- Ishdorj, G.; Li, L.; Gibson, S.B. Regulation of autophagy in hematological malignancies: Role of reactive oxygen species. Leuk Lymphoma 2012, 53, 26–33. [Google Scholar]
- Yu, L.; Alva, A.; Su, H.; Dutt, P.; Freundt, E.; Welsh, S.; Baehrecke, E.H.; Lenardo, M.J. Regulation of an ATG7-beclin 1 program of autophagic cell death by caspase-8. Science 2004, 304, 1500–1502. [Google Scholar]
- Reef, S.; Zalckvar, E.; Shifman, O.; Bialik, S.; Sabanay, H.; Oren, M.; Kimchi, A. A short mitochondrial form of p19ARF induces autophagy and caspase-independent cell death. Mol. Cell 2006, 22, 463–475. [Google Scholar]
- Cuervo, A.M. Autophagy and aging—When “all you can eat” is yourself. Sci. Aging Knowl. Environ 2003, 2003, 25. [Google Scholar]
- Codogno, P.; Meijer, A.J. Autophagy and signaling: Their role in cell survival and cell death. Cell Death Differ 2005, 12, 1509–1518. [Google Scholar]
- Yuan, H.; Perry, C.N.; Huang, C.; Iwai-Kanai, E.; Carreira, R.S.; Glembotski, C.C.; Gottlieb, R.A. LPS-induced autophagy is mediated by oxidative signaling in cardiomyocytes and is associated with cytoprotection. Am. J. Physiol. Heart Circ. Physiol 2009, 296, H470–H479. [Google Scholar]
- Chen, Y.; McMillan-Ward, E.; Kong, J.; Israels, S.J.; Gibson, S.B. Oxidative stress induces autophagic cell death independent of apoptosis in transformed and cancer cells. Cell Death Differ 2008, 15, 171–182. [Google Scholar]
- Chen, Y.; McMillan-Ward, E.; Kong, J.; Israels, S.J.; Gibson, S.B. Mitochondrial electron-transport-chain inhibitors of complexes I and II induce autophagic cell death mediated by reactive oxygen species. J. Cell Sci 2007, 120, 4155–4166. [Google Scholar]
- Chen, Y.; Azad, M.B.; Gibson, S.B. Superoxide is the major reactive oxygen species regulating autophagy. Cell Death Differ 2009, 16, 1040–1052. [Google Scholar]
- Yang, J.; Wu, L.J.; Tashino, S.; Onodera, S.; Ikejima, T. Reactive oxygen species and nitric oxide regulate mitochondria-dependent apoptosis and autophagy in evodiamine-treated human cervix carcinoma HeLa cells. Free Radic. Res 2008, 42, 492–504. [Google Scholar]
- Bolisetty, S.; Traylor, A.M.; Kim, J.; Joseph, R.; Ricart, K.; Landar, A.; Agarwal, A. Heme oxygenase-1 inhibits renal tubular macroautophagy in acute kidney injury. J. Am. Soc. Nephrol 2010, 21, 1702–1712. [Google Scholar]
- Morse, D.; Lin, L.; Choi, A.M.; Ryter, S.W. Heme oxygenase-1, a critical arbitrator of cell death pathways in lung injury and disease. Free Radic. Biol. Med 2009, 47, 1–12. [Google Scholar]
- Byun, Y.J.; Kim, S.K.; Kim, Y.M.; Chae, G.T.; Jeong, S.W.; Lee, S.B. Hydrogen peroxide induces autophagic cell death in C6 glioma cells via BNIP3-mediated suppression of the mTOR pathway. Neurosci. Lett 2009, 461, 131–135. [Google Scholar]
- Zhang, H.; Kong, X.; Kang, J.; Su, J.; Li, Y.; Zhong, J.; Sun, L. Oxidative stress induces parallel autophagy and mitochondria dysfunction in human glioma U251 cells. Toxicol. Sci 2009, 110, 376–388. [Google Scholar]
- McClung, J.M.; Judge, A.R.; Powers, S.K.; Yan, Z. p38 MAPK links oxidative stress to autophagy-related gene expression in cachectic muscle wasting. Am. J. Physiol. Cell Physiol 2010, 298, C542–549. [Google Scholar]
- Wong, C.H.; Iskandar, K.B.; Yadav, S.K.; Hirpara, J.L.; Loh, T.; Pervaiz, S. Simultaneous induction of non-canonical autophagy and apoptosis in cancer cells by ROS-dependent ERK and JNK activation. PLoS One 2010, 5, e9996. [Google Scholar]
- Wang, S.H.; Shih, Y.L.; Kuo, T.C.; Ko, W.C.; Shih, C.M. Cadmium toxicity toward autophagy through ROS-activated GSK-3beta in mesangial cells. Toxicol. Sci 2009, 108, 124–131. [Google Scholar]
- Huang, J.; Canadien, V.; Lam, G.Y.; Steinberg, B.E.; Dinauer, M.C.; Magalhaes, M.A.; Glogauer, M.; Grinstein, S.; Brumell, J.H. Activation of antibacterial autophagy by NADPH oxidases. Proc. Natl. Acad. Sci. USA 2009, 106, 6226–6231. [Google Scholar]
- Mitroulis, I.; Kourtzelis, I.; Kambas, K.; Rafail, S.; Chrysanthopoulou, A.; Speletas, M.; Ritis, K. Regulation of the autophagic machinery in human neutrophils. Eur. J. Immunol 2010, 40, 1461–1472. [Google Scholar]
- Huang, J.; Brumell, J.H. NADPH oxidases contribute to autophagy regulation. Autophagy 2009, 5, 887–889. [Google Scholar]
- Sanjuan, M.A.; Dillon, C.P.; Tait, S.W.; Moshiach, S.; Dorsey, F.; Connell, S.; Komatsu, M.; Tanaka, K.; Cleveland, J.L.; Withoff, S.; et al. Toll-like receptor signalling in macrophages links the autophagy pathway to phagocytosis. Nature 2007, 450, 1253–1257. [Google Scholar]
- Kissova, I.; Deffieu, M.; Manon, S.; Camougrand, N. Uth1p is involved in the autophagic degradation of mitochondria. J. Biol. Chem 2004, 279, 39068–39074. [Google Scholar]
- Lemasters, J.J. Selective mitochondrial autophagy, or mitophagy, as a targeted defense against oxidative stress, mitochondrial dysfunction, and aging. Rejuvenation Res 2005, 8, 3–5. [Google Scholar]
- Narendra, D.P.; Jin, S.M.; Tanaka, A.; Suen, D.F.; Gautier, C.A.; Shen, J.; Cookson, M.R.; Youle, R.J. PINK1 is selectively stabilized on impaired mitochondria to activate Parkin. PLoS Biol 2010, 8, e1000298. [Google Scholar]
- Jin, S.M.; Lazarou, M.; Wang, C.; Kane, L.A.; Narendra, D.P.; Youle, R.J. Mitochondrial membrane potential regulates PINK1 import and proteolytic destabilization by PARL. J. Cell Biol 2010, 191, 933–942. [Google Scholar]
- Narendra, D.; Tanaka, A.; Suen, D.F.; Youle, R.J. Parkin is recruited selectively to impaired mitochondria and promotes their autophagy. J. Cell Biol 2008, 183, 795–803. [Google Scholar]
- Vives-Bauza, C.; Zhou, C.; Huang, Y.; Cui, M.; de Vries, R.L.; Kim, J.; May, J.; Tocilescu, M.A.; Liu, W.; Ko, H.S.; et al. PINK1-dependent recruitment of Parkin to mitochondria in mitophagy. Proc. Natl. Acad. Sci. USA 2010, 107, 378–383. [Google Scholar]
- Matsuda, N.; Sato, S.; Shiba, K.; Okatsu, K.; Saisho, K.; Gautier, C.A.; Sou, Y.S.; Saiki, S.; Kawajiri, S.; Sato, F.; et al. PINK1 stabilized by mitochondrial depolarization recruits Parkin to damaged mitochondria and activates latent Parkin for mitophagy. J. Cell Biol 2010, 189, 211–221. [Google Scholar]
- Geisler, S.; Holmstrom, K.M.; Skujat, D.; Fiesel, F.C.; Rothfuss, O.C.; Kahle, P.J.; Springer, W. PINK1/Parkin-mediated mitophagy is dependent on VDAC1 and p62/SQSTM1. Nat. Cell Biol 2010, 12, 119–131. [Google Scholar]
- Egan, D.F.; Shackelford, D.B.; Mihaylova, M.M.; Gelino, S.; Kohnz, R.A.; Mair, W.; Vasquez, D.S.; Joshi, A.; Gwinn, D.M.; Taylor, R.; et al. Phosphorylation of ULK1 (hATG1) by AMP-activated protein kinase connects energy sensing to mitophagy. Science 2011, 331, 456–461. [Google Scholar]
- Nishida, K.; Yamaguchi, O.; Otsu, K. Crosstalk between autophagy and apoptosis in heart disease. Circ. Res 2008, 103, 343–351. [Google Scholar]
- Youle, R.J.; van der Bliek, A.M. Mitochondrial fission, fusion, and stress. Science 2012, 337, 1062–1065. [Google Scholar]
- Benard, G.; Karbowski, M. Mitochondrial fusion and division: Regulation and role in cell viability. Semin. Cell Dev. Biol 2009, 20, 365–374. [Google Scholar]
- Pletjushkina, O.Y.; Lyamzaev, K.G.; Popova, E.N.; Nepryakhina, O.K.; Ivanova, O.Y.; Domnina, L.V.; Chernyak, B.V.; Skulachev, V.P. Effect of oxidative stress on dynamics of mitochondrial reticulum. Biochim. Biophys. Acta 2006, 1757, 518–524. [Google Scholar]
- Fan, X.; Hussien, R.; Brooks, G.A. H2O2-induced mitochondrial fragmentation in C2C12 myocytes. Free Radic. Biol. Med 2010, 49, 1646–1654. [Google Scholar]
- Jendrach, M.; Mai, S.; Pohl, S.; Voth, M.; Bereiter-Hahn, J. Short- and long-term alterations of mitochondrial morphology, dynamics and mtDNA after transient oxidative stress. Mitochondrion 2008, 8, 293–304. [Google Scholar]
- Makino, A.; Scott, B.T.; Dillmann, W.H. Mitochondrial fragmentation and superoxide anion production in coronary endothelial cells from a mouse model of type 1 diabetes. Diabetologia 2010, 53, 1783–1794. [Google Scholar]
- Barsoum, M.J.; Yuan, H.; Gerencser, A.A.; Liot, G.; Kushnareva, Y.; Graber, S.; Kovacs, I.; Lee, W.D.; Waggoner, J.; Cui, J.; et al. Nitric oxide-induced mitochondrial fission is regulated by dynamin-related GTPases in neurons. EMBO J 2006, 25, 3900–3911. [Google Scholar]
- Liot, G.; Bossy, B.; Lubitz, S.; Kushnareva, Y.; Sejbuk, N.; Bossy-Wetzel, E. Complex II inhibition by 3-NP causes mitochondrial fragmentation and neuronal cell death via an NMDA- and ROS-dependent pathway. Cell Death Differ 2009, 16, 899–909. [Google Scholar]
- Brooks, C.; Wei, Q.; Cho, S.G.; Dong, Z. Regulation of mitochondrial dynamics in acute kidney injury in cell culture and rodent models. J. Clin. Invest 2009, 119, 1275–1285. [Google Scholar]
- Funk, J.A.; Schnellmann, R.G. Persistent disruption of mitochondrial homeostasis after acute kidney injury. Am. J. Physiol. Renal Physiol 2012, 302, F853–F864. [Google Scholar]
- Yu, T.; Sheu, S.S.; Robotham, J.L.; Yoon, Y. Mitochondrial fission mediates high glucose-induced cell death through elevated production of reactive oxygen species. Cardiovasc. Res 2008, 79, 341–351. [Google Scholar]
- Giedt, R.J.; Yang, C.; Zweier, J.L.; Matzavinos, A.; Alevriadou, B.R. Mitochondrial fission in endothelial cells after simulated ischemia/reperfusion: Role of nitric oxide and reactive oxygen species. Free Radic. Biol. Med 2012, 52, 348–356. [Google Scholar]
- Solesio, M.E.; Prime, T.A.; Logan, A.; Murphy, M.P.; Del Mar Arroyo-Jimenez, M.; Jordan, J.; Galindo, M.F. The mitochondria-targeted anti-oxidant MitoQ reduces aspects of mitochondrial fission in the 6-OHDA cell model of Parkinson’s disease. Biochim. Biophys. Acta 2012, 1832, 174–182. [Google Scholar]
- Estaquier, J.; Arnoult, D. Inhibiting Drp1-mediated mitochondrial fission selectively prevents the release of cytochrome c during apoptosis. Cell Death Differ 2007, 14, 1086–1094. [Google Scholar]
- Frank, S.; Gaume, B.; Bergmann-Leitner, E.S.; Leitner, W.W.; Robert, E.G.; Catez, F.; Smith, C.L.; Youle, R.J. The role of dynamin-related protein 1, a mediator of mitochondrial fission, in apoptosis. Dev. Cell 2001, 1, 515–525. [Google Scholar]
- Lee, Y.J.; Jeong, S.Y.; Karbowski, M.; Smith, C.L.; Youle, R.J. Roles of the mammalian mitochondrial fission and fusion mediators Fis1, Drp1, and Opa1 in apoptosis. Mol. Biol. Cell 2004, 15, 5001–5011. [Google Scholar]
- Sugioka, R.; Shimizu, S.; Tsujimoto, Y. Fzo1, a protein involved in mitochondrial fusion, inhibits apoptosis. J. Biol. Chem 2004, 279, 52726–52734. [Google Scholar]
- James, D.I.; Parone, P.A.; Mattenberger, Y.; Martinou, J.C. hFis1, a novel component of the mammalian mitochondrial fission machinery. J. Biol. Chem 2003, 278, 36373–36379. [Google Scholar]
- Olichon, A.; Baricault, L.; Gas, N.; Guillou, E.; Valette, A.; Belenguer, P.; Lenaers, G. Loss of OPA1 perturbates the mitochondrial inner membrane structure and integrity, leading to cytochrome c release and apoptosis. J. Biol. Chem 2003, 278, 7743–7746. [Google Scholar]
- Parone, P.A.; Da Cruz, S.; Tondera, D.; Mattenberger, Y.; James, D.I.; Maechler, P.; Barja, F.; Martinou, J.C. Preventing mitochondrial fission impairs mitochondrial function and leads to loss of mitochondrial DNA. PLoS One 2008, 3, e3257. [Google Scholar]
- Benard, G.; Bellance, N.; James, D.; Parrone, P.; Fernandez, H.; Letellier, T.; Rossignol, R. Mitochondrial bioenergetics and structural network organization. J. Cell Sci 2007, 120, 838–848. [Google Scholar]
- Frank, M.; Duvezin-Caubet, S.; Koob, S.; Occhipinti, A.; Jagasia, R.; Petcherski, A.; Ruonala, M.O.; Priault, M.; Salin, B.; Reichert, A.S. Mitophagy is triggered by mild oxidative stress in a mitochondrial fission dependent manner. Biochim. Biophys. Acta 2012, 1823, 2297–2310. [Google Scholar]
- Park, J.; Lee, J.; Choi, C. Mitochondrial network determines intracellular ROS dynamics and sensitivity to oxidative stress through switching inter-mitochondrial messengers. PLoS One 2011, 6, e23211. [Google Scholar]
- Jou, M.J. Pathophysiological and pharmacological implications of mitochondria-targeted reactive oxygen species generation in astrocytes. Adv. Drug Deliv. Rev 2008, 60, 1512–1526. [Google Scholar]
- Yoon, Y.; Galloway, C.A.; Jhun, B.S.; Yu, T. Mitochondrial dynamics in diabetes. Antioxid. Redox Signal 2011, 14, 439–457. [Google Scholar]
- Duncan, J.G. Mitochondrial dysfunction in diabetic cardiomyopathy. Biochim. Biophys. Acta 2011, 1813, 1351–1359. [Google Scholar]
- Ong, S.B.; Hausenloy, D.J. Mitochondrial morphology and cardiovascular disease. Cardiovasc. Res 2010, 88, 16–29. [Google Scholar]
- Romanello, V.; Guadagnin, E.; Gomes, L.; Roder, I.; Sandri, C.; Petersen, Y.; Milan, G.; Masiero, E.; del Piccolo, P.; Foretz, M.; et al. Mitochondrial fission and remodelling contributes to muscle atrophy. EMBO J 2010, 29, 1774–1785. [Google Scholar]
- Ong, S.B.; Subrayan, S.; Lim, S.Y.; Yellon, D.M.; Davidson, S.M.; Hausenloy, D.J. Inhibiting mitochondrial fission protects the heart against ischemia/reperfusion injury. Circulation 2010, 121, 2012–2022. [Google Scholar]
- Chen, L.; Gong, Q.; Stice, J.P.; Knowlton, A.A. Mitochondrial OPA1, apoptosis, and heart failure. Cardiovasc. Res 2009, 84, 91–99. [Google Scholar]
- Nakamura, K.; Nemani, V.M.; Azarbal, F.; Skibinski, G.; Levy, J.M.; Egami, K.; Munishkina, L.; Zhang, J.; Gardner, B.; Wakabayashi, J.; et al. Direct membrane association drives mitochondrial fission by the Parkinson disease-associated protein alpha-synuclein. J. Biol. Chem 2011, 286, 20710–20726. [Google Scholar]
- Wang, X.; Su, B.; Lee, H.G.; Li, X.; Perry, G.; Smith, M.A.; Zhu, X. Impaired balance of mitochondrial fission and fusion in Alzheimer’s disease. J. Neurosci 2009, 29, 9090–9103. [Google Scholar]
- Babior, B.M.; Kipnes, R.S.; Curnutte, J.T. Biological defense mechanisms. The production by leukocytes of superoxide, a potential bactericidal agent. J. Clin. Invest 1973, 52, 741–744. [Google Scholar]
- Babior, B.M.; Peters, W.A. The O2-producing enzyme of human neutrophils. Further properties. J. Biol. Chem 1981, 256, 2321–2323. [Google Scholar]
- Palmer, R.M.; Ferrige, A.G.; Moncada, S. Nitric oxide release accounts for the biological activity of endothelium-derived relaxing factor. Nature 1987, 327, 524–526. [Google Scholar]
- Gruetter, C.A.; Barry, B.K.; McNamara, D.B.; Gruetter, D.Y.; Kadowitz, P.J.; Ignarro, L. Relaxation of bovine coronary artery and activation of coronary arterial guanylate cyclase by nitric oxide, nitroprusside and a carcinogenic nitrosoamine. J. Cyclic Nucleotide Res 1979, 5, 211–224. [Google Scholar]
- Furchgott, R.F.; Carvalho, M.H.; Khan, M.T.; Matsunaga, K. Evidence for endothelium-dependent vasodilation of resistance vessels by acetylcholine. Blood Vessels 1987, 24, 145–149. [Google Scholar]
- Leclercq, B.; Jaimes, E.A.; Raij, L. Nitric oxide synthase and hypertension. Curr. Opin. Nephrol. Hypertens 2002, 11, 185–189. [Google Scholar]
- Tabima, D.M.; Frizzell, S.; Gladwin, M.T. Reactive oxygen and nitrogen species in pulmonary hypertension. Free Radic. Biol. Med 2012, 52, 1970–1986. [Google Scholar]
- Tousoulis, D.; Kampoli, A.M.; Tentolouris, C.; Papageorgiou, N.; Stefanadis, C. The role of nitric oxide on endothelial function. Curr. Vasc. Pharmacol 2012, 10, 4–18. [Google Scholar]
- Jaimes, E.A.; Hua, P.; Tian, R.X.; Raij, L. Human glomerular endothelium: Interplay among glucose, free fatty acids, angiotensin II, and oxidative stress. Am. J. Physiol. Renal Physiol 2010, 298, F125–F132. [Google Scholar]
- Radomski, M.W.; Moncada, S. Regulation of vascular homeostasis by nitric oxide. Thromb Haemost 1993, 70, 36–41. [Google Scholar]
- Frazziano, G.; Champion, H.C.; Pagano, P.J. NADPH oxidase-derived ROS and the regulation of pulmonary vessel tone. Am. J. Physiol. Heart Circ. Physiol 2012, 302, H2166–H2177. [Google Scholar]
- Satoh, K.; Berk, B.C.; Shimokawa, H. Vascular-derived reactive oxygen species for homeostasis and diseases. Nitric Oxide 2011, 25, 211–215. [Google Scholar]
- Erusalimsky, J.D.; Moncada, S. Nitric oxide and mitochondrial signaling: From physiology to pathophysiology. Arterioscler. Thromb. Vasc. Biol 2007, 27, 2524–2531. [Google Scholar]
- Napoli, C.; Ignarro, L.J. Nitric oxide and pathogenic mechanisms involved in the development of vascular diseases. Arch. Pharm. Res 2009, 32, 1103–1108. [Google Scholar]
- Zhou, M.S.; Jaimes, E.A.; Raij, L. Vascular but not cardiac remodeling is associated with superoxide production in angiotensin II hypertension. J. Hypertens 2005, 23, 1737–1743. [Google Scholar]
- Rigoulet, M.; Yoboue, E.D.; Devin, A. Mitochondrial ROS generation and its regulation: Mechanisms involved in H2O2 signaling. Antioxid. Redox Signal 2011, 14, 459–468. [Google Scholar]
- Groeger, G.; Quiney, C.; Cotter, T.G. Hydrogen peroxide as a cell-survival signaling molecule. Antioxid. Redox Signal 2009, 11, 2655–2671. [Google Scholar]
- Miller, A.A.; Drummond, G.R.; Sobey, C.G. Reactive oxygen species in the cerebral circulation: Are they all bad? Antioxid. Redox Signal 2006, 8, 1113–1120. [Google Scholar]
- Kamsler, A.; Segal, M. Hydrogen peroxide as a diffusible signal molecule in synaptic plasticity. Mol. NeuroBiol 2004, 29, 167–178. [Google Scholar]
- Kochevar, I.E. Singlet oxygen signaling: From intimate to global. Sci. STKE 2004, 2004, pe7. [Google Scholar]
- Turpaev, K.T. Reactive oxygen species and regulation of gene expression. Biochemistry (Mosc. ) 2002, 67, 281–292. [Google Scholar]
- Kim, P.K.; Zamora, R.; Petrosko, P.; Billiar, T.R. The regulatory role of nitric oxide in apoptosis. Int. Immunopharmacol 2001, 1, 1421–1441. [Google Scholar]
- Martin, L.D.; Krunkosky, T.M.; Voynow, J.A.; Adler, K.B. The role of reactive oxygen and nitrogen species in airway epithelial gene expression. Environ. Health Perspect 1998, 106, 1197–1203. [Google Scholar]
- Ho, J.J.; Man, H.S.; Marsden, P.A. Nitric oxide signaling in hypoxia. J. Mol. Med. (Berl. ) 2012, 90, 217–231. [Google Scholar]
- Martinez-Ruiz, A.; Cadenas, S.; Lamas, S. Nitric oxide signaling: Classical, less classical, and nonclassical mechanisms. Free Radic. Biol. Med 2011, 51, 17–29. [Google Scholar]
- Bryan, N.S.; Bian, K.; Murad, F. Discovery of the nitric oxide signaling pathway and targets for drug development. Front. Biosci 2009, 14, 1–18. [Google Scholar]
- Afanas’ev, I.B. Signaling functions of free radicals superoxide & nitric oxide under physiological & pathological conditions. Mol. Biotechnol 2007, 37, 2–4. [Google Scholar]
- Buetler, T.M.; Krauskopf, A.; Ruegg, U.T. Role of superoxide as a signaling molecule. News Physiol. Sci 2004, 19, 120–123. [Google Scholar]
- Veal, E.; Day, A. Hydrogen peroxide as a signaling molecule. Antioxid. Redox Signal 2011, 15, 147–151. [Google Scholar]
- Stone, J.R.; Yang, S. Hydrogen peroxide: A signaling messenger. Antioxid. Redox Signal 2006, 8, 243–270. [Google Scholar]
- Rhee, S.G. Redox signaling: Hydrogen peroxide as intracellular messenger. Exp. Mol. Med 1999, 31, 53–59. [Google Scholar]
- Lander, H.M. An essential role for free radicals and derived species in signal transduction. FASEB J 1997, 11, 118–124. [Google Scholar]
- Hensley, K.; Robinson, K.A.; Gabbita, S.P.; Salsman, S.; Floyd, R.A. Reactive oxygen species, cell signaling, and cell injury. Free Radic. Biol. Med 2000, 28, 1456–1462. [Google Scholar]
- Chen, X.L.; Kunsch, C. Induction of cytoprotective genes through Nrf2/antioxidant response element pathway: A new therapeutic approach for the treatment of inflammatory diseases. Curr. Pharm. Des 2004, 10, 879–891. [Google Scholar]
- Itoh, K.; Chiba, T.; Takahashi, S.; Ishii, T.; Igarashi, K.; Katoh, Y.; Oyake, T.; Hayashi, N.; Satoh, K.; Hatayama, I.; et al. An Nrf2/small Maf heterodimer mediates the induction of phase II detoxifying enzyme genes through antioxidant response elements. Biochem. Biophys. Res. Commun 1997, 236, 313–322. [Google Scholar]
- Thimmulappa, R.K.; Mai, K.H.; Srisuma, S.; Kensler, T.W.; Yamamoto, M.; Biswal, S. Identification of Nrf2-regulated genes induced by the chemopreventive agent sulforaphane by oligonucleotide microarray. Cancer Res 2002, 62, 5196–5203. [Google Scholar]
- Chorley, B.N.; Campbell, M.R.; Wang, X.; Karaca, M.; Sambandan, D.; Bangura, F.; Xue, P.; Pi, J.; Kleeberger, S.R.; Bell, D.A. Identification of novel NRF2-regulated genes by ChIP-Seq: Influence on retinoid X receptor alpha. Nucleic Acids Res 2012, 40, 7416–7429. [Google Scholar]
- Taylor, R.C.; Acquaah-Mensah, G.; Singhal, M.; Malhotra, D.; Biswal, S. Network inference algorithms elucidate Nrf2 regulation of mouse lung oxidative stress. PLoS Comput. Biol 2008, 4, e1000166. [Google Scholar]
- Numazawa, S.; Yoshida, T. Nrf2-dependent gene expressions: A molecular toxicological aspect. J. Toxicol. Sci 2004, 29, 81–89. [Google Scholar]
- Reddy, N.M.; Kleeberger, S.R.; Yamamoto, M.; Kensler, T.W.; Scollick, C.; Biswal, S.; Reddy, S.P. Genetic dissection of the Nrf2-dependent redox signaling-regulated transcriptional programs of cell proliferation and cytoprotection. Physiol. Genomics 2007, 32, 74–81. [Google Scholar]
- Cheng, X.; Siow, R.C.; Mann, G.E. Impaired redox signaling and antioxidant gene expression in endothelial cells in diabetes: A role for mitochondria and the nuclear factor-E2-related factor 2-Kelch-like ECH-associated protein 1 defense pathway. Antioxid. Redox Signal 2011, 14, 469–487. [Google Scholar]
- Van Muiswinkel, F.L.; Kuiperij, H.B. The Nrf2-ARE Signalling pathway: Promising drug target to combat oxidative stress in neurodegenerative disorders. Curr. Drug Targets CNS Neurol. Disord 2005, 4, 267–281. [Google Scholar]
- Kaspar, J.W.; Niture, S.K.; Jaiswal, A.K. Nrf2:INrf2 (Keap1) signaling in oxidative stress. Free Radic. Biol. Med 2009, 47, 1304–1309. [Google Scholar]
- Apopa, P.L.; He, X.; Ma, Q. Phosphorylation of Nrf2 in the transcription activation domain by casein kinase 2 (CK2) is critical for the nuclear translocation and transcription activation function of Nrf2 in IMR-32 neuroblastoma cells. J. Biochem. Mol. Toxicol 2008, 22, 63–76. [Google Scholar]
- Numazawa, S.; Ishikawa, M.; Yoshida, A.; Tanaka, S.; Yoshida, T. Atypical protein kinase C mediates activation of NF-E2-related factor 2 in response to oxidative stress. Am. J. Physiol. Cell Physiol 2003, 285, C334–C342. [Google Scholar]
- Itoh, K.; Wakabayashi, N.; Katoh, Y.; Ishii, T.; Igarashi, K.; Engel, J.D.; Yamamoto, M. Keap1 represses nuclear activation of antioxidant responsive elements by Nrf2 through binding to the amino-terminal Neh2 domain. Genes Dev 1999, 13, 76–86. [Google Scholar]
- Dinkova-Kostova, A.T.; Holtzclaw, W.D.; Kensler, T.W. The role of Keap1 in cellular protective responses. Chem. Res. Toxicol 2005, 18, 1779–1791. [Google Scholar]
- Kobayashi, A.; Kang, M.I.; Okawa, H.; Ohtsuji, M.; Zenke, Y.; Chiba, T.; Igarashi, K.; Yamamoto, M. Oxidative stress sensor Keap1 functions as an adaptor for Cul3-based E3 ligase to regulate proteasomal degradation of Nrf2. Mol. Cell. Biol 2004, 24, 7130–7139. [Google Scholar]
- Zhang, D.D.; Lo, S.C.; Cross, J.V.; Templeton, D.J.; Hannink, M. Keap1 is a redox-regulated substrate adaptor protein for a Cul3-dependent ubiquitin ligase complex. Mol. Cell. Biol 2004, 24, 10941–10953. [Google Scholar]
- Zhang, D.D. Mechanistic studies of the Nrf2-Keap1 signaling pathway. Drug Metab Rev 2006, 38, 769–789. [Google Scholar]
- Tong, K.I.; Kobayashi, A.; Katsuoka, F.; Yamamoto, M. Two-site substrate recognition model for the Keap1-Nrf2 system: A hinge and latch mechanism. Biol. Chem 2006, 387, 1311–1320. [Google Scholar]
- Kensler, T.W.; Wakabayashi, N.; Biswal, S. Cell survival responses to environmental stresses via the Keap1-Nrf2-ARE pathway. Annu. Rev. Pharmacol. Toxicol 2007, 47, 89–116. [Google Scholar]
- Villeneuve, N.F.; Lau, A.; Zhang, D.D. Regulation of the Nrf2-Keap1 antioxidant response by the ubiquitin proteasome system: An insight into cullin-ring ubiquitin ligases. Antioxid. Redox Signal 2010, 13, 1699–1712. [Google Scholar]
- Itoh, K.; Mimura, J.; Yamamoto, M. Discovery of the negative regulator of Nrf2, Keap1: A historical overview. Antioxid. Redox Signal 2010, 13, 1665–1678. [Google Scholar]
- Baird, L.; Dinkova-Kostova, A.T. The cytoprotective role of the Keap1-Nrf2 pathway. Arch. Toxicol 2011, 85, 241–272. [Google Scholar]
- Tkachev, V.O.; Menshchikova, E.B.; Zenkov, N.K. Mechanism of the Nrf2/Keap1/ARE signaling system. Biochemistry (Mosc. ) 2011, 76, 407–422. [Google Scholar]
- Dhakshinamoorthy, S.; Porter, A.G. Nitric oxide-induced transcriptional up-regulation of protective genes by Nrf2 via the antioxidant response element counteracts apoptosis of neuroblastoma cells. J. Biol. Chem 2004, 279, 20096–20107. [Google Scholar]
- Wilson, L.A.; Gemin, A.; Espiritu, R.; Singh, G. ets-1 is transcriptionally up-regulated by H2O2 via an antioxidant response element. FASEB J 2005, 19, 2085–2087. [Google Scholar]
- Ashino, T.; Yamanaka, R.; Yamamoto, M.; Shimokawa, H.; Sekikawa, K.; Iwakura, Y.; Shioda, S.; Numazawa, S.; Yoshida, T. Negative feedback regulation of lipopolysaccharide-induced inducible nitric oxide synthase gene expression by heme oxygenase-1 induction in macrophages. Mol. Immunol 2008, 45, 2106–2115. [Google Scholar]
- Buckley, B.J.; Marshall, Z.M.; Whorton, A.R. Nitric oxide stimulates Nrf2 nuclear translocation in vascular endothelium. Biochem. Biophys. Res. Commun 2003, 307, 973–979. [Google Scholar]
- Fourquet, S.; Guerois, R.; Biard, D.; Toledano, M.B. Activation of NRF2 by nitrosative agents and H2O2 involves KEAP1 disulfide formation. J. Biol. Chem 2010, 285, 8463–8471. [Google Scholar]
- Motohashi, H.; Yamamoto, M. Nrf2-Keap1 defines a physiologically important stress response mechanism. Trends Mol. Med 2004, 10, 549–557. [Google Scholar]
- Sen, R.; Baltimore, D. Multiple nuclear factors interact with the immunoglobulin enhancer sequences. Cell 1986, 46, 705–716. [Google Scholar]
- Ghosh, S.; May, M.J.; Kopp, E.B. NF-kappa B and Rel proteins: Evolutionarily conserved mediators of immune responses. Annu. Rev. Immunol 1998, 16, 225–260. [Google Scholar]
- Farrow, S.N. Death receptors, NF-kappa B activation and apoptosis: The potential for therapeutic intervention. Biochem. Soc. Trans 1999, 27, 812–814. [Google Scholar]
- Van Antwerp, D.J.; Martin, S.J.; Verma, I.M.; Green, D.R. Inhibition of TNF-induced apoptosis by NF-kappa B. Trends Cell Biol 1998, 8, 107–111. [Google Scholar]
- Sonenshein, G.E. Rel/NF-kappa B transcription factors and the control of apoptosis. Semin. Cancer Biol 1997, 8, 113–119. [Google Scholar]
- Francis, D.A.; Sen, R.; Rothstein, T.L. Receptor-specific regulation of NF-kappa B, c-Myc and Fas-mediated apoptosis in primary B cells. Curr. Top. MicroBiol. Immunol 1997, 224, 83–90. [Google Scholar]
- Wang, H.; Cho, C.H. Effect of NF-kappaB signaling on apoptosis in chronic inflammation-associated carcinogenesis. Curr. Cancer Drug Targets 2010, 10, 593–599. [Google Scholar]
- Piotrowska, A.; Izykowska, I.; Podhorska-Okolow, M.; Zabel, M.; Dziegiel, P. The structure of NF-kappaB family proteins and their role in apoptosis. Postepy Hig. Med. Dosw. (Online) 2008, 62, 64–74. [Google Scholar]
- Sen, R. Control of B lymphocyte apoptosis by the transcription factor NF-kappaB. Immunity 2006, 25, 871–883. [Google Scholar]
- Kucharczak, J.; Simmons, M.J.; Fan, Y.; Gelinas, C. To be, or not to be: NF-kappaB is the answer—Role of Rel/NF-kappaB in the regulation of apoptosis. Oncogene 2003, 22, 8961–8982. [Google Scholar]
- Barkett, M.; Gilmore, T.D. Control of apoptosis by Rel/NF-kappaB transcription factors. Oncogene 1999, 18, 6910–6924. [Google Scholar]
- DiDonato, J.A.; Mercurio, F.; Karin, M. NF-kappaB and the link between inflammation and cancer. Immunol. Rev 2012, 246, 379–400. [Google Scholar]
- Ben-Neriah, Y.; Karin, M. Inflammation meets cancer, with NF-kappaB as the matchmaker. Nat. Immunol 2011, 12, 715–723. [Google Scholar]
- Baker, R.G.; Hayden, M.S.; Ghosh, S. NF-kappaB, inflammation, and metabolic disease. Cell Metab 2011, 13, 11–22. [Google Scholar]
- Sanz, A.B.; Sanchez-Nino, M.D.; Ramos, A.M.; Moreno, J.A.; Santamaria, B.; Ruiz-Ortega, M.; Egido, J.; Ortiz, A. NF-kappaB in renal inflammation. J. Am. Soc. Nephrol 2010, 21, 1254–1262. [Google Scholar]
- Lawrence, T. The nuclear factor NF-kappaB pathway in inflammation. Cold Spring Harb. Perspect. Biol 2009, 1, a001651. [Google Scholar]
- Lawrence, T.; Fong, C. The resolution of inflammation: Anti-inflammatory roles for NF-kappaB. Int. J. Biochem. Cell Biol 2010, 42, 519–523. [Google Scholar]
- Gerondakis, S.; Grumont, R.; Gugasyan, R.; Wong, L.; Isomura, I.; Ho, W.; Banerjee, A. Unravelling the complexities of the NF-kappaB signalling pathway using mouse knockout and transgenic models. Oncogene 2006, 25, 6781–6799. [Google Scholar]
- Monaco, C.; Andreakos, E.; Kiriakidis, S.; Mauri, C.; Bicknell, C.; Foxwell, B.; Cheshire, N.; Paleolog, E.; Feldmann, M. Canonical pathway of nuclear factor kappa B activation selectively regulates proinflammatory and prothrombotic responses in human atherosclerosis. Proc. Natl. Acad. Sci. USA 2004, 101, 5634–5639. [Google Scholar]
- Song, X.Q.; Lv, L.X.; Li, W.Q.; Hao, Y.H.; Zhao, J.P. The interaction of nuclear factor-kappa B and cytokines is associated with schizophrenia. Biol. Psychiatry 2009, 65, 481–488. [Google Scholar]
- Yamamoto, Y.; Gaynor, R.B. Therapeutic potential of inhibition of the NF-kappaB pathway in the treatment of inflammation and cancer. J. Clin. Invest 2001, 107, 135–142. [Google Scholar]
- Yamamoto, Y.; Gaynor, R.B. Role of the NF-kappaB pathway in the pathogenesis of human disease states. Curr. Mol. Med 2001, 1, 287–296. [Google Scholar]
- Hoffmann, A.; Natoli, G.; Ghosh, G. Transcriptional regulation via the NF-kappaB signaling module. Oncogene 2006, 25, 6706–6716. [Google Scholar]
- Yamamoto, Y.; Gaynor, R.B. IkappaB kinases: Key regulators of the NF-kappaB pathway. Trends Biochem. Sci 2004, 29, 72–79. [Google Scholar]
- Hansen, S.K.; Guerrini, L.; Blasi, F. Differential DNA sequence specificity and regulation of HIV-1 enhancer activity by cRel-RelA transcription factor. J. Biol. Chem 1994, 269, 22230–22237. [Google Scholar]
- Brown, A.M.; Linhoff, M.W.; Stein, B.; Wright, K.L.; Baldwin, A.S., Jr; Basta, P.V.; Ting, J.P. Function of NF-kappa B/Rel binding sites in the major histocompatibility complex class II invariant chain promoter is dependent on cell-specific binding of different NF-kappa B/Rel subunits. Mol. Cell Biol. 1994, 14, 2926–2935. [Google Scholar]
- Plaksin, D.; Baeuerle, P.A.; Eisenbach, L. KBF1 (p50 NF-kappa B homodimer) acts as a repressor of H-2Kb gene expression in metastatic tumor cells. J. Exp. Med 1993, 177, 1651–1662. [Google Scholar]
- Kone, B.C.; Schwobel, J.; Turner, P.; Mohaupt, M.G.; Cangro, C.B. Role of NF-kappa B in the regulation of inducible nitric oxide synthase in an MTAL cell line. Am. J. Physiol 1995, 269, F718–F729. [Google Scholar]
- Pahan, K.; Sheikh, F.G.; Liu, X.; Hilger, S.; McKinney, M.; Petro, T.M. Induction of nitric-oxide synthase and activation of NF-kappaB by interleukin-12 p40 in microglial cells. J. Biol. Chem 2001, 276, 7899–7905. [Google Scholar]
- Fouad, D.; Siendones, E.; Costan, G.; Muntane, J. Role of NF-kappaB activation and nitric oxide expression during PGE protection against d-galactosamine-induced cell death in cultured rat hepatocytes. Liver Int 2004, 24, 227–236. [Google Scholar]
- Griscavage, J.M.; Wilk, S.; Ignarro, L.J. Inhibitors of the proteasome pathway interfere with induction of nitric oxide synthase in macrophages by blocking activation of transcription factor NF-kappa B. Proc. Natl. Acad. Sci. USA 1996, 93, 3308–3312. [Google Scholar]
- Lee, K.M.; Kang, B.S.; Lee, H.L.; Son, S.J.; Hwang, S.H.; Kim, D.S.; Park, J.S.; Cho, H.J. Spinal NF-kB activation induces COX-2 upregulation and contributes to inflammatory pain hypersensitivity. Eur. J. NeuroSci 2004, 19, 3375–3381. [Google Scholar]
- Schmedtje, J.F., Jr; Ji, Y.S.; Liu, W.L.; DuBois, R.N.; Runge, M.S. Hypoxia induces cyclooxygenase-2 via the NF-kappaB p65 transcription factor in human vascular endothelial cells. J. Biol. Chem. 1997, 272, 601–608. [Google Scholar]
- Jobin, C.; Morteau, O.; Han, D.S.; Balfour Sartor, R. Specific NF-kappaB blockade selectively inhibits tumour necrosis factor-alpha-induced COX-2 but not constitutive COX-1 gene expression in HT-29 cells. Immunology 1998, 95, 537–543. [Google Scholar]
- Schreck, R.; Rieber, P.; Baeuerle, P.A. Reactive oxygen intermediates as apparently widely used messengers in the activation of the NF-kappa B transcription factor and HIV-1. EMBO J 1991, 10, 2247–2258. [Google Scholar]
- Schreck, R.; Meier, B.; Mannel, D.N.; Droge, W.; Baeuerle, P.A. Dithiocarbamates as potent inhibitors of nuclear factor kappa B activation in intact cells. J. Exp. Med 1992, 175, 1181–1194. [Google Scholar]
- Schreck, R.; Grassmann, R.; Fleckenstein, B.; Baeuerle, P.A. Antioxidants selectively suppress activation of NF-kappa B by human T-cell leukemia virus type I Tax protein. J. Virol 1992, 66, 6288–6293. [Google Scholar]
- Lander, H.M.; Sehajpal, P.; Levine, D.M.; Novogrodsky, A. Activation of human peripheral blood mononuclear cells by nitric oxide-generating compounds. J. Immunol 1993, 150, 1509–1516. [Google Scholar]
- Shibanuma, M.; Kuroki, T.; Nose, K. Inhibition by N-acetyl-l-cysteine of interleukin-6 mRNA induction and activation of NF kappa B by tumor necrosis factor alpha in a mouse fibroblastic cell line, Balb/3T3. FEBS Lett 1994, 353, 62–66. [Google Scholar]
- Manna, S.K.; Zhang, H.J.; Yan, T.; Oberley, L.W.; Aggarwal, B.B. Overexpression of manganese superoxide dismutase suppresses tumor necrosis factor-induced apoptosis and activation of nuclear transcription factor-kappaB and activated protein-1. J. Biol. Chem 1998, 273, 13245–13254. [Google Scholar]
- Meyer, M.; Schreck, R.; Baeuerle, P.A. H2O2 and antioxidants have opposite effects on activation of NF-kappa B and AP-1 in intact cells: AP-1 as secondary antioxidant-responsive factor. EMBO J 1993, 12, 2005–2015. [Google Scholar]
- Kim, S.J.; Hwang, S.G.; Shin, D.Y.; Kang, S.S.; Chun, J.S. p38 kinase regulates nitric oxide-induced apoptosis of articular chondrocytes by accumulating p53 via NFkappa B-dependent transcription and stabilization by serine 15 phosphorylation. J. Biol. Chem 2002, 277, 33501–33508. [Google Scholar]
- Lee, S.; Chung, J.; Ha, I.S.; Yi, K.; Lee, J.E.; Kang, H.G.; Choi, I.; Oh, K.H.; Kim, J.Y.; Surh, C.D.; et al. Hydrogen peroxide increases human leukocyte adhesion to porcine aortic endothelial cells via NFkappaB-dependent up-regulation of VCAM-1. Int. Immunol 2007, 19, 1349–1359. [Google Scholar]
- Zhang, J.; Johnston, G.; Stebler, B.; Keller, E.T. Hydrogen peroxide activates NFkappaB and the interleukin-6 promoter through NFkappaB-inducing kinase. Antioxid. Redox Signal 2001, 3, 493–504. [Google Scholar]
- Waldow, T.; Witt, W.; Weber, E.; Matschke, K. Nitric oxide donor-induced persistent inhibition of cell adhesion protein expression and NFkappaB activation in endothelial cells. Nitric Oxide 2006, 15, 103–113. [Google Scholar]
- Nam, H.Y.; Choi, B.H.; Lee, J.Y.; Lee, S.G.; Kim, Y.H.; Lee, K.H.; Yoon, H.K.; Song, J.S.; Kim, H.J.; Lim, Y. The role of nitric oxide in the particulate matter (PM2.5)-induced NFkappaB activation in lung epithelial cells. Toxicol. Lett 2004, 148, 95–102. [Google Scholar]
- Tsai, S.H.; Lin-Shiau, S.Y.; Lin, J.K. Suppression of nitric oxide synthase and the down-regulation of the activation of NFkappaB in macrophages by resveratrol. Br. J. Pharmacol 1999, 126, 673–680. [Google Scholar]
- Schreck, R.; Albermann, K.; Baeuerle, P.A. Nuclear factor kappa B: An oxidative stress-responsive transcription factor of eukaryotic cells (a review). Free Radic. Res. Commun 1992, 17, 221–237. [Google Scholar]
- Packer, L.; Suzuki, Y.J. Vitamin E and alpha-lipoate: Role in antioxidant recycling and activation of the NF-kappa B transcription factor. Mol. Aspects Med 1993, 14, 229–239. [Google Scholar]
- Suzuki, Y.J.; Packer, L. Inhibition of NF-kappa B DNA binding activity by alpha-tocopheryl succinate. Biochem. Mol. Biol. Int 1993, 31, 693–700. [Google Scholar]
- Kretz-Remy, C.; Mehlen, P.; Mirault, M.E.; Arrigo, A.P. Inhibition of I kappa B-alpha phosphorylation and degradation and subsequent NF-kappa B activation by glutathione peroxidase overexpression. J. Cell Biol 1996, 133, 1083–1093. [Google Scholar]
- Aravindan, N.; Mohan, S.; Herman, T.S.; Natarajan, M. Nitric oxide-mediated inhibition of NFkappaB regulates hyperthermia-induced apoptosis. J. Cell Biochem 2009, 106, 999–1009. [Google Scholar]
- Gilad, E.; Wong, H.R.; Zingarelli, B.; Virag, L.; O’Connor, M.; Salzman, A.L.; Szabo, C. Melatonin inhibits expression of the inducible isoform of nitric oxide synthase in murine macrophages: Role of inhibition of NFkappaB activation. FASEB J 1998, 12, 685–693. [Google Scholar]
- Matthews, J.R.; Botting, C.H.; Panico, M.; Morris, H.R.; Hay, R.T. Inhibition of NF-kappaB DNA binding by nitric oxide. Nucleic Acids Res 1996, 24, 2236–2242. [Google Scholar]
- Peng, H.B.; Libby, P.; Liao, J.K. Induction and stabilization of I kappa B alpha by nitric oxide mediates inhibition of NF-kappa B. J. Biol. Chem 1995, 270, 14214–14219. [Google Scholar]
- Taylor, B.S.; Kim, Y.M.; Wang, Q.; Shapiro, R.A.; Billiar, T.R.; Geller, D.A. Nitric oxide down-regulates hepatocyte-inducible nitric oxide synthase gene expression. Arch. Surg 1997, 132, 1177–1183. [Google Scholar]
- Sekkai, D.; Aillet, F.; Israel, N.; Lepoivre, M. Inhibition of NF-kappaB and HIV-1 long terminal repeat transcriptional activation by inducible nitric oxide synthase 2 activity. J. Biol. Chem 1998, 273, 3895–3900. [Google Scholar]
- Ohkita, M.; Takaoka, M.; Shiota, Y.; Nojiri, R.; Matsumura, Y. Nitric oxide inhibits endothelin-1 production through the suppression of nuclear factor kappa B. Clin. Sci. (Lond. ) 2002, 103, 68S–71S. [Google Scholar]
- Perkins, N.D.; Gilmore, T.D. Good cop, bad cop: The different faces of NF-kappaB. Cell Death Differ 2006, 13, 759–772. [Google Scholar]
- Xie, Q.W.; Kashiwabara, Y.; Nathan, C. Role of transcription factor NF-kappa B/Rel in induction of nitric oxide synthase. J. Biol. Chem 1994, 269, 4705–4708. [Google Scholar]
- Muhl, H.; Pfeilschifter, J. Amplification of nitric oxide synthase expression by nitric oxide in interleukin 1 beta-stimulated rat mesangial cells. J. Clin. Invest 1995, 95, 1941–1946. [Google Scholar]
- Zouki, C.; Jozsef, L.; Ouellet, S.; Paquette, Y.; Filep, J.G. Peroxynitrite mediates cytokine-induced IL-8 gene expression and production by human leukocytes. J. Leukoc Biol 2001, 69, 815–824. [Google Scholar]
- Colasanti, M.; Persichini, T.; Menegazzi, M.; Mariotto, S.; Giordano, E.; Caldarera, C.M.; Sogos, V.; Lauro, G.M.; Suzuki, H. Induction of nitric oxide synthase mRNA expression. Suppression by exogenous nitric oxide. J. Biol. Chem 1995, 270, 26731–26733. [Google Scholar]
- Park, S.K.; Lin, H.L.; Murphy, S. Nitric oxide regulates nitric oxide synthase-2 gene expression by inhibiting NF-kappaB binding to DNA. Biochem. J 1997, 322, 609–613. [Google Scholar]
- Chen, F.; Lu, Y.; Castranova, V.; Rojanasakul, Y.; Miyahara, K.; Shizuta, Y.; Vallyathan, V.; Shi, X.; Demers, L.M. Nitric oxide inhibits HIV tat-induced NF-kappaB activation. Am. J. Pathol 1999, 155, 275–284. [Google Scholar]
- Matthews, J.R.; Wakasugi, N.; Virelizier, J.L.; Yodoi, J.; Hay, R.T. Thioredoxin regulates the DNA binding activity of NF-kappa B by reduction of a disulphide bond involving cysteine 62. Nucleic Acids Res 1992, 20, 3821–3830. [Google Scholar]
- Mitomo, K.; Nakayama, K.; Fujimoto, K.; Sun, X.; Seki, S.; Yamamoto, K. Two different cellular redox systems regulate the DNA-binding activity of the p50 subunit of NF-kappa B in vitro. Gene 1994, 145, 197–203. [Google Scholar]
- Satriano, J.; Schlondorff, D. Activation and attenuation of transcription factor NF-κB in mouse glomerular mesangial cells in response to tumor necrosis factor-alpha, immunoglobulin G, and adenosine 3′:5′-cyclic monophosphate. Evidence for involvement of reactive oxygen species. J. Clin. Invest 1994, 94, 1629–1636. [Google Scholar]
- Schreck, R.; Baeuerle, P.A. Assessing oxygen radicals as mediators in activation of inducible eukaryotic transcription factor NF-kappa B. Methods Enzymol 1994, 234, 151–163. [Google Scholar]
- Kabe, Y.; Ando, K.; Hirao, S.; Yoshida, M.; Handa, H. Redox regulation of NF-kappaB activation: Distinct redox regulation between the cytoplasm and the nucleus. Antioxid. Redox Signal 2005, 7, 395–403. [Google Scholar]
- Nishi, T.; Shimizu, N.; Hiramoto, M.; Sato, I.; Yamaguchi, Y.; Hasegawa, M.; Aizawa, S.; Tanaka, H.; Kataoka, K.; Watanabe, H.; et al. Spatial redox regulation of a critical cysteine residue of NF-kappa B in vivo. J. Biol. Chem 2002, 277, 44548–44556. [Google Scholar]
- Toledano, M.B.; Leonard, W.J. Modulation of transcription factor NF-kappa B binding activity by oxidation-reduction in vitro. Proc. Natl. Acad. Sci. USA 1991, 88, 4328–4332. [Google Scholar]
- Pineda-Molina, E.; Klatt, P.; Vazquez, J.; Marina, A.; Garcia de Lacoba, M.; Perez-Sala, D.; Lamas, S. Glutathionylation of the p50 subunit of NF-kappaB: A mechanism for redox-induced inhibition of DNA binding. Biochemistry 2001, 40, 14134–14142. [Google Scholar]
- Kamata, H.; Manabe, T.; Oka, S.; Kamata, K.; Hirata, H. Hydrogen peroxide activates IkappaB kinases through phosphorylation of serine residues in the activation loops. FEBS Lett 2002, 519, 231–237. [Google Scholar]
- Vandermoere, F.; El Yazidi-Belkoura, I.; Adriaenssens, E.; Lemoine, J.; Hondermarck, H. The antiapoptotic effect of fibroblast growth factor-2 is mediated through nuclear factor-kappaB activation induced via interaction between Akt and IkappaB kinase-beta in breast cancer cells. Oncogene 2005, 24, 5482–5491. [Google Scholar]
- Jamaluddin, M.; Wang, S.; Boldogh, I.; Tian, B.; Brasier, A.R. TNF-alpha-induced NF-kappaB/RelA Ser(276) phosphorylation and enhanceosome formation is mediated by an ROS-dependent PKAc pathway. Cell Signal 2007, 19, 1419–1433. [Google Scholar]
- Go, Y.M.; Gipp, J.J.; Mulcahy, R.T.; Jones, D.P. H2O2-dependent activation of GCLC-ARE4 reporter occurs by mitogen-activated protein kinase pathways without oxidation of cellular glutathione or thioredoxin-1. J. Biol. Chem 2004, 279, 5837–5845. [Google Scholar]
- Kamata, H.; Manabe, T.; Kakuta, J.; Oka, S.; Hirata, H. Multiple redox regulation of the cellular signaling system linked to AP-1 and NFkappaB: Effects of N-acetylcysteine and H2O2 on the receptor tyrosine kinases, the MAP kinase cascade, and IkappaB kinases. Ann. N. Y. Acad. Sci 2002, 973, 419–422. [Google Scholar]
- Vayalil, P.K.; Iles, K.E.; Choi, J.; Yi, A.K.; Postlethwait, E.M.; Liu, R.M. Glutathione suppresses TGF-beta-induced PAI-1 expression by inhibiting p38 and JNK MAPK and the binding of AP-1, SP-1, and Smad to the PAI-1 promoter. Am. J. Physiol. Lung Cell Mol. Physiol 2007, 293, L1281–L1292. [Google Scholar]
- Finocchietto, P.V.; Franco, M.C.; Holod, S.; Gonzalez, A.S.; Converso, D.P.; Antico Arciuch, V.G.; Serra, M.P.; Poderoso, J.J.; Carreras, M.C. Mitochondrial nitric oxide synthase: A masterpiece of metabolic adaptation, cell growth, transformation, and death. Exp. Biol. Med. (Maywood) 2009, 234, 1020–1028. [Google Scholar]
- Carreras, M.C.; Poderoso, J.J. Mitochondrial nitric oxide in the signaling of cell integrated responses. Am. J. Physiol. Cell Physiol 2007, 292, C1569–C1580. [Google Scholar]
- Gorlach, A.; Diebold, I.; Schini-Kerth, V.B.; Berchner-Pfannschmidt, U.; Roth, U.; Brandes, R.P.; Kietzmann, T.; Busse, R. Thrombin activates the hypoxia-inducible factor-1 signaling pathway in vascular smooth muscle cells: Role of the p22(phox)-containing NADPH oxidase. Circ. Res 2001, 89, 47–54. [Google Scholar]
- Shatrov, V.A.; Sumbayev, V.V.; Zhou, J.; Brune, B. Oxidized low-density lipoprotein (oxLDL) triggers hypoxia-inducible factor-1alpha (HIF-1alpha) accumulation via redox-dependent mechanisms. Blood 2003, 101, 4847–4849. [Google Scholar]
- Li, C.Q.; Robles, A.I.; Hanigan, C.L.; Hofseth, L.J.; Trudel, L.J.; Harris, C.C.; Wogan, G.N. Apoptotic signaling pathways induced by nitric oxide in human lymphoblastoid cells expressing wild-type or mutant p53. Cancer Res 2004, 64, 3022–3029. [Google Scholar]
- Rainwater, R.; Parks, D.; Anderson, M.E.; Tegtmeyer, P.; Mann, K. Role of cysteine residues in regulation of p53 function. Mol. Cell Biol 1995, 15, 3892–3903. [Google Scholar]
- Yang, F.; von Knethen, A.; Brune, B. Modulation of nitric oxide-evoked apoptosis by the p53-downstream target p21(WAF1/CIP1). J. Leukoc. Biol 2000, 68, 916–922. [Google Scholar]
- Giaccia, A.J.; Kastan, M.B. The complexity of p53 modulation: Emerging patterns from divergent signals. Genes Dev 1998, 12, 2973–2983. [Google Scholar]
- Li, J.M.; Fan, L.M.; George, V.T.; Brooks, G. Nox2 regulates endothelial cell cycle arrest and apoptosis via p21cip1 and p53. Free Radic. Biol. Med 2007, 43, 976–986. [Google Scholar]
- Liu, B.; Chen, Y.; St Clair, D.K. ROS and p53: A versatile partnership. Free Radic. Biol. Med 2008, 44, 1529–1535. [Google Scholar]
- Sun, Y.; Oberley, L.W. Redox regulation of transcriptional activators. Free Radic. Biol. Med 1996, 21, 335–348. [Google Scholar]
- Roebuck, K.A. Oxidant stress regulation of IL-8 and ICAM-1 gene expression: Differential activation and binding of the transcription factors AP-1 and NF-kappaB (Review). Int. J. Mol. Med 1999, 4, 223–230. [Google Scholar]
- Wu, X.; Bishopric, N.H.; Discher, D.J.; Murphy, B.J.; Webster, K.A. Physical and functional sensitivity of zinc finger transcription factors to redox change. Mol. Cell. Biol 1996, 16, 1035–1046. [Google Scholar]
- Akiba, S.; Chiba, M.; Mukaida, Y.; Sato, T. Involvement of reactive oxygen species and SP-1 in fibronectin production by oxidized LDL. Biochem. Biophys. Res. Commun 2003, 310, 491–497. [Google Scholar]
- Hampton, M.B.; Orrenius, S. Dual regulation of caspase activity by hydrogen peroxide: Implications for apoptosis. FEBS Lett 1997, 414, 552–556. [Google Scholar]
- Higuchi, M.; Honda, T.; Proske, R.J.; Yeh, E.T. Regulation of reactive oxygen species-induced apoptosis and necrosis by caspase 3-like proteases. Oncogene 1998, 17, 2753–2760. [Google Scholar]
- Moungjaroen, J.; Nimmannit, U.; Callery, P.S.; Wang, L.; Azad, N.; Lipipun, V.; Chanvorachote, P.; Rojanasakul, Y. Reactive oxygen species mediate caspase activation and apoptosis induced by lipoic acid in human lung epithelial cancer cells through Bcl-2 down-regulation. J. Pharmacol. Exp. Ther 2006, 319, 1062–1069. [Google Scholar]
- Fadeel, B.; Ahlin, A.; Henter, J.I.; Orrenius, S.; Hampton, M.B. Involvement of caspases in neutrophil apoptosis: Regulation by reactive oxygen species. Blood 1998, 92, 4808–4818. [Google Scholar]
- Izeradjene, K.; Douglas, L.; Tillman, D.M.; Delaney, A.B.; Houghton, J.A. Reactive oxygen species regulate caspase activation in tumor necrosis factor-related apoptosis-inducing ligand-resistant human colon carcinoma cell lines. Cancer Res 2005, 65, 7436–7445. [Google Scholar]
- Vlahopoulos, S.; Boldogh, I.; Casola, A.; Brasier, A.R. Nuclear factor-kappaB-dependent induction of interleukin-8 gene expression by tumor necrosis factor alpha: Evidence for an antioxidant sensitive activating pathway distinct from nuclear translocation. Blood 1999, 94, 1878–1889. [Google Scholar]
- Kurdi, M.; Booz, G.W. Evidence that IL-6-type cytokine signaling in cardiomyocytes is inhibited by oxidative stress: Parthenolide targets JAK1 activation by generating ROS. J. Cell Physiol 2007, 212, 424–431. [Google Scholar]
- Schulze-Osthoff, K.; Beyaert, R.; Vandevoorde, V.; Haegeman, G.; Fiers, W. Depletion of the mitochondrial electron transport abrogates the cytotoxic and gene-inductive effects of TNF. EMBO J 1993, 12, 3095–3104. [Google Scholar]
- De Keulenaer, G.W.; Ushio-Fukai, M.; Yin, Q.; Chung, A.B.; Lyons, P.R.; Ishizaka, N.; Rengarajan, K.; Taylor, W.R.; Alexander, R.W.; Griendling, K.K. Convergence of redox-sensitive and mitogen-activated protein kinase signaling pathways in tumor necrosis factor-alpha-mediated monocyte chemoattractant protein-1 induction in vascular smooth muscle cells. Arterioscler. Thromb. Vasc. Biol 2000, 20, 385–391. [Google Scholar]
- Shimada, T.; Watanabe, N.; Hiraishi, H.; Terano, A. Redox regulation of interleukin-8 expression in MKN28 cells. Dig. Dis. Sci 1999, 44, 266–273. [Google Scholar]
- Lo, Y.Y.; Cruz, T.F. Involvement of reactive oxygen species in cytokine and growth factor induction of c-fos expression in chondrocytes. J. Biol. Chem 1995, 270, 11727–11730. [Google Scholar]
- Maulik, N.; Das, D.K. Redox signaling in vascular angiogenesis. Free Radic. Biol. Med 2002, 33, 1047–1060. [Google Scholar]
- Eyries, M.; Collins, T.; Khachigian, L.M. Modulation of growth factor gene expression in vascular cells by oxidative stress. Endothelium 2004, 11, 133–139. [Google Scholar]
- Sundaresan, M.; Yu, Z.X.; Ferrans, V.J.; Irani, K.; Finkel, T. Requirement for generation of H2O2 for platelet-derived growth factor signal transduction. Science 1995, 270, 296–299. [Google Scholar]
- Morgan, M.J.; Liu, Z.G. Reactive oxygen species in TNFalpha-induced signaling and cell death. Mol. Cells 2010, 30, 1–12. [Google Scholar]
- Circu, M.L.; Aw, T.Y. Reactive oxygen species, cellular redox systems, and apoptosis. Free Radic. Biol. Med 2010, 48, 749–762. [Google Scholar]
- Fruehauf, J.P.; Meyskens, F.L., Jr. Reactive oxygen species: A breath of life or death? Clin. Cancer Res. 2007, 13, 789–794. [Google Scholar]
- Simon, H.U.; Haj-Yehia, A.; Levi-Schaffer, F. Role of reactive oxygen species (ROS) in apoptosis induction. Apoptosis 2000, 5, 415–418. [Google Scholar]
- Kizaki, M.; Xian, M.; Sagawa, M.; Ikeda, Y. Induction of apoptosis via the modulation of reactive oxygen species (ROS) production in the treatment of myeloid leukemia. Curr. Pharm. Biotechnol 2006, 7, 323–329. [Google Scholar]
- Leon, L.; Jeannin, J.F.; Bettaieb, A. Post-translational modifications induced by nitric oxide (NO): Implication in cancer cells apoptosis. Nitric. Oxide 2008, 19, 77–83. [Google Scholar]
- Tarr, J.M.; Eggleton, P.; Winyard, P.G. Nitric oxide and the regulation of apoptosis in tumour cells. Curr. Pharm. Des 2006, 12, 4445–4468. [Google Scholar]
- Li, C.Q.; Wogan, G.N. Nitric oxide as a modulator of apoptosis. Cancer Lett 2005, 226, 1–15. [Google Scholar]
- Brune, B. The intimate relation between nitric oxide and superoxide in apoptosis and cell survival. Antioxid. Redox Signal 2005, 7, 497–507. [Google Scholar]
- Vieira, H.; Kroemer, G. Mitochondria as targets of apoptosis regulation by nitric oxide. IUBMB Life 2003, 55, 613–616. [Google Scholar]
- Brune, B.; Zhou, J.; von Knethen, A. Nitric oxide, oxidative stress, and apoptosis. Kidney Int. Suppl. 2003, S22–24. [Google Scholar]
- Park, J.B. Phagocytosis induces superoxide formation and apoptosis in macrophages. Exp. Mol. Med 2003, 35, 325–335. [Google Scholar]
- Valko, M.; Leibfritz, D.; Moncol, J.; Cronin, M.T.; Mazur, M.; Telser, J. Free radicals and antioxidants in normal physiological functions and human disease. Int. J. Biochem. Cell Biol 2007, 39, 44–84. [Google Scholar]
- Ghosh, N.; Ghosh, R.; Mandal, S.C. Antioxidant protection: A promising therapeutic intervention in neurodegenerative disease. Free Radic. Res 2011, 45, 888–905. [Google Scholar]
- Iborra, M.; Moret, I.; Rausell, F.; Bastida, G.; Aguas, M.; Cerrillo, E.; Nos, P.; Beltran, B. Role of oxidative stress and antioxidant enzymes in Crohn’s disease. Biochem. Soc. Trans 2011, 39, 1102–1106. [Google Scholar]
- Dumont, M.; Lin, M.T.; Beal, M.F. Mitochondria and antioxidant targeted therapeutic strategies for Alzheimer’s disease. J. Alzheimers Dis 2010, 20, S633–S643. [Google Scholar]
- De Vries, H.E.; Witte, M.; Hondius, D.; Rozemuller, A.J.; Drukarch, B.; Hoozemans, J.; van Horssen, J. Nrf2-induced antioxidant protection: A promising target to counteract ROS-mediated damage in neurodegenerative disease? Free Radic. Biol. Med 2008, 45, 1375–1383. [Google Scholar]
- De Araujo, D.P.; de Lobato, R.F.; Cavalcanti, J.R.; Sampaio, L.R.; Araujo, P.V.; Silva, M.C.; Neves, K.R.; Fonteles, M.M.; Sousa, F.C.; Vasconcelos, S.M. The contributions of antioxidant activity of lipoic acid in reducing neurogenerative progression of Parkinson’s disease: A review. Int. J. NeuroSci 2011, 121, 51–57. [Google Scholar]
- Aliev, G.; Obrenovich, M.E.; Reddy, V.P.; Shenk, J.C.; Moreira, P.I.; Nunomura, A.; Zhu, X.; Smith, M.A.; Perry, G. Antioxidant therapy in Alzheimer’s disease: Theory and practice. Mini Rev. Med. Chem 2008, 8, 1395–1406. [Google Scholar]
- Zarjou, A.; Sanders, P.W.; Mehta, R.L.; Agarwal, A. Enabling innovative translational research in acute kidney injury. Clin. Transl. Sci 2012, 5, 93–101. [Google Scholar]
- Aziz, M.H.; Kumar, R.; Ahmad, N. Cancer chemoprevention by resveratrol: In vitro and in vivo studies and the underlying mechanisms (review). Int. J. Oncol 2003, 23, 17–28. [Google Scholar]
- Turan, B.; Vassort, G. Vitamin E in oxidant stress-related cardiovascular pathologies: Focus on experimental studies. Curr. Pharm. Des 2011, 17, 2155–2169. [Google Scholar]
- Turan, B. Role of antioxidants in redox regulation of diabetic cardiovascular complications. Curr. Pharm. Biotechnol 2010, 11, 819–836. [Google Scholar]
- Violi, F.; Cangemi, R. Antioxidants and cardiovascular disease. Curr. Opin. Investig Drugs 2005, 6, 895–900. [Google Scholar]
- Maxwell, S.R.; Lip, G.Y. Free radicals and antioxidants in cardiovascular disease. Br. J. Clin. Pharmacol 1997, 44, 307–317. [Google Scholar]
- Tylicki, L.; Rutkowski, B.; Horl, W.H. Antioxidants: A possible role in kidney protection. Kidney Blood Press Res 2003, 26, 303–314. [Google Scholar]
- Day, B.J. Antioxidants as potential therapeutics for lung fibrosis. Antioxid. Redox Signal 2008, 10, 355–370. [Google Scholar]
- Kinnula, V.L.; Crapo, J.D. Superoxide dismutases in the lung and human lung diseases. Am. J. Respir Crit. Care Med 2003, 167, 1600–1619. [Google Scholar]
- Russell, G.A. Antioxidants and neonatal lung disease. Eur. J. Pediatr 1994, 153, S36–S41. [Google Scholar]
- Kamat, C.D.; Gadal, S.; Mhatre, M.; Williamson, K.S.; Pye, Q.N.; Hensley, K. Antioxidants in central nervous system diseases: Preclinical promise and translational challenges. J. Alzheimers Dis 2008, 15, 473–493. [Google Scholar]
- Durante, W. Targeting heme oxygenase-1 in vascular disease. Curr. Drug Targets 2010, 11, 1504–1516. [Google Scholar]
- Subramanian, S.; Kalyanaraman, B.; Migrino, R.Q. Mitochondrially targeted antioxidants for the treatment of cardiovascular diseases. Recent Pat. Cardiovasc. Drug Discov 2010, 5, 54–65. [Google Scholar]
- Sawyer, D.B. Oxidative stress in heart failure: What are we missing? Am. J. Med. Sci 2011, 342, 120–124. [Google Scholar]
- Wu, J.; Hecker, J.G.; Chiamvimonvat, N. Antioxidant enzyme gene transfer for ischemic diseases. Adv. Drug Deliv. Rev 2009, 61, 351–363. [Google Scholar]
- Golden, T.R.; Patel, M. Catalytic antioxidants and neurodegeneration. Antioxid. Redox Signal 2009, 11, 555–570. [Google Scholar]
- Poljsak, B.; Milisav, I. The neglected significance of “antioxidative stress”. Oxid. Med. Cell Longev 2012, 2012, 480895. [Google Scholar]
- Firuzi, O.; Miri, R.; Tavakkoli, M.; Saso, L. Antioxidant therapy: Current status and future prospects. Curr. Med. Chem 2011, 18, 3871–3888. [Google Scholar]
- Mecocci, P.; Polidori, M.C. Antioxidant clinical trials in mild cognitive impairment and Alzheimer’s disease. Biochim. Biophys. Acta 2012, 1822, 631–638. [Google Scholar]
- Shamseer, L.; Adams, D.; Brown, N.; Johnson, J.A.; Vohra, S. Antioxidant micronutrients for lung disease in cystic fibrosis. Cochrane Database Syst. Rev. 2010. [Google Scholar] [CrossRef]
- Nunez-Cordoba, J.M.; Martinez-Gonzalez, M.A. Antioxidant vitamins and cardiovascular disease. Curr. Top. Med. Chem 2011, 11, 1861–1869. [Google Scholar]
- Bonnefont-Rousselot, D.; Collin, F. Melatonin: Action as antioxidant and potential applications in human disease and aging. Toxicology 2010, 278, 55–67. [Google Scholar]
- Lee, H.P.; Zhu, X.; Casadesus, G.; Castellani, R.J.; Nunomura, A.; Smith, M.A.; Lee, H.G.; Perry, G. Antioxidant approaches for the treatment of Alzheimer’s disease. Expert Rev. Neurother 2010, 10, 1201–1208. [Google Scholar]
- Farbstein, D.; Kozak-Blickstein, A.; Levy, A.P. Antioxidant vitamins and their use in preventing cardiovascular disease. Molecules 2010, 15, 8098–8110. [Google Scholar]
- Levonen, A.L.; Vahakangas, E.; Koponen, J.K.; Yla-Herttuala, S. Antioxidant gene therapy for cardiovascular disease: Current status and future perspectives. Circulation 2008, 117, 2142–2150. [Google Scholar]
- Davis, C.A.; Nick, H.S.; Agarwal, A. Manganese superoxide dismutase attenuates Cisplatin-induced renal injury: Importance of superoxide. J. Am. Soc. Nephrol 2001, 12, 2683–2690. [Google Scholar]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).