Clinicopathological Significance of NMIIA Overexpression in Human Gastric Cancer
Abstract
:1. Introduction
2. Results
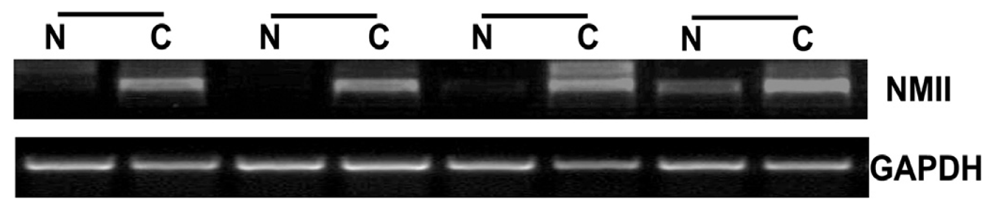
2.1. Overexpression of NMIIA in Human GC
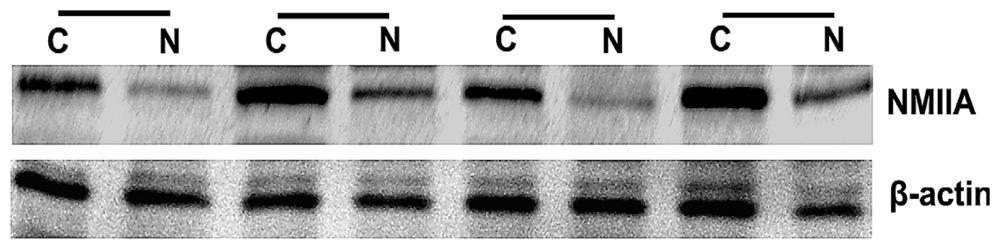
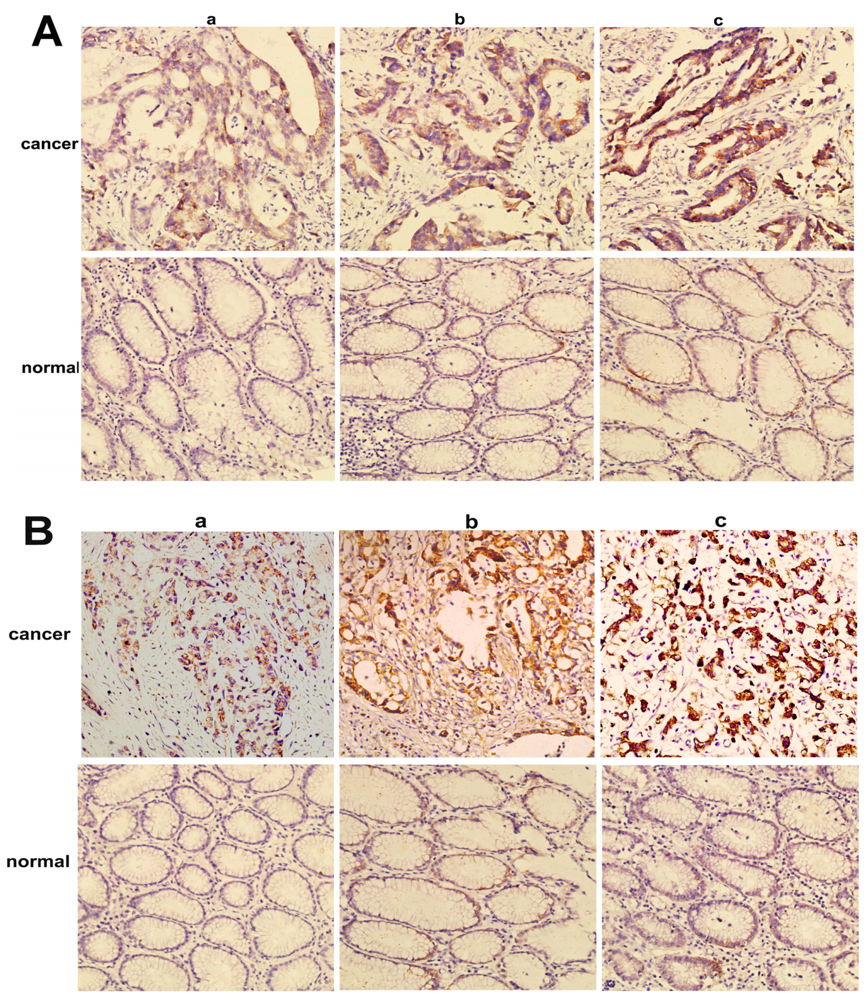
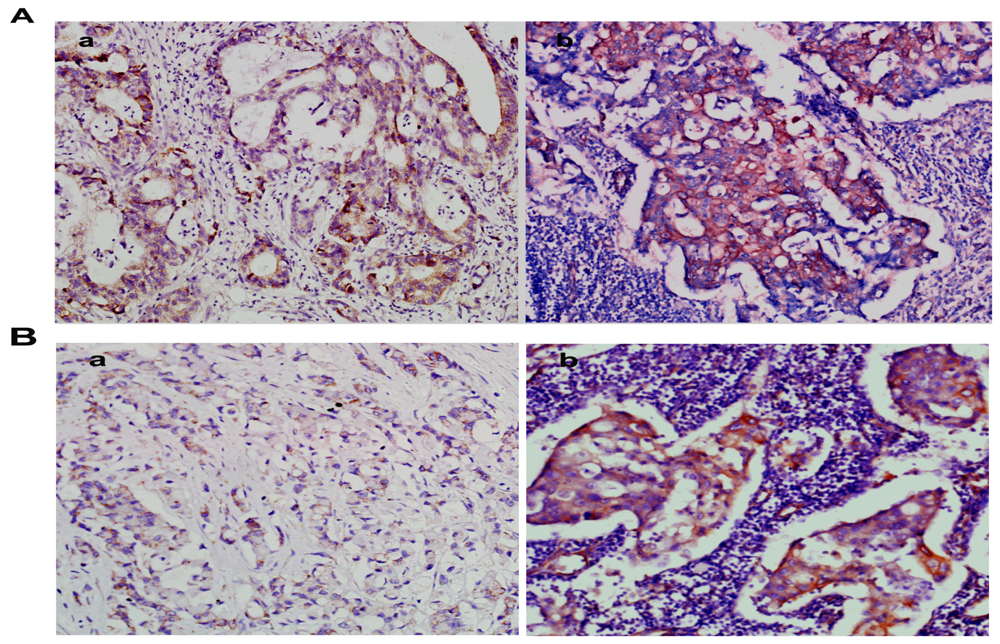
2.2. Overexpression of NMIIA Protein in Human GC
2.3. NMIIA Overexpression in Relation to Clinicopathologic Parameters
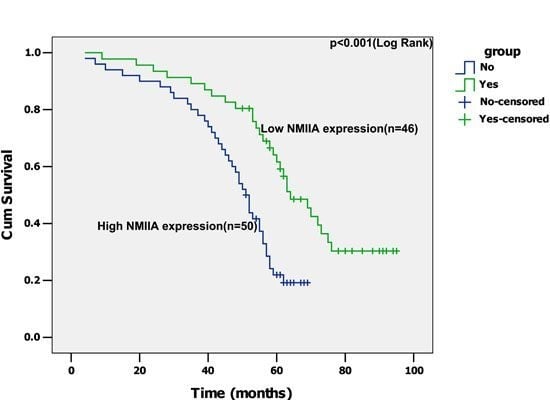
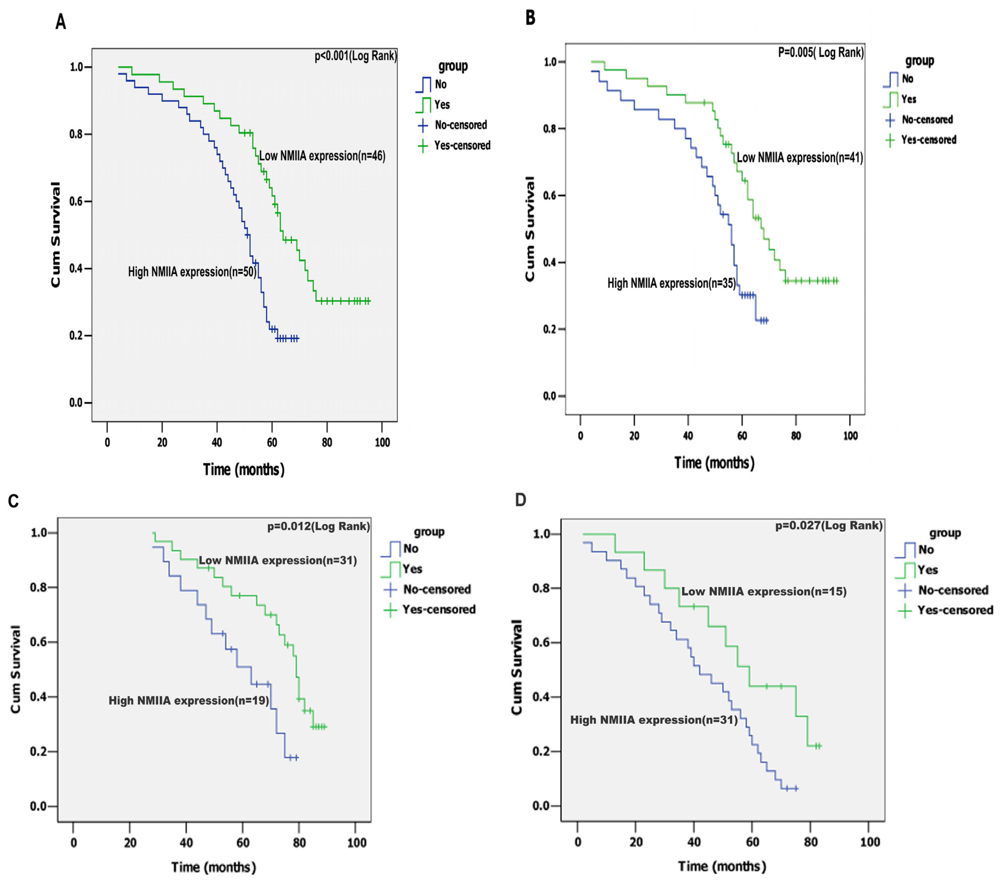
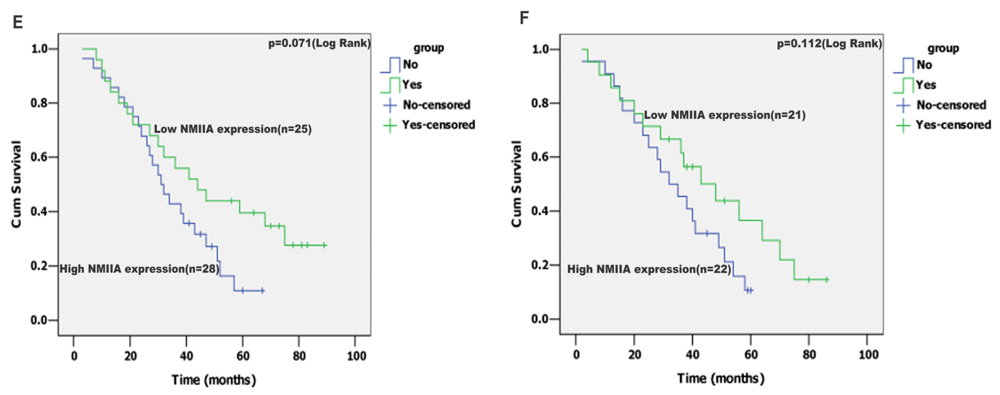
2.4. Univariate Analysis
2.5. Multivariate Analysis
3. Discussion
4. Materials and Methods
4.1. Tissue Specimens
4.2. RNA Extraction and Reverse-Transcriptase PCR
4.3. Protein Extraction and Western Blot Analysis
4.4. Immunohistochemistry
4.5. Statistical Analysis
5. Conclusions
Acknowledgments
- Conflict of InterestThe authors declare no conflict of interest.
References
- Jemal, A.; Bray, F.; Center, M.M.; Ferlay, J.; Ward, E.; Forman, D. Global cancer statistics. CA: Cancer J. Clin 2011, 61, 69–90. [Google Scholar]
- Sereno, M.; Aguayo, C.; Guillén Ponce, C.; Gómez-Raposo, C.; Zambrana, F.; Gómez-López, M.; Casado, E. Gastric tumours in hereditary cancer syndromes: Clinical features, molecular biology and strategies for prevention. Clin. Transl. Oncol 2011, 13, 599–610. [Google Scholar]
- Tahara, E. Molecular mechanism of stomach carcinogenesis. J. Cancer Res. Clin. Oncol 1993, 119, 265–272. [Google Scholar]
- Barreto-Zuñiga, R.; Maruyama, M.; Kato, Y.; Aizu, K.; Ohta, H.; Takekoshi, T.; Bernal, S.F. Significance of Helicobacter pylori infection as a risk factor in gastric cancer: Serological and histological studies. J. Gastroenterol 1997, 32, 289–294. [Google Scholar]
- Kim, K.M.; Kwon, M.S.; Hong, S.J.; Min, K.O.; Seo, E.J.; Lee, K.Y.; Choi, S.W.; Rhyu, M.G. Genetic classification of intestinal-type and diffuse-type gastric cancers based on chromosomal loss and microsatellite instability. Virchows Arch 2003, 443, 491–500. [Google Scholar]
- Dicken, B.J.; Bigam, D.L.; Cass, C.; Mackey, J.R.; Joy, A.A.; Hamilton, S.M. Gastric adenocarcinoma: Review and considerations for future directions. Ann. Surg 2005, 241, 27–39. [Google Scholar]
- Zhang, B.; Cao, W.; Zhang, F.; Zhang, L.; Niu, R.; Niu, Y.; Fu, L.; Hao, X.; Cao, X. Protein interacting with C alpha kinase 1 (PICK1) is involved in promoting tumor growth and correlates with poor prognosis of human breast cancer. Cancer Sci 2010, 101, 1536–1542. [Google Scholar]
- Simons, M.; Wang, M.; McBride, O.W.; Kawamoto, S.; Yamakawa, K.; Gdula, D.; Adelstein, R.S.; Weir, L. Human nonmuscle myosin heavy chains are encoded by two genes located on different chromosomes. Circ. Res 1991, 69, 530–539. [Google Scholar]
- Toothaker, L.E.; Gonzalez, D.A.; Tung, N.; Lemons, R.S.; le Beau, M.M.; Arnaout, M.A.; Clayton, L.K.; Tenen, D.G. Cellular myosin heavy chain in human leukocytes: Isolation of 5′cDNA clones, characterization of the protein, chromosomal localization, and upregulation during myeloid differentiation. Blood 1991, 78, 1826–1833. [Google Scholar]
- Vicente-Manzanares, M.; Ma, X.; Adelstein, R.S.; Horwitz, A.R. Non-muscle myosin II takes centre stage in cell adhesion and migration. Nat. Rev. Mol. Cell Biol 2009, 11, 778–790. [Google Scholar]
- Conti, M.A.; Adelstein, R.S. Nonmuscle myosin II moves in new directions. J. Cell Sci 2008, 121, 11–18. [Google Scholar]
- Dulyaninova, N.G.; House, R.P.; Betapudi, V.; Bresnick, A.R. Myosin-IIA heavy-chain phosphorylation regulates the motility of MDA-MB-231 carcinoma cells. Mol. Biol. Cell 2007, 18, 3144–3155. [Google Scholar]
- Maeda, J.; Hirano, T.; Ogiwara, A.; Akimoto, S.; Kawakami, T.; Fukui, Y.; Oka, T.; Gong, Y.; Guo, R.; Inada, H.; et al. Proteomic analysis of stage I primary lung adenocarcinoma aimed at individualisation of postoperative therapy. Br. J. Cancer 2008, 98, 596–603. [Google Scholar]
- Xia, Z.K.; Yuan, Y.C.; Yin, N.; Yin, B.L.; Tan, Z.P.; Hu, Y.R. Nonmuscle myosin IIA is associated with poor prognosis of esophageal squamous cancer. Dis. Esophagus 2012, 25, 427–436. [Google Scholar]
- Liang, S.; He, L.; Zhao, X.; Miao, Y.; Gu, Y.; Guo, C.; Xue, Z.; Dou, W.; Hu, F.; Wu, K.; et al. MicroRNA let-7f inhibits tumor invasion and metastasis by targeting MYH9 in human gastric cancer. PLoS One 2011, 6, e18409. [Google Scholar]
- Betapudi, V.; Licate, L.S.; Egelhoff, T.T. Distinct roles of nonmuscle myosin II isoforms in the regulation of MDA-MB-231 breast cancer cell spreading and migration. Cancer Res 2006, 66, 4725–4733. [Google Scholar]
- Minamiya, Y.; Nakagawa, T.; Saito, H.; Matsuzaki, I.; Taguchi, K.; Ito, M.; Ogawa, J. Increased expression of myosin light chain kinase mRNA is related to metastasis in non-small cell lung cancer. Tumour Biol 2005, 26, 153–157. [Google Scholar]
- Garrett, S.C.; Varney, K.M.; Weber, D.J.; Bresnick, A.R. S100A4, a mediator of metastasis. J. Biol. Chem 2005, 281, 677–680. [Google Scholar]
- Zaidel-Bar, R.; Ballestrem, C.; Kam, Z.; Geiger, B. Early molecular events in the assembly of matrix adhesions at the leading edge of migrating cells. J. Cell Sci 2003, 116, 4605–4613. [Google Scholar]
- Babbin, B.A.; Koch, S.; Bachar, M.; Conti, M.A.; Parkos, C.A.; Adelstein, R.S.; Nusrat, A.; Ivanov, A.I. Non-muscle myosin IIA differentially regulates intestinal epithelial cell restitution and matrix invasion. Am. J. Pathol 2009, 174, 436–448. [Google Scholar]
- Li, Y.; Friedmann, D.R.; Mhatre, A.N.; Lalwani, A.K. MYH9-SiRNA and MYH9 mutant alleles: Expression in cultured cell lines and their effects upon cell structure and function. Cell Motil. Cytoskeleton 2008, 65, 393–405. [Google Scholar]
- Yamazaki, D.; Kurisu, S.; Takenawa, T. Regulation of cancer cell motility through actin reorganization. Cancer Sci 2005, 96, 379–386. [Google Scholar]
- Even-Ram, S.; Doyle, A.D.; Conti, M.A.; Matsumoto, K.; Adelstein, R.S.; Yamada, K.M. Myosin IIA regulates cell motility and actomyosin-microtubule crosstalk. Nat. Cell Biol 2007, 9, 299–309. [Google Scholar]
| n | SI (mean ± SE) | Mean rank | U | p * value | |
|---|---|---|---|---|---|
| Benign epithelium | 30 | 3.20 ± 0.26 | 16.92 | 42.50 | <0.001 |
| Gastric cancer | 96 | 9.52 ± 0.29 | 78.06 | - | - |
| Benign epithelium | 30 | 3.20 ± 0.26 | 18.92 | 102.50 | <0.001 |
| Gastric cancer (intestinal) | 53 | 8.91 ± 0.31 | 55.07 | - | - |
| Benign epithelium | 30 | 3.20 ± 0.26 | 15.57 | 2.00 | <0.001 |
| Gastric cancer (diffuse) | 43 | 9.93 ± 0.24 | 51.95 | - | - |
| Characteristics | NMIIA | p value | |
|---|---|---|---|
| Low (%) | High (%) | ||
| Gender | 1.000 | ||
| Male | 28 (47.45) | 31 (52.55) | - |
| Female | 18 (48.64) | 19 (51.36) | - |
| Age (years) | 1.000 | ||
| <55 | 26 (47.27) | 29 (52.73) | - |
| ≥55 | 20 (48.78) | 21 (51.22) | - |
| Tumor size (cm) | 0.412 | ||
| <5 | 23 (53.48) | 20 (46.52) | - |
| ≥5 | 23 (43.39) | 30 (56.61) | - |
| Tumor location | 0.548 | ||
| Upper body or whole | 21 (44.68) | 26 (55.32) | - |
| Lower or middle body | 25 (51.02) | 24 (48.98) | - |
| Tumor differentiation | 0.078 | ||
| Well | 20 (62.50) | 12 (37.50) | - |
| Moderate | 15 (46.87) | 17 (53.13) | - |
| Poor | 11 (34.37) | 21 (65.63) | - |
| Lauren classification | 1.000 | ||
| Intestinal type | 25 (47.16) | 28 (52.84) | |
| Diffuse type | 21 (48.83) | 22 (51.17) | |
| Depth of wall invasion | 0.026 | ||
| T1 | 14 (70.00) | 6 (30.00) | - |
| T2 | 12 (42.85) | 9 (57.15) | - |
| T3 | 10 (47.61) | 11 (52.39) | - |
| T4 | 10 (29.41) | 24 (70.59) | - |
| Mean no. of tLNs | 18 | 20 | 0.545 |
| Mean no. of mLNs | 7 | 13 | <0.001 |
| Lymph node metastasis | 0.024 | ||
| Absent (N0) | 28 (60.86) | 18 (39.14) | - |
| Present (N1–3) | 18 (36.00) | 32 (64.00) | - |
| Distant metastasis | 0.025 | ||
| Absent (M0) | 41 (53.94) | 35 (46.06) | - |
| Present (M1) | 5 (25.00) | 15 (75.00) | - |
| TNM stage | - | - | 0.005 |
| I + II | 31 (62.00) | 19 (38.00) | - |
| III + IV | 15 (32.60) | 31 (67.40) | - |
| Variable | Univariate | Multivariae | ||||
|---|---|---|---|---|---|---|
| HR | CI (95%) | p value | HR | CI (95%) | p value | |
| Gender | 1.242 | 0.742–2.079 | 0.409 | - | - | - |
| Age | 0.997 | 0.973–1.022 | 0.815 | - | - | - |
| Tumor size | 1.007 | 0.995–1.020 | 0.266 | - | - | - |
| Tumor location | 1.176 | 0.721–1.917 | 0.516 | - | - | - |
| Lauren classification | 1.085 | 0.899–1.310 | 0.393 | - | - | - |
| Tumor differentiation | 1.063 | 0.792–1.426 | 0.684 | - | - | - |
| Depth of wall invasion | 1.818 | 1.431–2.309 | <0.001 | 0.990 | 0.579–1.690 | 0.969 |
| Lymph node metastasis | 4.031 | 2.289–7.099 | <0.001 | 2.849 | 1.388–5.850 | 0.004 |
| Distant metastasis | 4.069 | 2.262–7.320 | <0.001 | 1.920 | 0.991–3.722 | 0.053 |
| TNM stage | 3.875 | 2.301–6.525 | <0.001 | 2.474 | 0.432–5.028 | 0.535 |
| NMIIA expression | 2.530 | 1.485–4.311 | 0.001 | 2.031 | 1.113–3.706 | 0.021 |
© 2012 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Liu, D.; Zhang, L.; Shen, Z.; Tan, F.; Hu, Y.; Yu, J.; Li, G. Clinicopathological Significance of NMIIA Overexpression in Human Gastric Cancer. Int. J. Mol. Sci. 2012, 13, 15291-15304. https://doi.org/10.3390/ijms131115291
Liu D, Zhang L, Shen Z, Tan F, Hu Y, Yu J, Li G. Clinicopathological Significance of NMIIA Overexpression in Human Gastric Cancer. International Journal of Molecular Sciences. 2012; 13(11):15291-15304. https://doi.org/10.3390/ijms131115291
Chicago/Turabian StyleLiu, Dongning, Lei Zhang, Zhiyong Shen, Fei Tan, Yanfeng Hu, Jiang Yu, and Guoxin Li. 2012. "Clinicopathological Significance of NMIIA Overexpression in Human Gastric Cancer" International Journal of Molecular Sciences 13, no. 11: 15291-15304. https://doi.org/10.3390/ijms131115291
APA StyleLiu, D., Zhang, L., Shen, Z., Tan, F., Hu, Y., Yu, J., & Li, G. (2012). Clinicopathological Significance of NMIIA Overexpression in Human Gastric Cancer. International Journal of Molecular Sciences, 13(11), 15291-15304. https://doi.org/10.3390/ijms131115291