Natural Alkaloids as Antiviral Agents Against RNA Viruses: A Comprehensive and Mechanistic Review
Abstract
1. Introduction
Overview of RNA Viruses and Their Global Health Impact
2. Virus Classification in the Genomic Era
- I: dsDNA (e.g., Herpes simplex virus)
- II: ssDNA → dsDNA before transcription (e.g., Parvovirus)
- III: dsRNA; mRNA transcribed from RNA genome (e.g., Rotavirus)
- IV: (+)ssRNA; genome functions as mRNA (e.g., Picornavirus, Coronavirus, Flavivirus)
- V: (−)ssRNA; mRNA transcribed from RNA genome (e.g., Rabies virus, Influenza virus, Ebola virus)
- VI: ssRNA with reverse transcriptase (e.g., HIV)
- VII: dsDNA with reverse transcriptase (e.g., Hepatitis B virus)
3. Alkaloid Biosynthetic Pathways and Structural Diversity
3.1. Classification of Alkaloids
3.2. Host Hijacking and Viral Translation
3.3. Alkaloid Antiviral Modes of Action
4. Quaternary Alkaloids: Distinct Mechanisms and Pharmacology
| Compound & Origin | References | Study Type | Virus | Mechanisms | Antiviral Information Reported in the Original Studies (IC50, EC50, SI, Docking Scores, Qualitative Observations) * |
|---|---|---|---|---|---|
| 1. Berberine and derivates (From: Berberidaceae/Ranunculaceae family)  | Wu et al. 2011 [115] | In vitro and in vivo | IAV | Berberine exerted strong inhibition on the inflammatory substances production | IC50 = 0.025 g/L Decrease in mice mortality from 90% to 55% |
| Wang et al. 2018 [116] | In vitro | EV71 | Might inhibit MEK/ERK, suppresses autophagy (AKT, JNK, PI3KIII) | IC50: 7.12–14.8 μM (Compound 2d) and 7.43–10.25 μM (berberine) | |
| Shao et al. 2020 [117] | In vitro and in silico | HIV-1, clade B | It binds in the pocket of NHR and CHR of gp41. | IC50: 5.5–10.25 μg/mL | |
| Ratanakomol et al. 2021 [118] | In vitro | DENV, ZIKV, CHIKV | Potential AMPK activation, lipid metabolism disruption, direct virucidal activity | IC50: DENV: 42.87 μM; ZIKV: 11.42 μM; CHIKV: 14.21 μM | |
| Botwina et al. 2020 [119] | In vitro | IAV (H3N2) | Inhibits MAPK/ERK | IC50: MDCK = 52 μM; A549 = 17 μM; LET1 = 4 μM; Human airway epithelial (HAE) = 16 μM | |
| Enkhtaivan et al. 2017 [120] | In vitro and in silico | IAV | Competitive neuraminidase inhibition | Berberine derivatives IC50 H1N1: 0.87–1.63 µg/mL H3N2: 1.15–2.98 µg/mL | |
| Varghese et al. 2016 [121] | In vitro | CHIKV | Might be affecting one or several host factors important for CHIKV replication | IC50: 1.9 ± 0.9 μM | |
| Nguyen, C. Q. et al. (2021) [122] | In vitro and in silico | ZIKV | Potential candidate to inhibit NS2B-NS3 protease | Compound 4d: Selectivity index (SI): 15.3 IC50 = 5.3 ± 1.9 µM | |
| 2. Chelerythrine (From: Papaveraceae family)  | Españo, E. et al. (2022) [123] | In vitro, in silico | ZIKV | Potential entry/attachment inhibition | EC50 = 692.4 nM SI = 6.0 |
| Loe, M. et al. 2023 [124] | In vitro, in vivo | ZIKV | Potent inhibitor of ZIKV infection that targets the ZIKV NS4B protein | Chelerythrine chloride IC50 = 0.2513 µM. A 1.73 and 2.0 log 10 reduction in RNA copies/mL | |
| Guo, W. et al. 2020 [125] | In vitro (plant) | Tobacco Mosaic Virus (TMV) | Inactivation/proliferation inhibition | Chelerythrine at 0.5 mg/mL: 72.67% inactivation, (corresponding to 1.4 mM **) | |
| 3. Dehydrocorydaline (From: Papaveraceae family)  | Orhan, I. et al. 2007 [126] | In vitro | Parainfluenza-3 | N/A | CPE inhibitory concentration 16 µg/mL (≈ 40–44 µM ***) |
| 4. Oxymatrine (From: Fabaceae family)  | Dai, J. et al. 2018 [127] | In vitro and in vivo | IAV multiple strains (H1N1, H9N2, H5N1, H3N2) | Could significantly decrease the promoter activity of TLR signaling pathways TLR4, p38 MAPK, NF-κB | EC50: 5.91–23.67 µg/mL (strain-dependent) (= 22–90 µM *) |
| Zhi et al. 2024 [128] | In vitro | H9N2 Avian Influenza Virus (AIV) | TLR signaling pathways TLR3, NF-κB, IRF-3 | Dose-dependent effects on the cell survival rate | |
| Chen, N. et al. (2016) [129] | In vitro | HCV | Proliferation inhibition | Inhibition of cell proliferation: up to 85.4% at 72 h, 12mg/mL; mRNA expression 0.59 ± 0.12 vs. control | |
| 5. Oxysophoridine (From: Fabaceae family)  | Majnooni, M. et al. (2021) [33] | Review | SARS-CoV-2 | Nucleotide biosynthesis inhibitor | EC50 = 0.31 μM, |
| 6. Palmatine (From: Ranunculaceae; Rutaceae family)  | W. Zhang, 2024 [130] | In vitro | Infectious Bronchitis virus | Inactivating the virus, inhibiting its replication, modulating NF-κB/IRF7/JAK-STAT signaling pathways, and regulating apoptosis | IC50 = 7.76 µM Selection index (SI) was 86.74 |
| Fan Jia et al. 2010 [131] | In vitro | WNV | Palmatine could significantly inhibit the activity of NS2B-NS3 protease and that the inhibition was reversible | WNV: EC50: 3.6 µM, IC50: 96 µM | |
| Yi-Jung Ho et al. 2019 [132] | In vitro | ZIKV JEV | Inhibits Zika virus infection by disrupting virus binding, entry, and stability | Inhibited ZIKV binding by 95% and ZIKV entry by 69% Palmatine from 20–80 mM decreased JEV RNA levels. | |
| 7. Sanguinarine (From: Papaveraceae family)  | Qiyun Ke et al. 2023 [133] | In vitro and in silico | PRRSV | Targeting internalization, replication, and release stages of the viral life cycle | Sanguinarine inhibits the propagation of PRRSV in a dose-dependent manner |
5. Discussion
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ACE2 | Angiotensin Converting Enzyme 2 |
| ADMET | Absorption, Distribution, Metabolism, Excretion, and Toxicity |
| AIV | Avian Influenza Virus |
| AMPK | AMP-activated Protein Kinase |
| CAM | Chorioallantoic Membrane |
| CC50 | 50% Cytotoxic Concentration |
| CEB | Cepharanthine (context dependent, but in your text CEP is used) |
| CEP/PD-001 | Cepharanthine/PharmaDrug oral formulation PD-001 |
| CHIKV | Chikungunya Virus |
| CI | Confidence Interval |
| CPE | Cytopathic Effect |
| COVID-19 | Coronavirus Disease 2019 |
| CYP450 | Cytochrome P450 Enzyme System |
| DENV | Dengue Virus |
| DNJ | Deoxynojirimycin |
| EC50 | 50% Effective Concentration |
| EBOV/Ebola | Ebola Virus |
| ELISA | Enzyme-Linked Immunosorbent Assay (appears implied in mechanistic work) |
| ER | Endoplasmic Reticulum |
| ERK | Extracellular Signal-Regulated Kinase |
| FDA | U.S. Food and Drug Administration |
| GP | Glycoprotein |
| HAE | Human Airway Epithelium |
| HBV | Hepatitis B Virus (if referenced) |
| HCV | Hepatitis C Virus |
| HCoV | Human Coronavirus |
| HIV | Human Immunodeficiency Virus |
| hRSV | Human Respiratory Syncytial Virus |
| IC50 | 50% Inhibitory Concentration |
| IFN | Interferon |
| IL | Interleukin |
| IRF | Interferon Regulatory Factor |
| JAK | Janus Kinase |
| JEV | Japanese Encephalitis Virus |
| JNK | c-Jun N-terminal Kinase |
| LDH | Lactate Dehydrogenase |
| LEDGF/p75 | Lens Epithelium Derived Growth Factor |
| MAPK | Mitogen-Activated Protein Kinase |
| MERS-CoV | Middle East Respiratory Syndrome Coronavirus |
| MOI | Multiplicity of Infection |
| Mpro (3CLpro) | Main Protease of SARS-CoV-2 |
| MTT | 3-(4,5-Dimethylthiazol-2-yl)-2,5-Diphenyltetrazolium Bromide |
| NA | Neuraminidase |
| NBD | Nucleotide Binding Domain |
| NDV | Newcastle Disease Virus |
| NF-κB | Nuclear Factor kappa-light-chain-enhancer of activated B cells |
| NS proteins | Non-structural viral proteins (e.g., NS2B, NS3, NS4B, NS5) |
| PA | Polymerase Acidic Protein (Influenza) |
| PEDV | Porcine Epidemic Diarrhea Virus |
| PK | Pharmacokinetics |
| qPCR/RT-qPCR | Quantitative Real-Time Polymerase Chain Reaction |
| RdRp | RNA-dependent RNA Polymerase |
| RNP | Ribonucleoprotein |
| ROS | Reactive Oxygen Species |
| RSV/BRSV | Respiratory Syncytial Virus/Bovine RSV |
| SI | Selectivity Index |
| SARS-CoV-2 | Severe Acute Respiratory Syndrome Coronavirus 2 |
| TLR | Toll-Like Receptor |
| TMPRSS2 | Transmembrane Protease Serine 2 |
| TRPML | Transient Receptor Potential Mucolipin |
| TGEV | Transmissible Gastroenteritis Virus |
| US NLM | United States National Library of Medicine |
| VSV | Vesicular Stomatitis Virus |
| WNV | West Nile Virus |
| ZIKV | Zika Virus |
References
- Forterre, P.; Prangishvili, D. The origin of viruses. Res. Microbiol. 2009, 160, 466–472. [Google Scholar] [CrossRef] [PubMed]
- Baltimore, D. Expression of animal virus genomes. Bacteriol. Rev. 1971, 35, 235–241. [Google Scholar] [CrossRef]
- Gebreyes, W.A.; Dupouy-Camet, J.; Newport, M.J.; Oliveira, C.J.B.; Schlesinger, L.S.; Saif, Y.M.; Kariuki, S.; Saif, L.J.; Saville, W.; Wittum, T.; et al. The Global One Health Paradigm: Challenges and Opportunities for Tackling Infectious Diseases at the Human, Animal, and Environment Interface in Low-Resource Settings. PLoS Neglected Trop. Dis. 2014, 8, e3257. [Google Scholar] [CrossRef]
- Ludwig, B.; Kraus, F.B.; Allwinn, R.; Doerr, H.W.; Preiser, W. Viral zoonoses—A threat under control? Intervirology 2003, 46, 71–78. [Google Scholar] [CrossRef]
- Tao, K.; Tzou, P.L.; Nouhin, J.; Gupta, R.K.; de Oliveira, T.; Kosakovsky Pond, S.L.; Fera, D.; Shafer, R.W. The biological and clinical significance of emerging SARS-CoV-2 variants. Nat. Rev. Genet. 2021, 22, 757–773. [Google Scholar] [CrossRef]
- V’kovski, P.; Kratzel, A.; Steiner, S.; Stalder, H.; Thiel, V. Coronavirus biology and replication: Implications for SARS-CoV-2. Nat. Rev. Microbiol. 2021, 19, 155–170. [Google Scholar] [CrossRef]
- Mlcochova, P.; Kemp, S.A.; Dhar, M.S.; Papa, G.; Meng, B.; Ferreira, I.A.T.M.; Datir, R.; Collier, D.A.; Albecka, A.; Singh, S.; et al. SARS-CoV-2 B.1.617.2 Delta variant replication and immune evasion. Nature 2021, 599, 114–119. [Google Scholar] [CrossRef]
- Karim, S.S.A.; Karim, Q.A. Omicron SARS-CoV-2 variant: A new chapter in the COVID-19 pandemic. Lancet 2021, 398, 2126–2128. [Google Scholar] [CrossRef] [PubMed]
- Ewen, C. Omicron Variant Puts. Nature 2021, 600, 21. [Google Scholar] [CrossRef] [PubMed]
- Carrasco-Hernandez, R.; Jácome, R.; Vidal, Y.L.; de León, S.P. Are RNA viruses candidate agents for the next global pandemic? A review. ILAR J. 2017, 58, 343–358. [Google Scholar] [CrossRef] [PubMed]
- Bimrew, S.; Abera, M. Review on Classification and Nomenclature of Viruses. Am. J. Life Sci. 2023, 11, 11–23. [Google Scholar] [CrossRef]
- Simmonds, P.; Adams, M.J.; Benkő, M.; Breitbart, M.; Brister, J.R.; Carstens, E.B.; Davison, A.J.; Delwart, E.; Gorbalenya, A.E.; Harrach, B.; et al. Consensus statement: Virus taxonomy in the age of metagenomics. Nat. Rev. Microbiol. 2017, 15, 161–168. [Google Scholar] [CrossRef]
- Silva, J.M.F.; Melo, F.L.; Elena, S.F.; Candresse, T.; Sabanadzovic, S.; Tzanetakis, I.E.; Blouin, A.G.; Villamor, D.E.V.; Mollov, D.; Constable, F.; et al. Virus classification based on in-depth sequence analyses and development of demarcation criteria using the Betaflexiviridae as a case study. J. Gen. Virol. 2022, 103, 001806. [Google Scholar] [CrossRef]
- Bhambhani, S.; Kondhare, K.R.; Giri, A.P. Diversity in Chemical Structures and Biological Properties of Plant Alkaloids. Molecules 2021, 26, 3374. [Google Scholar] [CrossRef]
- Nguyen, T.D.; Dang, T.T.T. Cytochrome P450 Enzymes as Key Drivers of Alkaloid Chemical Diversification in Plants. Front. Plant Sci. 2021, 12, 682181. [Google Scholar] [CrossRef] [PubMed]
- Kishimoto, S.; Sato, M.; Tsunematsu, Y.; Watanabe, K. Evaluation of biosynthetic pathway and engineered biosynthesis of alkaloids. Molecules 2016, 21, 1078. [Google Scholar] [CrossRef] [PubMed]
- Masci, V.L.; Bernardini, S.; Modesti, L.; Ovidi, E.; Tiezzi, A. Medicinal Plants as a Source of Alkaloids. Microorg. Sustain. 2019, 15, 85–113. [Google Scholar] [CrossRef]
- Dey, P.; Kundu, A.; Kumar, A.; Gupta, M.; Lee, B.M.; Bhakta, T.; Dash, S.; Kim, H.S. Analysis of Alkaloids (Indole Alkaloids, Isoquinoline Alkaloids, Tropane Alkaloids); Elsevier Inc.: Amsterdam, The Netherlands, 2020. [Google Scholar] [CrossRef]
- Wang, X.; Zhu, J.; Zhang, D.; Liu, G. Ribosomal control in RNA virus-infected cells. Front. Microbiol. 2022, 13, 1026887. [Google Scholar] [CrossRef]
- Cesaro, T.; Michiels, T. Inhibition of PKR by Viruses. Front. Microbiol. 2021, 12, 757238. [Google Scholar] [CrossRef]
- Lee, H.R.; Choi, U.Y.; Hwang, S.W.; Kim, S.; Jung, J.U. Viral inhibition of PRR-Mediated innate immune response: Learning from KSHV evasion strategies. Mol. Cells 2016, 39, 777–782. [Google Scholar] [CrossRef] [PubMed]
- Firth, A.E.; Brierley, I. Non-canonical translation in RNA viruses. J. Gen. Virol. 2012, 93, 1385–1409. [Google Scholar] [CrossRef] [PubMed]
- Faisal, S.; Badshah, S.L.; Kubra, B.; Emwas, A.H.; Jaremko, M. Alkaloids as potential antivirals. A comprehensive review. Nat. Prod. Bioprospecting 2023, 13, 4. [Google Scholar] [CrossRef] [PubMed]
- Abookleesh, F.L.; Al-Anzi, B.S.; Ullah, A. Potential Antiviral Action of Alkaloids. Molecules 2022, 27, 903. [Google Scholar] [CrossRef] [PubMed]
- Seteyen, A.L.S.; Girard-Valenciennes, E.; Septembre-Malaterre, A.; Gasque, P.; Guiraud, P.; Sélambarom, J. Anti-Alphaviral Alkaloids: Focus on Some Isoquinolines, Indoles and Quinolizidines. Molecules 2022, 27, 5080. [Google Scholar] [CrossRef]
- Gonzalez, B.L.; de Oliveira, N.C.; Ritter, M.R.; Tonin, F.S.; Melo, E.B.; Sanches, A.C.C.; Fernandez-Llimos, F.; Petruco, M.V.; de Mello, J.C.P.; Chierrito, D.; et al. The naturally-derived alkaloids as a potential treatment for COVID-19: A scoping review. Phytother. Res. 2022, 36, 2686–2709. [Google Scholar] [CrossRef]
- Kittakoop, P.; Darshana, D.; Sangsuwan, R.; Mahidol, C. Alkaloids and Alkaloid-Like Compounds are Potential Scaffolds of Antiviral Agents against SARS-CoV-2 (COVID-19) Virus. Heterocycles 2022, 105, 115. [Google Scholar] [CrossRef]
- Sharma, D.; Sharma, N.; Manchanda, N.; Prasad, S.K.; Sharma, P.C.; Thakur, V.K.; Rahman, M.M.; Dhobi, M. Bioactivity and In Silico Studies of Isoquinoline and Related Alkaloids as Promising Antiviral Agents: An Insight. Biomolecules 2022, 13, 17. [Google Scholar] [CrossRef]
- Xian, Y.; Zhang, J.; Bian, Z.; Zhou, H.; Zhang, Z.; Lin, Z.; Xu, H. Bioactive natural compounds against human coronaviruses: A review and perspective. Acta Pharm. Sin. B 2020, 10, 1163–1174. [Google Scholar] [CrossRef]
- Liao, Y.; Ye, Y.; Liu, M.; Liu, Z.; Wang, J.; Li, B.; Huo, L.; Zhuang, Y.; Chen, L.; Chen, J.; et al. Identification of N- and C-3-Modified Laudanosoline Derivatives as Novel Influenza PAN Endonuclease Inhibitors. J. Med. Chem. 2023, 66, 188–219. [Google Scholar] [CrossRef]
- George, A.; Reddy, A.G.K.; Satyanarayana, G.; Raghavendra, N.K. 1,2,3,4-Tetrahydroisoquinolines as inhibitors of HIV-1 integrase and human LEDGF/p75 interaction. Chem. Biol. Drug Des. 2018, 91, 1133–1140. [Google Scholar] [CrossRef]
- Wang, X.; Burdzhiev, N.T.; Hu, H.; Li, Y.; Li, J.; Lozanova, V.V.; Kandinska, M.I.; Wang, M. Novel Tetrahydroisoquinoline-Based Heterocyclic Compounds Efficiently Inhibit SARS-CoV-2 Infection In Vitro. Viruses 2023, 15, 502. [Google Scholar] [CrossRef] [PubMed]
- Majnooni, M.B.; Fakhri, S.; Bahrami, G.; Naseri, M.; Farzaei, M.H.; Echeverr, J. Alkaloids as Potential Phytochemicals against SARS-CoV-2: Approaches to the Associated Pivotal Mechanisms. Evid.-Based Complement. Altern. Med. 2021, 2021, 6632623. [Google Scholar] [CrossRef] [PubMed]
- Ogunyemi, O.M.; A Gyebi, G.; A Elfiky, A.; O Afolabi, S.; Ogunro, O.B.; Adegunloye, A.P.; Ibrahim, I.M. Alkaloids and flavonoids from African phytochemicals as potential inhibitors of SARS-CoV-2 RNA-dependent RNA polymerase: An in silico perspective. Antivir. Chem. Chemother. 2020, 28, 2040206620984076. [Google Scholar] [CrossRef] [PubMed]
- Cheng, F.; Murray, J.L.; Zhao, J.; Sheng, J.; Zhao, Z.; Rubin, D.H. Systems Biology-Based Investigation of Cellular Antiviral Drug Targets Identified by Gene-Trap Insertional Mutagenesis. PLoS Comput. Biol. 2016, 12, e1005074. [Google Scholar] [CrossRef] [PubMed]
- Zhou, P.; Lao, Z.; Long, H.; Pan, P.; Liao, F.; Zheng, W.; Li, Z.; Dai, J.; Liu, H.; Jiang, Y.; et al. The in vitro and in vivo antiviral effects of aloperine against Zika virus infection. J. Med. Virol. 2023, 95, e28913. [Google Scholar] [CrossRef]
- Merindol, N.; Martins, L.L.B.; Elfayres, G.; Custeau, A.; Berthoux, L.; Evidente, A.; Desgagné-Penix, I. Amaryllidaceae Alkaloids Screen Unveils Potent Anticoronaviral Compounds and Associated Structural Determinants. ACS Pharmacol. Transl. Sci. 2024, 7, 3527–3539. [Google Scholar] [CrossRef]
- Huang, L.; Yuen, T.T.-T.; Ye, Z.; Liu, S.; Zhang, G.; Chu, H.; Yue, J. Berbamine inhibits SARS-CoV-2 infection by compromising TRPMLs-mediated endolysosomal trafficking of ACE2. Signal Transduct. Target. Ther. 2021, 6, 2–4. [Google Scholar] [CrossRef]
- Wang, C.; Wang, T.; Dai, J.; Han, Y.; Hu, R.; Li, N.; Yang, Z.; Wang, J. Canthin-6-one analogs block Newcastle disease virus proliferation via suppressing the Akt and ERK pathways. Poult. Sci. 2024, 103, 103944. [Google Scholar] [CrossRef]
- Verma, D.; Mitra, D.; Kamboj, A.; Mahakur, B.; Chaudhary, P.; Shrivastav, R.; Janmeda, P.; Pant, K.; Mohapatra, P.K.D. Canthin-6-One 9-O-Beta-Glucopyranoside: A natural inhibitor of SARS-CoV-2 Canthin-6-One 9-O-Beta-Glucopyranoside: An inhibitor of SARS-CoV-2 (COVID 19) proteases PLpro and Mpro/3CLpro. Res. Sq. 2020, 2, 1–16. [Google Scholar] [CrossRef]
- Zhang, M.-Q.; Jia, X.; Cheng, C.-Q.; Wang, Y.-X.; Li, Y.-Y.; Kong, L.-D.; Li, Q.-Q.; Xie, F.; Yu, Y.-L.; He, Y.-T.; et al. Capsaicin functions as a selective degrader of STAT3 to enhance host resistance to viral infection. Acta Pharmacol. Sin. 2023, 44, 2253–2264. [Google Scholar] [CrossRef]
- Marois, I.; Cloutier, A.; Meunier, I.; Weingartl, H.M.; Cantin, A.M.; Richter, M.V. Inhibition of influenza virus replication by targeting broad host cell pathways. PLoS ONE 2014, 9, e110631. [Google Scholar] [CrossRef]
- Trischitta, P.; Tamburello, M.P.; Venuti, A.; Pennisi, R. Pseudovirus-Based Systems for Screening Natural Antiviral Agents: A Comprehensive Review. Int. J. Mol. Sci. 2024, 25, 5188. [Google Scholar] [CrossRef]
- Xia, B.; Zheng, L.; Li, Y.; Sun, W.; Liu, Y.; Li, L.; Pang, J.; Chen, J.; Li, J.; Cheng, H. The brief overview, antivirus and anti-SARS-CoV-2 activity, quantitative methods, and pharmacokinetics of cepharanthine: A potential small-molecule drug against COVID-19. Front. Pharmacol. 2023, 14, 1098972. [Google Scholar] [CrossRef]
- Liu, K.; Hong, B.; Wang, S.; Lou, F.; You, Y.; Hu, R.; Shafqat, A.; Fan, H.; Tong, Y. Pharmacological Activity of Cepharanthine. Molecules 2023, 28, 5019. [Google Scholar] [CrossRef]
- Ren, P.-X.; Shang, W.-J.; Yin, W.-C.; Ge, H.; Wang, L.; Zhang, X.-L.; Li, B.-Q.; Li, H.-L.; Xu, Y.-C.; Xu, E.H.; et al. A multi-targeting drug design strategy for identifying potent anti-SARS-CoV-2 inhibitors. Acta Pharmacol. Sin. 2022, 43, 483–493. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Xu, M.; Lee, E.M.; Gorshkov, K.; Shiryaev, S.A.; He, S.; Sun, W.; Cheng, Y.-S.; Hu, X.; Tharappel, A.M.; et al. Emetine inhibits Zika and Ebola virus infections through two molecular mechanisms: Inhibiting viral replication and decreasing viral entry. Cell Discov. 2018, 4, 31. [Google Scholar] [CrossRef]
- Ka, S.; Merindol, N.; Sow, A.A.; Singh, A.; Landelouci, K.; Plourde, M.B.; Pépin, G.; Masi, M.; Di Lecce, R.; Evidente, A.; et al. Amaryllidaceae Alkaloid Cherylline Inhibits the Replication of Dengue and Zika Viruses. Antimicrob. Agents Chemother. 2021, 65, e0039821. [Google Scholar] [CrossRef] [PubMed]
- Ren, J.; Zeng, W.; Jiang, C.; Li, C.; Zhang, C.; Cao, H.; Li, W.; He, Q. Inhibition of Porcine Epidemic Diarrhea Virus by Cinchonine via Inducing Cellular Autophagy. Front. Cell. Infect. Microbiol. 2022, 12, 856711. [Google Scholar] [CrossRef]
- Hegazy, A.; Soltane, R.; Alasiri, A.; Mostafa, I.; Metwaly, A.M.; Eissa, I.H.; Mahmoud, S.H.; Allayeh, A.K.; Shama, N.M.A.; Khalil, A.A.; et al. Anti-rheumatic colchicine phytochemical exhibits potent antiviral activities against avian and seasonal Influenza A viruses (IAVs) via targeting different stages of IAV replication cycle. BMC Complement. Med. Ther. 2024, 24, 49. [Google Scholar] [CrossRef] [PubMed]
- Bailly, B.; Richard, C.-A.; Sharma, G.; Wang, L.; Johansen, L.; Cao, J.; Pendharkar, V.; Sharma, D.-C.; Galloux, M.; Wang, Y.; et al. Targeting human respiratory syncytial virus transcription anti-termination factor M2-1 to inhibit in vivo viral replication. Sci. Rep. 2016, 6, 25806. [Google Scholar] [CrossRef]
- Diot, C.; Richard, C.-A.; Risso-Ballester, J.; Martin, D.; Fix, J.; Eléouët, J.-F.; Sizun, C.; Rameix-Welti, M.-A.; Galloux, M. Hardening of Respiratory Syncytial Virus Inclusion Bodies by Cyclopamine Proceeds through Perturbation of the Interactions of the M2-1 Protein with RNA and the P Protein. Int. J. Mol. Sci. 2023, 24, 13862. [Google Scholar] [CrossRef]
- Fix, J.; Descamps, D.; Galloux, M.; Ferret, C.; Bouguyon, E.; Zohari, S.; Näslund, K.; Hägglund, S.; Altmeyer, R.; Valarcher, J.-F.; et al. Screening antivirals with a mCherry-expressing recombinant bovine respiratory syncytial virus: A proof of concept using cyclopamine. Vet. Res. 2023, 54, 36. [Google Scholar] [CrossRef]
- Li, K.; Wang, H.; Chen, T.; Wang, X.; Wang, X.; Zhong, M.; Gao, X.; Hao, Z. Dehydroevodiamine inhibits PEDV through regulateing ERK1/2 MAPK pathway in Vero cells. Virology 2024, 598, 110166. [Google Scholar] [CrossRef]
- Hussain, S.; Miller, J.L.; Harvey, D.J.; Gu, Y.; Rosenthal, P.B.; Zitzmann, N.; McCauley, J.W. Strain-specific antiviral activity of iminosugars against human influenza A viruses. J. Antimicrob. Chemother. 2015, 70, 136–152. [Google Scholar] [CrossRef]
- Perera, N.; Brun, J.; Alonzi, D.S.; Tyrrell, B.E.; Miller, J.L.; Zitzmann, N. Antiviral effects of deoxynojirimycin (DNJ)-based iminosugars in dengue virus-infected primary dendritic cells. Antivir. Res. 2022, 199, 105269. [Google Scholar] [CrossRef]
- Bhushan, G.; Lim, L.; Bird, I.; Chothe, S.K.; Nissly, R.H.; Kuchipudi, S.V. Iminosugars With Endoplasmic Reticulum α-Glucosidase Inhibitor Activity Inhibit ZIKV Replication and Reverse Cytopathogenicity in vitro. Front. Microbiol. 2020, 11, 531. [Google Scholar] [CrossRef] [PubMed]
- Khandelwal, N.; Chander, Y.; Rawat, K.D.; Riyesh, T.; Nishanth, C.; Sharma, S.; Jindal, N.; Tripathi, B.N.; Barua, S.; Kumar, N. Emetine inhibits replication of RNA and DNA viruses without generating drug-resistant virus variants. Antivir. Res. 2017, 144, 196–204. [Google Scholar] [CrossRef] [PubMed]
- Valipour, M. Different Aspects of Emetine’s Capabilities as a Highly Potent SARS-CoV-2 Inhibitor against COVID-19. ACS Pharmacol. Transl. Sci. 2022, 5, 387–399. [Google Scholar] [CrossRef] [PubMed]
- Bleasel, M.D.; Peterson, G.M. Emetine, ipecac, ipecac alkaloids and analogues as potential antiviral agents for coronaviruses. Pharmaceuticals 2020, 13, 51. [Google Scholar] [CrossRef]
- Wei, W.; Du, H.; Shao, C.; Zhou, H.; Lu, Y.; Yu, L.; Wan, H.; He, Y. Screening of antiviral components of Ma Huang Tang and investigation on the ephedra alkaloids efficacy on influenza virus type A. Front. Pharmacol. 2019, 10, 961. [Google Scholar] [CrossRef]
- Yang, S.; Yang, X.; Wang, Z.; Li, W.; Cao, R.; Zhong, W. Fangchinoline Inhibits Zika Virus by Disrupting Virus Internalization. ACS Infect. Dis. 2024, 10, 4066–4072. [Google Scholar] [CrossRef] [PubMed]
- Wan, Z.; Lu, Y.; Liao, Q.; Wu, Y.; Chen, X. Fangchinoline inhibits human immunodeficiency virus type 1 replication by interfering with gp160 proteolytic processing. PLoS ONE 2012, 7, e39225. [Google Scholar] [CrossRef]
- Zhang, Q.Y.; Li, J.-Q.; Li, Q.; Zhang, Y.; Zhang, Z.-R.; Li, X.-D.; Zhang, H.-Q.; Deng, C.-L.; Yang, F.-X.; Xu, Y.; et al. Identification of fangchinoline as a broad-spectrum enterovirus inhibitor through reporter virus based high-content screening. Virol. Sin. 2024, 39, 301–308. [Google Scholar] [CrossRef]
- Hegazy, A.; Mahmoud, S.H.; Elshaier, Y.A.M.M.; Shama, N.M.A.; Nasr, N.F.; Ali, M.A.; El-Shazly, A.M.; Mostafa, I.; Mostafa, A. Antiviral activities of plant-derived indole and β-carboline alkaloids against human and avian influenza viruses. Sci. Rep. 2023, 13, 1612. [Google Scholar] [CrossRef] [PubMed]
- Dahal, S.; Clayton, K.; Cabral, T.; Cheng, R.; Jahanshahi, S.; Ahmed, C.; Koirala, A.; Ocando, A.V.; Malty, R.; Been, T.; et al. On a path toward a broad-spectrum anti-viral: Inhibition of HIV-1 and coronavirus replication by SR kinase inhibitor harmine. J. Virol. 2023, 97, e0039623. [Google Scholar] [CrossRef]
- Kaur, P.; Thiruchelvan, M.; Lee, R.C.H.; Chen, H.; Chen, K.C.; Ng, M.L.; Chu, J.J.H. Inhibition of Chikungunya virus replication by harringtonine, a novel antiviral that suppresses viral protein expression. Antimicrob. Agents Chemother. 2013, 57, 155–167. [Google Scholar] [CrossRef] [PubMed]
- Lai, Z.Z.; Ho, Y.J.; Lu, J.W. Harringtonine inhibits zika virus infection through multiple mechanisms. Molecules 2020, 25, 4082. [Google Scholar] [CrossRef]
- Yang, Z.; Fu, X.; Zhao, Y.; Li, X.; Long, J.; Zhang, L. Molecular insights into the inhibition mechanism of harringtonine against essential proteins associated with SARS-CoV-2 entry. Int. J. Biol. Macromol. 2023, 240, 124352. [Google Scholar] [CrossRef]
- Dong, H.J.; Wang, Z.-H.; Meng, W.; Li, C.-C.; Hu, Y.-X.; Zhou, L.; Wang, X.-J. The natural compound homoharringtonine presents broad antiviral activity in vitro and in vivo. Viruses 2018, 10, 601. [Google Scholar] [CrossRef]
- Gong, M.; Li, S.; Xie, Y.; Zhao, F.; Shao, J.; Zhang, Y.; Wang, W.; Chang, H. Inhibitory effects of homoharringtonine on foot and mouth disease virus in vitro. J. Med. Virol. 2019, 91, 1595–1601. [Google Scholar] [CrossRef]
- Harisha, K.R.; Kailaje, V.; Kondreddi, R.R.; Gudla, C.S.; Singh, S.; Ramakrishnaiah, S.; Isloor, S.; Narayanan, S.; Shandil, R.K.; Rudramurthy, G.R. Homoharringtonine Inhibits CVS-11 and Clinical Isolates of Rabies Virus In Vitro: Identified via High-Throughput Screening of an FDA-Approved Drug Library. Viruses 2025, 17, 945. [Google Scholar] [CrossRef]
- Neerukonda, S.N. A Review on SARS-CoV-2 Virology, Pathophysiology. Pathogens 2020, 9, 426. [Google Scholar] [CrossRef]
- Jie, C.; Luo, Z.; Chen, H.; Wang, M.; Yan, C.; Mao, Z.-F.; Xiao, G.-K.; Kurihara, H.; Li, Y.-F.; He, R.-R. Indirubin, a bisindole alkaloid from Isatis indigotica, reduces H1N1 susceptibility in stressed mice by regulating MAVS signaling. Oncotarget 2017, 8, 105615–105629. [Google Scholar] [CrossRef]
- Chang, S.J.; Chang, Y.C.; Lu, K.Z.; Tsou, Y.Y.; Lin, C.W. Antiviral activity of Isatis indigotica extract and its derived indirubin against Japanese encephalitis virus. Evid.-Based Complement. Altern. Med. 2012, 2012, 925830. [Google Scholar] [CrossRef]
- Medina-Moreno, S.; Dowling, T.C.; Zapata, J.C.; Le, N.M.; Sausville, E.; Bryant, J.; Redfield, R.R.; Heredia, A. Targeting of CDK9 with indirubin 3′-monoxime safely and durably reduces HIV viremia in chronically infected humanized mice. PLoS ONE 2017, 12, e0183425. [Google Scholar] [CrossRef] [PubMed]
- Jin, Y.H.; Min, J.S.; Jeon, S.; Lee, J.; Kim, S.; Park, T.; Park, D.; Jang, M.S.; Park, C.M.; Song, J.H.; et al. Lycorine, a non-nucleoside RNA dependent RNA polymerase inhibitor, as potential treatment for emerging coronavirus infections. Phytomedicine 2021, 86, 153440. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Lao, Z.; Xu, J.; Li, Z.; Long, H.; Li, D.; Lin, L.; Liu, X.; Yu, L.; Liu, W.; et al. Antiviral activity of lycorine against Zika virus in vivo and in vitro. Virology 2020, 546, 88–97. [Google Scholar] [CrossRef] [PubMed]
- Li, N.; Wang, Z.; Wang, R.; Zhang, Z.-R.; Zhang, Y.-N.; Deng, C.-L.; Zhang, B.; Shang, L.-Q.; Ye, H.-Q. In Vitro Inhibition of Alphaviruses by Lycorine. Virol. Sin. 2021, 36, 1465–1474. [Google Scholar] [CrossRef]
- Narayanan, A.; Narwal, M.; Majowicz, S.A.; Varricchio, C.; Toner, S.A.; Ballatore, C.; Brancale, A.; Murakami, K.S.; Jose, J. Identification of SARS-CoV-2 inhibitors targeting Mpro and PLpro using in-cell-protease assay. Commun. Biol. 2022, 5, 169. [Google Scholar] [CrossRef]
- Fielding, B.C.; da Silva Maia Bezerra Filho, C.; Ismail, N.S.M.; de Sousa, D.P. Alkaloids: Therapeutic potential against human coronaviruses. Molecules 2020, 25, 5496. [Google Scholar] [CrossRef]
- Pan, Q.M.; Li, Y.H.; Hua, J.; Huang, F.P.; Wang, H.S.; Liang, D. Antiviral Matrine-Type Alkaloids from the Rhizomes of Sophora tonkinensis. J. Nat. Prod. 2015, 78, 1683–1688. [Google Scholar] [CrossRef]
- Qiao, W.T.; Yao, X.; Lu, W.-H.; Zhang, Y.-Q.; Malhi, K.K.; Li, H.-X.; Li, J.-L. Matrine exhibits antiviral activities against PEDV by directly targeting Spike protein of the virus and inducing apoptosis via the MAPK signaling pathway. Int. J. Biol. Macromol. 2024, 270, 132408. [Google Scholar] [CrossRef] [PubMed]
- McMahon, J.B.; Currens, M.J.; Gulakowski, R.J.; Buckheit, R.W.; Lackman-Smith, C.; Hallock, Y.F.; Boyd, M.R. Michellamine B, a novel plant alkaloid, inhibits human immunodeficiency virus-induced cell killing by at least two distinct mechanism. Antimicrob. Agents Chemother. 1995, 39, 484–488. [Google Scholar] [CrossRef]
- Yang, Y.; Yang, P.; Huang, C.; Wu, Y.; Zhou, Z.; Wang, X.; Wang, S. Inhibitory effect on SARS-CoV-2 infection of neferine by blocking Ca2+-dependent membrane fusion. J. Med. Virol. 2021, 93, 5825–5832. [Google Scholar] [CrossRef]
- Yang, D.D.; Chutiwitoonchai, N.; Wang, F.; Tian, P.; Sureram, S.; Lei, X.; Mahidol, C.; Ruchirawat, S.; Kittakoop, P. Effects of organic salts of virucidal and antiviral compounds from Nelumbo nucifera and Kaempferia parviflora against SARS-CoV-2. Sci. Rep. 2025, 15, 6380. [Google Scholar] [CrossRef]
- Kumar, A.; Sharma, M.; Richardson, C.D.; Kelvin, D.J. Potential of Natural Alkaloids From Jadwar (Delphinium denudatum) as Inhibitors Against Main Protease of COVID-19: A Molecular Modeling Approach. Front. Mol. Biosci. 2022, 9, 898874. [Google Scholar] [CrossRef]
- Nag, A.; Chowdhury, R.R. Piperine, an alkaloid of black pepper seeds can effectively inhibit the antiviral enzymes of Dengue and Ebola viruses, an in silico molecular docking study. Virusdisease 2020, 31, 308–315. [Google Scholar] [CrossRef] [PubMed]
- Pareek, A.; Kumar, R.; Mudgal, R.; Neetu, N.; Sharma, M.; Kumar, P.; Tomar, S. Alphavirus antivirals targeting RNA-dependent RNA polymerase domain of nsP4 divulged using surface plasmon resonance. FEBS J. 2022, 289, 4901–4924. [Google Scholar] [CrossRef]
- Li, D.; Chengzhi, C.; Xiaoyin, C.; Zhongping, W. Pseudoephedrine protects mice from infection of H1N1 virus. Int. J. Clin. Virol. 2020, 4, 14–20. [Google Scholar] [CrossRef]
- Yu, S.; Chen, Y.; Xiang, Y.; Lin, H.; Wang, M.; Ye, W.; Zhang, P.; Chen, H.; Lin, G.; Zhu, Y.; et al. Pseudoephedrine and its derivatives antagonize wild and mutated severe acute respiratory syndrome-CoV-2 viruses through blocking virus invasion and antiinflammatory effect. Phytother. Res. 2021, 35, 5847–5860. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Pei, R.-J.; Li, H.; Ma, X.-N.; Zhou, Y.; Zhu, F.-H.; He, P.-L.; Tang, W.; Zhang, Y.-C.; Xiong, J.; et al. Identification of SARS-CoV-2 entry inhibitors among already approved drugs. Acta Pharmacol. Sin. 2021, 42, 1347–1353. [Google Scholar] [CrossRef]
- Persoons, L.; Vanderlinden, E.; Vangeel, L.; Wang, X.; Do, N.D.T.; Foo, S.-Y.C.; Leyssen, P.; Neyts, J.; Jochmans, D.; Schols, D.; et al. Broad spectrum anti-coronavirus activity of a series of anti-malaria quinoline analogues. Antivir. Res. 2021, 193, 105127. [Google Scholar] [CrossRef]
- Mamidala, E.; Davella, R.; Gurrapu, S.; Shivakrishna, P. In Silico Identification of Clinically Approved Medicines Against the main Protease of SARS-CoV-2—A Causative Agent of COVID-19. Int. J. Life Sci. Pharma Res. 2022, 11, 107–122. [Google Scholar] [CrossRef]
- Malakar, S.; Sreelatha, L.; Dechtawewat, T.; Noisakran, S.; Yenchitsomanus, P.-T.; Chu, J.J.H.; Limjindaporn, T. Drug repurposing of quinine as antiviral against dengue virus infection. Virus Res. 2018, 255, 171–178. [Google Scholar] [CrossRef] [PubMed]
- D’alessandro, S.; Scaccabarozzi, D.; Signorini, L.; Perego, F.; Ilboudo, D.P.; Ferrante, P.; Delbue, S. The use of antimalarial drugs against viral infection. Microorganisms 2020, 8, 85. [Google Scholar] [CrossRef]
- Große, M.; Ruetalo, N.; Layer, M.; Hu, D.; Businger, R.; Rheber, S.; Setz, C.; Rauch, P.; Auth, J.; Fröba, M.; et al. Quinine inhibits infection of human cell lines with SARS-CoV-2. Viruses 2021, 13, 647. [Google Scholar] [CrossRef]
- Lin, S.; Wang, X.; Guo, H.; Dai, N.; Tang, R.W.-L.; Lee, H.C.; Leung, K.W.; Dong, T.T.-X.; Webb, S.E.; Miller, A.L.; et al. The Ethanol Extract of Evodiae Fructus and Its Ingredient, Rutaecarpine, Inhibit Infection of SARS-CoV-2 and Inflammatory Responses. Int. J. Mol. Sci. 2023, 24, 762. [Google Scholar] [CrossRef] [PubMed]
- Bhattacharjee, A.; Chaudhuri, R.; Dash, J.J.; Saha, M.; Choudhury, L.; Roy, S. Pre-treatment with Scopolamine Naturally Suppresses Japanese Encephalitis Viral Load in Embryonated Chick Through Regulation of Multiple Signaling Pathways. Appl. Biochem. Biotechnol. 2021, 193, 1654–1674. [Google Scholar] [CrossRef] [PubMed]
- Jin, Z.; Yang, L.; Ding, G.; Yang, G.; Han, Y.; Zhang, X.; Li, W. Sophocarpine against enterovirus 71 in vitro. Exp. Ther. Med. 2017, 14, 3792–3797. [Google Scholar] [CrossRef]
- Ren, G.; Ding, G.; Zhang, H.; Wang, H.; Jin, Z.; Yang, G.; Han, Y.; Zhang, X.; Li, G.; Li, W. Antiviral activity of sophoridine against enterovirus 71 in vitro. J. Ethnopharmacol. 2019, 236, 124–128. [Google Scholar] [CrossRef]
- Lixia, H.; Jun, C.; Song, H.; FaHu, Y.; Jinwen, T. Neuroprotective effect of (-)-tetrahydropalmatine in Japanese encephalitis virus strain GP-78 infected mouse model. Microb. Pathog. 2018, 114, 197–203. [Google Scholar] [CrossRef]
- Kim, D.E.; Min, J.S.; Jang, M.S.; Lee, J.Y.; Shin, Y.S.; Park, C.M.; Song, J.H.; Kim, H.R.; Kim, S.; Jin, Y.-H.; et al. Natural bis-benzylisoquinoline alkaloids-tetrandrine, fangchinoline, and cepharanthine, inhibit human coronavirus oc43 infection of mrc-5 human lung cells. Biomolecules 2019, 9, 696. [Google Scholar] [CrossRef]
- Liu, J.; Wang, F.; Wang, X.; Fan, S.; Li, Y.; Xu, M.; Hu, H.; Liu, K.; Zheng, B.; Wang, L.; et al. Antiviral effects and tissue exposure of tetrandrine against SARS-CoV-2 infection and COVID-19. MedComm 2023, 4, e206. [Google Scholar] [CrossRef]
- Islam, M.T.; Sarkar, C.; El-Kersh, D.M.; Jamaddar, S.; Uddin, S.J.; Shilpi, J.A.; Mubarak, M.S. Natural products and their derivatives against coronavirus: A review of the non-clinical and pre-clinical data. Phytother. Res. 2020, 34, 2471–2492. [Google Scholar] [CrossRef]
- Yang, C.W.; Lee, Y.-Z.; Kang, I.-J.; Barnard, D.L.; Jan, J.-T.; Lin, D.; Huang, C.-W.; Yeh, T.-K.; Chao, Y.-S.; Lee, S.-J. Identification of phenanthroindolizines and phenanthroquinolizidines as novel potent anti-coronaviral agents for porcine enteropathogenic coronavirus transmissible gastroenteritis virus and human severe acute respiratory syndrome coronavirus. Antivir. Res. 2010, 88, 160–168. [Google Scholar] [CrossRef]
- Wang, Y.; Lee, S.; Ha, Y.; Lam, W.; Chen, S.-R.; Dutschman, G.E.; Gullen, E.A.; Grill, S.P.; Cheng, Y.; Fürstner, A.; et al. Tylophorine Analogs Allosterically Regulates Heat Shock Cognate Protein 70 And Inhibits Hepatitis C Virus Replication. Sci. Rep. 2017, 7, 10037. [Google Scholar] [CrossRef]
- Mani, J.S.; Johnson, J.B.; Steel, J.C.; Broszczak, D.A.; Neilsen, P.M.; Walsh, K.B.; Naiker, M. Natural product-derived phytochemicals as potential agents against coronaviruses: A review. Virus Res. 2020, 284, 197989. [Google Scholar] [CrossRef] [PubMed]
- Akan, E.; Chang-Liu, C.M.; Watanabe, J.; Ishizawa, K.; Woloschak, G.E. The effects of vinblastine on the expression of human immunodeficiency virus type 1 long terminal repeat. Leuk. Res. 1997, 21, 459–464. [Google Scholar] [CrossRef] [PubMed]
- Qing, Z.-X.; Yang, P.; Tang, Q.; Cheng, P.; Liu, X.-B.; Zheng, Y.-J.; Liu, Y.-S.; Zeng, J.-G. Isoquinoline Alkaloids and Their Antiviral, Antibacterial, and Antifungal Activities and Structure-activity Relationship. Curr. Org. Chem. 2017, 21, 1920–1934. [Google Scholar] [CrossRef]
- Zheng, X.; Wu, F.; Lin, X.; Shen, L.; Feng, Y. Developments in drug delivery of bioactive alkaloids derived from traditional Chinese medicine. Drug Deliv. 2018, 25, 398–416. [Google Scholar] [CrossRef]
- Gidwani, B.; Bhairam, M.; Shukla, S.S.; Verma, H.; Pandey, R.K. Herbal Bioenhancers in Pharmaceutical Drug Delivery: Mechanisms, Challenges, and Future Innovations. Chem. Biodivers. 2025, 22, e202500760. [Google Scholar] [CrossRef] [PubMed]
- Onoue, S.; Yamada, S.; Chan, H.K. Nanodrugs: Pharmacokinetics and safety. Int. J. Nanomed. 2014, 9, 1025–1037. [Google Scholar] [CrossRef]
- Gabrielsen, B.; Monath, T.P.; Huggins, J.W.; Kefauver, D.F.; Pettit, G.R.; Groszek, G.; Hollingshead, M.; Kirsi, J.J.; Shannon, W.M.; Schubert, E.M.; et al. Antiviral (RNA) activity of selected Amaryllidaceae isoquinoline const ituents and synthesis of related substances. J. Nat. Prod. 1992, 55, 1569–1581. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Li, J.Q.; Kim, Y.J.; Wu, J.; Wang, Q.; Hao, Y. In vivo and in vitro antiviral effects of berberine on influenza virus. Chin. J. Integr. Med. 2011, 17, 444–452. [Google Scholar] [CrossRef]
- Wang, Y.X.; Yang, L.; Wang, H.-Q.; Zhao, X.-Q.; Liu, T.; Li, Y.-H.; Zeng, Q.-X.; Li, Y.-H.; Song, D.-Q. synthesis and evolution of berberine derivatives as a new class of antiviral agents against enterovirus 71 through the MEK/ERK pathway and autophagy. Molecules 2018, 23, 2084. [Google Scholar] [CrossRef] [PubMed]
- Shao, J.; Zeng, D.; Tian, S.; Liu, G.; Fu, J. Identification of the natural product berberine as an antiviral drug. AMB Express 2020, 10, 164. [Google Scholar] [CrossRef]
- Ratanakomol, T.; Roytrakul, S.; Wikan, N.; Smith, D.R. Berberine inhibits dengue virus through dual mechanisms. Molecules 2021, 26, 5501. [Google Scholar] [CrossRef]
- Botwina, P.; Owczarek, K.; Rajfur, Z.; Ochman, M.; Urlik, M.; Nowakowska, M.; Szczubiałka, K.; Pyrc, K. Berberine Hampers Influenza A Replication through. Viruses 2020, 12, 344. [Google Scholar] [CrossRef]
- Enkhtaivan, G.; Muthuraman, P.; Kim, D.H.; Mistry, B. Discovery of berberine based derivatives as anti-influenza agent through blocking of neuraminidase. Bioorg. Med. Chem. 2017, 25, 5185–5193. [Google Scholar] [CrossRef]
- Varghese, F.S.; Kaukinen, P.; Gläsker, S.; Bespalov, M.; Hanski, L.; Wennerberg, K.; Kümmerer, B.M.; Ahola, T. Discovery of berberine, abamectin and ivermectin as antivirals against chikungunya and other alphaviruses. Antivir. Res. 2016, 126, 117–124. [Google Scholar] [CrossRef]
- Nguyen, C.Q.; Nguyen, T.H.M.; Nguyen, T.T.T.; Bui, T.B.H.; Nguyen, T.T.; Huynh, N.T.; Le, T.D.; Nguyen, T.M.P.; Nguyen, D.T.; Nguyen, M.T.; et al. Designs, Synthesis, Docking Studies, and Biological Evaluation of Novel Berberine Derivatives Targeting Zika Virus. J. Chem. 2021, 2021, 5567111. [Google Scholar] [CrossRef]
- Españo, E.; Kim, J.; Lee, C.; Webster, R.G.; Webby, R.J. Chelerythrine as an anti-Zika virus agent: Therapeutic potential and mode of action. bioRxiv 2022. [Google Scholar] [CrossRef]
- Loe, M.W.C.; Lee, R.C.H.; Chin, W.-X.; Min, N.; Teo, Z.Y.; Ho, S.X.; Yi, B.; Chu, J.J.H. Chelerythrine chloride inhibits Zika virus infection by targeting the viral NS4B protein. Antivir. Res. 2023, 219, 105732. [Google Scholar] [CrossRef] [PubMed]
- Guo, W.; Lu, X.; Liu, B.; Yan, H.; Feng, J. Anti-TMV activity and mode of action of three alkaloids isolated from Chelidonium majus. Pest Manag. Sci. 2021, 77, 510–517. [Google Scholar] [CrossRef]
- Orhan, I.; Özçelik, B.; Karaoǧlu, T.; Şener, B. Antiviral and antimicrobial profiles of selected isoquinoline alkaloids from Fumaria and Corydalis species. Z. Naturforschung Sect. C J. Biosci. 2007, 62, 19–26. [Google Scholar] [CrossRef] [PubMed]
- Dai, J.P.; Wang, Q.-W.; Su, Y.; Gu, L.-M.; Deng, H.-X.; Chen, X.-X.; Li, W.-Z.; Li, K.-S. Oxymatrine inhibits influenza a virus replication and inflammation via TLR4, p38 MAPK and NF-κB pathways. Int. J. Mol. Sci. 2018, 19, 965. [Google Scholar] [CrossRef]
- Regulation, I. Oxymatrine Modulation of TLR3 Signaling: A Dual-Action. Molecules 2024, 29, 1945. [Google Scholar] [CrossRef]
- Chen, N.; Liu, Y.H.; Liu, X.J.; Chen, Y.R.; Guo, Y.H.; Liu, M. Oxymatrine inhibits target cell infection in the HCVcc system. Zhonghua Gan Zang Bing Za Zhi 2016, 24, 40–45. [Google Scholar] [CrossRef]
- Zhang, W.; Chen, Y.; Yang, F.; Zhang, H.; Su, T.; Wang, J.; Zhang, Y.; Song, X. Antiviral effect of palmatine against infectious bronchitis virus through regulation of NF-κB/IRF7/JAK-STAT signalling pathway and apoptosis. Br. Poult. Sci. 2024, 65, 119–128. [Google Scholar] [CrossRef]
- Jia, F.; Zou, G.; Fan, J.; Yuan, Z. Identification of palmatine as an inhibitor of West Nile virus. Arch. Virol. 2010, 155, 1325–1329. [Google Scholar] [CrossRef]
- Ho, Y.J.; Lu, J.W.; Huang, Y.L.; Lai, Z.Z. Palmatine inhibits Zika virus infection by disrupting virus binding, entry, and stability. Biochem. Biophys. Res. Commun. 2019, 518, 732–738. [Google Scholar] [CrossRef] [PubMed]
- Ke, Q.; Duan, K.; Cheng, Y.; Xu, S.; Xiao, S.; Fang, L. Sanguinarine Exhibits Antiviral Activity against Porcine Reproductive and Respiratory Syndrome Virus via Multisite Inhibition Mechanisms. Viruses 2023, 15, 688. [Google Scholar] [CrossRef]
- Bonincontro, G.; Scuderi, S.A.; Marino, A.; Simonetti, G. Synergistic Effect of Plant Compounds in Combination with Conventional Antimicrobials against Biofilm of Staphylococcus aureus, Pseudomonas aeruginosa, and Candida spp. Pharmaceuticals 2023, 16, 1531. [Google Scholar] [CrossRef]
- Ng, Y.P.; Or, T.C.T.; Ip, N.Y. Plant alkaloids as drug leads for Alzheimer’s disease. Neurochem. Int. 2015, 89, 260–270. [Google Scholar] [CrossRef]
- Hukkanen, J.; Jacob, P.; Benowitz, N.L. Metabolism and disposition kinetics of nicotine. Pharmacol. Rev. 2005, 57, 79–115. [Google Scholar] [CrossRef]
- Sweilam, S.H.; Alqarni, M.H.; Youssef, F.S. Antimicrobial Alkaloids from Marine-Derived Fungi as Drug Leads versus COVID-19 Infection: A Computational Approach to Explore their Anti-COVID-19 Activity and ADMET Properties. Evid.-Based Complement. Altern. Med. 2022, 2022, 5403757. [Google Scholar] [CrossRef] [PubMed]
- Linnakoski, R.; Reshamwala, D.; Veteli, P.; Cortina-Escribano, M.; Vanhanen, H.; Marjomäki, V. Antiviral agents from fungi: Diversity, mechanisms and potential applications. Front. Microbiol. 2018, 9, 2325. [Google Scholar] [CrossRef]
- Zhou, H.; Wang, W.; Cai, L.; Yang, T. Potentiation and Mechanism of Berberine as an Antibiotic Adjuvant Against Multidrug-Resistant Bacteria. Infect. Drug Resist. 2023, 16, 7313–7326. [Google Scholar] [CrossRef]
- Tilaoui, M.; Mouse, H.A.; Zyad, A. Update and New Insights on Future Cancer Drug Candidates From Plant-Based Alkaloids. Front. Pharmacol. 2021, 12, 719694. [Google Scholar] [CrossRef] [PubMed]
- Sandor, A.M.; Sturdivant, M.S.; Ting, J.P.Y. Influenza Virus and SARS-CoV-2 Vaccines. J. Immunol. 2021, 206, 2509–2520. [Google Scholar] [CrossRef]

| Compound & Origin | References | Study Type | Virus | Mechanisms | Values |
|---|---|---|---|---|---|
| 1. 1.2.3.4-Tetrahydroisoquinolines derivatives (From: e.g., Papaveraceae family)  | Y. Liao et al. 2023 [30] | In vitro, in silico and in vivo | IAV H1N1 H5N1 H3N2 Influenza B virus (IBV) | Mechanistic studies demonstrated that compound 35 could bind tightly to the PAN endonuclease of RNA-dependent RNA polymerase, thus blocking the viral replication to exert antiviral activity. | IC50 for compound 35 = 0.20 ± 0.01 µM EC50 = 0.88 µM SI = 113.1 |
| George, A. et al. (2018) [31] | In vitro | HIV-1 | Inhibit the LEDGF/p75- IN | Compound 6d: IC50 of ~10 μM | |
| Wang, X. et al. (2023) [32] | In vitro and in silico | SARS-CoV-2 | Tt mainly inhibited the post-entry viral replication in both Vero E6 and Calu-3 cells. | Compound trans-1 EC50 = 2.78 µM SI > 71.94 | |
| 2. 7-Methoxycryptopleurine (From: Menispermaceae family)  | Majnooni, M. et al. (2001) [33] | Review | SARS-CoV-2 | Blocking the S and N proteins, 3CLpro inhibitor | EC50 = 58 nM |
| 3. 10-Hydroxyusambarensine (From: Strychnos usambarensis, Loganiaceae family) 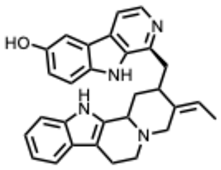 | Ogunyemi, O.M. et al. (2020) [34] | In silico | SARS-CoV-2 | Strong binding affinity to the RNA-dependent RNA polymerase (RdRp) | AutoDock version 4.2 programVina score: 10.1 |
| 4. Ajmaline (From: Rauwolfia serpentina, Apocynaceae family) 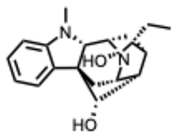 | Cheng, F. et al. (2016) [35] | Computational biology | EBOV | Ajmaline predicted to up-regulates expression of several important Ebola-related genes, such as MERTK, FURIN, TYRO3, FURIN, and CTSB | q = 0.002 |
| 5. Aloperine and derivatives (From: Sophora alopecuroides/Sophora flavescens, Fabaceae family) 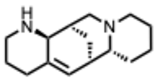 | Cheng, F. et al. (2016) [35] | In vitro and in vivo | SARS-CoV-2 | Inhibiting host cathepsin B activity and anti-cytokine effects | Compound 8a: EC 50 = 39.1 µM SI > 6.8 |
| Zhou, P. et al. (2023) [36] | In vitro, in silico and in vivo | ZIKV | Targeting its RNA-dependent RNA polymerase (RdRp) | EC50: from 2.98–6.036 µM depending on the cells line SI: 31.72–66.95 depending on the cells line Aloperine administration resulted in an improved survival rate in mice and reduced viremia | |
| 6. Amarbellisine (From: Amaryllidaceae family) 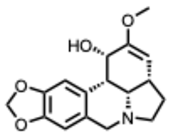 | Merindol, N. et al. (2024) [37] | In vitro | HCoV−OC43 HCoV-OC43 | Potential inhibition of viral replication | SI = 60 EC50 = 0.2 µM |
| 7. Berbamine (From: Berberis spp., Berberidaceae family)  | Huang, L. et al. (2021) [38] | In vitro | SARS-CoV-2 | Compromising TRPMLs-mediated endolysosomal trafficking of ACE2 | SARS-CoV-2: 2.35 ± 0.92 µM |
| 8. Canthin-6-one and derivatives (From: Rutaceae family)  | Wang, C. et al. (2024) [39] | In vitro | Newcastle disease virus (NDV) | Entry inhibition via Akt/ERK pathway; COX-2 induction | Analogue compounds: IC50 = 5.26–11.76 μM |
| Verma, D. et al. (2020) [40] | In silico | SARS-CoV-2 | Predicted binding to Mpro and PLpro, potential protease inhibition | Canthin-6-one 9-O-β-glucopyranoside PLpro: −9.4 kcal/mol Mpro: −8.5 kcal/mol | |
| 9. Capsaicin (From: Capsicum spp., Solanaceae family)  | Zhang, M. et al. (2023) [41] | In vitro and in vivo | Encephalomyocarditis virus Vesicular stomatitis virus IAV H1N1 | Capsaicin directly binds STAT3, promoting its lysosomal degradation | NA |
| Marois, I. et al. (2014) [42] | In vitro | IAV | Capsaicin reduced influenza PA gene expression by ~48% | The EC50 was between 44.69–55.17 µM depending on the different strains of influenza virus | |
| Trischitta, P. et al. (2024) [43] | In vitro | Lassa Virus | Inhibits GP-mediated membrane fusion during viral entry | EC50 = 6.9–10.0 µmol/L | |
| 10. Cepharanthine (From: Stephania spp., Menispermaceae family) 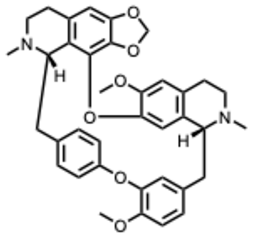 | Xia, B. et al. (2023) [44] | Review | SARS-CoV-2 | Inhibit viral entry and post-entry steps and attenuate the potential inflammatory effects | IC50 = 28.51 ng/mL |
| Liu, K. et al. (2023) [45] | Review | 1. SARS-CoV-2 2. SARS-CoV pseudovirus 3. MERS-CoV pseudovirus 4. HCoV-OC43 5. PEDV 6. SARS-CoV 7. EBOV 8. ZIKV 9. PRRSV 10. HIV-1 | Multiple potential mechanisms of actions depending on the virus | 1. EC50 = 0.15 M 2. EC50 ≈ 0.0417 µM 3. EC50 ≈ 0.14 µM 4. IC50 ≈ 0.83 µM 5. EC50 ≈ 2.53 µM; in vivo: 11.1 mg/kg oral dose reduced viral load 6. EC50 ≈ 0.79 µM 7. IC50 ≈ 0.42 µM 8. IC50 ≈ 2.19 µM 9. 10 µM reduced TCID50 ~5.6-fold; NF-κB inhibition 10. EC50 = 0.026 M | |
| 11. Cephaeline (From: Cephaelis ipecacuanha, Rubiaceae family) 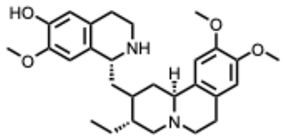 | Ren, P. et al. (2022) [46] | In vitro and in silico | SARS-CoV-2 | Targeting the host ribosome, and viral RNA, RdRp as well as N protein to interfere with the translating, propagating, replicating, and assembling process of the virus. | EC50 = 0.01 µM |
| Yang, S. et al. (2018) [47] | In vitro, in silico and in vivo | ZIKV EBOV | Acts on the ZIKV RdRp NS5, host cell’s lysosome, and 40S ribosomal subunit | IC50 values of less than 42 nM IC50 = 16.9 nM with 95% CI of 10.7–25.8 nM | |
| 12. Cherylline (From: Amaryllidaceae family)  | Ka, S. et al. (2021) [48] | In vitro and in silico | DENV ZIKV | Post-entry inhibition of RNA replication | EC50 = 8.8 µM SI = 28 EC50 = 20.3 µM SI = 12 |
| 13. Cinchonine (From: Cinchona spp., Rubiaceae family)  | Ren, J. et al. (2022) [49] | In vitro | Porcine Epidemic Diarrhea Virus (PEDV) | Induction of autophagy, inhibiting early (adsorption/entry) and replication stages | Dose-dependent suppression of viral mRNA and N protein; ~100 µM nearly complete inhibition; RT-qPCR and TCID50 assays show significant reduction |
| 14. Colchicine (From: Colchicaceae family)  | Hegazy, A. et al. (2024) [50] | In vitro and in silico | IAV H5N1 H1N1 | Inhibition of viral adsorption and replication; docking supports NA/M2 binding | IC50 = 0.111 µg/mL IC50 = 0.326 µg/mL |
| 15. Conessine (From: Apocynaceae family) 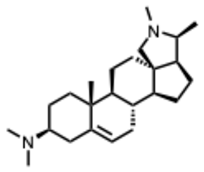 | Majnooni, M. et al. (2021) [33] | Review | SARS-CoV-2 | Mpro inhibitor | EC50 = 2.34 μM, |
| 16. Cyclopamine (From: Melanthiaceae family)  | Bailly, C. et al. (2016) [51] | In vitro and In vivo | Human RSV (hRSV) | Disrupts inclusion bodies via M2-1 protein interaction, impairs RdRp complex | IC50 ≈ 380 nM; CC50 > 320 µM |
| Diot, C. et al. (2023) [52] | In vitro and in silico | hRSV Bovine RSV (BRSV) | Hardens inclusion bodies, interferes with M2-1–P–RNA dynamics | Dose-dependent inhibition | |
| Fix, J. et al. (2023) [53] | In vitro | BRSV | Similar M2-1-mediated mechanism to hRSV | EC50 = 76 nM | |
| 17. Dehydroevodiamine (From: Rutaceae family)  | Li, K. et al. (2024) [54] | In vitro | PEDV | Inhibition of the PEDV replication stage, and its downregulation oft he ERK1/2 MAPH pathway | 48 h PI: IC50 = 3.574 ± 0.566 µg/mL SI = 3.503 |
| 18. Deoxynojirimycin and derivatives (From: e.g., Morus alba, Moraceae family) 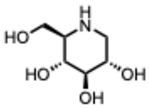 | Hussain, M. et al. (2015) [55] | In vitro | IAV H3N2 strains | α-glucosidase inhibition, impaired viral glycoprotein folding | NN-DNJ IC50 = 0.5–2.5 µM depending on the viral strain |
| Perera, N. et al. (2022) [56] | In vitro | DENV | Blocks viral release; inhibits glycoprotein maturation via α-glucosidase inhibition | IC50 of DNJ-iminosugars 48 h PI: 2THO-DNJ 1.6 ± 0.8 µM EOO-DNJ 3.1 ± 1.3 µM NN-DNJ 3.3 ± 1.5 µM | |
| Bhushan, G. et al. (2020) [57] | In vitro | ZIKV | Impairs viral replication by disrupting glycoprotein folding via ER glucosidase inhibition | At 1 µM, DNJ significantly reduced ZIKV RNA levels in supernatants compared to vehicle control (p ≤ 0.0318) | |
| 19. Emetine (From: Rubiaceae family)  | Yang, S. et al. (2018) [47] | In vitro and in vivo | ZIKV EBOV | Inhibits NS5 polymerase activity and disrupts lysosomal function; also inhibits EBOV entry | IC50 = 52.9 nM (95% CI: 35.4–73.2 nM) SJL mice; 1 mg/kg/day ~10-fold reduction in blood viremia at 7-day PI IC50 = 16.9 nM (95% CI:10.7–25.8 nM) 67% survival (4/6 mice) with IP emetine dosing |
| Khandelwal, N. et al. (2017) [58] | In vitro and in ovo | PPRV NDV | Inhibits viral polymerase and entry; reduces viral RNA/protein synthesis | NDV: Reduced virus production 2-3 log in NDV infected cells | |
| Valipour, M. (2022) [59] | Review | SARS-CoV-2 | Likely affects both virus- and host-based targets (translation machinery, NF-κB, etc.) | EC50 = 0.007 µM EC50 = 0.46 µM IC50 = 0.52 µM EC50 = 0.000147 µM EC50 = 0.00771 µM | |
| Bleasel, M. D. (2020) [60] | Commentary | SARS-CoV MERS-CoV | Broad-spectrum coronavirus inhibition suggested | EC50 SARS: 0.054 µM; EC50 MERS: 0.014 µM | |
| 20. Ephedrine and derivatives (From: Ephedraceae family)  | Wei, Y. et al. (2019) [61] | In vitro and in vivo | IAV (H1N1) | Modulation of TLR3/4/7 signaling, reducing TNF-α and increasing IFN-β | EC50 = 5.66–10.96 µg/mL depending on the drug delivery way |
| 21. Fangchinoline (From: Menispermaceae family) 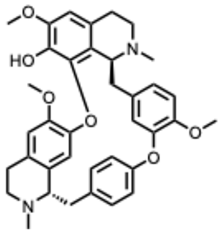 | Yang, S. et al. (2024) [62] | In vitro and in vivo | ZIKA | Inhibits viral internalization | EC50: 0.86 ± 0.47 μM |
| Wan, Z. et al. (2012) [63] | In vitro | HIV-1 | Inhibits gp160 proteolytic processing, blocking envelope maturation | EC50: 0.8–1.7 µM depending on the HIV-1 strains | |
| Zhang, Q. Y. et al. (2024) [64] | In vitro | Enterovirus | Inhibits early-stage infection; VP1 mutations (E145G, V258I) reduce sensitivity | At concentration of 10 µmol/L FAN resulted in about 600-fold reduction in viral titers | |
| 22. Harmaline (From: Nitrariaceae/Malpighiaceae family) 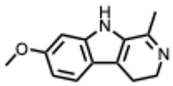 | Hegazy, A. et al. (2023) [65] | In vitro | IAV H1N1 and H5N1 | Not specified: measured via viral inhibition/cytopathic effect assay | H1N1: Harmaline: IC50 = 0.056 µg/mL H5N1: Harmaline: IC50 = 3.42 µg/mL |
| 23. Harmine (From: Nitrariaceae/Malpighiaceae family) 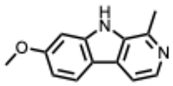 | Hegazy, A. et al. (2023) [65] | In vitro | IAV H1N1 and H5N1 | Not specified: measured via viral inhibition/cytopathic effect assay | H1N1: Harmine IC50 = 0.033 µg/mL H5N1: Harmine IC50 = 0.023 µg/mL |
| Dahal, S. et al. (2023) [66] | In vitro | HCoV-229E SARS-CoV-2 variants HIV-1 | Post-entry inhibition of viral replication via SR kinase inhibition; reduced viral protein expression and subgenomic RNAs. | Dose-dependent inhibition | |
| 24. Harringtonine (From: Cephalotaxaceae family)  | Kaur, P. et al. (2013) [67] | In vitro | CHIKV ZIKV | Inhibits viral protein synthesis by targeting host translation machinery | IC50 = 0.24 µM (plaque assay); EC50 = 0.29 µM (immunofluorescence assay) |
| Lai, Z. et al. (2020) [68] | In vitro | ZIKV | Inhibits multiple stages: binding, entry, replication, release; also virucidal and prophylactic | ~625 nM caused ~3-log reduction in viral RNA and titers | |
| Yang, Y. et al. (2023) [69] | In silico | SARS-CoV-2 | Binds to spike RBD, TMPRSS2, and RBD–ACE2 complex to inhibit viral entry | High binding affinity in silico with RBD and TMPRSS2 | |
| 25. Hernandezine (From: Menispermaceae/Ranunculaceae family)  | Majnooni, M. et al. (2021) [33] | Review | SARS-CoV-2 | Blocking the calcium transition | EC50 = 10 μM, |
| 26. Homoharringtonine (From: Cephalotaxaceae family)  | Dong. H. J. et al. (2018) [70] | In vitro | PEDV | Acts at early replication stages; additive effect with hydroxychloroquine (HCQ) or peptide tHR2 | HHT (150 nM) reduced viral titers by ~3.5-fold; at 300 nM, approx. 40-fold reduction (TCID50 assay) |
| Gong, M. et al. (2019) [71] | In vitro | Foot-and-mouth disease virus (FMDV) | Inhibits early stages of replication | Dose-dependent inhibition | |
| Harisha, K. R. et al. (2025) [72] | In vitro | Rabies virus (RABV) | Post-entry and antiviral across both lab-adapted and clinical isolates | EC50 ≈ 0.3 µM (BHK-21); EC50 ≈ 0.4 µM (Neuro-2a); cell–cell spread inhibited at ≈ 1.0 µM | |
| Neerukonda, S. N. et al. (2020) [73] | Review | SARS-CoV-2 | Likely suppresses viral replication by targeting phosphorylated eIF4E and inhibiting host protein translation | EC50 ≈ 2.10 µM in Vero E6 cells | |
| 27. Indirubin (From: Brassicaceae family)  | Mok, C. K. P. et al. (2014) [74] | In vitro | H5N1 | Anti-inflammatory and antiviral via indirubin derivatives. Delays virus replication; reduces pro-inflammatory cytokines (IP-10) | Indirubin derivatives strongly suppress the pro-inflammatory cytokines including IP-10 (CXCL10), one of the key factors which contribute to the lung inflammation during H5N1 virus infection. |
| Chang, S.-J. et al. (2012) [75] | In vitro and in vivo | JEV | Blocks viral attachment and has virucidal activity | EC50 = 0.006–0.105 mg/mL (concentration-dependent inhibition) | |
| Jie, C. et al. (2017) [76] | In vivo | IAV (H1N1) | Promotes MAVS-mediated IFN-β production and protects mitochondrial antiviral signaling. | Reduced lung NP levels, improved survival and lung pathology; enhanced IFN-β and IFITM3 signaling | |
| Medina-Moreno, S. et al. (2017) [77] | In vivo | HIV-1 | Inhibits CDK9 to suppress HIV proviral transcription | Significant reduction in viremia at 5 mg/kg/day dosing | |
| 28. Lycorine and derivatives (From: Amaryllidaceae family)  | Jin, Y.-H. et al. (2021) [78] | In vitro | 1. MERS-CoV | RdRp inhibition | IC50 = 1.406 ± 0.260 µM |
| In vitro | 2. SARS-CoV | RdRp inhibition | IC50 = 1.021 ± 0.025 µM | ||
| In vitro | 3. SARS-CoV-2 | RdRp inhibition | IC50 = 0.878 ± 0.022 µM | ||
| Chen, H. et al. (2020) [79] | In vitro and in vivo | ZIKV | Post-entry inhibition of RdRp activity | EC50 = 0.22–0.39 µM in different cell lines; CC50 = 4.4–21 µM; RdRp IC50 = 25 µM; 10 mg/kg in mice protected against lethality (~83%) | |
| Li, N. et al. (2021) [80] | In vitro | CHIKV | Inhibits viral translation post-entry | EC50 ≈ 10 µM | |
| Narayanan, A. et al. (2022) [81] | In vitro | SARS-CoV-2 | Mpro inhibition; 88% reduction in viral spread in cell culture | EC50 = 0.01 µM (Lycorine HCl against Mpro; SI = 1878) | |
| Fielding, B. C. et al. (2020) [82] | In vitro | SARS-CoV-2 | Likely host-targeted antiviral modulation | EC50 = 300 nM; SI ≈ 130 | |
| 29. Matrine and derivatives (From: Fabaceae family)  | Pan, Q.-M. et al. (2015) [83] | In vitro | IAV H3N2 | Inhibits viral replication | For Matrine type alkaloids: IC50 = 63.07–242.46 µM |
| Qiao, W.-T. et al. (2024) [84] | In vivo | PEDV | Inhibits PEDV attachment and entry to cells. | EC50 = 0.09 µM (SI = 358.9) | |
| 30. Michellamine B (From: Ancistrocladaceae family)  | McMahon, J. B. et al. (1995) [85] | In vitro | HIV-1 | Inhibits reverse transcriptase activity and viral-induced cellular fusion | EC50 = 1–20 µM Complete inhibition of cytopathic effects; RT enzymatic inhibition; IC50 ≈ 10 µM for fusion |
| 31. Neferine (From: Nelumbonaceae family) 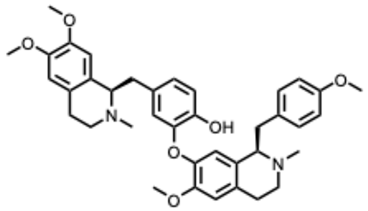 | Yang, Y. et al. (2021) [86] | In vitro | SARS-CoV-2 Pseudovirus | Blocks host Ca2+-dependent membrane fusion → viral entry | EC50 = 0.13–0.41 µM |
| Yang, D. et al. (2025) [87] | In vitro | SARS-CoV-2 | Enhanced virucidal activity by salt form | Neferine free base: EC50 = 36.01 µM; Neferine salt: EC50 = 4.78 µM | |
| 32. Panicutine (From: Ranunculaceae family)  | Kumar, A et al. (2022) [87] | In silico | SARS-CoV-2 | Potential Mpro inhibition | Binding energy: −7.4 kcal/mol |
| 33. Piperine (From: Piperaceae family)  | Nag, A. & Chowdhury, S. (2020) [88] | In silico | DENV EBOV | Potential inhibition of enzymatic targets | Stronger binding than ribavirin |
| Pareek, A. et al. (2022) [89] | In vitro | CHIKV | RdRp inhibition, reduces viral replication | Kd = 0.08 µM (RdRp); EC50 = 6.68 µM | |
| 34. Pseudoephedrine (From: Ephedraceae family) 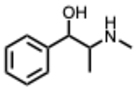 | Deng, L et al. (2020) [90] | Review | IAV H1N1 | Blunts cytokine storm, reduces lung inflammation, and inhibits virus replication | Increased life span in infected mice treated by PE |
| Yu, S. et al. (2021) [91] | In vitro | SARS-CoV-2 | Blocks viral entry and reduces inflammatory response | Inhibits spike pseudovirus entry; EC50 < 20 µM | |
| 35. Quinidine (From: Rubiaceae family)  | Yang et al. (2021) [92] | In vitro | SARS-CoV-2 | Moderate antiviral activity among approved drugs in a high-throughput screen | IC50 = 0.42 µM (compound 6g); IC50 = 1.41 µM (compound 7k) |
| Persoons, L. et al. (2021) [93] | In vitro | SARS-CoV-2 pseudovirus/HCoV-229E/OC43 | Showed some broad-spectrum activity but only at relatively high doses | EC50 = 6 µM (SARS-CoV-2); EC50 = 0.2–9.4 µM (other HCoVs) | |
| Mamidala, E. et al. (2022) [94] | In silico | SARS-CoV-2 main protease (Mpro) | Binds SARS-CoV-2 protease with predicted inhibitory affinity | Strong docking affinity | |
| 36. Quinine (From: Rubiaceae family) 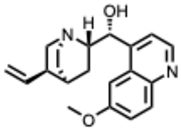 | Malakar, S. et al. (2018) [95] | In vitro | DENV | Significant inhibition of DENV replication across multiple serotypes | ~80% reduction vs. control |
| D’Alessandro, S. et al. (2020) [96] | Review | IAV | Prophylactic plaque reduction at non-cytotoxic doses | NR | |
| Große, M. et al. (2021) [97] | In vitro | SARS-CoV-2 | Complete viral inhibition at ≥50 µM; wider effective range depending on MOI and cell line | IC50 ≈ 25 µM (range: ~3.7–50 µM) | |
| 37. Rutaecarpine (From: Rutaceae family)  | Lin, S. et al. (2023) [98] | In vitro and in silico | SARS-CoV-2 pseudovirus | Disrupts binding between spike protein and ACE2 receptor | IC50 ≈ 30 µM Omicron variant: IC50 ≈ 15 µM Binding energy ≈ −8.6 kJ/mol (wild-type), −10.2 kJ/mol (omicron) |
| 38. Scopolamine (From: Solanaceae family)  | Bhattacharjee, A. et al. (2021) [99] | In ovo and in silico | JEV | Binds to the JEV NS5 protein and modulates TLR and IFN signaling pathways, potentiating antiviral innate immunity | Significant reduction in the viral load in CAM (p < 0.0001) and brain tissues (p < 0.0001) of the embryonated chick eggs when pre-treated with scopolamine hydrobromide |
| 39. Sophocarpine (From: Fabaceae family)  | Jin, Z. et al. (2017) [100] | In vitro | Enterovirus 71 (E71) | Inhibits viral attachment, penetration, and RNA replication | IC50 = 350 µg/mL CC50 = 1346 µg/mL |
| 40. Sophoridine (From: Fabaceae family)  | Ren, G. et al. (2019) [101] | In vitro | E71 | Inhibits viral adsorption when added before infection | IC50 = 61.39 µg/mL |
| 41. Strychnine (From: Loganiaceae family) 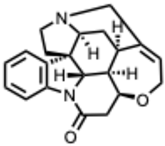 | Hegazy, A. et al. (2023) [65] | In vitro | IAV H5N1 H1N1 | Inhibits viral adsorption to host cells | IC50 = 11.85 µg/mL SI > 843 IC50 = 0.06 µg/mL; SI ≈ 167,000 |
| 42. Tetrahydropalmatine (From: Papaveraceae family) 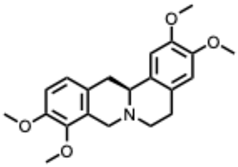 | Lixia, H. et al. (2018) [102] | In vivo | JEV | Protects against neuronal apoptosis, reduces reactive oxygen/nitrogen species, lowers proinflammatory mediators, exhibits antiviral effects in brain tissues | Reduction in viral load in brain and CAM via neuroprotective and anti-inflammatory effects |
| 43. Tetrandrine (From: Menispermaceae family)  | Kim, D. E. et al. (2019) [103] | In vitro | Human Coronavirus 0C43 | Prevents viral replication and viral protein expression; effective mainly when administered pre- or during infection; also activates p38 MAPK signaling | IC50 ≈ 0.33 µM; SI > 40 |
| Liu, J. et al. (2023) [104] | In vitro and in vivo | SARS-CoV-2 | Blocks viral entry at early stage by interfering with endosomal trafficking; exhibits favorable lung biodistribution when inhaled | EC50 = 0.40–5.03 µM depending on a cell line | |
| 44. Tylophorine and analogues (From: Apocynaceae family)  | Fielding et al. (2020) [81] | In vitro | SARS-CoV | Blocks viral replication and cytopathic effects; potent inhibition of virus-induced apoptosis | EC50 = 5–340 nM (synthetic derivatives); 8–1468 nM (natural) |
| T. I. M. et al. (2021) [105] | In vitro | SARS-CoV SARS-CoV-2 | Inhibits viral RNA replication and NF-κB activation via JAK2 signaling pathway | IC50 = 58 nM (tylophorine), IC50 = 20 nM (7-methoxycryptopleurine) | |
| Yang et al. (2010) [106] | In vitro | Transmissible gastroenteritis virus (TGEV) | Targets viral RNA/RNP complex and inhibits NF-κB-mediated pro-inflammatory signaling | Substantial suppression of viral RNA replication (~4-log reduction) | |
| Wang, Y. et al. (2017) [107] | In vitro | Hepatitis C | Binds Hsc70 NBD, enhances ATPase activity → disrupts viral replication | Synthetic analogues of tylophorine DCB-3503 and rac-cryptopleurine EC50 = 30nM EC50 =300nM | |
| 45. Tryptanthrine (From: Brassicaceae family)  | Mani, J. S. et al. (2020) [108] | Review | SARS-CoV-2 & Other HCoVs | Blocks coronaviral replication in early and late stages via inhibition of RdRp and papain-like protease | EC50 = 1.52 µM IC50 = 0.06 μM |
| 46. Vilmorrianone (From: Ranunculaceae family)  | Kumar, A et al. (2022) [87] | In silico | SARS-CoV-2 | Potential Mpro inhibition | Binding energy: −7.0 kcal/mol |
| 47. Vinblastine (From: Apocynaceae family)  | Akan, E. et al. (1997) [109] | In vitro | HIV-1 | Vinblastine induced transcription through the HIV-1 long terminal repeat, suggesting modulation of NF-κB pathways, not antiviral action | ~9–10-fold induction |
| Alkaloid | In Silico | In Vitro | In Vivo | Virus families | Main Targets |
|---|---|---|---|---|---|
| 1,2,3,4-Tetrahydroisoquinoline derivatives | ✓ | ✓ | ✓ | Orthomyxo, Retro, CoV | RdRp (PAN), IN |
| 7-Methoxycryptopleurine | – | ✓ | – | CoV | Blocking the S and N proteins, 3CLpro inhibitor |
| 10-Hydroxyusambarensine | ✓ | – | – | CoV | RdRp |
| Ajmaline | ✓ | – | – | Filo | Host gene regulation |
| Aloperine | – | ✓ | ✓ | Flavi, CoV | RdRp/Cathepsin B |
| Amarbellisine | – | ✓ | – | CoV | Potential inhibition of viral replication |
| Berbamine | – | ✓ | – | CoV | Endolysosomal trafficking |
| Berberine | ✓ | ✓ | ✓ | Orthomyxo, Flavi, Alpha, Retro | Host pathways/Entry |
| Canthin-6-one derivatives | ✓ | ✓ | – | Paramyxo, CoV | Proteases/Entry |
| Capsaicin | – | ✓ | ✓ | Orthomyxo, Picorna | STAT3/Host immunity |
| Cepharanthine | ✓ | ✓ | ✓ | CoV, Flavi, Alpha, Retro | Entry/Host pathways |
| Cephaeline | ✓ | ✓ | ✓ | CoV, Flavi | Ribosome/RdRp |
| Cherylline | ✓ | ✓ | – | Flavi | RdRp |
| Cinchonine | – | ✓ | – | CoV | Autophagy |
| Colchicine | ✓ | ✓ | – | Orthomyxo | Adsorption/Replication |
| Conessine | – | ✓ | – | CoV | Mpro inhibitor |
| Cyclopamine | – | ✓ | ✓ | Pneumoviridae | Inclusion bodies (M2-1) |
| Dehydroevodiamine | – | ✓ | – | CoV | ERK/MAPK |
| Deoxynojirimycin derivatives | – | ✓ | – | Orthomyxo, Flavi | α-glucosidase |
| Emetine | ✓ | ✓ | ✓ | CoV, Flavi, Filo, Retro | Translation/Entry |
| Ephedrine | – | ✓ | ✓ | Orthomyxo | TLR signaling |
| Fangchinoline | – | ✓ | ✓ | Flavi, Retro, Picorna | Viral internalization |
| Harmaline | – | ✓ | – | Orthomyxo | – |
| Harmine | – | ✓ | – | Orthomyxo, CoV, Retro | SR kinase |
| Harringtonine | ✓ | ✓ | – | Alpha, Flavi, CoV | Translation |
| Hernandezine | – | ✓ | – | CoV | Blocking calcium transition |
| Homoharringtonine | – | ✓ | ✓ | CoV, Flavi, Rhabdo | Translation |
| Indirubin | – | ✓ | ✓ | Orthomyxo, Flavi, Retro | MAVS/Cytokines |
| Lycorine | ✓ | ✓ | ✓ | CoV, Flavi, Alpha | RdRp |
| Matrine | – | ✓ | ✓ | Orthomyxo, Arteri | Viral entry |
| Michellamine B | – | ✓ | – | Retro | Reverse transcriptase |
| Neferine | – | ✓ | – | CoV | Ca2+-dependent fusion |
| Oxysophoridine | – | ✓ | – | CoV | Nucleotide biosynthesis inhibitor |
| Panicutine | ✓ | Protease (Mpro) | |||
| Piperine | ✓ | ✓ | – | Flavi, Alpha | RdRp |
| Pseudoephedrine | – | ✓ | ✓ | Orthomyxo, CoV | Entry/Inflammation |
| Quinidine | ✓ | ✓ | – | CoV | Protease (Mpro) |
| Quinine | – | ✓ | – | Flavi, CoV | – |
| Rutaecarpine | ✓ | ✓ | – | CoV | Spike–ACE2 |
| Scopolamine | ✓ | ✓ | – | Flavi | NS5/Innate immunity |
| Sophocarpine | – | ✓ | – | Picorna | Viral attachment |
| Sophoridine | – | ✓ | – | Picorna | Viral adsorption |
| Strychnine | – | ✓ | – | Orthomyxo | Viral adsorption |
| Tetrahydropalmatine | – | – | ✓ | Flavi | Neuroprotection |
| Tetrandrine | ✓ | ✓ | ✓ | CoV, Orthomyxo | Entry/Endosomes |
| Tylophorine & analogues | – | ✓ | – | CoV, Alpha | RdRp/NF-κB |
| Tryptanthrine | – | ✓ | – | CoV | RdRp/PLpro |
| Vilmorrianone | ✓ | – | – | CoV | Protease (Mpro) |
| Vinblastine | – | ✓ | – | Retro | Host transcription (No antiviral activity reported; host transcription modulation) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Leka, K.; Mamede, L.; Vandeberg, E.; Garigliany, M.-M.; Ledoux, A. Natural Alkaloids as Antiviral Agents Against RNA Viruses: A Comprehensive and Mechanistic Review. Molecules 2026, 31, 539. https://doi.org/10.3390/molecules31030539
Leka K, Mamede L, Vandeberg E, Garigliany M-M, Ledoux A. Natural Alkaloids as Antiviral Agents Against RNA Viruses: A Comprehensive and Mechanistic Review. Molecules. 2026; 31(3):539. https://doi.org/10.3390/molecules31030539
Chicago/Turabian StyleLeka, Kristi, Lúcia Mamede, Elyn Vandeberg, Mutien-Marie Garigliany, and Allison Ledoux. 2026. "Natural Alkaloids as Antiviral Agents Against RNA Viruses: A Comprehensive and Mechanistic Review" Molecules 31, no. 3: 539. https://doi.org/10.3390/molecules31030539
APA StyleLeka, K., Mamede, L., Vandeberg, E., Garigliany, M.-M., & Ledoux, A. (2026). Natural Alkaloids as Antiviral Agents Against RNA Viruses: A Comprehensive and Mechanistic Review. Molecules, 31(3), 539. https://doi.org/10.3390/molecules31030539






