Bioactive Metabolites from Terrestrial and Marine Actinomycetes
Abstract
1. Introduction
2. Ecology of Actinomycetes
2.1. Soil Actinomycetes
2.2. Endophytic Actinomycetes
2.3. Actinomycetes in Compost
2.4. Marine Actinomycetes
3. Taxonomy and Classification
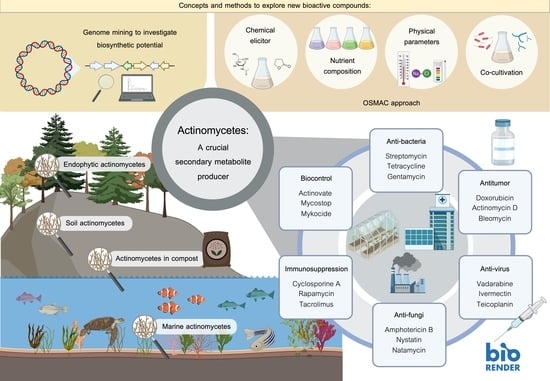
4. A Crucial Secondary Metabolite Producer
5. Biological Activity of Secondary Metabolites from Actinomycetes and Their Applications
5.1. Antibacterial Agents
5.2. Antifungal Agents
5.3. Immunosuppressive Agent
5.4. Biocontrol Agents
| Commercial Product Name | Organism As Active Substance | Registered As a Microbial Pesticide | Targeted Pest/Pathogen/Disease |
|---|---|---|---|
| Actinovate, Novozymes BioAg Inc., Milwaukee, WI, USA | S. lydicus WYEC 108 | Canada, USA | Soilborne diseases, viz. Pythium, Fusarium, Phytophthora, Rhizoctonia, and Verticillium; foliar diseases such as powdery and downy mildew, Botrytis, Alternaria, Postia, Geotrichum, and Sclerotinia |
| Mycostop, Verdera Oy, Espoo, Finland | Streptomyces K61 | EU, Canada, USA | Damping off caused by Alternaria, R. solani, Fusarium, Phytophthora, Pythium wilt, and root diseases |
| Mykocide, KIBC Co., Ltd., Yongin, Gyeonggi-do, Republic of Korea | S. colombiensis | Republic of Korea | Powdery mildews, grey mold, and brown patch |
| Bactophil | Streptomyces albus | Ukraine | Seed germination diseases |
5.5. Antitumor Compounds
5.6. Antiviral Agents
5.7. Other Activities
6. Concepts and Methods to Explore New Bioactive Compounds
6.1. Exploring New Habitats or Extreme Environments As A Source for Novel Strains
6.2. Genome Mining to Investigate Biosynthetic Potential
6.3. OSMAC Approach
6.4. Co-Cultivation Technique
6.5. Using Chemical Elicitors
6.5.1. γ-. Butyrolactones and Related Regulators
6.5.2. N-Acetylglucosamine (GlcNAc)
7. Future Perspectives on Actinomycetes
8. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Barka, E.A.; Vatsa, P.; Sanchez, L.; Gaveau-Vaillant, N.; Jacquard, C.; Klenk, H.P.; Clement, C.; Ouhdouch, Y.; van Wezel, G.P. Taxonomy, Physiology, and Natural Products of Actinobacteria. Microbiol. Mol. Biol. Rev. 2016, 80, 1–43. [Google Scholar] [CrossRef] [PubMed]
- Kieser, T.; Bibb, M.J.; Buttner, M.J.; Chater, K.F.; Hopwood, D.A. Practical Streptomyces Genetics; The John Innes Foundation: Norwich, UK, 2000. [Google Scholar]
- Tiwari, K.; Gupta, R.K. Rare actinomycetes: A potential storehouse for novel antibiotics. Crit. Rev. Biotechnol. 2012, 32, 108–132. [Google Scholar] [CrossRef] [PubMed]
- Ding, T.; Yang, L.-J.; Zhang, W.-D.; Shen, Y.-H. The secondary metabolites of rare actinomycetes: Chemistry and bioactivity. RSC Adv. 2019, 9, 21964–21988. [Google Scholar] [CrossRef] [PubMed]
- Al-Fadhli, A.A.; Threadgill, M.D.; Mohammed, F.; Sibley, P.; Al-Ariqi, W.; Parveen, I. Macrolides from rare actinomycetes: Structures and bioactivities. Int. J. Antimicrob. Agents 2022, 59, 106523. [Google Scholar] [CrossRef] [PubMed]
- Prudence, S.M.M.; Addington, E.; Castaño-Espriu, L.; Mark, D.R.; Pintor-Escobar, L.; Russell, A.H.; McLean, T.C. Advances in actinomycete research: An ActinoBase review of 2019. Microbiology 2020, 166, 683–694. [Google Scholar] [CrossRef] [PubMed]
- Qinyuan, L.; Xiu, C.; Yi, J.; Chenglin, J. Morphological Identification of Actinobacteria. In Actinobacteria; Dharumadurai, D., Yi, J., Eds.; IntechOpen: Rijeka, Croatia, 2016; pp. 59–86. [Google Scholar]
- Goodfellow, M.; Williams, S.T. Ecology of actinomycetes. Annu. Rev. Microbiol. 1983, 37, 189–216. [Google Scholar] [CrossRef]
- van der Meij, A.; Worsley, S.F.; Hutchings, M.I.; van Wezel, G.P. Chemical ecology of antibiotic production by actinomycetes. FEMS Microbiol. Rev. 2017, 41, 392–416. [Google Scholar] [CrossRef]
- Matsumoto, A.; Takahashi, Y. Endophytic actinomycetes: Promising source of novel bioactive compounds. J. Ant. 2017, 70, 514–519. [Google Scholar] [CrossRef] [PubMed]
- Helmke, E.; Weyland, H. Rhodococcus marinonascens sp. nov., an actinomycete from the sea. Int. J. Syst. Bacteriol. 1984, 34, 127–138. [Google Scholar] [CrossRef]
- Masand, M.; Jose, P.A.; Menghani, E.; Jebakumar, S.R.D. Continuing hunt for endophytic actinomycetes as a source of novel biologically active metabolites. World J. Microbiol. 2015, 31, 1863–1875. [Google Scholar] [CrossRef]
- Kumar, S.; Solanki, D.S.; Parihar, K.; Tak, A.; Gehlot, P.; Pathak, R.; Singh, S.K. Actinomycetes isolates of arid zone of Indian Thar Desert and efficacy of their bioactive compounds against human pathogenic bacteria. Biol. Futur. 2021, 72, 431–440. [Google Scholar] [CrossRef]
- Mohammadipanah, F.; Wink, J. Actinobacteria from arid and desert habitats: Diversity and biological activity. Front. Microbiol. 2016, 6, 1541. [Google Scholar] [CrossRef]
- Silva, L.J.; Crevelin, E.J.; Souza, D.T.; Lacerda-Júnior, G.V.; de Oliveira, V.M.; Ruiz, A.L.T.G.; Rosa, L.H.; Moraes, L.A.B.; Melo, I.S. Actinobacteria from Antarctica as a source for anticancer discovery. Sci. Rep. 2020, 10, 13870. [Google Scholar] [CrossRef]
- Zenova, G.M.; Manucharova, N.A.; Zvyagintsev, D.G. Extremophilic and extremotolerant actinomycetes in different soil types. Eurasian Soil Sci. 2011, 44, 417–436. [Google Scholar] [CrossRef]
- Bhatti, A.A.; Haq, S.; Bhat, R.A. Actinomycetes benefaction role in soil and plant health. Microb. Pathog. 2017, 111, 458–467. [Google Scholar] [CrossRef]
- Aamir, M.; Rai, K.K.; Zehra, A.; Dubey, M.K.; Samal, S.; Yadav, M.; Upadhyay, R.S. Endophytic actinomycetes in bioactive compounds production and plant defense system. In Microbial Endophytes; Kumar, A., Singh, V.K., Eds.; Woodhead Publishing: Sawston, UK, 2020; pp. 189–229. [Google Scholar] [CrossRef]
- Chaurasia, A.; Meena, B.R.; Tripathi, A.N.; Pandey, K.K.; Rai, A.B.; Singh, B. Actinomycetes: An unexplored microorganisms for plant growth promotion and biocontrol in vegetable crops. World J. Microbiol. 2018, 34, 132. [Google Scholar] [CrossRef] [PubMed]
- Nonthakaew, N.; Panbangred, W.; Songnuan, W.; Intra, B. Plant growth-promoting properties of Streptomyces spp. isolates and their impact on mung bean plantlets’ rhizosphere microbiome. Front. Microbiol 2022, 13, 967415. [Google Scholar] [CrossRef] [PubMed]
- Intra, B.; Mungsuntisuk, I.; Nihira, T.; Igarashi, Y.; Panbangred, W. Identification of actinomycetes from plant rhizospheric soils with inhibitory activity against Colletotrichum spp., the causative agent of anthracnose disease. BMC Res. Notes 2011, 4, 98. [Google Scholar] [CrossRef] [PubMed]
- Bao, Y.; Dolfing, J.; Guo, Z.; Chen, R.; Wu, M.; Li, Z.; Lin, X.; Feng, Y. Important ecophysiological roles of non-dominant Actinobacteria in plant residue decomposition, especially in less fertile soils. Microbiome 2021, 9, 84. [Google Scholar] [CrossRef]
- Guan, T.W.; Lin, Y.J.; Ou, M.Y.; Chen, K.B. Isolation and diversity of sediment bacteria in the hypersaline aiding lake, China. PLoS ONE 2020, 15, e0236006. [Google Scholar] [CrossRef]
- Ahmed, V.; Verma, M.K.; Gupta, S.; Mandhan, V.; Chauhan, N.S. Metagenomic Profiling of Soil Microbes to Mine Salt Stress Tolerance Genes. Front. Microb. 2018, 9, 159. [Google Scholar] [CrossRef]
- Rungin, S.; Indananda, C.; Suttiviriya, P.; Kruasuwan, W.; Jaemsaeng, R.; Thamchaipenet, A. Plant growth enhancing effects by a siderophore-producing endophytic streptomycete isolated from a Thai jasmine rice plant (Oryza sativa L. cv. KDML105). Antonie Van Leeuwenhoek 2012, 102, 463–472. [Google Scholar] [CrossRef]
- Janso, J.E.; Carter, G.T. Biosynthetic Potential of Phylogenetically Unique Endophytic Actinomycetes from Tropical Plants. Appl. Environ. Microbiol. 2010, 76, 4377–4386. [Google Scholar] [CrossRef] [PubMed]
- Hayakawa, M. Studies on the isolation and distribution of rare actinomycetes in soil. Actinomycetologica 2008, 22, 12–19. [Google Scholar] [CrossRef]
- Hemati, A.; Nazari, M.; Asgari Lajayer, B.; Smith, D.L.; Astatkie, T. Lignocellulosics in plant cell wall and their potential biological degradation. Folia Microbiol. 2022, 67, 671–681. [Google Scholar] [CrossRef]
- Hemati, A.; Aliasgharzad, N.; Khakvar, R.; Delangiz, N.; Asgari Lajayer, B.; van Hullebusch, E.D. Bioaugmentation of thermophilic lignocellulose degrading bacteria accelerate the composting process of lignocellulosic materials. Biomass Conv. Bioref. 2022, 12, 1–15. [Google Scholar] [CrossRef]
- Hemati, A.; Aliasgharzad, N.; Khakvar, R.; Khoshmanzar, E.; Asgari Lajayer, B.; van Hullebusch, E.D. Role of lignin and thermophilic lignocellulolytic bacteria in the evolution of humification indices and enzymatic activities during compost production. Waste Manag. 2021, 119, 122–134. [Google Scholar] [CrossRef] [PubMed]
- de Gannes, V.; Eudoxie, G.; Hickey, W.J. Prokaryotic successions and diversity in composts as revealed by 454-pyrosequencing. Bioresour. Technol. 2013, 133, 573–580. [Google Scholar] [CrossRef] [PubMed]
- Takaku, H.; Kodaira, S.; Kimoto, A.; Nashimoto, M.; Takagi, M. Microbial communities in the garbage composting with rice hull as an amendment revealed by culture-dependent and -independent approaches. J. Biosci. Bioeng. 2006, 101, 42–50. [Google Scholar] [CrossRef]
- Setyati, W.A.; Pringgenies, D.; Soenardjo, N.; Pramesti, R. Actinomycetes of secondary metabolite producers from mangrove sediments, Central Java, Indonesia. Vet. World 2021, 14, 2620. [Google Scholar] [CrossRef]
- Khalifa, S.A.M.; Elias, N.; Farag, M.A.; Chen, L.; Saeed, A.; Hegazy, M.E.F.; Moustafa, M.S.; El-Wahed, A.A.; Al-Mousawi, S.M.; Musharraf, S.G.; et al. Marine Natural Products: A Source of Novel Anticancer Drugs. Mar. Drugs 2019, 17, 491. [Google Scholar] [CrossRef]
- Jensen, P.R.; Dwight, R.; Fenical, W. Distribution of actinomycetes in near-shore tropical marine sediments. Appl. Environ. Microbiol. 1991, 57, 1102–1108. [Google Scholar] [CrossRef] [PubMed]
- Jensen, P.R.; Mafnas, C. Biogeography of the marine actinomycete Salinispora. Environ. Microbiol. 2006, 8, 1881–1888. [Google Scholar] [CrossRef]
- Bienhold, C.; Zinger, L.; Boetius, A.; Ramette, A. Diversity and Biogeography of Bathyal and Abyssal Seafloor Bacteria. PLoS ONE 2016, 11, e0148016. [Google Scholar] [CrossRef] [PubMed]
- Betancur, L.A.; Naranjo-Gaybor, S.J.; Vinchira-Villarraga, D.M.; Moreno-Sarmiento, N.C.; Maldonado, L.A.; Suarez-Moreno, Z.R.; Acosta-González, A.; Padilla-Gonzalez, G.F.; Puyana, M.; Castellanos, L.; et al. Marine Actinobacteria as a source of compounds for phytopathogen control: An integrative metabolic-profiling/bioactivity and taxonomical approach. PLoS ONE 2017, 12, e0170148. [Google Scholar] [CrossRef]
- Mahmoud, H.M.; Kalendar, A.A. Coral-associated Actinobacteria: Diversity, abundance, and biotechnological potentials. Front. Microb. 2016, 7, 204. [Google Scholar] [CrossRef]
- Sarmiento-Vizcaíno, A.; González, V.; Braña, A.F.; Palacios, J.J.; Otero, L.; Fernández, J.; Molina, A.; Kulik, A.; Vázquez, F.; Acuña, J.L.; et al. Pharmacological Potential of Phylogenetically Diverse Actinobacteria Isolated from Deep-Sea Coral Ecosystems of the Submarine Avilés Canyon in the Cantabrian Sea. Microb. Ecol. 2017, 73, 338–352. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Ye, Y.; Wang, R.; Zhang, Y.; Wu, C.; Debnath, S.C.; Ma, Z.; Wang, J.; Wu, M. Streptomyces nigra sp. nov. Is a Novel Actinobacterium Isolated From Mangrove Soil and Exerts a Potent Antitumor Activity in Vitro. Front. Microbiol. 2018, 9, 1587. [Google Scholar] [CrossRef]
- Kemung, H.M.; Tan, L.T.H.; Chan, K.G.; Ser, H.L.; Law, J.W.F.; Lee, L.H.; Goh, B.H. Streptomyces sp. Strain MUSC 125 from Mangrove Soil in Malaysia with Anti-MRSA, Anti-Biofilm and Antioxidant Activities. Molecules 2020, 25, 3545. [Google Scholar] [CrossRef]
- Lin, X.; Hetharua, B.; Lin, L.; Xu, H.; Zheng, T.; He, Z.; Tian, Y. Mangrove Sediment Microbiome: Adaptive Microbial Assemblages and Their Routed Biogeochemical Processes in Yunxiao Mangrove National Nature Reserve, China. Microb. Ecol. 2019, 78, 57–69. [Google Scholar] [CrossRef]
- Xu, D.; Han, L.; Li, C.; Cao, Q.; Zhu, D.; Barrett, N.H.; Harmody, D.; Chen, J.; Zhu, H.; McCarthy, P.J.; et al. Bioprospecting deep-sea actinobacteria for novel anti-infective natural products. Front. Microbiol. 2018, 9, 787. [Google Scholar] [CrossRef] [PubMed]
- Olano, C.; Méndez, C.; Salas, J.A. Antitumor compounds from marine actinomycetes. Mar. Drugs 2009, 7, 210–248. [Google Scholar] [CrossRef]
- Subramani, R.; Aalbersberg, W. Marine actinomycetes: An ongoing source of novel bioactive metabolites. Microbiol. Res. 2012, 167, 571–580. [Google Scholar] [CrossRef] [PubMed]
- Anderson, A.S.; Wellington, E.M.H. The taxonomy of Streptomyces and related genera. Int. J. Syst. Evol. Microbiol. 2001, 51, 797–814. [Google Scholar] [CrossRef]
- Ngamcharungchit, C.; Kanto, H.; Také, A.; Intra, B.; Matsumoto, A.; Panbangred, W.; Inahashi, Y. Amycolatopsis iheyensis sp. nov., isolated from soil on Iheya Island, Japan. Int. J. Syst. Evol. Microbiol. 2023, 73, 005757. [Google Scholar] [CrossRef] [PubMed]
- Citarella, R.V.; Colwell, R.R. Polyphasic Taxonomy of the Genus Vibrio: Polynucleotide Sequence Relationships Among Selected Vibrio Species. J. Bacteriol. 1970, 104, 434. [Google Scholar] [CrossRef]
- Vandamme, P.; Pot, B.; Gillis, M.; De Vos, P.; Kersters, K.; Swings, J. Polyphasic taxonomy, a consensus approach to bacterial systematics. Microbiol. Rev. 1996, 60, 407–438. [Google Scholar] [CrossRef]
- Wayne, L.G.; Brenner, D.J.; Colwell, R.R.; Grimont, P.A.D.; Kandler, O.; Krichevsky, M.I.; Moore, L.H.; Moore, W.E.C.; Murray, R.G.E.; Stackebrandt, E.; et al. Report of the Ad Hoc Committee on Reconciliation of Approaches to Bacterial Systematics. Int. J. Syst. Evol. Microbiol. 1987, 37, 463–464. [Google Scholar] [CrossRef]
- Meier-Kolthoff, J.P.; Auch, A.F.; Klenk, H.P.; Göker, M. Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinform. 2013, 14, 60. [Google Scholar] [CrossRef]
- Tindall, B.J.; Rosselló-Móra, R.; Busse, H.J.; Ludwig, W.; Kämpfer, P. Notes on the characterization of prokaryote strains for taxonomic purposes. Int. J. Syst. Evol. Microbiol. 2010, 60, 249–266. [Google Scholar] [CrossRef]
- Intra, B.; Matsumoto, A.; Inahashi, Y.; Ōmura, S.; Takahashi, Y.; Panbangred, W. Actinokineospora bangkokensis sp. nov., isolated from rhizospheric soil. Int. J. Syst. Evol. Microbiol. 2013, 63, 2655–2660. [Google Scholar] [CrossRef] [PubMed]
- Intra, B.; Euanorasetr, J.; Také, A.; Inahashi, Y.; Mori, M.; Panbangred, W.; Matsumoto, A. Saccharopolyspora rhizosphaerae sp. nov., an actinomycete isolated from rhizosphere soil in Thailand. Int. J. Syst. Evol. Microbiol. 2019, 69, 1299–1305. [Google Scholar] [CrossRef] [PubMed]
- Intra, B.; Matsumoto, A.; Inahashi, Y.; Ōmura, S.; Panbangred, W.; Takahashi, Y. Streptosporangium jomthongense sp. nov., an actinomycete isolated from rhizospheric soil and emendation of the genus Streptosporangium. Int. J. Syst. Evol. Microbiol. 2014, 64, 2400–2406. [Google Scholar] [CrossRef]
- Wattanasuepsin, W.; Intra, B.; Také, A.; Inahashi, Y.; Euanorasetr, J.; Ōmura, S.; Matsumoto, A.; Panbangred, W. Saccharomonospora colocasiae sp. Nov., an actinomycete isolated from the rhizosphere of Colocasia esculenta. Int. J. Syst. Evol. Microbiol. 2017, 67, 4572–4577. [Google Scholar] [CrossRef]
- Intra, B.; Panbangred, W.; Inahashi, Y.; Také, A.; Mori, M.; Ōmura, S.; Matsumoto, A. Micromonospora pelagivivens sp. nov., a new species of the genus Micromonospora isolated from deep-sea sediment in Japan. Int. J. Syst. Evol. Microbiol. 2020, 70, 3069–3075. [Google Scholar] [CrossRef]
- Bérdy, J. Thoughts and facts about antibiotics: Where we are now and where we are heading? J. Antibiot. 2012, 65, 385–395. [Google Scholar] [CrossRef]
- Challis, G.L.; Hopwood, D.A. Synergy and contingency as driving forces for the evolution of multiple secondary metabolite production by Streptomyces species. Proc. Nat. Acad. Sci. USA 2003, 100, 14555–14561. [Google Scholar] [CrossRef] [PubMed]
- Medema, M.H.; Kottmann, R.; Yilmaz, P.; Cummings, M.; Biggins, J.B.; Blin, K.; De Bruijn, I.; Chooi, Y.H.; Claesen, J.; Coates, R.C.; et al. Minimum Information about a Biosynthetic Gene cluster. Nat. Chem. Biol. 2015, 11, 625–631. [Google Scholar] [CrossRef]
- Cdc. Antibiotic Resistance Threats Report In The United States 2019. Available online: https://www.cdc.gov/drugresistance/pdf/threats-report/2019-ar-threats-report-508.pdf (accessed on 16 June 2023).
- De Simeis, D.; Serra, S. Actinomycetes: A Never-Ending Source of Bioactive Compounds—An Overview on Antibiotics Production. Antibiotics 2021, 10, 483. [Google Scholar] [CrossRef]
- Euanorasetr, J.; Nilvongse, A.; Tantimavanich, S.; Nihira, T.; Igarashi, Y.; Panbangred, W. Identification and characterization of soil-isolated Streptomyces SJE177 producing actinomycin. Southeast Asian J. Trop. Med. Public Health 2010, 41, 1177–1187. [Google Scholar]
- Euanorasetr, J.; Intra, B.; Mongkol, P.; Chankhamhaengdecha, S.; Tuchinda, P.; Mori, M.; Shiomi, K.; Nihira, T.; Panbangred, W. Spirotetronate antibiotics with anti-Clostridium activity from Actinomadura sp. 2EPS. W. J. Microbiol. Biotech. 2015, 31, 391–398. [Google Scholar] [CrossRef] [PubMed]
- Iorio, M.; Cruz, J.; Simone, M.; Bernasconi, A.; Brunati, C.; Sosio, M.; Donadio, S.; Maffioli, S.I. Antibacterial Paramagnetic Quinones from Actinoallomurus. J. Nat. Prod. 2017, 80, 819–827. [Google Scholar] [CrossRef] [PubMed]
- Kodani, S.; Komaki, H.; Ishimura, S.; Hemmi, H.; Ohnishi-Kameyama, M. Isolation and structure determination of a new lantibiotic cinnamycin B from Actinomadura atramentaria based on genome mining. J. Ind. Microbiol. Biotechnol. 2016, 43, 1159–1165. [Google Scholar] [CrossRef]
- Shin, B.; Kim, B.Y.; Cho, E.; Oh, K.B.; Shin, J.; Goodfellow, M.; Oh, D.C. Actinomadurol, an Antibacterial Norditerpenoid from a Rare Actinomycete, Actinomadura sp. KC 191. J. Nat. Prod. 2016, 79, 1886–1890. [Google Scholar] [CrossRef] [PubMed]
- Bauermeister, A.; Calil, F.A.; Pinto, F.d.C.L.; Medeiros, T.C.T.; Almeida, L.C.; Silva, L.J.; de Melo, I.S.; Zucchi, T.D.; Costa-Lotufo, L.V.; Moraes, L.A.B. Pradimicin-IRD from Amycolatopsis sp. IRD-009 and its antimicrobial and cytotoxic activities. Nat. Prod. Res. 2019, 33, 1713–1720. [Google Scholar] [CrossRef] [PubMed]
- Beemelmanns, C.; Ramadhar, T.R.; Kim, K.H.; Klassen, J.L.; Cao, S.; Wyche, T.P.; Hou, Y.; Poulsen, M.; Bugni, T.S.; Currie, C.R.; et al. Macrotermycins A-D, Glycosylated Macrolactams from a Termite-Associated Amycolatopsis sp. M39. Org. Lett. 2017, 19, 1000–1003. [Google Scholar] [CrossRef]
- Khan, A.; Said, M.S.; Borade, B.R.; Gonnade, R.; Barvkar, V.; Kontham, R.; Dastager, S.G. Enceleamycins A-C, Furo-Naphthoquinones from Amycolatopsis sp. MCC0218: Isolation, Structure Elucidation, and Antimicrobial Activity. J. Nat. Prod. 2022, 85, 1267–1273. [Google Scholar] [CrossRef]
- Hashizume, H.; Sawa, R.; Yamashita, K.; Nishimura, Y.; Igarashi, M. Structure and antibacterial activities of new cyclic peptide antibiotics, pargamicins B, C and D, from Amycolatopsis sp. ML1-hF4. J. Ant. 2017, 70, 699–704. [Google Scholar] [CrossRef]
- Zhang, J.; Liang, X.; Zhang, S.; Song, Z.; Wang, C.; Xu, Y. Maipomycin A, a Novel Natural Compound With Promising Anti-biofilm Activity Against Gram-Negative Pathogenic Bacteria. Front. Microbiol. 2021, 11, 598024. [Google Scholar] [CrossRef]
- Kohda, Y.; Sakamoto, S.; Umekita, M.; Kimura, T.; Kubota, Y.; Arisaka, R.; Shibuya, Y.; Muramatsu, H.; Sawa, R.; Dan, S.; et al. Isolation of new derivatives of the 20-membered macrodiolide bispolide from Kitasatospora sp. MG372-hF19. J. Antibiot. 2021, 75, 77–85. [Google Scholar] [CrossRef]
- Uzair, B.; Menaa, F.; Khan, B.A.; Mohammad, F.V.; Ahmad, V.U.; Djeribi, R.; Menaa, B. Isolation, purification, structural elucidation and antimicrobial activities of kocumarin, a novel antibiotic isolated from actinobacterium Kocuria marina CMG S2 associated with the brown seaweed Pelvetia canaliculata. Microbiol. Res. 2018, 206, 186–197. [Google Scholar] [CrossRef] [PubMed]
- Gong, T.; Zhen, X.; Li, X.L.; Chen, J.J.; Chen, T.J.; Yang, J.L.; Zhu, P. Tetrocarcin Q, a New Spirotetronate with a Unique Glycosyl Group from a Marine-Derived Actinomycete Micromonospora carbonacea LS276. Mar. Drugs 2018, 16, 74. [Google Scholar] [CrossRef] [PubMed]
- Shin, Y.H.; Bae, S.; Sim, J.; Hur, J.; Jo, S.I.; Shin, J.; Suh, Y.G.; Oh, K.B.; Oh, D.C. Nicrophorusamides A and B, Antibacterial Chlorinated Cyclic Peptides from a Gut Bacterium of the Carrion Beetle Nicrophorus concolor. J. Nat. Prod. 2017, 80, 2962–2968. [Google Scholar] [CrossRef] [PubMed]
- Gui, C.; Zhang, S.; Zhu, X.; Ding, W.; Huang, H.; Gu, Y.C.; Duan, Y.; Ju, J. Antimicrobial Spirotetronate Metabolites from Marine-Derived Micromonospora harpali SCSIO GJ089. J. Nat. Prod. 2017, 80, 1594–1603. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Xie, L.; Zhao, W.; Zhou, J.; Jiang, H.; Liu, W.; Jiang, H.; Lin, F. Two new rakicidin derivatives from marine Micromonospora chalcea FIM-R160609. Nat. Prod. Res. 2022, 36, 1–8. [Google Scholar] [CrossRef]
- Hifnawy, M.S.; Hassan, H.M.; Mohammed, R.; Fouda, M.M.; Sayed, A.M.; Hamed, A.A.; AbouZid, S.F.; Rateb, M.E.; Alhadrami, H.A.; Abdelmohsen, U.R. Induction of Antibacterial Metabolites by Co-Cultivation of Two Red-Sea-Sponge-Associated Actinomycetes Micromonospora sp. UR56 and Actinokinespora sp. EG49. Mar. Drugs 2020, 18, 243. [Google Scholar] [CrossRef] [PubMed]
- Adnani, N.; Chevrette, M.G.; Adibhatla, S.N.; Zhang, F.; Yu, Q.; Braun, D.R.; Nelson, J.; Simpkins, S.W.; McDonald, B.R.; Myers, C.L.; et al. Coculture of Marine Invertebrate-Associated Bacteria and Interdisciplinary Technologies Enable Biosynthesis and Discovery of a New Antibiotic, Keyicin. ACS Chem. Biol. 2017, 12, 3093–3102. [Google Scholar] [CrossRef]
- Tan, Y.; Hu, Y.; Wang, Q.; Zhou, H.; Wang, Y.; Gan, M. Tetrocarcins N and O, glycosidic spirotetronates from a marine-derived Micromonospora sp. identified by PCR-based screening. RSC Adv. 2016, 6, 91773–91778. [Google Scholar] [CrossRef]
- Pérez-Bonilla, M.; Oves-Costales, D.; De La Cruz, M.; Kokkini, M.; Martín, J.; Vicente, F.; Genilloud, O.; Reyes, F. Phocoenamicins B and C, New Antibacterial Spirotetronates Isolated from a Marine Micromonospora sp. Mar. Drugs 2018, 16, 95. [Google Scholar] [CrossRef]
- Williams, D.E.; Dalisay, D.S.; Chen, J.; Polishchuck, E.A.; Patrick, B.O.; Narula, G.; Ko, M.; Av-Gay, Y.; Li, H.; Magarvey, N.; et al. Aminorifamycins and Sporalactams Produced in Culture by a Micromonospora sp. Isolated from a Northeastern-Pacific Marine Sediment Are Potent Antibiotics. Org. Lett. 2017, 19, 766–769. [Google Scholar] [CrossRef]
- Cheng, Z.; Zhang, Q.; Peng, J.; Zhao, X.; Ma, L.; Zhang, C.; Zhu, Y. Genomics-Driven Discovery of Benzoxazole Alkaloids from the Marine-Derived Micromonospora sp. SCSIO 07395. Molecules 2023, 28, 821. [Google Scholar] [CrossRef]
- Zhou, Q.; Luo, G.C.; Zhang, H.; Tang, G.L. Discovery of 16-Demethylrifamycins by Removing the Predominant Polyketide Biosynthesis Pathway in Micromonospora sp. Strain TP-A0468. Appl. Environ. Microbiol. 2019, 85, e02597-18. [Google Scholar] [CrossRef]
- Lasch, C.; Gummerlich, N.; Myronovskyi, M.; Palusczak, A.; Zapp, J.; Luzhetskyy, A. Loseolamycins: A Group of New Bioactive Alkylresorcinols Produced after Heterologous Expression of a Type III PKS from Micromonospora endolithica. Molecules 2020, 25, 4594. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Wen, Z.; Liu, L.; Zhu, X.; Shen, B.; Yan, X.; Duan, Y.; Huang, Y. Yangpumicins F and G, Enediyne Congeners from Micromonospora yangpuensis DSM 45577. J. Nat. Prod. 2019, 82, 2483–2488. [Google Scholar] [CrossRef] [PubMed]
- Shi, T.; Li, Y.-J.; Wang, Z.-M.; Wang, Y.-F.; Wang, B.; Shi, D.-Y. New Pyrroline Isolated from Antarctic Krill-Derived Actinomycetes Nocardiopsis sp. LX-1 Combining with Molecular Networking. Mar. Drugs 2023, 21, 127. [Google Scholar] [CrossRef] [PubMed]
- Siddharth, S.; Aswathanarayan, J.B.; Kuruburu, M.G.; Madhunapantula, S.R.V.; Vittal, R.R. Diketopiperazine derivative from marine actinomycetes Nocardiopsis sp. SCA30 with antimicrobial activity against MRSA. Arch. Microbiol. 2021, 203, 6173–6181. [Google Scholar] [CrossRef]
- Braña, A.F.; Sarmiento-Vizcaíno, A.; Pérez-Victoria, I.; Otero, L.; Fernández, J.; Palacios, J.J.; Martín, J.; De La Cruz, M.; Díaz, C.; Vicente, F.; et al. Branimycins B and C, Antibiotics Produced by the Abyssal Actinobacterium Pseudonocardia carboxydivorans M-227. J. Nat. Prod. 2017, 80, 569–573. [Google Scholar] [CrossRef] [PubMed]
- Matroodi, S.; Siitonen, V.; Baral, B.; Yamada, K.; Akhgari, A.; Metsä-Ketelä, M. Genotyping-Guided Discovery of Persiamycin A From Sponge-Associated Halophilic Streptomonospora sp. PA3. Front Microbiol. 2020, 11, 1237. [Google Scholar] [CrossRef]
- Pereira, F.; Almeida, J.R.; Paulino, M.; Grilo, I.R.; Macedo, H.; Cunha, I.; Sobral, R.G.; Vasconcelos, V.; Gaudêncio, S.P. Antifouling Napyradiomycins from Marine-Derived Actinomycetes Streptomyces aculeolatus. Mar. Drugs 2020, 18, 63. [Google Scholar] [CrossRef]
- Jia, J.; Zhang, C.; Liu, Y.; Huang, Y.; Bai, Y.; Hang, X.; Zeng, L.; Zhu, D.; Bi, H. Armeniaspirol A: A novel anti-Helicobacter pylori agent. Microb. Biotechnol. 2022, 15, 442–454. [Google Scholar] [CrossRef]
- Rodríguez Estévez, M.; Gummerlich, N.; Myronovskyi, M.; Zapp, J.; Luzhetskyy, A. Benzanthric Acid, a Novel Metabolite From Streptomyces albus Del14 Expressing the Nybomycin Gene Cluster. Front. Chem. 2020, 7, 896. [Google Scholar] [CrossRef] [PubMed]
- Manikkam, R.; Murthy, S.; Palaniappan, S.; Kaari, M.; Sahu, A.K.; Said, M.; Ganesan, V.; Kannan, S.; Ramasamy, B.; Thirugnanasambandan, S. Antibacterial and Anti-HIV Metabolites from Marine Streptomyces albus MAB56 Isolated from Andaman and Nicobar Islands, India. Appl. Biochem. Biotechnol. 2023, 195, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Chung, B.; Kwon, O.S.; Shin, J.; Oh, K.B. Antibacterial Activity and Mode of Action of Lactoquinomycin A from Streptomyces bacillaris. Mar. Drugs 2020, 19, 7. [Google Scholar] [CrossRef]
- Singh, R.; Dubey, A.K. Isolation and Characterization of a New Endophytic Actinobacterium Streptomyces californicus Strain ADR1 as a Promising Source of Anti-Bacterial, Anti-Biofilm and Antioxidant Metabolites. Microorganisms 2020, 8, 929. [Google Scholar] [CrossRef] [PubMed]
- Shaala, L.A.; Youssef, D.T.A.; Alzughaibi, T.A.; Elhady, S.S. Antimicrobial Chlorinated 3-Phenylpropanoic Acid Derivatives from the Red Sea Marine Actinomycete Streptomyces coelicolor LY001. Mar. Drugs 2020, 18, 450. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y.; Chen, N.; Li, J.; Su, J.C.; Yang, J.; Zhang, C.X.; Lin, H.W.; Zhou, Y. Antimicrobial Chlorinated Carbazole Alkaloids from the Sponge-Associated Actinomycete Streptomyces diacarni LHW51701. Chin. J. Chem. 2021, 39, 1188–1192. [Google Scholar] [CrossRef]
- Takehana, Y.; Umekita, M.; Hatano, M.; Kato, C.; Sawa, R.; Igarashi, M. Fradiamine A, a new siderophore from the deep-sea actinomycete Streptomyces fradiae MM456M-mF7. J. Antibiot. 2017, 70, 611–615. [Google Scholar] [CrossRef]
- Chen, Z.; Ou, P.; Liu, L.; Jin, X. Anti-MRSA Activity of Actinomycin X2 and Collismycin A Produced by Streptomyces globisporus WA5-2-37 From the Intestinal Tract of American Cockroach (Periplaneta americana). Front. Microbiol. 2020, 11, 555. [Google Scholar] [CrossRef]
- Kaweewan, I.; Komaki, H.; Hemmi, H.; Kodani, S. Isolation and structure determination of a new thiopeptide globimycin from Streptomyces globisporus subsp. globisporus based on genome mining. Tetrahedron Lett. 2018, 59, 409–414. [Google Scholar] [CrossRef]
- Saleem, M.; Hassan, A.; Li, F.; Lu, Q.; Ponomareva, L.V.; Parkin, S.; Sun, C.; Thorson, J.S.; Shaaban, K.A.; Sajid, I. Bioprospecting of desert actinobacteria with special emphases on griseoviridin, mitomycin C and a new bacterial metabolite producing Streptomyces sp. PU-KB10–4. BMC Microbiol. 2023, 23, 69. [Google Scholar] [CrossRef]
- Harunari, E.; Imada, C.; Igarashi, Y. Konamycins A and B and Rubromycins CA1 and CA2, Aromatic Polyketides from the Tunicate-Derived Streptomyces hyaluromycini MB-PO13T. J. Nat. Prod. 2019, 82, 1609–1615. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.; Song, Y.; Li, X.; Wang, X.; Ling, C.; Qin, X.; Zhou, Z.; Li, Q.; Wei, X.; Ju, J. Abyssomicin Monomers and Dimers from the Marine-Derived Streptomyces koyangensis SCSIO 5802. J. Nat. Prod. 2018, 81, 1892–1898. [Google Scholar] [CrossRef] [PubMed]
- Cho, E.; Kwon, O.S.; Chung, B.; Lee, J.; Sun, J.; Shin, J.; Oh, K.B. Antibacterial Activity of Chromomycins from a Marine-Derived Streptomyces microflavus. Mar. Drugs 2020, 18, 522. [Google Scholar] [CrossRef]
- Martinet, L.; Naômé, A.; Rezende, L.C.; Tellatin, D.; Pignon, B.; Docquier, J.-D.; Sannio, F.; Baiwir, D.; Mazzucchelli, G.; Frédérich, M. Lunaemycins, new cyclic hexapeptide antibiotics from the cave moonmilk-dweller Streptomyces lunaelactis MM109T. Int. J. Mol. Sci. 2023, 24, 1114. [Google Scholar] [CrossRef]
- Yang, L.; Hou, L.; Li, H.; Li, W. Antibiotic angucycline derivatives from the deepsea-derived Streptomyces lusitanus. Nat. Prod. Res. 2020, 34, 3444–3450. [Google Scholar] [CrossRef] [PubMed]
- Sujarit, K.; Mori, M.; Dobashi, K.; Shiomi, K.; Pathom-Aree, W.; Lumyong, S. New Antimicrobial Phenyl Alkenoic Acids Isolated from an Oil Palm Rhizosphere-Associated Actinomycete, Streptomyces palmae CMU-AB204T. Microorganisms 2020, 8, 350. [Google Scholar] [CrossRef]
- Xu, D.; Tian, E.; Kong, F.; Hong, K. Bioactive Molecules from Mangrove Streptomyces qinglanensis 172205. Mar. Drugs 2020, 18, 255. [Google Scholar] [CrossRef]
- Heo, C.-S.; Kang, J.S.; Kwon, J.-H.; Anh, C.V.; Shin, H.J. Pyrrole-Containing Alkaloids from a Marine-Derived Actinobacterium Streptomyces zhaozhouensis and Their Antimicrobial and Cytotoxic Activities. Mar. Drugs 2023, 21, 167. [Google Scholar] [CrossRef] [PubMed]
- Bo, S.T.; Xu, Z.F.; Yang, L.; Cheng, P.; Tan, R.X.; Jiao, R.H.; Ge, H.M. Structure and biosynthesis of mayamycin B, a new polyketide with antibacterial activity from Streptomyces sp. 120454. J. Antibiot. 2018, 71, 601–605. [Google Scholar] [CrossRef]
- Liang, Y.; Xie, X.; Chen, L.; Yan, S.; Ye, X.; Anjum, K.; Huang, H.; Lian, X.; Zhang, Z. Bioactive Polycyclic Quinones from Marine Streptomyces sp. 182SMLY. Mar. Drugs 2016, 14, 10. [Google Scholar] [CrossRef]
- Safaei, N.; Mast, Y.; Steinert, M.; Huber, K.; Bunk, B.; Wink, J. Angucycline-like Aromatic Polyketide from a Novel Streptomyces Species Reveals Freshwater Snail Physa acuta as Underexplored Reservoir for Antibiotic-Producing Actinomycetes. Antibiotics 2020, 10, 22. [Google Scholar] [CrossRef]
- Wang, T.; Li, F.; Lu, Q.; Wu, G.; Jiang, Z.; Liu, S.; Habden, X.; Razumova, E.A.; Osterman, I.A.; Sergiev, P.V.; et al. Diversity, novelty, antimicrobial activity, and new antibiotics of cultivable endophytic actinobacteria isolated from psammophytes collected from Taklamakan Desert. J. Pharm. Anal. 2021, 11, 241–250. [Google Scholar] [CrossRef]
- Guerrero-Garzón, J.F.; Zehl, M.; Schneider, O.; Rückert, C.; Busche, T.; Kalinowski, J.; Bredholt, H.; Zotchev, S.B. Streptomyces spp. From the Marine Sponge Antho dichotoma: Analyses of Secondary Metabolite Biosynthesis Gene Clusters and Some of Their Products. Front. Microbiol. 2020, 11, 437. [Google Scholar] [CrossRef]
- Shen, X.; Wang, X.; Huang, T.; Deng, Z.; Lin, S. Naphthoquinone-Based Meroterpenoids from Marine-Derived Streptomyces sp. B9173. Biomolecules 2020, 10, 1187. [Google Scholar] [CrossRef]
- Carretero-Molina, D.; Ortiz-López, F.J.; Martín, J.; Oves-Costales, D.; Díaz, C.; De La Cruz, M.; Cautain, B.; Vicente, F.; Genilloud, O.; Reyes, F. New Napyradiomycin Analogues from Streptomyces sp. Strain CA-271078. Mar. Drugs 2019, 18, 22. [Google Scholar] [CrossRef]
- Hu, X.; Hu, X.; Hu, X.; Li, S.; Li, L.; Yu, L.; Liu, H.; You, X.; Wang, Z.; Li, L.; et al. Cytotoxic and Antibacterial Cervinomycins B1-4 from a Streptomyces Species. J. Nat. Prod. 2019, 82, 2337–2342. [Google Scholar] [CrossRef] [PubMed]
- Dardić, D.; Lauro, G.; Bifulco, G.; Laboudie, P.; Sakhaii, P.; Bauer, A.; Vilcinskas, A.; Hammann, P.E.; Plaza, A. Svetamycins A-G, Unusual Piperazic Acid-Containing Peptides from Streptomyces sp. J. Org. Chem. 2017, 82, 6032–6043. [Google Scholar] [CrossRef] [PubMed]
- Lü, Y.; Shao, M.; Wang, Y.; Qian, S.; Wang, M.; Wang, Y.; Li, X.; Bao, Y.; Deng, C.; Yue, C.; et al. Zunyimycins B and C, New Chloroanthrabenzoxocinones Antibiotics against Methicillin-Resistant Staphylococcus aureus and Enterococci from Streptomyces sp. FJS31-2. Molecules 2017, 22, 251. [Google Scholar] [CrossRef]
- Kitani, S.; Ueguchi, T.; Igarashi, Y.; Leetanasaksakul, K.; Thamchaipenet, A.; Nihira, T. Rakicidin F, a new antibacterial cyclic depsipeptide from a marine sponge-derived Streptomyces sp. J. Ant. 2017, 71, 139–141. [Google Scholar] [CrossRef]
- Yang, Z.; Shao, L.; Wang, M.; Rao, M.; Ge, M.; Xu, Y. Two novel quinomycins discovered by UPLC-MS from Stretomyces sp. HCCB11876. J. Ant. 2019, 72, 164–168. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.X.; Ding, R.; Jiang, S.T.; Tang, J.S.; Hu, D.; Chen, G.D.; Lin, F.; Hong, K.; Yao, X.S.; Gao, H. Aldgamycins J-O, 16-Membered Macrolides with a Branched Octose Unit from Streptomycetes sp. and Their Antibacterial Activities. J. Nat. Prod. 2016, 79, 2446–2454. [Google Scholar] [CrossRef]
- Zhou, B.; Ji, Y.Y.; Zhang, H.J.; Shen, L. Gephyyamycin and cysrabelomycin, two new angucyclinone derivatives from the Streptomyces sp. HN-A124. Nat. Prod. Res. 2021, 35, 2117–2122. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Qi, H.; Zhang, H.; Zhang, S.-Y.; Zhang, C.-H.; Zhang, L.-Q.; Xiang, W.-S.; Wang, J.-D. Anulamycins A–F, Cinnamoyl-Containing Peptides from a Lake Sediment Derived Streptomyces. J. Nat. Prod. 2023, 86, 357–367. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Chen, L.; Zhang, X.; Liang, Y.; Anjum, K.; Chen, L.; Lian, X.Y. Bioactive Bafilomycins and a New N-Arylpyrazinone Derivative from Marine-derived Streptomyces sp. HZP-2216E. Planta Med. 2017, 83, 1405–1411. [Google Scholar] [CrossRef] [PubMed]
- Voitsekhovskaia, I.; Paulus, C.; Dahlem, C.; Rebets, Y.; Nadmid, S.; Zapp, J.; Axenov-Gribanov, D.; Rückert, C.; Timofeyev, M.; Kalinowski, J.; et al. Baikalomycins A-C, New Aquayamycin-Type Angucyclines Isolated from Lake Baikal Derived Streptomyces sp. IB201691-2A. Microorganisms 2020, 8, 680. [Google Scholar] [CrossRef]
- Iniyan, A.M.; Sudarman, E.; Wink, J.; Kannan, R.R.; Vincent, S.G.P. Ala-geninthiocin, a new broad spectrum thiopeptide antibiotic, produced by a marine Streptomyces sp. ICN19. J. Antibiot. 2019, 72, 99–105. [Google Scholar] [CrossRef]
- Son, S.; Hong, Y.S.; Jang, M.; Heo, K.T.; Lee, B.; Jang, J.P.; Kim, J.W.; Ryoo, I.J.; Kim, W.G.; Ko, S.K.; et al. Genomics-Driven Discovery of Chlorinated Cyclic Hexapeptides Ulleungmycins A and B from a Streptomyces Species. J. Nat. Prod. 2017, 80, 3025–3031. [Google Scholar] [CrossRef]
- Son, S.; Ko, S.K.; Kim, S.M.; Kim, E.; Kim, G.S.; Lee, B.; Ryoo, I.J.; Kim, W.G.; Lee, J.S.; Hong, Y.S.; et al. Antibacterial Cyclic Lipopeptide Enamidonins with an Enamide-Linked Acyl Chain from a Streptomyces Species. J. Nat. Prod. 2018, 81, 2462–2469. [Google Scholar] [CrossRef]
- Sawa, R.; Kubota, Y.; Umekita, M.; Hatano, M.; Hayashi, C.; Igarashi, M. Quadoctomycin, a 48-membered macrolide antibiotic from Streptomyces sp. MM168-141F8. J. Antibiot. 2017, 71, 91–96. [Google Scholar] [CrossRef]
- Konwar, A.N.; Basak, S.; Devi, S.G.; Borah, J.C.; Thakur, D. Streptomyces sp. MNP32, a forest-dwelling Actinomycetia endowed with potent antibacterial metabolites. 3 Biotech 2023, 13, 257. [Google Scholar] [CrossRef]
- Zhang, L.; Shi, J.; Liu, C.L.; Xiang, L.; Ma, S.Y.; Li, W.; Jiao, R.H.; Tan, R.X.; Ge, H.M. New borrelidin derivatives from an endophytic Streptomyces sp. Tetrahedron Lett. 2018, 59, 4517–4520. [Google Scholar] [CrossRef]
- Guo, Z.K.; Wang, Y.C.; Tan, Y.Z.; Abulaizi, A.; Xiong, Z.J.; Zhang, S.Q.; Yang, Y.; Yang, L.Y.; Shi, J. Nagimycins A and B, Antibacterial Ansamycin-Related Macrolactams from Streptomyces sp. NA07423. Org. Lett. 2023, 25, 4203–4207. [Google Scholar] [CrossRef] [PubMed]
- Arn, F.; Frasson, D.; Kroslakova, I.; Rezzonico, F.; Pothie, J.F.; Riedl, R.; Sievers, M. Isolation and Identification of Actinomycetes Strains from Switzerland and their Biotechnological Potential. Chimia 2020, 74, 382–390. [Google Scholar] [CrossRef]
- Mazumdar, R.; Dutta, P.P.; Saikia, J.; Borah, J.C.; Thakur, D. Streptomyces sp. Strain PBR11, a Forest-Derived Soil Actinomycetia with Antimicrobial Potential. Microbiol. Spectr. 2023, 11, 17544. [Google Scholar] [CrossRef]
- Betancur, L.A.; Forero, A.M.; Vinchira-Villarraga, D.M.; Cárdenas, J.D.; Romero-Otero, A.; Chagas, F.O.; Pupo, M.T.; Castellanos, L.; Ramos, F.A. NMR-based metabolic profiling to follow the production of anti-phytopathogenic compounds in the culture of the marine strain Streptomyces sp. PNM-9. Microbiol. Res. 2020, 239, 126507. [Google Scholar] [CrossRef]
- Cheng, C.; Othman, E.M.; Reimer, A.; Grüne, M.; Kozjak-Pavlovic, V.; Stopper, H.; Hentschel, U.; Abdelmohsen, U.R. Ageloline A, new antioxidant and antichlamydial quinolone from the marine sponge-derived bacterium Streptomyces sp. SBT345. Tetrahedron Lett. 2016, 57, 2786–2789. [Google Scholar] [CrossRef]
- Cong, Z.; Huang, X.; Liu, Y.; Liu, Y.; Wang, P.; Liao, S.; Yang, B.; Zhou, X.; Huang, D.; Wang, J. Cytotoxic anthracycline and antibacterial tirandamycin analogues from a marine-derived Streptomyces sp. SCSIO 41399. J. Antibiot. 2019, 72, 45–49. [Google Scholar] [CrossRef]
- Tian, H.; Shafi, J.; Ji, M.; Bi, Y.; Yu, Z. Antimicrobial Metabolites from Streptomyces sp. SN0280. J. Nat. Prod. 2017, 80, 1015–1019. [Google Scholar] [CrossRef]
- Maiti, P.K.; Das, S.; Sahoo, P.; Mandal, S. Streptomyces sp. SM01 isolated from Indian soil produces a novel antibiotic picolinamycin effective against multi drug resistant bacterial strains. Sci. Rep. 2020, 10, 788. [Google Scholar] [CrossRef]
- Shi, Y.; Wang, X.; He, N.; Xie, Y.; Hong, B. Rescrutiny of the sansanmycin biosynthetic gene cluster leads to the discovery of a novel sansanmycin analogue with more potency against Mycobacterium tuberculosis. J. Antibiot. 2019, 72, 769–774. [Google Scholar] [CrossRef]
- Pratiwi, R.H.; Hidayat, I.; Hanafi, M.; Mangunwardoyo, W. Isolation and structure elucidation of phenazine derivative from Streptomyces sp. strain UICC B-92 isolated from Neesia altissima (Malvaceae). Iran. J. Microbiol. 2020, 12, 127. [Google Scholar] [CrossRef]
- Jiang, Y.J.; Zhang, D.S.; Zhang, H.J.; Li, J.Q.; Ding, W.J.; Xu, C.D.; Ma, Z.J. Medermycin-Type Naphthoquinones from the Marine-Derived Streptomyces sp. XMA39. J. Nat. Prod. 2018, 81, 2120–2124. [Google Scholar] [CrossRef] [PubMed]
- Liu, T.; Ren, Z.; Chunyu, W.X.; Li, G.D.; Chen, X.; Zhang, Z.T.L.; Sun, H.B.; Wang, M.; Xie, T.P.; Wang, M.; et al. Exploration of Diverse Secondary Metabolites From Streptomyces sp. YINM00001, Using Genome Mining and One Strain Many Compounds Approach. Front. Microbiol. 2022, 13, 831174. [Google Scholar] [CrossRef] [PubMed]
- Newaz, A.W.; Yong, K.; Lian, X.Y.; Zhang, Z. Streptoindoles A–D, novel antimicrobial indole alkaloids from the marine-associated actinomycete Streptomyces sp. ZZ1118. Tetrahedron 2022, 104, 132598. [Google Scholar] [CrossRef]
- Zhang, D.; Yi, W.; Ge, H.; Zhang, Z.; Wu, B. Bioactive Streptoglutarimides A-J from the Marine-Derived Streptomyces sp. ZZ741. J. Nat. Prod. 2019, 82, 2800–2808. [Google Scholar] [CrossRef] [PubMed]
- Asolkar, R.N.; Singh, A.; Jensen, P.R.; Aalbersberg, W.; Carté, B.K.; Feussner, K.D.; Subramani, R.; DiPasquale, A.; Rheingold, A.L.; Fenical, W. Marinocyanins, cytotoxic bromo-phenazinone meroterpenoids from a marine bacterium from the streptomycete clade MAR4. Tetrahedron 2017, 73, 2234–2241. [Google Scholar] [CrossRef]
- Fujita, Y.; Kagoshima, Y.; Masuda, T.; Kizuka, M.; Ogawa, Y.; Endo, S.; Nishigoori, H.; Saito, K.; Takasugi, K.; Miura, M.; et al. Muraminomicins, new lipo-nucleoside antibiotics from Streptosporangium sp. SANK 60501-structure elucidations of muraminomicins and supply of the core component for derivatization. J. Antibiot. 2019, 72, 943–955. [Google Scholar] [CrossRef]
- Teta, R.; Marteinsson, V.T.; Longeon, A.; Klonowski, A.M.; Groben, R.; Bourguet-Kondracki, M.L.; Costantino, V.; Mangoni, A. Thermoactinoamide A, an Antibiotic Lipophilic Cyclopeptide from the Icelandic Thermophilic Bacterium Thermoactinomyces vulgaris. J. Nat. Prod. 2017, 80, 2530–2535. [Google Scholar] [CrossRef]
- Zhang, S.; Xie, Q.; Sun, C.; Tian, X.P.; Gui, C.; Qin, X.; Zhang, H.; Ju, J. Cytotoxic Kendomycins Containing the Carbacylic Ansa Scaffold from the Marine-Derived Verrucosispora sp. SCSIO 07399. J. Nat. Prod. 2019, 82, 3366–3371. [Google Scholar] [CrossRef]
- Gupte, M.; Kulkarni, P.; Ganguli, B.N. Antifungal antibiotics. Appl. Microbiol. Biotechnol. 2002, 58, 46–57. [Google Scholar] [CrossRef]
- Hazen, E.L.; Brown, R. Fungicidin, an Antibiotic Produced by a Soil Actinomycete. Proc. Soc. Exp. Biol. Med. 1951, 76, 93–97. [Google Scholar] [CrossRef]
- Georgopapadakou, N.H. Antifungals: Mechanism of action and resistance, established and novel drugs. Curr. Opin. Microbiol. 1998, 1, 547–557. [Google Scholar] [CrossRef]
- Ghannoum, M.A.; Rice, L.B. Antifungal agents: Mode of action, mechanisms of resistance, and correlation of these mechanisms with bacterial resistance. Clin. Microbiol. Rev. 1999, 12, 501–517. [Google Scholar] [CrossRef] [PubMed]
- Intra, B.; Euanorasetr, J.; Nihira, T.; Panbangred, W. Characterization of a gamma-butyrolactone synthetase gene homologue (stcA) involved in bafilomycin production and aerial mycelium formation in Streptomyces sp. SBI034. Appl. Microbiol. Biotechnol. 2016, 100, 2749–2760. [Google Scholar] [CrossRef] [PubMed]
- Intra, B.; Greule, A.; Bechthold, A.; Euanorasetr, J.; Paululat, T.; Panbangred, W. Thailandins A and B, New Polyene Macrolactone Compounds Isolated from Actinokineospora bangkokensis Strain 44EHWT, Possessing Antifungal Activity against Anthracnose Fungi and Pathogenic Yeasts. J. Agric. Food Chem. 2016, 64, 5171–5179. [Google Scholar] [CrossRef] [PubMed]
- Euanorasetr, J.; Junhom, M.; Tantimavanich, S.; Vorasin, O.; Munyoo, B.; Tuchinda, P.; Panbangred, W. Halogenated benzoate derivatives of altholactone with improved anti-fungal activity. J. Asian Nat. Prod. Res. 2016, 18, 462–474. [Google Scholar] [CrossRef]
- Bunyapaiboonsri, T.; Yoiprommarat, S.; Suriyachadkun, C.; Supothina, S.; Chanthaket, R.; Chutrakul, C.; Vichai, V. Actinomadurone, a polycyclic tetrahydroxanthone from Actinomadura sp. BCC 35430. Tetrahedron Lett. 2017, 58, 3223–3225. [Google Scholar] [CrossRef]
- Cheng, M.-J.; Chen, J.-J.; Wu, M.-D.; Leu, J.-Y.; Tseng, M. Antifungal Activities of Compounds Produced by Newly Isolated Acrocarpospora Strains. Antibiotics 2023, 12, 95. [Google Scholar] [CrossRef]
- Kim, H.R.; Kim, J.; Yu, J.S.; Lee, B.S.; Kim, K.H.; Kim, C.S. Isolation, structure elucidation, total synthesis, and biosynthesis of dermazolium A, an antibacterial imidazolium metabolite of a vaginal bacterium Dermabacter vaginalis. Arch. Pharm. Res. 2023, 46, 35–43. [Google Scholar] [CrossRef]
- Hara, S.; Ishikawa, N.; Hara, Y.; Nehira, T.; Sakai, K.; Gonoi, T.; Ishibashi, M. Nabscessins A and B, Aminocyclitol Derivatives from Nocardia abscessus IFM 10029T. J. Nat. Prod. 2017, 80, 565–568. [Google Scholar] [CrossRef]
- Kim, H.J.; Han, C.Y.; Park, J.S.; Oh, S.H.; Kang, S.H.; Choi, S.S.; Kim, J.M.; Kwak, J.H.; Kim, E.S. Nystatin-like Pseudonocardia polyene B1, a novel disaccharide-containing antifungal heptaene antibiotic. Sci. Rep. 2018, 8, 13584. [Google Scholar] [CrossRef]
- Li, Z.; Gao, X.; Kang, Z.; Huang, L.; Fan, D.; Yan, X. Saccharothrix yanglingensis Strain Hhs.015 Is a Promising Biocontrol Agent on Apple Valsa Canker. Plant Dis. 2016, 100, 510–514. [Google Scholar] [CrossRef]
- Thekkangil, A.; George, B.; Prakash, S.M.U.; Suchithra, T.V. Mechanism of Streptomyces albidoflavus STV1572a derived 1-heneicosanol as an inhibitor against squalene epoxidase of Trichophyton mentagrophytes. Microb. Pathog. 2021, 154, 104853. [Google Scholar] [CrossRef]
- Ding, N.; Jiang, Y.; Han, L.; Chen, X.; Ma, J.; Qu, X.; Mu, Y.; Liu, J.; Li, L.; Jiang, C.; et al. Bafilomycins and Odoriferous Sesquiterpenoids from Streptomyces albolongus Isolated from Elephas maximus Feces. J. Nat. Prod. 2016, 79, 799–805. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Fu, S.N.; Bao, Y.X.; Yang, Y.; Shen, H.F.; Lin, B.R.; Zhou, G.X. Kitamycin C, a new antimycin-type antibiotic from Streptomyces antibioticus strain 200-09. Nat. Prod. Res. 2017, 31, 1819–1824. [Google Scholar] [CrossRef] [PubMed]
- Pérez, M.; Schleissner, C.; Fernández, R.; Rodríguez, P.; Reyes, F.; Zuñiga, P.; De La Calle, F.; Cuevas, C. PM100117 and PM100118, new antitumor macrolides produced by a marine Streptomyces caniferus GUA-06-05-006A. J. Antibiot. 2016, 69, 388–394. [Google Scholar] [CrossRef] [PubMed]
- Vicente Dos Reis, G.; Abraham, W.R.; Grigoletto, D.F.; De Campos, J.B.; Marcon, J.; Da Silva, J.A.; Quecine, M.C.; De Azevedo, J.L.; Ferreira, A.G.; De Lira, S.P. Gloeosporiocide, a new antifungal cyclic peptide from Streptomyces morookaense AM25 isolated from the Amazon bulk soil. FEMS Microbiol. Lett. 2019, 366, 175. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Yu, Z.; Zhao, J.; Zhuang, X.; Cao, P.; Guo, X.; Liu, C.; Xiang, W. Community Composition, Antifungal Activity and Chemical Analyses of Ant-Derived Actinobacteria. Front. Microbiol. 2020, 11, 201. [Google Scholar] [CrossRef]
- Mojicevic, M.; D’Agostino, P.M.; Pavic, A.; Vojnovic, S.; Senthamaraikannan, R.; Vasiljevic, B.; Gulder, T.A.M.; Nikodinovic-Runic, J. Streptomyces sp. BV410 isolate from chamomile rhizosphere soil efficiently produces staurosporine with antifungal and antiangiogenic properties. Microbiol. Open 2020, 9, e986. [Google Scholar] [CrossRef]
- Shi, J.; Peng, D.; Peng, F.-F.; Zhang, Q.-B.; Duan, Y.-W.; Huang, Y. The Isolation and Structure Elucidation of Spirotetronate Lobophorins A, B, and H8 from Streptomyces sp. CB09030 and Their Biosynthetic Gene Cluster. Molecules 2023, 28, 3597. [Google Scholar] [CrossRef]
- Fang, Q.; Maglangit, F.; Mugat, M.; Urwald, C.; Kyeremeh, K.; Deng, H. Targeted Isolation of Indole Alkaloids from Streptomyces sp. CT37. Molecules 2020, 25, 1108. [Google Scholar] [CrossRef] [PubMed]
- Xiao, L.; Niu, H.J.; Qu, T.L.; Zhang, X.F.; Du, F.Y. Streptomyces sp. FX13 inhibits fungicide-resistant Botrytis cinerea in vitro and in vivo by producing oligomycin A. Pestic. Biochem. Physiol. 2021, 175, 104834. [Google Scholar] [CrossRef] [PubMed]
- Cao, P.; Li, C.; Wang, H.; Yu, Z.; Xu, X.; Wang, X.; Zhao, J.; Xiang, W. Community Structures and Antifungal Activity of Root-Associated Endophytic Actinobacteria in Healthy and Diseased Cucumber Plants and Streptomyces sp. HAAG3-15 as a Promising Biocontrol Agent. Microorganisms 2020, 8, 236. [Google Scholar] [CrossRef]
- Yu, Z.; Wang, L.; Yang, J.; Zhang, F.; Sun, Y.; Yu, M.; Yan, Y.; Ma, Y.T.; Huang, S.X. A new antifungal macrolide from Streptomyces sp. KIB-H869 and structure revision of halichomycin. Tetrahedron Lett. 2016, 57, 1375–1378. [Google Scholar] [CrossRef]
- Feng, X.-L.; Zhang, R.-Q.; Wang, D.-C.; Dong, W.-G.; Wang, Z.-X.; Zhai, Y.-J.; Han, W.-B.; Yin, X.; Tian, J.; Wei, J. Genomic and Metabolite Profiling Reveal a Novel Streptomyces Strain, QHH-9511, from the Qinghai-Tibet Plateau. Microbiol. Spectr. 2023, 11, e02764-02722. [Google Scholar] [CrossRef] [PubMed]
- Yi, W.; Qin, L.; Lian, X.Y.; Zhang, Z. New Antifungal Metabolites from the Mariana Trench Sediment-Associated Actinomycete Streptomyces sp. SY1965. Mar. Drugs 2020, 18, 385. [Google Scholar] [CrossRef]
- Herbrík, A.; Corretto, E.; Chroňáková, A.; Langhansová, H.; Petrásková, P.; Hrdý, J.; Čihák, M.; Krištůfek, V.; Bobek, J.; Petříček, M.; et al. A Human Lung-Associated Streptomyces sp. TR1341 Produces Various Secondary Metabolites Responsible for Virulence, Cytotoxicity and Modulation of Immune Response. Front. Microbiol. 2020, 10, 3028. [Google Scholar] [CrossRef]
- Dutta, J.; Thakur, D. Evaluation of Antagonistic and Plant Growth Promoting Potential of Streptomyces sp. TT3 Isolated from Tea (Camellia sinensis) Rhizosphere Soil. Curr. Microbiol. 2020, 77, 1829–1838. [Google Scholar] [CrossRef]
- Yamamoto, K.; Futamura, Y.; Uson-Lopez, R.A.; Aono, H.; Shimizu, T.; Osada, H. YO-001A, a new antifungal agent produced by Streptomyces sp. YO15-A001. J. Antibiot. 2019, 72, 986–990. [Google Scholar] [CrossRef]
- Gopalakrishnan, S.; Sharma, R.; Srinivas, V.; Naresh, N.; Mishra, S.P.; Ankati, S.; Pratyusha, S.; Govindaraj, M.; Gonzalez, S.V.; Nervik, S.; et al. Identification and Characterization of a Streptomyces albus Strain and Its Secondary Metabolite Organophosphate against Charcoal Rot of Sorghum. Plants 2020, 9, 1727. [Google Scholar] [CrossRef]
- Hoshino, S.; Wong, C.P.; Ozeki, M.; Zhang, H.; Hayashi, F.; Awakawa, T.; Asamizu, S.; Onaka, H.; Abe, I. Umezawamides, new bioactive polycyclic tetramate macrolactams isolated from a combined-culture of Umezawaea sp. and mycolic acid-containing bacterium. J. Antibiot. 2018, 71, 653–657. [Google Scholar] [CrossRef]
- Barreiro, C.; Prieto, C.; Sola-Landa, A.; Solera, E.; Martínez-Castro, M.; Pérez-Redondo, R.; García-Estrada, C.; Aparicio, J.F.; Fernández-Martínez, L.T.; Santos-Aberturas, J.; et al. Draft genome of Streptomyces tsukubaensis NRRL 18488, the producer of the clinically important immunosuppressant tacrolimus (FK506). J. Bacteriol. 2012, 194, 3756–3757. [Google Scholar] [CrossRef] [PubMed]
- Bierer, B.E.; Mattila, P.S.; Standaert, R.F.; Herzenberg, L.A.; Burakoff, S.J.; Crabtree, G.; Schreiber, S.L. Two distinct signal transmission pathways in T lymphocytes are inhibited by complexes formed between an immunophilin and either FK506 or rapamycin. Proc. Nat. Acad. Sci. USA 1990, 87, 9231–9235. [Google Scholar] [CrossRef] [PubMed]
- Ohta, Y.; Ikeda, M. Deodorization of pig feces by actinomycetes. Appl. Environ. Microbiol. 1978, 36, 487–491. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, Y.; Murata, A.; Shinsaku, H. Accelerated composting of cereal shochu-distillery wastes by actinomycetes. J. Ferment. Bioeng. 1995, 80, 421. [Google Scholar] [CrossRef]
- Mansour, F.A.; Mohamedin, A.H. Enzymes of Candida albicans cell-wall lytic system produced by Streptomyces thermodiastaticus. Acta Microbiol. Immunol. Hung. 2001, 48, 53–65. [Google Scholar] [CrossRef] [PubMed]
- Vurukonda, S.S.K.P.; Giovanardi, D.; Stefani, E. Plant Growth Promoting and Biocontrol Activity of Streptomyces spp. as Endophytes. Int. J. Mol. Sci. 2018, 19, 952. [Google Scholar] [CrossRef]
- Subramani, R.; Sipkema, D. Marine Rare Actinomycetes: A Promising Source of Structurally Diverse and Unique Novel Natural Products. Mar. Drugs 2019, 17, 249. [Google Scholar] [CrossRef]
- Lee, H.S.; Jeong, G.S. Salinosporamide A, a Marine-Derived Proteasome Inhibitor, Inhibits T Cell Activation through Regulating Proliferation and the Cell Cycle. Molecules 2020, 25, 5031. [Google Scholar] [CrossRef]
- Arcamone, F.; Cassinelli, G.; Fantini, G.; Grein, A.; Orezzi, P.; Pol, C.; Spalla, C. Adriamycin, 14-hydroxydaimomycin, a new antitumor antibiotic from S. peucetius var. caesius. Biotechnol. Bioeng. 1969, 11, 1101–1110. [Google Scholar] [CrossRef]
- Jagannathan, S.V.; Manemann, E.M.; Rowe, S.E.; Callender, M.C.; Soto, W. Marine Actinomycetes, New Sources of Biotechnological Products. Mar. Drugs 2021, 19, 365. [Google Scholar] [CrossRef] [PubMed]
- Igarashi, Y.; Shimasaki, R.; Miyanaga, S.; Oku, N.; Onaka, H.; Sakurai, H.; Saiki, I.; Kitani, S.; Nihira, T.; Wimonsiravude, W.; et al. Rakicidin D, an inhibitor of tumor cell invasion from marine-derived Streptomyces sp. J. Antibiot. 2010, 63, 563–565. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.; Chen, Y.; Ukaji, T.; Okada, S.; Umezawa, K. Isolation of ketomycin from Actinomycetes as an inhibitor of 2D and 3D cancer cell invasion. J. Antibiot. 2018, 72, 148–154. [Google Scholar] [CrossRef] [PubMed]
- Choi, B.K.; Lee, H.S.; Kang, J.S.; Shin, H.J. Dokdolipids A−C, Hydroxylated Rhamnolipids from the Marine-Derived Actinomycete Actinoalloteichus hymeniacidonis. Mar. Drugs 2019, 17, 237. [Google Scholar] [CrossRef] [PubMed]
- Kimura, T.; Iwatsuki, M.; Asami, Y.; Ishiyama, A.; Hokari, R.; Otoguro, K.; Matsumoto, A.; Sato, N.; Shiomi, K.; Takahashi, Y.; et al. Anti-trypanosomal compound, sagamilactam, a new polyene macrocyclic lactam from Actinomadura sp. K13-0306. J. Antibiot. 2016, 69, 818–824. [Google Scholar] [CrossRef] [PubMed]
- Igarashi, Y.; Matsuoka, N.; In, Y.; Kataura, T.; Tashiro, E.; Saiki, I.; Sudoh, Y.; Duangmal, K.; Thamchaipenet, A. Nonthmicin, a Polyether Polyketide Bearing a Halogen-Modified Tetronate with Neuroprotective and Antiinvasive Activity from Actinomadura sp. Org. Lett. 2017, 19, 1406–1409. [Google Scholar] [CrossRef]
- Lu, C.; Xie, F.; Shan, C.; Shen, Y. Two novel cyclic hexapeptides from the genetically engineered Actinosynnema pretiosum. Appl. Microbiol. Biotechnol. 2017, 101, 2273–2279. [Google Scholar] [CrossRef]
- Frattaruolo, L.; Lacret, R.; Cappello, A.R.; Truman, A.W. A Genomics-Based Approach Identifies a Thioviridamide-Like Compound with Selective Anticancer Activity. ACS Chem. Biol. 2017, 12, 2815–2822. [Google Scholar] [CrossRef]
- Hoshino, S.; Ozeki, M.; Awakawa, T.; Morita, H.; Onaka, H.; Abe, I. Catenulobactins A and B, Heterocyclic Peptides from Culturing Catenuloplanes sp. with a Mycolic Acid-Containing Bacterium. J. Nat. Prod. 2018, 81, 2106–2110. [Google Scholar] [CrossRef]
- Pournejati, R.; Gust, R.; Kircher, B.; Karbalaei-Heidari, H.R. Microindoline 581, an Indole Derivative from Microbacterium Sp. RP581 as A Novel Selective Antineoplastic Agent to Combat Hepatic Cancer Cells: Production, Optimization and Structural Elucidation. Iran. J. Pharm. Res. 2020, 19, 290–305. [Google Scholar] [CrossRef]
- Fu, G.; Wang, R.; Ding, J.; Qi, H.; Zhao, Z.; Chen, C.; Zhang, H.; Xue, Z.; Wang, J.; Wu, M. Micromonospora zhangzhouensis sp. nov., a Novel Actinobacterium Isolated from Mangrove Soil, Exerts a Cytotoxic Activity in vitro. Sci. Rep. 2020, 10, 3889. [Google Scholar] [CrossRef] [PubMed]
- Sarmiento-Vizcaíno, A.; Braña, A.F.; Pérez-Victoria, I.; Martín, J.; De Pedro, N.; De La Cruz, M.; Díaz, C.; Vicente, F.; Acuña, J.L.; Reyes, F.; et al. Paulomycin G, a New Natural Product with Cytotoxic Activity against Tumor Cell Lines Produced by Deep-Sea Sediment Derived Micromonospora matsumotoense M-412 from the Avilés Canyon in the Cantabrian Sea. Mar. Drugs 2017, 15, 271. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.J.; Zhang, S.Y.; Ye, Y.H.; Yu, Z.; Qi, H.; Zhang, H.; Xue, Z.L.; Wang, J.D.; Wu, M. Three New Isoflavonoid Glycosides from the Mangrove-Derived Actinomycete Micromonospora aurantiaca 110B. Mar. Drugs 2019, 17, 294. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Zhao, W.; Jiang, H.L.; Zhou, J.; Chen, X.M.; Lian, Y.Y.; Jiang, H.; Lin, F. Rakicidins G-I, cyclic depsipeptides from marine Micromonospora chalcea FIM 02-523. Tetrahedron 2018, 74, 4151–4154. [Google Scholar] [CrossRef]
- Nie, Y.L.; Wu, Y.D.; Wang, C.X.; Lin, R.; Xie, Y.; Fang, D.S.; Jiang, H.; Lian, Y.Y. Structure elucidation and antitumour activity of a new macrolactam produced by marine-derived actinomycete Micromonospora sp. FIM05328. Nat. Prod. Res. 2018, 32, 2133–2138. [Google Scholar] [CrossRef]
- Gao, M.Y.; Qi, H.; Li, J.S.; Zhang, H.; Zhang, J.; Wang, J.D.; Xiang, W.S. A new naphthalenepropanoic acid analog from the marine-derived actinomycetes Micromonospora sp. HS-HM-036. J. Asian Nat. Prod. Res. 2017, 19, 930–934. [Google Scholar] [CrossRef] [PubMed]
- Yan, X.; Chen, J.J.; Adhikari, A.; Yang, D.; Crnovcic, I.; Wang, N.; Chang, C.Y.; Rader, C.; Shen, B. Genome Mining of Micromonospora yangpuensis DSM 45577 as a Producer of an Anthraquinone-Fused Enediyne. Org. Lett. 2017, 19, 6192–6195. [Google Scholar] [CrossRef]
- Fukuda, T.; Takahashi, M.; Nagai, K.; Harunari, E.; Imada, C.; Tomoda, H. Isomethoxyneihumicin, a new cytotoxic agent produced by marine Nocardiopsis alba KM6-1. J. Antibiot. 2017, 70, 590–594. [Google Scholar] [CrossRef]
- Messaoudi, O.; Sudarman, E.; Bendahou, M.; Jansen, R.; Stadler, M.; Wink, J. Kenalactams A-E, Polyene Macrolactams Isolated from Nocardiopsis CG3. J. Nat. Prod. 2019, 82, 1081–1088. [Google Scholar] [CrossRef]
- Shaaban, K.A.; Shaaban, M.; Rahman, H.; Grün-Wollny, I.; Kämpfer, P.; Kelter, G.; Fiebig, H.H.; Laatsch, H. Karamomycins A-C: 2-Naphthalen-2-yl-thiazoles from Nonomuraea endophytica. J. Nat. Prod. 2019, 82, 870–877. [Google Scholar] [CrossRef]
- Yang, T.; Yamada, K.; Zhou, T.; Harunari, E.; Igarashi, Y.; Terahara, T.; Kobayashi, T.; Imada, C. Akazamicin, a cytotoxic aromatic polyketide from marine-derived Nonomuraea sp. J. Antibiot. 2019, 72, 202–209. [Google Scholar] [CrossRef] [PubMed]
- Gamaleldin, N.M.; Bakeer, W.; Sayed, A.M.; Shamikh, Y.I.; El-Gendy, A.O.; Hassan, H.M.; Horn, H.; Abdelmohsen, U.R.; Hozzein, W.N. Exploration of Chemical Diversity and Antitrypanosomal Activity of Some Red Sea-Derived Actinomycetes Using the OSMAC Approach Supported by LC-MS-Based Metabolomics and Molecular Modelling. Antibiotics 2020, 9, 629. [Google Scholar] [CrossRef] [PubMed]
- Anh, C.V.; Kwon, J.-H.; Kang, J.S.; Lee, H.-S.; Heo, C.-S.; Shin, H.J. New Angucycline Glycosides from a Marine-Derived Bacterium Streptomyces ardesiacus. Int. J. Mol. Sci. 2022, 23, 13779. [Google Scholar] [CrossRef] [PubMed]
- Kaari, M.; Joseph, J.; Manikkam, R.; Kalyanasundaram, R.; Sivaraj, A.; Anbalmani, S.; Murthy, S.; Sahu, A.K.; Said, M.; Dastager, S.G. A Novel Finding: 2, 4-Di-tert-butylphenol from Streptomyces bacillaris ANS2 Effective Against Mycobacterium tuberculosis and Cancer Cell Lines. Appl. Biochem. Biotechnol. 2023, 195, 1–14. [Google Scholar] [CrossRef]
- Gui, C.; Yuan, J.; Mo, X.; Huang, H.; Zhang, S.; Gu, Y.C.; Ju, J. Cytotoxic Anthracycline Metabolites from a Recombinant Streptomyces. J. Nat. Prod. 2018, 81, 1278–1289. [Google Scholar] [CrossRef]
- Kifer, D.; Mužinić, V.; Klaric, M.Š. Antimicrobial potency of single and combined mupirocin and monoterpenes, thymol, menthol and 1,8-cineole against Staphylococcus aureus planktonic and biofilm growth. J. Antibiot. 2016, 69, 689–696. [Google Scholar] [CrossRef]
- Xiao, F.; Li, H.; Xu, M.; Li, T.; Wang, J.; Sun, C.; Hong, K.; Li, W. Staurosporine Derivatives Generated by Pathway Engineering in a Heterologous Host and Their Cytotoxic Selectivity. J. Nat. Prod. 2018, 81, 1745–1751. [Google Scholar] [CrossRef]
- Kaweewan, I.; Komaki, H.; Hemmi, H.; Hoshino, K.; Hosaka, T.; Isokawa, G.; Oyoshi, T.; Kodani, S. Isolation and structure determination of a new cytotoxic peptide, curacozole, from Streptomyces curacoi based on genome mining. J. Antibiot. 2019, 72, 1–7. [Google Scholar] [CrossRef]
- Ortiz-López, F.J.; Alcalde, E.; Sarmiento-Vizcaíno, A.; Díaz, C.; Cautain, B.; García, L.A.; Blanco, G.; Reyes, F. New 3-Hydroxyquinaldic Acid Derivatives from Cultures of the Marine Derived Actinomycete Streptomyces cyaneofuscatus M-157. Mar. Drugs 2018, 16, 371. [Google Scholar] [CrossRef]
- Kimata, S.; Matsuda, T.; Suizu, Y.; Hayakawa, Y. Prodigiosin R2, a new prodigiosin from the roseophilin producer Streptomyces griseoviridis 2464-S5. J. Antibiot. 2018, 71, 393–396. [Google Scholar] [CrossRef]
- Liu, S.H.; Wang, W.; Wang, K.B.; Zhang, B.; Li, W.; Shi, J.; Jiao, R.H.; Tan, R.X.; Ge, H.M. Heterologous Expression of a Cryptic Giant Type I PKS Gene Cluster Leads to the Production of Ansaseomycin. Org. Lett. 2019, 21, 3785–3788. [Google Scholar] [CrossRef] [PubMed]
- Koomsiri, W.; Inahashi, Y.; Kimura, T.; Shiomi, K.; Takahashi, Y.; Omura, S.; Thamchaipenet, A.; Nakashima, T. Bisoxazolomycin A: A new natural product from ‘Streptomyces subflavus subsp. irumaensis’ AM-3603. J. Antibiot. 2017, 70, 1142–1145. [Google Scholar] [CrossRef]
- Zheng, D.; Ding, N.; Jiang, Y.; Zhang, J.; Ma, J.; Chen, X.; Liu, J.; Han, L.; Huang, X. Albaflavenoid, a new tricyclic sesquiterpenoid from Streptomyces violascens. J. Antibiot. 2016, 69, 773–775. [Google Scholar] [CrossRef] [PubMed]
- Zheng, D.; Han, L.; Qu, X.; Chen, X.; Zhong, J.; Bi, X.; Liu, J.; Jiang, Y.; Jiang, C.; Huang, X. Cytotoxic Fusicoccane-Type Diterpenoids from Streptomyces violascens Isolated from Ailuropoda melanoleuca Feces. J. Nat. Prod. 2017, 80, 837–844. [Google Scholar] [CrossRef]
- Lv, Q.; Fan, Y.; Tao, G.; Fu, P.; Zhai, J.; Ye, B.; Zhu, W. Sekgranaticin, a SEK34b-Granaticin Hybrid Polyketide from Streptomyces sp. 166. J. Org Chem. 2019, 84, 9087–9092. [Google Scholar] [CrossRef]
- Liang, Y.; Chen, L.; Ye, X.; Anjum, K.; Lian, X.Y.; Zhang, Z. New streptophenazines from marine Streptomyces sp. 182SMLY. Nat. Prod. Res. 2017, 31, 411–417. [Google Scholar] [CrossRef]
- Liu, C.X.; Liu, S.H.; Zhao, J.W.; Zhang, J.; Wang, X.J.; Li, J.S.; Wang, J.D.; Xiang, W.S. A new spectinabilin derivative with cytotoxic activity from ant-derived Streptomyces sp. 1H-GS5. J. Asian Nat. Prod. Res. 2017, 19, 924–929. [Google Scholar] [CrossRef]
- Zhou, B.; Qin, L.L.; Ding, W.J.; Ma, Z.J. Cytotoxic indolocarbazoles alkaloids from the Streptomyces sp. A65. Tetrahedron 2018, 74, 726–730. [Google Scholar] [CrossRef]
- Wang, X.; Elshahawi, S.I.; Ponomareva, L.V.; Ye, Q.; Liu, Y.; Copley, G.C.; Hower, J.C.; Hatcher, B.E.; Kharel, M.K.; Van Lanen, S.G.; et al. Structure Determination, Functional Characterization, and Biosynthetic Implications of Nybomycin Metabolites from a Mining Reclamation Site-Associated Streptomyces. J. Nat. Prod. 2019, 82, 3469–3476. [Google Scholar] [CrossRef]
- Saito, S.; Fujimaki, T.; Panbangred, W.; Sawa, R.; Igarashi, Y.; Imoto, M. Antarlides F–H, new members of the antarlide family produced by Streptomyces sp. BB47. J. Antibiot. 2017, 70, 595–600. [Google Scholar] [CrossRef]
- Han, X.; Liu, Z.; Zhang, Z.; Zhang, X.; Zhu, T.; Gu, Q.; Li, W.; Che, Q.; Li, D. Geranylpyrrol A and Piericidin F from Streptomyces sp. CHQ-64 ΔrdmF. J. Nat. Prod. 2017, 80, 1684–1687. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.N.; Zhang, H.J.; Li, J.Q.; Ding, W.J.; Ma, Z.J. Bioactive Indolocarbazoles from the Marine-Derived Streptomyces sp. DT-A61. J. Nat. Prod. 2018, 81, 949–956. [Google Scholar] [CrossRef] [PubMed]
- Abdelfattah, M.S.; Elmallah, M.I.Y.; Mohamed, A.A.; Ishibashi, M. Sharkquinone, a new ana-quinonoid tetracene derivative from marine-derived Streptomyces sp. EGY1 with TRAIL resistance-overcoming activity. J. Nat. Med. 2017, 71, 564–569. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.H.; Lu, M.C.; Chung, H.M.; Weng, C.F.; Su, J.H.; Yang, Y.T.; Su, Y.D.; Chang, Y.C.; Kuo, J.; Wu, Y.C.; et al. Bafilomycin M, a new cytotoxic bafilomycin produced by a Streptomyces sp. isolated from a marine sponge Theonella sp. Tetrahedron Lett. 2016, 57, 4863–4865. [Google Scholar] [CrossRef]
- Chen, H.; Cai, K.; Yao, R. A new macrolactam derivative from the marine actinomycete HF-11225. J. Antibiot. 2018, 71, 477–479. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.J.; Li, J.Q.; Zhang, H.J.; Ding, W.J.; Ma, Z.J. Cyclizidine-Type Alkaloids from Streptomyces sp. HNA39. J. Nat. Prod. 2018, 81, 394–399. [Google Scholar] [CrossRef] [PubMed]
- Lu, C.; Li, J.M.; Qi, H.; Zhang, H.; Zhang, J.; Xiang, W.S.; Wang, J.D.; Wang, X.J. Two new lankacidin-related metabolites from Streptomyces sp. HS-NF-1178. J. Antibiot. 2018, 71, 397–401. [Google Scholar] [CrossRef]
- Zhao, X.L.; Wang, H.; Xue, Z.L.; Li, J.S.; Qi, H.; Zhang, H.; Zhao, T.; Wang, J.D.; Xiang, W.S. Two new glutarimide antibiotics from Streptomyces sp. HS-NF-780. J. Antibiot. 2019, 72, 241–245. [Google Scholar] [CrossRef]
- Yixizhuoma; Ishikawa, N.; Abdelfattah, M.S.; Ishibashi, M. Elmenols C-H, new angucycline derivatives isolated from a culture of Streptomyces sp. IFM 11490. J. Antibiot. 2017, 70, 601–606. [Google Scholar] [CrossRef][Green Version]
- Son, S.; Jang, M.; Lee, B.; Hong, Y.S.; Ko, S.K.; Jang, J.H.; Ahn, J.S. Ulleungdin, a Lasso Peptide with Cancer Cell Migration Inhibitory Activity Discovered by the Genome Mining Approach. J. Nat. Prod. 2018, 81, 2205–2211. [Google Scholar] [CrossRef]
- Son, S.; Ko, S.K.; Jang, M.; Lee, J.K.; Kwon, M.C.; Kang, D.H.; Ryoo, I.J.; Lee, J.S.; Hong, Y.S.; Kim, B.Y.; et al. Polyketides and Anthranilic Acid Possessing 6-Deoxy-α-l-talopyranose from a Streptomyces Species. J. Nat. Prod. 2017, 80, 1378–1386. [Google Scholar] [CrossRef] [PubMed]
- Yang, F.X.; Hou, G.X.; Luo, J.; Yang, J.; Yan, Y.; Huang, S.X. New phenoxazinone-related alkaloids from strain Streptomyces sp. KIB-H1318. J. Antibiot. 2018, 71, 1040–1043. [Google Scholar] [CrossRef] [PubMed]
- Dong, M.; Cao, P.; Ma, Y.T.; Luo, J.; Yan, Y.; Li, R.T.; Huang, S.X. A new actinomycin Z analogue with an additional oxygen bridge between chromophore and β-depsipentapeptide from Streptomyces sp. KIB-H714. Nat. Prod. Res. 2019, 33, 219–225. [Google Scholar] [CrossRef] [PubMed]
- Guo, L.; Xie, Z.P.; Yang, Q.; Feng, L.L.; Zhang, L.; Zhang, Y.Z.; Li, X.N.; Pescitelli, G.; Zhang, S.M. Kiamycins B and C, unusual bridged angucyclinones from a marine sediment-derived Streptomyces sp. Tetrahedron Lett. 2018, 59, 2176–2180. [Google Scholar] [CrossRef]
- Kawahara, T.; Izumikawa, M.; Kozone, I.; Hashimoto, J.; Kagaya, N.; Koiwai, H.; Komatsu, M.; Fujie, M.; Sato, N.; Ikeda, H.; et al. Neothioviridamide, a Polythioamide Compound Produced by Heterologous Expression of a Streptomyces sp. Cryptic RiPP Biosynthetic Gene Cluster. J. Nat. Prod. 2018, 81, 264–269. [Google Scholar] [CrossRef]
- Lu, D.D.; Ren, J.W.; Du, Q.Q.; Song, Y.J.; Lin, S.Q.; Li, X.; Li, E.W.; Xie, W.D. p-Terphenyls and actinomycins from a Streptomyces sp. associated with the larva of mud dauber wasp. Nat. Prod. Res. 2021, 35, 1869–1873. [Google Scholar] [CrossRef]
- Song, Y.J.; Zheng, H.B.; Peng, A.H.; Ma, J.H.; Lu, D.D.; Li, X.; Zhang, H.Y.; Xie, W.D. Strepantibins A-C: Hexokinase II Inhibitors from a Mud Dauber Wasp Associated Streptomyces sp. J. Nat. Prod. 2019, 82, 1114–1119. [Google Scholar] [CrossRef]
- Cheng, P.; Xu, K.; Chen, Y.C.; Wang, T.T.; Chen, Y.; Yang, C.L.; Ma, S.Y.; Liang, Y.; Ge, H.M.; Jiao, R.H. Cytotoxic aromatic polyketides from an insect derived Streptomyces sp. NA4286. Tetrahedron Lett. 2019, 60, 1706–1709. [Google Scholar] [CrossRef]
- Lu, C.; Zhao, Y.; Jia, W.-Q.; Zhang, H.; Qi, H.; Xiang, W.-S.; Wang, J.-D.; Wang, X.-J. A new anthracycline-type metabolite from Streptomyces sp. NEAU-L3. J. Antibiot. 2017, 70, 1026–1028. [Google Scholar] [CrossRef]
- Kawahara, T.; Fujiwara, T.; Kagaya, N.; Shin-Ya, K. JBIR-150, a novel 20-membered polyene macrolactam from marine-derived Streptomyces sp. OPMA00071. J. Antibiot. 2018, 71, 390–392. [Google Scholar] [CrossRef]
- Abbas, M.; Elshahawi, S.I.; Wang, X.; Ponomareva, L.V.; Sajid, I.; Shaaban, K.A.; Thorson, J.S. Puromycins B-E, Naturally Occurring Amino-Nucleosides Produced by the Himalayan Isolate Streptomyces sp. PU-14G. J. Nat. Prod. 2018, 81, 2560–2566. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Chai, W.; Wang, W.; Song, T.; Lian, X.Y.; Zhang, Z. Cytotoxic Bagremycins from Mangrove-Derived Streptomyces sp. Q22. J. Nat. Prod. 2017, 80, 1450–1456. [Google Scholar] [CrossRef] [PubMed]
- Hayakawa, Y.; Akimoto, M.; Ishikawa, A.; Izawa, M.; Shin-Ya, K. Curromycin A as a GRP78 downregulator and a new cyclic dipeptide from Streptomyces sp. J. Antibiot. 2016, 69, 187–188. [Google Scholar] [CrossRef] [PubMed]
- Nogawa, T.; Okano, A.; Lim, C.L.; Futamura, Y.; Shimizu, T.; Takahashi, S.; Ibrahim, D.; Osada, H. Opantimycin A, a new metabolite isolated from Streptomyces sp. RK88-1355. J. Antibiot. 2017, 70, 222–225. [Google Scholar] [CrossRef]
- Cheng, C.; Othman, E.M.; Fekete, A.; Krischke, M.; Stopper, H.; Edrada-Ebel, R.A.; Mueller, M.J.; Hentschel, U.; Abdelmohsen, U.R. Strepoxazine A, a new cytotoxic phenoxazin from the marine sponge-derived bacterium Streptomyces sp. SBT345. Tetrahedron Lett. 2016, 57, 4196–4199. [Google Scholar] [CrossRef]
- Liu, Z.; Ma, L.; Zhang, L.; Zhang, W.; Zhu, Y.; Chen, Y.; Zhang, W.; Zhang, C. Functional characterization of the halogenase SpmH and discovery of new deschloro-tryptophan dimers. Org. Biomol. Chem. 2019, 17, 1053–1057. [Google Scholar] [CrossRef] [PubMed]
- Peng, J.; Zhang, Q.; Jiang, X.; Ma, L.; Long, T.; Cheng, Z.; Zhang, C.; Zhu, Y. New piericidin derivatives from the marine-derived Streptomyces sp. SCSIO 40063 with cytotoxic activity. Nat. Prod. Res. 2022, 36, 2458–2464. [Google Scholar] [CrossRef]
- Jiang, Y.J.; Gan, L.S.; Ding, W.J.; Chen, Z.; Ma, Z.J. Cytotoxic gephyromycins from the Streptomyces sp. SS13I. Tetrahedron Lett. 2017, 58, 3747–3750. [Google Scholar] [CrossRef]
- Zhang, D.; Jiang, Y.; Li, J.; Ding, W.; Chen, Z.; Ma, Z. Thioquinomycins A-D, novel naphthothiophenediones from the marine-derived Streptomyces sp. SS17F. Tetrahedron 2018, 74, 6150–6154. [Google Scholar] [CrossRef]
- Xu, C.D.; Zhang, H.J.; Ma, Z.J. Pyrimidine Nucleosides from Streptomyces sp. SSA28. J. Nat. Prod. 2019, 82, 2509–2516. [Google Scholar] [CrossRef]
- Lee, B.; Lee, G.-E.; Hwang, G.J.; Heo, K.T.; Lee, J.K.; Jang, J.-P.; Hwang, B.Y.; Jang, J.-H.; Cho, Y.-Y.; Hong, Y.-S. Rubiflavin G, photorubiflavin G, and photorubiflavin E: Novel pluramycin derivatives from Streptomyces sp. W2061 and their anticancer activity against breast cancer cells. J. Antibiot. 2023, 76, 1–7. [Google Scholar] [CrossRef]
- Xu, Y.; Hu, B.Y.; Qin, Y.; Zhuang, L.; Yang, Y.B.; Zhao, L.X. Jiangchuanmycin, a New Pyrrolizidine Analog from Streptomyces sp. YIM S01863. Chem. Biodivers. 2023, 20, e202201240. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.; Wang, Y.; Yang, Y.; Chen, H. Shellmycin A-D, Novel Bioactive Tetrahydroanthra-γ-Pyrone Antibiotics from Marine Streptomyces sp. Shell-016. Mar. Drugs 2020, 18, 58. [Google Scholar] [CrossRef] [PubMed]
- El Sayed, K.A. Natural Products as Antiviral Agents. Stud. Nat. Prod. Chem. 2000, 24, 473–572. [Google Scholar] [CrossRef]
- Patridge, E.; Gareiss, P.; Kinch, M.S.; Hoyer, D. An analysis of FDA-approved drugs: Natural products and their derivatives. Drug Discov. Today 2016, 21, 204–207. [Google Scholar] [CrossRef]
- Farmer, P.B.; Suhadolnik, R.J. Nucleoside antibiotics. Biosynthesis of arabinofuranosyladenine by Streptomyces antibioticus. Biochemistry 1972, 11, 911–916. [Google Scholar] [CrossRef]
- Sayed, A.M.; Alhadrami, H.A.; El-Gendy, A.O.; Shamikh, Y.I.; Belbahri, L.; Hassan, H.M.; Abdelmohsen, U.R.; Rateb, M.E. Microbial Natural Products as Potential Inhibitors of SARS-CoV-2 Main Protease (Mpro). Microorganisms 2020, 8, 970. [Google Scholar] [CrossRef]
- Raveh, A.; Delekta, P.C.; Dobry, C.J.; Peng, W.; Schultz, P.J.; Blakely, P.K.; Tai, A.W.; Matainaho, T.; Irani, D.N.; Sherman, D.H.; et al. Discovery of potent broad spectrum antivirals derived from marine actinobacteria. PLoS ONE 2013, 8, e82318. [Google Scholar] [CrossRef]
- Jakubiec-Krzesniak, K.; Rajnisz-Mateusiak, A.; Guspiel, A.; Ziemska, J.; Solecka, J. Secondary Metabolites of Actinomycetes and their Antibacterial, Antifungal and Antiviral Properties. Pol. J. Microbiol. 2018, 67, 259–272. [Google Scholar] [CrossRef]
- Rawal, K.R.; Bariwal, J.; Singh, V. Chemistry and Bioactivities of Aristeromycins: An Overview. Curr. Top. Med. Chem. 2016, 16, 3258–3273. [Google Scholar] [CrossRef]
- Selim, M.S.M.; Abdelhamid, S.A.; Mohamed, S.S. Secondary metabolites and biodiversity of actinomycetes. J. Genet. Eng. Biotechnol. 2021, 19, 72. [Google Scholar] [CrossRef]
- Abdelmohsen, U.R.; Bayer, K.; Hentschel, U. Diversity, abundance and natural products of marine sponge-associated actinomycetes. Nat. Prod. Rep. 2014, 31, 381–399. [Google Scholar] [CrossRef] [PubMed]
- Caly, L.; Druce, J.D.; Catton, M.G.; Jans, D.A.; Wagstaff, K.M. The FDA-approved drug ivermectin inhibits the replication of SARS-CoV-2 in vitro. Antiviral Res. 2020, 178, 104787. [Google Scholar] [CrossRef] [PubMed]
- Euanorasetr, J.; Intra, B.; Thunmrongsiri, N.; Limthongkul, J.; Ubol, S.; Anuegoonpipat, A.; Kurosu, T.; Ikuta, K.; Nihira, T.; Panbangred, W. In vitro antiviral activity of spirotetronate compounds against dengue virus serotype 2. J. Gen. Appl. Microbiol. 2019, 65, 197–203. [Google Scholar] [CrossRef] [PubMed]
- Keller, L.; Oueis, E.; Kaur, A.; Safaei, N.; Kirsch, S.H.; Gunesch, A.P.; Haid, S.; Rand, U.; Čičin-Šain, L.; Fu, C. Persicamidines—Unprecedented Sesquarterpenoids with Potent Antiviral Bioactivity against Coronaviruses. Angew. Chem. Int. Ed. 2023, 62, e202214595. [Google Scholar] [CrossRef]
- Shuai, H.; Myronovskyi, M.; Nadmid, S.; Luzhetskyy, A. Identification of a Biosynthetic Gene Cluster Responsible for the Production of a New Pyrrolopyrimidine Natural Product-Huimycin. Biomolecules 2020, 10, 1074. [Google Scholar] [CrossRef]
- Zhang, Y.; Fang, W.; Wang, K.; Zhang, Z.; Wu, Z.; Shi, L.; Liu, F.; Wan, Z.; Liu, M. Napyradiomycin A4 and Its Relate Compounds, a New Anti-PRV Agent and Their Antibacterial Activities, from Streptomyces kebangsaanensis WS-68302. Molecules 2023, 28, 640. [Google Scholar] [CrossRef]
- Liu, M.; Ren, M.; Zhang, Y.; Wan, Z.; Wang, Y.; Wu, Z.; Wang, K.; Fang, W.; Yang, X. Antiviral activity of benzoheterocyclic compounds from soil-derived Streptomyces jiujiangensis NBERC-24992. Molecules 2023, 28, 878. [Google Scholar] [CrossRef]
- Hao, X.; Li, S.; Wang, G.; Li, J.; Peng, Z.; Zhang, Y.; Yu, L.; Gan, M. Zelkovamycins F and G, Cyclopeptides with Cα-Methyl-threonine Residues, from an Endophytic Kitasatospora sp. J. Nat. Prod. 2022, 85, 1715–1722. [Google Scholar] [CrossRef]
- Kimura, T.; Suga, T.; Kameoka, M.; Ueno, M.; Inahashi, Y.; Matsuo, H.; Iwatsuki, M.; Shigemura, K.; Shiomi, K.; Takahashi, Y.; et al. New tetrahydroquinoline and indoline compounds containing a hydroxy cyclopentenone, virantmycin B and C, produced by Streptomyces sp. AM-2504. J. Antibiot. 2019, 72, 169–173. [Google Scholar] [CrossRef]
- Kim, S.H.; Ha, T.K.Q.; Oh, W.K.; Shin, J.; Oh, D.C. Antiviral Indolosesquiterpenoid Xiamycins C-E from a Halophilic Actinomycete. J. Nat. Prod. 2016, 79, 51–58. [Google Scholar] [CrossRef] [PubMed]
- Fang, Y.; Wang, J.; Tang, Y.; Guo, Z.; Bai, J.; Wu, L.; Su, J.; Cen, S.; Yu, L.; Zhang, D. Geninthiocins E and F, two new cyclic thiopeptides with antiviral activities from soil-derived Streptomyces sp. CPCC 200267 using OSMAC strategy. J. Antibiot. 2023, 76, 101–104. [Google Scholar] [CrossRef]
- Li, F.; Lu, S.; Xie, X.; Fan, S.; Chen, D.; Wu, S.; He, J. Antiviral properties of extracts of Streptomyces sp. SMU 03 isolated from the feces of Elephas maximus. Fitoterapia 2020, 143, 104600. [Google Scholar] [CrossRef] [PubMed]
- Saito, S.; Funayama, K.; Kato, W.; Okuda, M.; Kawamoto, M.; Matsubara, T.; Sato, T.; Sato, A.; Otsuguro, S.; Sasaki, M. Dihydromaniwamycin E, a Heat-Shock Metabolite from Thermotolerant Streptomyces sp. JA74, Exhibiting Antiviral Activity against Influenza and SARS-CoV-2 Viruses. J. Nat. Prod. 2022, 85, 2583–2591. [Google Scholar] [CrossRef] [PubMed]
- Igarashi, Y.; Yu, L.; Ikeda, M.; Oikawa, T.; Kitani, S.; Nihira, T.; Bayanmunkh, B.; Panbangred, W. Jomthonic Acid A, a Modified Amino Acid from a Soil-Derived Streptomyces. J. Nat. Prod. 2012, 75, 986–990. [Google Scholar] [CrossRef] [PubMed]
- Yu, L.; Oikawa, T.; Kitani, S.; Nihira, T.; Bayanmunkh, B.; Panbangred, W.; Igarashi, Y. Jomthonic acids B and C, two new modified amino acids from Streptomyces sp. J. Antibiot. 2014, 67, 345–347. [Google Scholar] [CrossRef]
- Inahashi, Y.; Iwatsuki, M.; Ishiyama, A.; Matsumoto, A.; Hirose, T.; Oshita, J.; Sunazuka, T.; Panbangred, W.; Takahashi, Y.; Kaiser, M.; et al. Actinoallolides A–E, New Anti-trypanosomal Macrolides, Produced by an Endophytic Actinomycete, Actinoallomurus fulvus MK10-036. Org. Lett. 2015, 17, 864–867. [Google Scholar] [CrossRef]
- Akiyama, H.; Oku, N.; Harunari, E.; Panbangred, W.; Igarashi, Y. Complete NMR assignment and absolute configuration of k4610422, a norditerpenoid inhibitor of testosterone-5α-reductase originally from Streptosporangium: Rediscovery from a thermophilic Actinomadura. J. Antibiot. 2020, 73, 60–65. [Google Scholar] [CrossRef]
- Lyddiard, D.; Jones, G.L.; Greatrex, B.W. Keeping it simple: Lessons from the golden era of antibiotic discovery. FEMS Microbiol. Lett. 2016, 363, 84. [Google Scholar] [CrossRef]
- Majumder, M.A.A.; Rahman, S.; Cohall, D.; Bharatha, A.; Singh, K.; Haque, M.; Gittens-St Hilaire, M. Antimicrobial Stewardship: Fighting Antimicrobial Resistance and Protecting Global Public Health. Infect. Drug Resist. 2020, 13, 4713. [Google Scholar] [CrossRef]
- Baltz, R.H. Renaissance in antibacterial discovery from actinomycetes. Curr. Opin. Pharm. 2008, 8, 557–563. [Google Scholar] [CrossRef] [PubMed]
- Lazzarini, A.; Cavaletti, L.; Toppo, G.; Marinelli, F. Rare genera of actinomycetes as potential producers of new antibiotics. Antonie Van Leeuwenhoek 2000, 78, 399–405. [Google Scholar] [CrossRef] [PubMed]
- Kohli, I.; Joshi, N.C.; Mohapatra, S.; Varma, A. Extremophile—An Adaptive Strategy for Extreme Conditions and Applications. Curr. Genom. 2020, 21, 96–110. [Google Scholar] [CrossRef] [PubMed]
- Subramani, R.; Aalbersberg, W. Culturable rare Actinomycetes: Diversity, isolation and marine natural product discovery. Appl. Microbiol. Biotechnol. 2013, 97, 9291–9321. [Google Scholar] [CrossRef]
- Mahajan, G.B.; Balachandran, L. Sources of antibiotics: Hot springs. Biochem. Pharmacol. 2017, 134, 35–41. [Google Scholar] [CrossRef]
- Thawai, C.; Thamsathit, W.; Kudo, T. Planosporangium thailandense sp. nov., isolated from soil from a Thai hot spring. Int. J. Syst. Evol. Microbiol. 2013, 63, 1051–1055. [Google Scholar] [CrossRef]
- Bull, A.T.; Idris, H.; Sanderson, R.; Asenjo, J.; Andrews, B.; Goodfellow, M. High altitude, hyper-arid soils of the Central-Andes harbor mega-diverse communities of actinobacteria. Extremophiles 2017, 22, 47–57. [Google Scholar] [CrossRef]
- Sun, Y.; Shi, Y.-L.; Wang, H.; Zhang, T.; Yu, L.-Y.; Sun, H.; Zhang, Y.-Q. Diversity of Bacteria and the Characteristics of Actinobacteria Community Structure in Badain Jaran Desert and Tengger Desert of China. Front. Microbiol. 2018, 9, 1068. [Google Scholar] [CrossRef]
- Li, W.J.; Zhang, Y.G.; Zhang, Y.Q.; Tang, S.K.; Xu, P.; Xu, L.H.; Jiang, C.L. Streptomyces sodiiphilus sp. nov., a novel alkaliphilic actinomycete. Int. J. Syst. Evol. Microbiol. 2005, 55, 1329–1333. [Google Scholar] [CrossRef]
- Hui, M.L.-Y.; Tan, L.T.H.; Letchumanan, V.; He, Y.-W.; Fang, C.-M.; Chan, K.-G.; Law, J.W.F.; Lee, L.H. The Extremophilic Actinobacteria: From Microbes to Medicine. Antibiotics 2021, 10, 682. [Google Scholar] [CrossRef]
- Singh, R.; Dubey, A.K. Diversity and Applications of Endophytic Actinobacteria of Plants in Special and Other Ecological Niches. Front. Microbiol. 2018, 9, 1767. [Google Scholar] [CrossRef] [PubMed]
- Bentley, S.D.; Chater, K.F.; Cerdeno-Tarraga, A.M.; Chalis, G.L.; Thomson, N.R.; James, K.D.; Harris, D.E.; Quail, M.A.; Kieser, H.; Harper, D.; et al. Complete genome sequence of the model actinomycete Streptomyces coelicolor A3(2). Nature 2002, 417, 141–147. [Google Scholar] [CrossRef] [PubMed]
- Hoskisson, P.A.; van Wezel, G.P. Streptomyces coelicolor. Trends Microbiol. 2019, 27, 468–469. [Google Scholar] [CrossRef] [PubMed]
- Blin, K.; Shaw, S.; Kloosterman, A.M.; Charlop-Powers, Z.; Van Wezel, G.P.; Medema, M.H.; Weber, T. antiSMASH 6.0: Improving cluster detection and comparison capabilities. Nucleic Acids Res. 2021, 49, W29–W35. [Google Scholar] [CrossRef]
- Weber, T.; Rausch, C.; Lopez, P.; Hoof, I.; Gaykova, V.; Huson, D.H.; Wohlleben, W. CLUSEAN: A computer-based framework for the automated analysis of bacterial secondary metabolite biosynthetic gene clusters. J. Biotechnol. 2009, 140, 13–17. [Google Scholar] [CrossRef]
- Skinnider, M.A.; Johnston, C.W.; Gunabalasingam, M.; Merwin, N.J.; Kieliszek, A.M.; MacLellan, R.J.; Li, H.; Ranieri, M.R.M.; Webster, A.L.H.; Cao, M.P.T.; et al. Comprehensive prediction of secondary metabolite structure and biological activity from microbial genome sequences. Nat. Comm. 2020, 11, 6058. [Google Scholar] [CrossRef]
- Li, M.H.T.; Ung, P.M.U.; Zajkowski, J.; Garneau-Tsodikova, S.; Sherman, D.H. Automated genome mining for natural products. BMC Bioinform. 2009, 10, 185. [Google Scholar] [CrossRef] [PubMed]
- Starcevic, A.; Zucko, J.; Simunkovic, J.; Long, P.F.; Cullum, J.; Hranueli, D. ClustScan: An integrated program package for the semi-automatic annotation of modular biosynthetic gene clusters and in silico prediction of novel chemical structures. Nucleic Acids Res. 2008, 36, 6882–6892. [Google Scholar] [CrossRef]
- Li, L.; Jiang, W.; Lu, Y. New strategies and approaches for engineering biosynthetic gene clusters of microbial natural products. Biotechnol. Adv. 2017, 35, 936–949. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Zhao, Y.; Huang, C.; Luo, Y. Recent Advances in Silent Gene Cluster Activation in Streptomyces. Front. Bioeng. Biotechnol. 2021, 9, 88. [Google Scholar] [CrossRef]
- Chou, W.K.W.; Fanizza, I.; Uchiyama, T.; Komatsu, M.; Ikeda, H.; Cane, D.E. Genome mining in Streptomyces avermitilis: Cloning and characterization of SAV_76, the synthase for a new sesquiterpene, avermitilol. J. Am. Chem. Soc. 2010, 132, 8850. [Google Scholar] [CrossRef] [PubMed]
- Pan, R.; Bai, X.; Chen, J.; Zhang, H.; Wang, H. Exploring structural diversity of microbe secondary metabolites using OSMAC strategy: A literature review. Front. Microbiol. 2019, 10, 294. [Google Scholar] [CrossRef]
- Romano, S.; Jackson, S.A.; Patry, S.; Dobson, A.D.W. Extending the “One Strain Many Compounds” (OSMAC) Principle to Marine Microorganisms. Mar. Drugs 2018, 16, 244. [Google Scholar] [CrossRef] [PubMed]
- Asamizu, S.; Ozaki, T.; Teramoto, K.; Satoh, K.; Onaka, H. Killing of Mycolic Acid-Containing Bacteria Aborted Induction of Antibiotic Production by Streptomyces in Combined-Culture. PLoS ONE 2015, 10, e0142372. [Google Scholar] [CrossRef] [PubMed]
- Xia, H.; Li, X.; Li, Z.; Zhan, X.; Mao, X.; Li, Y. The Application of Regulatory Cascades in Streptomyces: Yield Enhancement and Metabolite Mining. Front. Microbiol. 2020, 11, 406. [Google Scholar] [CrossRef] [PubMed]
- Tawfike, A.; Attia, E.Z.; Desoukey, S.Y.; Hajjar, D.; Makki, A.A.; Schupp, P.J.; Edrada-Ebel, R.A.; Abdelmohsen, U.R. New bioactive metabolites from the elicited marine sponge-derived bacterium Actinokineospora spheciospongiae sp. nov. AMB Express 2019, 9, 12. [Google Scholar] [CrossRef] [PubMed]
- Yu, M.; Li, Y.; Banakar, S.P.; Liu, L.; Shao, C.; Li, Z.; Wang, C. New metabolites from the Co-culture of marine-derived actinomycete Streptomyces rochei MB037 and fungus Rhinocladiella similis 35. Front. Microbiol. 2019, 10, 915. [Google Scholar] [CrossRef]
- Tyurin, A.P.; Alferova, V.A.; Korshun, V.A. Chemical Elicitors of Antibiotic Biosynthesis in Actinomycetes. Microorganisms 2018, 6, 52. [Google Scholar] [CrossRef]
- Zong, G.; Fu, J.; Zhang, P.; Zhang, W.; Xu, Y.; Cao, G.; Zhang, R. Use of elicitors to enhance or activate the antibiotic production in Streptomyces. Crit. Rev. Biotechnol. 2022, 42, 1260–1283. [Google Scholar] [CrossRef]
- Choi, S.U.; Lee, C.K.; Hwang, Y.I.; Kinosita, H.; Nihira, T. γ-Butyrolactone autoregulators and receptor proteins in non-Streptomyces actinomycetes producing commercially important secondary metabolites. Arch. Microbiol. 2003, 180, 303–307. [Google Scholar] [CrossRef]
- Onaka, H. Novel antibiotic screening methods to awaken silent or cryptic secondary metabolic pathways in actinomycetes. J. Ant. 2017, 70, 865–870. [Google Scholar] [CrossRef] [PubMed]
- Dashti, Y.; Grkovic, T.; Abdelmohsen, U.R.; Hentschel, U.; Quinn, R.J. Actinomycete Metabolome Induction/Suppression with N-Acetylglucosamine. J. Nat. Prod. 2017, 80, 828–836. [Google Scholar] [CrossRef] [PubMed]
- Mast, Y.; Stegmann, E. Actinomycetes: The Antibiotics Producers. Antibiotics 2019, 8, 105. [Google Scholar] [CrossRef]
- Jose, P.A.; Jha, B. New Dimensions of Research on Actinomycetes: Quest for Next Generation Antibiotics. Front. Microbiol. 2016, 7, 1295. [Google Scholar] [CrossRef] [PubMed]
- Gadelhak, G.G.; El-Tarabily, K.A.; Al-Kaabi, F.K. Insect Control Using Chitinolytic Soil Actinomycetes as Bio control Agents. Int. J. Agri. Biol. 2005, 7, 627–633. [Google Scholar]
- Lam, K.S. Discovery of novel metabolites from marine actinomycetes. Curr. Opin. Microbiol. 2006, 9, 245–251. [Google Scholar] [CrossRef]
- Vijayabharathi, R.; Kumari, B.R.; Sathya, A.; Srinivas, V.; Abhishek, R.; Sharma, H.; Gopalakrishnan, S. Biological activity of entomopathogenic actinomycetes against lepidopteran insects (Noctuidae: Lepidoptera). Can. J. Plant Sci. 2014, 94, 759–769. [Google Scholar] [CrossRef]
- Choi, S.-S.; Kim, H.-J.; Lee, H.-S.; Kim, P.; Kim, E.-S. Genome mining of rare actinomycetes and cryptic pathway awakening. Process. Biochem. 2015, 50, 1184–1193. [Google Scholar] [CrossRef]
- Peng, Q.; Gao, G.; Lü, J.; Long, Q.; Chen, X.; Zhang, F.; Xu, M.; Liu, K.; Wang, Y.; Deng, Z.; et al. Engineered Streptomyces lividans Strains for Optimal Identification and Expression of Cryptic Biosynthetic Gene Clusters. Front. Microbiol. 2018, 9, 3042. [Google Scholar] [CrossRef]
- Lee, N.; Hwang, S.; Kim, J.; Cho, S.; Palsson, B.; Cho, B.K. Mini review: Genome mining approaches for the identification of secondary metabolite biosynthetic gene clusters in Streptomyces. Comput. Struct. Biotechnol. J. 2020, 18, 1548–1556. [Google Scholar] [CrossRef]
- Gomez-Escribano, J.P.; Bibb, M.J. Engineering Streptomyces coelicolor for heterologous expression of secondary metabolite gene clusters. Microb. Biotechnol. 2011, 4, 207–215. [Google Scholar] [CrossRef] [PubMed]
- Mao, D.; Okada, B.K.; Wu, Y.; Xu, F.; Seyedsayamdost, M.R. Recent advances in activating silent biosynthetic gene clusters in bacteria. Curr. Opin. Microbiol. 2018, 45, 156–163. [Google Scholar] [CrossRef] [PubMed]
- Salcedo, R.G.; Olano, C.; Gómez, C.; Fernández, R.; Braña, A.F.; Méndez, C.; Calle, F.; Salas, J.A. Characterization and engineering of the biosynthesis gene cluster for antitumor macrolides PM100117 and PM100118 from a marine actinobacteria: Generation of a novel improved derivative. Microb. Cell Factories 2016, 15, 44. [Google Scholar] [CrossRef] [PubMed]
- Hoshino, S.; Onaka, H.; Abe, I. Activation of silent biosynthetic pathways and discovery of novel secondary metabolites in actinomycetes by co-culture with mycolic acid-containing bacteria. J. Ind. Microbiol. Biotechnol. 2018, 46, 363–374. [Google Scholar] [CrossRef] [PubMed]
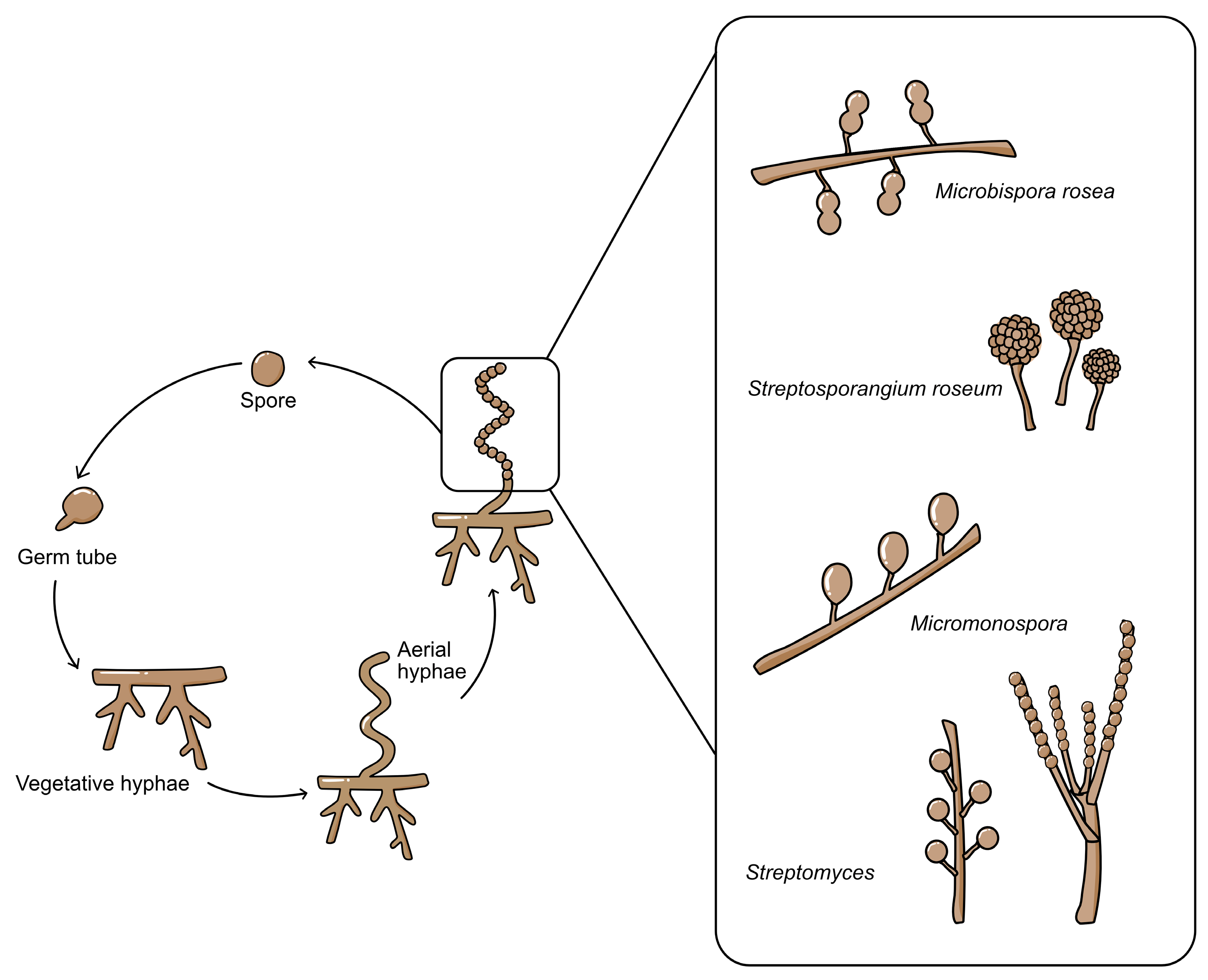
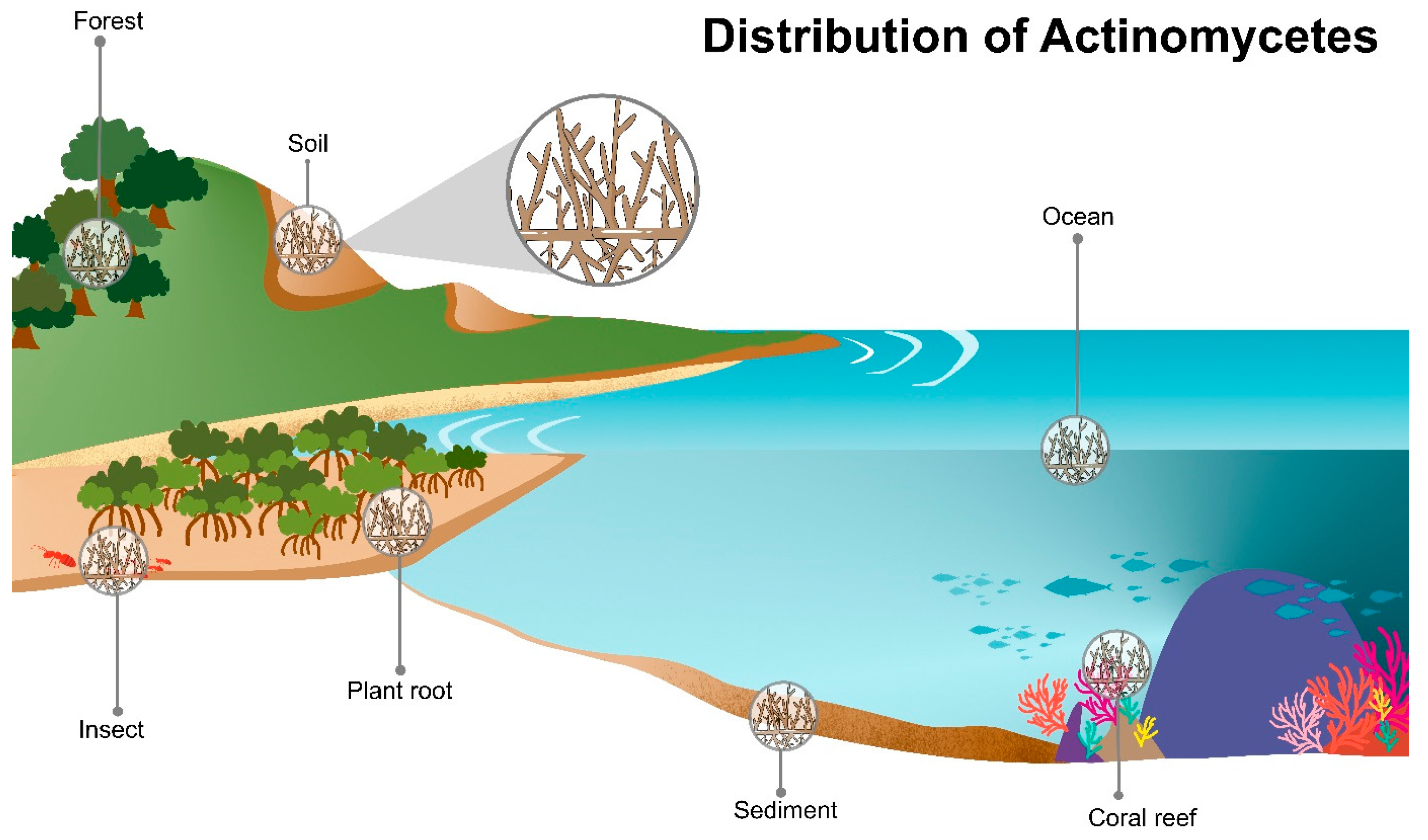


| Organism (s) | Compound Name (s) | Reference (s) |
|---|---|---|
| Actinoallomurus sp. ID145113, 145206, 145754 | Paramagnetoquinones A–C | [66] |
| Actinomadura atramentaria NBRC 14695 | Cinnamycin B | [67] |
| Actinomadura sp. KC191 | Actinomadurol | [68] |
| Amycolatopsis sp. IRD-009 | Pradimicin-IRD | [69] |
| Amycolatopsis sp. M39 | Macrotermycins A & C | [70] |
| Amycolatopsis sp. MCC0218 | Enceleamycins A–C | [71] |
| Amycolatopsis sp. ML1-hF4 | Pargamicins B–D | [72] |
| Kibdelosporangium phytohabitans XY-R10 | Maipomycin A (MaiA) | [73] |
| Kitasatospora sp. MG372-hF19 | Biospolides C-E | [74] |
| Kocuria marina CMG S2 | Kocumarin | [75] |
| Micromonospora carbonacea LS276 | Tetrocarcin Q | [76,77] |
| Micromonospora harpali SCSIO GJ089 | Icrosporanate A, Tetrocarcin P, Microsporanates B–F | [78] |
| Micromonospora chalcea FIM 02 | Rakicidins G–I | [79] |
| Micromonospora sp. UR56 + Actinokineospora sp. EG49 | Phenazine-1-carboxylic acid, Aestivophoenin C, Methyl saphenate | [80] |
| Micromonospora sp. WMMB-235 + Rhodococcus sp. WMMA-185 | Keyicin | [81] |
| Micromonospora sp. 5-297 | Tetrocarcins N | [82] |
| Micromonospora sp. CA-214671 | Phocoenamicin B | [83] |
| Micromonospora sp. RJA4480 | Sporalactam B | [84] |
| Micromonospora sp. SCSIO 07395 | Microechmycin A | [85] |
| Micromonospora sp. TP-A0468 | 16-demethylrifamycin S | [86] |
| Micromonospora endolithica | loseolamycins A1, A2 | [87] |
| Micromonospora yangpuensis DSM 45577 | Yangpumicins F, G | [88] |
| Nocardiopsis sp. LX-1 | Nocarpyrroline A, Isoflavonoid E | [89] |
| Nocardiopsis sp. SCA30 | 1-acetyl-4-4(hydroxyphenyl)piperazine | [90] |
| Pseudonocardia carboxydivorans M227 | Branimycins B, C | [91] |
| Streptomonospora sp. PA3 | Persiamycin A | [92] |
| Streptomyces aculeolatus PTM-029 | Napyradiomycins (3,7,9,10,12) | [93,94] |
| Streptomyces aculeolatus PTM-420 | Napyradiomycins (1,2,4–6,8,11) | [93] |
| Streptomyces albus 4N24 | Benzanthric acid | [95] |
| Streptomyces albus MAB56 | 12-methyltetradecanoic acid, Palmitic acid, Tridecanoic acid | [96] |
| Streptomyces armeniacus DSM 43125 | Armeniaspirol analogues 1–6, 9–12 | [94] |
| Streptomyces bacillaris MBTC38 | Lactoquinomycin A | [97] |
| Streptomyces californicus ADR1 | Methanoazulen-9-ol, decahydro-2, 2, 4, 8-tetramethyl-stereoisomer (Sesquiterpene) | [98] |
| Streptomyces coelicolor LY001 | 3-(3,5-dichloro-4-hydroxyphenyl) propanoic acid, 3-(3,5-dichloro-4-hydroxyphenyl) propanoic acid methyl ester, 3-(3-chloro-4-hydroxyphenyl) propanoic acid | [99] |
| Streptomyces diacarni LHW51701 | Chlocarbazomycins (CCBs) A–D | [100] |
| Streptomyces fradiae MM456M-mF7 | Fradiamines B | [101] |
| Streptomyces globisporus sp. WA5-2-37 | Actinomycin X2, Collismycin A | [102] |
| Streptomyces globisporus subsp. globisporus | Globimycin | [103] |
| Streptomyces griseoviridis PU-KB10–4 | Mitomycin C | [104] |
| Streptomyces hyaluromycini MB-PO13 | Rubromycins CA1, CA2 | [105] |
| Streptomyces koyangensis SCSIO 5802 | Neoabyssomicins F, G | [106] |
| Streptomyces malachitospinus ITD-35 | 3-octanone, neopentyl, Isothiocyanate, 2-methyl butyl isothiocyanate | [13] |
| Streptomyces microflavus (MBTI36) | Chromomycin A9, Ap, A2, A3 | [107] |
| Streptomyces lunaelactis MM109T | Lunaemycins A, B1, D | [108] |
| Streptomyces lusitanus OUCT16-27 | Grincamycin L | [109] |
| Streptomyces palmae CMU-AB204T | Phenyl alkenoic acids C, D Phenyl alkenoic acids E, F | [110] |
| Streptomyces qinglanensis 172205 | 15R-17,18-dehydroxantholipin | [111] |
| Streptomyces zhaozhouensis 208DD-064 | Streptopyrroles B, C | [112] |
| Streptomyces sp. 120454 | Mayamycin B | [113] |
| Streptomyces sp. 182SMLY | N-acetyl-N-demethylmayamycin | [114] |
| Streptomyces sp. 7NS3 | Emycin A | [115] |
| Streptomyces sp. 8P21H-1 | Desulphurzing, Griseoviridin, Griseoviridin | [116] |
| Streptomyces sp. AD-3-6 | Nybomycins D | [117] |
| Streptomyces sp. ADI91-18, ADI95-16 | Linearmycins | [117] |
| Streptomyces sp. ADI97-07 | Clavams | [117] |
| Streptomyces sp. ADI98-12 | Griselimycin, Kirromycin | [117] |
| Streptomyces sp. B9173 | Flaviogeranin B1, B, Flaviogeranin D, Flaviogeranin C2 | [118] |
| Streptomyces sp. CA-271078 | Napyradiomycins 2, D1 | [119] |
| Streptomyces sp. CPCC 204980 | Cervinomycins B1–B4 | [120] |
| Streptomyces sp. DSM14386 | Svetamycins C, G | [121] |
| Streptomyces sp. FJS31-2 | Zunyimycins B, C | [122] |
| Streptomyces sp. GKU 220 | Rakicidin F | [123] |
| Stretomyces sp. HCCB11876 | Quinomycins I, J | [124] |
| Streptomyces sp. HK-2006-1 | Aldgamycin O | [125] |
| Streptomyces sp. HN-A124 | Cysrabelomycin | [126] |
| Streptomyces sp. Hu103 | Anulamycins A-D | [127] |
| Streptomyces sp. HZP-2216E | 23-O-butyrylbafilomycin D | [128] |
| Streptomyces sp. IB201691-2A | Baikalomycins A–C | [129] |
| Streptomyces sp. ICN19 | Ala-geninthiocin | [130] |
| Streptomyces sp. KCB13F003 | Ulleungmycin A, B | [131] |
| Streptomyces sp. KCB14A132 | Enamidonins B, C | [132] |
| Streptomyces sp. MM168-141F8 | Quadoctomycin | [133] |
| Streptomyces sp. MNP32 | Phenol, 3,5-bis(1,1-dimethylethyl)-1,1′-Biphenyl]-2,3′-diol, 3,4′,5,6′-tetrakis(1,1-dimethylethyl) | [134] |
| Streptomyces sp. MUSC 125 | Thiophene, 2-butyl-5-ethyl, 1-Heptyn-3-ol, 8-[N-Aziridylethylamino]-2-6,dimethyloctene-2, Pyrrolo[1,2-a]pyrazine-1,4-dion,hexahydro, and 2,4-Dihydroxy-6-propylbenzoic acid | [42] |
| Streptomyces sp. NA06554 | Borrelidin J | [135] |
| Streptomyces sp. NA07423 | Nagimycins A, B | [136] |
| Streptomyces sp. OPOK_MB_B11 | Azalomycin F4a 2-ethylpentyl ester | [137] |
| Streptomyces sp. PBR11 | 1-Tetradecanol phenol, 2,5-bis(1,1-dimethylethyl n-pentadecanol, 1-nonadecene, and pyrrolo[1,2-a]pyrazine-1,4-dione hexahydro-3-(phenylmethyl). | [138] |
| Streptomyces sp. PNM-9 | 2-methyl-N-(2′-phenylethyl)-butanamide 3-methyl-N-(2′-phenylethyl)-butanamide | [139] |
| Streptomyces sp. SBT345 | Ageloline A | [140] |
| Streptomyces sp. SCSIO 41399 | Isotirandamycin B | [141] |
| Streptomyces sp. sima1_6 | Echinomycin | [137] |
| Streptomyces sp. SN0280 | Streptoone A | [142] |
| Streptomyces sp. SM01 | Picolinamycin | [143] |
| Streptomyces sp. SS | Sansanmycin Q | [144] |
| Streptomyces sp. Stup16_B49.2 | Echinomycin | [137] |
| Streptomyces sp. Stup16_B146 | TPU-0037-C | [137] |
| Streptomyces sp. Stup18_J70 | Bafilomycin A1 derivative | [137] |
| Streptomyces sp. UICC B-92 | 4-O-glucosyl,1-carboxyl-phenazine | [145] |
| Streptomyces sp. W367A | W367A | [137] |
| Streptomyces sp. XMA39 | Strepoxepinmycins A–D | [146] |
| Streptomyces sp. YINM00001 | Peperodione, Peperophthalene | [147] |
| Streptomyces sp. ZZ1118 | Streptoindoles A–D | [148] |
| Streptomyces sp. ZZ741 | Streptoglutarimides A–J | [149] |
| Streptomycete clade MAR4 CNY-960, CNS-284 | Marinocyanins A–F | [150] |
| Streptosporangium sp. SANK 60,501 | Muraminomicins A, B, C, D, E1, E2, F | [151] |
| Thermoactinomyces vulgaris ISCAR 2354 | Thermoactinoamide A | [152] |
| Verrucosispora sp. FIM06-0036 | 2-ethylhexyl 1H-imidazole-4-carboxylate | [100] |
| Verrucosispora sp. SCSIO 07399 | Kendomycins B–D | [153] |
| Organism (s) | Compound Name (s) | Reference (s) |
|---|---|---|
| Actinokineospora bangkokensis 44EHWT | Thailandins A & B | [159] |
| Actinomadura sp. BCC 35,430 | Actinomadurone | [161] |
| Amycolatopsis sp. M39 | Macrotermycins A & D | [70] |
| Acrocarpospora punica 04107M | Acrocarposporins A, B, D, and E | [162] |
| Dermabacter vaginalis AD1-86 | Dermazolium A | [163] |
| Micromonospora sp. UR56 + Actinokineospora sp. EG49 | Phenazine-1-carboxylic acid, Aestivophoenin c, Methyl saphenate | [80] |
| Nocardia abscessus IFM 10029 | Nabscessins A, B | [164] |
| Nocardiopsis sp. LX-1 | Isoflavonoid E | [89] |
| Pseudonocardia autotrophica KCTC9441 | Polyene B1 | [165] |
| Saccharothrix yanglingensis Hhs.015 | 10-deoxyfungichromin (WH02) | [166] |
| Streptomyces albidoflavus STV1572a | 1-heneicosanol | [167] |
| Streptomyces albolongus YIM 101047 | 19-methoxybafilomycin C1 amide | [168] |
| Streptomyces antibioticus strain 200-09 | Kitamycin C | [169] |
| Streptomyces caniferus GUA-06-05-006A | PM100117, PM100118 | [170] |
| Streptomyces griseoviridis PU-KB10–4 | Mitomycin C | [104] |
| Streptomyces hyaluromycini MB-PO13 | Rubromycins CA1, CA2 | [105] |
| Streptomyces morookaense AM25 | Gloeosporiocide | [171] |
| Streptomyces sp. 1H-XA2 | Furamycins I, II | [172] |
| Streptomyces sp. ADI91-18 | Linearmycins | [117] |
| Streptomyces sp. ADI95-16 | Linearmycins | [117] |
| Streptomyces sp. ADI96-02 | Cycloheximide, Galbonolides | [117] |
| Streptomyces sp. ADI96-15 | Candicidin | [117] |
| Streptomyces sp. ADI97-07 | Galbonolides | [117] |
| Streptomyces sp. ADI98-12 | Candicidin | [117] |
| Streptomyces sp. BV410 | Staurosporine | [173] |
| Streptomyces sp. CB09030 | LOB A, B, H8 | [174] |
| Streptomyces sp. CT37 | Legonimide 1, 1H-indole-3-carbaldehyde | [175] |
| Streptomyces sp. FX13 | Oligomycin A | [176] |
| Streptomyces sp. HAAG3-15 | Azalomycin B | [177] |
| Streptomyces sp. KIB-H869 | Hygrolidin-type macrolide | [178] |
| Streptomyces sp. MUSC 125 | Thiophene, 2-butyl-5-ethyl, 1-Heptyn-3-ol, Pyrrolo[1,2-a] pyrazine-1,4-dion,hexahydro, 9,9-Dimethyl-3, 7-diazabicyclo[3.3.1]nonane, 2,4-Dihydroxy-6-propylbenzoic acid | [42] |
| Streptomyces sp. PBR11 | 1-tetradecanol, n-pentadecanol, 1-nonadecene, pyrrolo [1,2-a]pyrazine-1,4-dione hexahydro-3-(phenylmethyl) | [138] |
| Streptomyces sp. QHH-9511 | 6-deoxy-13-hydroxy-8,11-dione Dihydrogranaticin A Granaticin A, B | [179] |
| Streptomyces sp. SN0280 | Streptoone B | [142] |
| Streptomyces sp. SY1965 | Streptothiazomycin A, Streptodiketopiperazines A, B, [2-hydroxy-1-(hydroxymethyl)ethyl]-2-methoxybenzamide, salicylamide, 4-hydroxymethyl benzoate, Spoxazomicin C | [180] |
| Streptomyces sp. TR1341 | Filipin, Fungichromin, Actinomycin X2 | [181] |
| Streptomyces sp. TT3 | Actinorhodin | [182] |
| Streptomyces sp. YO15-A001 | YO-001A | [183] |
| Streptomyces albus CAI-21 | Organophosphate | [184] |
| Streptomyces coelicolor LY001 | diketopiperazine alkaloids cyclo (l-Phe-trans-4-OH-l-Pro), cyclo(l-Phe-cis-4-OH-d-Pro) | [99] |
| Streptomyces palmae CMU-AB204T | Phenyl alkenoic acids A,B, Anguinomycin A, Leptomycin A, Actinopyrone A | [110] |
| Streptomyces qinglanensis 172205 | 15R-17,18-dehydroxantholipin | [111] |
| Streptomycete clade MAR4 CNY-960 and CNS-284 | Marinocyanins A | [150] |
| Umezawaea sp. RD066910 + Tsukamurella pulmonis TP-B0596 | Umezawamides A | [185] |
| Organism (s) | Compound Name (s) | Reference (s) |
|---|---|---|
| Actinoalloteichus hymeniacidonis 179DD-027 | Dokdolipid B | [198] |
| Actinomadura sp. K13-0306 | Sagamilactam | [199] |
| Actinomadura sp. K4S16 | Nonthmicin, Ecteinamycin | [200] |
| Actinosynnema pretiosum HGF052::asm18 | Actinosynneptide A, B | [201] |
| Amycolatopsis sp. IRD-009 | Pradimicin-IRD | [69] |
| Amycolatopsis alba DSM 44,262 | Thioalbamide | [202] |
| Catenuloplanes sp. RD067331 + Tsukamurella pulmonis TP-B059 | Catenulobactin B | [203] |
| Micromonospora sp. UR56 + Actinokineospora sp. EG49 | Phenazine-1-carboxylic acid, Aestivophoenin c, Methyl saphenate | [80,204] |
| Micromonospora carbonacea LS276 | Tetrocarcin Q | [76] |
| Micromonospora zhangzhouensis HM134T | 7E,11E)-6-hydroxy-1-isopropyl-11-(methoxycarbonyl)-4-methylene-1,2,3,4,4a,5,6,9,10,12a-decahydrobenzo[10]annulene-7-carboxylic acid | [205] |
| Micromonospora matsumotoense M-412 | Paulomycin G | [206] |
| Micromonospora aurantiaca 110B | Isoflavonoid glycosides 1-3 | [207] |
| Micromonospora chalcea FIM 02-523 | Rakicidins G–I | [208] |
| Micromonospora sp. FIM05328 | FW05328-1 | [209] |
| Micromonospora sp. HS-HM-036 | Naphthalenepropanoic acid analog | [210] |
| Micromonospora yangpuensis DSM 45577 | Yangpumicin A | [211] |
| Micromonospora yangpuensis DSM 45577 | Yangpumicins F, G | [88] |
| Nocardiopsis alba KM6-1 | Isomethoxyneihumicin | [212] |
| Nocardiopsis sp. CG3 | Kenalactams A, E | [213] |
| Nonomuraea endophytica GW58/450 | Karamomycins B, C | [214] |
| Nonomuraea sp. AKA32 | Akazamicin | [215] |
| Saccharomonospora sp. UR22 + Dietzia sp. UR66 | Saccharomonosporine A | [216] |
| Streptomonospora sp. PA3 | Persiamycin A | [92] |
| Streptomyces albolongus YIM 101047 | 19-methoxybafilomycin C1 amide, 21-deoxybafilomycin A1 | [168] |
| Streptomyces ardesiacus 156VN-095 | Urdamycins W, X, Grincamycin U | [217] |
| Streptomyces bacillaris | 2,4-di-tert-butylphenol | [218] |
| Streptomycete clade MAR4 CNY-960 and CNS-284 | Marinocyanins A–F | [150] |
| Streptomyces qinglanensis 172205 | 15R-17,18-dehydroxantholipin, (3E,5E,7E)-3-methyldeca-3,5,7-triene-2,9-dione, Qinlactone A–B | [111] |
| Streptomyces cacaoi subsp. asoensis H2S5 | Trienomycins J–L | [219] |
| Streptomyces caniferus GUA-06-05-006A | PM100117, PM100118 | [220] |
| Streptomyces coelicolor M1146 | Staurosporines M1, M2 | [221] |
| Streptomyces curacoi NBRC 12761 | Curacozole | [222] |
| Streptomyces cyaneofuscatus M-157 | 3-Hydroxyquinaldic acid derivative 1 | [223] |
| Streptomyces griseoviridis 2464-S5 | Prodigiosin R2 | [224] |
| Streptomyces seoulensis A01 | Ansaseomycins A, B | [225] |
| Streptomyces subflavus subsp. irumaensis AM-3603 | Bisoxazolomycin A | [226] |
| Streptomyces violascens YIM 100225 | Albaflavenoid | [227] |
| Streptomyces violascens YIM100212 | Fusicomycin A, B, Isofusicomycin A | [228] |
| Streptomyces sp. 166 | Sekgranaticin | [229] |
| Streptomyces sp. 182SMLY | N-acetyl-N-demethylmayamycin, Streptoanthraquinone A | [230] |
| Streptomyces sp. 1H-GS5 | Spectinabilin derivative 1 | [231] |
| Streptomyces sp. A65 | Streptocarbazoles C, 2040-epi-K252d, 0-epi-K252d | [232] |
| Streptomyces sp. AD-3 | Nybomycins D | [233] |
| Streptomyces sp. ADI92-24 | Tomaymycin | [117] |
| Streptomyces sp. ADI95-17 | Naringenin | [117] |
| Streptomyces sp. ADI96-02 | Echinomycin | [117] |
| Streptomyces sp. ADI96-15 | Antimycin, Lobophorins | [117] |
| Streptomyces sp. ADI97-07 | Neocarzilin, Antimycin | [117] |
| Streptomyces sp. ADI98-10 | Actinomycin | [117] |
| Streptomyces sp. B9173 | Flaviogeranin B1, B, C2, D | [118] |
| Streptomyces sp. BB47 | Antarlides F–H | [234] |
| Streptomyces sp. CHQ-64 ΔrdmF | Geranylpyrrol A, Piericidin F | [235] |
| Streptomyces sp. CMAA 1527 | Cinerubin B | [15] |
| Streptomyces sp. CMAA 1653 | Actinomycin V | [15,120] |
| Streptomyces sp. CPCC 204,980 | Cervinomycins B1-B4 | [120] |
| Streptomyces sp. DT-A61 | 9-hydroxy-K252c, 3-hydroxy-K252c, 3-hydroxy-7-methoxy-K252c, Nacetylholyrine A, 3-hydroxyholyrine A, 3′-O-demethyl-4′-N-demethyl-4′-N-acetyl-4′-epi-staurosporine, Streptocarbazoles D, E | [236] |
| Streptomyces sp. EGY1 | Sharkquinone | [237] |
| Streptomyces sp. GIC10-1 | Bafilomycin M | [238] |
| Streptomyces sp. GKU 220 | Rakicidin F | [123] |
| Streptomyces sp. HF-11225 | Nivelactam B | [239] |
| Streptomyces sp. HN-A124 | Cysrabelomycin | [126] |
| Streptomyces sp. HNA39 | Cyclizidines B–I | [240] |
| Streptomyces sp. HS-NF-1178 | 218-seco-lankacidinols A, B | [241] |
| Streptomyces sp. HS-NF-780 | 9-methylstreptimidone 2-α-d-glucopyranoside, ydroxyiso-9-methylstreptimidone | [242] |
| Streptomyces sp. HZP-2216E | 23-O-butyrylbafilomycin D | [81,128] |
| Streptomyces sp. ICN19 | Ala-geninthiocin | [130] |
| Streptomyces sp. IFM 11490 | Elmenols G | [243] |
| Streptomyces sp. KCB13F003 | Ulleungdin | [244] |
| Streptomyces sp. KCB13F030 | Ulleungoside | [245] |
| Streptomyces sp. KIB-H1318 | Phenoxazinone-related alkaloid 2 | [246] |
| Streptomyces sp. KIB-H714 | Actinomycin Z6 | [247] |
| Streptomyces sp. M268 | Kiamycin B | [248] |
| Streptomyces sp. MSB090213SC12 | Neothioviridamide | [249] |
| Streptomyces sp. MUSC 125 | 8-[N-Aziridylethylamino]-2-6,dimethyloctene-2 | [42] |
| Streptomyces sp. N1510.2 | Strepantibin D | [250] |
| Streptomyces sp. N1510.2 | Strepantibins A–C | [251] |
| Streptomyces sp. NA4286 | Murayaquinone D | [252] |
| Streptomyces sp. NEAU-L3 | Tetracenoquinocin A | [253] |
| Streptomyces sp. OPMA00071 | JBIR-150 | [254] |
| Streptomyces sp. PU-14G | Puromycins C, D | [255] |
| Streptomyces sp. PU-KB10-4 | 4-hydroxycinnamide | [104] |
| Streptomyces sp. Q22 | Bagremycins C | [256] |
| Streptomyces sp. RAI364 | Curromycin A | [257] |
| Streptomyces sp. RK88 | Opantimycin A | [258] |
| Streptomyces sp. SBT345 | Strepoxazine A | [259] |
| Streptomyces sp. SCSIO 03032 | Indimicins F, G, Spiroindimicins G, H | [260] |
| Streptomyces sp. SCSIO 1666/17C4 (engineered) | l-rhodinose-l-rhodinose-2-deoxy-l-fucose-10-decarbomethoxy-ε-rhodomycinone | [219] |
| Streptomyces sp. SCSIO 40063 | Piericidins A5, G1 | [261] |
| Streptomyces sp. SCSIO 41399 | Aranciamycin K, Isotirandamycin B | [141] |
| Streptomyces sp. SS13I | Ephyromycins B, C | [262] |
| Streptomyces sp. SS17F | Thioquinomycins A, C, D | [263] |
| Streptomyces sp. SSA28 | Cytosaminomycin E | [264] |
| Streptomyces sp. TR1341 | Filipin, Fungichromin, Actinomycin X2 | [181] |
| Streptomyces sp. XMA39 | Strepoxepinmycins A–D | [146] |
| Streptomyces sp. W2061 | Rubiflavin G Photorubiflavin G, E | [265] |
| Streptomyces sp. YIM S01863 | Jiangchuanmycin | [266] |
| Streptomyces sp. ZZ741 | Streptoglutarimides H | [149] |
| Streptomyces OPOK_MB_A9 | Milbemycin-α8 | [137] |
| Streptomyces sp. shell-016 | Shellmycin A–D | [267] |
| Umezawaea sp. RD066910 + Tsukamurella pulmonis TP-B0596 | Umezawamides A & B | [185] |
| Verrucosispora sp. SCSIO 07399 | Kendomycins B-D | [153] |
| Organism (s) | Compound Name (s) | Reference (s) |
|---|---|---|
| Actinomadura sp. 2EPS | Decatromicins | [278] |
| Kibdelosporangium persicum | Persicamidines A–E | [279] |
| Kutzneria albida DSM 43870 | Huimycin | [106,280] |
| Streptomyces kebangsaanensis WS-68302 | Napyradiomycin A4, A80915 H | [281] |
| Streptomyces jiujiangensis NBERC-24992 | Virantmycins D–G | [282] |
| Streptomyces bacillaris | Zelkovamycins F, G | [283] |
| Streptomyces koyangensis SCSIO 5802 | Neoabyssomicins F, G | [106,284] |
| Streptomyces sp. AM-2504 | Virantmycins B | [284,285] |
| Streptomyces sp. CPCC 200267 | Geninthiocins E, F | [286] |
| Streptomyces sp. HK18 | Xiamycins D | [285,287] |
| Streptomyces sp. JA74 | Dihydromaniwamycin E | [288] |
| Streptomyces sp. SMU 03 | dichloromethane extracts (DCME) | [287] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ngamcharungchit, C.; Chaimusik, N.; Panbangred, W.; Euanorasetr, J.; Intra, B. Bioactive Metabolites from Terrestrial and Marine Actinomycetes. Molecules 2023, 28, 5915. https://doi.org/10.3390/molecules28155915
Ngamcharungchit C, Chaimusik N, Panbangred W, Euanorasetr J, Intra B. Bioactive Metabolites from Terrestrial and Marine Actinomycetes. Molecules. 2023; 28(15):5915. https://doi.org/10.3390/molecules28155915
Chicago/Turabian StyleNgamcharungchit, Chananan, Nutsuda Chaimusik, Watanalai Panbangred, Jirayut Euanorasetr, and Bungonsiri Intra. 2023. "Bioactive Metabolites from Terrestrial and Marine Actinomycetes" Molecules 28, no. 15: 5915. https://doi.org/10.3390/molecules28155915
APA StyleNgamcharungchit, C., Chaimusik, N., Panbangred, W., Euanorasetr, J., & Intra, B. (2023). Bioactive Metabolites from Terrestrial and Marine Actinomycetes. Molecules, 28(15), 5915. https://doi.org/10.3390/molecules28155915