Identification and Structural Characterization of Novel Chondroitin/Dermatan Sulfate Hexassacharide Domains in Human Decorin by Ion Mobility Tandem Mass Spectrometry
Abstract
:1. Introduction
2. Results and Discussion
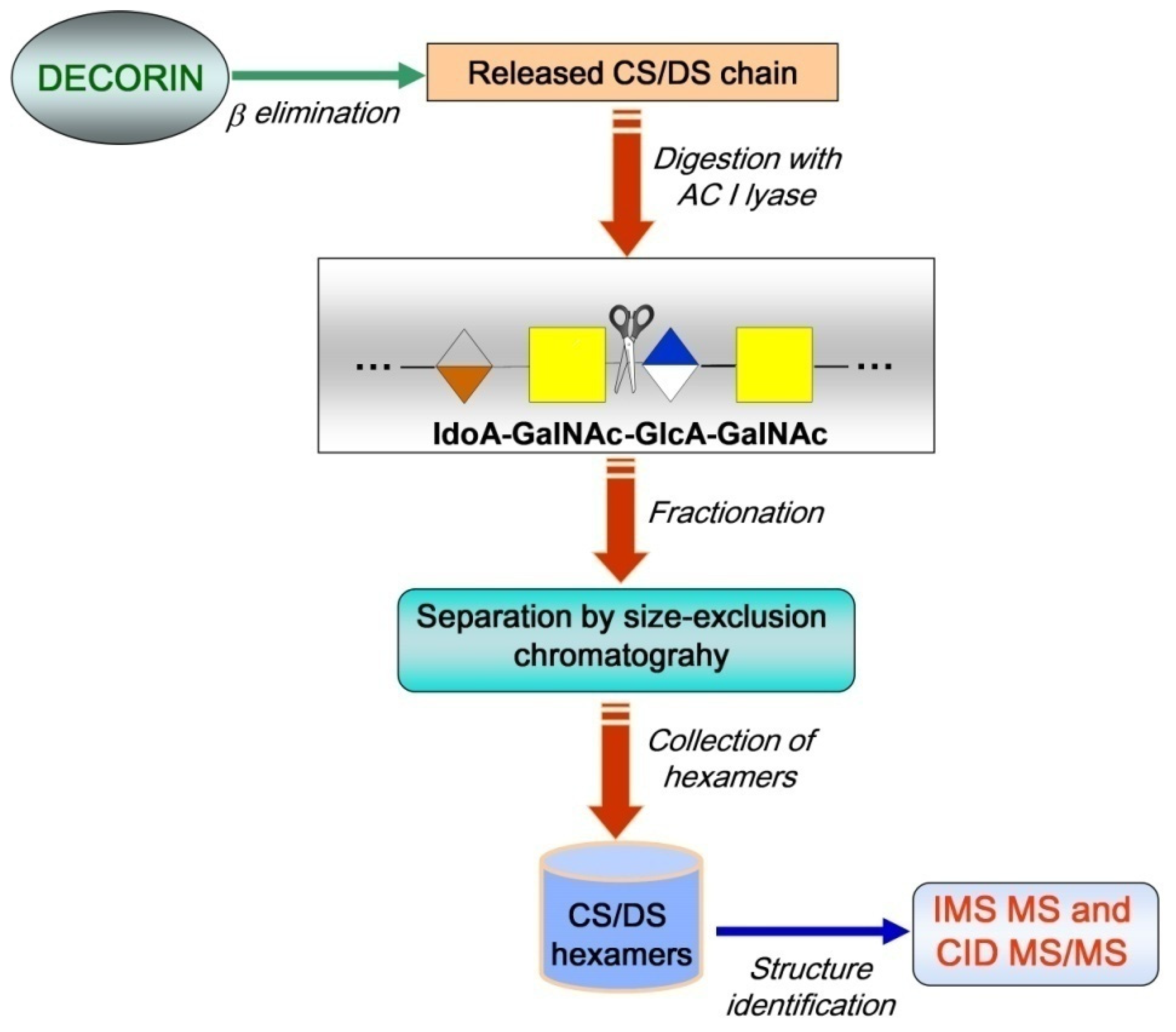
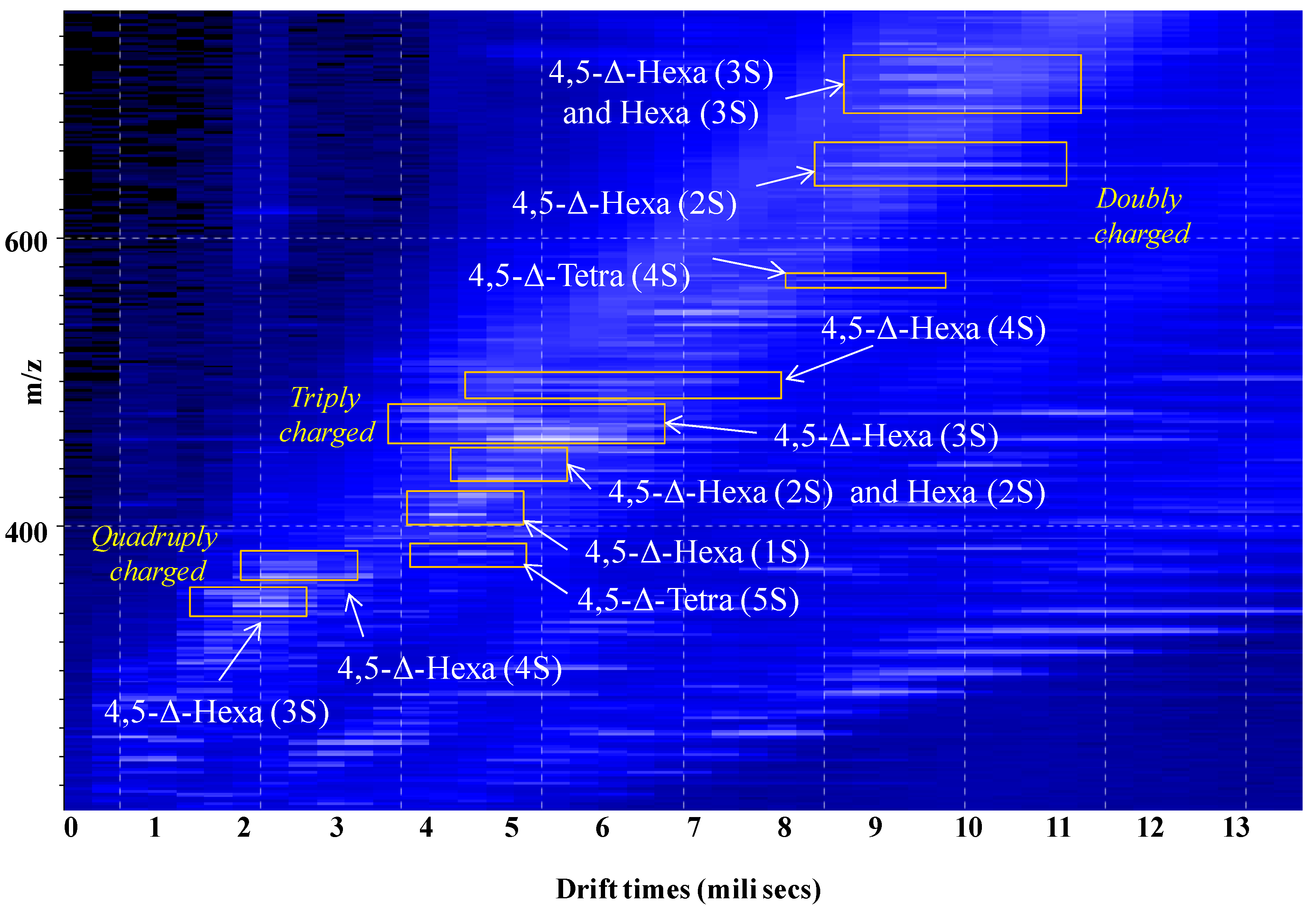
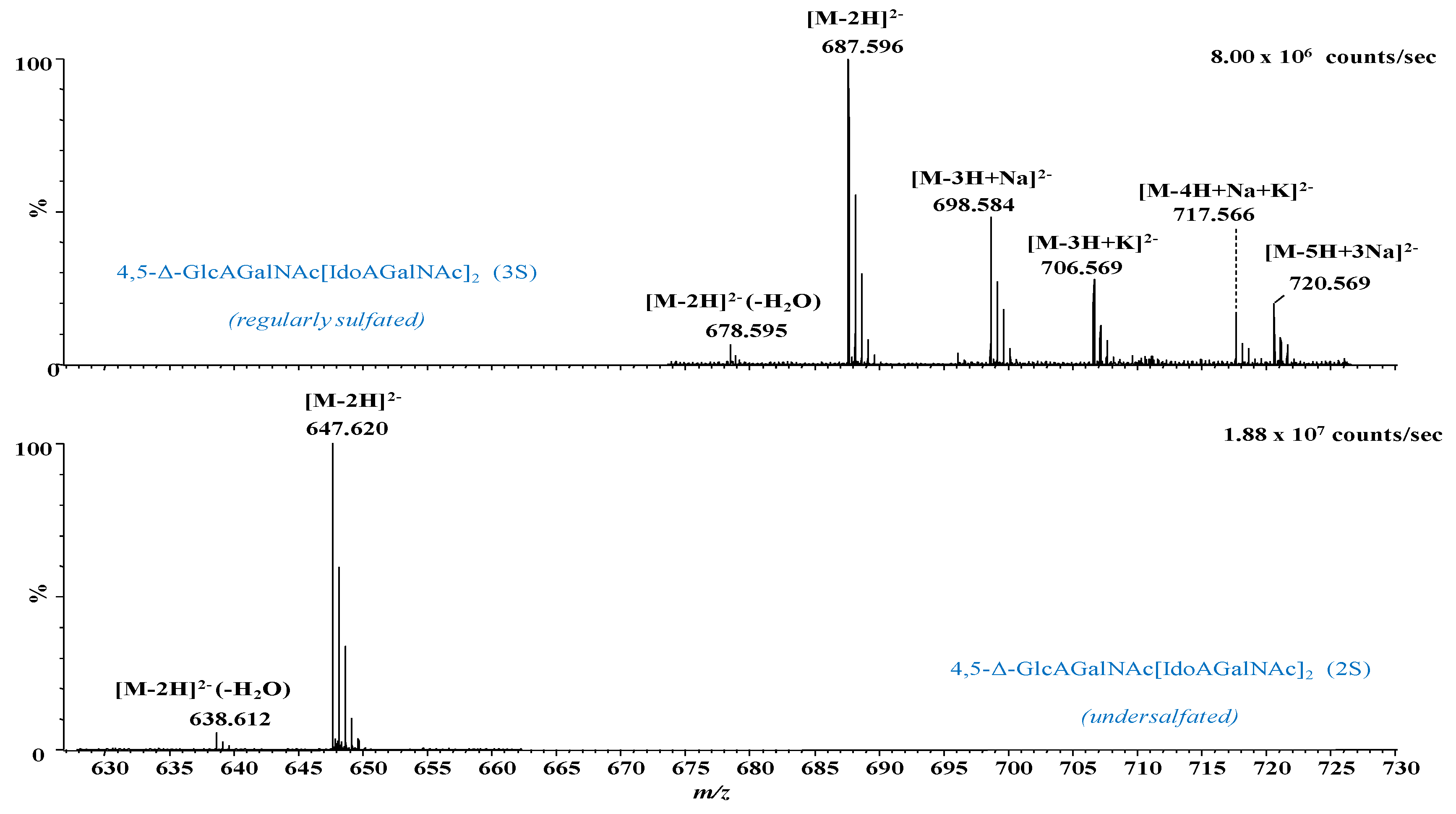
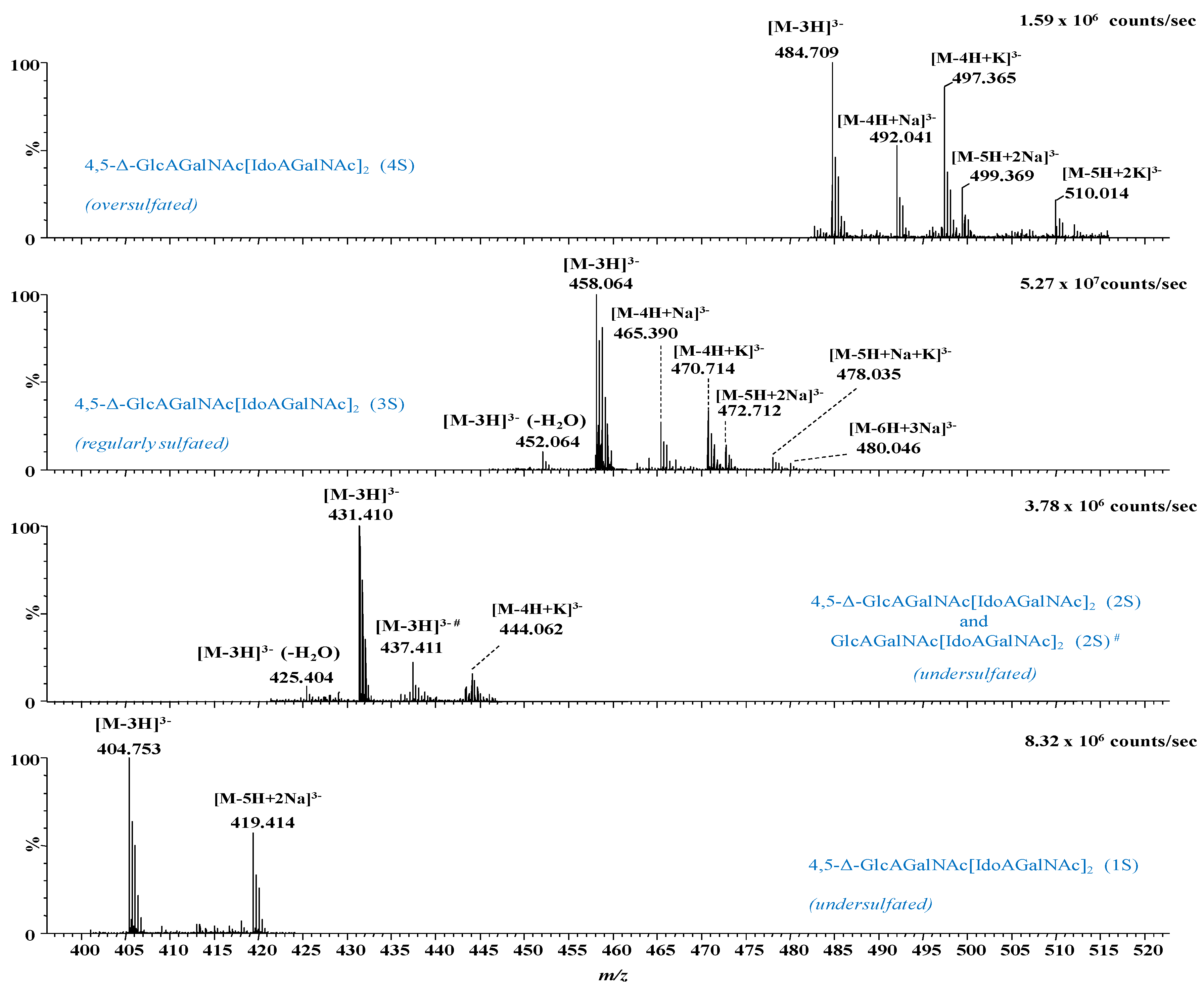
2.1. IMS MS Analysis of Hexasaccharide Domains in CS/DS from HEK293 DCN
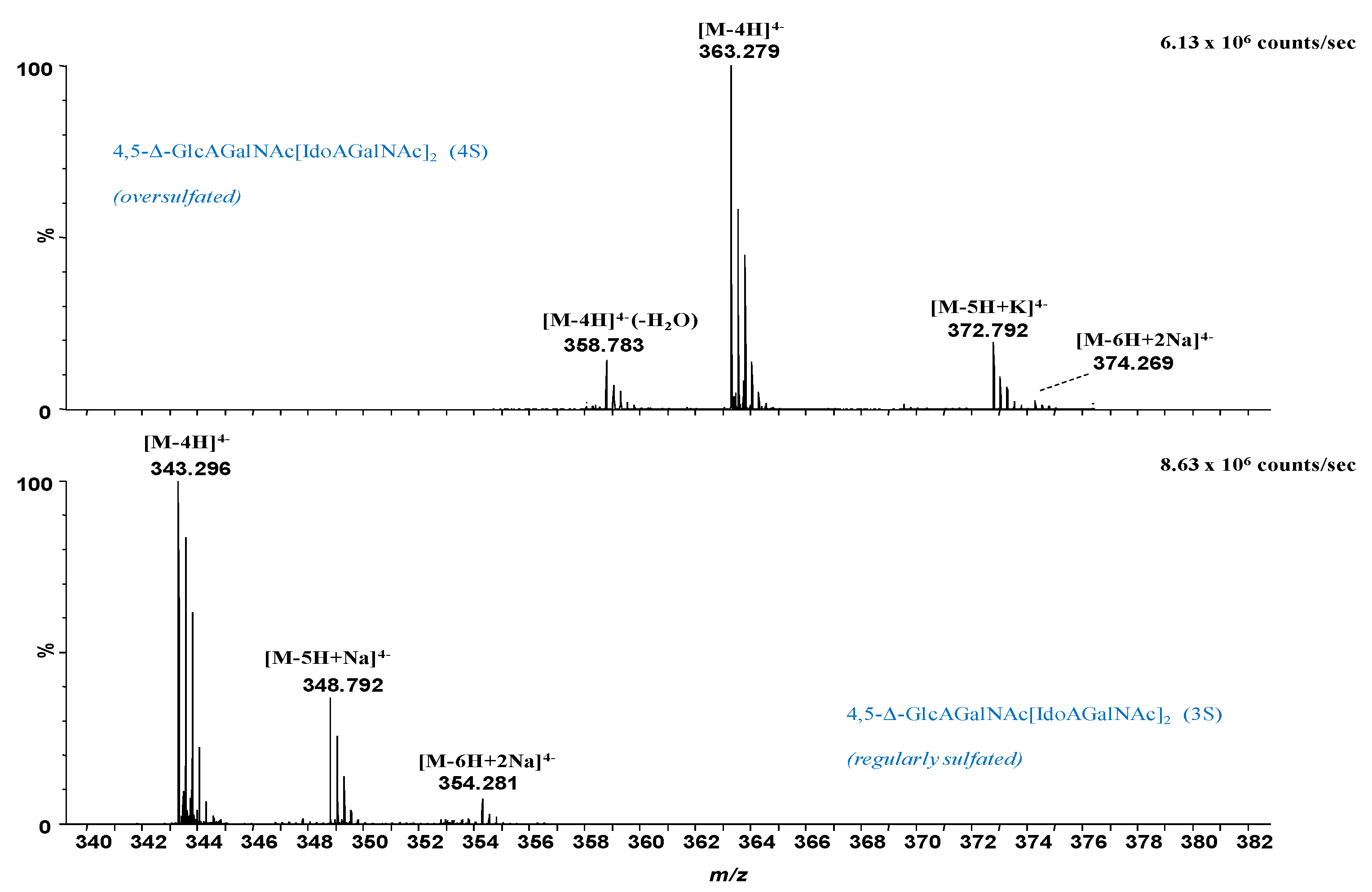
2.2. IMS CID MS/MS for the Determination of Misregulations in the Sulfation Code
3. Materials and Methods
3.1. CS/DS Hexasaccharide Fraction
3.2. IMS MS and CID Tandem MS
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Wei, J.; Hu, M.; Huang, K.; Lin, S.; Du, H. Roles of proteoglycans and glycosaminoglycans in cancer development and progression. Int. J. Mol. Sci. 2020, 21, 5983. [Google Scholar] [CrossRef] [PubMed]
- Cortes, M.; Cortes, L.K.; Schwartz, N.B. Mapping proteoglycan function using novel genetic strategies. Methods Mol. Biol. 2022, 2303, 731–752. [Google Scholar] [CrossRef] [PubMed]
- Song, Y.; Zhang, F.; Linhardt, R.J. Glycosaminoglycans. Adv. Exp. Med. Biol. 2021, 1325, 103–116. [Google Scholar] [CrossRef] [PubMed]
- Ghatak, S.; Maytin, E.V.; Mack, J.A.; Hascall, V.C.; Atanelishvili, I.; Rodriguez, R.M.; Markwald, R.R.; Misra, S. Roles of proteoglycans and glycosaminoglycans in wound healing and fibrosis. Int. J. Cell Biol. 2015, 2015, 834893. [Google Scholar] [CrossRef]
- Vallet, S.D.; Clerc, O.; Ricard-Blum, S. Glycosaminoglycan-protein interactions: The first draft of the glycosaminoglycan interactome. J. Histochem. Cytochem. 2020, 69, 93–104. [Google Scholar] [CrossRef]
- Mizumoto, S.; Yamada, S. The specific role of dermatan sulfate as an instructive glycosaminoglycan in tissue development. Int. J. Mol. Sci. 2022, 23, 7485. [Google Scholar] [CrossRef]
- Bougatef, H.; Ghlissi, Z.; Kallel, R.; Ben Amor, I.; Boudawara, T.; Gargouri, J.; Sahnoun, Z.; Volpi, N.; Sila, A.; Bougatef, A. Chondroitin/dermatan sulfate purified from corb (Sciaena umbra) skin and bone: In vivo assessment of anticoagulant activity. Int. J. Biol. Macromol. 2020, 164, 131–139. [Google Scholar] [CrossRef]
- Zhang, B.; Chi, L. Chondroitin sulfate/dermatan sulfate-protein interactions and their biological functions in human diseases: Implications and analytical tools. Front. Cell Dev. Biol. 2021, 9, 693563. [Google Scholar] [CrossRef]
- Ren, Q.; Wang, J.; Liu, C.; Meng, L.-X.; Qian, R.-K.; Gao, H.-J.; Qin, W.; Zhou, C.-J.; Qiao, S.; Wang, H.-Y.; et al. Exploring the sulfate patterns of chondroitin sulfate/dermatan sulfate and keratan sulfate in human pancreatic cancer. J. Pharm. Biomed. Anal. 2021, 205, 114339. [Google Scholar] [CrossRef]
- Li, G.; Li, L.; Joo, E.J.; Son, J.W.; Kim, Y.J.; Kang, J.; Lee, K.B.; Zhang, F.; Linhardt, R.J. Glycosaminoglycans and glycolipids as potential biomarkers in lung cancer. Glycoconj. J. 2017, 34, 661–669. [Google Scholar] [CrossRef]
- Yu, Y.; Duan, J.; Leach, I.F.E.; Toida, T.; Higashi, K.; Zhang, H.; Zhang, F.; Amster, I.J.; Linhardt, R.J. Sequencing the dermatan sulfate chain of decorin. J. Am. Chem. Soc. 2017, 139, 16986–16995. [Google Scholar] [CrossRef] [PubMed]
- da Costa, D.S.; Reis, R.L.; Pashkuleva, I. Sulfation of glycosaminoglycans and its implications in human health and disorders. Annu. Rev. Biomed. Eng. 2017, 19, 1–26. [Google Scholar] [CrossRef] [PubMed]
- Flangea, C.; Petrescu, A.J.; Seidler, D.G.; Munteanu, C.V.A.; Zamfir, A.D. Identification of an unusually sulfated tetrasaccharide chondroitin/dermatan motif in mouse brain by combining chip-nanoelectrospray multistage MS2-MS4 and high resolution MS. Electrophoresis 2013, 34, 1581–1592. [Google Scholar] [CrossRef]
- Robu, A.C.; Popescu, L.; Munteanu, C.V.; Seidler, D.G.; Zamfir, A.D. Orbitrap mass spectrometry characterization of hybrid chondroitin/dermatan sulfate hexasaccharide domains expressed in brain. Anal. Biochem. 2015, 485, 122–131. [Google Scholar] [CrossRef]
- Hayes, A.J.; Melrose, J. Neural tissue homeostasis and repair is regulated via CS and DS proteoglycan motifs. Front. Cell Dev. Biol. 2021, 9, 696640. [Google Scholar] [CrossRef] [PubMed]
- Li, H.-P.; Komuta, Y.; Kimura-Kuroda, J.; van Kuppevelt, T.H.; Kawano, H. Roles of chondroitin sulfate and dermatan sulfate in the formation of a lesion scar and axonal regeneration after traumatic injury of the mouse brain. J. Neurotrauma 2013, 30, 413–425. [Google Scholar] [CrossRef]
- Sugahara, K.; Mikami, T. Chondroitin/dermatan sulfate in the central nervous system. Curr. Opin. Struct. Biol. 2007, 17, 536–545. [Google Scholar] [CrossRef]
- Bilong, M.; Bayat, P.; Bourderioux, M.; Jérôme, M.; Giuliani, A.; Daniel, R. Mammal hyaluronidase activity on chondroitin sulfate and dermatan sulfate: Mass spectrometry analysis of oligosaccharide products. Glycobiology 2021, 31, 751–761. [Google Scholar] [CrossRef]
- Solakyildirim, K. Recent advances in glycosaminoglycan analysis by various mass spectrometry techniques. Anal. Bioanal. Chem. 2019, 411, 3731–3741. [Google Scholar] [CrossRef]
- Zamfir, A.D.; Flangea, C.; Sisu, E.; Serb, A.F.; Dinca, N.; Bruckner, P.; Seidler, D.G. Analysis of novel over- and undersulfated glycosaminoglycan sequences by enzyme cleavage and multiple stage mass spectrometry. Proteomics 2009, 9, 3435–3444. [Google Scholar] [CrossRef]
- Robu, A.C.; Popescu, L.; Seidler, D.G.; Zamfir, A.D. Chip-based high resolution tandem mass spectrometric determination of fibroblast growth factor—Chondroitin sulfate disaccharides noncovalent interaction. Biol. Mass Spectrom. 2018, 53, 624–634. [Google Scholar] [CrossRef] [PubMed]
- Zamfir, A.D. Microfluidics-mass spectrometry of protein-carbohydrate interactions: Applications to the development of therapeutics and biomarker discovery. Methods Mol. Biol. 2017, 1647, 109–128. [Google Scholar] [CrossRef]
- Miller, M.J.C.; Costello, C.E.; Malmström, A.; Zaia, J. A tandem mass spectrometric approach to determination of chondroitin/dermatan sulfate oligosaccharide glycoforms. Glycobiology 2006, 16, 502–513. [Google Scholar] [CrossRef] [PubMed]
- Gill, V.L.; Aich, U.; Rao, S.; Pohl, C.; Zaia, J. Disaccharide Analysis of Glycosaminoglycans Using Hydrophilic Interaction Chromatography and Mass Spectrometry. Anal. Chem. 2013, 85, 1138–1145. [Google Scholar] [CrossRef] [PubMed]
- Persson, A.; Nikpour, M.; Vorontsov, E.; Nilsson, J.; Larson, G. Domain mapping of chondroitin/dermatan sulfate glycosaminoglycans enables structural characterization of proteoglycans. Mol. Cell. Proteom. 2021, 20, 100074. [Google Scholar] [CrossRef]
- Tóth, G.; Pál, D.; Vékey, K.; Drahos, L.; Turiák, L. Stability and recovery issues concerning chondroitin sulfate disaccharide analysis. Anal. Bioanal. Chem. 2021, 413, 1779–1785. [Google Scholar] [CrossRef]
- Zamfir, A.D. Applications of capillary electrophoresis electrospray ionization mass spectrometry in glycosaminoglycan analysis. Electrophoresis 2016, 37, 973–986. [Google Scholar] [CrossRef]
- Zamfir, A.; Seidler, D.G.; Schönherr, E.; Kresse, H.; Peter-Katalinić, J. On-line sheathless capillary electrophoresis/nanoelectrospray ionization-tandem mass spectrometry for the analysis of glycosaminoglycan oligosaccharides. Electrophoresis 2004, 25, 2010–2016. [Google Scholar] [CrossRef]
- Zamfir, A.D.; Ica, R.; Sarbu, M. Liquid-phase separation methods hyphenated to electrospray ionization mass spectrometry for structural analysis of chondroitin/dermatan sulfate. In Carbohydrate Analysis by Modern Liquid Phase Separation Techniques; Elsevier: Amsterdam, The Netherlands, 2021; pp. 529–562. [Google Scholar]
- Sarbu, M.; Zhu, F.; Peter-Katalinić, J.; Clemmer, D.E.; Zamfir, A.D. Application of ion mobility tandem mass spectrometry to compositional and structural analysis of glycopeptides extracted from the urine of a patient diagnosed with Schindler disease. Rapid Commun. Mass Spectrom. 2015, 29, 1929–1937. [Google Scholar] [CrossRef]
- Sarbu, M.; Robu, A.C.; Ghiulai, R.M.; Vukelić, Ž.; Clemmer, D.E.; Zamfir, A.D. Electrospray ionization ion mobility mass spectrometry of human brain gangliosides. Anal. Chem. 2016, 88, 5166–5178. [Google Scholar] [CrossRef]
- Sarbu, M.; Vukelić, Ž.; Clemmer, D.E.; Zamfir, A.D. Ion mobility mass spectrometry provides novel insights into the expression and structure of gangliosides in normal adult human hippocampus. Analyst 2018, 143, 5234–5246. [Google Scholar] [CrossRef] [PubMed]
- Pallister, E.G.; Choo, M.S.F.; Walsh, I.; Tai, J.N.; Tay, S.J.; Yang, Y.S.; Ng, S.K.; Rudd, P.M.; Flitsch, S.L.; Nguyen-Khuong, T. Utility of ion-mobility spectrometry for deducing branching of multiply charged glycans and glycopeptides in a high-throughput positive ion LC-FLR-IMS-MS workflow. Anal. Chem. 2020, 92, 15323–15335. [Google Scholar] [CrossRef] [PubMed]
- Dufresne, M.; Guneysu, D.; Patterson, N.H.; Marcinkiewicz, M.M.; Regina, A.; Demeule, M.; Chaurand, P. Multimodal detection of GM2 and GM3 lipid species in the brain of mucopolysaccharidosis type II mouse by serial imaging mass spectrometry and immunohistochemistry. Anal. Bioanal. Chem. 2016, 409, 1425–1433. [Google Scholar] [CrossRef] [PubMed]
- Sarbu, M.; Fabris, D.; Vukelić, Ž.; Clemmer, D.E.; Zamfir, A.D. Ion mobility mass spectrometry reveals rare sialylated glycosphingolipid structures in human cerebrospinal fluid. Molecules 2022, 27, 743. [Google Scholar] [CrossRef] [PubMed]
- Miller, R.L.; Guimond, S.E.; Schwörer, R.; Zubkova, O.V.; Tyler, P.C.; Xu, Y.; Liu, J.; Chopra, P.; Boons, G.-J.; Grabarics, M.; et al. Shotgun ion mobility mass spectrometry sequencing of heparan sulfate saccharides. Nat. Commun. 2020, 11, 1481. [Google Scholar] [CrossRef]
- Wei, J.; Wu, J.; Tang, Y.; Ridgeway, M.E.; Park, M.A.; Costello, C.E.; Zaia, J.; Lin, C. Characterization and quantification of highly sulfated glycosaminoglycan isomers by gated-trapped ion mobility spectrometry negative electron transfer dissociation MS/MS. Anal. Chem. 2019, 91, 2994–3001. [Google Scholar] [CrossRef]
- Zhao, Y.; Yang, J.Y.; Thieker, D.F.; Xu, Y.; Zong, C.; Boons, G.-J.; Liu, J.; Woods, R.J.; Moremen, K.W.; Amster, I.J. A traveling wave ion mobility spectrometry (TWIMS) study of the robo1-heparan sulfate interaction. J. Am. Soc. Mass Spectrom. 2018, 29, 1153–1165. [Google Scholar] [CrossRef]
- Zaia, J. Glycosaminoglycan glycomics using mass spectrometry. Mol. Cell. Proteom. 2013, 12, 885–892. [Google Scholar] [CrossRef] [PubMed]
- Zamfir, A.; Seidler, D.G.; Kresse, H.; Peter-Katalinić, J. Structural investigation of chondroitin/dermatan sulfate oligosaccharides from human skin fibroblast decorin. Glycobiology 2003, 13, 733–742. [Google Scholar] [CrossRef]
- Domon, B.; Costello, C.E. A systematic nomenclature of carbohydrate fragmentation in FAB-MS/MS spectra of glycoconjugates. Glycoconj. J. 1988, 5, 397–409. [Google Scholar] [CrossRef]
- Costello, C.E.; Juhasz, P.; Perreault, H. Chapter 4 New mass spectral approaches to ganglioside structure determination. Prog. Brain Res. 1994, 101, 45–61. [Google Scholar] [CrossRef] [PubMed]


| No. | m/z exp | m/z Theor | Mass Accuracy (ppm) | Proposed Stucture | Sulfation | Molecular Ion |
|---|---|---|---|---|---|---|
| 1 | 305.040 | 305.037 | −8.7 | 4,5-Δ-GlcAGalNAcIdoAGalNAc (2S) | 2S | [M-3H]3− |
| 2 | 314.309 | 314.306 | −11.1 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 (1S) | 1S | [M-6H+2Na]4− |
| 3 | 323.308 | 323.304 | −13.2 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 (2S) | 2S | [M-4H]4− |
| 4 | 324.301 | 324.297 | −13.9 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 - H2O (2S) | 2S | [M-5H+Na]4− |
| 5 | 343.296 | 343.293 | −8.7 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 (3S) | 3S | [M-4H]4− |
| 6 | 348.792 | 348.789 | −10.1 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 (3S) | 3S | [M-5H+Na]4− |
| 7 | 352.785 | 352.782 | −9.2 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 (3S) | 3S | [M-3H+K]2− |
| 8 | 354.281 | 354.284 | 8.5 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 (3S) | 3S | [M-6H+2Na]4− |
| 9 | 358.783 | 358.780 | −9.8 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 - H2O (4S) | 4S | [M-4H]4− |
| 10 | 363.279 | 363.282 | 9.0 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 (4S) | 4S | [M-4H]4− |
| 11 | 372.774 | 372.771 | −8.1 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 (4S) | 4S | [M-5H+K]4− |
| 12 | 374.269 | 374.273 | 11.4 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 (4S) | 4S | [M-6H+2Na]4− |
| 13 | 384.990 | 384.994 | 11.3 | 4,5-Δ-GlcAGalNAcIdoAGalNAc (5S) | 5S | [M-3H]3− |
| 14 | 404.753 | 404.755 | 5.8 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 (1S) | 1S | [M-3H]3− |
| 15 | 419.414 | 419.410 | −9.5 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 (1S) | 1S | [M-5H+2Na]3− |
| 16 | 425.404 | 425.408 | 8.6 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 - H2O (2S) | 2S | [M-3H]3− |
| 17 | 431.410 | 431.408 | −5.4 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 (2S) | 2S | [M-3H]3− |
| 18 | 437.411 | 437.408 | −7.6 | GlcAGalNAc[IdoAGalNAc]2 (2S) | 2S | [M-3H]3− |
| 19 | 444.062 | 444.059 | −6.0 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 (2S) | 2S | [M-4H+K]3− |
| 20 | 449.064 | 449.060 | −8.9 | 4,5-Δ-GlcAGalNAcIdoAGalNAc - H2O (2S) | 2S | [M-2H]2− |
| 21 | 452.064 | 452.060 | −8.8 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 - H2O (3S) | 3S | [M-3H]3− |
| 22 | 458.064 | 458.060 | −8.7 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 (3S) | 3S | [M-3H]3− |
| 23 | 465.390 | 465.387 | −5.7 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 (3S) | 3S | [M-4H+Na]3− |
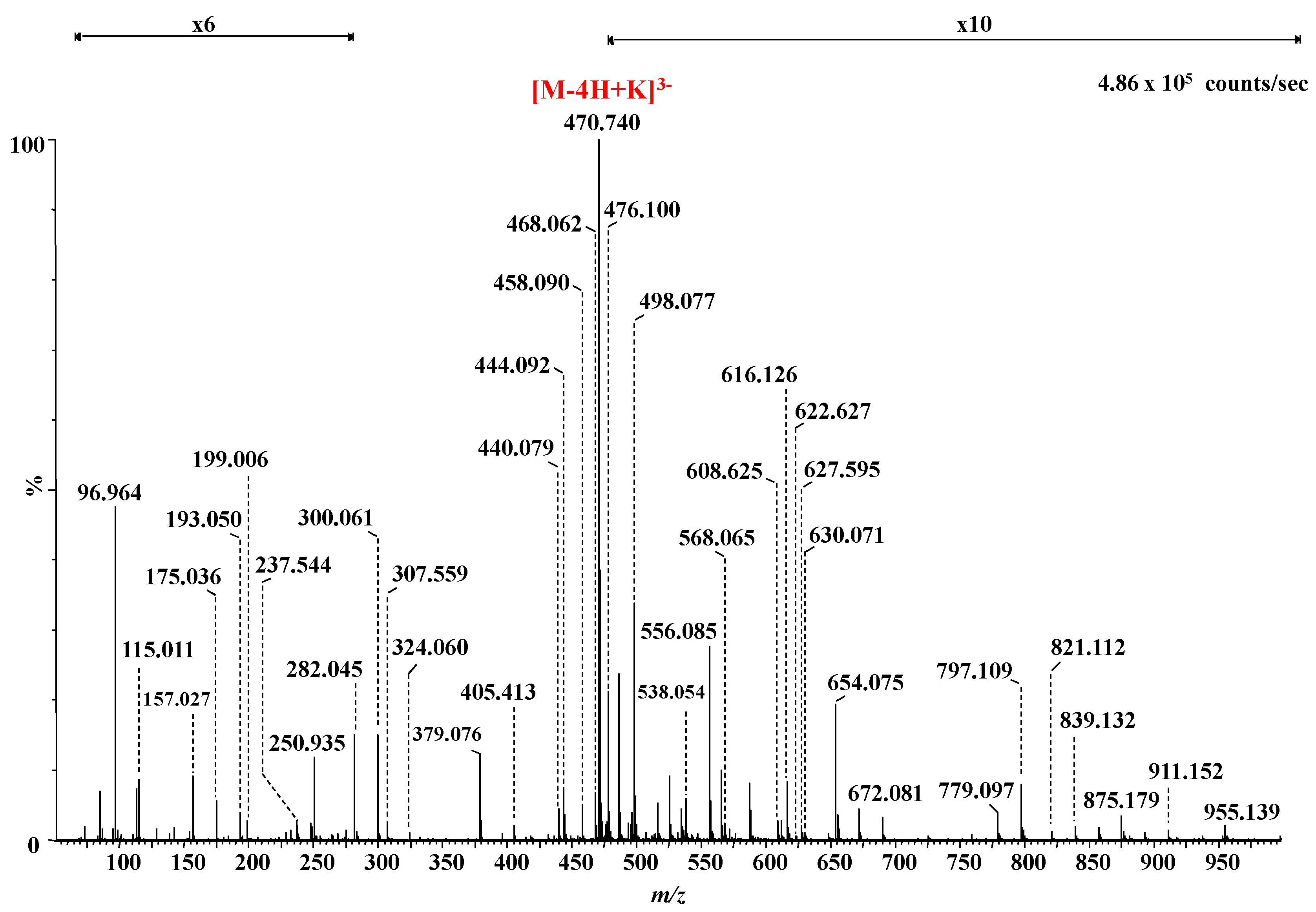
| 24 | 470.714 | 470.712 | −5.0 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 (3S) | 3S | [M-4H+K]3− |
| 25 | 472.712 | 472.715 | 5.6 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 (3S) | 3S | [M-5H+2Na]3− |
| 26 | 478.035 | 478.039 | 8.4 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 (3S) | 3S | [M-5H+Na+K]3− |
| 27 | 480.046 | 480.042 | −8.3 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 (3S) | 3S | [M-6H+3Na]3− |
| 28 | 484.709 | 484.712 | 6.9 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 (4S) | 4S | [M-3H]3− |
| 29 | 492.041 | 492.040 | −2.7 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 (4S) | 4S | [M-4H+Na]3− |
| 30 | 497.365 | 497.364 | −2.0 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 (4S) | 4S | [M-4H+K]3− |
| 31 | 499.369 | 499.367 | −4.0 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 (4S) | 4S | [M-5H+2Na]3− |
| 32 | 510.014 | 510.016 | 3.9 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 (4S) | 4S | [M-5H+2K]3− |
| 33 | 570.994 | 570.990 | −7.0 | 4,5-Δ-GlcAGalNAcIdoAGalNAc (4S) | 4S | [M-5H+3Na]2− |
| 34 | 629.617 | 629.619 | 3.2 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 (1S) | 1S | [M-4H+2Na]2− |
| 35 | 638.612 | 638.610 | −3.1 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 - H2O (2S) | 2S | [M-2H]2− |
| 36 | 647.620 | 647.616 | −7.0 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 (2S) | 2S | [M-2H]2− |
| 37 | 656.626 | 656.621 | −7.6 | GlcAGalNAc[IdoAGalNAc]2 (2S) | 2S | [M-2H]2− |
| 38 | 678.595 | 678.589 | −8.8 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 - H2O (3S) | 3S | [M-2H]2− |
| 39 | 687.596 | 687.594 | −2.9 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 (3S) | 3S | [M-2H]2− |
| 40 | 698.584 | 698.585 | 1.4 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 (3S) | 3S | [M-3H+Na]2− |
| 41 | 706.569 | 706.572 | 4.2 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 (3S) | 3S | [M-3H+K]2− |
| 42 | 717.566 | 717.563 | −4.9 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 (3S) | 3S | [M-4H+Na+K]2− |
| 43 | 720.569 | 720.567 | −2.8 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 (3S) | 3S | [M-5H+3Na]2− |
| 44 | 899.133 | 899.128 | −5.6 | 4,5-Δ-GlcAGalNAcIdoAGalNAc - 2H2O (2S) | 2S | [M-2H]2− |
| 45 | 1296.242 | 1296.239 | −2.3 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 (2S) | 2S | [M-H]− |
| 46 | 1376.198 | 1376.196 | −1.5 | 4,5-Δ-GlcAGalNAc[IdoAGalNAc]2 (3S) | 3S | [M-H]− |
| m/z | Charge State | Structure | The Type of Ion for the Structure in | |
|---|---|---|---|---|
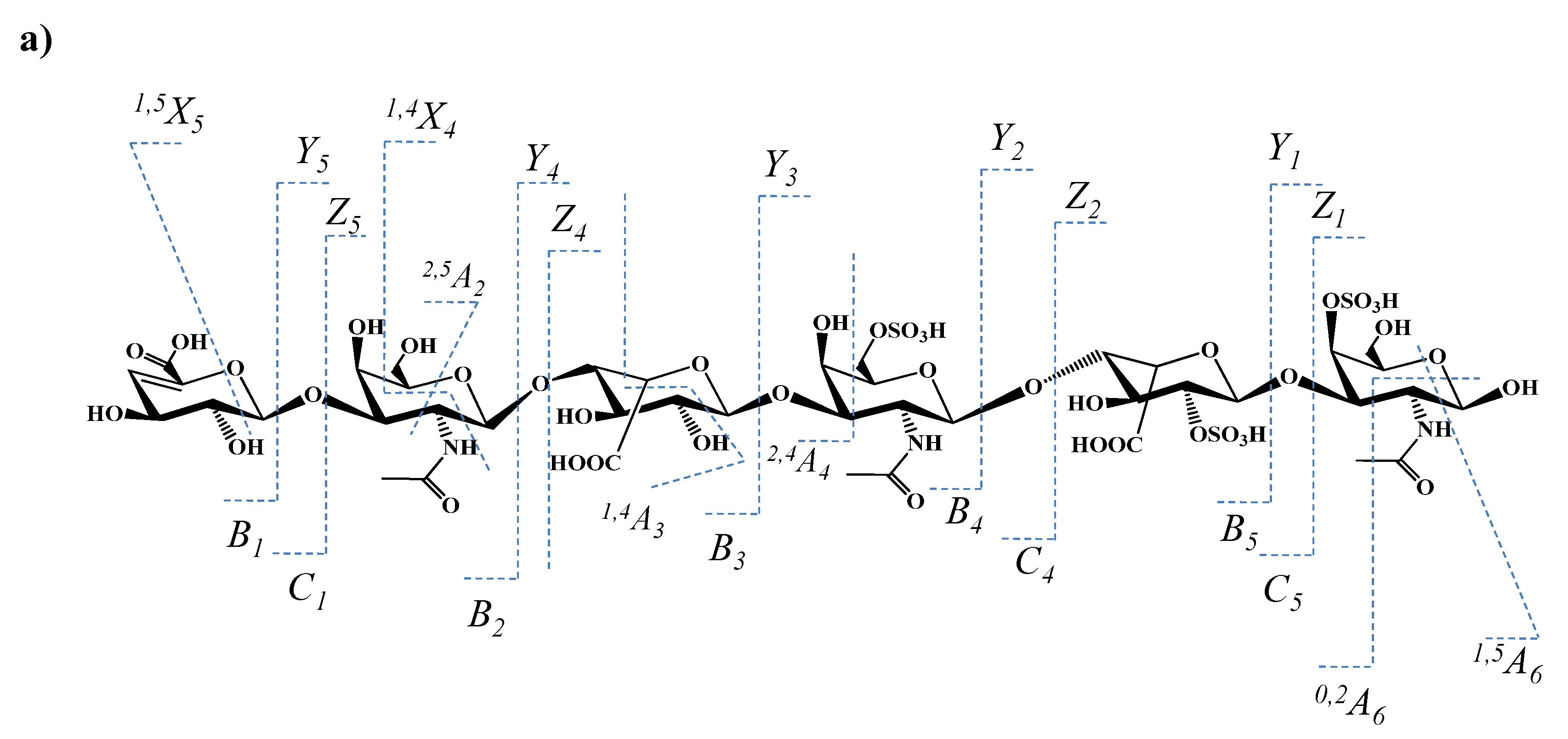
| Figure 8a | Figure 8b | |||
| 96.964 | 1 | HOSO3 | ||
| 115.011 | 1 | GalNAc | 2,5X0 | |
| 157.027 | 1 | 4,5-Δ-GlcA | B1 | |
| 175.036 | 1 | 4,5-Δ-GlcA | C1 | |
| 193.050 | 1 | IdoA (0S) | Y2/Z1 or Y4/Z3 | |
| 199.006 | 1 | GalNAc (1S) | 0,2A6/B5 or0,2A4/B3 | |
| 237.544 | 1 | IdoA (1S) b | Z2/Y1 | Z4/Y3 |
| 282.045 | 1 | GalNAc (1S) a | Z1 | Z5/Z4 |
| 300.061 | 1 | GalNAc (1S) a | Y1 | Y5/Z4 |
| 307.559 | 2 | IdoAGalNAcIdoA (1S) a | Z4/Y1 | B3 |
| 343.318 | 1 | IdoAGalNAc (1S) a | 0,2X1 | 2,5A2 |
| 379.076 | 2 | GalNAcIdoAGalNAc (2S) a | Y3 | Z5/Z2 |
| 405.413 | 3 | GalNAcIdoAGalNAcIdoAGalNAc (3S) a | Y5 | |
| 418.068 | 3 | GalNAcIdoAGalNAcIdoAGalNAc (3S) a | Y5+K | |
| 432.071 | 3 | GalNAcIdoAGalNAcIdoAGalNAc (3S) a | 2,5A6 +K | |
| 440.079 | 1 | 4,5-Δ-GlcAGalNAc (1S) or IdoAGalNAc (1S) a | Z4/Y2 | B2 |
| 444.092 | 3 | GlcAGalNAcIdoAGalNAcIdoAGalNAc (3S) a | 1,5A6 | 1,5A6 |
| 454.023 | 1 | IdoAGalNAc (2S) b | 0,3X1 | 0,3X3/Z2 |
| 458.090 | 3 | GlcAGalNAcIdoAGalNAcIdoAGalNAc (3S) a | [M-3H]3− | |
| 468.062 | 2 | IdoAGalNAcIdoAGalNAc (2S) a | Y4 | |
| 470.714 | 3 | 4,5-Δ-GlcAGalNAcIdoAGalNAcIdoAGalNAc (3S) a | [M-4H+K]3− | |
| 476.100 | 1 | IdoAGalNAc (1S) a | Y4/Z2 | Y3/Z1 orY4/Z2 |
| 498.077 | 2 | IdoAGalNAcIdoAGalNAc (3S) b | Z4 | C4 |
| 507.085 | 2 | IdoAGalNAcIdoAGalNAc (3S) b | Y4 | Y5/Z1 |
| 538.054 | 1 | 4,5-Δ-GlcAGalNAcIdoA (0S) | B3 | |
| IdoAGalNAc (2S) b | Z2 | Y4/Y2 | ||
| 543.091 | 2 | GalNAcIdoAGalNAcIdoAGalNAc (3S) a | 1,4X4 | |
| 556.085 | 2 | 4,5-Δ-GlcAGalNAcIdoAGalNAcIdoA (2S) a | B5 +K | |
| 565.092 | 2 | 4,5-Δ-GlcAGalNAcIdoAGalNAcIdoA (2S) a | C5 +K | |
| 568.065 | 2 | GalNAcIdoAGalNAcIdoAGalNAc (2S) | Y5 | |
| 608.625 | 2 | GalNAcIdoAGalNAcIdoAGalNAc (3S) a | Y5 | |
| 612.035 | 1 | IdoAGalNAcIdoA (2S) b | 1,4A5/C2 | 1,4A3 |
| 616.126 | 1 | 4,5-Δ-GlcAGalNAcIdoA (1S) a IdoAGalNAcIdoA (1S) a | - Z4/Y1 | B3 Z4/Y1 |
| 622.627 | 2 | 4,5-Δ-GlcAGalNAcIdoAGalNAcIdoAGalNAc (3S) a | 1,5X5 | |
| 627.595 | 2 | GalNAcIdoAGalNAcIdoAGalNAc (3S) a | Y5+K | |
| 630.071 | 2 | 4,5-Δ-GlcAGalNAcIdoAGalNAcIdoAGalNAc (3S) a | 2,5A6 | |
| 648.613 | 2 | GalNAcIdoAGalNAcIdoAGalNAc (3S) a | 2,5A6 +K | |
| 654.075 | 1 | IdoAGalNAcIdoA (1S) a | Z4/Y1+K | B3+K |
| 779.097 | 1 | GalNAcIdoAGalNAc (3S) b | 2,4X2 | 2,4X4/Y2 |
| 797.109 | 1 | GalNAcIdoAGalNAc (2S) a | Y3+K | Y5/Z2+K |
| 821.112 | 1 | GalNAcIdoAGalNAc (3S) b | Z3 | Y5/Y2 |
| 839.132 | 1 | GalNAcIdoAGalNAc (3S) b | Y3 | Y5/Z2 |
| 857.169 | 1 | 4,5-Δ-GlcAGalNAcIdoAGalNAc (1S) | B4+K | B4+K or Y5/Y1+K |
| 875.179 | 1 | 4,5-Δ-GlcAGalNAcIdoAGalNAc (1S) | C4+K | C4+K or Y5/Z1+K |
| 911.152 | 1 | IdoAGalNAcIdoAGalNAc (3S) b | 0,3X3 | C4/0,3A1 |
| 917.187 | 1 | IdoAGalNAcIdoAGalNAc (2S) a | Z4 | Z4 |
| 955.139 | 1 | IdoAGalNAcIdoAGalNAc (3S) b IdoAGalNAcIdoAGalNAc (2S) a | 2,4X3 Z4+K | 2,4X4/Z1 Z4+K |
| 997.154 | 1 | IdoAGalNAcIdoAGalNAc (3S) b | Z4 | Y5/Y1 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sarbu, M.; Ica, R.; Sharon, E.; Clemmer, D.E.; Zamfir, A.D. Identification and Structural Characterization of Novel Chondroitin/Dermatan Sulfate Hexassacharide Domains in Human Decorin by Ion Mobility Tandem Mass Spectrometry. Molecules 2022, 27, 6026. https://doi.org/10.3390/molecules27186026
Sarbu M, Ica R, Sharon E, Clemmer DE, Zamfir AD. Identification and Structural Characterization of Novel Chondroitin/Dermatan Sulfate Hexassacharide Domains in Human Decorin by Ion Mobility Tandem Mass Spectrometry. Molecules. 2022; 27(18):6026. https://doi.org/10.3390/molecules27186026
Chicago/Turabian StyleSarbu, Mirela, Raluca Ica, Edie Sharon, David E. Clemmer, and Alina D. Zamfir. 2022. "Identification and Structural Characterization of Novel Chondroitin/Dermatan Sulfate Hexassacharide Domains in Human Decorin by Ion Mobility Tandem Mass Spectrometry" Molecules 27, no. 18: 6026. https://doi.org/10.3390/molecules27186026
APA StyleSarbu, M., Ica, R., Sharon, E., Clemmer, D. E., & Zamfir, A. D. (2022). Identification and Structural Characterization of Novel Chondroitin/Dermatan Sulfate Hexassacharide Domains in Human Decorin by Ion Mobility Tandem Mass Spectrometry. Molecules, 27(18), 6026. https://doi.org/10.3390/molecules27186026





