Quantitative Retention (Structure)–Activity Relationships in Predicting the Pharmaceutical and Toxic Properties of Potential Pesticides
Abstract
1. Introduction
2. Results
3. Discussion
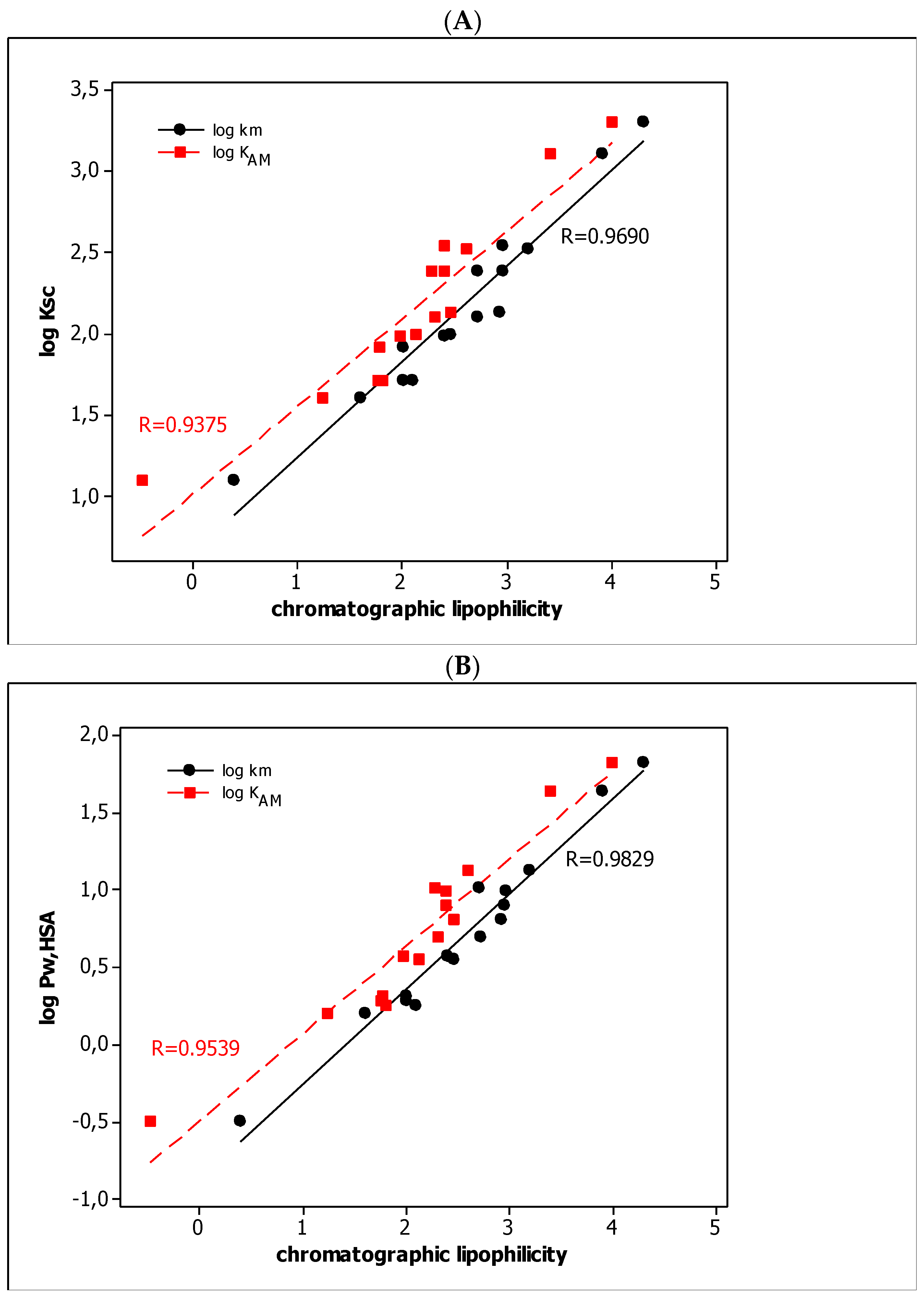
3.1. QRARs
3.2. QSARs
| No. | Equation |
|---|---|
| (3) | log Kp = −5.329(0.156) + 0.597(0.041)log km − 0.669(0.075)HBD − 0.345(0.061)HBA + 0.197(0.044)NRB n = 15; R = 0.9893; Radj. = 0.9850; sd = 0.110; PRESS = 0.276; MSE = 0.012; F = 116; p < 0.0001; VIF < 3.6; PRESScv = 0.276; MSEcv = 0.012 |
| (4) | log Kp = −4.900(0.218) + 0.552(0.060)log KAM − 0.714(0.118)HBD − 0.378(0.092)HBA + 0.213(0.067)NRB n = 15; R = 0.9747; Radj. = 0.9644; sd = 0.169; PRESS = 0.610; MSE = 0.0286; F = 48; p < 0.0001; VIF < 3.5; PRESScv = 0.610; MSEcv = 0.0286 |
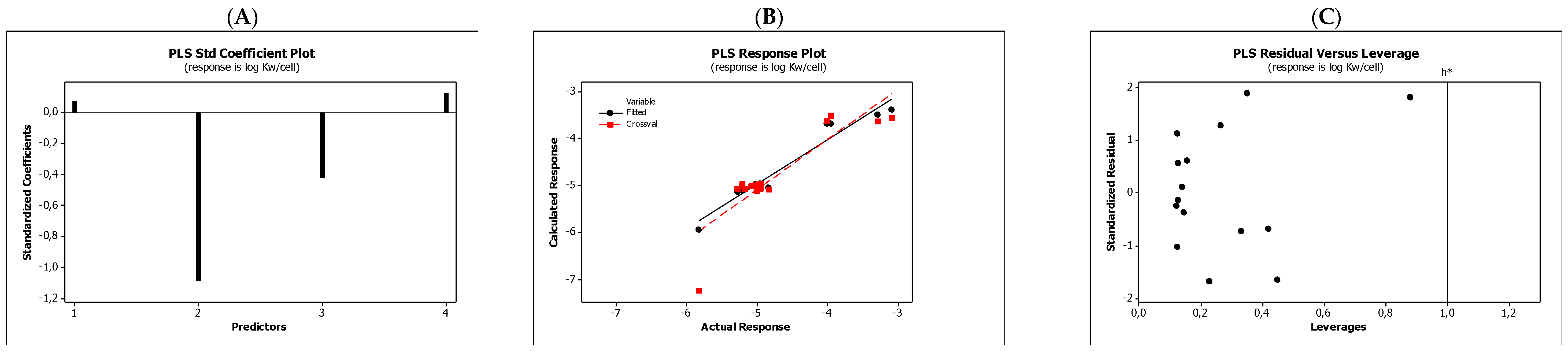
| (5) | log Kw/cell = −2.773(0.295) + 0.062(0.078)log km − 1.846(0.143)HBD − 0.368(0.115)HBA + 0.076(0.083)NRB n = 15; R = 0.9740; Radj = 0.9634; sd = 0.209; PRESS =4.064; MSE =0.0435; F = 47; p < 0.0001; VIF < 3.6; PRESScv = 4.064; MSEcv = 0.0435 |
| (6) | log Kw/cell = −2.762(0.262) + 0.079(0.073)log KAM − 1.867(0.142)HBD − 0.356(0.111)HBA + 0.064(0.081)NRB n = 15; R = 0.9753; Radj = 0.9652; sd = 0.203; PRESS = 4.889; MSE =0.0414; F = 49; p < 0.0001; VIF < 3.5; PRESScv = 4.889; MSEcv = 0.0400 |
| (7) | log KHSA = 4.619(0.350) + 0.251(0.093)log km + 0.106(0.170)HBD − 0.381(0.136)HBA + 0.150(0.100)NRB n = 15; R = 0.8953; Radj = 0.8498; sd = 0.248; PRESS = 1.078; MSE =0.0615; F = 11; p < 0.0001; VIF < 3.6; PRESScv = 1.078; MSEcv = 0.0615 |
| (8) | log KHSA = 4.843(0.351) + 0.204(0.097)log KAM +0.109(0.190)HBD − 0.415(0.148)HBA + 0.175(0.108)NRB n = 15; R = 0.8722; Radj = 0.8155; sd = 0.272; PRESS = 1.326; MSE =0.0741; F = 8; p < 0.0001; VIF < 3.5; PRESScv = 1.326; MSEcv = 0.0741 |
| (9) | log BB = −0.375(0.078) + 0.254(0.021)log km − 0.454(0.038)HBD + 0.041(0.030)HBA − 0.006(0.023)NRB n = 15; R = 0.9836; Radj = 0.9770; sd = 0.055; PRESS = 0.090; MSE = 0.0031; F = 75; p < 0.0001; VIF < 3.6; PRESScv = 0.090; MSEcv = 0.0031 |
| (10) | log BB = −0.205(0.073) + 0.242(0.020) log KAM − 0.479(0.039) HBD + 0.032(0.031) HBA − 0.003(0.022) NRB n = 15; R = 0.9831; Radj = 0.9762; sd = 0.056; PRESS = 0.069; MSE = 0,0032; F = 72; p < 0.0001; VIF < 3,5; PRESScv = 0,069; MSEcv = 0.0032 |
| (11) | LD50= 2870(411) − 253(109) log km + 650(199) HBD − 194(160) HBA − 245(116) NRB n = 15; R = 0.9330; Radj = 0.9048; sd = 291; PRESS = 2078292; MSE = 84613; F = 17; p < 0.0001; VIF < 3.6; PRESScv = 2078292; MSEcv = 76083 |
| (12) | LD50= 2714(367) − 251(102) log KAM +682(192) HBD − 192(155) HBA − 242(113) NRB n = 15; R = 0.9358; Radj = 0.9088; sd = 285; PRESS = 2017628; MSE = 81170; F = 18; p < 0.0001; VIF < 3.5; PRESScv = 2017628; MSEcv = 74652 |
4. Materials and Methods
4.1. Chromatographic Measurements
4.2. In Silico Parameters
4.3. Statistics
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Li, Q.; Lu, Y.; Wang, P.; Wang, T.; Suriyanarayanan, S.; Liang, R.; Baninla, Y.; Khan, K. Distribution, source, and risk of organochlorine pesticides (OCPs) and polychlorinated biphenyls (PCBs) in urban and rural soils around the Yellow and Bohai Seas, China. Environ. Pollut. 2018, 239, 233–241. [Google Scholar] [CrossRef] [PubMed]
- Syafrudin, M.; Kristanti, R.A.; Yuniarto, A.; Hadibarata, T.; Rhee, J.; Al-onazi, W.A.; Algarni, T.S.; Almarri, A.H.; Al-Mohaimeed, A.M. Pesticides in Drinking Water—A Review. Int. J. Environ. Res. Public Health 2021, 18, 468. [Google Scholar] [CrossRef] [PubMed]
- Umetsu, N.; Shirai, Y. Development of novel pesticides in the 21st century. J. Pestic. Sci. 2020, 45, 54–74. [Google Scholar] [CrossRef]
- Cimmino, A.; Zonno, M.C.; Andolfi, A.; Troise, C.; Motta, A.; Vurro, M.; Evidente, A. Agropyrenol, a phytotoxic fungal metabolite, and its derivatives: A structure−activity relationship study. J. Agric. Food Chem. 2013, 61, 1779−1783. [Google Scholar] [CrossRef]
- Toropov, A.A.; Toropova, A.P.; Marzo, M.; Dorne, J.L.; Georgiadis, N.; Benfenati, E. QSAR models for predicting acute toxicity of pesticides in rainbow trout using the CORAL software and EFSA’s OpenFoodTox database. Environ. Tox. Pharm. 2017, 53, 158–163. [Google Scholar] [CrossRef] [PubMed]
- Kaliszan, R. QSRR Quantitative structure−(chromatographic) retention relationships. Chem. Rev. 2007, 107, 3212−3246. [Google Scholar] [CrossRef] [PubMed]
- Kempińska, D.; Chmiel, T.; Kot-Wasik, A.; Mróz, A.; Mazerska, Z.; Namieśnik, J. State of the art and prospects of methods for determination of lipophilicity of chemical compounds. Trends Anal. Chem. 2019, 113, 54–73. [Google Scholar] [CrossRef]
- Héberger, K. Quantitative structure-(chromatographic) retention relationships. J. Chromatogr. A 2007, 1158, 273–305. [Google Scholar] [CrossRef]
- Andrić, F.; Héberger, K. Towards better understanding of lipophilicity: Assessment of in silico and chromatographic log P measures for pharmaceutically important compounds by nonparametric rankings. J. Pharm. Biomed. Anal. 2015, 115, 183–191. [Google Scholar] [CrossRef]
- Valko, K.; Nunhuck, S.; Bevan, C.; Abraham, M.H.; Reynolds, D.P. Fast Gradient HPLC Method to Determine Compounds Binding to Human Serum Albumin. Relationships with Octanol/Water and Immobilized Artificial Membrane Lipophilicity. J. Pharm. Sci. 2003, 92, 2236–2248. [Google Scholar] [CrossRef]
- Valko, K. Lipophilicity and biomimetic properties measured by HPLC to support drug discovery. J. Pharm. Biomed. Anal. 2016, 130, 35–54. [Google Scholar] [CrossRef] [PubMed]
- Ciura, K.; Dziomba, S.Z. Application of separation methods for In Vitro prediction of blood–brain barrier permeability—The state of the art. J. Pharm. Biomed. Anal. 2020, 177, 112891. [Google Scholar] [CrossRef] [PubMed]
- Milošević, N.; Janjić, N.; Milić, N.; Milanović, M.; Popović, J.; Antonowić, D. Pharmacokinetics and Toxicity Predictors of New s-Triazines, Herbicide Candidates, in Correlation with Chromatographic Retention Constants. J. Agr. Food Chem. 2014, 62, 8579–8585. [Google Scholar] [CrossRef] [PubMed]
- Tsopelas, F.; Giaginis, C.; Tsantili-Kakoulidou, A. Lipophilicity and biomimetic properties to support drug discovery. Expert Opin. Drug Discov. 2017, 12, 885–896. [Google Scholar] [CrossRef]
- Russo, G.; Grumetto, L.; Szucs, R.; Barbato, F.; Lynen, F. Screening therapeutics according to their uptake across the blood-brain barrier: A high throughput method based on immobilized artificial membrane liquid chromatography-diode-array-detection coupled to electrospray-time-of-flight mass spectrometry. Eur. J. Pharm. Biopharm. 2018, 127, 72–84. [Google Scholar] [CrossRef]
- Stergiopoulos, C.; Tsopelas, F.; Valko, K.; Ochsenkühn-Petropoulou, M. The use of biomimetic chromatography to predict acute aquatic toxicity of pharmaceutical compounds. Toxicol. Environ. Chem. 2021. [Google Scholar] [CrossRef]
- Escuder-Gilabert, L.; Molero-Monfort, M.; Villanueva-Camaňas, R.M.; Sagrado, S.; Medina-Hernández, M.J. Potential of biopartitioning micellar chromatography as an In Vitro technique for predicting drug penetration across the blood-brain barrier. J. Chromatogr. B Analyt. Technol. Biomed. Life Sci. 2004, 807, 193–201. [Google Scholar] [CrossRef]
- Riiz-Ángel, M.; Garcia-Álvarez-Coque, M.; Berthod, A. New Insights and Recent Developments in Micellar Liquid Chromatography. Sep. Pur. Rev. 2009, 38, 45–96. [Google Scholar] [CrossRef]
- Rambla-Alegre, M. Basic Principles of MLC. Chromatogr. Res. Int. 2012, 2012, 898520. [Google Scholar] [CrossRef]
- Kalyankar, T.M.; Kulkarni, P.D.; Wadher, S.J.; Pekamwar, S.S. Applications of Micellar Liquid Chromatography in Bioanalysis: A Review. J. App. Pharm. Sci. 2014, 4, 128–134. [Google Scholar] [CrossRef]
- Tsopelas, F.; Danias, P.; Pappa, A.; Tsantili-Kakoulidou, A. Biopartitioning micellar chromatography under different conditions: Insight into the retention mechanism and the potential to model biological processes. J. Chromatogr. A 2020, 1621, 461027. [Google Scholar] [CrossRef] [PubMed]
- Foley, J.P. Critical compilation of solute-micelle binding constants and related parameters from micellar liquid chromatographic measurements. Anal. Chim. Acta 1990, 231, 237–247. [Google Scholar] [CrossRef]
- Janicka, M. Correlations between Chromatographic Parameters and Bioactivity Predictors of Potential Herbicides. J. Chrom. Sci. 2014, 52, 676–684. [Google Scholar] [CrossRef][Green Version]
- Leeson, P.D. Molecular inflation, attrition and the rule of five. Adv. Drug Deliv. Rev. 2016, 101, 22–33. [Google Scholar] [CrossRef] [PubMed]
- Toutain, P.L.; Bousquet-Mélou, A. Volumes of distribution. J. Vet. Pharmacol. Therap. 2004, 27, 441–453. [Google Scholar] [CrossRef]
- Huang, L.; Wells, M.C.; Zhao, Z. A Practical Perspective on the evaluation of small Molecule CNS Penetration in Drug Discovery. Drug Metabol. Lett. 2019, 13, 78–94. [Google Scholar] [CrossRef]
- Norinder, U.; Haberlein, M. Computational approaches to the prediction of the blood brain distribution. Adv. Drug Deliv. Rev. 2002, 54, 291–313. [Google Scholar] [CrossRef]
- Clark, D.E. In Silico prediction of blood-brain barrier permeation. Drug Discov. Today 2003, 8, 927–933. [Google Scholar] [CrossRef]
- Geldenhuys, W.J.; Mohammad, A.S.; Adkins, C.E.; Lockman, P.R. Molecular determinants of blood–brain barrier permeation. Ther. Deliv. 2015, 6, 961–971. [Google Scholar] [CrossRef]
- Prausnitz, M.R.; Langer, R. Transdermal drug delivery. Nat. Biotechnol. 2008, 26, 1261−1268. [Google Scholar] [CrossRef]
- Chen, C.-P.; Chen, C.-C.; Huang, C.-W.; Chang, Y.-C. Evaluating Molecular Properties involved in Transport of Small Molecules in Stratum Corneum: A Quantitative Structure-Activity Relationship for Skin Permeability. Molecules 2018, 23, 911. [Google Scholar] [CrossRef] [PubMed]
- Aggarwal, M.; Fisher, P.; Hȕser, A.; Kluxen, F.M.; Parr-Dobrzanski, R.; Soufi, M.; Strupp, C.; Wiemann, C.; Billington, R. Assessment of an extended dataset of in vitro human dermal absorption studies on pesticides to determine default values, opportunities for read-across and influence of dilution on absorption. Reg. Tox. Pharmacol. 2015, 72, 58–70. [Google Scholar] [CrossRef] [PubMed]
- Kerns, E.; Di, L. Pharmaceutical profiling in drug discovery. Drug Disc. Today 2003, 8, 316–323. [Google Scholar] [CrossRef]
- Platts, J.A.; Abraham, M.H.; Hersey, A.; Butina, D. Estimation of Molecular Linear Free Energy Relationship Descriptors. 4. Correlation and Prediction of Cell Permeation. Pharm. Res. 2000, 17, 1013–1018. [Google Scholar] [CrossRef]
- Carter, D.C.; Ho, J.X. Strucyure of Serum Albumin. Adv. Protein Chem. 1994, 45, 153–176. [Google Scholar] [CrossRef]
- Kratochwill, N.; Huber, W.; Muller, F.; Kansy, M.; Gerber, P.R. Predicting plasma protein binding of drugs-revisited. Curr. Opin. Drug Discov. Dev. 2004, 4, 507–512. [Google Scholar]
- Hollósy, F.; Valkó, K.; Hersey, A.; Nunhuck, S.; Kéri, G.; Bevan, C. Estimation of Volume of Distribution in Humans from High Throughput HPLC-Based Measurements of Human Serum Albumin Binding and Immobilized Artificial Membrane Partitioning. J. Med. Chem. 2006, 49, 6958–6971. [Google Scholar] [CrossRef]
- Baur, P.; Marzouk, H.; Schőnherr, J.; Grayson, B.T. Partition Coefficients of Active Ingredients between Plant Cuticle and Adjuvants as Related to Rates of Foliar Uptake. J. Agric. Food. Chem. 1997, 45, 3659–3665. [Google Scholar] [CrossRef]
- Karlsson, J.O.G. Preclinical safety assessment of contrast media: Predictive value. Eur. Radiol. 1996, 6, S3–S7. [Google Scholar] [CrossRef]
- Quiñones-Torrelo, C.; Sagrado-Vives, S.; Villanueva-Camañas, R.M.; Medina-Hernández, M.J. An LD50 model for predicting psychotropic drug toxicity using biopartitioning micellar chromatography. Biomed. Chromatogr. 2001, 15, 31–40. [Google Scholar] [CrossRef]
- Wang, J.; Skolnik, S. Recent Advances in Physicochemical and ADMET Profiling in Drug Discovery. Chem. Biodiver. 2009, 6, 1887–1899. [Google Scholar] [CrossRef] [PubMed]
- Rácz, A.; Bajusz, D.; Miranda-Quintana, R.A.; Héberger, K. Machine learning models for classification task related to drug safety. Mol. Diver. 2021, 25, 1409–1424. [Google Scholar] [CrossRef]
- Lipinski, C.A.; Lombardo, F.; Dominy, B.W.; Feeney, P.J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug Deliv. Rev. 2001, 46, 3–26. [Google Scholar] [CrossRef]
- Lipinski, C.A. Rule of five in 2015 and beyond: Target and ligand structural limitations, ligand chemistry structure and drug discovery project decisions. Adv. Drug. Deliv. Rev. 2016, 101, 34–41. [Google Scholar] [CrossRef] [PubMed]
- Gramatica, P. On the development and Validation of QSAR Models. In Computational Toxicology; Reisfeld, B., Mayeno, A.N., Eds.; Springer: Berlin/Heidelberg, Germany; Humana Press: Totowa, NJ, USA, 2013; Volume II. [Google Scholar] [CrossRef]











| No | Group | Structure | log km | log KAM |
|---|---|---|---|---|
| 1 | 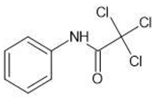 | 2.00 | 1.76 | |
| 2 |  | 2.46 | 2.13 | |
| 3 | 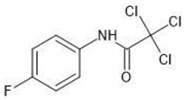 | 2.10 | 1.80 | |
| 4 |  | 2.72 | 2.31 | |
| 5 | I |  | 2.40 | 1.98 |
| 6 |  | 3.20 | 2.60 | |
| 7 |  | 2.96 | 2.39 | |
| 8 | 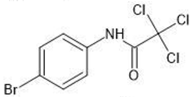 | 2.92 | 2.46 | |
| 9 |  | 0.40 | −0.48 | |
| 10 |  | 2.00 | 1.76 | |
| 11 | 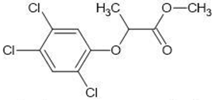 | 2.95 | 2.39 | |
| 12 | 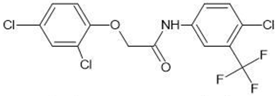 | 3.90 | 3.40 | |
| 13 | II |  | 4.30 | 4.00 |
| 14 |  | 1.60 | 1.23 | |
| 15 |  | 2.71 | 2.28 |
| Nr | log P | HBD | HBA | NRB | MW [g/mol] | TPSA [A2] | Vd [L/kg] | fu, plasma | fu, brain | log BB | log Kp | log Ksc | log Kw/cell | log Pw/HSA | log KHSA | log Pw/pc | LD50 [mg/kg] |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 2.285 | 1 | 2 | 2 | 238.50 | 29.10 | 2.1 | 0.0750 | 0.06 | −0.218 | −5.095 | 1.713 | −4.988 | 0.280 | 4.58 | 2.513 | 2600 |
| 2 | 2.937 | 1 | 2 | 2 | 252.52 | 29.10 | 2.3 | 0.0830 | 0.04 | −0.026 | −4.747 | 1.996 | −4.833 | 0.554 | 4.60 | 3.106 | 2000 |
| 3 | 2.309 | 1 | 2 | 2 | 256.49 | 29.10 | 2.1 | 0.1000 | 0.06 | −0.275 | −5.075 | 1.717 | −5.164 | 0.253 | 4.86 | 2.504 | 1800 |
| 4 | 2.971 | 1 | 2 | 2 | 272.94 | 29.10 | 2.4 | 0.0370 | 0.02 | −0.098 | −4.710 | 2.100 | −5.071 | 0.694 | 4.70 | 3.254 | 1900 |
| 5 | 2.810 | 1 | 2 | 2 | 272.94 | 29.10 | 2.1 | 0.0510 | 0.03 | −0.110 | −4.836 | 1.989 | −4.951 | 0.576 | 5.11 | 3.077 | 2000 |
| 6 | 3.755 | 1 | 2 | 2 | 307.39 | 29.10 | 2.8 | 0.0140 | 0.01 | 0.056 | −4.244 | 2.529 | −4.944 | 1.132 | 4.97 | 4.092 | 1800 |
| 7 | 3.488 | 1 | 2 | 2 | 307.39 | 29.10 | 2.4 | 0.0320 | 0.01 | −0.011 | −4.434 | 2.390 | −5.020 | 0.993 | 5.53 | 3.847 | 2300 |
| 8 | 2.980 | 1 | 2 | 2 | 317.39 | 29.10 | 2.4 | 0.0540 | 0.02 | −0.094 | −4.818 | 2.134 | −5.197 | 0.804 | 4.93 | 3.331 | 1400 |
| 9 | 1.345 | 0 | 3 | 3 | 232.49 | 29.54 | 1.5 | 0.3100 | 0.30 | −0.225 | −5.538 | 1.097 | −3.947 | −0.492 | 4.17 | 1.522 | 1600 |
| 10 | 2.917 | 0 | 3 | 4 | 235.06 | 35.53 | 1.9 | 0.0920 | 0.07 | 0.263 | −4.431 | 1.923 | −3.279 | 0.313 | 4.36 | 3.132 | 550 |
| 11 | 4.067 | 0 | 3 | 4 | 283.53 | 35.53 | 2.1 | 0.0610 | 0.02 | 0.513 | −3.731 | 2.541 | −3.081 | 0.904 | 4.94 | 4.400 | 580 |
| 12 | 4.936 | 1 | 3 | 5 | 398.59 | 38.33 | 3.8 | 0.0030 | 0.01 | 0.227 | −3.741 | 3.107 | −5.275 | 1.642 | 5.35 | 5.420 | 780 |
| 13 | 5.386 | 1 | 3 | 5 | 412.62 | 38.33 | 4.0 | 0.0029 | 0.01 | 0.345 | −3.504 | 3.308 | −5.213 | 1.823 | 5.39 | 5.858 | 720 |
| 14 | 2.600 | 1 | 5 | 5 | 347.24 | 50.80 | 1.6 | 0.2400 | 0.12 | −0.229 | −5.761 | 1.608 | −5.822 | 0.200 | 3.98 | 2.765 | 840 |
| 15 | 3.807 | 0 | 4 | 3 | 292.76 | 44.12 | 2.1 | 0.0690 | 0.02 | 0.448 | −4.540 | 2.386 | −3.999 | 1.017 | 4.17 | 4.240 | 750 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Janicka, M.; Śliwińska, A. Quantitative Retention (Structure)–Activity Relationships in Predicting the Pharmaceutical and Toxic Properties of Potential Pesticides. Molecules 2022, 27, 3599. https://doi.org/10.3390/molecules27113599
Janicka M, Śliwińska A. Quantitative Retention (Structure)–Activity Relationships in Predicting the Pharmaceutical and Toxic Properties of Potential Pesticides. Molecules. 2022; 27(11):3599. https://doi.org/10.3390/molecules27113599
Chicago/Turabian StyleJanicka, Małgorzata, and Anna Śliwińska. 2022. "Quantitative Retention (Structure)–Activity Relationships in Predicting the Pharmaceutical and Toxic Properties of Potential Pesticides" Molecules 27, no. 11: 3599. https://doi.org/10.3390/molecules27113599
APA StyleJanicka, M., & Śliwińska, A. (2022). Quantitative Retention (Structure)–Activity Relationships in Predicting the Pharmaceutical and Toxic Properties of Potential Pesticides. Molecules, 27(11), 3599. https://doi.org/10.3390/molecules27113599







