Applications and Restrictions of Integrated Genomic and Metabolomic Screening: An Accelerator for Drug Discovery from Actinomycetes?
Abstract
:1. Introduction
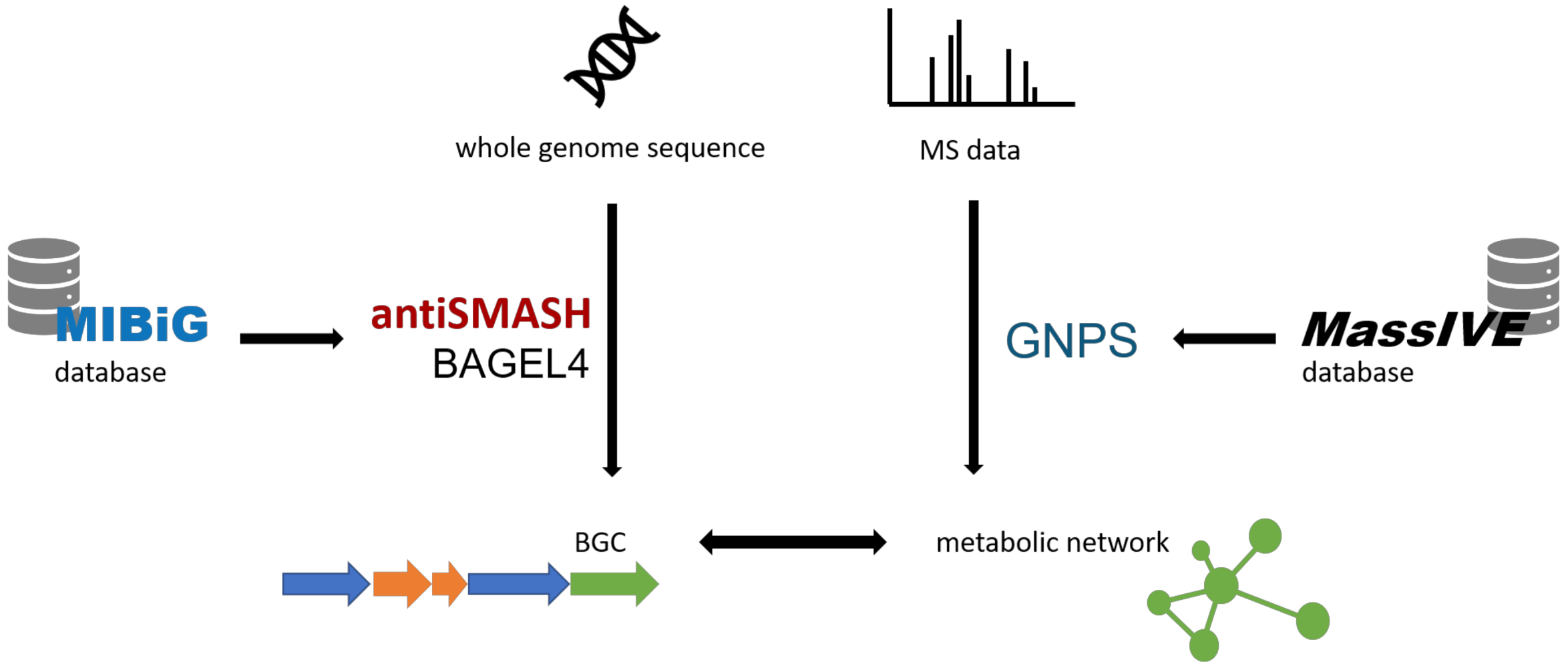
1.1. Bioinformatic Tools for Genome Mining
1.2. Bioinformatic Tools for Metabolomic Analysis
2. Applications of Genomic and Metabolomic Screening
2.1. Discovery of New Analogs
2.2. Exploring the Productive Spectrum
2.3. Elucidation of Biosynthesis
3. Conclusions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Takahashi, Y.; Nakashima, T. Actinomycetes, an inexhaustible source of naturally occurring antibiotics. Antibiotics 2018, 7, 45. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bérdy, J. Thoughts and facts about antibiotics: Where we are now and where we are heading. J. Antibiot. 2012, 65, 385–395. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wibowo, J.T.; Kellermann, M.Y.; Köck, M.; Putra, M.Y.; Murniasih, T.; Mohr, K.I.; Wink, J.; Praditya, D.F.; Steinmann, E.; Schupp, P.J. Anti-infective and antiviral activity of valinomycin and its analogues from a sea cucumber-associated bacterium, Streptomyces Sp. SV 21. Mar. Drugs 2021, 19, 81. [Google Scholar] [CrossRef] [PubMed]
- Běhal, V. Bioactive products from Streptomyces. In Advances in Applied Microbiology; Academic Press Inc.: Cambridge, MA, USA, 2000; pp. 113–156. [Google Scholar] [CrossRef]
- Hutchings, M.; Truman, A.; Wilkinson, B. Antibiotics: Past, present and future. In Current Opinion in Microbiology; Elsevier Ltd.: Amsterdam, The Netherlands, 2019; pp. 72–80. [Google Scholar] [CrossRef]
- Katz, L.; Baltz, R.H. Natural product discovery: Past, present, and future. J. Ind. Microbiol. Biotechnol. 2016, 43, 155–176. [Google Scholar] [CrossRef] [PubMed]
- Baltz, R.H. Natural product drug discovery in the genomic era: Realities, conjectures, misconceptions, and opportunities. J. Ind. Microbiol. Biotechnol. 2018, 46, 1–19. [Google Scholar] [CrossRef] [PubMed]
- Traxler, M.F.; Kolter, R. Natural products in soil microbe interactions and evolution. Nat. Prod. Rep. 2015, 32, 956–970. [Google Scholar] [CrossRef]
- Kenshole, E.; Herisse, M.; Michael, M.; Pidot, S.J. Natural product discovery through microbial genome mining. Curr. Opin. Chem. Biol. 2021, 60, 47–54. [Google Scholar] [CrossRef]
- Niu, G. Genomics-driven natural product discovery in actinomycetes. Trends Biotechnol. 2018, 36, 238–241. [Google Scholar] [CrossRef]
- Blin, K.; Shaw, S.; Steinke, K.; Villebro, R.; Ziemert, N.; Lee, S.Y.; Medema, M.H.; Weber, T. AntiSMASH 5.0: Updates to the secondary metabolite genome mining pipeline. Nucleic Acids Res. 2019, 47, W81–W87. [Google Scholar] [CrossRef] [Green Version]
- Van Heel, A.J.; de Jong, A.; Song, C.; Viel, J.H.; Kok, J.; Kuipers, O.P. BAGEL4: A user-friendly web server to thoroughly mine RiPPs and bacteriocins. Nucleic Acids Res. 2018, 46, W278–W281. [Google Scholar] [CrossRef]
- Van Heel, A.J.; De Jong, A.; Montalbá N-Ló Pez, M.; Kok, J.; Kuipers, O.P. BAGEL3: Automated identification of genes encoding bacteriocins and (non-)bactericidal posttranslationally modified peptides. Nucleic Acids Res. 2013, 41, W448–W453. [Google Scholar] [CrossRef] [PubMed]
- Kautsar, S.A.; Blin, K.; Shaw, S.; Navarro-Muñoz, J.C.; Terlouw, B.R.; Van Der Hooft, J.J.J.; Van Santen, J.A.; Tracanna, V.; Suarez Duran, H.G.; Pascal Andreu, V.; et al. MIBiG 2.0: A repository for biosynthetic gene clusters of known function. Nucleic Acids Res. 2020, 48, D454–D458. [Google Scholar] [CrossRef] [Green Version]
- Wang, M.; Carver, J.J.; Phelan, V.V.; Sanchez, L.M.; Garg, N.; Peng, Y.; Nguyen, D.D.; Watrous, J.; Kapono, C.A.; Luzzatto-Knaan, T.; et al. Sharing and community curation of mass spectrometry data with global natural products social molecular networking. Nat. Biotechnol. 2016, 34, 828–837. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Aron, A.T.; Gentry, E.C.; McPhail, K.L.; Nothias, L.F.; Nothias-Esposito, M.; Bouslimani, A.; Petras, D.; Gauglitz, J.M.; Sikora, N.; Vargas, F.; et al. Reproducible molecular networking of untargeted mass spectrometry data using GNPS. Nat. Protoc. 2020, 15, 1954–1991. [Google Scholar] [CrossRef]
- Blin, K.; Wolf, T.; Chevrette, M.G.; Lu, X.; Schwalen, C.J.; Kautsar, S.A.; Suarez Duran, H.G.; de Los Santos, E.L.C.; Kim, H.U.; Nave, M.; et al. AntiSMASH 4.0—Improvements in chemistry prediction and gene cluster boundary identification. Nucleic Acids Res. 2017, 45, W36–W41. [Google Scholar] [CrossRef]
- Walsh, C.; Tang, Y. Natural product biosynthesis: Chemical logic and enzymatic machinery. Chem. Rev. 2017, 117, 5226–5333. [Google Scholar]
- Van Agthoven, M.A.; Lam, Y.P.Y.; O’Connor, P.B.; Rolando, C.; Delsuc, M.A. Two-dimensional mass spectrometry: New perspectives for tandem mass spectrometry. Eur. Biophys. J. 2019, 48, 213–229. [Google Scholar] [CrossRef] [Green Version]
- Mohimani, H.; Gurevich, A.; Shlemov, A.; Mikheenko, A.; Korobeynikov, A.; Cao, L.; Shcherbin, E.; Nothias, L.F.; Dorrestein, P.C.; Pevzner, P.A. Dereplication of microbial metabolites through database search of mass spectra. Nat. Commun. 2018, 9, 1–12. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gurevich, A.; Mikheenko, A.; Shlemov, A.; Korobeynikov, A.; Mohimani, H.; Pevzner, P.A. Increased diversity of peptidic natural products revealed by modification-tolerant database search of mass spectra. Nat. Microbiol. 2018, 3, 319–327. [Google Scholar] [CrossRef] [PubMed]
- Van Der Hooft, J.J.J.; Mohimani, H.; Bauermeister, A.; Dorrestein, P.C.; Duncan, K.R.; Medema, M.H. Linking genomics and metabolomics to chart specialized metabolic diversity. Chem. Soc. Rev. 2020, 49, 3297–3314. [Google Scholar] [CrossRef]
- AbuSara, N.F.; Piercey, B.M.; Moore, M.A.; Shaikh, A.A.; Nothias, L.F.; Srivastava, S.K.; Cruz-Morales, P.; Dorrestein, P.C.; Barona-Gómez, F.; Tahlan, K. Comparative genomics and metabolomics analyses of clavulanic acid-producing Streptomyces species provides insight into specialized metabolism. Front. Microbiol. 2019, 10, 2550. [Google Scholar] [CrossRef]
- Duncan, K.R.; Crüsemann, M.; Lechner, A.; Sarkar, A.; Li, J.; Ziemert, N.; Wang, M.; Bandeira, N.; Moore, B.S.; Dorrestein, P.C.; et al. Molecular networking and pattern-based genome mining improves discovery of biosynthetic gene clusters and their products from Salinispora species. Chem. Biol. 2015, 22, 460–471. [Google Scholar] [CrossRef] [Green Version]
- Liu, L.-L.; Chen, Z.-F.; Liu, Y.; Tang, D.; Gao, H.-H.; Zhang, Q.; Gao, J.-M. Molecular networking-based for the target discovery of potent antiproliferative polycyclic macrolactam ansamycins from Streptomyces Cacaoi subsp. asoensis. Org. Chem. Front. 2020, 7, 4008–4018. [Google Scholar] [CrossRef]
- Sigrist, R.; Paulo, B.S.; Angolini, C.F.F.; De Oliveira, L.G. Mass spectrometry-guided genome mining as a tool to uncover novel natural products. J. Vis. Exp. 2020, 2020, e60825. [Google Scholar] [CrossRef]
- Liu, W.T.; Lamsa, A.; Wong, W.R.; Boudreau, P.D.; Kersten, R.; Peng, Y.; Moree, W.J.; Duggan, B.M.; Moore, B.S.; Gerwick, W.H.; et al. MS/MS-based networking and peptidogenomics guided genome mining revealed the stenothricin gene cluster in Streptomyces roseosporus. J. Antibiot. 2014, 67, 99–104. [Google Scholar] [CrossRef] [Green Version]
- Schimana, J.; Gebhardt, K.; Höltzel, A.; Schmid, D.G.; Süssmuth, R.; Müller, J.; Pukall, R.; Fiedler, H.-P. Arylomycins A and B, new biaryl-bridged lipopeptide antibiotics produced by Streptomyces Sp. tue 6075. I. Taxonomy, fermentation, isolation and biological activities. J. Antibiot. 2002, 55, 565–570. [Google Scholar] [CrossRef] [Green Version]
- Chatterjee, S.; Nadkarni, S.R.; Vijayakumar, E.K.S.; Patel, M.V.; Gangul, B.N.; Fehlhaber, H.-W.; Vertesy, L. Napsamycins, new Pseudomonas active antibiotics of the mureidomycin family from Streptomyces sp. HIL Y-82, 11372. J. Antibiot. 1994, 47, 595–598. [Google Scholar] [CrossRef] [Green Version]
- Rinken, M.; Lehmann, W.D.; König, W.A. Die Struktur von stenothricin—Korrektur eines früheren strukturvorschlags. Liebigs Ann. Chem. 1984, 1984, 1672–1684. [Google Scholar] [CrossRef]
- Paulus, C.; Rebets, Y.; Tokovenko, B.; Nadmid, S.; Terekhova, L.P.; Myronovskyi, M.; Zotchev, S.B.; Rückert, C.; Braig, S.; Zahler, S.; et al. New natural products identified by combined genomics-metabolomics profiling of marine Streptomyces Sp. MP131-18. Sci. Rep. 2017, 7, 1–11. [Google Scholar] [CrossRef]
- Jensen, P.R.; Moore, B.S.; Fenical, W. The marine actinomycete genus Salinispora: A model organism for secondary metabolite discovery. Nat. Prod. Rep. 2015, 32, 738–751. [Google Scholar] [CrossRef] [Green Version]
- Renner, M.K.; Shen, Y.C.; Cheng, X.C.; Jensen, P.R.; Frankmoelle, W.; Kauffman, C.A.; Fenical, W.; Lobkovsky, E.; Clardy, J. Cyclomarins A-C, new antiinflammatory cyclic peptides produced by a marine bacterium (Streptomyces Sp.). J. Am. Chem. Soc. 1999, 121, 11273–11276. [Google Scholar] [CrossRef]
- Gontang, E.A.; Fenical, W.; Jensen, P.R. Phylogenetic diversity of gram-negative positive bacteria cultured from marine sediments. Appl. Environ. Microbiol. 2007, 73, 3272–3282. [Google Scholar] [CrossRef] [Green Version]
- Williams, P.G.; Miller, E.D.; Asolkar, R.N.; Jensen, P.R.; Fenical, W. Arenicolides A-C, 26-Membered ring macrolides from the marine actinomycete Salinispora Arenicola. J. Org. Chem. 2007, 72, 5025–5034. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, Y.; Liu, Y.; Tang, F.; Bernot, K.M.; Schore, R.; Marcucci, G.; Caligiuri, M.A.; Zheng, P.; Liu, Y. echinomycin protects mice against relapsed acute myeloid leukemia without adverse effect on hematopoietic stem cells. Blood 2014, 124, 1127–1135. [Google Scholar] [CrossRef] [Green Version]
- McArthur, K.A.; Mitchell, S.S.; Tsueng, G.; Rheingold, A.; White, D.J.; Grodberg, J.; Lam, K.S.; Potts, B.C.M. Lynamicins A-E, chlorinated bisindole pyrrole antibiotics from a novel marine actinomycete. J. Nat. Prod. 2008, 71, 1732–1737. [Google Scholar] [CrossRef]
- Zhang, W.; Liu, Z.; Li, S.; Yang, T.; Zhang, Q.; Ma, L.; Tian, X.; Zhang, H.; Huang, C.; Zhang, S.; et al. Spiroindimicins A-D: New bisindole alkaloids from a deep-sea-derived actinomycete. Org. Lett. 2012, 14, 3364–3367. [Google Scholar] [CrossRef]
- Lindel, T.; Jensen, P.R.; Fenical, W. Lagunapyrones A-C: Cytotoxic acetogenins of a new skeletal class from a marine sediment bacterium. Tetrahedron Lett. 1996, 37, 1327–1330. [Google Scholar] [CrossRef]
- Ishaque, N.M.; Burgsdorf, I.; Limlingan Malit, J.J.; Saha, S.; Teta, R.; Ewe, D.; Kannabiran, K.; Hrouzek, P.; Steindler, L.; Costantino, V.; et al. Isolation, genomic and metabolomic characterization of Streptomyces Tendae VITAKN with quorum sensing inhibitory activity from Southern India. Microorganisms 2020, 8, 121. [Google Scholar] [CrossRef] [Green Version]
- Borthwick, A.D. 2,5-Diketopiperazines: Synthesis, reactions, medicinal chemistry, and bioactive natural products. Chem. Rev. 2012, 112, 3641–3716. [Google Scholar] [CrossRef]
- Yao, T.; Liu, J.; Liu, Z.; Li, T.; Li, H.; Che, Q.; Zhu, T.; Li, D.; Gu, Q.; Li, W. Genome mining of cyclodipeptide synthases unravels unusual TRNA-dependent diketopiperazine-terpene biosynthetic machinery. Nat. Commun. 2018, 9, 1–12. [Google Scholar] [CrossRef]
- Reading, C.; Cole, M. Clavulanic acid: A beta lactamase inhibiting beta lactam from Streptomyces Clavuligerus. Antimicrob. Agents Chemother. 1977, 11, 852–857. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Keberle, H. The biochemistry of desferrioxamine and its relation to iron metabolism. Ann. N. Y. Acad. Sci. 1964, 119, 758–768. [Google Scholar] [CrossRef]
- Galinski, E.A.; Pfeiffer, H.-P.; Truper, H.G. 1,4,5,6-tetrahydro-2-methyl-4-pyrimidinecarboxylic acid. A novel cyclic amino acid from halophilic phototrophic bacteria of the genus Ectothiorhodospira. Eur. J. Biochem. 1985, 149, 135–139. [Google Scholar] [CrossRef] [PubMed]
- Oliva, B.; O’Neill, A.; Wilson, J.M.; O’Hanlon, P.J.; Chopra, I. Antimicrobial properties and mode of action of the pyrrothine holomycin. Antimicrob. Agents Chemother. 2001, 45, 532–539. [Google Scholar] [CrossRef] [Green Version]
- Lauinger, L.; Li, J.; Shostak, A.; Cemel, I.A.; Ha, N.; Zhang, Y.; Merkl, P.E.; Obermeyer, S.; Stankovic-Valentin, N.; Schafmeier, T.; et al. Thiolutin is a zinc chelator that inhibits the Rpn11 and other JAMM metalloproteases. Nat. Chem. Biol. 2017, 13, 709–714. [Google Scholar] [CrossRef] [Green Version]
- Dillman, R.O. Pentostatin (Nipent®) in the treatment of chronic lymphocyte leukemia and hairy cell leukemia. Expert Rev. Anticancer Ther. 2004, 4, 27–36. [Google Scholar] [CrossRef]
- Zeng, W.; Jin, L.; Zhang, F.; Zhang, C.; Liang, W. Naringenin as a potential immunomodulator in therapeutics. Pharmacol. Res. 2018, 135, 122–126. [Google Scholar] [CrossRef]
- Tahlan, K.; Anders, C.; Wong, A.; Mosher, R.H.; Beatty, P.H.; Brumlik, M.J.; Griffin, A.; Hughes, C.; Griffin, J.; Barton, B.; et al. 5S clavam biosynthetic genes are located in both the clavam and paralog gene clusters in Streptomyces Clavuligerus. Chem. Biol. 2007, 14, 131–142. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Newton, G.G.F.; Abraham, E.P. Cephalosporin C, a new antibiotic containing sulphur and D-α-aminoadipic acid. Nature 1955, 175, 548. [Google Scholar] [CrossRef]
- Romero, J.; Liras, P.; Martín, J.F. Dissociation of cephamycin and clavulanic acid biosynthesis in streptomyces clavuligerus. Appl. Microbiol. Biotechnol. 1984, 20, 318–325. [Google Scholar] [CrossRef]
- Handayani, I.; Saad, H.; Ratnakomala, S.; Lisdiyanti, P.; Kusharyoto, W.; Krause, J.; Kulik, A.; Wohlleben, W.; Aziz, S.; Gross, H.; et al. Mining indonesian microbial biodiversity for novel natural compounds by a combined genome mining and molecular networking approach. Mar. Drugs 2021, 19, 316. [Google Scholar] [CrossRef] [PubMed]
- Gosse, J.T.; Ghosh, S.; Sproule, A.; Overy, D.; Cheeptham, N.; Boddy, C.N. Whole genome sequencing and metabolomic study of cave Streptomyces isolates ICC1 and ICC4. Front. Microbiol. 2019, 10, 1020. [Google Scholar] [CrossRef] [PubMed]
- Gourdeau, H.; McAlpine, J.B.; Ranger, M.; Simard, B.; Berger, F.; Beaudry, F.; Falardeau, P. Identification, characterization and potent antitumor activity of ECO-4601, a novel peripheral benzodiazepine receptor ligand. Cancer Chemother. Pharmacol. 2008, 61, 911–921. [Google Scholar] [CrossRef] [PubMed]
- McAlpine, J.B.; Banskota, A.H.; Charan, R.D.; Schlingmann, G.; Zazopoulos, E.; Piraee, M.; Janso, J.; Bernan, V.S.; Aouidate, M.; Farnet, C.M.; et al. Biosynthesis of diazepinomicin/ECO-4601, a Micromonospora secondary metabolite with a novel ring system. J. Nat. Prod. 2008, 71, 1585–1590. [Google Scholar] [CrossRef] [PubMed]
- Paulo, B.S.; Sigrist, R.; Angolini, C.F.F.; De Oliveira, L.G. New cyclodepsipeptide derivatives revealed by genome mining and molecular networking. ChemistrySelect 2019, 4, 7785–7790. [Google Scholar] [CrossRef]
- Pettit, G.R.; Tan, R.; Melody, N.; Kielty, J.M.; Pettit, R.K.; Herald, D.L.; Tucker, B.E.; Mallavia, L.P.; Doubek, D.L.; Schmidt, J.M. Antineoplastic agents. Part 409: Isolation and structure of montanastatin from a terrestrial actinomycete. Bioorganic Med. Chem. 1999, 7, 895–899. [Google Scholar] [CrossRef]
- Wu, C.Y.; Jan, J.T.; Ma, S.H.; Kuo, C.J.; Juan, H.F.; Cheng, Y.S.E.; Hsu, H.H.; Huang, H.C.; Wu, D.; Brik, A.; et al. Small molecules targeting severe acute respiratory syndrome human Coronavirus. Proc. Natl. Acad. Sci. USA 2004, 101, 10012–10017. [Google Scholar] [CrossRef] [Green Version]
- Iacobazzi, R.M.; Annese, C.; Azzariti, A.; D’accolti, L.; Franco, M.; Fusco, C.; la Piana, G.; Laquintana, V.; Denora, N. Antitumor potential of conjugable valinomycins bearing hydroxyl sites: In vitro studies. ACS Med. Chem. Lett. 2013, 4, 1189–1192. [Google Scholar] [CrossRef] [Green Version]
- Rinehart, K.L.; Shield, L.S. Chemistry of the ansamycin antibiotics. Fortschr. Chem. Org. Nat. 1976, 33, 231–307. [Google Scholar] [CrossRef]
| Compound | Strain | Activity | Reference |
|---|---|---|---|
| napsamycin analogs stenothricin analogs | Streptomyces roseosporus | antibiotic | [27] |
| spiroindimicins E and F lagunapyrones D and E | Streptomyces sp. MP131-18 | none | [31] |
| strecacansamycin A, B, C | Streptomyces cacaoi subsp. asoensis | antiproliferative | [25] |
| valinomycin derivatives | Streptomyces sp. CBMAI 2042 | antibiotic | [26] |
| cyclomarin analogs arenicolide analogs retimycin A | Salinispora sp. | suggested antiproliferative | [24] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Krause, J. Applications and Restrictions of Integrated Genomic and Metabolomic Screening: An Accelerator for Drug Discovery from Actinomycetes? Molecules 2021, 26, 5450. https://doi.org/10.3390/molecules26185450
Krause J. Applications and Restrictions of Integrated Genomic and Metabolomic Screening: An Accelerator for Drug Discovery from Actinomycetes? Molecules. 2021; 26(18):5450. https://doi.org/10.3390/molecules26185450
Chicago/Turabian StyleKrause, Janina. 2021. "Applications and Restrictions of Integrated Genomic and Metabolomic Screening: An Accelerator for Drug Discovery from Actinomycetes?" Molecules 26, no. 18: 5450. https://doi.org/10.3390/molecules26185450
APA StyleKrause, J. (2021). Applications and Restrictions of Integrated Genomic and Metabolomic Screening: An Accelerator for Drug Discovery from Actinomycetes? Molecules, 26(18), 5450. https://doi.org/10.3390/molecules26185450






