Effective Use of Empirical Data for Virtual Screening against APJR GPCR Receptor
Abstract
:1. Introduction
2. Results
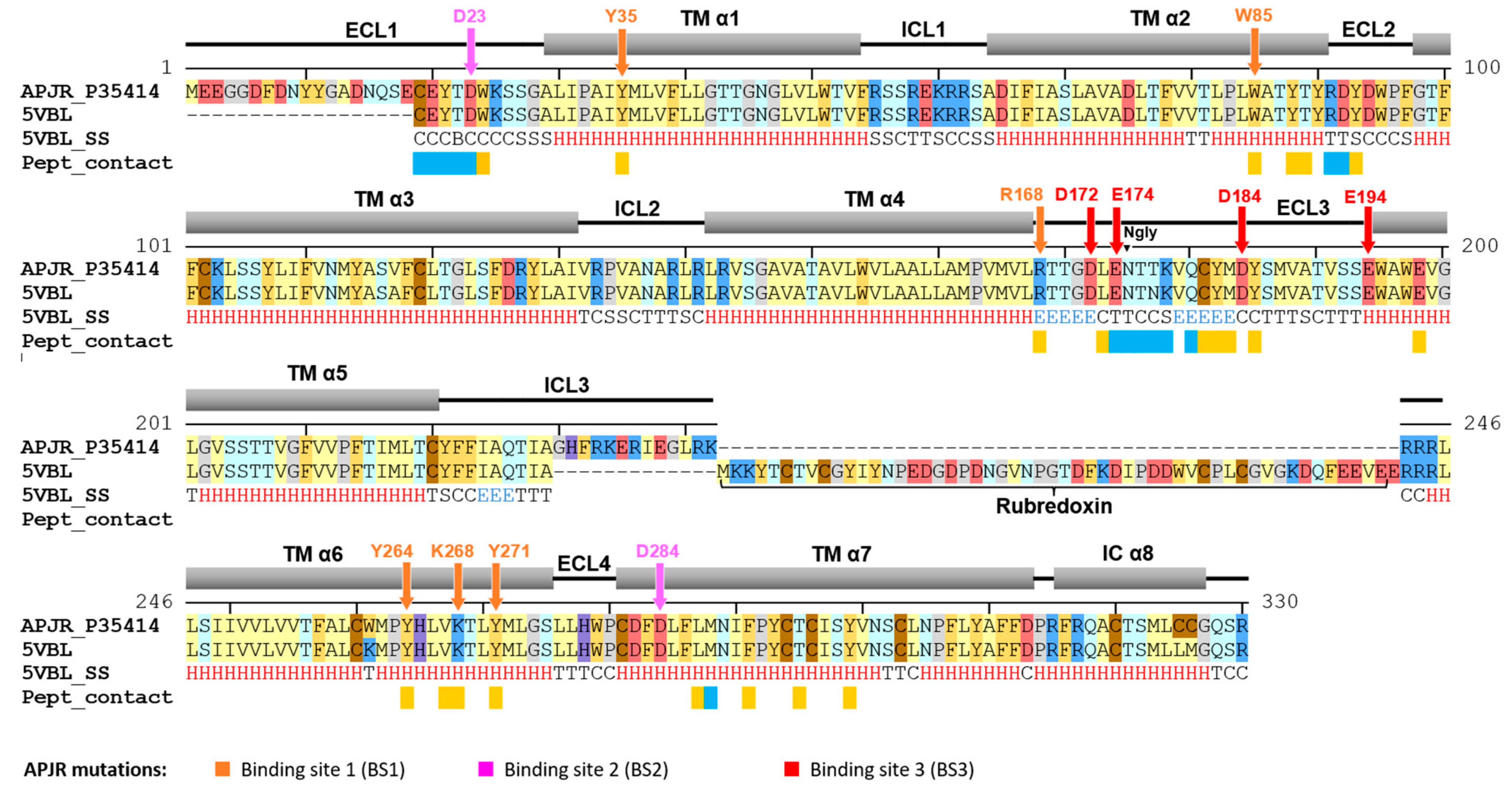
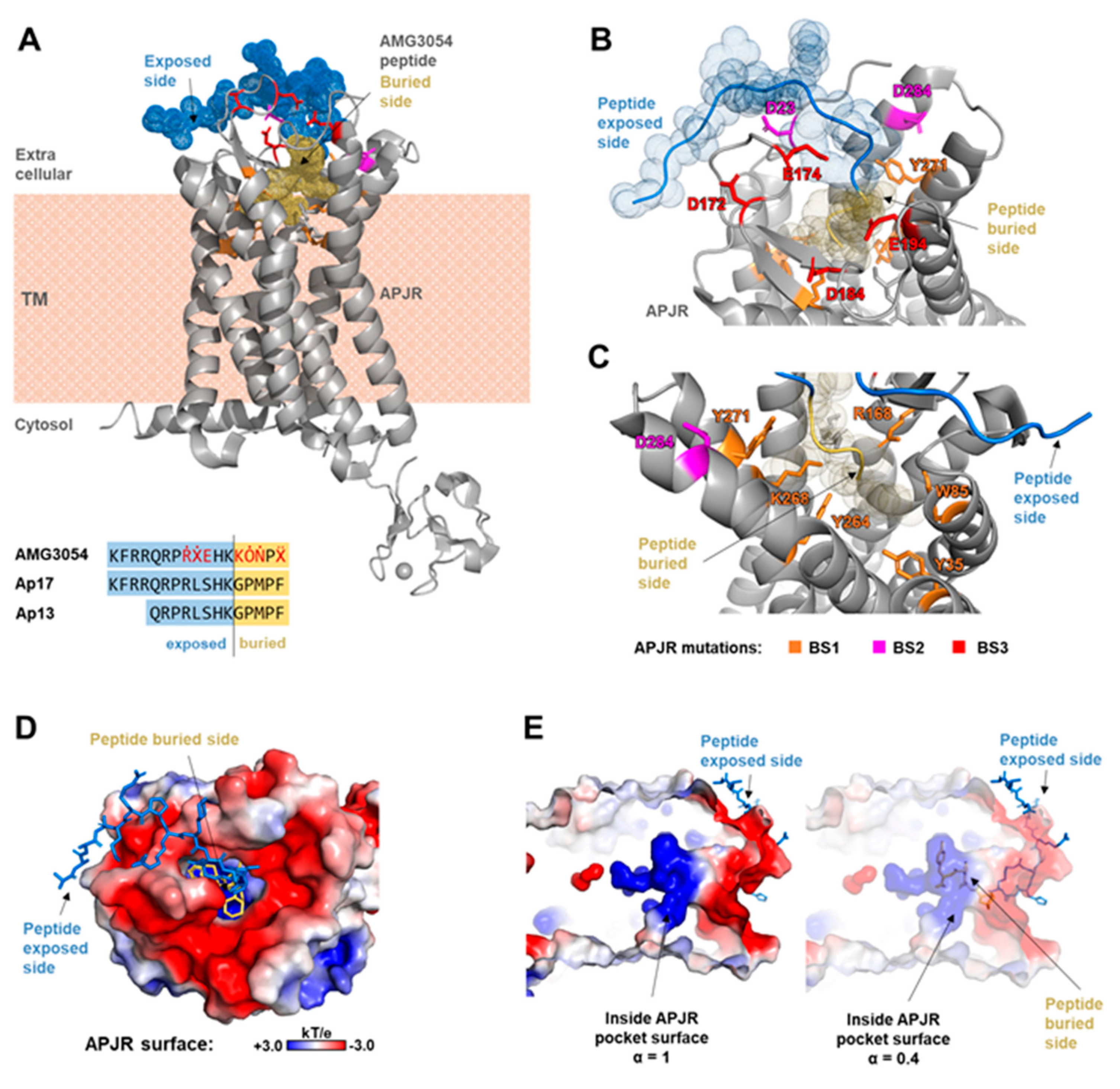
2.1. Receptor Structural Analysis
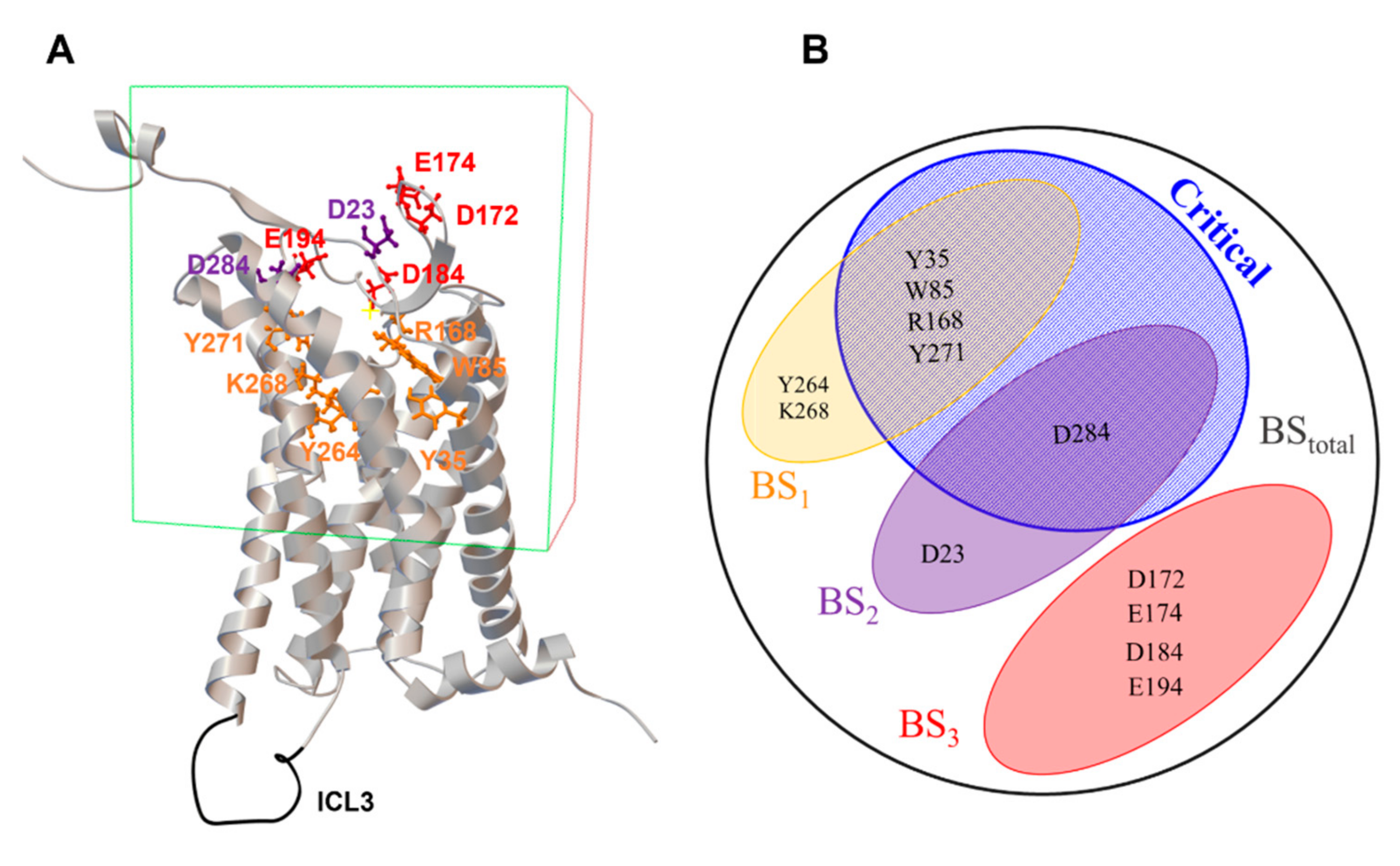
2.2. Receptor Preparation
2.3. Ligand Library Preparation
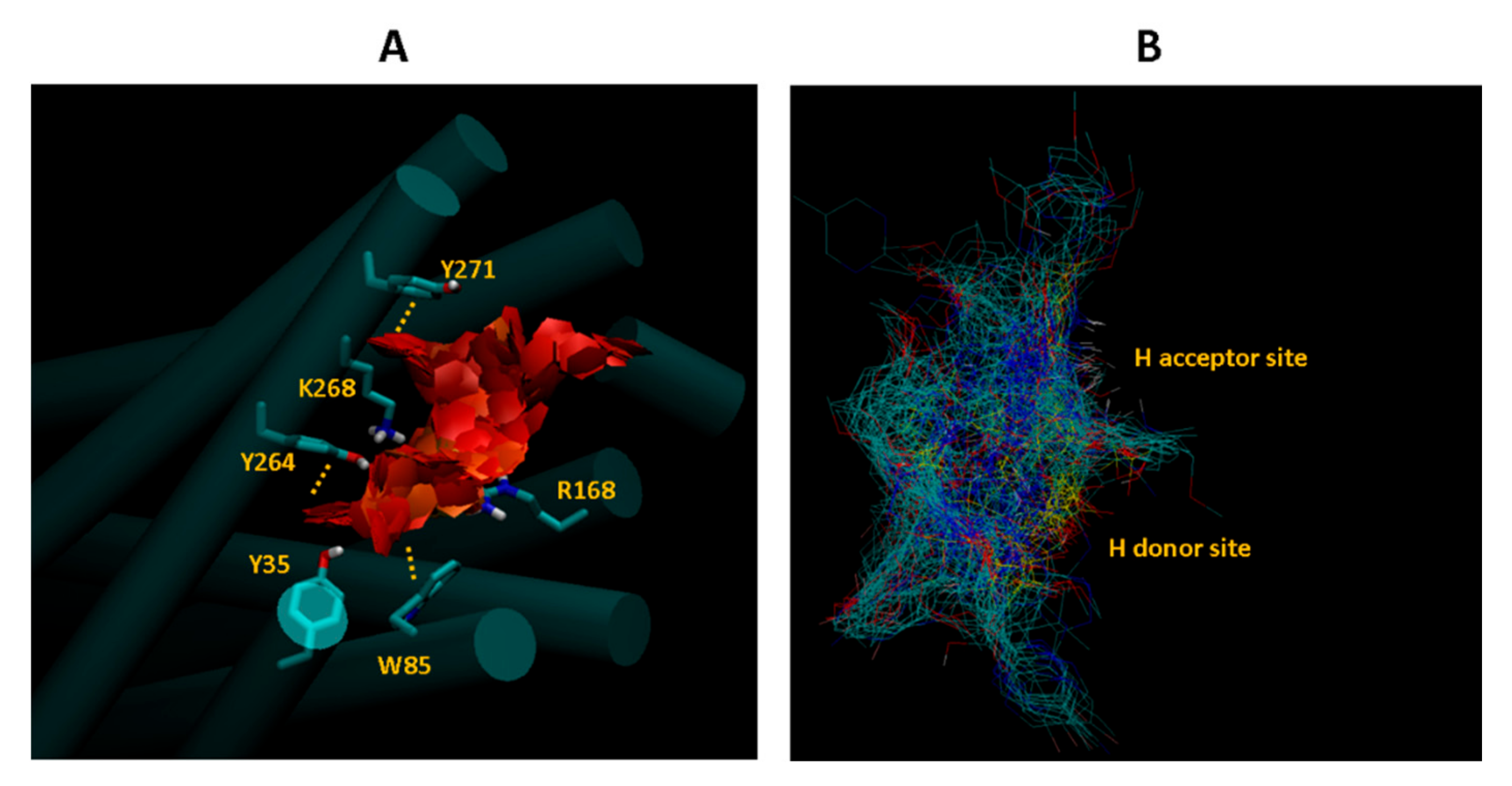
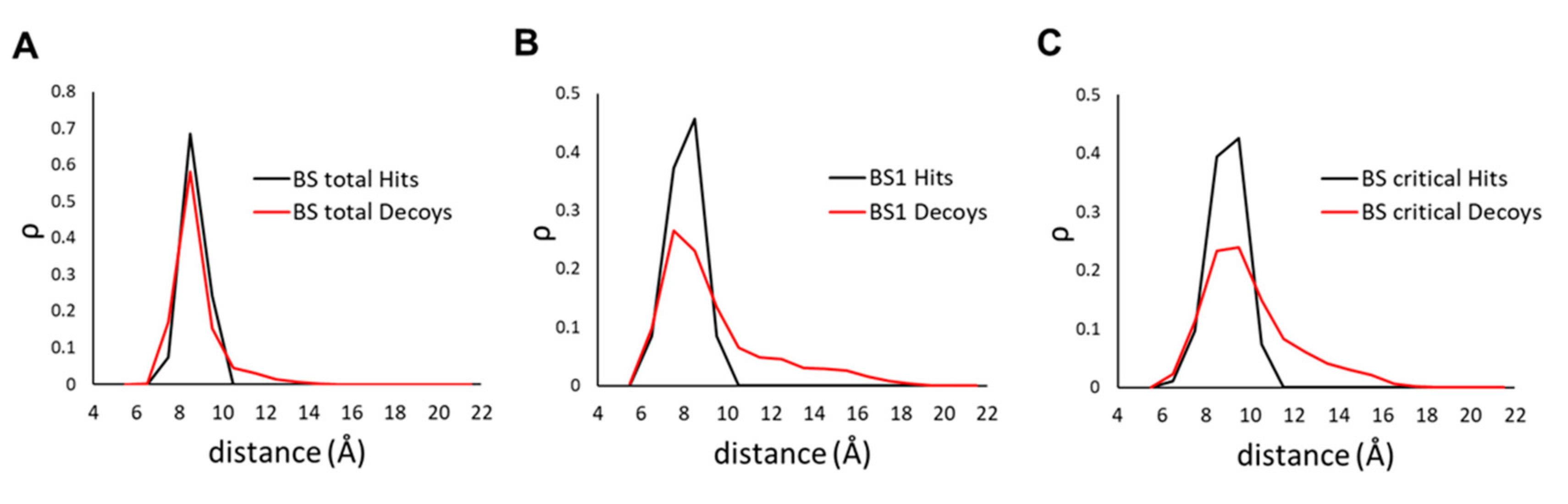
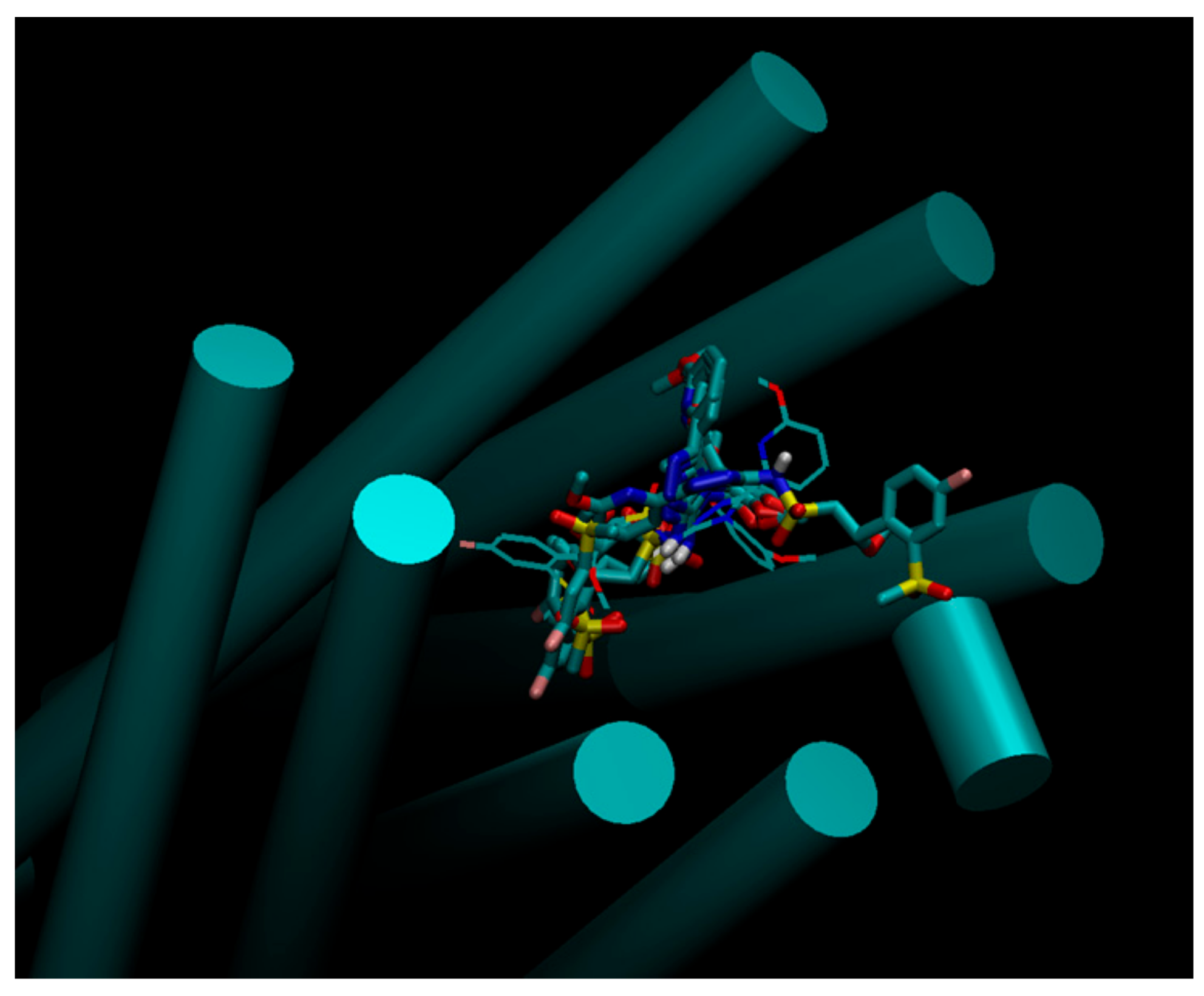
2.4. Docking Pose Analysis
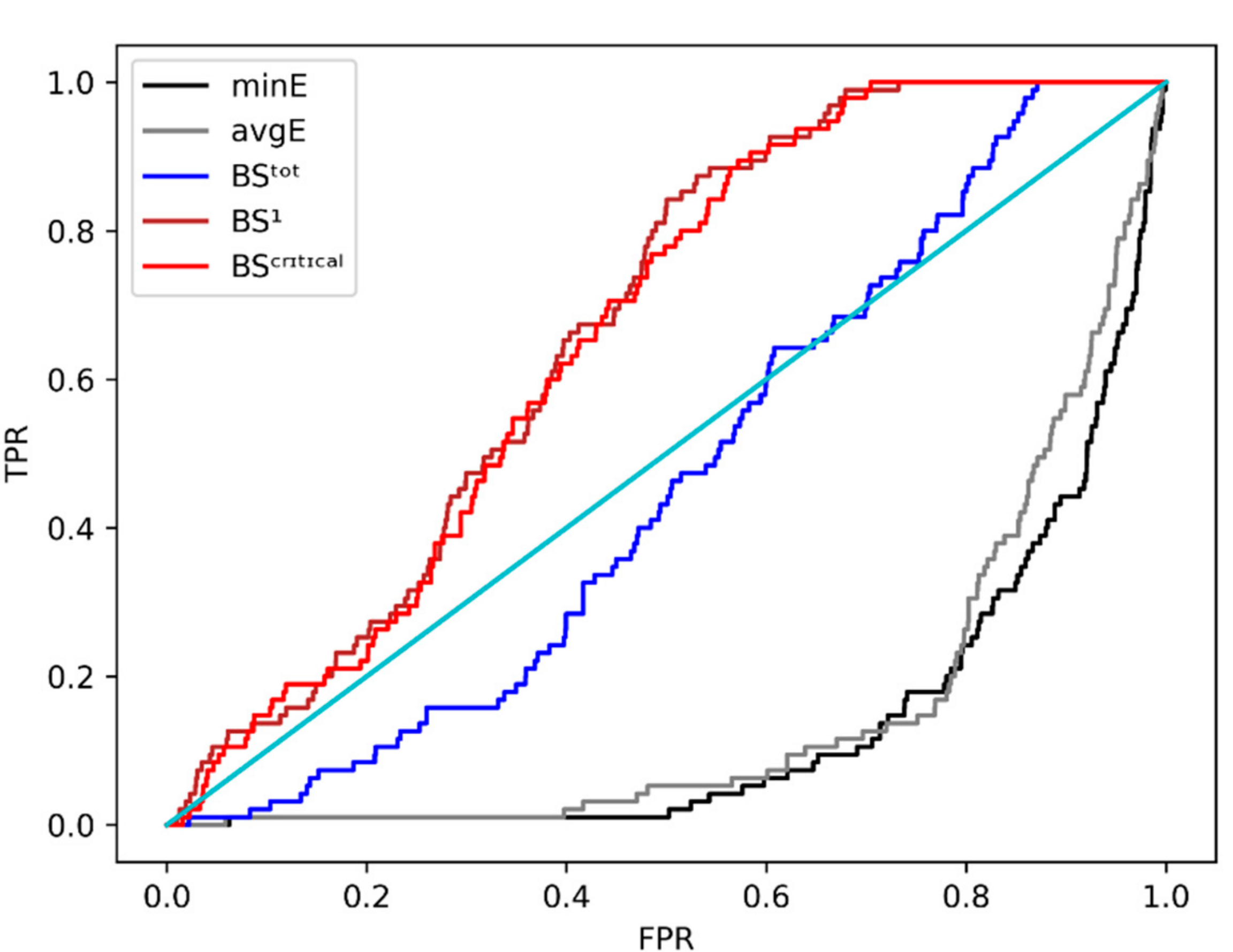
2.5. Fast Stage II rescoring
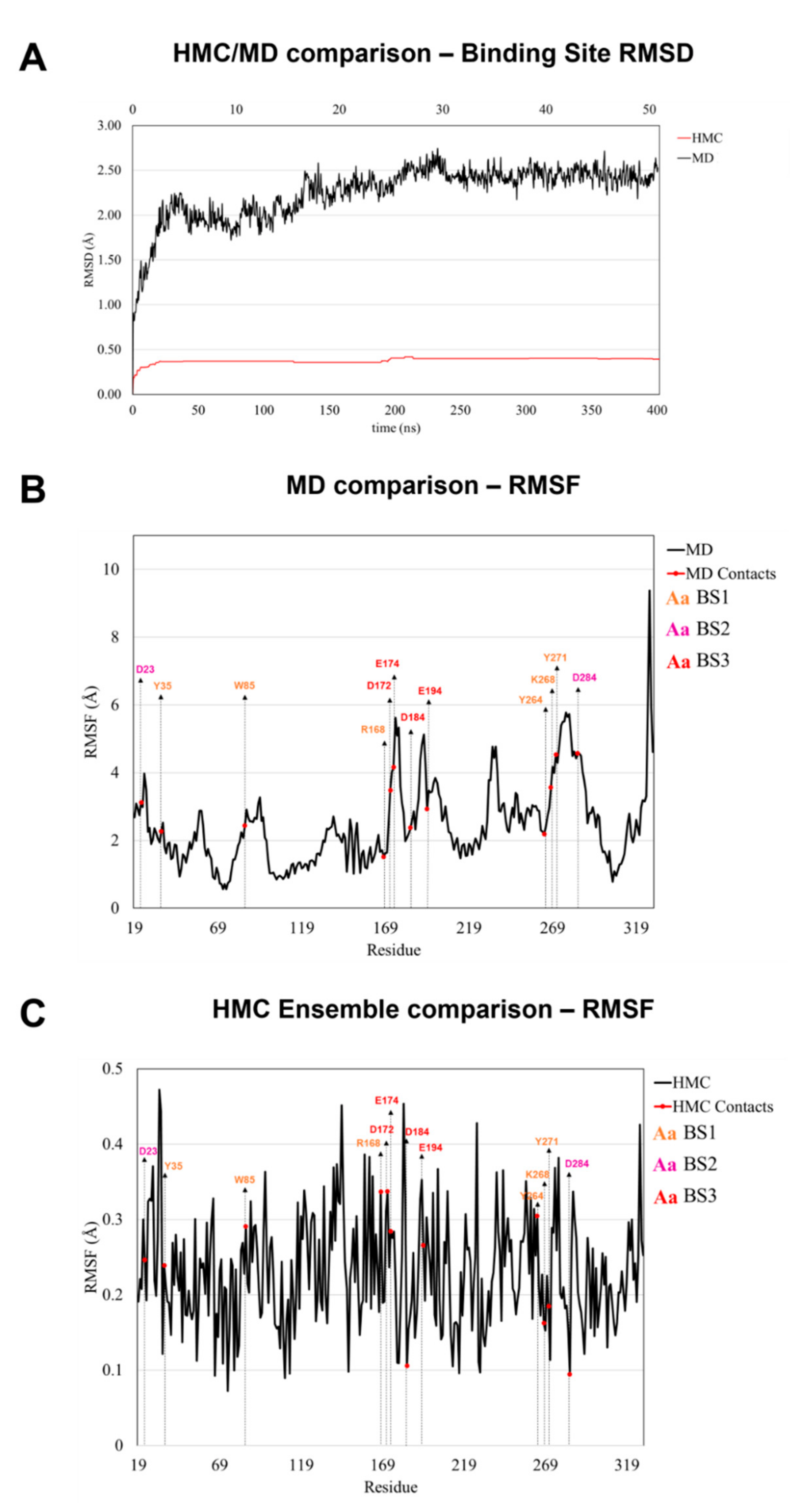
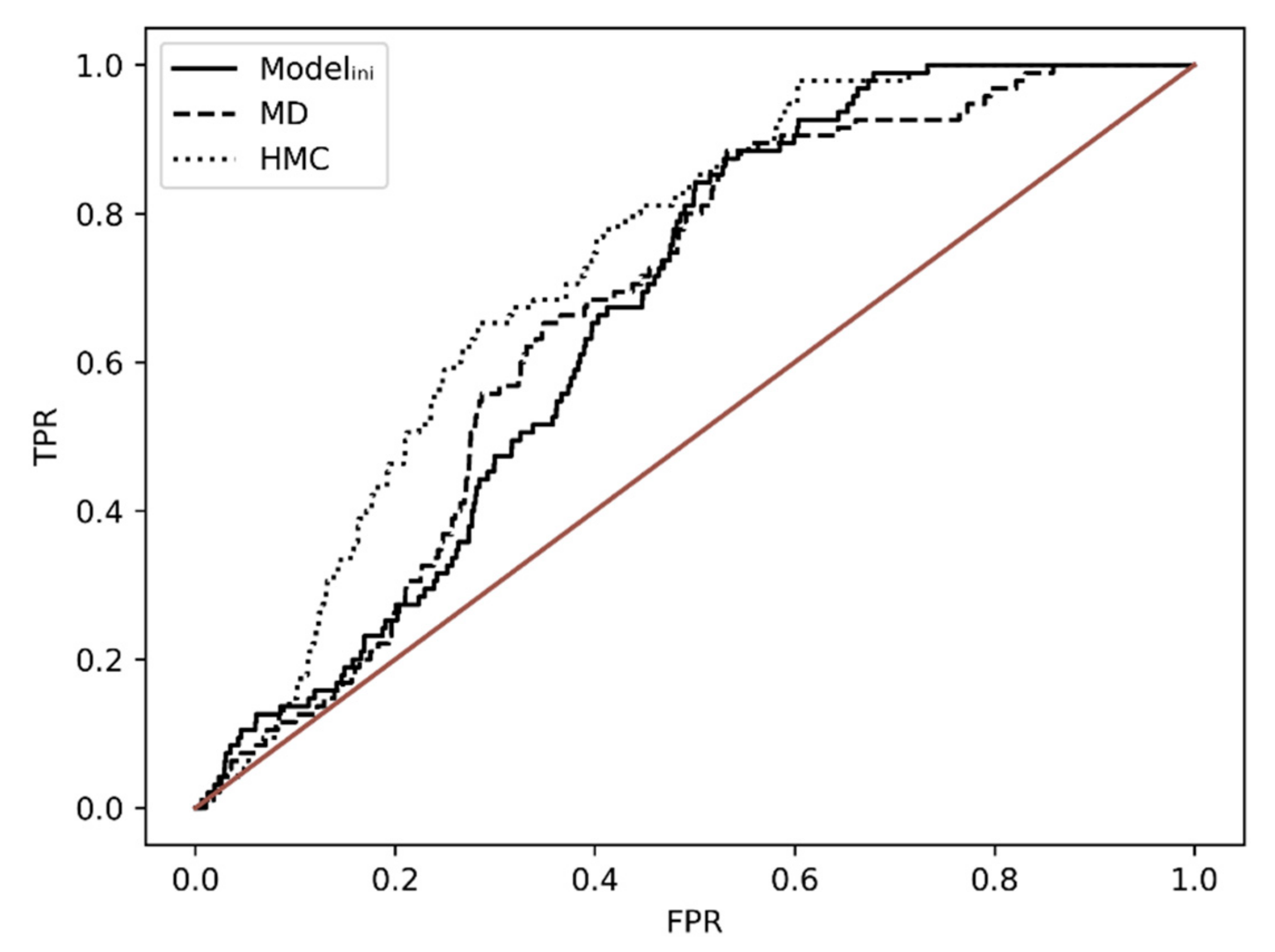
2.6. Ensemble Docking
3. Discussion
4. Materials and Methods
4.1. Molecular Modelling
4.2. Molecular Simulations
4.3. Library Assembly
4.4. Docking
4.5. Analysis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Sample Availability
Appendix A
References
- Briggs, R.; Kennelly, S.P.; O’Neill, D. Drug treatments in Alzheimer’s disease. Clin. Med. 2016, 16, 247. [Google Scholar] [CrossRef]
- Oboudiyat, C.; Glazer, H.; Seifan, A.; Greer, C.; Isaacson, R.S. Alzheimer’s disease. Semin. Neurol. 2013, 33, 313–329. [Google Scholar] [CrossRef] [PubMed]
- Ja, S.L.; González, H.M.; Léger, G.C. Alzheimer’s disease. Handb. Clin. Neurol. 2019, 167, 231–255. [Google Scholar] [CrossRef]
- Luo, H.; Han, L.; Xu, J. Apelin/APJ system: A novel promising target for neurodegenerative diseases. J. Cell Physiol. 2020, 235, 638–657. [Google Scholar] [CrossRef]
- Kumar, A.; Singh, A.; None, E. A review on Alzheimer’s disease pathophysiology and its management: An update. Pharmacol. Rep. 2015, 67, 195–203. [Google Scholar] [CrossRef] [PubMed]
- Masoumi, J.; Abbasloui, M.; Parvan, R.; Mohammadnejad, D.; Pavon-Djavid, G.; Barzegari, A.; Abdolalizadeh, J. Apelin, a promising target for Alzheimer disease prevention and treatment. Neuropeptides 2018, 70, 76–86. [Google Scholar] [CrossRef] [PubMed]
- Kurowska, P.; Barbe, A.; Różycka, M.; Chmielińska, J.; Dupont, J.; Rak, A. Apelin in Reproductive Physiology and Pathology of Different Species: A Critical Review. Int. J. Endocrinol. 2018, 2018, 1–12. [Google Scholar] [CrossRef]
- Chapman, N.A.; Dupré, D.J.; Rainey, J.K. The apelin receptor, Physiology, pathology, cell signalling, and ligand modulation of a peptide-activated class A GPCR. Biochem. Cell. Biol. 2014, 92, 431. [Google Scholar] [CrossRef] [Green Version]
- Wu, L.; Chen, L.; Li, L. Apelin/APJ system: A novel promising therapy target for pathological angiogenesis. Clin. Chim. Acta. 2017, 466, 78–84. [Google Scholar] [CrossRef]
- Wysocka, M.B.; Pietraszek-Gremplewicz, K.; Nowak, D. The Role of Apelin in Cardiovascular Diseases, Obesity and Cancer. Front. Physiol. 2018, 9, 557. [Google Scholar] [CrossRef]
- De Falco, M.; De Luca, L.; Onori, N.; Cavallotti, I.; Artigiano, F.; Esposito, V.; De Luca, B.; Laforgia, V.; Groeger, A.M.; De Luca, A. Apelin expression in normal human tissues. In Vivo 2002, 16, 333–336. [Google Scholar]
- O’Carroll, A.M.; Lolait, S.J.; Harris, L.E.; Pope, G.R. The apelin receptor APJ: Journey from an orphan to a multifaceted regulator of homeostasis. J. Endocrinol. 2013, 219, R13–R35. [Google Scholar] [CrossRef]
- Kleinz, M.J.; Skepper, J.N.; Davenport, A.P. Immunocytochemical localisation of the apelin receptor, APJ, to human cardiomyocytes, vascular smooth muscle and endothelial cells. Regul. Pept. 2005, 126, 233–240. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Yue, Y.; Ma, Y.; Zhang, Q.; Zhou, Q.; Song, Y.; Shen, Y.; Li, X.; Ma, X.; Li, C.; et al. Structural Basis for Apelin Control of the Human Apelin Receptor. Structure 2017, 25, 858–866.e4. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Iturrioz, X.; Gerbier, R.; Leroux, V.; Alvear-Perez, R.; Maigret, B.; Llorens-Cortes, C. By Interacting with the C-terminal Phe of Apelin, Phe255 and Trp259 in Helix VI of the Apelin Receptor Are Critical for Internalization. J. Biol. Chem. 2010, 285, 32627. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Duane, S.; Kennedy, A.D.; Pendleton, B.J.; Roweth, D. Hybrid Monte Carlo. Phys. Lett. B 1987, 195, 216–222. [Google Scholar] [CrossRef]
- Neal, R. MCMC using hamiltonian dynamics. In Handbook of Markov Chain Monte Carlo; Chapman & Hall/CRC: Boca Raton, FL, USA, 2011. [Google Scholar] [CrossRef] [Green Version]
- Sterling, T.; Irwin, J.J. ZINC 15–Ligand Discovery for Everyone. J. Chem. Inf. Model. 2015, 55, 2324–2337. [Google Scholar] [CrossRef]
- Irwin, J.J.; Sterling, T.; Mysinger, M.M.; Bolstad, E.S.; Coleman, R.G. ZINC: A free tool to discover chemistry for biology. J. Chem. Inf. Model. 2012, 52, 1757–1768. [Google Scholar] [CrossRef]
- Irwin, J.J.; Shoichet, B.K. ZINC—A free database of commercially available compounds for virtual screening. J. Chem. Inf. Model. 2005, 45, 177–182. [Google Scholar] [CrossRef] [Green Version]
- Paul, S.M.; Mytelka, D.; Dunwiddie, C.T.; Persinger, C.C.; Munos, B.H.; Lindborg, S.R.; Schacht, A.L. How to improve R&D productivity: The pharmaceutical industry’s grand challenge. Nat. Rev. Drug. Discov. 2010, 9, 203–214. [Google Scholar] [CrossRef]
- Chodera, J.D.; Mobley, D.L.; Shirts, M.R.; Dixon, R.W.; Branson, K.; Pande, V.S. Alchemical free energy methods for drug discovery: Progress and challenges. Curr. Opin. Struct. Biol. 2011, 21, 150–160. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Carlson, H.A.; Masukawa, K.M.; McCammon, A.J. Method for Including the Dynamic Fluctuations of a Protein in Computer-Aided Drug Design. J. Phys. Chem. A 1999, 103, 10213–10219. [Google Scholar] [CrossRef] [Green Version]
- Metz, C.E. Basic principles of ROC analysis. Semin. Nucl. Med. 1978, 8, 283–298. [Google Scholar] [CrossRef]
- Yang, P.; Read, C.; Kuc, R.E.; Buonincontri, G.; Southwood, M.; Torella, R.; Upton, P.D.; Crosby, A.; Sawiak, S.; Carpenter, T.A.; et al. Elabela/Toddler Is an Endogenous Agonist of the Apelin APJ Receptor in the Adult Cardiovascular System, and Exogenous Administration of the Peptide Compensates for the Downregulation of Its Expression in Pulmonary Arterial Hypertension. Circulation 2017, 135, 1160. [Google Scholar] [CrossRef]
- Trân, K.; Murza, A.; Sainsily, X.; Delile, E.; Couvineau, P.; Côté, J.; Coquerel, D.; Peloquin, M.; Auger-Messier, M.; Bouvier, M.; et al. Structure–Activity Relationship and Bioactivity of Short Analogues of ELABELA as Agonists of the Apelin Receptor. J. Med. Chem. 2021, 64, 602–615. [Google Scholar] [CrossRef]
- Gerbier, R.; Leroux, V.; Couvineau, P.; Alvear-Perez, R.; Maigret, B.; Llorens-Cortes, C.; Iturrioz, X. New structural insights into the apelin receptor: Identification of key residues for apelin binding. Faseb J. 2015, 29, 314–322. [Google Scholar] [CrossRef]
- Read, C.; Fitzpatrick, C.M.; Yang, P.; Kuc, R.E.; Maguire, J.; Glen, R.C.; Foster, R.; Davenport, A.P. Cardiac action of the first G protein biased small molecule apelin agonist. Biochem. Pharmacol. 2016, 116, 63–72. [Google Scholar] [CrossRef] [Green Version]
- Fischer, C. A patent review of apelin receptor (APJR) modulators (2014–2019). Expert Opin Pat. 2020, 30, 251–261. [Google Scholar] [CrossRef]
- McAnally, D.; Siddiquee, K.; Sharir, H.; Qi, F.; Phatak, S.; Li, J.-L.; Berg, E.; Fishman, J.; Smith, L. A Systematic Approach to Identify Biased Agonists of the Apelin Receptor through High-Throughput Screening. Slas Discov. Adv. Life Sci. R D 2017, 22, 867–878. [Google Scholar] [CrossRef] [Green Version]
- Chen, N.; Chen, X.; Chen, Y.; Fu, Z.; Heath, J.A.; Horne, D.B. Methods of Treating Heart Failure. US Patent 10,058,550, 28 August 2018. [Google Scholar]
- Liu, T.; Lin, Y.; Wen, X.; Jorissen, R.N.; Gilson, M.K. BindingDB: A web-accessible database of experimentally determined protein–ligand binding affinities. Nucleic Acids Res. 2007, 35, D198. [Google Scholar] [CrossRef] [Green Version]
- Imrie, F.; Bradley, A.R.; Deane, C.M. Generating Property-Matched Decoy Molecules Using Deep Learning. Biorxiv 2020, 1–7. [Google Scholar] [CrossRef]
- Ropp, P.J.; Kaminsky, J.C.; Yablonski, S.; Durrant, J.D. Dimorphite-DL: An open-source program for enumerating the ionization states of drug-like small molecules. J Cheminform. 2019, 11, 1–8. [Google Scholar] [CrossRef]
- Santos-Martins, D.; Solis-Vasquez, L.; Tillack, A.F.; Sanner, M.F.; Koch, A.; Forli, S. Accelerating AutoDock4 with GPUs and Gradient-Based Local Search. J. Chem. Theory Comput. 2021, 17, 1060–1073. [Google Scholar] [CrossRef]
- PubChem. Preparation of (R)—N-(4-(2,6-dimethoxyphenyl)-5-(6-methoxypyridin-2-yl)-4H-1,2,4-triazol-3-yl)-2-(4-fluoro-2-(methylsulfonyl)phenyl)-2-methoxyethanesulfonamide or (S)—N-(4-(2,6-dimethoxyphenyl)-5-(6-methoxypyridin-2-yl)-4H-1,2,4-triazol-3-yl)-2-(4-fluoro-2-(methylsulfonyl)phenyl)-2-methoxyethanesulfonamide. Available online: https://pubchem.ncbi.nlm.nih.gov/compound/122702151 (accessed on 30 May 2021).
- Sali, A.; Blundell, T.L. Comparative protein modelling by satisfaction of spatial restraints. J. Mol. Biol. 1993, 234, 779–815. [Google Scholar] [CrossRef]
- Williams, C.J.; Headd, J.J.; Moriarty, N.W.; Prisant, M.G.; Videau, L.L.; Deis, L.N.; Verma, V.; Keedy, D.A.; Hintze, B.; Chen, V.B.; et al. MolProbity: More and better reference data for improved all-atom structure validation. Protein Sci. 2018, 27, 293. [Google Scholar] [CrossRef] [PubMed]
- Lomize, M.A.; Pogozheva, I.D.; Joo, H.; Mosberg, H.I.; Lomize, A.L. OPM database and PPM web server: Resources for positioning of proteins in membranes. Nucleic Acids Res. 2012, 40, D370–D376. [Google Scholar] [CrossRef] [PubMed]
- Jo, S.; Kim, T.; Iyer, V.G.; Im, W. CHARMM-GUI: A web-based graphical user interface for CHARMM. J. Comput. Chem. 2008, 29, 1859–1865. [Google Scholar] [CrossRef] [PubMed]
- Brooks, B.R.; Brooks, C.L. 3rd, Mackerell, A.D. Jr, Nilsson, L.; Petrella, R.J.; Roux, B.; Karplus, M. CHARMM: The biomolecular simulation program. J. Comput. Chem. 2009, 30, 1545–1614. [Google Scholar] [CrossRef]
- Lee, J.; Cheng, X.; Swails, J.M.; Yeom, M.S.; Eastman, P.K.; Lemkul, J.; Wei, S.; Buckner, J.; Jeong, J.C.; Qi, Y.; et al. CHARMM-GUI Input Generator for NAMD, GROMACS, AMBER, OpenMM, and CHARMM/OpenMM Simulations Using the CHARMM36 Additive Force Field. J. Chem. Theory Comput. 2016, 12, 405–413. [Google Scholar] [CrossRef] [PubMed]
- Wu, E.L.; Cheng, X.; Jo, S.; Rui, H.; Song, K.C.; Dávila-Contreras, E.M.; Im, W. CHARMM-GUI Membrane Builder toward realistic biological membrane simulations. J. Comput. Chem. 2014, 35, 1997–2004. [Google Scholar] [CrossRef] [Green Version]
- Jo, S.; Lim, J.B.; Klauda, J.B.; Im, W. CHARMM-GUI Membrane Builder for mixed bilayers and its application to yeast membranes. Biophys. J. 2009, 97, 50–58. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jo, S.; Kim, T.; Im, W. Automated builder and database of protein/membrane complexes for molecular dynamics simulations. PLoS One 2007, 2, e880. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lee, J.; Patel, D.S.; Ståhle, J.; Park, S.-J.; Kern, N.R.; Kim, S.; Im, W. CHARMM-GUI Membrane Builder for Complex Biological Membrane Simulations with Glycolipids and Lipoglycans. J. Chem. Theory Comput. 2019, 15, 775–786. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Eastman, P.; Swails, J.; Chodera, J.D.; McGibbon, R.T.; Zhao, Y.; Beauchamp, K.A.; Wang, L.-P.; Simmonett, A.C.; Harrigan, M.P.; Stern, C.D.; et al. OpenMM 7: Rapid development of high performance algorithms for molecular dynamics. PLoS Comput. Biol. 2017, 13, e1005659. [Google Scholar] [CrossRef] [PubMed]
- Essmann, U.; Perera, L.; Berkowitz, M.L.; Darden, T.; Lee, H.; Pedersen, L.G. A smooth particle mesh Ewald method. J. Chem. Phys. 1995, 103, 8577–8593. [Google Scholar] [CrossRef] [Green Version]
- Maier, J.A.; Martinez, C.; Kasavajhala, K.; Wickstrom, L.; Hauser, K.E.; Simmerling, C. ff14SB: Improving the Accuracy of Protein Side Chain and Backbone Parameters from ff99SB. J. Chem. Theory Comput. 2015, 11, 3696–3713. [Google Scholar] [CrossRef] [Green Version]
- Kav, B.; Miettinen, M.S. Amber Lipid17 Simulations of POPC/POPS Membranes with CaCl2; Zenodo: Geneva, Switzerland, 2018. [Google Scholar] [CrossRef]
- Wang, J.; Wolf, R.M.; Caldwell, J.W.; Kollman, P.A.; Case, D.A. Development and testing of a general amber force field. J. Comput. Chem. 2004, 25, 1157–1174. [Google Scholar] [CrossRef]
- Betancourt, M.; Girolami, M. Hamiltonian Monte Carlo for Hierarchical Models. Curr. Trends Bayesian Methodol. Appl. 2015, 79, 2–4. [Google Scholar] [CrossRef] [Green Version]
- Fang, L.; Makkonen, E.; Todorović, M.; Rinke, P.; Chen, X. Efficient Amino Acid Conformer Search with Bayesian Optimization. J. Chem. Theory Comput. 2021, 17, 1955–1966. [Google Scholar] [CrossRef]
- O’Boyle, N.M.; Banck, M.; James, C.A.; Morley, C.; Vandermeersch, T.; Hutchison, G.R. Open Babel: An open chemical toolbox. J. Cheminform. 2011, 3, 1–14. [Google Scholar] [CrossRef] [Green Version]
- Open Babel. Available online: http://openbabel.org/wiki/Main_Page (accessed on 7 April 2021).
- Case, D.A.; Cheatham, T.E., III.; Darden, T.; Gohlke, H.; Luo, R.; Merz, K.M.; Onufriev, A.; Simmerling, C.; Wang, B.; Woods, R.J. The Amber Biomolecular Simulation Programs. J. Comput. Chem. 2005, 26, 1668. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Morris, G.M.; Goodsell, D.S.; Halliday, R.S.; Huey, R.; Hart, W.E.; Belew, R.K.; Olson, A.J. Automated docking using a Lamarckian genetic algorithm and an empirical binding free energy function. J. Comput. Chem. 1998, 19, 1639–1662. [Google Scholar] [CrossRef] [Green Version]
- Schrödinger, LLC. The PyMOL Molecular Graphics System, Version 1.8.; Schrödinger, LLC: New York, NY, USA, 2015. [Google Scholar]
- Humphrey, W.; Dalke, A.; Schulten, K. VMD: Visual molecular dynamics. J. Mol. Graph. 1996, 14, 33–38. [Google Scholar] [CrossRef]
- Jurrus, E.; Engel, D.; Star, K.; Monson, K.; Brandi, J.; Felberg, L.E.; Brookes, D.H.; Wilson, L.; Chen, J.; Liles, K.; et al. Improvements to the APBS biomolecular solvation software suite. Protein Sci. 2018, 27, 112–128. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Caswell, T.A.; Droettboom, M.; Lee, A.; de Andrade, E.S.; Hunter, J.; Hoffmann, T.; Firing, E.; Klymak, J.; Stansby, D.; Varoquaux, N.; et al. Matplotlib/Matplotlib: REL: v3.4.2.; Zenodo: Geneva, Switzerland, 2021. [Google Scholar] [CrossRef]

| AUC | AUC Ratio | |
|---|---|---|
| minE | 0.142 | 1 |
| avgE | 0.164 | 1.154 |
| BStot | 0.464 | 3.267 |
| BS1 | 0.664 | 4.676 |
| BScritical | 0.656 | 4.619 |
| AUC | AUC Ratio | |
|---|---|---|
| Modelini | 0.664 | 1.000 |
| MD | 0.665 | 1.001 |
| HMC | 0.725 | 1.091 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Manoliu, L.C.E.; Martin, E.C.; Milac, A.L.; Spiridon, L. Effective Use of Empirical Data for Virtual Screening against APJR GPCR Receptor. Molecules 2021, 26, 4894. https://doi.org/10.3390/molecules26164894
Manoliu LCE, Martin EC, Milac AL, Spiridon L. Effective Use of Empirical Data for Virtual Screening against APJR GPCR Receptor. Molecules. 2021; 26(16):4894. https://doi.org/10.3390/molecules26164894
Chicago/Turabian StyleManoliu, Laura C. E., Eliza C. Martin, Adina L. Milac, and Laurentiu Spiridon. 2021. "Effective Use of Empirical Data for Virtual Screening against APJR GPCR Receptor" Molecules 26, no. 16: 4894. https://doi.org/10.3390/molecules26164894
APA StyleManoliu, L. C. E., Martin, E. C., Milac, A. L., & Spiridon, L. (2021). Effective Use of Empirical Data for Virtual Screening against APJR GPCR Receptor. Molecules, 26(16), 4894. https://doi.org/10.3390/molecules26164894






