Red Ginseng Improves Exercise Endurance by Promoting Mitochondrial Biogenesis and Myoblast Differentiation
Abstract
1. Introduction
2. Results
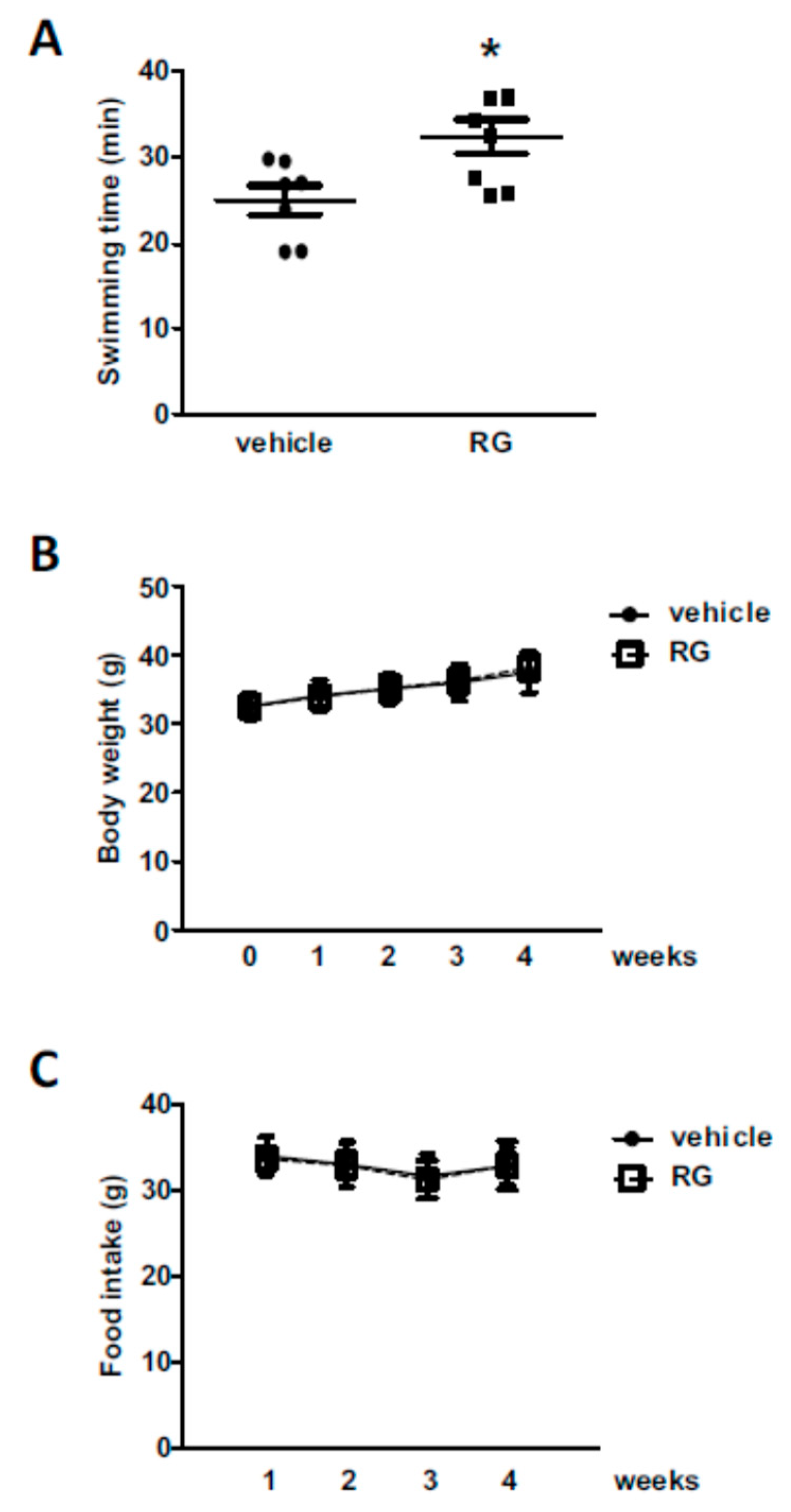
2.1. Administration of RG Improves Swimming Performance in Mice
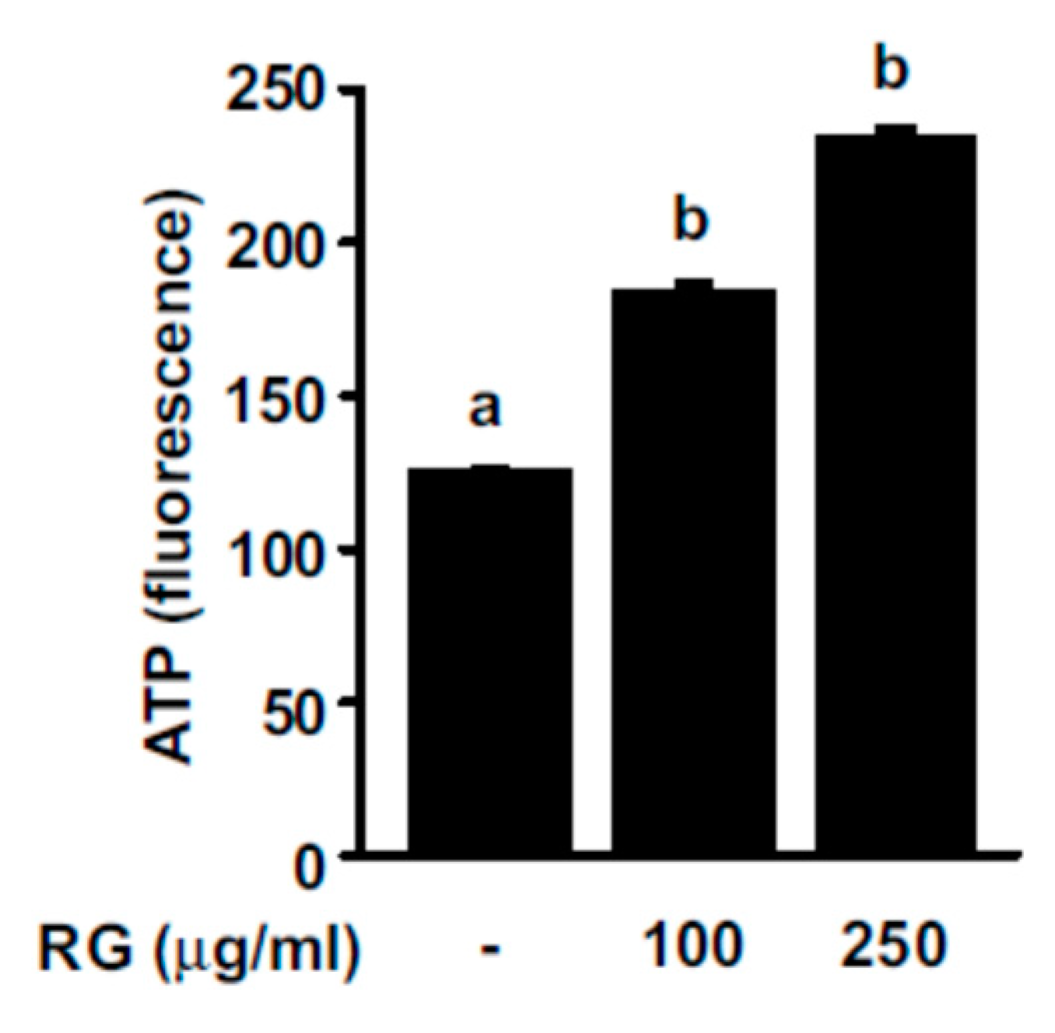
2.2. RG Increases ATP Production Levels in C2C12 Myoblasts
2.3. RG Promotes Mitochondrial Biogenesis in C2C12 Myoblasts
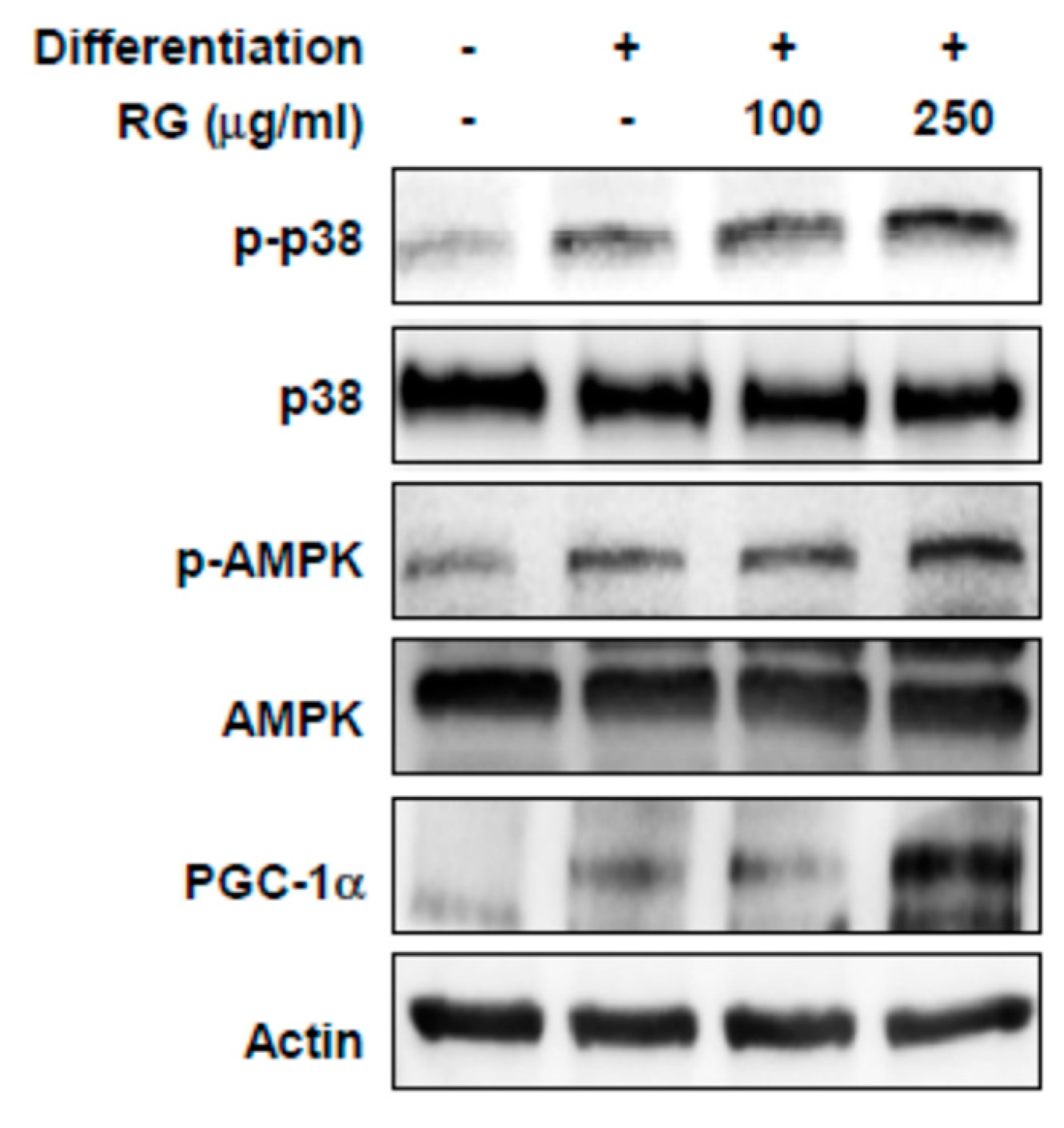
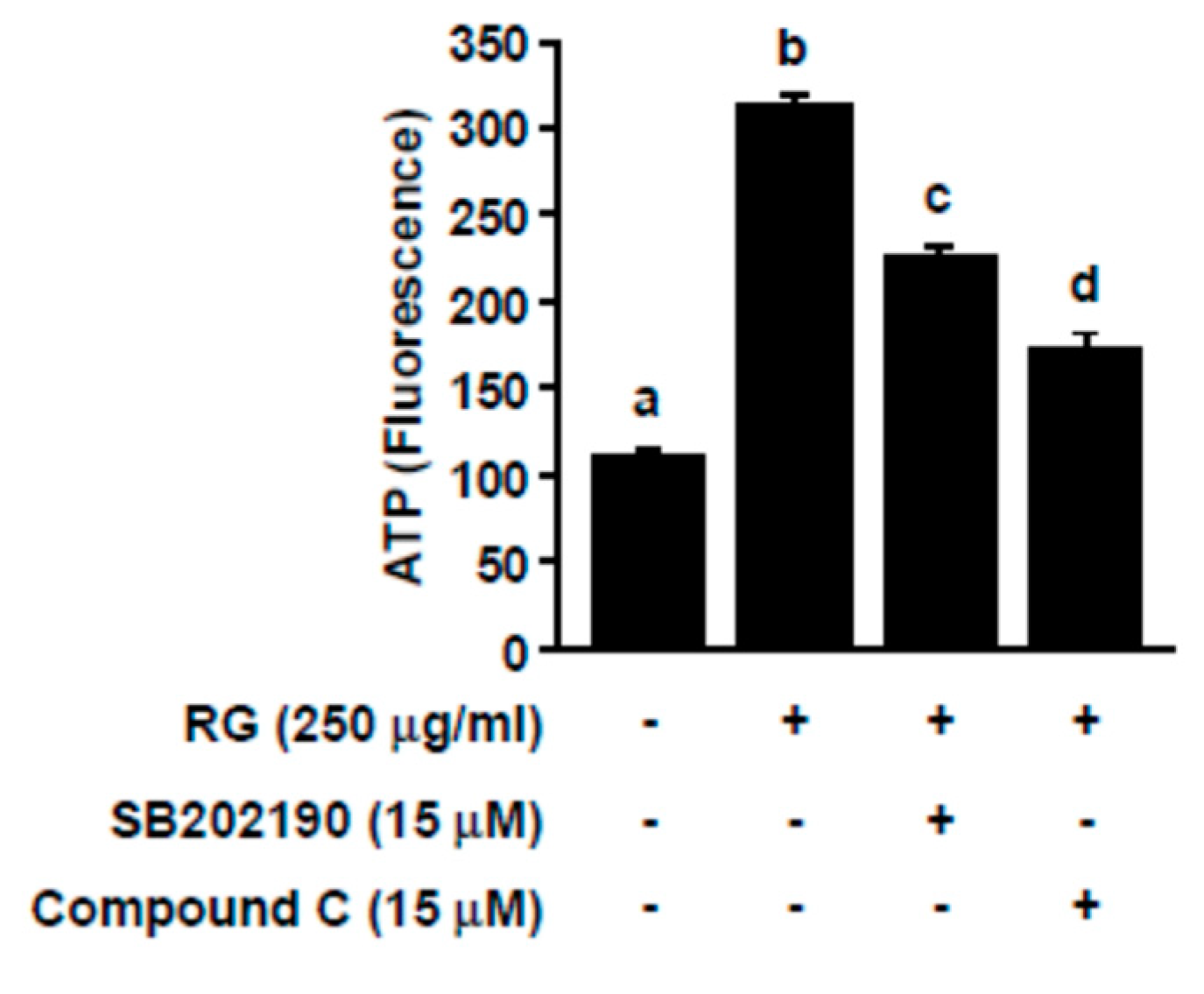
2.4. RG Upregulates p38, AMPK, and PGC-1α
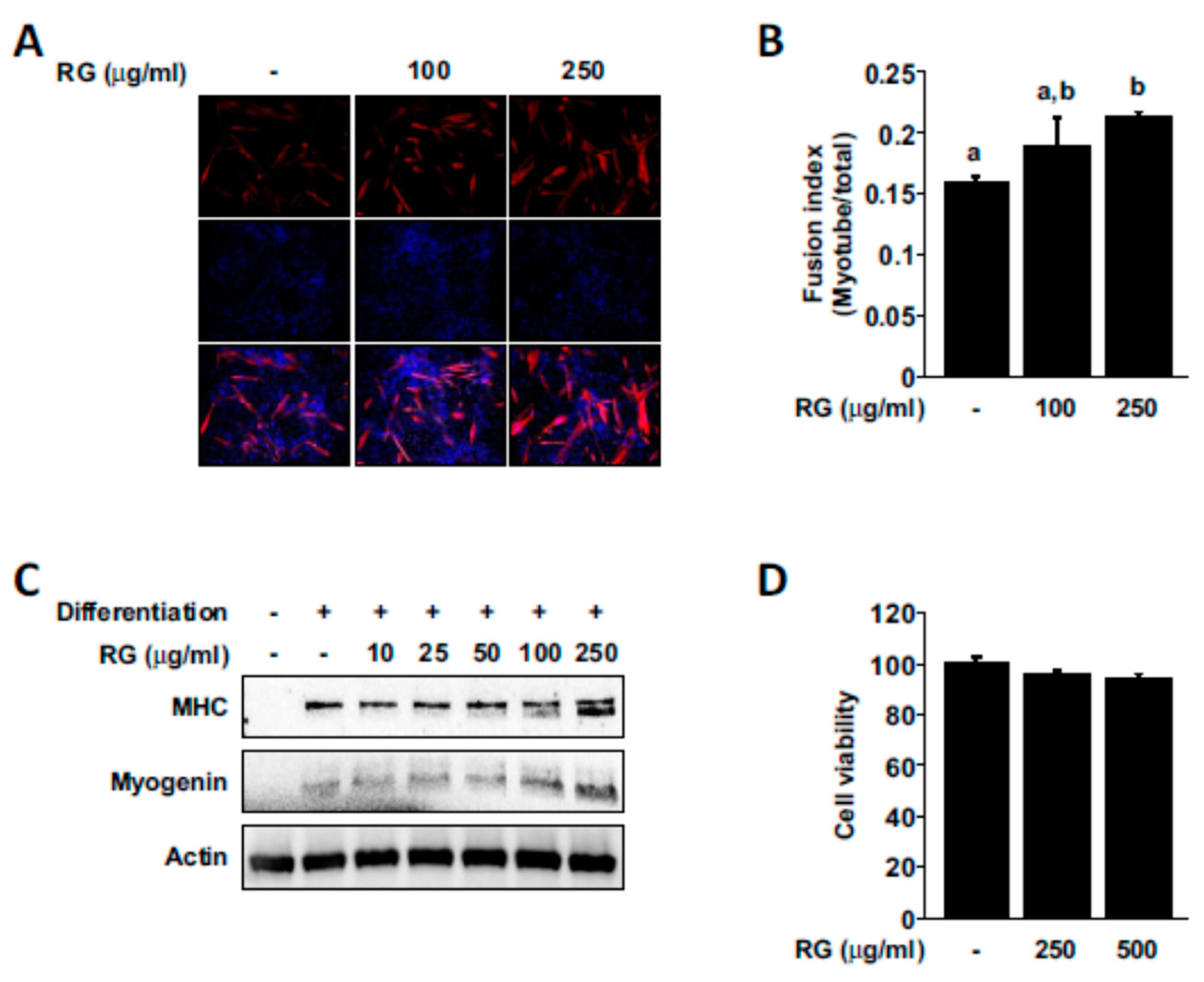
2.5. RG Promotes Myogenic Differentiation
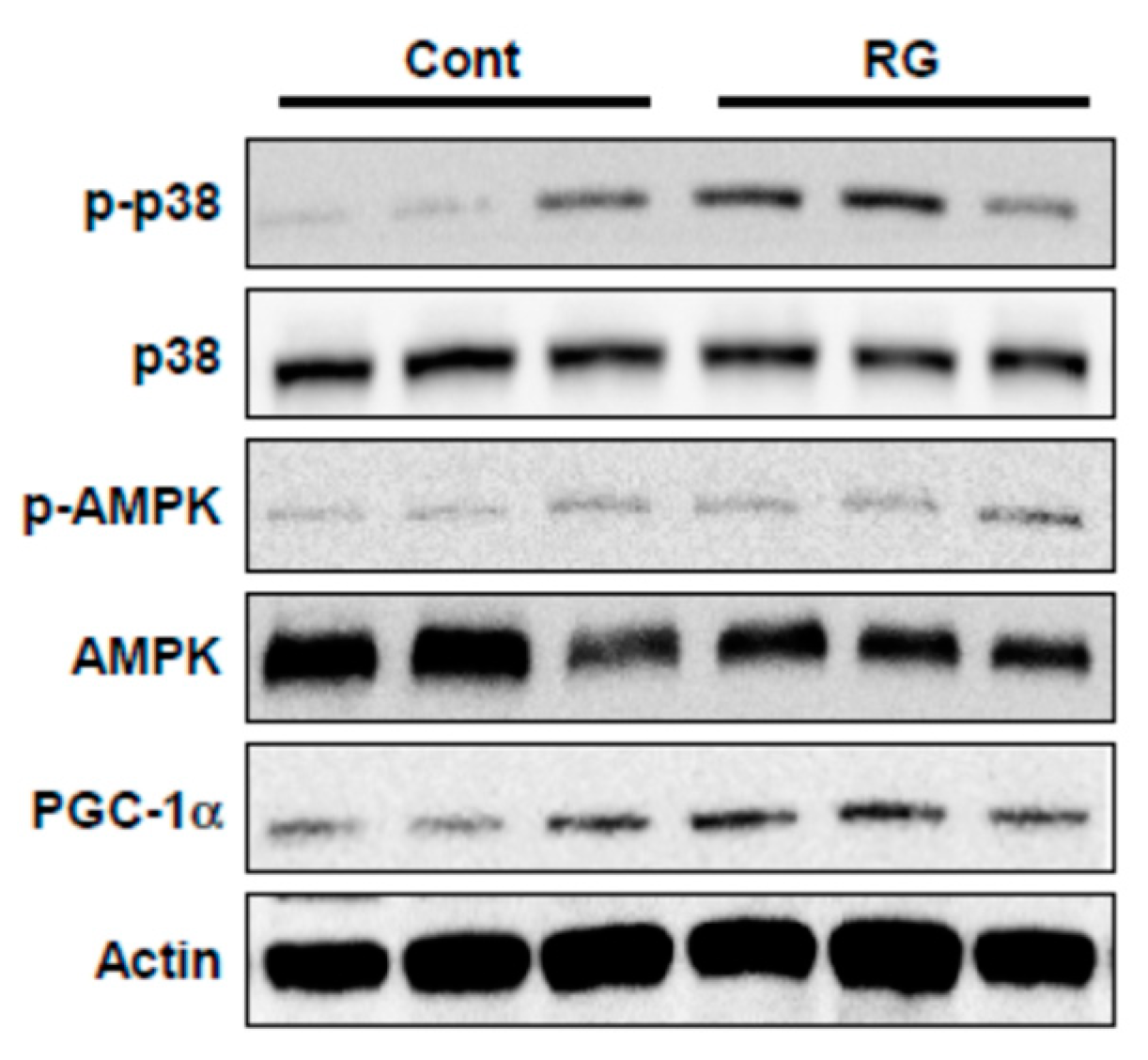
2.6. RG Administration Activates p38, AMPK, and PGC-1α Signaling In Vivo
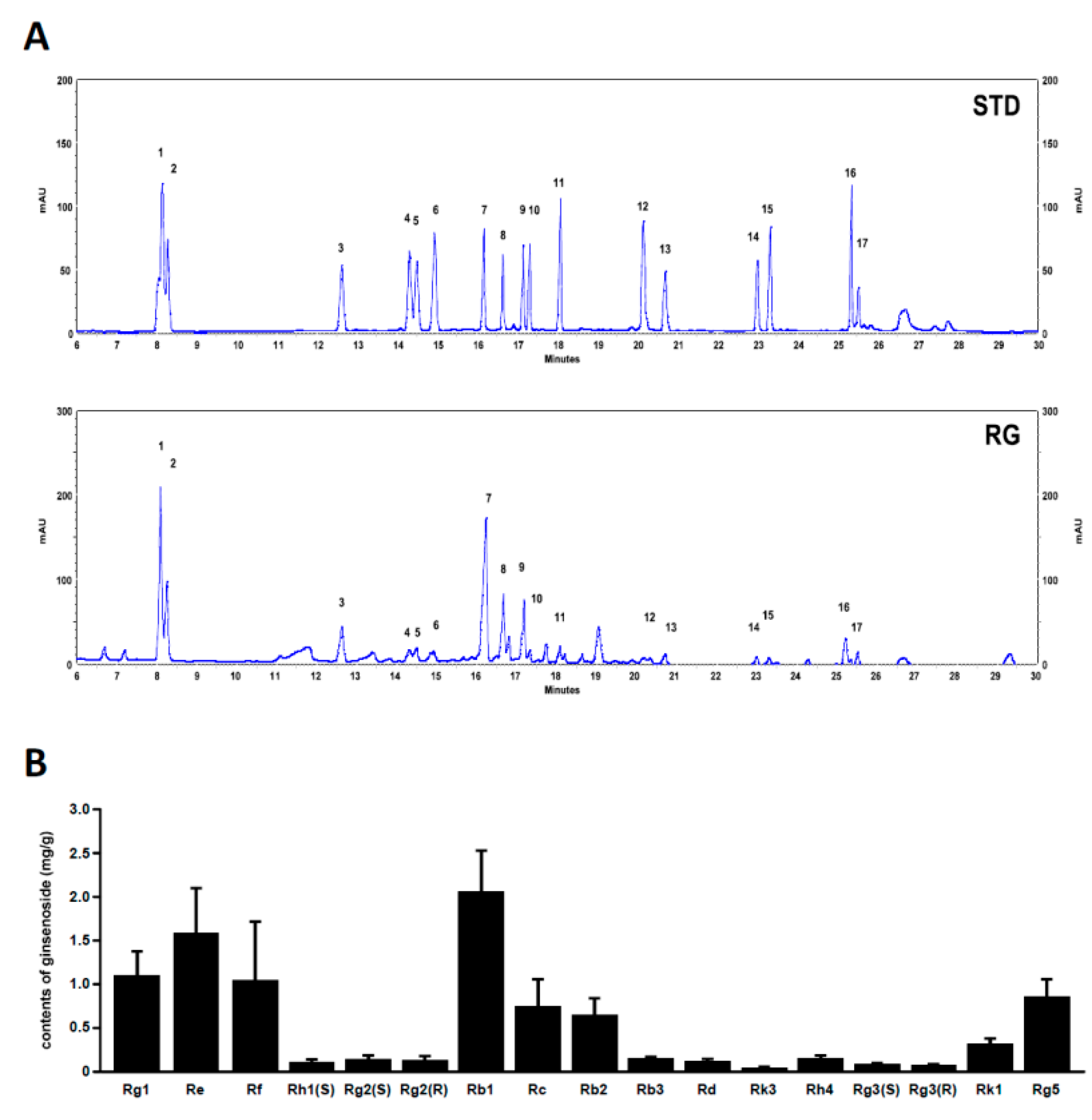
2.7. Analysis of Ginsenoside Profiles for RG
3. Discussion
4. Materials and Methods
4.1. Reagents
4.2. Preparation of Red Ginseng Extracts
4.3. Animals and Treatment
4.4. Exhaustive Swimming Exercise Performance Test
4.5. Cell Culture
4.6. Measurement of Adenosine Triphosphate Levels
4.7. Quantitative Real-Time PCR and Analyses of mtDNA Content
4.8. Immunoblotting
4.9. Immunofluorescence
4.10. Cell Viability
4.11. UPLC Analysis
4.12. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Xue, Q.-L.; Walston, J.D.; Fried, L.P.; Beamer, B.A. Prediction of Risk of Falling, Physical Disability, and Frailty by Rate of Decline in Grip Strength: The Women’s Health and Aging Study. Arch. Intern. Med. 2011, 171, 1119. [Google Scholar] [CrossRef] [PubMed]
- Marsh, A.P.; Rejeski, W.J.; Espeland, M.A.; Miller, M.E.; Church, T.S.; Fielding, R.A.; Gill, T.M.; Guralnik, J.M.; Newman, A.B.; Pahor, M. Muscle Strength and BMI as Predictors of Major Mobility Disability in the Lifestyle Interventions and Independence for Elders Pilot (LIFE-P). J. Gerontol. Ser. A Boil. Sci. Med. Sci. 2011, 66, 1376–1383. [Google Scholar] [CrossRef] [PubMed]
- Peng, P.; Hyder, O.; Firoozmand, A.; Kneuertz, P.; Schulick, R.D.; Huang, D.; Makary, M.; Hirose, K.; Edil, B.; Choti, M.A.; et al. Impact of sarcopenia on outcomes following resection of pancreatic adenocarcinoma. J. Gastrointest. Surg. 2012, 16, 1478–1486. [Google Scholar] [CrossRef] [PubMed]
- Newman, A.B.; Kupelian, V.; Visser, M.; Simonsick, E.M.; Goodpaster, B.H.; Kritchevsky, S.B.; Tylavsky, F.A.; Rubin, S.M.; Harris, T.B. Strength, but not muscle mass, is associated with mortality in the health, aging and body composition study cohort. J. Gerontol. Ser. A Boil. Sci. Med. Sci. 2006, 61, 72–77. [Google Scholar] [CrossRef] [PubMed]
- Landi, F.; Liperoti, R.; Russo, A.; Giovannini, S.; Tosato, M.; Capoluongo, E.; Bernabei, R.; Onder, G. Sarcopenia as a risk factor for falls in elderly individuals: Results from the ilSIRENTE study. Clin. Nutr. 2012, 31, 652–658. [Google Scholar] [CrossRef] [PubMed]
- Walston, J.D. Sarcopenia in older adults. Curr. Opin. Rheumatol. 2012, 24, 623–627. [Google Scholar] [CrossRef]
- Deane, C.S.; Wilkinson, D.J.; Phillips, B.E.; Smith, K.; Etheridge, T.; Atherton, P.J. “Nutraceuticals” in relation to human skeletal muscle and exercise. Am. J. Physiol. Endocrinol. Metab. 2017, 312, E282–E299. [Google Scholar] [CrossRef]
- Kang, C.H.; Chung, E.; Diffee, G.; Ji, L.L. Exercise training attenuates aging-associated mitochondrial dysfunction in rat skeletal muscle: Role of PGC-1 alpha. Exp. Gerontol. 2013, 48, 1343–1350. [Google Scholar] [CrossRef]
- Egan, B.; Zierath, J.R. Exercise Metabolism and the Molecular Regulation of Skeletal Muscle Adaptation. Cell Metab. 2013, 17, 162–184. [Google Scholar] [CrossRef]
- Kang, C.H.; Ji, L.L. Role of PGC-1 alpha signaling in skeletal muscle health and disease. Ann. N. Y. Acad. Sci. 2012, 1271, 110–117. [Google Scholar] [CrossRef]
- Ruas, J.L.; White, J.P.; Rao, R.R.; Kleiner, S.; Brannan, K.T.; Harrison, B.C.; Greene, N.P.; Wu, J.; Estall, J.L.; Irving, B.A.; et al. PGC-1 alpha Isoform Induced by Resistance Training Regulates Skeletal Muscle Hypertrophy. Cell 2012, 151, 1319–1331. [Google Scholar] [CrossRef] [PubMed]
- Tadaishi, M.; Miura, S.; Kai, Y.; Kano, Y.; Oishi, Y.; Ezaki, O. Skeletal Muscle-Specific Expression of PGC-1 alpha-b, an Exercise-Responsive Isoform, Increases Exercise Capacity and Peak Oxygen Uptake. PLoS ONE 2011, 6, e28290. [Google Scholar] [CrossRef] [PubMed]
- Jørgensen, S.B.; Richter, E.A.; Wojtaszewski, J.F.P. Role of AMPK in skeletal muscle metabolic regulation and adaptation in relation to exercise. J. Physiol. 2006, 574, 17–31. [Google Scholar] [CrossRef] [PubMed]
- Zong, H.; Ren, J.M.; Young, L.H.; Pypaert, M.; Mu, J.; Birnbaum, M.J.; Shulman, G.I. AMP kinase is required for mitochondrial biogenesis in skeletal muscle in response to chronic energy deprivation. Proc. Natl. Acad. Sci. USA 2002, 99, 15983–15987. [Google Scholar] [CrossRef]
- Kjøbsted, R.; Hingst, J.R.; Fentz, J.; Foretz, M.; Sanz, M.-N.; Pehmøller, C.; Shum, M.; Marette, A.; Mounier, R.; Treebak, J.T.; et al. AMPK in skeletal muscle function and metabolism. FASEB J. 2018, 32, 1741–1777. [Google Scholar] [CrossRef] [PubMed]
- Peralta, S.; Garcia, S.; Yin, H.Y.; Arguello, T.; Diaz, F.; Moraes, C.T. Sustained AMPK activation improves muscle function in a mitochondrial myopathy mouse model by promoting muscle fiber regeneration. Hum. Mol. Genet. 2016, 25, 3178–3191. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.-H. Pharmacological and medical applications of Panax ginseng and ginsenosides: A review for use in cardiovascular diseases. J. Ginseng Res. 2018, 42, 264–269. [Google Scholar] [CrossRef]
- Choi, J.; Kim, T.H.; Choi, T.Y.; Lee, M.S. Ginseng for Health Care: A Systematic Review of Randomized Controlled Trials in Korean Literature. PLoS ONE 2013, 8, e59978. [Google Scholar] [CrossRef]
- Kim, W.Y.; Kim, J.M.; Han, S.B.; Lee, S.K.; Kim, N.D.; Park, M.K.; Kim, C.K.; Park, J.H. Steaming of ginseng at high temperature enhances biological activity. J. Nat. Prod. 2000, 63, 1702–1704. [Google Scholar] [CrossRef]
- Lee, S.-A.; Jo, H.-K.; Im, B.-O.; Kim, S.-U.; Whang, W.-K.; Ko, S.-K. Changes in the Contents of Prosapogenin in the Red Ginseng (Panax ginseng) Depending on Steaming Batches. J. Ginseng Res. 2012, 36, 102–106. [Google Scholar] [CrossRef]
- Toft, A.D.; Jensen, L.B.; Bruunsgaard, H.; Ibfelt, T.; Halkjær-Kristensen, J.; Febbraio, M.; Pedersen, B.K. Cytokine response to eccentric exercise in young and elderly humans. Am. J. Physiol. Physiol. 2002, 283, C289–C295. [Google Scholar] [CrossRef] [PubMed]
- Voces, J.; De Oliveira, A.C.; Prieto, J.; Vila, L.; Perez, A.; Duarte, I.; Alvarez, A. Ginseng administration protects skeletal muscle from oxidative stress induced by acute exercise in rats. Braz. J. Med. Boil. Res. 2004, 37, 1863–1871. [Google Scholar] [CrossRef] [PubMed]
- Li, X.-T.; Chen, R.; Jin, L.-M.; Chen, H.-Y. Regulation on Energy Metabolism and Protection on Mitochondria of Panax Ginseng Polysaccharide. Am. J. Chin. Med. 2009, 37, 1139–1152. [Google Scholar] [CrossRef] [PubMed]
- Babiker, L.B.; Gadkariem, E.A.; Alashban, R.M.; Aljohar, H.I. Investigation of Stability of Korean Ginseng in Herbal Drug Product. Am. J. Appl. Sci. 2014, 11, 160–170. [Google Scholar] [CrossRef]
- Shin, M.-S.; Song, J.H.; Choi, P.; Lee, J.H.; Kim, S.-Y.; Shin, K.S.; Ham, J.; Kang, K.S. Stimulation of Innate Immune Function by Panax ginseng after Heat Processing. J. Agric. Food Chem. 2018, 66, 4652–4659. [Google Scholar] [CrossRef]
- Lee, S.M.; Kim, S.C.; Oh, J.; Kim, J.H.; Na, M. 20(R)-Ginsenoside Rf: A new ginsenoside from red ginseng extract. Phytochem. Lett. 2013, 6, 620–624. [Google Scholar] [CrossRef]
- Li, S.; Chan, Z. Evaluation of Antifatigue Effects of 20(S)-Ginsenoside Rg3 in Forced Swimming Mice. Indian J. Pharm. Sci. 2018, 80, 510–515. [Google Scholar] [CrossRef]
- Go, G.-Y.; Lee, S.-J.; Jo, A.; Lee, J.; Seo, N.-W.; Kang, J.-S.; Kim, S.-K.; Kim, S.-N.; Kim, Y.K.; Bae, G.-U. Ginsenoside Rg1 from Panax ginseng enhances myoblast differentiation and myotube growth. J. Ginseng Res. 2017, 41, 608–614. [Google Scholar] [CrossRef]
- Go, G.-Y.; Jo, A.; Seo, N.-W.; Kim, W.-Y.; Kim, Y.K.; So, E.-Y.; Chen, Q.; Kang, J.-S.; Bae, G.-U.; Lee, S.-J. Ginsenoside Rb1 and Rb2 upregulate Akt/mTOR signaling–mediated muscular hypertrophy and myoblast differentiation. J. Ginseng Res. 2019. [Google Scholar] [CrossRef]
- Jiang, R.; Wang, M.; Shi, L.; Zhou, J.; Ma, R.; Feng, K.; Chen, X.; Xu, X.; Li, X.; Li, T.; et al. Panax ginseng Total Protein Facilitates Recovery from Dexamethasone-Induced Muscle Atrophy through the Activation of Glucose Consumption in C2C12 Myotubes. BioMed Res. Int. 2019, 2019, 3719643. [Google Scholar] [CrossRef]
- Boonlert, W.; Benya-Aphikul, H.; Welbat, J.U.; Rodsiri, R. Ginseng Extract G115 Attenuates Ethanol-Induced Depression in Mice by Increasing Brain BDNF Levels. Nutrients 2017, 9, 931. [Google Scholar] [CrossRef] [PubMed]
- Seo, E.-Y.; Kim, W.-K. Red Ginseng Extract Reduced Metastasis of Colon Cancer Cells In Vitro and In Vivo. J. Ginseng Res. 2011, 35, 315–324. [Google Scholar] [CrossRef] [PubMed]
- Gonzales, E.L.T.; Jang, J.-H.; Mabunga, D.F.N.; Kim, J.-W.; Ko, M.J.; Cho, K.S.; Bahn, G.H.; Hong, M.; Ryu, J.H.; Kim, H.J.; et al. Supplementation of Korean Red Ginseng improves behavior deviations in animal models of autism. Food Nutr. Res. 2016, 60, 29245. [Google Scholar] [CrossRef] [PubMed]
- Huang, W.-C.; Hsu, Y.-J.; Wei, L.; Chen, Y.-J.; Huang, C.-C. Association of physical performance and biochemical profile of mice with intrinsic endurance swimming. Int. J. Med. Sci. 2016, 13, 892–901. [Google Scholar] [CrossRef]
- Flynn, J.M.; Meadows, E.; Fiorotto, M.; Klein, W.H. Myogenin Regulates Exercise Capacity and Skeletal Muscle Metabolism in the Adult Mouse. PLoS ONE 2010, 5, e13535. [Google Scholar] [CrossRef]
- Beaudart, C.; Rolland, Y.; Cruz-Jentoft, A.J.; Bauer, J.M.; Sieber, C.; Cooper, C.; Al-Daghri, N.; De Carvalho, I.A.; Bautmans, I.; Bernabei, R.; et al. Assessment of Muscle Function and Physical Performance in Daily Clinical Practice: A position paper endorsed by the European Society for Clinical and Economic Aspects of Osteoporosis, Osteoarthritis and Musculoskeletal Diseases (ESCEO). Calcif. Tissue Int. 2019, 105, 1–14. [Google Scholar] [CrossRef]
- Puigserver, P.; Rhee, J.; Lin, J.D.; Wu, Z.D.; Yoon, J.C.; Zhang, C.Y.; Krauss, S.; Mootha, V.K.; Lowell, B.B.; Spiegelman, B.M. Cytokine stimulation of energy expenditure through p38 MAP kinase activation of PPAR gamma coactivator-1. Mol. Cell 2001, 8, 971–982. [Google Scholar] [CrossRef]
- Jung, S.; Kim, K. Exercise-induced PGC-1alpha transcriptional factors in skeletal muscle. Integr. Med. Res. 2014, 3, 155–160. [Google Scholar] [CrossRef]
- Suwa, M.; Nakano, H.; Kumagai, S. Effects of chronic AICAR treatment on fiber composition, enzyme activity, UCP3, and PGC-1 in rat muscles. J. Appl. Physiol. 2003, 95, 960–968. [Google Scholar] [CrossRef]
- Jager, S.; Handschin, C.; Pierre, J.; Spiegelman, B.M. AMP-activated protein kinase (AMPK) action in skeletal muscle via direct phosphorylation of PGC-1 alpha. Proc. Natl. Acad. Sci. USA 2007, 104, 12017–12022. [Google Scholar] [CrossRef]
- Taherzadeh-Fard, E.; Saft, C.; Akkad, D.; Wieczorek, S.; Haghikia, A.; Chan, A.; Epplen, J.T.; Arning, L. PGC-1alpha downstream transcription factors NRF-1 and TFAM are genetic modifiers of Huntington disease. Mol. Neurodegener. 2011, 6, 32. [Google Scholar] [CrossRef]
- Wagatsuma, A.; Sakuma, K. Mitochondria as a Potential Regulator of Myogenesis. Sci. World J. 2013, 2013, 1–9. [Google Scholar] [CrossRef]
- Lin, Y.; Zhao, Y.; Li, R.; Gong, J.; Zheng, Y.; Wang, Y. PGC-1α is associated with C2C12 Myoblast differentiation. Open Life Sci. 2014, 9, 1030–1036. [Google Scholar] [CrossRef]
- Attele, A.S.; A Wu, J.; Yuan, C.S. Ginseng pharmacology: Multiple constituents and multiple actions. Biochem. Pharmacol. 1999, 58, 1685–1693. [Google Scholar] [CrossRef]
- Gui, Y.; Ryu, G.H. Effects of extrusion cooking on physicochemical properties of white and red ginseng (powder). J. Ginseng Res. 2014, 38, 146–153. [Google Scholar] [CrossRef]
- Zhang, H.-M.; Li, S.-L.; Zhang, H.; Wang, Y.; Zhao, Z.-L.; Chen, S.-L.; Xu, H.-X. Holistic quality evaluation of commercial white and red ginseng using a UPLC-QTOF-MS/MS-based metabolomics approach. J. Pharm. Biomed. Anal. 2012, 62, 258–273. [Google Scholar] [CrossRef]
- Ma, G.-D.; Chiu, C.-H.; Hsu, Y.-J.; Hou, C.-W.; Chen, Y.-M.; Huang, C.-C. Changbai Mountain Ginseng (Panax ginseng C.A. Mey) Extract Supplementation Improves Exercise Performance and Energy Utilization and Decreases Fatigue-Associated Parameters in Mice. Molecules 2017, 22, 237. [Google Scholar] [CrossRef]
- Li, L.; Pan, C.-S.; Yan, L.; Cui, Y.-C.; Liu, Y.-Y.; Mu, H.-N.; He, K.; Hu, B.-H.; Chang, X.; Sun, K.; et al. Ginsenoside Rg1 Ameliorates Rat Myocardial Ischemia-Reperfusion Injury by Modulating Energy Metabolism Pathways. Front. Physiol. 2018, 9, 78. [Google Scholar] [CrossRef]
- Xu, M.; Ma, Q.; Fan, C.; Chen, X.; Zhang, H.; Tang, M. Ginsenosides Rb1 and Rg1 protect primary cultured astrocytes against oxygen-glucose deprivation/reoxygenation-induced injury via improving mtochondrial fnction. Int. J. Mol. Sci. 2019, 20, 6086. [Google Scholar] [CrossRef]
- Tan, S.-J.; Li, N.; Zhou, F.; Dong, Q.-T.; Zhang, X.-D.; Chen, B.-C.; Yu, Z. Ginsenoside Rb1 improves energy metabolism in the skeletal muscle of an animal model of postoperative fatigue syndrome. J. Surg. Res. 2014, 191, 344–349. [Google Scholar] [CrossRef]
- Yang, Q.-Y.; Lai, X.-D.; Ouyang, J.; Yang, J.-D. Effects of Ginsenoside Rg3 on fatigue resistance and SIRT1 in aged rats. Toxicology 2018, 409, 144–151. [Google Scholar] [CrossRef] [PubMed]
Sample Availability: Samples of the compounds are not available. |

© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shin, E.J.; Jo, S.; Choi, S.; Cho, C.-W.; Lim, W.-C.; Hong, H.-D.; Lim, T.-G.; Jang, Y.J.; Jang, M.; Byun, S.; et al. Red Ginseng Improves Exercise Endurance by Promoting Mitochondrial Biogenesis and Myoblast Differentiation. Molecules 2020, 25, 865. https://doi.org/10.3390/molecules25040865
Shin EJ, Jo S, Choi S, Cho C-W, Lim W-C, Hong H-D, Lim T-G, Jang YJ, Jang M, Byun S, et al. Red Ginseng Improves Exercise Endurance by Promoting Mitochondrial Biogenesis and Myoblast Differentiation. Molecules. 2020; 25(4):865. https://doi.org/10.3390/molecules25040865
Chicago/Turabian StyleShin, Eun Ju, Seongin Jo, Sungbin Choi, Chang-Won Cho, Won-Chul Lim, Hee-Do Hong, Tae-Gyu Lim, Young Jin Jang, Mi Jang, Sanguine Byun, and et al. 2020. "Red Ginseng Improves Exercise Endurance by Promoting Mitochondrial Biogenesis and Myoblast Differentiation" Molecules 25, no. 4: 865. https://doi.org/10.3390/molecules25040865
APA StyleShin, E. J., Jo, S., Choi, S., Cho, C.-W., Lim, W.-C., Hong, H.-D., Lim, T.-G., Jang, Y. J., Jang, M., Byun, S., & Rhee, Y. (2020). Red Ginseng Improves Exercise Endurance by Promoting Mitochondrial Biogenesis and Myoblast Differentiation. Molecules, 25(4), 865. https://doi.org/10.3390/molecules25040865






